Disclosing the Interaction of Gold Nanoparticles with Aβ(1–40) Monomers through Replica Exchange Molecular Dynamics Simulations
Abstract
1. Introduction
2. Results
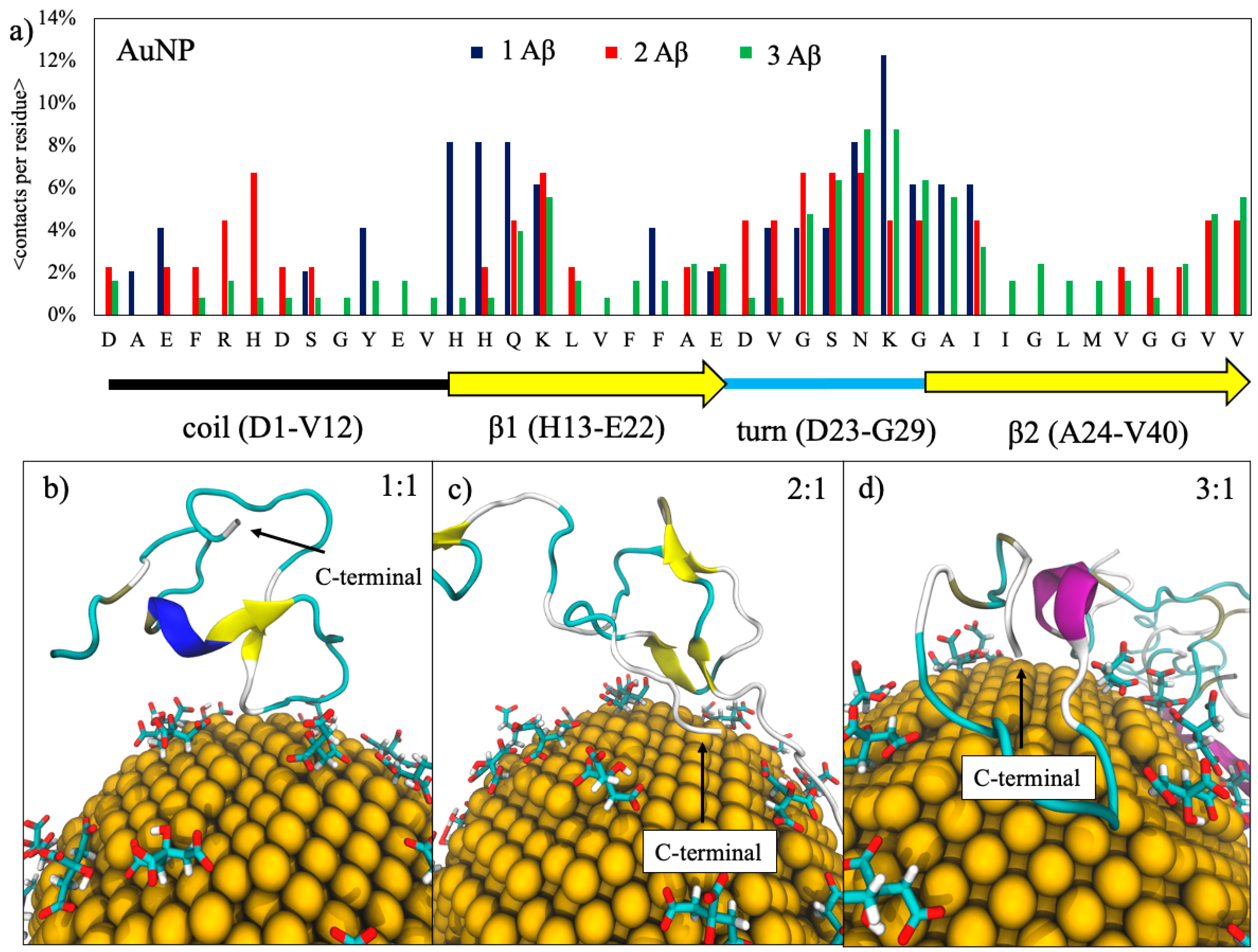
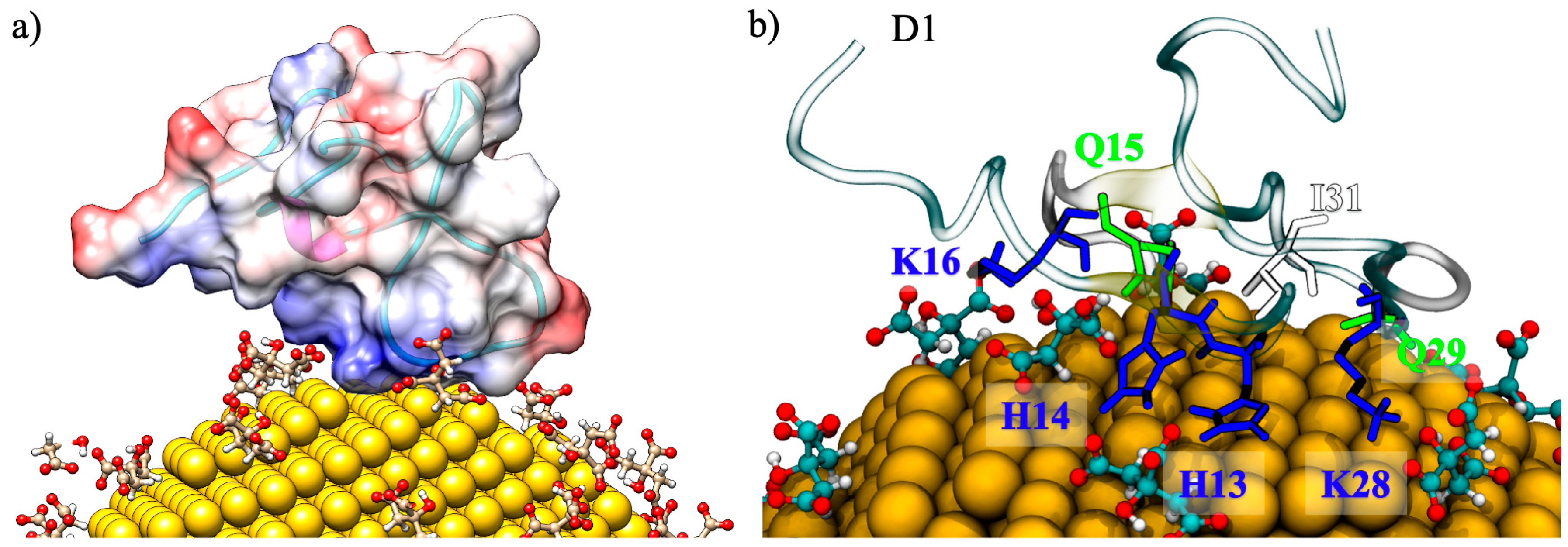
2.1. Binding Contacts
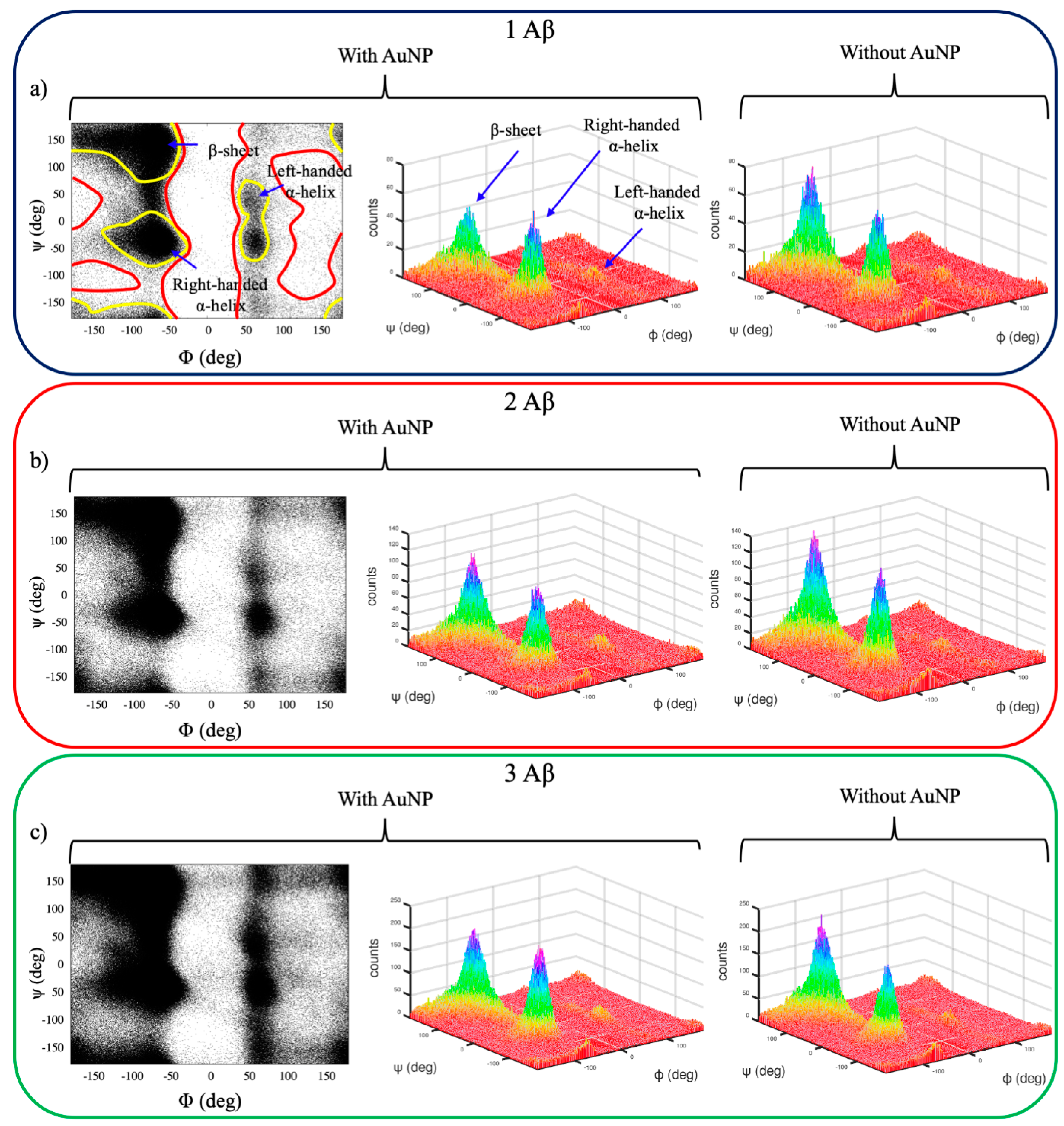
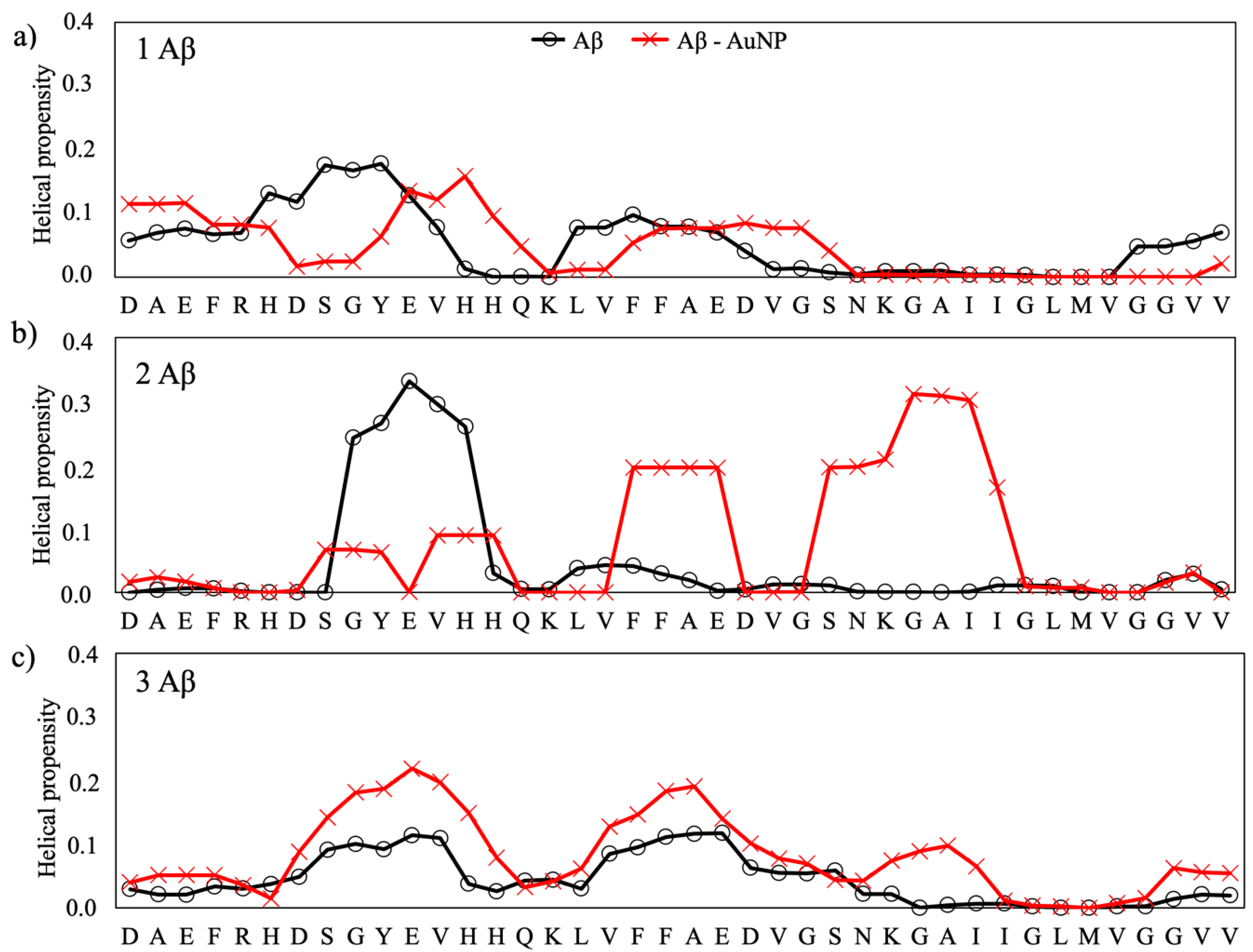
2.2. Structural Analysis
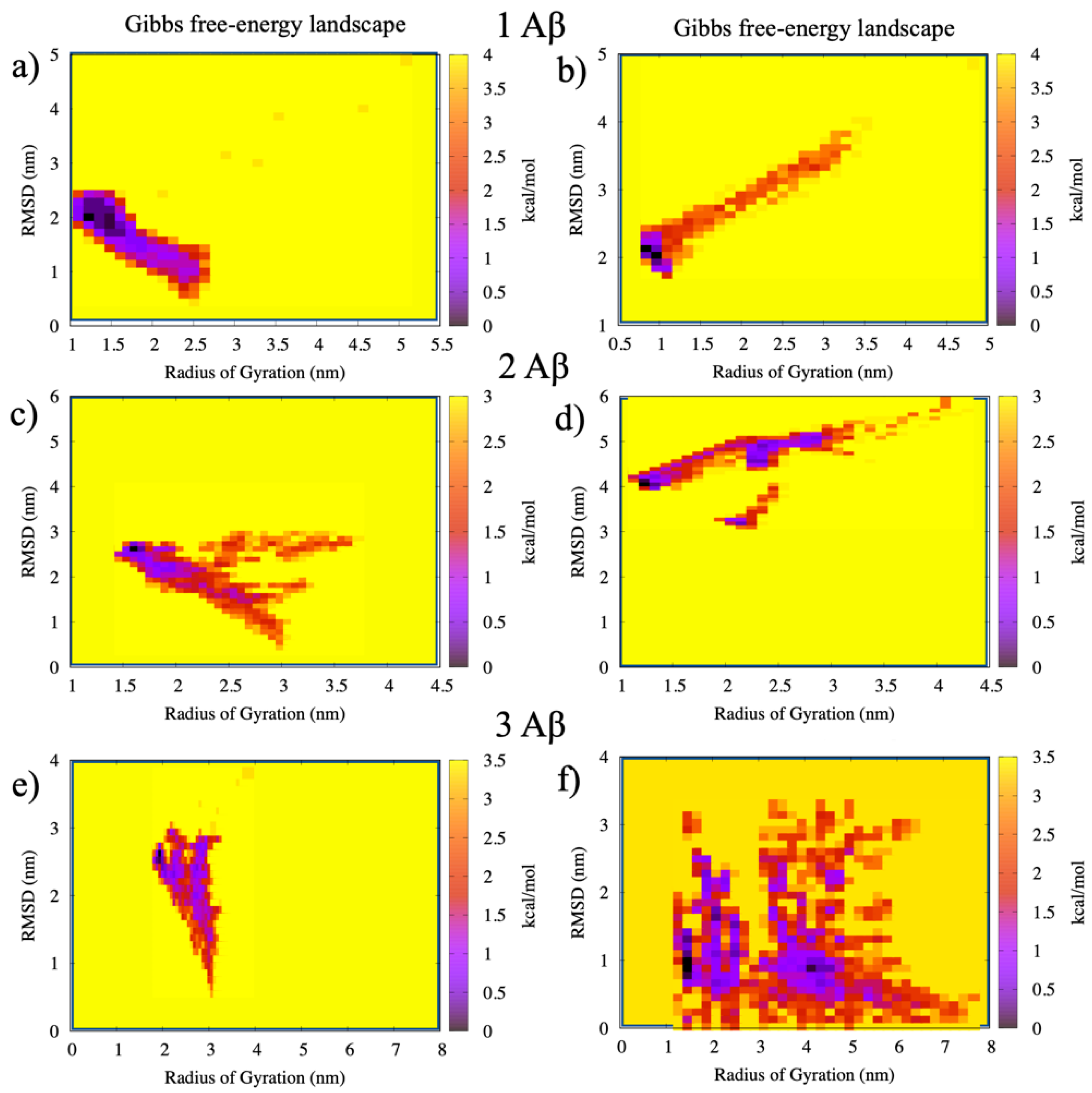
2.3. Free-Energy Calculations
3. Discussion
4. Materials and Methods
4.1. REST Simulations
4.2. Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Tycko, R. Amyloid Polymorphism: Structural Basis and Neurobiological Relevance. Neuron 2015, 86, 632–645. [Google Scholar] [CrossRef] [PubMed]
- Cummings, J.L. Alzheimer’s Disease. N. Engl. J. Med. 2004, 351, 56–67. [Google Scholar] [CrossRef] [PubMed]
- Hardy, J.; Selkoe, D.J. The amyloid hypothesis of Alzheimer’s disease: Progress and problems on the road to therapeutics. Science 2002, 297, 353–356. [Google Scholar] [CrossRef] [PubMed]
- Paravastu, A.K.; Leapman, R.D.; Yau, W.-M.; Tycko, R. Molecular structural basis for polymorphism in Alzheimer’s -amyloid fibrils. Proc. Natl. Acad. Sci. USA 2008, 105, 18349–18354. [Google Scholar] [CrossRef] [PubMed]
- Tycko, R. Molecular structure of amyloid fibrils: Insights from solid-state NMR. Q. Rev. Biophys. 2006, 39, 1–55. [Google Scholar] [CrossRef] [PubMed]
