De novo ssRNA Aptamers against the SARS-CoV-2 Main Protease: In Silico Design and Molecular Dynamics Simulation
Abstract
:1. Introduction
2. Results
2.1. Design of Aptamers against Mpro
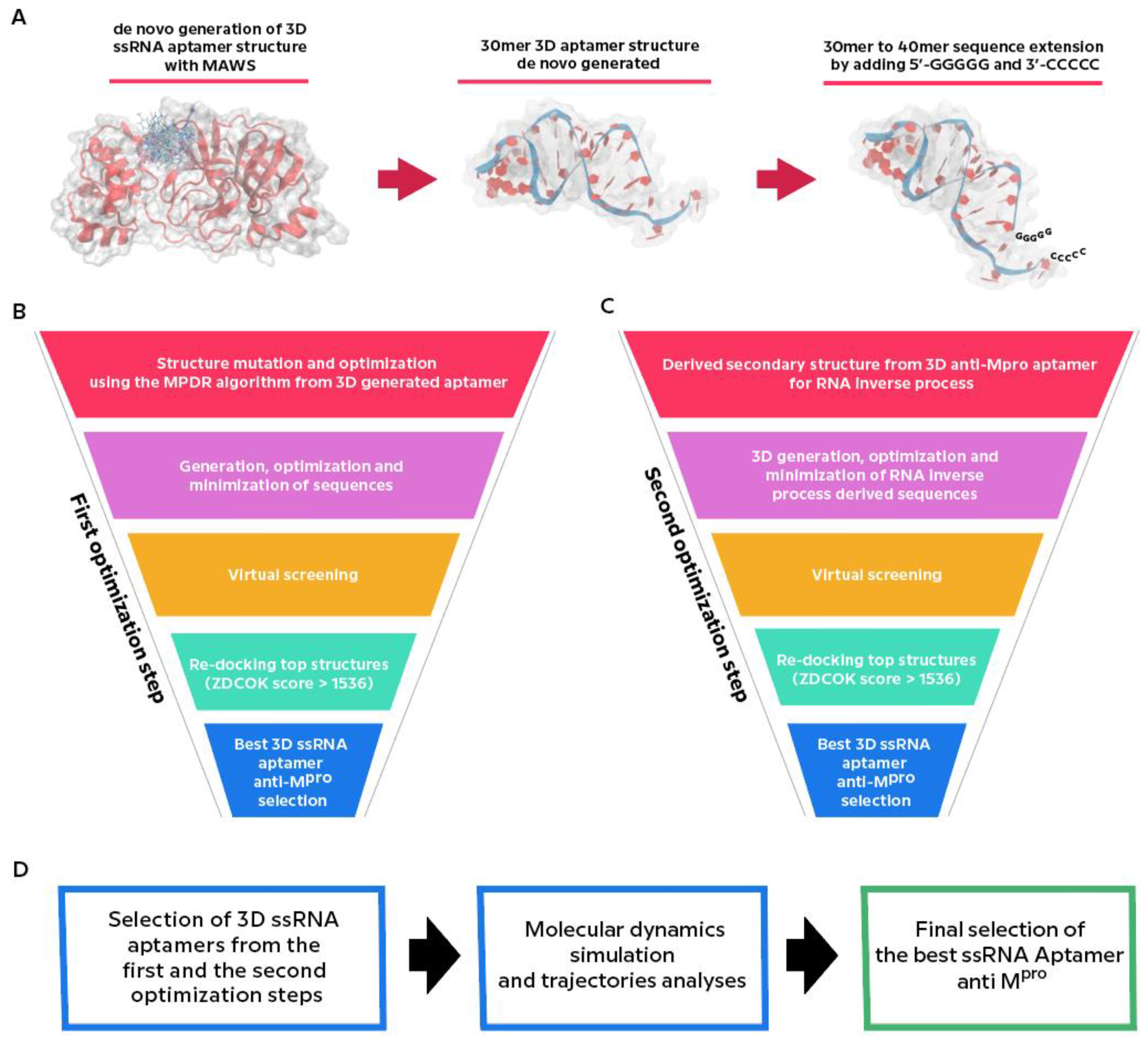
2.2. Generation of De novo ssRNA Aptamer and First Optimization Step
Molecular Docking Simulation of De novo ssRNA Optimized Aptamer against Mpro
2.3. Generation of the 3D ssRNA Aptamer Structures Starting from MAptapro Sequence: Second Optimization Step
2.3.1. Inverse RNA Processing
2.3.2. Molecular Docking Simulation of 3D ssRNA Aptamer Structures
2.4. Molecular Dynamics Simulations and Trajectory Analyses of MAptapro, MAptapro-RI1, and MAptapro-RI2 Aptamers
2.4.1. The Root-Mean-Square Deviation (RMSD)
2.4.2. The Radius of Gyration (Rg)
2.4.3. Number of Hydrogen-Bonds
2.5. Binding Free Energy Analysis of MAptapro, MAptapro-RI1, and MAptapro-RI2 Aptamers
3. Discussion
4. Materials and Methods
4.1. De novo Generation of 3D ssRNA Aptamer Structure against Mpro of SARS-CoV-2
4.2. Generation of the 3D ssRNA Aptamer Structure from a Given Secondary Structure: RNA Inverse Process
4.3. Molecular Docking Simulation
4.4. Molecular Dynamics Simulation
4.5. MD Simulation Analysis
4.6. Graphical Images
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- WHO. Coronavirus (COVID-19) Dashboard|WHO Coronavirus Disease (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 16 March 2021).
