Hierarchical Action of Mulberry miR156 in the Vegetative Phase Transition
Abstract
1. Introduction
2. Results
2.1. Mulberry miR156 Is Differentially Expressed in Juvenile and Mature Phase
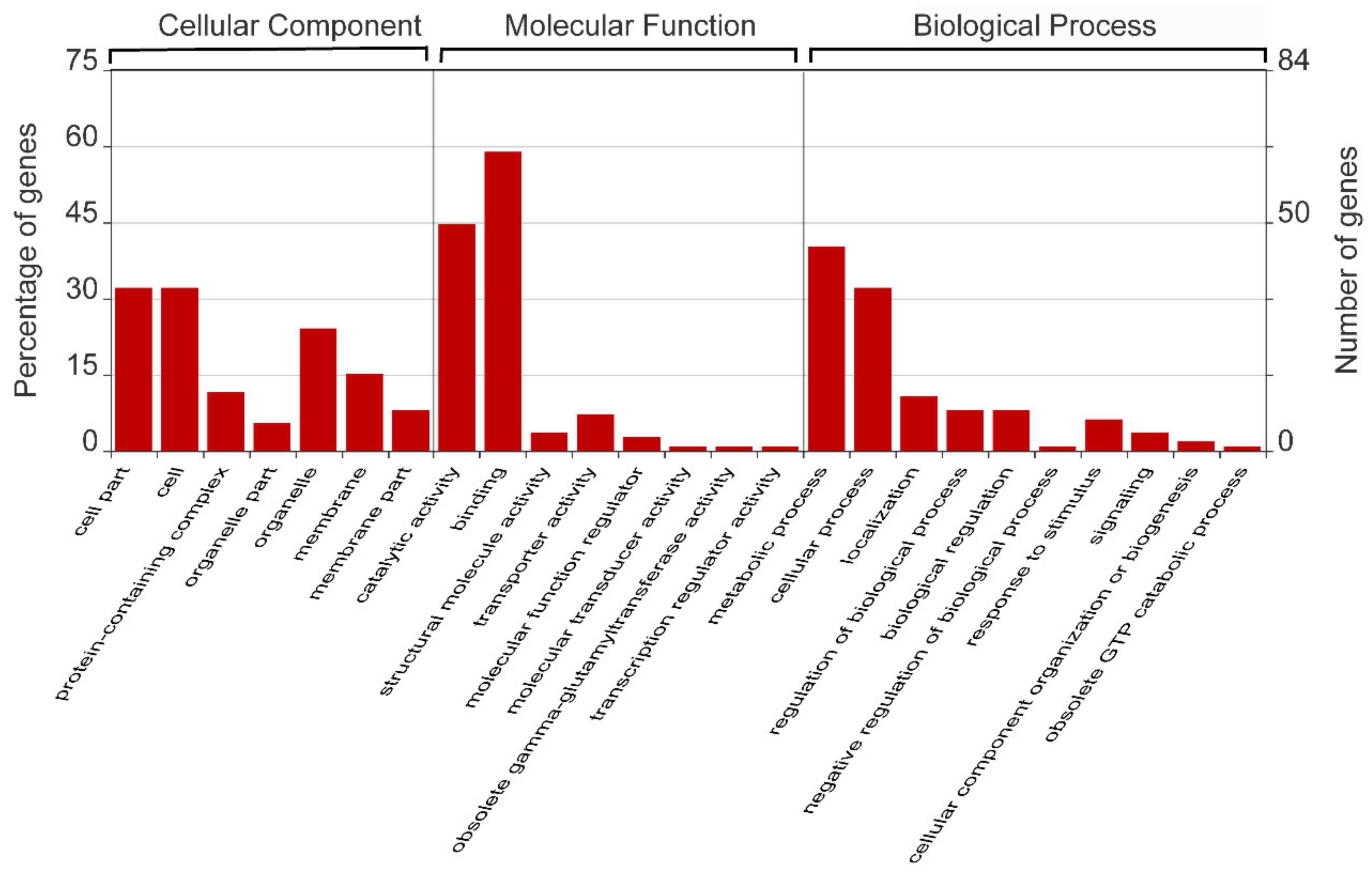
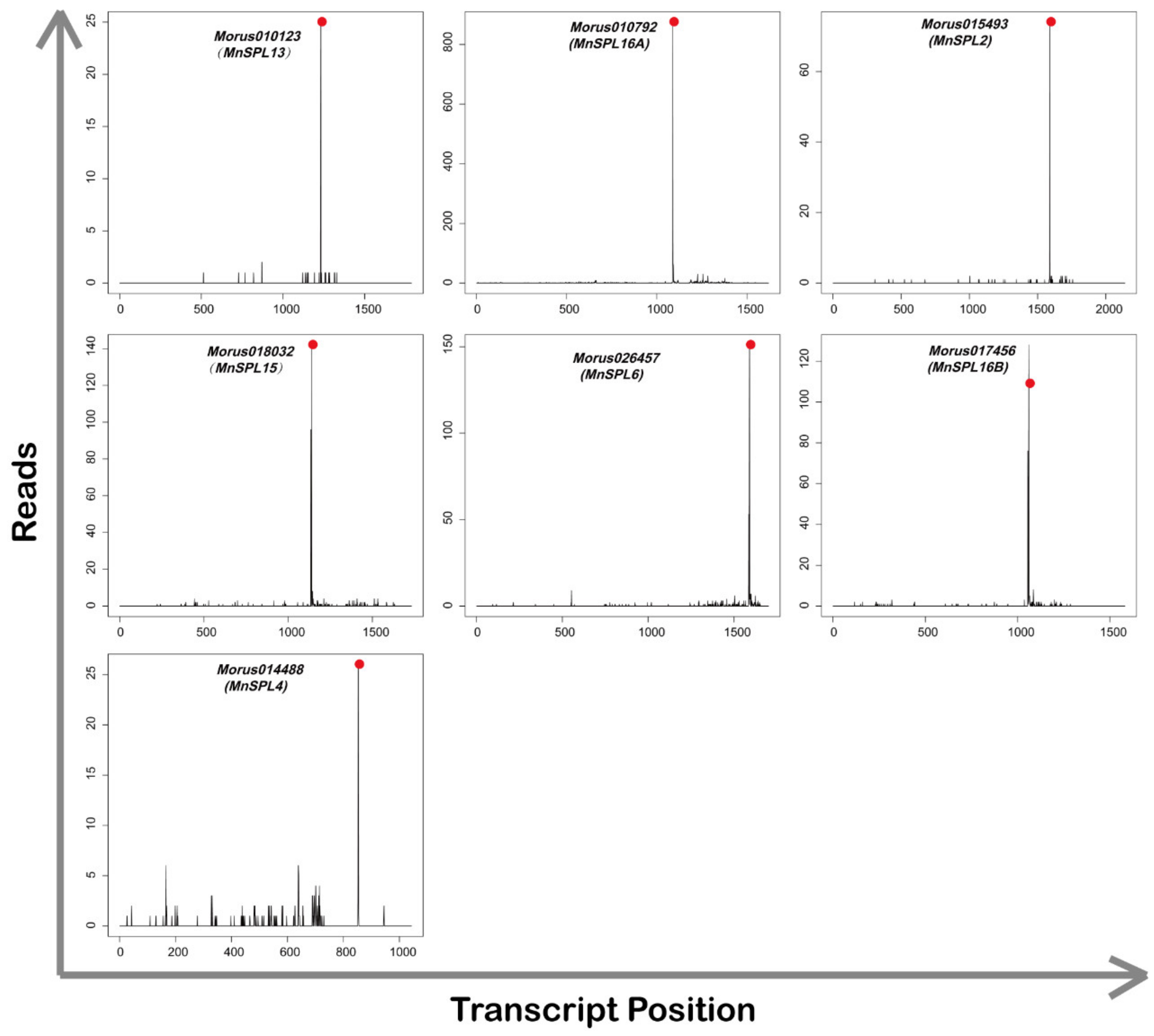
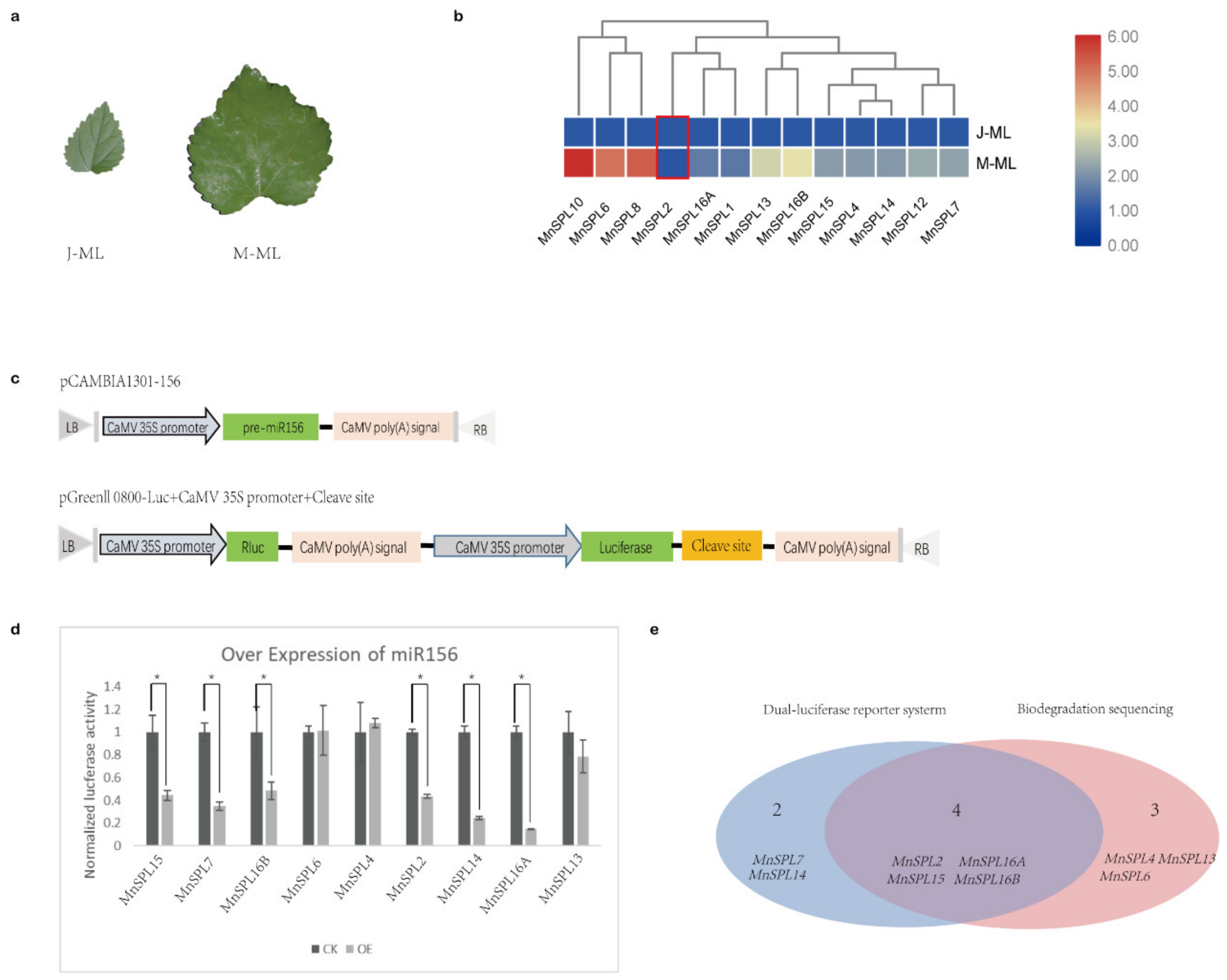
2.2. Identification of the miR156 Target Genes in Mulberry
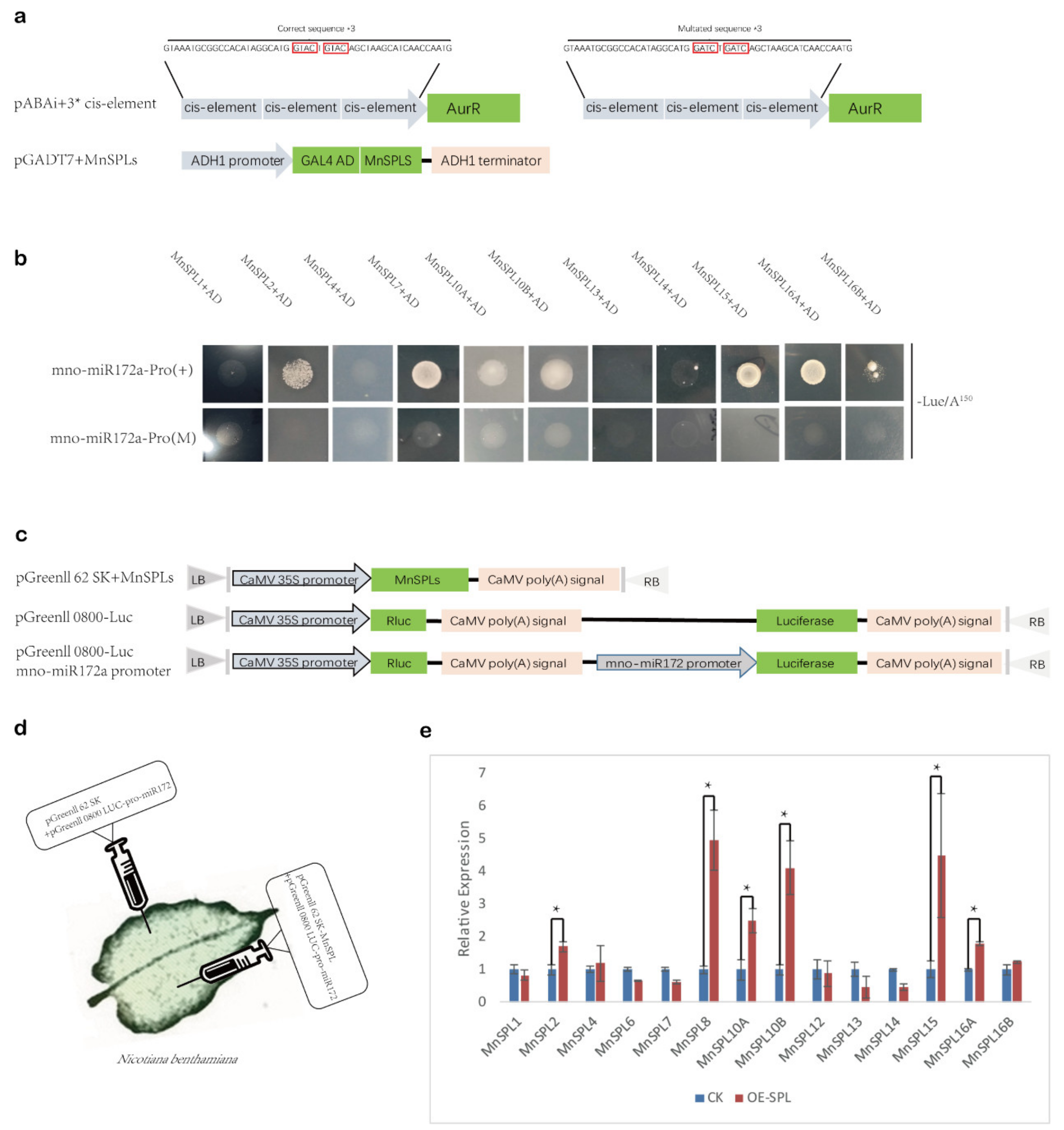
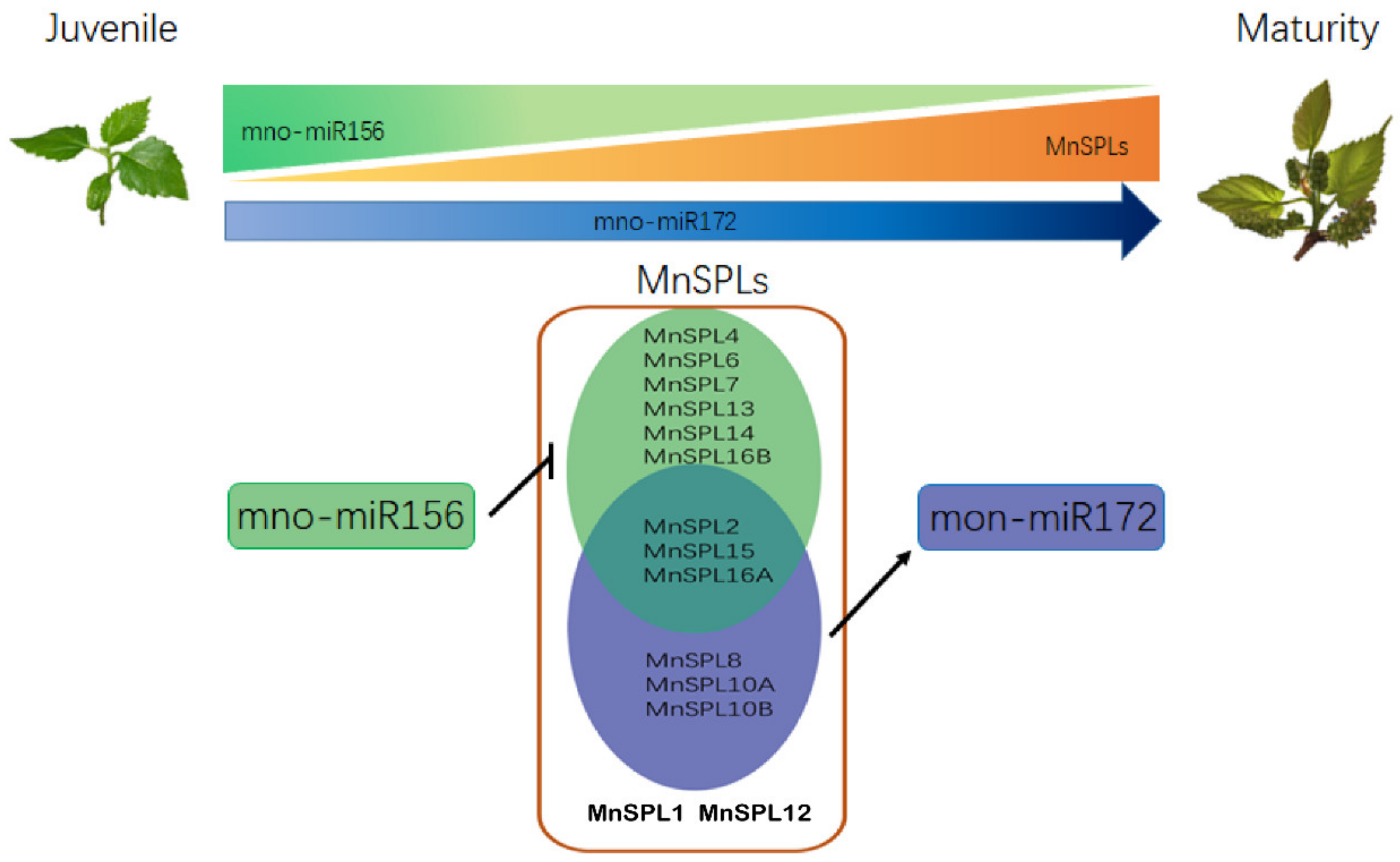
2.3. MnSPLs Bound to the Promoter of mno-miR172a and Promoted Its Expression in Mulberry
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Bioinformatics Analysis of miR156, SPL Genes, and miR172 in Mulberry
4.3. Epression Analysis of miRNAs and Genes
4.4. Transient Trnsgene Expression in Mulberry
4.5. Dual-Luciferase Repoter Assay
4.6. Yeast One-Hybrid Assay
4.7. Degradome Sequencing
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Baurle, I.; Dean, C. The timing of developmental transitions in plants. Cell 2006, 125, 655–664. [Google Scholar] [CrossRef] [PubMed]
- Poethig, R.S. Vegetative phase change and shoot maturation in plants. Curr. Top. Dev. Biol. 2013, 105, 125–152. [Google Scholar] [CrossRef]
- Wu, G.; Park, M.Y.; Conway, S.R.; Wang, J.W.; Weigel, D.; Poethig, R.S. The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 2009, 138, 750–759. [Google Scholar] [CrossRef]
- Yang, L.; Conway, S.R.; Poethig, R.S. Vegetative phase change is mediated by a leaf-derived signal that represses the transcription of miR156. Development 2011, 138, 245–249. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Wang, J.; Zhou, C. The role of miR156 in developmental transitions in Nicotiana tabacum. Sci. China Life Sci. 2015, 58, 253–260. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Xie, K.; Wu, C.; Xiong, L. Genomic organization, differential expression, and interaction of SQUAMOSA promoter-binding-like transcription factors and microRNA156 in rice. Plant Physiol. 2006, 142, 280–293. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Sun, S.; Jin, J.; Fu, D.; Yang, X.; Weng, X.; Xu, C.; Li, X.; Xiao, J.; Zhang, Q. Coordinated regulation of vegetative and reproductive branching in rice. Proc. Natl. Acad. Sci. USA 2015, 112, 15504–15509. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Cao, L.; Zhou, C.M.; Zhang, T.Q.; Lian, H.; Sun, Y.; Wu, J.; Huang, J.; Wang, G.; Wang, J.W. Sugar is an endogenous cue for juvenile-to-adult phase transition in plants. eLife 2013, 2, e00269. [Google Scholar] [CrossRef]
- Yang, L.; Xu, M.; Koo, Y.; He, J.; Poethig, R.S. Sugar promotes vegetative phase change in Arabidopsis thaliana by repressing the expression of MIR156A and MIR156C. eLife 2013, 2, e00260. [Google Scholar] [CrossRef]
- Lawson, E.J.; Poethig, R.S. Shoot development in plants: Time for a change. Trends Genet. 1995, 11, 263–268. [Google Scholar] [CrossRef]
- Wang, J.W.; Park, M.Y.; Wang, L.J.; Koo, Y.; Chen, X.Y.; Weigel, D.; Poethig, R.S. miRNA control of vegetative phase change in trees. PLoS Genet. 2011, 7, e1002012. [Google Scholar] [CrossRef]
- Ahsan, M.U.; Hayward, A.; Irihimovitch, V.; Fletcher, S.; Tanurdzic, M.; Pocock, A.; Beveridge, C.A.; Mitter, N. Juvenility and Vegetative Phase Transition in Tropical/Subtropical Tree Crops. Front. Plant Sci. 2019, 10, 729. [Google Scholar] [CrossRef]
- Shalom, L.; Shlizerman, L.; Zur, N.; Doron-Faigenboim, A.; Blumwald, E.; Sadka, A. Molecular characterization of Squamosa Promoter Binding Protein-Like (SPL) gene family from Citrus and the effect of fruit load on their expression. Front. Plant Sci. 2015, 6, 389. [Google Scholar] [CrossRef]