- Rosenman, D.J.; Connors, C.R.; Chen, W.; Wang, C.; García, A.E. Aβ Monomers Transiently Sample Oligomer and Fibril-Like Configurations: Ensemble Characterization Using a Combined MD/NMR Approach. J. Mol. Biol. 2013, 425, 3338–3359. [Google Scholar] [CrossRef]
- Rambaran, R.N.; Serpell, L.C. Amyloid fibrils. Prion 2008, 2, 112–117. [Google Scholar] [CrossRef]
- Cummings, J.; Lee, G.; Ritter, A.; Sabbagh, M.; Zhong, K. Alzheimer’s disease drug development pipeline: 2020. Transl. Res. Clin. Interv. 2020, 6, e12050. [Google Scholar] [CrossRef]
- Hamaguchi, T.; Ono, K.; Yamada, M. Curcumin and Alzheimer’s disease. CNS Neurosci. Ther. 2010, 16, 285–297. [Google Scholar] [CrossRef]
- Scarmeas, N. Physical Activity, Diet, and Risk of Alzheimer Disease. JAMA 2009, 302, 627–637. [Google Scholar] [CrossRef]
- Scarmeas, N.; Stern, Y.; Mayeux, R.; Luchsinger, J.A. Mediterranean Diet, Alzheimer Disease, and Vascular Mediation. Arch. Neurol. 2006, 63, 1709–1717. [Google Scholar] [CrossRef] [PubMed]
- Zorkina, Y.; Abramova, O.; Ushakova, V.; Morozova, A.; Zubkov, E.; Valikhov, M.; Melnikov, P.; Majouga, A.; Chekhonin, V. Nano Carrier Drug Delivery Systems for the Treatment of Neuropsychiatric Disorders: Advantages and Limitations. Molecules 2020, 25, 5294. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Ding, F.; Cai, R.; Javed, I.; Yang, W.; Zhang, Z.; Li, Y.; Davis, T.P.; Ke, P.C.; Chen, C. Amyloidosis inhibition, a new frontier of the protein corona. Nano Today 2020, 35, 100937. [Google Scholar] [CrossRef] [PubMed]
