Genome-Wide Identification and Analysis of the Polycomb Group Family in Medicago truncatula
Abstract
1. Introduction
2. Results
2.1. Identification of M. truncatula PcG Members
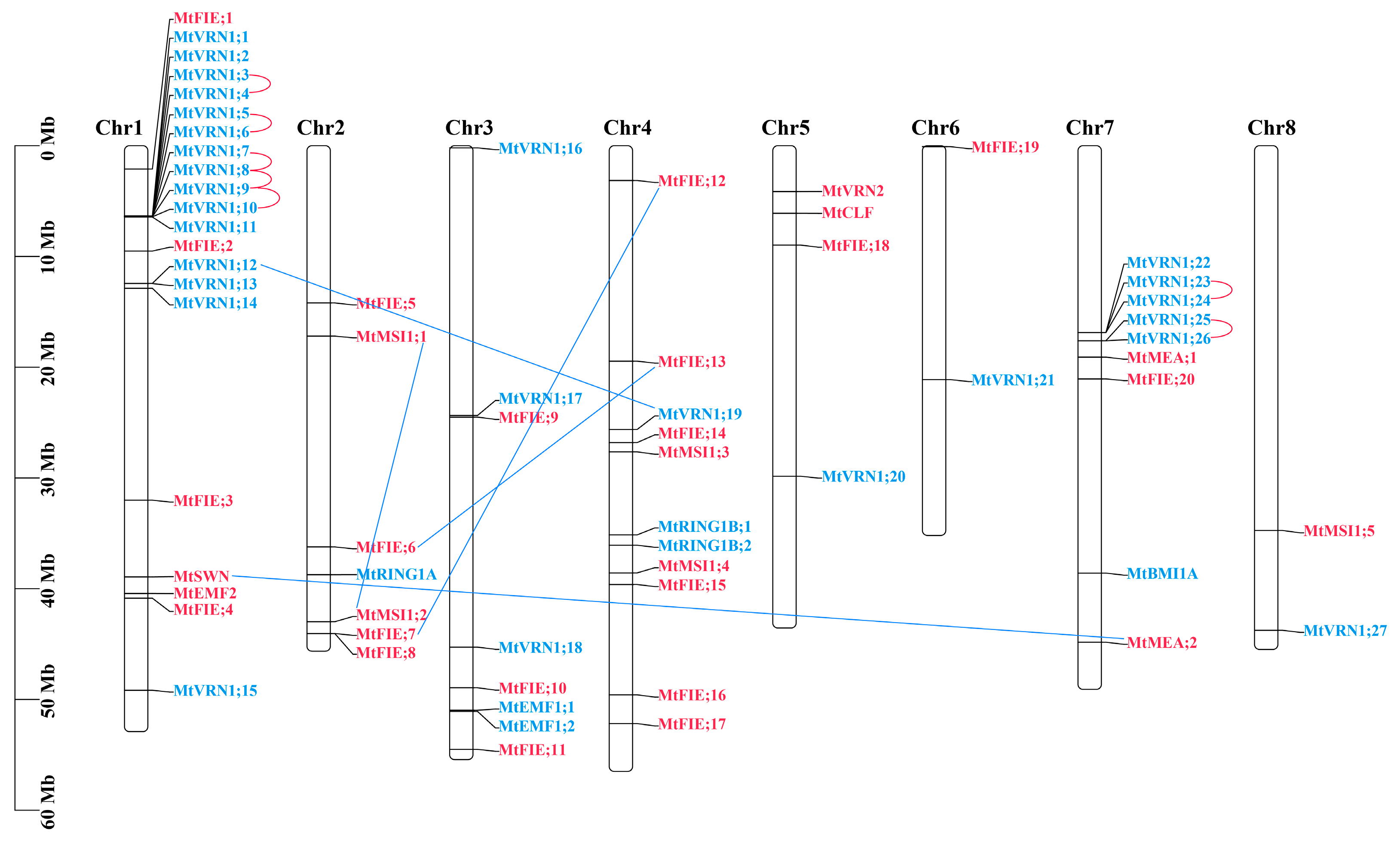
2.2. Gene Structure and Chromosome Location Analysis
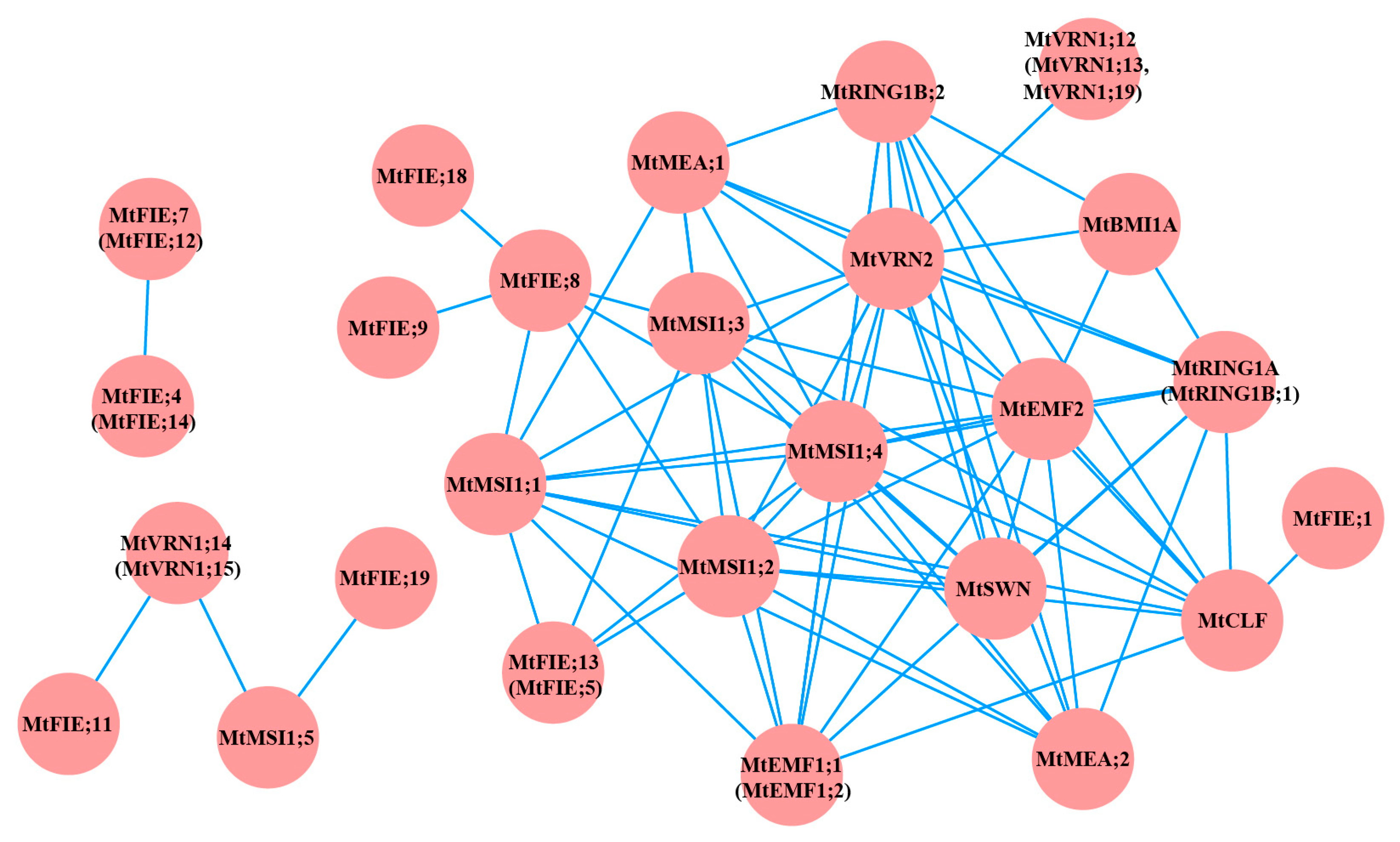
2.3. Protein–Protein Interaction Analysis
2.4. Phylogenetic Analysis and Synteny Analysis
2.5. Expression Pattern Analysis
2.5.1. Microarray Expression Data and Transcriptome Sequencing Data
2.5.2. Co-Expression and GO Enrichment Analysis of PcG Genes
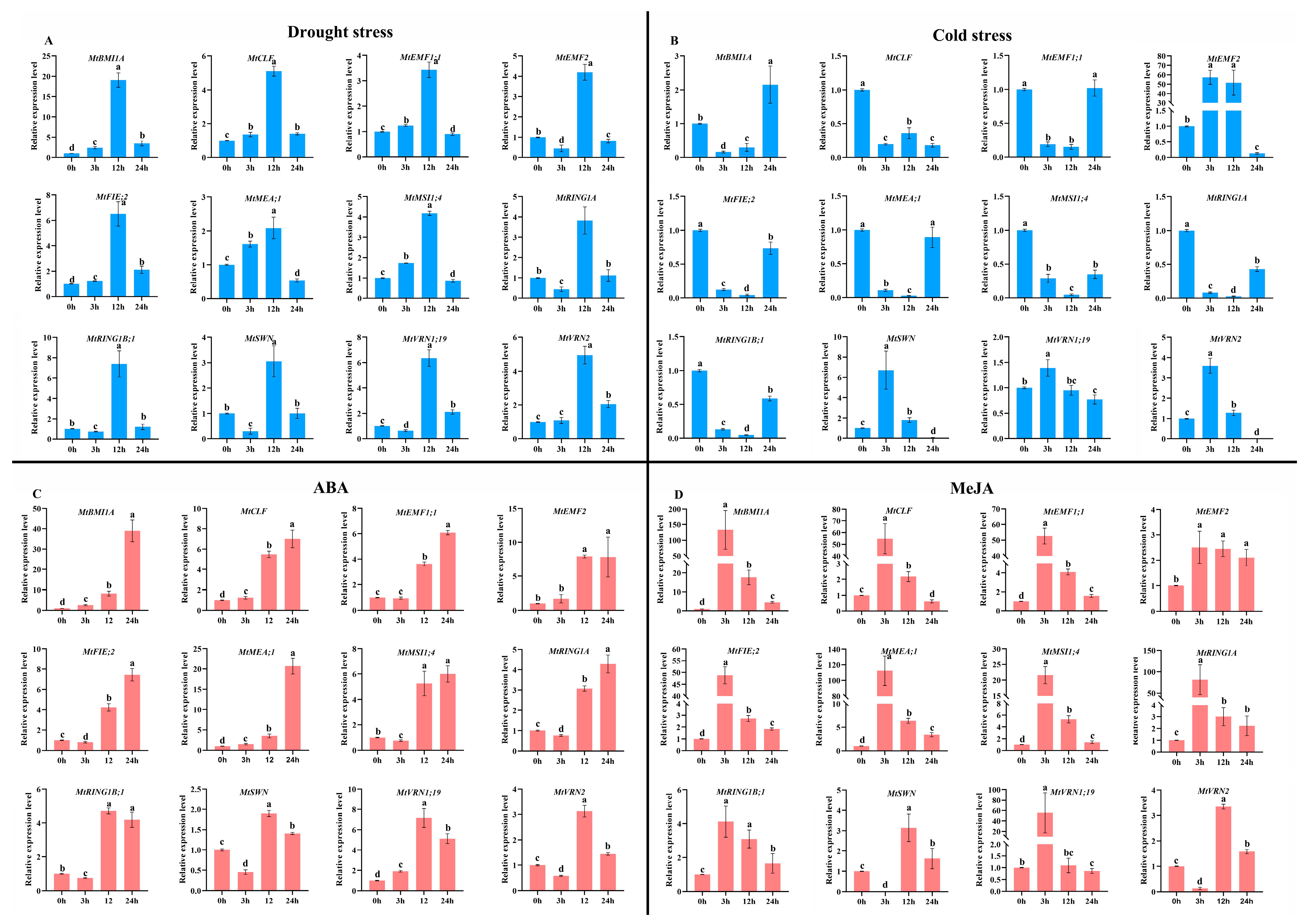
2.5.3. The Expression Patterns of PcG Genes in qRT-PCR Analysis
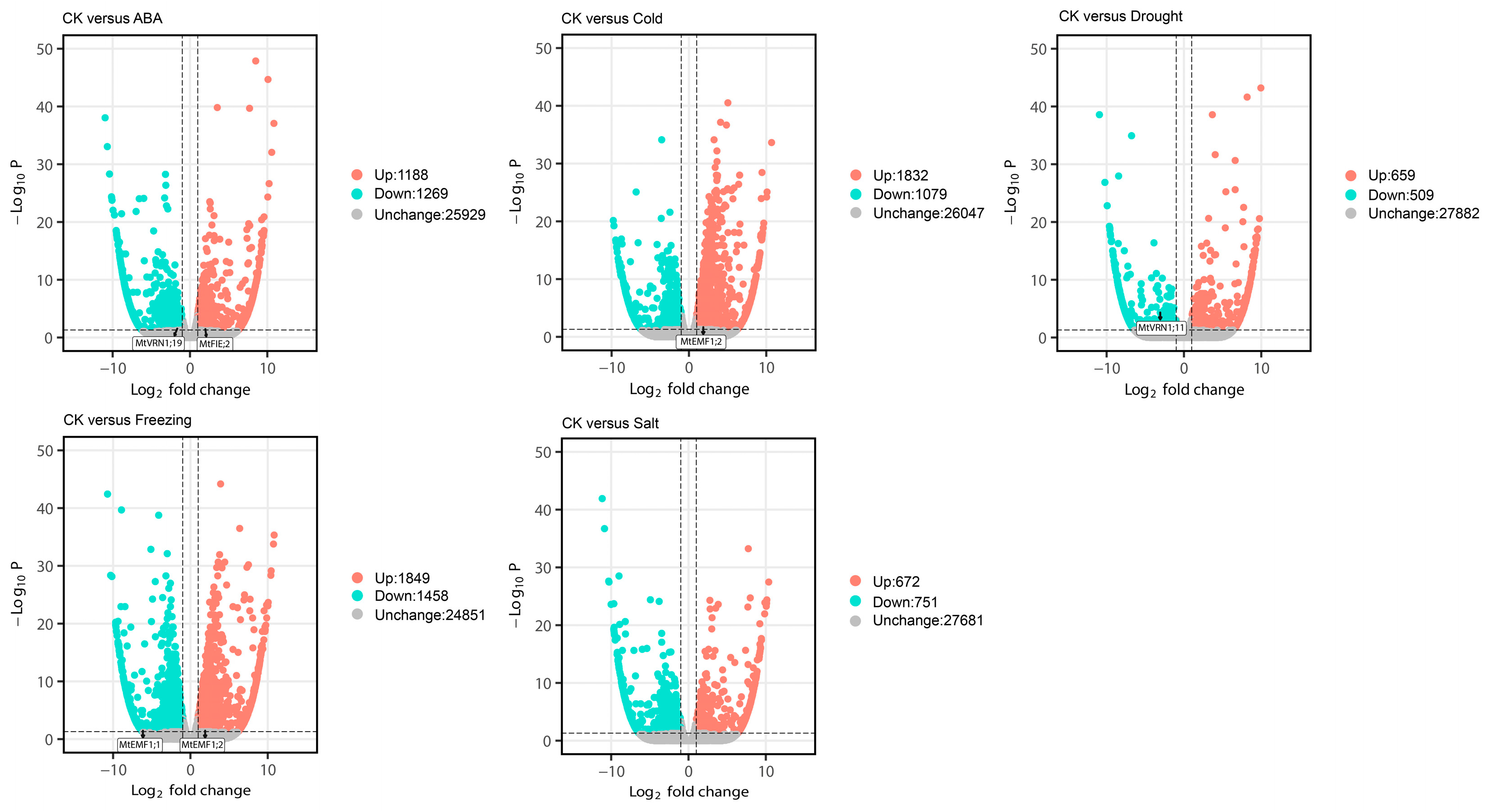
2.6. Identification of Differentially Expressed PcG Genes Based on RNA-seq Data
3. Discussion
3.1. Identification of M. truncatula PcG Members and Structure Analysis
3.2. Evolutionary Relationship Analysis of PcG Proteins in M. truncatula
3.3. Expression Pattern Analysis of PcG Genes in M. truncatula
4. Materials and Methods
4.1. Materials, Growth Conditions, Treatment, and Sampling
4.2. Download and Analysis of Genome Data
4.3. Identification of PcG Genes in M. truncatula
4.4. Basic Analysis of PcG Proteins
4.5. Sequence Analysis and Chromosome Location
4.6. Prediction of Protein Interaction Networks
4.7. Phylogenetic and Synteny Analysis
4.8. PcG Genes Expression Analysis
4.9. Co-Expression and GO Enrichment Analysis of PcG Genes
4.10. RNA Extraction and qRT-PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- De la Paz Sanchez, M.; Aceves-García, P.; Petrone, E.; Steckenborn, S.; Vega-León, R.; Álvarez-Buylla, E.R.; Garay-Arroyo, A.; García-Ponce, B. The impact oolycomb group (PcG) and Trithorax group (TrxG) epigenetic factors in plant plasticity. New Phytol. 2015, 208, 684–694. [Google Scholar] [CrossRef]
- Mozgova, I.; Hennig, L. The polycomb group protein regulatory network. Annu. Rev. Plant Biol. 2015, 66, 269–296. [Google Scholar] [CrossRef]
- Lewis, E.B. A gene complex controlling segmentation in Drosophila. Nature 1978, 276, 565–570. [Google Scholar] [CrossRef]
- Yoshida, N.; Yanai, Y.; Chen, L.; Kato, Y.; Hiratsuka, J.; Miwa, T.; Sung, Z.R.; Takahashi, S. EMBRYONIC FLOWER2, a novel polycomb group protein homolog, mediates shoot development and flowering in Arabidopsis. Plant Cell 2001, 13, 2471–2481. [Google Scholar] [CrossRef]
- Li, W.; Wang, Z.; Li, J.; Yang, H.; Cui, S.; Wang, X.; Ma, L. Overexpression of AtBMI1C, a polycomb group protein gene, accelerates flowering in Arabidopsis. PLoS ONE 2011, 6, e21364. [Google Scholar] [CrossRef]
- Zhong, J.; Peng, Z.; Peng, Q.; Cai, Q.; Peng, W.; Chen, M.; Yao, J. Regulation of plant height in ricy tholycomb group genes OsEMF2b, OsFIE2 and OsCLF. Plant Sci. 2018, 267, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Wagner, D. Polycomb repression in the regulation of growth and development in Arabidopsis. Curr. Opin. Plant Biol. 2015, 23, 15–24. [Google Scholar] [CrossRef]
- Yan, B.; Lv, Y.; Zhao, C.; Wang, X. Knowing when to silence: Roles of polycomb-group proteins in SAM maintenance, root development, and developmental phase transition. Int. J. Mol. Sci. 2020, 21, 5871. [Google Scholar] [CrossRef] [PubMed]
- Alexandre, C.; Möller-Steinbach, Y.; Schönrock, N.; Gruissem, W.; Hennig, L. Arabidopsis MSI1 is required for negative regulation of the response trought stress. Mol. Plant 2009, 2, 675–687. [Google Scholar] [CrossRef] [PubMed]
- Kleinmanns, J.A.; Schubert, D. Polycomb and Trithorax group protein-mediated control of stress responses in plants. Biol. Chem. 2014, 395, 1291–1300. [Google Scholar] [CrossRef]
- Liu, C.; Cheng, J.; Zhuang, Y.; Ye, L.; Li, Z.; Wang, Y.; Qi, M.; Xu, L.; Zhang, Y. Polycomb repressive complex 2 attenuates ABA-induced senescence in Arabidopsis. Plant J. 2019, 97, 368–377. [Google Scholar] [CrossRef]
- Ringrose, L. Polycomb comes of age: Genome-wide profiling of target sites. Curr. Opin. Cell Biol. 2007, 19, 290–297. [Google Scholar] [CrossRef]
- Zhang, X.; Clarenz, O.; Cokus, S.; Bernatavichute, Y.V.; Pellegrini, M.; Goodrich, J.; Jacobsen, S.E. Whole-genome analysis of histone H3 lysine 27 trimethylation in Arabidopsis. PLoS Biol. 2007, 5, e129. [Google Scholar] [CrossRef]
- Margueron, R.; Reinberg, D. Tholycomb complex PRC2 and its mark in life. Nature 2011, 469, 343–349. [Google Scholar] [CrossRef]
- Jiao, H.; Xie, Y.; Li, Z. Current understanding of plant Polycomb group proteins and the repressive histone H3 Lysine 27 trimethylation. Biochem. Soc. Trans. 2020, 48, 1697–1706. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Molitor, A.; Liu, C.; Shen, W.-H. The Arabidopsis PRC1-like ring-finger proteins are necessary for repression ombryonic traituring vegetative growth. Cell Res. 2010, 20, 1332–1344. [Google Scholar] [CrossRef] [PubMed]
- Calonje, M. PRC1 marks thifference in plant PcG repression. Mol. Plant 2014, 7, 459–471. [Google Scholar] [CrossRef]
- Mozgova, I.; Köhler, C.; Hennig, L. Keeping the gate closed: Functions of the polycomb repressive complex PRC2 in development. Plant J. 2015, 83, 121–132. [Google Scholar] [CrossRef]
