Metabolic Dynamics in Short- and Long-Term Microgravity in Human Primary Macrophages
Abstract
:1. Introduction
2. Results
2.1. Identification of Metabolic Effects in Altered Gravity
2.2. Identification of Pools of Gravity-Sensitive Metabolites
2.3. Intra-Experiment Comparison Revealed Potential Gravity-Sensitive Metabolic Networks
2.4. An Inter-Experiment Gravity Effect Comparison Identified a Large Gravisensitive Cluster
3. Discussion
4. Materials and Methods
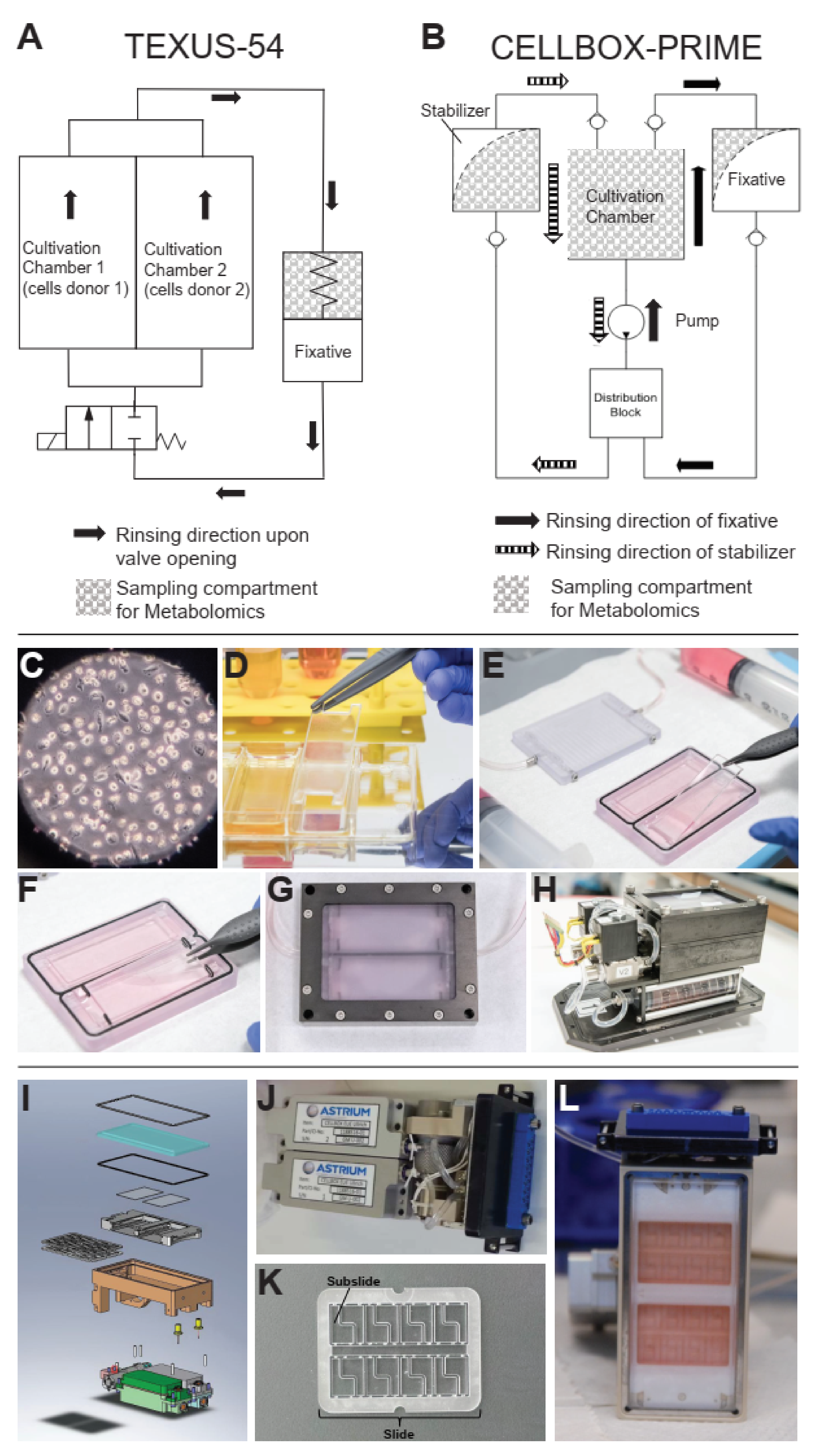
4.1. CELLBOX-PRIME/SpaceX CRS3
4.1.1. Preparation of Primary Human Macrophages
4.1.2. Hardware Concept
4.1.3. Experiment Integration and Upload
4.1.4. Experiment Design and Sampling
4.2. TEXUS-54
4.2.1. Isolation and Cryopreservation of Human Monocytes
4.2.2. Preparation of Primary Human Macrophages from Cryopreserved Human Monocytes
4.2.3. Experiment Hardware
4.2.4. Experiment Design, Conduction, and Sampling
- (1)
- Flight-baseline (BL) sample: cell culture supernatant was separated after the hypergravity phase of the suborbital ballistic flight, directly before the microgravity phase, 50 s after launch.