- Brancolini, G.; Maschio, M.C.; Cantarutti, C.; Corazza, A.; Fogolari, F.; Bellotti, V.; Corni, S.; Esposito, G. Citrate stabilized gold nanoparticles interfere with amyloid fibril formation: D76N and ΔN6 β2-microglobulin variants. Nanoscale 2018, 10, 4793–4806. [Google Scholar] [CrossRef] [PubMed]
- Cendrowska, U.; Silva, P.J.; Ait-Bouziad, N.; Müller, M.; Guven, Z.P.; Vieweg, S.; Chiki, A.; Radamaker, L.; Kumar, S.T.; Fändrich, M.; et al. Unraveling the complexity of amyloid polymorphism using gold nanoparticles and cryo-EM. Proc. Natl. Acad. Sci. USA 2020, 117, 6866–6874. [Google Scholar] [CrossRef]
- Tavanti, F.; Pedone, A.; Menziani, M.C.; Alexander-Katz, A. Computational Insights into the Binding of Monolayer-Capped Gold Nanoparticles onto Amyloid-β Fibrils. ACS Chem. Neurosci. 2020, 11, 3153–3160. [Google Scholar] [CrossRef]
- Corni, S.; Bussi, G.; Di Felice, R.; Corni, S. Fibrillation-prone conformations of the amyloid-β-42 peptide at the gold/water interface. Nanoscale 2017, 9, 2279–2290. [Google Scholar] [CrossRef]
- Tavanti, F.; Pedone, A.; Menziani, M.C. Computational Insight into the Effect of Natural Compounds on the Destabilization of Preformed Amyloid-β (1–40) Fibrils. Molecules 2018, 23, 1320. [Google Scholar] [CrossRef]
- Tavanti, F.; Pedone, A.; Menziani, M.C. Insights into the Effect of Curcumin and (–)-Epigallocatechin-3-Gallate on the Aggregation of Aβ(1–40) Monomers by Means of Molecular Dynamics. Int. J. Mol. Sci. 2020, 21, 5462. [Google Scholar] [CrossRef]