- Derkacheva, M.; Hennig, L. Variations on a theme: Polycomb group proteins in plants. J. Exp. Bot. 2014, 65, 2769–2784. [Google Scholar] [CrossRef]
- Berke, L.; Snel, B. The plant Polycomb repressive complex 1 (PRC1) existed in the ancestor of seed plants and has a complex duplication history. BMC Evol. Biol. 2015, 15, 44. [Google Scholar] [CrossRef]
- Molitor, A.; Shen, W.-H. The polycomb complex PRC1: Composition and function in plants. J. Genet. Genom. 2013, 40, 231–238. [Google Scholar] [CrossRef]
- Latrasse, D.; Germann, S.; Houba-Hérin, N.; Dubois, E.; Bui-Prodhomme, D.; Hourcade, D.; Juul-Jensen, T.; Le Roux, C.; Majira, A.; Simoncello, N. Control of flowering and cell faty LIF2, an RNA binding partner of the polycomb complex component LHP1. PLoS ONE 2011, 6, e16592. [Google Scholar] [CrossRef]
- Calonje, M.; Sanchez, R.; Chen, L.; Sung, Z.R. EMBRYONIC FLOWER1 participates in polycomb group–mediated AG gene silencing in Arabidopsis. Plant Cell 2008, 20, 277–291. [Google Scholar] [CrossRef]
- Levy, Y.Y.; Mesnage, S.; Mylne, J.S.; Gendall, A.R.; Dean, C. Multiple roles of Arabidopsis VRN1 in vernalization and flowering time control. Science 2002, 297, 243–246. [Google Scholar] [CrossRef] [PubMed]
- Pu, L.; Sung, Z.R. PcG and trxG in plants–friends or foes. Trends Genet. 2015, 31, 252–262. [Google Scholar] [CrossRef] [PubMed]
- Mayama, T.; Ohtsubo, E.; Tsuchimoto, S. Isolation and expression analysis of petunia CURLY LEAF-like genes. Plant Cell Physiol. 2003, 44, 811–819. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chanvivattana, Y.; Bishopp, A.; Schubert, D.; Stock, C.; Moon, Y.-H.; Sung, Z.R.; Goodrich, J. Interaction oolycomb-group proteins controlling flowering in Arabidopsis. Development 2004, 131, 5263–5276. [Google Scholar] [CrossRef]
- Grossniklaus, U.; Vielle-Calzada, J.-P.; Hoeppner, M.A.; Gagliano, W.B. Maternal control ombryogenesiy MEDEA, a polycomb group gene in Arabidopsis. Science 1998, 280, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Goodrich, J.; Puangsomlee, P.; Martin, M.; Long, D.; Meyerowitz, E.M.; Coupland, G. A Polycomb-group gene regulates homeotic genxpression in Arabidopsis. Nature 1997, 386, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Schubert, D.; Primavesi, L.; Bishopp, A.; Roberts, G.; Doonan, J.; Jenuwein, T.; Goodrich, J. Silencing by plant Polycomb-group genes requireispersed trimethylation of histone H3 at lysine 27. EMBO J. 2006, 25, 4638–4649. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Kim, Y.J.; Müller, R.; Yumul, R.E.; Liu, C.; Pan, Y.; Cao, X.; Goodrich, J.; Chen, X. AGAMOUS terminates floral stem cell maintenance in Arabidopsis by directly repressing WUSCHEL through recruitment oolycomb Group proteins. Plant Cell 2011, 23, 3654–3670. [Google Scholar] [CrossRef]
- Aichinger, E.; Villar, C.B.; Di Mambro, R.; Sabatini, S.; Köhler, C. The CHD3 chromatin remodeleICKLE and polycomb group proteins antagonistically regulate meristem activity in the Arabidopsis root. Plant Cell 2011, 23, 1047–1060. [Google Scholar] [CrossRef]
- Li, C.; Chen, C.; Gao, L.; Yang, S.; Nguyen, V.; Shi, X.; Siminovitch, K.; Kohalmi, S.E.; Huang, S.; Wu, K. The Arabidopsis SWI2/SNF2 chromatin remodeler BRAHMA regulates polycomb function during vegetativevelopment and directly activates the flowering repressor gene SVP. PLoS Genet. 2015, 11, e1004944. [Google Scholar] [CrossRef] [PubMed]
- Shu, J.; Chen, C.; Thapa, R.K.; Bian, S.; Nguyen, V.; Yu, K.; Yuan, Z.C.; Liu, J.; Kohalmi, S.E.; Li, C. Genome-wide occupancy of histone H3K27 methyltransferases CURLY LEAF and SWINGER in Arabidopsis seedlings. Plant Direct. 2019, 3, e00100. [Google Scholar] [CrossRef] [PubMed]
- Spillane, C.; Schmid, K.J.; Laoueillé-Duprat, S.; Pien, S.; Escobar-Restrepo, J.-M.; Baroux, C.; Gagliardini, V.; Page, D.R.; Wolfe, K.H.; Grossniklaus, U. Positivarwinian selection at the imprinted MEDEA locus in plants. Nature 2007, 448, 349–352. [Google Scholar] [CrossRef]
- Gendall, A.R.; Levy, Y.Y.; Wilson, A.; Dean, C. The VERNALIZATION 2 gene mediates thpigenetic regulation of vernalization in Arabidopsis. Cell 2001, 107, 525–535. [Google Scholar] [CrossRef]
- Luo, M.; Bilodeau, P.; Koltunow, A.; Dennis, E.S.; Peacock, W.J.; Chaudhury, A.M. Genes controlling fertilization-independent seed development in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 1999, 96, 296–301. [Google Scholar] [CrossRef]
- Liu, M.-S.; Chen, L.-F.O.; Lin, C.-H.; Lai, Y.-M.; Huang, J.-Y.; Sung, Z.R. Molecular and functional characterization oroccoli EMBRYONIC FLOWER 2 genes. Plant Cell Physiol. 2012, 53, 1217–1231. [Google Scholar] [CrossRef]
- Jaudal, M.; Zhang, L.; Che, C.; Hurley, D.G.; Thomson, G.; Wen, J.; Mysore, K.S.; Putterill, J. Mt VRN 2 is a Polycomb VRN 2-like gene which represses the transition to flowering in the model legum edicago truncatula. Plant J. 2016, 86, 145–160. [Google Scholar] [CrossRef] [PubMed]
- Diallo, A.; Kane, N.; Agharbaoui, Z.; Badawi, M.; Sarhan, F. Heterologouxpression of wheat VERNALIZATION 2 (TaVRN2) gene in Arabidopsis delays flowering and enhances freezing tolerance. PLoS ONE 2010, 5, e8690. [Google Scholar] [CrossRef]