- Stasi, C.; Fallani, S.; Voller, F.; Silvestri, C. Treatment for COVID-19: An Overview. Eur. J. Pharm. 2020, 889. [Google Scholar] [CrossRef]
- Rahman, M.R.; Banik, A.; Chowdhury, I.M.; Sajib, E.H.; Sarkar, S. Identification of Potential Antivirals against SARS-CoV-2 Using Virtual Screening Method. Inform. Med. Unlocked 2021, 23, 100531. [Google Scholar] [CrossRef]
- Deb, P.; Molla, M.M.A.; Rahman, K.M.S.-U. An Update to Monoclonal Antibody as Therapeutic Option against COVID-19. Biosaf. Health 2021. [Google Scholar] [CrossRef]
- Castells, M.C.; Phillips, E.J. Maintaining Safety with SARS-CoV-2 Vaccines. N. Engl. J. Med. 2021, 384, 643–649. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjee, A.; Saha, M.; Halder, A.; Debnath, A.; Mukherjee, O. Therapeutics and Vaccines: Strengthening Our Fight Against the Global Pandemic COVID-19. Curr. Microbiol. 2021, 78, 435–448. [Google Scholar] [CrossRef] [PubMed]
- Izda, V.; Jeffries, M.A.; Sawalha, A.H. COVID-19: A Review of Therapeutic Strategies and Vaccine Candidates. Clin. Immunol. 2021, 222, 108634. [Google Scholar] [CrossRef]
- Dai, L.; Gao, G.F. Viral Targets for Vaccines against COVID-19. Nat. Rev. Immunol. 2021, 21, 73–82. [Google Scholar] [CrossRef]
- Zhou, B.; Thi Nhu Thao, T.; Hoffmann, D.; Taddeo, A.; Ebert, N.; Labroussaa, F.; Pohlmann, A.; King, J.; Steiner, S.; Kelly, J.N.; et al. SARS-CoV-2 Spike D614G Change Enhances Replication and Transmission. Nature 2021, 1–8. [Google Scholar] [CrossRef]
- SARS-CoV-2 Variants of Concern|CDC. Available online: https://www.cdc.gov/coronavirus/2019-ncov/cases-updates/variant-surveillance/variant-info.html (accessed on 16 March 2021).
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic Characterisation and Epidemiology of 2019 Novel Coronavirus: Implications for Virus Origins and Receptor Binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef] [Green Version]
- Yoshimoto, F.K. A Biochemical Perspective of the Nonstructural Proteins (NSPs) and the Spike Protein of SARS CoV-2. Protein J. 2021, 40, 260–295. [Google Scholar] [CrossRef]
- Li, J.; Zhan, P.; Liu, X. Targeting the Entry Step of SARS-CoV-2: A Promising Therapeutic Approach. Signal Transduct. Target. Ther. 2020, 5, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Harrison, A.G.; Lin, T.; Wang, P. Mechanisms of SARS-CoV-2 Transmission and Pathogenesis. Trends Immunol. 2020, 41, 1100–1115. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.Y.; Zhao, R.; Gao, L.J.; Gao, X.F.; Wang, D.P.; Cao, J.M. SARS-CoV-2: Structure, Biology, and Structure-Based Therapeutics Development. Front. Cell. Infect. Microbiol. 2020, 10, 724. [Google Scholar] [CrossRef] [PubMed]
- Finkel, Y.; Mizrahi, O.; Nachshon, A.; Weingarten-Gabbay, S.; Morgenstern, D.; Yahalom-Ronen, Y.; Tamir, H.; Achdout, H.; Stein, D.; Israeli, O.; et al. The Coding Capacity of SARS-CoV-2. Nature 2021, 589, 125–130. [Google Scholar] [CrossRef] [PubMed]
- Mousavizadeh, L.; Ghasemi, S. Genotype and Phenotype of COVID-19: Their Roles in Pathogenesis. J. Microbiol. Immunol. Infect. 2020, 54, 159–163. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Lin, D.; Sun, X.; Curth, U.; Drosten, C.; Sauerhering, L.; Becker, S.; Rox, K.; Hilgenfeld, R. Crystal Structure of SARS-CoV-2 Main Protease Provides a Basis for Design of Improved a-Ketoamide Inhibitors. Science 2020, 368, 409–412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, H.; Yang, M.; Ding, Y.; Liu, Y.; Lou, Z.; Zhou, Z.; Sun, L.; Mo, L.; Ye, S.; Pang, H.; et al. The Crystal Structures of Severe Acute Respiratory Syndrome Virus Main Protease and Its Complex with an Inhibitor. Proc. Natl. Acad. Sci. USA 2003, 100, 13190–13195. [Google Scholar] [CrossRef] [Green Version]