- Laio, A.; Parrinello, M. Escaping free-energy minima. Proc. Natl. Acad. Sci. USA 2002, 99, 12562–12566. [Google Scholar] [CrossRef]
- Hansmann, U.H.E. Parallel tempering algorithm for conformational studies of biological molecules. Chem. Phys. Lett. 1997, 281, 140–150. [Google Scholar] [CrossRef]
- Do, T.N.; Choy, W.-Y.; Karttunen, M.M. Accelerating the Conformational Sampling of Intrinsically Disordered Proteins. J. Chem. Theory Comput. 2014, 10, 5081–5094. [Google Scholar] [CrossRef] [PubMed]
- Granata, D.; Camilloni, C.; Vendruscolo, M.; Laio, A. Characterization of the free-energy landscapes of proteins by NMR-guided metadynamics. Proc. Natl. Acad. Sci. USA 2013, 110, 6817–6822. [Google Scholar] [CrossRef] [PubMed]
- Tavanti, F.; Pedone, A.; Menziani, M.C. A closer look into the ubiquitin corona on gold nanoparticles by computational studies. New J. Chem. 2014, 39, 2474–2482. [Google Scholar] [CrossRef]
- Tan, S.F.; Raj, S.; Bisht, G.; Annadata, H.V.; Chu, H.-S.; Král, P.; Mirsaidov, U. Nanoparticle Interactions Guided by Shape-Dependent Hydrophobic Forces. Adv. Mater. 2018, 30, e1707077. [Google Scholar] [CrossRef]
- Lin, W.; Insley, T.; Tuttle, M.D.; Zhu, L.; Berthold, D.A.; Král, P.; Rienstra, M.; Murphy, C.J. Control of Protein Orientation on Gold Nanoparticles. J. Phys. Chem. C 2015, 119, 21035–21043. [Google Scholar] [CrossRef]