- Karsai, I.; Szűcs, P.; Mészáros, K.; Filichkina, T.; Hayes, P.; Skinner, J.; Láng, L.; Bedő, Z. The Vrn-H2 locus is a majoeterminant of flowering time in a facultative× winter growth habit barley (Hordeum vulgare L.) mapping population. Theor. Appl. Genet. 2005, 110, 1458–1466. [Google Scholar] [CrossRef] [PubMed]
- Köhler, C.; Hennig, L.; Bouveret, R.; Gheyselinck, J.; Grossniklaus, U.; Gruissem, W. Arabidopsi SI1 is a component of thEA/FIE Polycomb group complex and required for seed development. EMBO J. 2003, 22, 4804–4814. [Google Scholar] [CrossRef] [PubMed]
- Makarevich, G.; Leroy, O.; Akinci, U.; Schubert, D.; Clarenz, O.; Goodrich, J.; Grossniklaus, U.; Köhler, C. Different Polycomb group complexes regulate common target genes in Arabidopsis. EMBO Rep. 2006, 7, 947–952. [Google Scholar] [CrossRef]
- Reyes, J.C.; Grossniklaus, U. Diverse functions oolycomb group proteinuring plant development. Semin. Cell Dev. Biol. 2003, 14, 77–84. [Google Scholar] [CrossRef]
- Ohad, N.; Yadegari, R.; Margossian, L.; Hannon, M.; Michaeli, D.; Harada, J.J.; Goldberg, R.B.; Fischer, R.L. Mutations in FIE, a WD polycomb group gene, allow endosperm development without fertilization. Plant Cell 1999, 11, 407–415. [Google Scholar] [CrossRef]
- Ach, R.A.; Taranto, P.; Gruissem, W. A conserved family of WD-40 proteininds to the retinoblastoma protein in both plants and animals. Plant Cell 1997, 9, 1595–1606. [Google Scholar] [PubMed]
- Hu, Y.; Gao, Y.; Xu, H.; Sun, J.; Li, Y.; Yan, Y. Characterization of the fertilization independent endosperm (FIE) gene from soybean. Afr. J. Biotechnol. 2012, 11, 7795–7802. [Google Scholar]
- Bouveret, R.; Schönrock, N.; Gruissem, W.; Hennig, L. Regulation of flowering timy Arabidopsis SI1. Development 2006, 133, 1693–1702. [Google Scholar] [CrossRef][Green Version]
- Hennig, L.; Derkacheva, M. Diversity oolycomb group complexes in plants: Same rules, different players? Trends Genet. 2009, 25, 414–423. [Google Scholar] [CrossRef]
- Bemer, M.; Grossniklaus, U. Dynamic regulation oolycomb group activity during plant development. Curr. Opin. Plant Biol. 2012, 15, 523–529. [Google Scholar] [CrossRef]
- Chandler, J.; Wilson, A.; Dean, C. Arabidopsis mutants showing an altered response to vernalization. Plant J. 1996, 10, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Wood, C.C.; Robertson, M.; Tanner, G.; Peacock, W.J.; Dennis, E.S.; Helliwell, C.A. The Arabidopsis thaliana vernalization response requires a polycomb-like protein complex that also includes VERNALIZATION INSENSITIVE 3. Proc. Natl. Acad. Sci. USA 2006, 103, 14631–14636. [Google Scholar] [CrossRef] [PubMed]
- Lucia, F.D.; Crevillen, P.; Jones, A.M.; Greb, T.; Dean, C. A PHD-polycomb repressive complex 2 triggers thpigenetic silencing of FLuring vernalization. Proc. Natl. Acad. Sci. USA 2008, 105, 16831–16836. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Zhu, T.; Sung, Z.R. Epigenetic regulation of gene programy EMF1 and EMF2 in Arabidopsis. Plant Physiol. 2010, 152, 516–528. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.-H.; Chen, L.-J.; Sung, Z.R. Genetic regulation of shoot development in Arabidopsis: Role of the EMF genes. Dev. Biol. 1995, 169, 421–435. [Google Scholar] [CrossRef]
- Branca, A.; Paape, T.D.; Zhou, P.; Briskine, R.; Farmer, A.D.; Mudge, J.; Bharti, A.K.; Woodward, J.E.; May, G.D.; Gentzbittel, L. Whole-genome nucleotidiversity, recombination, and linkagisequilibrium in the model legum edicago truncatula. Proc. Natl. Acad. Sci. USA 2011, 108, E864–E870. [Google Scholar] [CrossRef]
- Orłowska, A.; Igielski, R.; Łagowska, K.; Kępczyńska, E. Identification of LEC1, L1L and Polycomb Repressive Complex 2 genes and theixpression during the induction phase o edicago truncatula Gaertn. somatic embryogenesis. Plant Cell Tissue Organ. Cult. 2017, 129, 119–132. [Google Scholar] [CrossRef]
- Orłowska, A.; Kępczyńska, E. Identification oolycomb Repressive Complex1, Trithorax group genes and their simultaneouxpression with WUSCHEL, WUSCHEL-related Homeobox5 and SHOOT MERISTEMLESS during the induction phase of somatic embryogenesis in Medicago truncatula Gaertn. Plant Cell Tissue Organ Cult. 2018, 134, 345–356. [Google Scholar] [CrossRef]
- Che, C. Functional analysis of candidate flowering time genes from the model legum edicago truncatula. Ph.D. thesis, The University of Auckland, Auckland, New Zealand, 2017. [Google Scholar]
- Jaudal, M.; Thomson, G.; Zhang, L.; Che, C.; Wen, J.; Mysore, K.S.; Tadege, M.; Putterill, J. Forward and reverse screens to identify genes that control vernalization and flowering time in Medicago truncatula. In The Model Legumedicago Truncatula, 3rd ed.; Frans, J.D.B., Ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2019; pp. 189–196. [Google Scholar]
- Köhler, C.; Hennig, L. Regulation of cell identity by plant Polycomb and trithorax group proteins. Curr. Opin. Genet. Dev. 2010, 20, 541–547. [Google Scholar] [CrossRef]
- Strejčková, B.; Čegan, R.; Pecinka, A.; Milec, Z.; Šafář, J. Identification of polycomb repressive complex 1 and 2 core components in hexaploid bread wheat. BMC Plant Biol. 2020, 20, 175. [Google Scholar] [CrossRef]