- Jiang, Y.; Peng, J.; Wang, M.; Su, W.; Gan, X.; Jing, Y.; Yang, X. The Role of EjSPL3, EjSPL4, EjSPL5, and EjSPL9 in Regulating Flowering in Loquat (Eriobotrya japonica Lindl.). Int. J. Mol. Sci. 2019, 21, 248. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.P.; Ou, T.T.; Wang, C.J. Mulberry (sang shèn zǐ) and its bioactive compounds, the chemoprevention effects and molecular mechanisms in vitro and in vivo. J. Tradit. Complement. Med. 2013, 3, 7–15. [Google Scholar] [CrossRef] [PubMed]
- He, N.; Zhang, C.; Qi, X.; Zhao, S.; Tao, Y.; Yang, G.; Lee, T.H.; Wang, X.; Cai, Q.; Li, D.; et al. Draft genome sequence of the mulberry tree Morus notabilis. Nat. Commun. 2013, 4, 2445. [Google Scholar] [CrossRef]
- Yuan, Q.; Zhao, L. The Mulberry (Morus alba L.) Fruit-A Review of Characteristic Components and Health Benefits. J. Agric. Food Chem. 2017, 65, 10383–10394. [Google Scholar] [CrossRef]
- Chen, H.; Pu, J.; Liu, D.; Yu, W.; Shao, Y.; Yang, G.; Xiang, Z.; He, N. Anti-Inflammatory and Antinociceptive Properties of Flavonoids from the Fruits of Black Mulberry (Morus nigra L.). PLoS ONE 2016, 11, e0153080. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Yu, W.; Chen, G.; Meng, S.; Xiang, Z.; He, N. Antinociceptive and Antibacterial Properties of Anthocyanins and Flavonols from Fruits of Black and Non-Black Mulberries. Molecules 2017, 23, 4. [Google Scholar] [CrossRef]
- Liu, C.; Xu, Y.; Feng, Y.; Long, D.; Cao, B.; Xiang, Z.; Zhao, A. Ectopic Expression of Mulberry G-Proteins Alters Drought and Salt Stress Tolerance in Tobacco. Int. J. Mol. Sci. 2018, 20, 89. [Google Scholar] [CrossRef]
- Yamagishi, N.; Kishigami, R.; Yoshikawa, N. Reduced generation time of apple seedlings to within a year by means of a plant virus vector: A new plant-breeding technique with no transmission of genetic modification to the next generation. Plant Biotechnol. J. 2014, 12, 60–68. [Google Scholar] [CrossRef]