- Sen, S.; Vuković, L.; Král, P. Computational screening of nanoparticles coupling to Aβ40 peptides and fibrils. Sci. Rep. 2019, 9, 1–8. [Google Scholar] [CrossRef]
- Brancolini, G.; Lopez, H.; Corni, S.; Tozzini, V. Low-Resolution Models for the Interaction Dynamics of Coated Gold Nanoparticles with β2-microglobulin. Int. J. Mol. Sci. 2019, 20, 3866. [Google Scholar] [CrossRef]
- Ramachandran, G.; Ramakrishnan, C.; Sasisekharan, V. Stereochemistry of polypeptide chain configurations. J. Mol. Biol. 2009, 7, 95–99. [Google Scholar] [CrossRef]
- Cheon, M.; Hall, C.K.; Chang, I. Structural Conversion of Aβ17–42 Peptides from Disordered Oligomers to U-Shape Protofilaments via Multiple Kinetic Pathways. PLoS Comput. Biol. 2015, 11, e1004258. [Google Scholar] [CrossRef]
- Festa, G.; Mallamace, F.; Sancesario, G.M.; Corsaro, C.; Mallamace, D.; Fazio, E.; Arcidiacono, L.; Sakai, V.G.; Senesi, R.; Preziosi, E.; et al. Aggregation States of Aβ1–40, Aβ1–42 and Aβp3–42 Amyloid Beta Peptides: A SANS Study. Int. J. Mol. Sci. 2019, 20, 4126. [Google Scholar] [CrossRef] [PubMed]
- Carballo-Pacheco, M.; Strodel, B. Advances in the Simulation of Protein Aggregation at the Atomistic Scale. J. Phys. Chem. B 2016, 120, 2991–2999. [Google Scholar] [CrossRef] [PubMed]
- Teleanu, D.M.; Chircov, C.; Grumezescu, A.M.; Volceanov, A.; Teleanu, R.I. Blood-Brain Delivery Methods Using Nanotechnology. Pharmaceutics 2018, 10, 269. [Google Scholar] [CrossRef] [PubMed]