- (2)
- Flight-microgravity (µg) sample: cell culture supernatant was separated after 325 s of microgravity, 375 s after Launch.
- (3)
- 1 g hardware (H/W) ground control BL-time: samples were in the ground-module and cell culture supernatant was separated concomitantly to the Flight-BL sample.
- (4)
- 1 g hardware (H/W) ground control µg -time: samples were in the ground-module and cell culture supernatant was separated concomitantly to the Flight-µg sample.
4.2.5. Rocket Flight Profile
4.3. Metabolomic Analysis
4.4. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Airbus DS | Airbus Defense and Space |
| TEXUS | German: Technologische Experimente unter Schwerelosigkeit |
| DLR | German Aerospace Center |
| MORABA DLR | Mobile Rocket Base |
| SSC | Swedish Space Cooperation |
References
- Kimzey, S.L. Hematology and immunology studies. In Biomedical Results from Skylab; NASA: Washington, DC, USA, 1977. [Google Scholar]
- Cogoli, A.; Tschopp, A.; Fuchs-Bislin, P. Cell sensitivity to gravity. Science 1984, 225, 228–230. [Google Scholar] [CrossRef]
- Cogoli-Greuter, M.; Meloni, M.A.; Sciola, L.; Spano, A.; Pippia, P.; Monaco, G.; Cogoli, A. Movements and interactions of leukocytes in microgravity. J. Biotechnol. 1996, 47, 279–287. [Google Scholar] [CrossRef]
- NASA Manned Spacecraft Center. Gemini Summary Conference; NASA: Houston, TX, USA, 1967. [Google Scholar]
- Konstantinova, I.V.; Antropova, Y.N.; Legenkov, V.I.; Zazhirey, V.D. Study of reactivity of blood lymphoid cells in crew members of the Soyuz-6, Soyuz-7 and Soyuz-8 spaceships before and after flight(blatogenesis delay of lymphocytes). Space Biol. Med. 1973, 7, 48–55. [Google Scholar]
- Mao, X.; Pecaut, M.; Stodieck, L.; Ferguson, V.; Bateman, T.; Bouxsein, M.; Gridley, D. Biological and metabolic response in STS-135 space-flown mouse skin. Free. Radic. Res. 2014, 48, 890–897. [Google Scholar] [CrossRef]
- Pecaut, M.J.; Mao, X.W.; Bellinger, D.L.; Jonscher, K.R.; Stodieck, L.S.; Ferguson, V.L.; Bateman, T.A.; Mohney, R.P.; Gridley, D.S. Is spaceflight-induced immune dysfunction linked to systemic changes in metabolism? PLoS ONE 2017, 12, e0174174. [Google Scholar] [CrossRef] [Green Version]
- Schmidt, M.A.; Goodwin, T.J. Personalized medicine in human space flight: Using omics based analyses to develop individualized countermeasures that enhance astronaut safety and performance. Metabolomics 2013, 9, 1134–1156. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adrian, A.; Schoppmann, K.; Sromicki, J.; Brungs, S.; von der Wiesche, M.; Hock, B.; Kolanus, W.; Hemmersbach, R.; Ullrich, O. The oxidative burst reaction in mammalian cells depends on gravity. Cell Commun. Signal. 2013, 11, 98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thiel, C.S.; de Zélicourt, D.; Tauber, S.; Adrian, A.; Franz, M.; Simmet, D.M.; Schoppmann, K.; Hauschild, S.; Krammer, S.; Christen, M.; et al. Rapid adaptation to microgravity in mammalian macrophage cells. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thiel, C.S.; Hauschild, S.; Huge, A.; Tauber, S.; Lauber, B.A.; Polzer, J.; Paulsen, K.; Lier, H.; Engelmann, F.; Schmitz, B.; et al. Dynamic gene expression response to altered gravity in human T cells. Sci. Rep. 2017, 7, 1–22. [Google Scholar] [CrossRef] [Green Version]
- Thiel, C.S.; Tauber, S.; Christoffel, S.; Huge, A.; Lauber, B.A.; Polzer, J.; Paulsen, K.; Lier, H.; Engelmann, F.; Schmitz, B.; et al. Rapid coupling between gravitational forces and the transcriptome in human myelomonocytic U937 cells. Sci. Rep. 2018, 8, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Thiel, C.S.; Huge, A.; Hauschild, S.; Tauber, S.; Lauber, B.A.; Polzer, J.; Paulsen, K.; Lier, H.; Engelmann, F.; Schmitz, B.; et al. Stability of gene expression in human T cells in different gravity environments is clustered in chromosomal region 11p15.4. NPJ Microgravity 2017, 3, 1–20. [Google Scholar] [CrossRef] [Green Version]