- Xia, B.; Kang, X. Activation and Maturation of SARS-CoV Main Protease. Protein Cell 2011, 2, 282–290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abian, O.; Ortega-Alarcon, D.; Jimenez-Alesanco, A.; Ceballos-Laita, L.; Vega, S.; Reyburn, H.T.; Rizzuti, B.; Velazquez-Campoy, A. Structural Stability of SARS-CoV-2 3CLpro and Identification of Quercetin as an Inhibitor by Experimental Screening. Int. J. Biol. Macromol. 2020, 164, 1693–1703. [Google Scholar] [CrossRef]
- Jin, Z.; Du, X.; Xu, Y.; Deng, Y.; Liu, M.; Zhao, Y.; Zhang, B.; Li, X.; Zhang, L.; Peng, C.; et al. Structure of Mpro from SARS-CoV-2 and Discovery of Its Inhibitors. Nature 2020, 582, 289–293. [Google Scholar] [CrossRef] [Green Version]
- Bzówka, M.; Mitusińska, K.; Raczyńska, A.; Samol, A.; Tuszyński, J.A.; Góra, A. Structural and Evolutionary Analysis Indicate That the Sars-COV-2 Mpro Is a Challenging Target for Small-Molecule Inhibitor Design. Int. J. Mol. Sci. 2020, 21, 3099. [Google Scholar] [CrossRef]
- Dai, W.; Zhang, B.; Jiang, X.M.; Su, H.; Li, J.; Zhao, Y.; Xie, X.; Jin, Z.; Peng, J.; Liu, F.; et al. Structure-Based Design of Antiviral Drug Candidates Targeting the SARS-CoV-2 Main Protease. Science 2020, 368, 1331–1335. [Google Scholar] [CrossRef] [Green Version]
- Rut, W.; Groborz, K.; Zhang, L.; Sun, X.; Zmudzinski, M.; Pawlik, B.; Wang, X.; Jochmans, D.; Neyts, J.; Młynarski, W.; et al. SARS-CoV-2 Mpro Inhibitors and Activity-Based Probes for Patient-Sample Imaging. Nat. Chem. Biol. 2021, 17, 222–228. [Google Scholar] [CrossRef]
- Pillaiyar, T.; Manickam, M.; Namasivayam, V.; Hayashi, Y.; Jung, S.H. An Overview of Severe Acute Respiratory Syndrome-Coronavirus (SARS-CoV) 3CL Protease Inhibitors: Peptidomimetics and Small Molecule Chemotherapy. J. Med. Chem. 2016, 59, 6595–6628. [Google Scholar] [CrossRef]
- Kuo, C.-J.; Liang, P.-H. Characterization and Inhibition of the Main Protease of Severe Acute Respiratory Syndrome Coronavirus. ChemBioEng Rev. 2015, 2, 118–132. [Google Scholar] [CrossRef]
- Ding, L.; Zhang, X.X.; Wei, P.; Fan, K.; Lai, L. The Interaction between Severe Acute Respiratory Syndrome Coronavirus 3C-like Proteinase and a Dimeric Inhibitor by Capillary Electrophoresis. Anal. Biochem. 2005, 343, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Wei, P.; Fan, K.; Chen, H.; Ma, L.; Huang, C.; Tan, L.; Xi, D.; Li, C.; Liu, Y.; Cao, A.; et al. The N-Terminal Octapeptide Acts as a Dimerization Inhibitor of SARS Coronavirus 3C-like Proteinase. Biochem. Biophys. Res. Commun. 2006, 339, 865–872. [Google Scholar] [CrossRef]
- Hayashi, T.; Matsuda, T.; Nagata, T.; Katahira, M.; Kinoshita, M. Mechanism of Protein-RNA Recognition: Analysis Based on the Statistical Mechanics of Hydration. Phys. Chem. Chem. Phys. 2018, 20, 9167–9180. [Google Scholar] [CrossRef]
- Zhou, J.; Rossi, J. Aptamers as Targeted Therapeutics: Current Potential and Challenges. Nat. Rev. Drug Discov. 2017, 16, 181–202. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Zhou, S.; Cai, Y.; Tang, F. Nucleic Acid Aptamer Application in Diagnosis and Therapy of Colorectal Cancer Based on Cell-SELEX Technology. NPJ Precis. Oncol. 2017, 1, 37. [Google Scholar] [CrossRef] [Green Version]
- Zhu, G.; Niu, G.; Chen, X. Aptamer-Drug Conjugates. Bioconjug. Chem. 2015, 26, 2186–2197. [Google Scholar] [CrossRef] [Green Version]
- Kulabhusan, P.K.; Hussain, B.; Yüce, M. Current Perspectives on Aptamers as Diagnostic Tools and Therapeutic Agents. Pharmaceutics 2020, 12, 646. [Google Scholar] [CrossRef] [PubMed]
- Buglak, A.A.; Samokhvalov, A.V.; Zherdev, A.V.; Dzantiev, B.B. Methods and Applications of in Silico Aptamer Design and Modeling. Int. J. Mol. Sci. 2020, 21, 8420. [Google Scholar] [CrossRef] [PubMed]
- Team:Heidelberg/Software/Maws—2015.Igem.Org. Available online: http://2015.igem.org/Team:Heidelberg/software/maws (accessed on 18 March 2021).