- Schmid, N.; Eichenberger, A.P.; Choutko, A.; Riniker, S.; Winger, M.; Mark, A.E.; Van Gunsteren, W.F. Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur. Biophys. J. 2011, 40, 843–856. [Google Scholar] [CrossRef] [PubMed]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Abraham, M.; Van Der Spoel, D.; Lindahl, E.; Hess, B.; Spoel, D.; van der Lindahl, E. GROMACS User Manual Version 5.0.4. 2014. Available online: www.gromacs.org (accessed on 21 December 2020). [CrossRef]
- Malde, A.K.; Zuo, L.; Breeze, M.; Stroet, M.; Poger, D.; Nair, P.C.; Oostenbrink, C.; Mark, A.E. An Automated Force Field Topology Builder (ATB) and Repository: Version 1.0. J. Chem. Theory Comput. 2011, 7, 4026–4037. [Google Scholar] [CrossRef]
- Koziara, K.B.; Stroet, M.; Malde, A.K.; Mark, A.E. Testing and validation of the Automated Topology Builder (ATB) version 2.0: Prediction of hydration free enthalpies. J. Comput. Mol. Des. 2014, 28, 221–233. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Richard, T.; Papastamoulis, Y.; Waffo-Teguo, P.; Monti, J.-P. 3D NMR structure of a complex between the amyloid beta peptide (1–40) and the polyphenol ε-viniferin glucoside: Implications in Alzheimer’s disease. Biochim. et Biophys. Acta (BBA)-Gen. Subj. 2013, 1830, 5068–5074. [Google Scholar] [CrossRef]
- Kyrychenko, A.; Korsun, O.M.; Gubin, I.I.; Kovalenko, S.M.; Kalugin, O.N. Atomistic Simulations of Coating of Silver Nanoparticles with Poly(vinylpyrrolidone) Oligomers: Effect of Oligomer Chain Length. J. Phys. Chem. C 2015, 119, 7888–7899. [Google Scholar] [CrossRef]