- Thiel, C.S.; Paulsen, K.; Bradacs, G.; Lust, K.; Tauber, S.; Dumrese, C.; Hilliger, A.; Schoppmann, K.; Biskup, J.; Golz, N.; et al. Rapid alterations of cell cycle control proteins in human T lymphocytes in microgravity. Cell Commun. Signal. 2012, 10, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Mangala, L.S.; Zhang, Y.; He, Z.; Emami, K.; Ramesh, G.T.; Story, M.; Rohde, L.H.; Wu, H. Effects of simulated microgravity on expression profile of microRNA in human lymphoblastoid cells. J. Biol. Chem. 2011, 286, 32483–32490. [Google Scholar] [CrossRef] [Green Version]
- Paulsen, K.; Thiel, C.; Timm, J.; Schmidt, P.M.; Huber, K.; Tauber, S.; Hemmersbach, R.; Seibt, D.; Kroll, H.; Grote, K.-H.; et al. Microgravity-induced alterations in signal transduction in cells of the immune system. Acta Astronaut. 2010, 67, 1116–1125. [Google Scholar] [CrossRef]
- Tauber, S.; Lauber, B.A.; Paulsen, K.; Layer, L.E.; Lehmann, M.; Hauschild, S.; Shepherd, N.R.; Polzer, J.; Segerer, J.; Thiel, C.S.; et al. Cytoskeletal stability and metabolic alterations in primary human macrophages in long-term microgravity. PLoS ONE 2017, 12, e0175599. [Google Scholar] [CrossRef] [Green Version]
- Choukér, A. Stress Challenges and Immunity in Space; Springer: Berlin/Heidelberg, Germany, 2019. [Google Scholar]
- Haas, R.; Zelezniak, A.; Iacovacci, J.; Kamrad, S.; Townsend, S.; Ralser, M. Designing and interpreting ‘multi-omic’experiments that may change our understanding of biology. Curr. Opin. Syst. Biol. 2017, 6, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Kell, D.B.; Oliver, S.G. The metabolome 18 years on: A concept comes of age. Metabolomics 2016, 12, 148. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Bartok, B.; Oler, E.; Liang, K.Y.H.; Budinski, Z.; Berjanskii, M.; Guo, A.; Cao, X.; Wilson, M. MarkerDB: An online database of molecular biomarkers. Nucleic Acids Res. 2020, 49, D1259–D1267. [Google Scholar] [CrossRef] [PubMed]
- Guy, C.; Kaplan, F.; Kopka, J.; Selbig, J.; Hincha, D.K. Metabolomics of temperature stress. Physiol. Plant. 2008, 132, 220–235. [Google Scholar] [CrossRef]
- Kaplan, F.; Kopka, J.; Haskell, D.W.; Zhao, W.; Schiller, K.C.; Gatzke, N.; Sung, D.Y.; Guy, C.L. Exploring the temperature-stress metabolome of Arabidopsis. Plant Physiol. 2004, 136, 4159–4168. [Google Scholar] [CrossRef] [Green Version]
- Wilinski, D.; Winzeler, J.; Duren, W.; Persons, J.L.; Holme, K.J.; Mosquera, J.; Khabiri, M.; Kinchen, J.M.; Freddolino, P.L.; Karnovsky, A. Rapid metabolic shifts occur during the transition between hunger and satiety in Drosophila melanogaster. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Vasilakou, E.; Van Loosdrecht, M.C.M.; Wahl, S.A. Escherichia coli metabolism under short-term repetitive substrate dynamics: Adaptation and trade-offs. Microb. Cell Factories 2020, 19, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Garrett-Bakelman, F.E.; Darshi, M.; Green, S.J.; Gur, R.C.; Lin, L.; Macias, B.R.; McKenna, M.J.; Meydan, C.; Mishra, T.; Nasrini, J.; et al. The NASA Twins Study: A multidimensional analysis of a year-long human spaceflight. Science 2019, 364, eaau8650. [Google Scholar] [CrossRef] [PubMed]