- Ni, J.; Ma, X.; Feng, Y.; Tian, Q.; Wang, Y.; Xu, N.; Tang, J.; Wang, G. Updating and interaction of polycomb repressive complex 2 components in maize (Zea mays). Planta 2019, 250, 573–588. [Google Scholar] [CrossRef]
- Mozgová, I.; Muñoz-Viana, R.; Hennig, L. PRC2 represses hormone-induced somatic embryogenesis in vegetative tissue of Arabidopsis thaliana. PLoS Genet. 2017, 13, e1006562. [Google Scholar] [CrossRef]
- Scott, M.S.; Calafell, S.J.; Thomas, D.Y.; Hallett, M.T. Refining protein subcellular localization. PLoS Comput. Biol. 2005, 1, e66. [Google Scholar] [CrossRef] [PubMed]
- Schatlowski, N.; Stahl, Y.; Hohenstatt, M.L.; Goodrich, J.; Schubert, D. The CURLY LEAF interacting protein BLISTER controlxpression of polycomb-group target genes and cellulaifferentiation of Arabidopsis thaliana. Plant Cell 2010, 22, 2291–2305. [Google Scholar] [CrossRef][Green Version]
- Xu, L.; Shen, W.-H. Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr. Biol. 2008, 18, 1966–1971. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Huang, Y.; Lu, J. Cloning, subcellular localization, and expression analysis of VvRING1a from grapevine. Hortic. Environ. Biotechnol. 2019, 60, 733–739. [Google Scholar] [CrossRef]
- Deng, L.; Zhang, S.; Wang, G.; Fan, S.; Li, M.; Chen, W.; Tu, B.; Tan, J.; Wang, Y.; Ma, B. Down-regulation of OsEMF2b caused semi-sterility due to anther and pollen development defects in rice. Front. Plant Sci. 2017, 8, 1998. [Google Scholar] [CrossRef] [PubMed]
- Lü, J.; Suo, H.; Yi, R.; Ma, Q.; Nian, H. Glyma11g13220, a homolog of the vernalization pathway gene VERNALIZATION 1 from soybean [Glycine max (L.) Merr.], promotes flowering in Arabidopsis thaliana. BMC Plant Biol. 2015, 15, 232. [Google Scholar] [CrossRef]
- Oliva, M.; Butenko, Y.; Hsieh, T.-F.; Hakim, O.; Katz, A.; Smorodinsky, N.I.; Michaeli, D.; Fischer, R.L.; Ohad, N. FIE, a nucleaRC2 protein, forms cytoplasmic complexes in Arabidopsis thaliana. J. Exp. Bot. 2016, 67, erw373. [Google Scholar] [CrossRef]
- Li, S.; Zhou, B.; Peng, X.; Kuang, Q.; Huang, X.; Yao, J.; Du, B.; Sun, M.X. OsFIE2 plays an essential role in the regulation of rice vegetative and reproductivevelopment. New Phytol. 2014, 201, 66–79. [Google Scholar] [CrossRef]
- Liu, D.-D.; Dong, Q.-L.; Sun, C.; Wang, Q.-L.; You, C.-X.; Yao, Y.-X.; Hao, Y.-J. Functional characterization of an apple apomixis-related MhFIE gene in reproduction development. Plant Sci. 2012, 185, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Guitton, A.-E.; Page, D.R.; Chambrier, P.; Lionnet, C.; Faure, J.-E.; Grossniklaus, U.; Berger, F. Identification of new members of Fertilisation Independent Seed Polycomb Group pathway involved in the control of seed development in Arabidopsis thaliana. Development 2004, 131, 2971–2981. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.-J.; Diao, Z.-Y.; Specht, C.; Sung, Z.R. Moleculavolution of VEF-domain-containing PcG genes in plants. Mol. Plant 2009, 2, 738–754. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Aubert, D.; Chen, L.; Moon, Y.-H.; Martin, D.; Castle, L.A.; Yang, C.-H.; Sung, Z.R. EMF1, a novel protein involved in the control of shoot architecture and flowering in Arabidopsis. Plant Cell 2001, 13, 1865–1875. [Google Scholar] [CrossRef] [PubMed]
- Kapazoglou, A.; Tondelli, A.; Papaefthimiou, D.; Ampatzidou, H.; Francia, E.; Stanca, M.A.; Bladenopoulos, K.; Tsaftaris, A.S. Epigenetic chromatin modifiers in barley: IV. The study oarley polycomb group (PcG) geneuring seed development and in response txternal ABA. BMC Plant Biol. 2010, 10, 73. [Google Scholar] [CrossRef]
- Luo, M.; Platten, D.; Chaudhury, A.; Peacock, W.; Dennis, E.S. Expression, imprinting, and evolution of rice homologs of the polycomb group genes. Mol. Plant 2009, 2, 711–723. [Google Scholar] [CrossRef]
- Ling, L.; Zhang, W.; An, Y.; Du, B.; Wang, D.; Guo, C. Genome-wide analysis of the TCP transcription factor genes in five legume genomes and their response to salt and drought stresses. Funct. Integr. Genom. 2020, 4, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Hu, R.; Gu, T.; Han, J.; Qiu, D.; Su, P.; Feng, J.; Chang, J.; Yang, G.; He, G. Genome-wide identification and expression profiling of trihelix gene family under abiotic stresses in wheat. BMC Genom. 2019, 20, 287. [Google Scholar] [CrossRef] [PubMed]
- Bancroft, I. Duplicate and diverge: Thvolution of plant genome microstructure. Trends Genet. 2001, 17, 89–93. [Google Scholar] [CrossRef]
- Kayum, M.A.; Park, J.-I.; Nath, U.K.; Saha, G.; Biswas, M.K.; Kim, H.-T.; Nou, I.-S. Genome-wide characterization and expression profiling oDI family gene reveals function as abiotic and biotic stress tolerance in Chinese cabbage (Brassica rapa ssp. pekinensis). BMC Genom. 2017, 18, 885. [Google Scholar] [CrossRef]
- Marques-Bonet, T.; Ryder, O.A.; Eichler, E.E. Sequencing primate genomes: What have we learned? Annu. Rev. Genom. Hum. Genet. 2009, 10, 355–386. [Google Scholar] [CrossRef]