- Jiao, F.; Luo, R.; Dai, X.; Liu, H.; Yu, G.; Han, S.; Lu, X.; Su, C.; Chen, Q.; Song, Q.; et al. Chromosome-Level Reference Genome and Population Genomic Analysis Provide Insights into the Evolution and Improvement of Domesticated Mulberry (Morus alba). Mol. Plant 2020, 13, 1001–1012. [Google Scholar] [CrossRef] [PubMed]
- Jia, L.; Zhang, D.; Qi, X.; Ma, B.; Xiang, Z.; He, N. Identification of the conserved and novel miRNAs in Mulberry by high-throughput sequencing. PloS ONE 2014, 9, e104409. [Google Scholar] [CrossRef]
- Wu, P.; Han, S.; Zhao, W.; Chen, T.; Zhou, J.; Li, L. Genome-wide identification of abiotic stress-regulated and novel microRNAs in mulberry leaf. Plant Physiol Biochem 2015, 95, 75–82. [Google Scholar] [CrossRef]
- Poethig, R.S. The past, present, and future of vegetative phase change. Plant Physiol. 2010, 154, 541–544. [Google Scholar] [CrossRef] [PubMed]
- Axtell, M.J.; Bowman, J.L. Evolution of plant microRNAs and their targets. Trends Plant Sci. 2008, 13, 343–349. [Google Scholar] [CrossRef]
- Zheng, C.; Ye, M.; Sang, M.; Wu, R. A Regulatory Network for miR156-SPL Module in Arabidopsis thaliana. Int. J. Mol. Sci. 2019, 20, 166. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Hu, T.; Zhao, J. Developmental Functions of miR156-Regulated Squamosa Promoter Binding Protein-Like (SPL) Genes in Arabidopsis thaliana. PLoS Genet. 2016, 12, e1006263. [Google Scholar] [CrossRef] [PubMed]
- Barrera-Rojas, C.H.; Rocha, G.H.B.; Polverari, L.; Pinheiro Brito, D.A.; Batista, D.S.; Notini, M.M.; da Cruz, A.C.F.; Morea, E.G.O.; Sabatini, S.; Otoni, W.C.; et al. miR156-targeted SPL10 controls Arabidopsis root meristem activity and root-derived de novo shoot regeneration via cytokinin responses. J. Exp. Bot. 2020, 71, 934–950. [Google Scholar] [CrossRef] [PubMed]
