A Positive Regulatory Feedback Loop between EKLF/KLF1 and TAL1/SCL Sustaining the Erythropoiesis
Abstract
:1. Introduction
2. Result
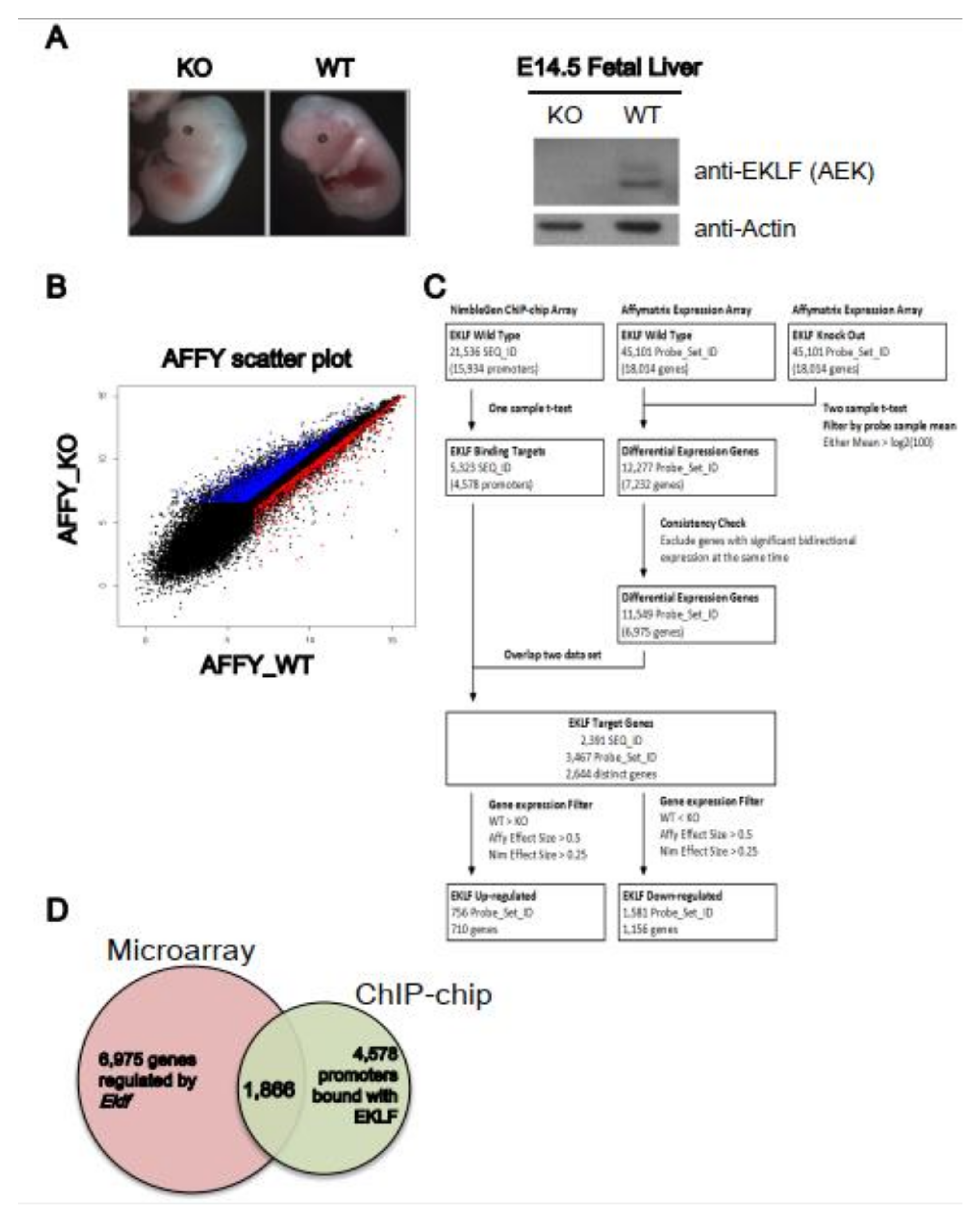
2.1. Genome-Wide Identification of EKLF Target Genes by Microarray