- Michaletti, A.; Gioia, M.; Tarantino, U.; Zolla, L. Effects of microgravity on osteoblast mitochondria: A proteomic and metabolomics profile. Sci. Rep. 2017, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Costantini, D.; Overi, D.; Casadei, L.; Cardinale, V.; Nevi, L.; Carpino, G.; Di Matteo, S.; Safarikia, S.; Valerio, M.; Melandro, F.; et al. Simulated microgravity promotes the formation of tridimensional cultures and stimulates pluripotency and a glycolytic metabolism in human hepatic and biliary tree stem/progenitor cells. Sci. Rep. 2019, 9, 5559. [Google Scholar] [CrossRef] [Green Version]
- Perez de Souza, L.; Alseekh, S.; Brotman, Y.; Fernie, A.R. Network based strategies in metabolomics data analysis and interpretation: From molecular networking to biological interpretation. Expert Rev. Proteom. 2020, 17, 243–255. [Google Scholar] [CrossRef]
- Thiel; Christoffel; Tauber; Vahlensieck; Zélicourt; Layer; Lauber; Polzer; Ullrich, Rapid Cellular Perception of Gravitational Forces in Human Jurkat T Cells and Transduction into Gene Expression Regulation. Int. J. Mol. Sci. 2020, 21, 514. [CrossRef] [Green Version]
- Thiel, C.S.; Tauber, S.; Lauber, B.; Polzer, J.; Seebacher, C.; Uhl, R.; Neelam, S.; Zhang, Y.; Levine, H.; Ullrich, O. Rapid morphological and cytoskeletal response to microgravity in human primary macrophages. Int. J. Mol. Sci. 2019, 20, 2402. [Google Scholar] [CrossRef] [Green Version]
- Wolfe, J.W.; Rummel, J.D. Long-term effects of microgravity and possible countermeasures. Adv. Space Res. 1992, 12, 281–284. [Google Scholar] [CrossRef]
- Po, A.; Giuliani, A.; Masiello, M.G.; Cucina, A.; Catizone, A.; Ricci, G.; Chiacchiarini, M.; Tafani, M.; Ferretti, E.; Bizzarri, M. Phenotypic transitions enacted by simulated microgravity do not alter coherence in gene transcription profile. NPJ Microgravity 2019, 5, 1–13. [Google Scholar] [CrossRef]
- Wu, G. Amino acids: Metabolism, functions, and nutrition. Amino Acids 2009, 37, 1–17. [Google Scholar] [CrossRef]
- Bröer, S.; Bröer, A. Amino acid homeostasis and signalling in mammalian cells and organisms. Biochem. J. 2017, 474, 1935–1963. [Google Scholar] [CrossRef] [Green Version]
- Calder, P.C. Branched-chain amino acids and immunity. J. Nutr. 2006, 136, 288S–293S. [Google Scholar] [CrossRef]
- Grohmann, U.; Mondanelli, G.; Belladonna, M.L.; Orabona, C.; Pallotta, M.T.; Iacono, A.; Puccetti, P.; Volpi, C. Amino-acid sensing and degrading pathways in immune regulation. Cytokine Growth Factor Rev. 2017, 35, 37–45. [Google Scholar] [CrossRef]
- Kedia-Mehta, N.; Finlay, D.K. Competition for nutrients and its role in controlling immune responses. Nat. Commun. 2019, 10, 2123. [Google Scholar] [CrossRef]
- Li, P.; Yin, Y.-L.; Li, D.; Kim, S.W.; Wu, G. Amino acids and immune function. Br. J. Nutr. 2007, 98, 237–252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McGaha, T.L.; Huang, L.; Lemos, H.; Metz, R.; Mautino, M.; Prendergast, G.C.; Mellor, A.L. Amino acid catabolism: A pivotal regulator of innate and adaptive immunity. Immunol. Rev. 2012, 249, 135–157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ullrich, O.; Huber, K.; Lang, K. Signal transduction in cells of the immune system in microgravity. Cell Commun. Signal. 2008, 6, 1–6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Massey, K.A.; Blakeslee, C.H.; Pitkow, H.S. A review of physiological and metabolic effects of essential amino acids. Amino Acids 1998, 14, 271–300. [Google Scholar] [CrossRef]
- Bachmair, A.; Finley, D.; Varshavsky, A. In vivo half-life of a protein is a function of its amino-terminal residue. Science 1986, 234, 179–186. [Google Scholar] [CrossRef] [PubMed]
- Koren, I.; Timms, R.T.; Kula, T.; Xu, Q.; Li, M.Z.; Elledge, S.J. The eukaryotic proteome is shaped by E3 ubiquitin ligases targeting C-terminal degrons. Cell 2018, 173, 1622–1635. [Google Scholar] [CrossRef] [Green Version]