- Brancolini, G.; Corazza, A.; Vuano, M.; Fogolari, F.; Mimmi, M.C.; Bellotti, V.; Stoppini, M.; Corni, S.; Esposito, G. Probing the Influence of Citrate-Capped Gold Nanoparticles on an Amyloidogenic Protein. ACS Nano 2015, 9, 2600–2613. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, H.J.C.; Postma, J.P.M.; Van Gunsteren, W.F.; Hermans, J. Interaction Models for Water in Relation to Protein Hydration. In Excited States in Organic Chemistry and Biochemistry; Springer Science and Business Media LLC: Heidelerg/Germany, 1981; pp. 331–342. [Google Scholar]
- Darden, A.T. York, D.M.; Pedersen, L. Particle mesh Ewald: AnN⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Smith, A.K.; Lockhart, C.; Klimov, D.K. Does Replica Exchange with Solute Tempering Efficiently Sample Aβ Peptide Conformational Ensembles? J. Chem. Theory Comput. 2016, 12, 5201–5214. [Google Scholar] [CrossRef]
- Joosten, R.P.; Beek, T.A.H.T.; Krieger, E.; Hekkelman, M.L.; Hooft, R.W.W.; Schneider, R.; Sander, C.; Vriend, G. A series of PDB related databases for everyday needs. Nucleic Acids Res. 2010, 39, D411–D419. [Google Scholar] [CrossRef]
- Dolinsky, T.J.; Czodrowski, P.; Li, H.; Nielsen, J.E.; Jensen, J.H.; Klebe, G.; Baker, N.A. PDB2PQR: Expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 2007, 35, W522–W525. [Google Scholar] [CrossRef]