- Team:Athens/Software—2019.Igem.Org. Available online: https://2019.igem.org/Team:Athens/Software (accessed on 18 March 2021).
- Bavi, R.; Liu, Z.; Han, Z.; Zhang, H.; Gu, Y. In Silico Designed RNA Aptamer against Epithelial Cell Adhesion Molecule for Cancer Cell Imaging. Biochem. Biophys. Res. Commun. 2019, 509, 937–942. [Google Scholar] [CrossRef] [PubMed]
- Ahirwar, R.; Nahar, S.; Aggarwal, S.; Ramachandran, S.; Maiti, S.; Nahar, P. In Silico Selection of an Aptamer to Estrogen Receptor Alpha Using Computational Docking Employing Estrogen Response Elements as Aptamer-Alike Molecules. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Rhinehardt, K.L.; Srinivas, G.; Mohan, R.V. Molecular Dynamics Simulation Analysis of Anti-MUC1 Aptamer and Mucin 1 Peptide Binding. J. Phys. Chem. B 2015, 119, 6571–6583. [Google Scholar] [CrossRef]
- Xiao, J.; Salsbury, F.R. Molecular Dynamics Simulations of Aptamer-Binding Reveal Generalized Allostery in Thrombin. J. Biomol. Struct. Dyn. 2017, 35, 3354–3369. [Google Scholar] [CrossRef]
- Ruan, M.; Seydou, M.; Noel, V.; Piro, B.; Maurel, F.; Barbault, F. Molecular Dynamics Simulation of a RNA Aptasensor. J. Phys. Chem. B 2017, 121, 4071–4080. [Google Scholar] [CrossRef]
- Hofacker, I.L.; Fontana, W.; Stadler, P.F.; Bonhoeffer, L.S.; Tacker, M.; Schuster, P. Fast Folding and Comparison of RNA Secondary Structures. Mon. Chem. Chem. Mon. 1994, 125, 167–188. [Google Scholar] [CrossRef]
- Lorenz, R.; Bernhart, S.H.; Höner zu Siederdissen, C.; Tafer, H.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. ViennaRNA Package 2.0. Algorithms Mol. Biol. 2011, 6, 26. [Google Scholar] [CrossRef]
- Goyal, B.; Goyal, D. Targeting the Dimerization of the Main Protease of Coronaviruses: A Potential Broad-Spectrum Therapeutic Strategy. ACS Comb. Sci. 2020, 22, 297–305. [Google Scholar] [CrossRef]
- Greaney, A.J.; Starr, T.N.; Gilchuk, P.; Zost, S.J.; Binshtein, E.; Loes, A.N.; Hilton, S.K.; Huddleston, J.; Eguia, R.; Crawford, K.H.D.; et al. Complete Mapping of Mutations to the SARS-CoV-2 Spike Receptor-Binding Domain That Escape Antibody Recognition. Cell Host Microbe 2021, 29, 44–57.e9. [Google Scholar] [CrossRef] [PubMed]
- Gurung, A.B.; Ali, M.A.; Lee, J.; Farah, M.A.; Al-Anazi, K.M. Unravelling Lead Antiviral Phytochemicals for the Inhibition of SARS-CoV-2 Mpro Enzyme through in Silico Approach. Life Sci. 2020, 255, 117831. [Google Scholar] [CrossRef] [PubMed]
- Narkhede, R.R.; Pise, A.V.; Cheke, R.S.; Shinde, S.D. Recognition of Natural Products as Potential Inhibitors of COVID-19 Main Protease (Mpro): In-Silico Evidences. Nat. Prod. Bioprospect. 2020, 10, 297–306. [Google Scholar] [CrossRef]
- Ma, C.; Sacco, M.D.; Hurst, B.; Townsend, J.A.; Hu, Y.; Szeto, T.; Zhang, X.; Tarbet, B.; Marty, M.T.; Chen, Y.; et al. Boceprevir, GC-376, and Calpain Inhibitors II, XII Inhibit SARS-CoV-2 Viral Replication by Targeting the Viral Main Protease. Cell Res. 2020, 30, 678–692. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Kumar, V.; Lee, K.W. A Computational Drug Repurposing Approach in Identifying the Cephalosporin Antibiotic and Anti-Hepatitis C Drug Derivatives for COVID-19 Treatment. Comput. Biol. Med. 2021, 130. [Google Scholar] [CrossRef]