- Lin, H.-C.; Yeh, C.-W.; Chen, Y.-F.; Lee, T.-T.; Hsieh, P.-Y.; Rusnac, D.V.; Lin, S.-Y.; Elledge, S.J.; Zheng, N.; Yen, H.-C.S. C-terminal end-directed protein elimination by CRL2 ubiquitin ligases. Mol. Cell 2018, 70, 602–613. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Varshavsky, A. N-degron and C-degron pathways of protein degradation. Proc. Natl. Acad. Sci. USA 2019, 116, 358–366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Timms, R.T.; Koren, I. Tying up loose ends: The N-degron and C-degron pathways of protein degradation. Biochem. Soc. Trans. 2020, 48, 1557–1567. [Google Scholar] [CrossRef]
- Berko, D.; Tabachnick-Cherny, S.; Shental-Bechor, D.; Cascio, P.; Mioletti, S.; Levy, Y.; Admon, A.; Ziv, T.; Tirosh, B.; Goldberg, A.L. The direction of protein entry into the proteasome determines the variety of products and depends on the force needed to unfold its two termini. Mol. Cell 2012, 48, 601–611. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flick, K.; Kaiser, P. Protein degradation and the stress response. Semin. Cell Dev. Biol. 2012, 23, 515–522. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Okumura, F.; Fujiki, Y.; Oki, N.; Osaki, K.; Nishikimi, A.; Fukui, Y.; Nakatsukasa, K.; Kamura, T. Cul5-type ubiquitin ligase KLHDC1 contributes to the elimination of truncated SELENOS produced by failed UGA/Sec decoding. iScience 2020, 23, 1009700. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thiel, C.S.; Hauschild, S.; Tauber, S.; Paulsen, K.; Raig, C.; Raem, A.; Biskup, J.; Gutewort, A.; Hürlimann, E.; Unverdorben, F.; et al. Identification of reference genes in human myelomonocytic cells for gene expression studies in altered gravity. BioMed Res. Int. 2015, 2015, 363575. [Google Scholar] [CrossRef] [PubMed]
- Rhee, H.J.; Kim, E.-J.; Lee, J.K. Physiological polyamines: Simple primordial stress molecules. J. Cell. Mol. Med. 2007, 11, 685–7033. [Google Scholar] [CrossRef] [Green Version]
- Kusano, T.; Berberich, T.; Tateda, C.; Takahashi, Y. Polyamines: Essential factors for growth and survival. Planta 2008, 228, 367–381. [Google Scholar] [CrossRef]
- Kumar, N.; Basundra, R.; Maiti, S. Elevated polyamines induce c-MYC overexpression by perturbing quadruplex-WC duplex equilibrium. Nucleic Acids Res. 2009, 37, 3321–3331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, T.; Gallo, M.A.; Klinge, C.M.; Thomas, T.J. Polyamine-mediated conformational perturbations in DNA alter the binding of estrogen receptor to poly(dG-m5dC).poly(dG-m5dC) and a plasmid containing the estrogen response element. J. Steroid Biochem. Mol. Biol. 1995, 54, 89–99. [Google Scholar] [CrossRef]
- Ray, R.M.; Zimmerman, B.J.; McCormack, S.A.; Patel, T.B.; Johnson, L.R. Polyamine depletion arrests cell cycle and induces inhibitors p21(Waf1/Cip1), p27(Kip1), and p53 in IEC-6 cells. Am. J. Physiol. 1999, 276, C684–C691. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, Q.W.; Tabor, C.W.; Tabor, H. Deletion mutations in the speED operon: Spermidine is not essential for the growth of Escherichia coli. Gene 1993, 126, 115–117. [Google Scholar] [CrossRef]
- Das, K.C.; Misra, H.P. Hydroxyl radical scavenging and singlet oxygen quenching properties of polyamines. Mol. Cell. Biochem. 2004, 262, 127–133. [Google Scholar] [CrossRef] [PubMed]
- Ha, H.C.; Sirisoma, N.S.; Kuppusamy, P.; Zweier, J.L.; Woster, P.M.; Casero, R.A. The natural polyamine spermine functions directly as a free radical scavenger. Proc. Natl. Acad. Sci. USA 1998, 95, 11140–11145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rider, J.E.; Hacker, A.; Mackintosh, C.A.; Pegg, A.E.; Woster, P.M.; Casero, R.A. Spermine and spermidine mediate protection against oxidative damage caused by hydrogen peroxide. Amino Acids 2007, 33, 231–240. [Google Scholar] [CrossRef] [PubMed]