- Huang, X.; Lu, Z.; Wang, X.; Ouyang, Y.; Chen, W.; Xie, K.; Wang, D.; Luo, M.; Luo, J.; Yao, J. Imprinted gene OsFIE1 modulates rice seed development by influencing nutrient metabolism and modifying genome H3K27me3. Plant J. 2016, 87, 305–317. [Google Scholar] [CrossRef]
- Yan, L.; Loukoianov, A.; Blechl, A.; Tranquilli, G.; Ramakrishna, W.; SanMiguel, P.; Bennetzen, J.L.; Echenique, V.; Dubcovsky, J. The wheat VRN2 gene is a flowering repressoown-regulated by vernalization. Science 2004, 303, 1640–1644. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.; Yan, L. Duplicated, deleted and translocated VRN2 genes in hexaploid wheat. Euphytica 2016, 208, 277–284. [Google Scholar] [CrossRef]
- Sharma, V.; RANJAN, T.; KUMAR, P.; PAL, A.K.; JHA, V.K.; SAHNI, S.; PRASAD, B.D. Protein–protein interaction detection: Methods and analysis. Plant Biotechnol. 2017, 1, 391–411. [Google Scholar]
- Bratzel, F.; López-Torrejón, G.; Koch, M.; Del Pozo, J.C.; Calonje, M. Keeping cell identity in Arabidopsis requireRC1 RING-finger homologs that catalyze H2A monoubiquitination. Curr. Biol. 2010, 20, 1853–1859. [Google Scholar] [CrossRef]
- Luo, M.; Bilodeau, P.; Dennis, E.S.; Peacock, W.J.; Chaudhury, A. Expression and parent-of-origin effects for FIS2, MEA, and FIE in thndosperm and embryo oeveloping Arabidopsis seeds. Proc. Natl. Acad. Sci. USA 2000, 97, 10637–10642. [Google Scholar] [CrossRef] [PubMed]
- Spillane, C.; MacDougall, C.; Stock, C.; Köhler, C.; Vielle-Calzada, J.; Nunes, S.M.; Grossniklaus, U.; Goodrich, J. Interaction of the Arabidopsis polycomb group proteins FIE and MEA mediates their common phenotypes. Curr. Biol. 2000, 10, 1535–1538. [Google Scholar] [CrossRef]
- Yadegari, R.; Kinoshita, T.; Lotan, O.; Cohen, G.; Katz, A.; Choi, Y.; Katz, A.; Nakashima, K.; Harada, J.J.; Goldberg, R.B. Mutations in the FIE and MEA genes that encode interacting polycomb proteins cause parent-of-origin effects on seed development by distinct mechanisms. Plant Cell 2000, 12, 2367–2381. [Google Scholar] [CrossRef] [PubMed]
- Katz, A.; Oliva, M.; Mosquna, A.; Hakim, O.; Ohad, N. FIE and CURLY LEAF polycomb proteins interact in the regulation of homeobox genxpression during sporophytevelopment. Plant J. 2004, 37, 707–719. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Tyson, M.D.; Jackson, S.S.; Yadegari, R. Partially redundant functions of two SET-domain polycomb-group proteins in controlling initiation of seed development in Arabidopsis. Proc. Natl. Acad. Sci. USA 2006, 103, 13244–13249. [Google Scholar] [CrossRef]
- Derkacheva, M.; Steinbach, Y.; Wildhaber, T.; Mozgova, I.; Mahrez, W.; Nanni, P.; Bischof, S.; Gruissem, W.; Hennig, L. Arabidopsis SI1 connects LHP1 tRC2 complexes. EMBO J. 2013, 32, 2073–2085. [Google Scholar] [CrossRef]
- Shen, L.; Thong, Z.; Gong, X.; Shen, Q.; Gan, Y.; Yu, H. The putativRC1 RING-finger protein AtRING1A regulates flowering through repressing MADS AFFECTING FLOWERING genes in Arabidopsis. Development 2014, 141, 1303–1312. [Google Scholar] [CrossRef]
- Förderer, A.; Zhou, Y.; Turck, F. The age of multiplexity: Recruitment and interactions oolycomb complexes in plants. Curr. Opin. Plant Biol. 2016, 29, 169–178. [Google Scholar] [CrossRef]
- Kidd, K.K.; Sgaramella-Zonta, L.A. Phylogenetic analysis: Concepts and methods. Am. J. Hum. Genet. 1971, 23, 235. [Google Scholar] [PubMed]
- Jordan, F.M. Comparative phylogenetic methods and the study of pattern and process in kinship. In Kinship Systems: Change and Reconstruction, 3rd ed.; Patrick, M.C., Ian, K., Rachel, H., Eds.; University of Utah Press: Salt Lake City, UT, USA, 2013; pp. 43–58. [Google Scholar]
- Young, N.D.; Debellé, F.; Oldroyd, G.E.; Geurts, R.; Cannon, S.B.; Udvardi, M.K.; Benedito, V.A.; Mayer, K.F.; Gouzy, J.; Schoof, H. Thedicago genome provides insight into thvolution of rhizobial symbioses. Nature 2011, 480, 520–524. [Google Scholar] [CrossRef] [PubMed]
- Whitcomb, S.J.; Basu, A.; Allis, C.D.; Bernstein, E. Polycomb Group proteins: An evolutionary perspective. Trends Genet. 2007, 23, 494–502. [Google Scholar] [CrossRef] [PubMed]
- Pien, S.; Grossniklaus, U. Polycomb group and trithorax group proteins in Arabidopsis. BBA-Gene Struct. Expr. 2007, 1769, 375–382. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Jiang, L.; Liu, B.-Y.; Tan, C.-F.; Chen, D.-H.; Shen, W.-H.; Ruan, Y. Evolution and conservation of polycomb repressive complex 1 core components and putative associated factors in the green lineage. BMC Genom. 2019, 20, 533. [Google Scholar] [CrossRef]
- Chen, D.; Molitor, A.M.; Xu, L.; Shen, W.-H. Arabidopsis PRC1 core component AtRING1 regulates stem cell-determining carpel development mainly through repression of class I KNOX genes. BMC Biol. 2016, 14, 112. [Google Scholar] [CrossRef]
- Chen, D.-h.; Huang, Y.; Ruan, Y.; Shen, W.-H. Thvolutionary landscape oRC1 core components in green lineage. Planta 2016, 243, 825–846. [Google Scholar] [CrossRef]
- Hennig, L.; Taranto, P.; Walser, M.; Schönrock, N.; Gruissem, W. Arabidopsi SI1 is required fopigenetic maintenance of reproductivevelopment. Development 2003, 130, 2555–2565. [Google Scholar] [CrossRef]