- Hattori, S.I.; Higashi-Kuwata, N.; Hayashi, H.; Allu, S.R.; Raghavaiah, J.; Bulut, H.; Das, D.; Anson, B.J.; Lendy, E.K.; Takamatsu, Y.; et al. A Small Molecule Compound with an Indole Moiety Inhibits the Main Protease of SARS-CoV-2 and Blocks Virus Replication. Nat. Commun. 2021, 12, 1–12. [Google Scholar] [CrossRef]
- Jiménez-Alberto, A.; Ribas-Aparicio, R.M.; Aparicio-Ozores, G.; Castelán-Vega, J.A. Virtual Screening of Approved Drugs as Potential SARS-CoV-2 Main Protease Inhibitors. Comput. Biol. Chem. 2020, 88, 107325. [Google Scholar] [CrossRef]
- Ton, A.; Gentile, F.; Hsing, M.; Ban, F.; Cherkasov, A. Rapid Identification of Potential Inhibitors of SARS-CoV-2 Main Protease by Deep Docking of 1.3 Billion Compounds. Mol. Inf. 2020, 39, 2000028. [Google Scholar] [CrossRef] [Green Version]
- Hu, W.-P.; Kumar, J.V.; Huang, C.-J.; Chen, W.-Y. Computational Selection of RNA Aptamer against Angiopoietin-2 and Experimental Evaluation. BioMed Res. Int. 2015, 2015, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Lao, Y.H.; Chiang, H.Y.; Yang, D.K.; Peck, K.; Chen, L.C. Selection of Aptamers Targeting the Sialic Acid Receptor of Hemagglutinin by Epitope-Specific SELEX. Chem. Commun. 2014, 50, 8719–8722. [Google Scholar] [CrossRef]
- Baig, I.A.; Moon, J.Y.; Lee, S.C.; Ryoo, S.W.; Yoon, M.Y. Development of SsDNA Aptamers as Potent Inhibitors of Mycobacterium Tuberculosis Acetohydroxyacid Synthase. Biochim. Biophys. Acta Proteins Proteom. 2015, 1854, 1338–1350. [Google Scholar] [CrossRef]
- Oliviero, G.; Stornaiuolo, M.; D’Atri, V.; Nici, F.; Yousif, A.M.; D’Errico, S.; Piccialli, G.; Mayol, L.; Novellino, E.; Marinelli, L.; et al. Screening Platform toward New Anti-HIV Aptamers Set on Molecular Docking and Fluorescence Quenching Techniques. Anal. Chem. 2016, 88, 2327–2334. [Google Scholar] [CrossRef]
- Tseng, C.-Y.; Ashrafuzzaman, M.; Mane, J.Y.; Kapty, J.; Mercer, J.R.; Tuszynski, J.A. Entropic Fragment-Based Approach to Aptamer Design. Chem. Biol. Drug Des. 2011, 78, 1–13. [Google Scholar] [CrossRef]
- Chushak, Y.; Stone, M.O. In Silico Selection of RNA Aptamers. Nucleic Acids Res. 2009, 37, 87. [Google Scholar] [CrossRef] [Green Version]
- Rockey, W.M.; Hernandez, F.J.; Huang, S.Y.; Cao, S.; Howell, C.A.; Thomas, G.S.; Liu, X.Y.; Lapteva, N.; Spencer, D.M.; McNamara, J.O.; et al. Rational Truncation of an RNA Aptamer to Prostate-Specific Membrane Antigen Using Computational Structural Modeling. Nucleic Acid Ther. 2011, 21, 299–314. [Google Scholar] [CrossRef] [Green Version]
- Rabal, O.; Pastor, F.; Villanueva, H.; Soldevilla, M.M.; Hervas-Stubbs, S.; Oyarzabal, J. In Silico Aptamer Docking Studies: From a Retrospective Validation to a Prospective Case Study’TIM3 Aptamers Binding. Mol. Nucleic Acids 2016, 5, e376. [Google Scholar] [CrossRef] [PubMed]
- Vorobyeva, M.; Davydova, A.; Vorobjev, P.; Pyshnyi, D.; Venyaminova, A. Key Aspects of Nucleic Acid Library Design for in Vitro Selection. Int. J. Mol. Sci. 2018, 19, 470. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, Y.; Song, J.; Wei, X.; Huang, M.; Sun, M.; Zhu, L.; Lin, B.; Shen, H.; Zhu, Z.; Yang, C. Discovery of Aptamers Targeting the Receptor-Binding Domain of the SARS-CoV-2 Spike Glycoprotein. Anal. Chem. 2020, 92, 9895–9900. [Google Scholar] [CrossRef] [PubMed]