- Iyer, R.; Delcour, A.H. Complex inhibition of OmpF and OmpC bacterial porins by polyamines. J. Biol. Chem. 1997, 272, 18595–18601. [Google Scholar] [CrossRef] [Green Version]
- Yamaguchi, K.; Takahashi, Y.; Berberich, T.; Imai, A.; Miyazaki, A.; Takahashi, T.; Michael, A.; Kusano, T. The polyamine spermine protects against high salt stress in Arabidopsis thaliana. FEBS Lett. 2006, 580, 6783–6788. [Google Scholar] [CrossRef] [Green Version]
- Jung, I.L.; Kim, I.G. Transcription of ahpC, katG, and katE genes in Escherichia coli is regulated by polyamines: Polyamine-deficient mutant sensitive to H2O2-induced oxidative damage. Biochem. Biophys. Res. Commun. 2003, 301, 915–922. [Google Scholar] [CrossRef]
- Krüger, A.; Vowinckel, J.; Mülleder, M.; Grote, P.; Capuano, F.; Bluemlein, K.; Ralser, M. Tpo1-mediated spermine and spermidine export controls cell cycle delay and times antioxidant protein expression during the oxidative stress response. EMBO Rep. 2013, 14, 1113–1119. [Google Scholar] [CrossRef]
- Tkachenko, A.G.; Nesterova, L.Y. Polyamines as modulators of gene expression under oxidative stress in Escherichia coli. Biochem. Biokhimiia 2003, 68, 850–856. [Google Scholar] [CrossRef]
- Miller-Fleming, L.; Olin-Sandoval, V.; Campbell, K.; Ralser, M. Remaining mysteries of molecular biology: The role of polyamines in the cell. J. Mol. Biol. 2015, 427, 3389–3406. [Google Scholar] [CrossRef] [PubMed]
- Ilaiwy, A.; Quintana, M.T.; Bain, J.R.; Muehlbauer, M.J.; Brown, D.I.; Stansfield, W.E.; Willis, M.S. Cessation of biomechanical stretch model of C2C12 cells models myocyte atrophy and anaplerotic changes in metabolism using non-targeted metabolomics analysis. Int. J. Biochem. Cell Biol. 2016, 79, 80–92. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Globus, R.K.; Morey-Holton, E. Hindlimb unloading: Rodent analog for microgravity. J. Appl. Physiol. 2016, 120, 1196–1206. [Google Scholar] [CrossRef]
- Krebs, H.A.; Lund, P. Aspects of the regulation of the metabolism of branched-chain amino acids. Adv. Enzym. Regul. 1977, 15, 375–394. [Google Scholar] [CrossRef]
- Hörl, W.H.; Kittel, R.; Heidland, A. Effects of high doses of leucine and ketoleucine on glycogen and protein metabolism in acute uremia. Am. J. Clin. Nutr. 1980, 33, 1468–1475. [Google Scholar] [CrossRef] [Green Version]
- Walser, M. Role of branched-chain ketoacids in protein metabolism. Kidney Int. 1990, 38, 595–604. [Google Scholar] [CrossRef] [Green Version]
- Sgaravatti, A.M.; Rosa, R.B.; Schuck, P.F.; Ribeiro, C.A.J.; Wannmacher, C.M.D.; Wyse, A.T.S.; Dutra-Filho, C.S.; Wajner, M. Inhibition of brain energy metabolism by the alpha-keto acids accumulating in maple syrup urine disease. Biochim. Biophys. Acta 2003, 1639, 232–238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, M.; Nakai, N.; Ishigure, K.; Nonami, T.; Nagasaki, M.; Obayashi, M.; Li, Z.; Sato, Y.; Fujitsuka, N.; Murakami, T.; et al. The α-ketoisocaproate catabolism in human and rat livers. Biochem. Biophys. Res. Commun. 2000, 276, 1080–1084. [Google Scholar] [CrossRef] [PubMed]
- Ævarsson, A.; Chuang, J.L.; Wynn, R.M.; Turley, S.; Chuang, D.T.; Hol, W.G.J. Crystal structure of human branched-chain α-ketoacid dehydrogenase and the molecular basis of multienzyme complex deficiency in maple syrup urine disease. Structure 2000, 8, 277–291. [Google Scholar] [CrossRef] [Green Version]