- Nanda, S.; Hussain, S. Genome-wide identification of the SPL gene family in Dichanthelium oligosanthes. Bioinformation 2019, 15, 165–171. [Google Scholar] [CrossRef]
- Zhu, T.; Liu, Y.; Ma, L.; Wang, X.; Zhang, D.; Han, Y.; Ding, Q.; Ma, L. Genome-wide identification, phylogeny and expression analysis of the SPL gene family in wheat. BMC Plant Biol. 2020, 20, 420. [Google Scholar] [CrossRef]
- Shao, F.; Lu, Q.; Wilson, I.W.; Qiu, D. Genome-wide identification and characterization of the SPL gene family in Ziziphus jujuba. Gene 2017, 627, 315–321. [Google Scholar] [CrossRef]
- Yu, N.; Niu, Q.W.; Ng, K.H.; Chua, N.H. The role of miR156/SPLs modules in Arabidopsis lateral root development. Plant J. 2015, 83, 673–685. [Google Scholar] [CrossRef]
- Wang, J.W. Regulation of flowering time by the miR156-mediated age pathway. J. Exp. Bot. 2014, 65, 4723–4730. [Google Scholar] [CrossRef] [PubMed]
- Chen, X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 2004, 303, 2022–2025. [Google Scholar] [CrossRef] [PubMed]
- Aukerman, M.J.; Sakai, H. Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell 2003, 15, 2730–2741. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.H.; Seo, P.J.; Kang, S.K.; Park, C.M. miR172 signals are incorporated into the miR156 signaling pathway at the SPL3/4/5 genes in Arabidopsis developmental transitions. Plant Mol. Biol. 2011, 76, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Lu, S. Molecular characterization of the SPL gene family in Populus trichocarpa. BMC Plant Biol. 2014, 14, 131. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Tang, G.; Yan, J.; Gu, Y.; Qiao, M.; Fan, R.; Mao, Y.; Tang, X. Construction of short tandem target mimic (STTM) to block the functions of plant and animal microRNAs. Methods 2012, 58, 118–125. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhang, J.; Yan, J.; Gou, F.; Mao, Y.; Tang, G.; Botella, J.R. Short tandem target mimic rice lines uncover functions of miRNAs in regulating important agronomic traits. Proc. Natl. Acad. Sci. USA 2017, 114, 5277–5282. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yang, Z.; Zeng, Q.; Wang, S.; Luo, Y.; Huang, Y.; Xin, Y.; He, N. Abnormal expression of bHLH3 disrupts a flavonoid homeostasis network, causing differences in pigment composition among mulberry fruits. Hortic. Res. 2020, 7, 83. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Zhu, P.; Fan, W.; Feng, Y.; Kou, M.; Hu, J.; Zhao, A. Functional analysis of drought and salt tolerance mechanisms of mulberry RACK1 gene. Tree Physiol. 2019, 39, 2055–2069. [Google Scholar] [CrossRef] [PubMed]
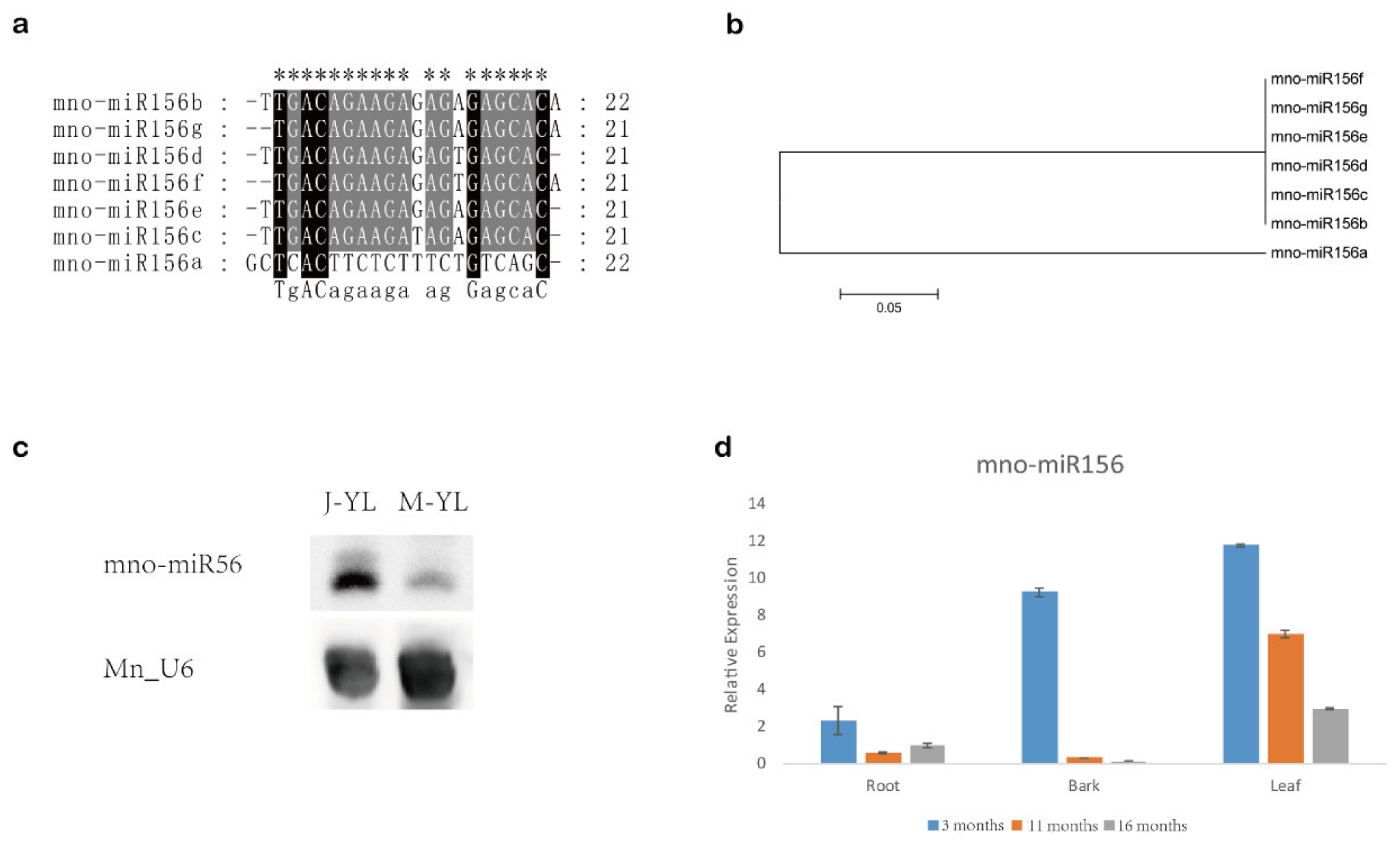


| Name | Sequence | Length | Read in Leaf | Read in Bark | Read in Male Flower |
|---|---|---|---|---|---|
| mno-miR156a | GCTCACTTCTCTTTCTGTCAGC | 22 | 0 | 46 | 4 |
| mno-miR156b | TTGACAGAAGAGAGAGAGCACA | 22 | 6 | 2 | 337 |
| mno-miR156c | TTGACAGAAGATAGAGAGCAC | 21 | 12,787 | 117,270 | 768,488 |
| mno-miR156d | TTGACAGAAGAGAGTGAGCAC | 21 | 2942 | 14,574 | 8723 |
| mno-miR156e | TTGACAGAAGAGAGAGAGCAC | 21 | 2099 | 1277 | 155,199 |
| mno-miR156f | TGACAGAAGAGAGTGAGCACA | 21 | 22 | 378 | 65 |
| mno-miR156g | TGACAGAAGAGAGAGAGCACA | 21 | 8 | 4 | 1198 |
| mno-miR172a | AGAATCTTGATGATGCTGCAT | 21 | 19,265 | 8484 | 1245 |
| mno-miR172b | AGAATCTTGATGATGCTGCAG | 21 | 143 | 13 | 137 |
| mno-miR172c | GGAATCTTGATGATGCTGCAT | 21 | 15 | 266 | 6 |
| mno-miR172d | GGAGCATCATCAAGATTCACA | 21 | 11 | 6 | 4 |
| mno-miR172e | AGAATCCTGATGATGCTGCAT | 21 | 3 | 2 | 0 |
| mno-miR172f | GTAGCATCATCAAGATTCACA | 21 | 1 | 6 | 0 |