- Bacha, U.; Barrila, J.; Velazquez-Campoy, A.; Leavitt, S.A.; Freire, E. Identification of Novel Inhibitors of the SARS Coronavirus Main Protease 3CLpro. Biochemistry 2004, 43, 4906–4912. [Google Scholar] [CrossRef]
- Chou, C.Y.; Chang, H.C.; Hsu, W.C.; Lin, T.Z.; Lin, C.H.; Chang, G.G. Quaternary Structure of the Severe Acute Respiratory Syndrome (SARS) Coronavirus Main Protease. Biochemistry 2004, 43, 14958–14970. [Google Scholar] [CrossRef]
- RCSB PDB: Homepage. Available online: https://www.rcsb.org/ (accessed on 18 March 2021).
- GitHub—Igemathens 2019/MPDR. Available online: https://github.com/igemathens2019/MPDR (accessed on 18 March 2021).
- RNApdbee. Available online: http://rnapdbee.cs.put.poznan.pl/ (accessed on 18 March 2021).
- Zok, T.; Antczak, M.; Zurkowski, M.; Popenda, M.; Blazewicz, J.; Adamiak, R.W.; Szachniuk, M. RNApdbee 2.0: Multifunctional Tool for RNA Structure Annotation. Nucleic Acids Res. 2018, 46, W30–W35. [Google Scholar] [CrossRef] [Green Version]
- RNAinverse Web Server. Available online: http://rna.tbi.univie.ac.at/cgi-bin/RNAWebSuite/RNAinverse.cgi (accessed on 18 March 2021).
- Gruber, A.R.; Bernhart, S.H.; Lorenz, R. The ViennaRNA Web Services. Methods Mol. Biol. 2015, 1269, 307–326. [Google Scholar] [CrossRef] [PubMed]
- Boniecki, M.J.; Lach, G.; Dawson, W.K.; Tomala, K.; Lukasz, P.; Soltysinski, T.; Rother, K.M.; Bujnicki, J.M. SimRNA: A Coarse-Grained Method for RNA Folding Simulations and 3D Structure Prediction. Nucleic Acids Res. 2015, 44. [Google Scholar] [CrossRef] [PubMed]
- Stasiewicz, J.; Mukherjee, S.; Nithin, C.; Bujnicki, J.M. QRNAS: Software Tool for Refinement of Nucleic Acid Structures. BMC Struct. Biol. 2019, 19, 1–11. [Google Scholar] [CrossRef]
- Iwakiri, J.; Hamada, M.; Asai, K.; Kameda, T. Improved Accuracy in RNA-Protein Rigid Body Docking by Incorporating Force Field for Molecular Dynamics Simulation into the Scoring Function. J. Chem. Theory Comput. 2016, 12, 4688–4697. [Google Scholar] [CrossRef]
- Yan, Y.; Zhang, D.; Zhou, P.; Li, B.; Huang, S.Y. HDOCK: A Web Server for Protein-Protein and Protein-DNA/RNA Docking Based on a Hybrid Strategy. Nucleic Acids Res. 2017, 45, W365–W373. [Google Scholar] [CrossRef]
- Alhossary, A.; Handoko, S.D.; Mu, Y.; Kwoh, C.K. Fast, Accurate, and Reliable Molecular Docking with QuickVina 2. Bioinformatics 2015, 31, 2214–2216. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hassan, N.M.; Alhossary, A.A.; Mu, Y.; Kwoh, C.K. Protein-Ligand Blind Docking Using QuickVina-W with Inter-Process Spatio-Temporal Integration. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Pierce, B.G.; Wiehe, K.; Hwang, H.; Kim, B.H.; Vreven, T.; Weng, Z. ZDOCK Server: Interactive Docking Prediction of Protein-Protein Complexes and Symmetric Multimers. Bioinformatics 2014, 30, 1771–1773. [Google Scholar] [CrossRef]
- HDOCK Server. Available online: http://hdock.phys.hust.edu.cn/ (accessed on 18 March 2021).