- Kastritis, P.L.; Gavin, A.-C. Enzymatic complexes across scales. Essays Biochem. 2018, 62, 501–514. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kohn, F.; Hauslage, J.; Hanke, W. Membrane fluidity changes, a basic mechanism of interaction of gravity with cells? Microgravity Sci. Technol. 2017, 29, 337–342. [Google Scholar] [CrossRef]
- Holecek, M. Branched-chain amino acids in health and disease: Metabolism, alterations in blood plasma, and as supplements. Nutr. Metab. 2018, 15, 33. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carlsson, H.; Rollborn, N.; Herman, S.; Freyhult, E.; Svenningsson, A.; Burman, J.; Kultima, K. Metabolomics of cerebrospinal fluid from healthy subjects reveal metabolites associated with ageing. Metabolites 2021, 11, 126. [Google Scholar] [CrossRef] [PubMed]
- Strollo, F.; Vernikos, J. Aging-like metabolic and adrenal changes in microgravity: State of the art in preparation for Mars. Neurosci. Biobehav. Rev. 2021, 126, 236–242. [Google Scholar] [CrossRef]
- Riley, D.A.; Ellis, S.; Slocum, G.R.; Sedlak, F.R.; Bain, J.L.; Krippendorf, B.B.; Lehman, C.T.; Macias, M.Y.; Thompson, J.L.; Vijayan, K.; et al. In-flight and postflight changes in skeletal muscles of SLS-1 and SLS-2 spaceflown rats. J. Appl. Physiol. 1996, 81, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Radugina, E.A.; Almeida, E.A.C.; Blaber, E.; Poplinskaya, V.A.; Markitantova, Y.V.; Grigoryan, E.N. Exposure to microgravity for 30 days onboard Bion M1 caused muscle atrophy and impaired regeneration in murine femoral Quadriceps. Life Sci. Space Res. 2018, 16, 18–25. [Google Scholar] [CrossRef]
- Okada, R.; Fujita, S.-i.; Suzuki, R.; Hayashi, T.; Tsubouchi, H.; Kato, C.; Sadaki, S.; Kanai, M.; Fuseya, S.; Inoue, Y.; et al. Transcriptome analysis of gravitational effects on mouse skeletal muscles under microgravity and artificial 1 g onboard environment. Sci. Rep. 2021, 11, 9168. [Google Scholar] [CrossRef]
- Moosavi, D.; Wolovsky, D.; Depompeis, A.; Uher, D.; Lennington, D.; Bodden, R.; Garber, C.E. The effects of spaceflight microgravity on the musculoskeletal system of humans and animals, with an emphasis on exercise as a countermeasure: A systematic scoping review. Physiol. Res. 2021, 70, 119–151. [Google Scholar] [CrossRef]
- Stein, T.P.; Leskiw, M.J.; Schluter, M.D. Diet and nitrogen metabolism during spaceflight on the shuttle. J. Appl. Physiol. 1996, 81, 82–97. [Google Scholar] [CrossRef]
- Stein, T.P.; Leskiw, M.J.; Schluter, M.D.; Donaldson, M.R.; Larina, I. Protein kinetics during and after long-duration spaceflight on MIR. Am. J. Physiol. Endocrinol. Metab. 1999, 276, E1014–E1021. [Google Scholar] [CrossRef]
- Stein, T.P.; Schluter, M.D. Plasma amino acids during human spaceflight. Aviat. space Environ. Med. 1999, 70, 250–255. [Google Scholar]
- Pajares, M.; Jiménez-Moreno, N.; Dias, I.H.K.; Debelec, B.; Vucetic, M.; Fladmark, K.E.; Basaga, H.; Ribaric, S.; Milisav, I.; Cuadrado, A. Redox control of protein degradation. Redox Biol. 2015, 6, 409–420. [Google Scholar] [CrossRef] [Green Version]
- Jeong, A.J.; Kim, Y.J.; Lim, M.H.; Lee, H.; Noh, K.; Kim, B.-H.; Chung, J.W.; Cho, C.-H.; Kim, S.; Ye, S.-K. Microgravity induces autophagy via mitochondrial dysfunction in human Hodgkin’s lymphoma cells. Sci. Rep. 2018, 8, 14646. [Google Scholar] [CrossRef] [Green Version]
- Wu, M.Y.; Lu, J.H. Autophagy and macrophage functions: Inflammatory response and phagocytosis. Cells 2019, 9, 70. [Google Scholar] [CrossRef] [Green Version]
- Mosser, D.M.; Edwards, J.P. Exploring the full spectrum of macrophage activation. Nat. Rev. Immunol. 2008, 8, 958–969. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, N.; Vogel, S.N.; Van Way, C.; Papasian, C.J.; Qureshi, A.A.; Morrison, D.C. The proteasome. Immunol. Res. 2005, 31, 243–260. [Google Scholar] [CrossRef] [PubMed]