- Yu, J.; Sun, H.; Zhang, J.; Hou, Y.; Zhang, T.; Kang, J.; Wang, Z.; Yang, Q.; Long, R. Analysis of aldo–keto reductase gene family and their responses to salt, drought, and abscisic acid stresses in Medicago truncatula. Int. J. Mol. Sci. 2020, 21, 754. [Google Scholar] [CrossRef]
- Sun, Q.; Yu, S.; Guo, Z. Calmodulin-Like (CML) gene family in Medicago truncatula: Genome-wide identification, characterization and expression analysis. Int. J. Mol. Sci. 2020, 21, 7142. [Google Scholar] [CrossRef]
- Pestana-Calsa, M.C.; Pacheco, C.M.; Castro, R.C.d.; Almeida, R.R.d.; Lira, N.P.V.d.; Calsa Junior, T. Cell wall, lignin and fatty acid-related transcriptome in soybean: Achieving genxpression patterns foioenergy legume. Genet. Mol. Biol. 2012, 35, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.-D.; Zhou, L.-J.; Fang, M.-J.; Dong, Q.-L.; An, X.-H.; You, C.-X.; Hao, Y.-J. Polycomb-group protein SlMSI1 represses thxpression of fruit-ripening genes to prolong shelf life in tomato. Sci. Rep. 2016, 6, 1–9. [Google Scholar]
- Picó, S.; Ortiz-Marchena, M.I.; Merini, W.; Calonje, M. Deciphering the role oOLYCOMB REPRESSIVE COMPLEX1 variants in regulating the acquisition of flowering competence in Arabidopsis. Plant Physiol. 2015, 168, 1286–1297. [Google Scholar] [CrossRef]
- Merini, W.; Romero-Campero, F.J.; Gomez-Zambrano, A.; Zhou, Y.; Turck, F.; Calonje, M. The Arabidopsis Polycomb repressive complex 1 (PRC1) components AtBMI1A, B, and C impact gene networks throughout all stages of plant development. Plant Physiol. 2017, 173, 627–641. [Google Scholar] [CrossRef]
- Bratzel, F.; Yang, C.; Angelova, A.; López-Torrejón, G.; Koch, M.; del Pozo, J.C.; Calonje, M. Regulation of the new Arabidopsis imprinted gene AtBMI1C requires the interplay oifferent epigenetic mechanisms. Mol. Plant 2012, 5, 260–269. [Google Scholar] [CrossRef]
- Steinbach, Y.; Hennig, L. Arabidopsis MSI1 functions in photoperiodic flowering time control. Front. Plant Sci. 2014, 5, 77. [Google Scholar] [CrossRef]
- Park, H.-Y.; Han, J.-S.; Woo, D.-H.; Seok, H.-Y.; Lee, J.-M.; Moon, B.Y.; Park, P.B.; Moon, Y.-H. Interaction between EMF1 and EMF1IP1, 6, 9 plays an important roluring early growth stage in Arabidopsis development. In Proceedings of the 3rd Asian and Oceanian Conference on Photobiology, Beijing, China, 17–20 November 2006. [Google Scholar]
- Qin, F.; Sakuma, Y.; Tran, L.-S.P.; Maruyama, K.; Kidokoro, S.; Fujita, Y.; Fujita, M.; Umezawa, T.; Sawano, Y.; Miyazono, K.-I. Arabidopsis DREB2A-interacting proteins function as RING E3 ligases and negatively regulate plant drought stress–responsive genxpression. Plant Cell 2008, 20, 1693–1707. [Google Scholar] [CrossRef]
- Pecrix, Y.; Staton, S.E.; Sallet, E.; Lelandais-Brière, C.; Moreau, S.; Carrere, S.; Blein, T.; Jardinaud, M.-F.; Latrasse, D.; Zouine, M. Whole-genome landscape o edicago truncatula symbiotic genes. Nature Plants 2018, 4, 1017–1025. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses oig biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Min, X.; Wu, H.; Zhang, Z.; Wei, X.; Jin, X.; Ndayambaza, B.; Wang, Y.; Liu, W. Genome-wide identification and characterization of the aquaporin gene family in Medicago truncatula. J. Plant Biochem. Biotechnol. 2019, 28, 320–335. [Google Scholar] [CrossRef]
- Sato, S.; Nakamura, Y.; Kaneko, T.; Asamizu, E.; Kato, T.; Nakao, M.; Sasamoto, S.; Watanabe, A.; Ono, A.; Kawashima, K. Genome structure of the legume, Lotus japonicus. DNA Res. 2008, 15, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Hou, D.; Cheng, Z.; Xie, L.; Li, X.; Li, J.; Mu, S.; Gao, J. The R2R3MYB gene family in Phyllostachydulis: Genome-wide analysis and identification of stress oevelopment-related R2R3MYBs. Front. Plant Sci. 2018, 9, 738. [Google Scholar] [CrossRef]
- Sun, Y.; Wu, Z.; Wang, Y.; Yang, J.; Wei, G.; Chou, M. Identification of phytocyanin gene family in legume plants and their involvement in nodulation o edicago truncatula. Plant Cell Physiol. 2019, 60, 900–915. [Google Scholar] [CrossRef] [PubMed]










| Gene Name | Gene ID | Location | ORF 1 Length (bp) | Number of Exons | MW 2 (kDa) | pI 3 Value | GRAVY 4 |
|---|---|---|---|---|---|---|---|
| MtBMI1A | Medtr7g096210.1 | 38,583,482–38,583,911 | 1290 | 7 | 47.37 | 9.24 | −0.576 |
| MtEMF1;1 | Medtr3g110082.1 | 50,968,972–50,970,140 | 3507 | 7 | 129.51 | 8.92 | −0.826 |
| MtEMF1;2 | Medtr3g110132.1 | 51,072,108–51,073,292 | 3555 | 5 | 131.39 | 9.02 | −0.839 |
| MtRING1A | Medtr2g090595.1 | 38,732,918–38,733,422 | 1515 | 10 | 58.11 | 4.77 | −1.259 |
| MtRING1B;1 | Medtr4g088520.1 | 35,141,049–35,141,489 | 1323 | 10 | 49.82 | 5.05 | −0.961 |
| MtRING1B;2 | Medtr4g091110.1 | 36,070,448–36,070,700 | 759 | 6 | 29.13 | 3.97 | −1.125 |
| MtVRN1;1 | Medtr1g021270.1 | 6,335,039–6,335,635 | 1791 | 6 | 66.67 | 8.79 | −0.379 |
| MtVRN1;2 | Medtr1g021290.1 | 21,113,921–21,114,394 | 1044 | 3 | 39.36 | 6.56 | −0.501 |
| MtVRN1;3 | Medtr1g021320.1 | 6,356,167–6,357,009 | 2529 | 8 | 94.10 | 8.70 | −0.476 |
| MtVRN1;4 | Medtr1g021330.1 | 6,366,047–6,366,432 | 1158 | 4 | 43.38 | 8.67 | −0.434 |
| MtVRN1;5 | Medtr1g021360.1 | 6,377,524–6,378,002 | 1437 | 5 | 54.49 | 8.91 | −0.542 |
| MtVRN1;6 | Medtr1g021380.1 | 6,390,073–6,390,423 | 1053 | 4 | 39.57 | 6.83 | −0.426 |