- CHARMM-GUI. Available online: http://www.charmm-gui.org/ (accessed on 18 March 2021).
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of Simple Potential Functions for Simulating Liquid Water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- The Amber Molecular Dynamics Package. Available online: https://ambermd.org/ (accessed on 18 March 2021).
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. Ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from Ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zgarbová, M.; Otyepka, M.; Šponer, J.; Mládek, A.; Banáš, P.; Cheatham, T.E.; Jurečka, P. Refinement of the Cornell et Al. Nucleic Acids Force Field Based on Reference Quantum Chemical Calculations of Glycosidic Torsion Profiles. J. Chem. Theory Comput. 2011, 7, 2886–2902. [Google Scholar] [CrossRef] [PubMed]
- Krepl, M.; Havrila, M.; Stadlbauer, P.; Banas, P.; Otyepka, M.; Pasulka, J.; Stefl, R.; Sponer, J. Can We Execute Stable Microsecond-Scale Atomistic Simulations of Protein-RNA Complexes? J. Chem. Theory Comput. 2015, 11, 1220–1243. [Google Scholar] [CrossRef] [PubMed]
- Darden, T.; York, D.; Pedersen, L. Particle Mesh Ewald: An N·log(N) Method for Ewald Sums in Large Systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef] [Green Version]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A Smooth Particle Mesh Ewald Method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef] [Green Version]
- Case, D.A.; Aktulga, H.M.; Belfon, K.; Ben-Shalom, I.Y.; Brozell, S.R.; Cerutti, D.S.; Cheatham, T.E.; Cruzeiro, V.W.D.; Darden, T.A.; Duke, R.E.; et al. Amber 2020, University of California, San Francisco. Available online: https://ambermd.org (accessed on 18 March 2021).
- Grant, B.J.; Skjærven, L.; Yao, X.-Q. The Bio3D Packages for Structural Bioinformatics. Protein Sci. 2021, 30, 20–30. [Google Scholar] [CrossRef] [PubMed]
- R: A Language and Environment for Statistical Computing. Available online: https://www.gbif.org/tool/81287/r-a-language-and-environment-for-statistical-computing (accessed on 18 March 2021).
- R: The R Project for Statistical Computing. Available online: https://www.r-project.org/ (accessed on 18 March 2021).
- Miller, B.R.; McGee, T.D.; Swails, J.M.; Homeyer, N.; Gohlke, H.; Roitberg, A.E. MMPBSA.Py: An Efficient Program for End-State Free Energy Calculations. J. Chem. Theory Comput. 2012, 8, 3314–3321. [Google Scholar] [CrossRef]
- Xu, L.; Sun, H.; Li, Y.; Wang, J.; Hou, T. Assessing the Performance of MM/PBSA and MM/GBSA Methods. 3. The Impact of Force Fields and Ligand Charge Models. J. Phys. Chem. B 2013, 117, 8408–8421. [Google Scholar] [CrossRef]
- Sun, H.; Li, Y.; Tian, S.; Xu, L.; Hou, T. Assessing the Performance of MM/PBSA and MM/GBSA Methods. 4. Accuracies of MM/PBSA and MM/GBSA Methodologies Evaluated by Various Simulation Protocols Using PDBbind Data Set. Phys. Chem. Chem. Phys. 2014, 16, 16719–16729. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, D.T.; Case, D.A. On Finding Stationary States on Large-Molecule Potential Energy Surfaces. J. Phys. Chem. 1985, 89, 4020–4026. [Google Scholar] [CrossRef]











| ID | ZDOCK Score | HDOCK Score | QVINA-W Score (kcal/mol) | Docking Hbond of QVINA-W Complex |
|---|---|---|---|---|
| MAptapro | 1738.34 | −340 | −17.4 | 6 |
| MApta1pro | 1735.42 | −264 | −19.6 | 5 |
| MApta2pro | 1712.23 | −258 | −14.2 | 5 |