- Ikemoto, M.; Nikawa, T.; Takeda, S.I.; Watanabe, C.; Kitano, T.; Baldwin, K.M.; Izumi, R.; Nonaka, I.; Towatari, T.; Teshima, S. Space shuttle flight (STS-90) enhances degradation of rat myosin heavy chain in association with activation of ubiquitin-proteasome pathway. FASEB J. 2001, 15, 1279–1281. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ullrich, O.; Ciftci, Ö.; Hass, R. Proteasome activation by poly-ADP-ribose-polymerase in human myelomonocytic cells after oxidative stress. Free. Radic. Biol. Med. 2000, 29, 995–1004. [Google Scholar] [CrossRef]
- Cogoli, A. Space flight and the immune system. Vaccine 1993, 11, 496–503. [Google Scholar] [CrossRef]
- Kaur, I.; Simons, E.R.; Castro, V.A.; Ott, C.M.; Pierson, D.L. Changes in monocyte functions of astronauts. Brain Behav. Immun. 2005, 19, 547–554. [Google Scholar] [CrossRef]
- Ludtka, C.; Silberman, J.; Moore, E.; Allen, J.B. Macrophages in microgravity: The impact of space on immune cells. NPJ Microgravity 2021, 7, 13. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Tian, H.; Wang, P.; Li, L.; Zhang, Z.; Zhang, J.; Zhao, Y. Spaceflight and simulated microgravity suppresses macrophage development via altered RAS/ERK/NFκB and metabolic pathways. Cell. Mol. Immunol. 2021, 18, 1489–1502. [Google Scholar] [CrossRef] [Green Version]
- Luedemann, A.; Strassburg, K.; Erban, A.; Kopka, J. TagFinder for the quantitative analysis of gas chromatography--mass spectrometry (GC-MS)-based metabolite profiling experiments. Bioinformatics 2008, 24, 732–737. [Google Scholar] [CrossRef] [PubMed]
- Kopka, J.; Schauer, N.; Krueger, S.; Birkemeyer, C.; Usadel, B.; Bergmüller, E.; Dörmann, P.; Weckwerth, W.; Gibon, Y.; Stitt, M.; et al. [email protected]: The golm metabolome database. Bioinformatics 2004, 21, 1635–1638. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wishart, D.; Tzur, D.; Knox, C. HMDB: The human metabolome database. Nucleic Acids Res. 2007, 35, D521–D526. [Google Scholar] [CrossRef]
- Frolkis, A.; Knox, C.; Lim, E.; Jewison, T.; Law, V.; Hau, D.D.; Liu, P.; Gautam, B.; Ly, S.; Guo, A.C.; et al. SMPDB: The small molecule pathway database. Nucleic Acids Res. 2009, 38 (Suppl. 1), D480–D487. [Google Scholar] [CrossRef] [Green Version]










| Selected Metabolites Occurring in All Datasets | TEXUS-54 hypg | TEXUS-54 µg | CELLBOX-PRIME | |
|---|---|---|---|---|
| Yellow Cluster | 3-Hydroxybutyric acid | + | ++ | ++ |
| L-Cysteine | + | + | ||
| L-Phenylalanine | + | |||
| L-Valine | + | + | ||
| Threose | + | + | ||
| Pink Cluster | L-Arginine | ++ | ++ | |
| myo-Inositole | ++ | ++ | ||
| L-Methionine | + | |||
| L-Proline | ++ | ++ | + | |
| L-Aspartic Acid | ||||
| Glycine | + | ++ | ||
| Ketoleucine | ++ | ++ | ||
| Ornithine | + | + |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thiel, C.S.; Vahlensieck, C.; Bradley, T.; Tauber, S.; Lehmann, M.; Ullrich, O. Metabolic Dynamics in Short- and Long-Term Microgravity in Human Primary Macrophages. Int. J. Mol. Sci. 2021, 22, 6752. https://doi.org/10.3390/ijms22136752
Thiel CS, Vahlensieck C, Bradley T, Tauber S, Lehmann M, Ullrich O. Metabolic Dynamics in Short- and Long-Term Microgravity in Human Primary Macrophages. International Journal of Molecular Sciences. 2021; 22(13):6752. https://doi.org/10.3390/ijms22136752
Chicago/Turabian StyleThiel, Cora S., Christian Vahlensieck, Timothy Bradley, Svantje Tauber, Martin Lehmann, and Oliver Ullrich. 2021. "Metabolic Dynamics in Short- and Long-Term Microgravity in Human Primary Macrophages" International Journal of Molecular Sciences 22, no. 13: 6752. https://doi.org/10.3390/ijms22136752