| MtVRN1;7 | Medtr1g021400.1 | 6,397,039–6,397,407 | 1107 | 4 | 41.12 | 8.52 | −0.516 |
| MtVRN1;8 | Medtr1g021410.1 | 6,403,159–6,403,520 | 1086 | 4 | 41.12 | 9.10 | −0.615 |
| MtVRN1;9 | Medtr1g021435.1 | 6,414,120–6,414,492 | 1119 | 4 | 41.83 | 8.78 | −0.430 |
| MtVRN1;10 | Medtr1g021440.1 | 6,419,474–6,419,873 | 1200 | 4 | 44.91 | 8.06 | −0.594 |
| MtVRN1;11 | Medtr1g021500.1 | 6,438,573–6,439,064 | 1476 | 6 | 54.50 | 8.78 | −0.617 |
| MtVRN1;12 | Medtr1g034210.1 | 12,425,305–12,425,675 | 1113 | 5 | 42.30 | 8.38 | −0.699 |
| MtVRN1;13 | Medtr1g034240.1 | 12,442,470–12,442,903 | 1302 | 4 | 49.15 | 8.93 | −0.577 |
| MtVRN1;14 | Medtr1g035460.1 | 12,873,695–12,874,091 | 1191 | 6 | 46.28 | 8.68 | −0.842 |
| MtVRN1;15 | Medtr1g108780.1 | 49,164,002–49,164,463 | 1386 | 6 | 53.61 | 8.73 | −0.666 |
| MtVRN1;16 | Medtr3g005420.1 | 198,959–199,501 | 1629 | 6 | 62.39 | 9.33 | −0.544 |
| MtVRN1;17 | Medtr3g061320.1 | 24,354,366–24,354,645 | 840 | 4 | 32.68 | 9.69 | −0.510 |
| MtVRN1;18 | Medtr3g098790.1 | 45,269,198–45,269,526 | 987 | 4 | 38.00 | 9.87 | −0.427 |
| MtVRN1;19 | Medtr4g068320.1 | 25,619,387–25,619,816 | 1290 | 4 | 48.52 | 8.75 | −0.536 |
| MtVRN1;20 | Medtr5g070440.1 | 29,839,127–29,839,401 | 825 | 3 | 31.93 | 9.54 | −0.570 |
| MtVRN1;21 | Medtr6g061070.1 | 21,113,921–21,114,394 | 1422 | 7 | 54.38 | 6.84 | −0.642 |
| MtVRN1;22 | Medtr7g050560.1 | 16,862,100–16,862,415 | 948 | 5 | 36.27 | 9.70 | −0.387 |
| MtVRN1;23 | Medtr7g050580.1 | 16,867,490–16,867,805 | 315 | 5 | 36.08 | 9.72 | −0.610 |
| MtVRN1;24 | Medtr7g050590.1 | 16,873,037–16,873,334 | 894 | 3 | 34.41 | 9.99 | −0.631 |
| MtVRN1;25 | Medtr7g050710.1 | 17,605,025–17,605,349 | 975 | 4 | 37.06 | 9.91 | −0.544 |
| MtVRN1;26 | Medtr7g050720.1 | 17,612,279–17,612,599 | 963 | 4 | 36.77 | 9.50 | −0.457 |
| MtVRN1;27 | Medtr8g103940.1 | 43,749,832–43,750,330 | 1497 | 7 | 57.36 | 9.15 | −0.679 |
| MtSWN | Medtr1g086980.1 | 38,931,514–38,932,145 | 2496 | 17 | 93.50 | 7.97 | −0.775 |
| MtCLF | Medtr5g016870.1 | 6,095,959–6,096,822 | 2592 | 18 | 96.55 | 8.60 | −0.819 |
| MtMEA;1 | Medtr7g055660.1 | 19,084,301–19,085,033 | 2199 | 16 | 84.19 | 8.18 | −0.799 |
| MtMEA;2 | Medtr7g109560.1 | 44,830,300–44,831,064 | 2295 | 16 | 87.40 | 7.23 | −0.813 |
| MtEMF2 | Medtr1g090240.1 | 40,419,285–40,419,977 | 2079 | 17 | 77.93 | 6.41 | −0.348 |
| MtVRN2 | Medtr5g013150.1 | 4,143,460–4,143,938 | 1437 | 16 | 54.56 | 8.30 | −0.567 |
| MtFIE;1 | Medtr1g011610.2 | 2,109,762–2,110,664 | 2709 | 18 | 99.01 | 6.45 | −0.695 |
| MtFIE;2 | Medtr1g028310.1 | 9,510,205–9,510,577 | 1119 | 13 | 41.61 | 6.21 | −0.119 |
| MtFIE;3 | Medtr1g072140.1 | 32,005,164–32,005,480 | 951 | 2 | 34.85 | 8.69 | −0.275 |
| MtFIE;4 | Medtr1g090827.1 | 40,854,644–40,855,860 | 3651 | 3 | 136.27 | 6.45 | −0.186 |
| MtFIE;5 | Medtr2g436510.1 | 14,197,679–14,198,159 | 1443 | 17 | 53.14 | 9.23 | −0.485 |
| MtFIE;6 | Medtr2g086140.1 | 36,225,284–36,226,276 | 2979 | 10 | 108.23 | 8.04 | −0.328 |
| MtFIE;7 | Medtr2g102267.1 | 44,039,290–44,039,607 | 954 | 10 | 36.20 | 5.31 | −0.348 |
| MtFIE;8 | Medtr2g102277.2 | 44,045,802–44,046,723 | 2766 | 24 | 104.06 | 4.93 | −0.292 |
| MtFIE;9 | Medtr3g061450.1 | 24,506,254–24,506,567 | 942 | 11 | 35.17 | 6.34 | −0.383 |
| MtFIE;10 | Medtr3g106080.1 | 48,932,864–48,933,533 | 2010 | 19 | 74.24 | 6.50 | −0.438 |
| MtFIE;11 | Medtr3g116500.1 | 54,500,761–54,501,139 | 1137 | 6 | 41.04 | 7.58 | −0.217 |
| MtFIE;12 | Medtr4g011950.1 | 3,145,419–3,146,350 | 2796 | 24 | 105.08 | 4.87 | −0.327 |
| MtFIE;13 | Medtr4g053675.1 | 19,445,007–19,446,038 | 3096 | 10 | 112.64 | 6.62 | −0.358 |
| MtFIE;14 | Medtr4g071130.1 | 26,806,737–26,807,943 | 3621 | 4 | 135.47 | 6.49 | −0.221 |
| MtFIE;15 | Medtr4g095058.1 | 39,607,610–39,608,188 | 1737 | 14 | 64.92 | 6.39 | −0.595 |
| MtFIE;16 | Medtr4g119620.1 | 49,575,751–49,576,077 | 981 | 2 | 35.88 | 6.78 | −0.350 |
| MtFIE;17 | Medtr4g125710.1 | 52,171,088–52,171,409 | 966 | 2 | 34.39 | 5.55 | −0.075 |
| MtFIE;18 | Medtr5g022690.1 | 8,972,131–8,972,619 | 1467 | 5 | 55.02 | 6.08 | −0.646 |
| MtFIE;19 | Medtr6g004040.1 | 91,010–91,770 | 2283 | 18 | 83.11 | 7.93 | −0.306 |
| MtFIE;20 | Medtr7g058460.1 | 21,059,701–21,060,485 | 2355 | 17 | 85.49 | 6.36 | −0.566 |
| MtMSI1;1 | Medtr2g039250.1 | 17,196,265–17,196,765 | 1503 | 15 | 55.61 | 6.11 | −0.627 |
| MtMSI1;2 | Medtr2g100090.1 | 42,972,259–42,972,738 | 1440 | 12 | 52.71 | 5.26 | −0.406 |
| MtMSI1;3 | Medtr4g073080.1 | 27,640,107–27,640,512 | 1218 | 6 | 45.75 | 4.66 | −0.536 |
| MtMSI1;4 | Medtr4g096880.1 | 38,572,495–38,572,918 | 1272 | 5 | 48.22 | 4.75 | −0.569 |
| MtMSI1;5 | Medtr8g080700.1 | 34,739,363–34,739,832 | 1410 | 12 | 52.04 | 4.97 | −0.586 |
| Gene Name | Protein Secondary Structure | Subcellular Location Prediction | ||||
|---|---|---|---|---|---|---|
| α-Helix (%) | Extended Strand (%) | Β-Sheet (%) | Random coil (%) | Plant-mPLoc | WoLF PSORT | |
| MtBMI1A | 31.24 | 7.93 | 2.10 | 58.74 | Nucleus | Nuclear |
| MtEMF1;1 | 17.64 | 7.45 | 1.71 | 73.20 | Nucleus | Nuclear |
| MtEMF1;2 | 18.58 | 9.29 | 2.28 | 69.85 | Nucleus | Nuclear |
| MtRING1A | 44.64 | 7.34 | 2.98 | 45.04 | Nucleus | Nuclear |