| MApta3pro | 1768.38 | −246 | −13.3 | 4 |
| MApta4pro | 1674.48 | −268 | −14.6 | 3 |
| MApta5pro | 1678.51 | −265 | −13.7 | 3 |
| MApta6pro | 1684.85 | −266 | −14.1 | 2 |
| MApta7pro | 1661.02 | −261 | −10.8 | 2 |
| MApta8pro | 1664.22 | −258 | −13.4 | 2 |
| MApta9pro | 1615.67 | −254 | −16.0 | 2 |
| ID | ZDOCK Score | Hdock Score | QVINA-W Score (kcal/mol) | Docking Hbond of QVINA-W Complex |
|---|---|---|---|---|
| MAptapro-IR1 | 1835.17 | −290 | −15.69 | 3 |
| MAptapro-IR2 | 1756.89 | −307 | −13.61 | 3 |
| ID_Complex | Acceptor | Donor | Occupancy (%) | Distance (Å) | Angle (°) |
|---|---|---|---|---|---|
| MAptapro | G_342@N3 | ASN_274@ND2 | 52.6 | 2.88 | 158.71 |
| C_317@O2 | ASN_277@N | 49.8 | 2.85 | 153.89 | |
| G_342@O2’ | ASN_274@ND2 | 42.5 | 2.87 | 150.81 | |
| C_346@OP1 | ARG_222@NH2 | 41.8 | 2.78 | 158.71 | |
| G_345@OP1 | ARG_222@NH2 | 38.9 | 2.78 | 158.35 | |
| G_328@OP2 | ARG_4@NH1 | 38.5 | 2.78 | 159.16 | |
| U_329@O2 | ARG_4@NH2 | 36.6 | 2.84 | 151.91 | |
| U_330@O4 | ALA_285@N | 34.6 | 2.86 | 162.63 | |
| GLN_273@O | A_339@O2’ | 33.2 | 2.71 | 159.51 | |
| C_317@O2’ | GLY_278@N | 33.0 | 2.90 | 155.15 | |
| MAptapro-RI1 | U_323@OP2 | ARG_4@NH2 | 89.9 | 2.78 | 161.16 |
| A_339@OP1 | ARG_279@NH2 | 80.9 | 2.80 | 161.66 | |
| C_322@OP1 | ARG_4@NH2 | 73.1 | 2.81 | 158.14 | |
| SER_139@O | G_318@N2 | 62.8 | 2.83 | 161.40 | |
| A_339@OP2 | ARG_279@NE | 49.3 | 2.86 | 158.23 | |
| ASN_274@O | G_329@N2 | 43.3 | 2.87 | 164.27 | |
| G_328@O6 | MET_276@N | 42.6 | 2.87 | 161.74 | |
| GLN_273@O | G_329@O2’ | 39.7 | 2.78 | 147.54 | |
| U_323@OP1 | ARG_4@NH1 | 38.6 | 2.84 | 159.93 | |
| LEU_220@O | A_340@N6 | 34.0 | 2.89 | 159.49 | |
| MAptapro-RI2 | G_335@OP1 | TYR_118@OH | 98.2 | 2.65 | 163.77 |
| A_333@OP1 | SER_121@OG | 70.2 | 2.68 | 163.72 | |
| A_316@O2’ | TYR_237@OH | 70.1 | 2.82 | 160.94 | |
| G_335@O4’ | SER_123@OG | 62.8 | 2.79 | 161.29 | |
| G_318@OP1 | MET_276@N | 47.3 | 2.85 | 151.86 | |
| C_338@OP1 | ARG_4@NH2 | 36.3 | 2.79 | 155.40 | |
| G_318@OP1 | ASN_277@N | 36.3 | 2.85 | 160.22 | |
| THR_169@O | G_320@N2 | 34.3 | 2.84 | 153.97 |
| System | ΔGgas | ΔGsolv | ΔGPB Bind | TΔS | ΔGPB Bind-TΔS |
|---|---|---|---|---|---|
| MAptapro/Mpro | 140.77 (2.15) | −215.31 (25.92) | −74.54 (0.56) | −72.62 (5.33) | −1.92 (5.89) |
| MAptapro-IR1/Mpro | 184.97 (3.11) | −281.22 (2.99) | −96.25 (0.54) | −66.32 (5.17) | −29.93 (5.71) |
| MAptapro-IR2/Mpro | 294.60 (2.12) | −369.64 (1.94) | −75.04 (0.52) | −69.65 (3.69) | −5.39 (4.21) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morena, F.; Argentati, C.; Tortorella, I.; Emiliani, C.; Martino, S. De novo ssRNA Aptamers against the SARS-CoV-2 Main Protease: In Silico Design and Molecular Dynamics Simulation. Int. J. Mol. Sci. 2021, 22, 6874. https://doi.org/10.3390/ijms22136874
Morena F, Argentati C, Tortorella I, Emiliani C, Martino S. De novo ssRNA Aptamers against the SARS-CoV-2 Main Protease: In Silico Design and Molecular Dynamics Simulation. International Journal of Molecular Sciences. 2021; 22(13):6874. https://doi.org/10.3390/ijms22136874
Chicago/Turabian StyleMorena, Francesco, Chiara Argentati, Ilaria Tortorella, Carla Emiliani, and Sabata Martino. 2021. "De novo ssRNA Aptamers against the SARS-CoV-2 Main Protease: In Silico Design and Molecular Dynamics Simulation" International Journal of Molecular Sciences 22, no. 13: 6874. https://doi.org/10.3390/ijms22136874