| Target Acc. | Expectation | Target Start | Target End | Alignment | Target Aligned Fragment | Inhibition | Molecular Function (Go) | Gene Family |
|---|---|---|---|---|---|---|---|---|
| Morus010792 | 0 | 1554 | 1574 | ::::::::::::::::::::: | GUGCUCUCUAUCUUCUGUCAA | Cleavage | Binding | SPL |
| Morus010123 | 1 | 2201 | 2221 | :::::::::::::::::::: | GUGCUCUCUAUCUUCUGUCAU | Cleavage | Binding | SPL |
| Morus015493 | 1.5 | 2135 | 2155 | ::::::::: ::::::::::: | GUGCUCUCUCUCUUCUGUCAA | Cleavage | Binding | SPL |
| Morus018032 | 1.5 | 2589 | 2609 | ::::::::: ::::::::::: | GUGCUCUCUCUCUUCUGUCAA | Cleavage | Binding | SPL |
| Morus014488 | 1.5 | 2018 | 2038 | :::::::: ::::::::::: | UUGCUCUCUCUCUUCUGUCAA | Cleavage | Binding | SPL |
| Morus021290 | 2 | 1586 | 1606 | : .:::::::::::::::.: | UUUUUCUCUAUCUUCUGUCGA | Cleavage | / | / |
| Morus017456 | 2.5 | 1220 | 1240 | ::::::::: :::::::::: | GUGCUCUCUCUCUUCUGUCAU | Cleavage | Binding | SPL |
| Morus026457 | 2.5 | 2801 | 2821 | ::::::::: :::::::::: | GUGCUCUCUCUCUUCUGUCAU | Cleavage | Binding | SPL |
| Morus026689 | 2.5 | 549 | 569 | :::::.:.:::.::::::.:. | GUGCUUUUUAUUUUCUGUUAG | Cleavage | Binding | WD40 repeat-like |
| Morus002371 | 2.5 | 3369 | 3389 | :::::.:::::.:::: :::: | GUGCUUUCUAUUUUCUAUCAA | Cleavage | Binding | WD40 repeat-like |
| Morus011698 | 2.5 | 3694 | 3714 | : :.:.:::.::::::::..: | GAGUUUUCUGUCUUCUGUUGA | Cleavage | Binding | WD40 repeat-like |
| Morus017876 | 2.5 | 3883 | 3903 | :.:.:::.:.::::::.:: | UAGUUUUCUGUUUUCUGUUAA | Cleavage | catalytic activity | FBP |
| Morus006566 | 2.5 | 1116 | 1136 | :.:: .::::::.:::::: | CCGUUCAUUAUCUUUUGUCAA | Cleavage | / | / |
| Gene_ID1 | Gene_ID2 | Gene Name | T_Start | T_Stop | T_Slice | D_RawReads | D_Pval |
|---|---|---|---|---|---|---|---|
| Morus010123 | XM_010088675.2 | MnSPL13 | 1221 | 1241 | 1232 | 25 | 0.00 |