- Baker, N.A.; Sept, D.; Joseph, S.; Holst, M.J.; McCammon, J.A. Electrostatics of nanosystems: Application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 2001, 98, 10037–10041. [Google Scholar] [CrossRef]

| System | Ratio 1:1 | Ratio 2:1 | Ratio 3:1 |
|---|---|---|---|
| Monomers only [19] | 1 monomer | 2 monomers | 3 monomers |
| AuNP | 1 monomer + 1 AuNP | 2 monomers + 1 AuNP | 3 monomers + 1 AuNP |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tavanti, F.; Pedone, A.; Menziani, M.C. Disclosing the Interaction of Gold Nanoparticles with Aβ(1–40) Monomers through Replica Exchange Molecular Dynamics Simulations. Int. J. Mol. Sci. 2021, 22, 26. https://doi.org/10.3390/ijms22010026
Tavanti F, Pedone A, Menziani MC. Disclosing the Interaction of Gold Nanoparticles with Aβ(1–40) Monomers through Replica Exchange Molecular Dynamics Simulations. International Journal of Molecular Sciences. 2021; 22(1):26. https://doi.org/10.3390/ijms22010026
Chicago/Turabian StyleTavanti, Francesco, Alfonso Pedone, and Maria Cristina Menziani. 2021. "Disclosing the Interaction of Gold Nanoparticles with Aβ(1–40) Monomers through Replica Exchange Molecular Dynamics Simulations" International Journal of Molecular Sciences 22, no. 1: 26. https://doi.org/10.3390/ijms22010026
APA StyleTavanti, F., Pedone, A., & Menziani, M. C. (2021). Disclosing the Interaction of Gold Nanoparticles with Aβ(1–40) Monomers through Replica Exchange Molecular Dynamics Simulations. International Journal of Molecular Sciences, 22(1), 26. https://doi.org/10.3390/ijms22010026