| MtRING1B;1 | 40.91 | 7.95 | 2.73 | 48.41 | Nucleus | Nuclear |
| MtRING1B;2 | 50.79 | 11.90 | 5.16 | 32.14 | Nucleus | Nuclear |
| MtVRN1;1 | 20.81 | 20.97 | 5.54 | 52.68 | Chloroplast/Nucleus | Nuclear |
| MtVRN1;2 | 24.50 | 23.05 | 4.32 | 48.13 | Cell wall/Nucleus | Nuclear |
| MtVRN1;3 | 28.74 | 18.76 | 5.23 | 47.27 | Nucleus | Nuclear |
| MtVRN1;4 | 22.60 | 16.36 | 4.16 | 56.88 | Chloroplast/Nucleus | Nuclear |
| MtVRN1;5 | 24.48 | 23.01 | 6.07 | 46.44 | Chloroplast/Nucleus | Nuclear |
| MtVRN1;6 | 19.71 | 20.29 | 6.86 | 53.14 | Nucleus | Nuclear |
| MtVRN1;7 | 19.02 | 22.01 | 5.71 | 53.26 | Cell wall/Nucleus | Nuclear |
| MtVRN1;8 | 18.28 | 22.99 | 4.16 | 54.57 | Cell wall/Nucleus | Nuclear |
| MtVRN1;9 | 17.74 | 20.70 | 6.72 | 54.84 | Cell wall/Nucleus | Nuclear |
| MtVRN1;10 | 20.55 | 18.30 | 4.76 | 56.39 | Nucleus | Nuclear |
| MtVRN1;11 | 24.85 | 16.29 | 2.65 | 56.21 | Nucleus | Nuclear |
| MtVRN1;12 | 22.16 | 20.27 | 10.00 | 47.57 | Nucleus | Chloroplast |
| MtVRN1;13 | 20.32 | 18.94 | 6.70 | 54.04 | Nucleus | Chloroplast |
| MtVRN1;14 | 15.40 | 21.46 | 5.30 | 57.83 | Nucleus | Nuclear |
| MtVRN1;15 | 14.75 | 22.34 | 6.07 | 56.83 | Nucleus | Nuclear |
| MtVRN1;16 | 22.14 | 23.43 | 5.90 | 48.52 | Nucleus | Chloroplast |
| MtVRN1;17 | 14.70 | 25.45 | 9.32 | 50.54 | Chloroplast/Nucleus | Chloroplast |
| MtVRN1;18 | 13.41 | 24.39 | 5.79 | 56.40 | Chloroplast/Nucleus | Nuclear |
| MtVRN1;19 | 22.84 | 19.11 | 6.29 | 51.75 | Chloroplast/Nucleus | Nuclear |
| MtVRN1;20 | 17.88 | 27.74 | 7.30 | 47.08 | Nucleus | Nuclear |
| MtVRN1;21 | 15.01 | 23.89 | 7.19 | 53.91 | Nucleus | Nuclear |
| MtVRN1;22 | 13.33 | 23.49 | 8.25 | 54.92 | Chloroplast/Nucleus | Cytoplasmic |
| MtVRN1;23 | 15.24 | 25.08 | 7.62 | 52.06 | Chloroplast/Nucleus | Nuclear |
| MtVRN1;24 | 15.49 | 24.24 | 7.07 | 53.20 | Nucleus | Nuclear |
| MtVRN1;25 | 14.20 | 23.77 | 6.79 | 55.25 | Nucleus | Nuclear |
| MtVRN1;26 | 13.75 | 24.69 | 9.06 | 52.50 | Nucleus | Cytoplasmic |
| MtVRN1;27 | 14.66 | 21.89 | 5.62 | 57.83 | Nucleus | Nuclear |
| MtSWN | 29.96 | 10.71 | 5.90 | 53.43 | Nucleus | Nuclear |
| MtCLF | 30.94 | 10.43 | 4.98 | 53.65 | Nucleus | Nuclear |
| MtMEA;1 | 32.24 | 12.02 | 4.10 | 51.64 | Nucleus | Nuclear |
| MtMEA;2 | 31.94 | 13.22 | 3.27 | 51.57 | Nucleus | Nuclear |
| MtEMF2 | 26.59 | 21.39 | 5.78 | 46.24 | Nucleus | Nuclear |
| MtVRN2 | 33.05 | 12.13 | 5.02 | 49.79 | Nucleus | Nuclear |
| MtFIE;1 | 29.82 | 16.63 | 7.32 | 46.23 | Nucleus | Nuclear |
| MtFIE;2 | 10.48 | 35.22 | 11.29 | 43.01 | Nucleus | Cytoplasmic |
| MtFIE;3 | 6.01 | 44.94 | 13.61 | 35.44 | Nucleus | Chloroplast |
| MtFIE;4 | 32.07 | 23.60 | 8.80 | 35.53 | Nucleus | Nuclear |
| MtFIE;5 | 11.04 | 31.67 | 9.17 | 48.12 | Nucleus | Nuclear |
| MtFIE;6 | 18.65 | 22.98 | 6.35 | 52.02 | Nucleus | Nuclear |
| MtFIE;7 | 12.62 | 41.64 | 14.20 | 31.55 | Nucleus | Nuclear |
| MtFIE;8 | 35.40 | 25.95 | 9.66 | 28.99 | Nucleus | Cytoplasmic |
| MtFIE;9 | 7.35 | 38.98 | 9.90 | 43.77 | Nucleus | Cytoplasmic |
| MtFIE;10 | 31.84 | 23.17 | 6.28 | 38.71 | Nucleus | Nuclear |
| MtFIE;11 | 15.34 | 36.51 | 11.38 | 36.77 | Endoplasmic reticulum | Nuclear |
| MtFIE;12 | 34.16 | 25.99 | 10.42 | 29.43 | Endoplasmic reticulum/Golgi apparatus | Chloroplast |
| MtFIE;13 | 20.76 | 21.34 | 5.92 | 51.99 | Nucleus | Nuclear |
| MtFIE;14 | 31.51 | 23.71 | 8.71 | 36.07 | Nucleus | Chloroplast |
| MtFIE;15 | 17.82 | 29.58 | 9.34 | 43.25 | Nucleus | Nuclear |
| MtFIE;16 | 3.68 | 45.09 | 14.42 | 36.81 | Nucleus | Nuclear |
| MtFIE;17 | 10.59 | 40.50 | 12.46 | 36.45 | Nucleus | Cytoplasmic |
| MtFIE;18 | 32.38 | 28.89 | 10.45 | 28.28 | Nucleus | Nuclear |
| MtFIE;19 | 26.32 | 19.74 | 6.32 | 47.63 | Nucleus | Chloroplast |
| MtFIE;20 | 28.19 | 17.22 | 5.74 | 48.85 | Nucleus | Nuclear |
| MtMSI1;1 | 16.40 | 26.00 | 5.00 | 52.60 | Nucleus | Nuclear |
| MtMSI1;2 | 12.94 | 28.60 | 2.92 | 55.53 | Nucleus | Cytoplasmic |
| MtMSI1;3 | 12.59 | 30.62 | 5.93 | 50.86 | Nucleus | Nuclear |
| MtMSI1;4 | 14.89 | 29.79 | 5.67 | 49.65 | Nucleus | Nuclear |
| MtMSI1;5 | 16.20 | 24.00 | 5.54 | 54.16 | Nucleus | Mitochondrial |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, Y.; Zhang, J.; Sun, Z.; Tang, Y.; Wu, Y. Genome-Wide Identification and Analysis of the Polycomb Group Family in Medicago truncatula. Int. J. Mol. Sci. 2021, 22, 7537. https://doi.org/10.3390/ijms22147537
Zhao Y, Zhang J, Sun Z, Tang Y, Wu Y. Genome-Wide Identification and Analysis of the Polycomb Group Family in Medicago truncatula. International Journal of Molecular Sciences. 2021; 22(14):7537. https://doi.org/10.3390/ijms22147537
Chicago/Turabian StyleZhao, Yuanyuan, Junchao Zhang, Zhanmin Sun, Yixiong Tang, and Yanmin Wu. 2021. "Genome-Wide Identification and Analysis of the Polycomb Group Family in Medicago truncatula" International Journal of Molecular Sciences 22, no. 14: 7537. https://doi.org/10.3390/ijms22147537
APA StyleZhao, Y., Zhang, J., Sun, Z., Tang, Y., & Wu, Y. (2021). Genome-Wide Identification and Analysis of the Polycomb Group Family in Medicago truncatula. International Journal of Molecular Sciences, 22(14), 7537. https://doi.org/10.3390/ijms22147537