| Morus010792 | XM_010114085.2 | MnSPL16A | 1078 | 1098 | 1089 | 875 | 0.00 |
| Morus015493 | XM_024169654.1 | MnSPL2 | 1452 | 1472 | 1463 | 68 | 0.00 |
| Morus018032 | XM_010104258.2 | MnSPL15 | 1129 | 1149 | 1140 | 142 | 0.00 |
| Morus026457 | XM_024175201.1 | MnSPL6 | 1580 | 1600 | 1591 | 151 | 0.00 |
| Morus017456 | XM_024173253.1 | MnSPL16B | 758 | 778 | 769 | 140 | 0.01 |
| Morus014488 | XM_010109861.2 | MnSPL4 | 842 | 862 | 853 | 26 | 0.01 |
| Morus018550 | XM_024174390.1 | MnGR-RBP7 | 587 | 607 | 598 | 6 | 0.36 |
| Morus012399 | XM_010089645.2 | MnrsmG | 171 | 191 | 182 | 1 | 0.87 |
| Morus014475 | XM_024170153.1 | MnRH13 | 1613 | 1640 | 1631 | 1 | 0.97 |
| Morus021827 | XM_024177195.1 | MnCABP5 | 446 | 464 | 455 | 1 | 0.99 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, H.; Luo, Y.; Ma, B.; Hu, J.; Lv, Z.; Wei, W.; Hao, H.; Yuan, J.; He, N. Hierarchical Action of Mulberry miR156 in the Vegetative Phase Transition. Int. J. Mol. Sci. 2021, 22, 5550. https://doi.org/10.3390/ijms22115550
Li H, Luo Y, Ma B, Hu J, Lv Z, Wei W, Hao H, Yuan J, He N. Hierarchical Action of Mulberry miR156 in the Vegetative Phase Transition. International Journal of Molecular Sciences. 2021; 22(11):5550. https://doi.org/10.3390/ijms22115550
Chicago/Turabian StyleLi, Hongshun, Yiwei Luo, Bi Ma, Jianqiong Hu, Zhiyuan Lv, Wuqi Wei, Haiye Hao, Jianglian Yuan, and Ningjia He. 2021. "Hierarchical Action of Mulberry miR156 in the Vegetative Phase Transition" International Journal of Molecular Sciences 22, no. 11: 5550. https://doi.org/10.3390/ijms22115550
APA StyleLi, H., Luo, Y., Ma, B., Hu, J., Lv, Z., Wei, W., Hao, H., Yuan, J., & He, N. (2021). Hierarchical Action of Mulberry miR156 in the Vegetative Phase Transition. International Journal of Molecular Sciences, 22(11), 5550. https://doi.org/10.3390/ijms22115550