Hybridization and Promoter-Based ChIP-Chip Analyses
2.2. Functional and Pathway Analysis of EKLF Target Genes
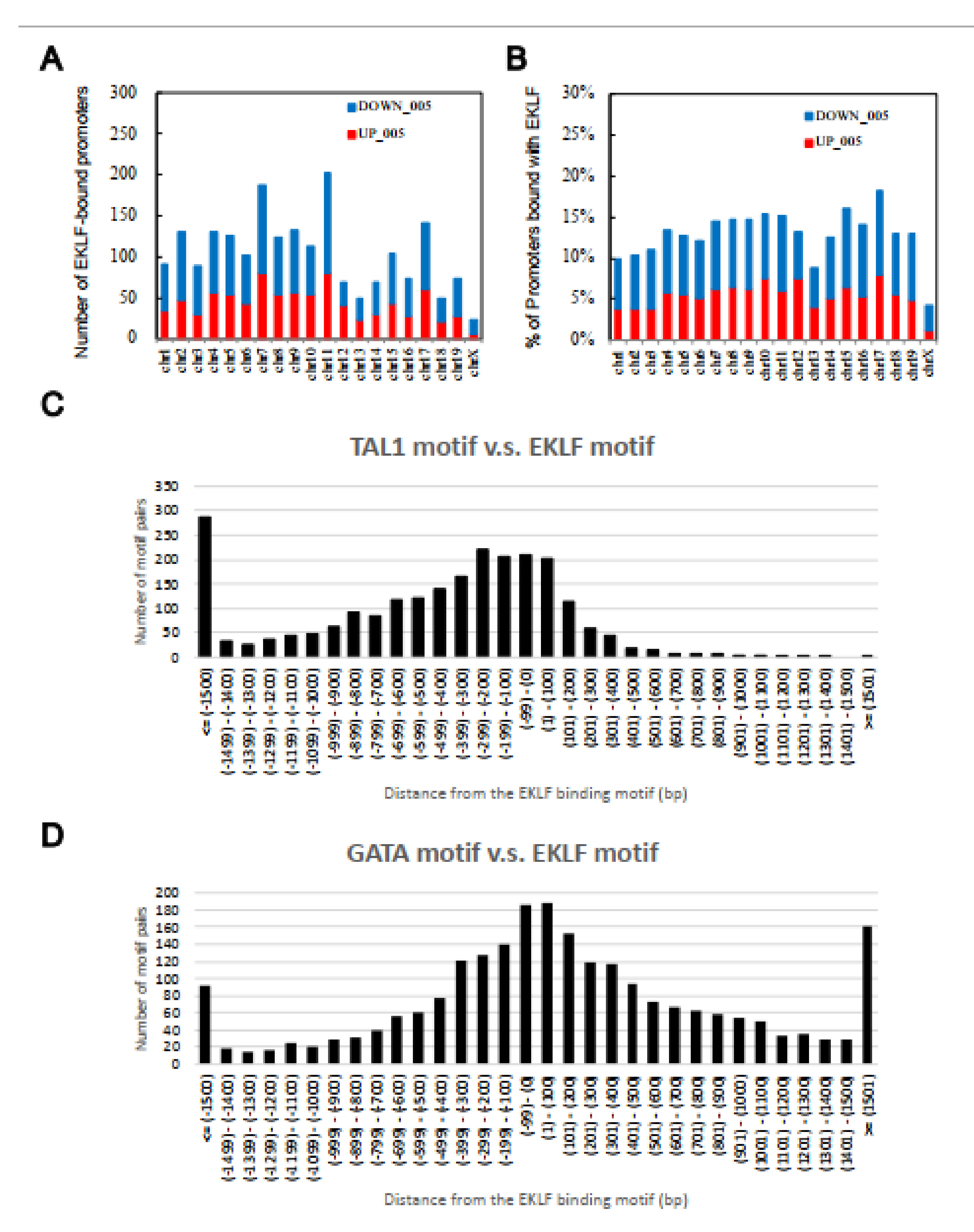
2.3. Identification of Potential Transcription Factors Co-Regulating the EKLF Targets
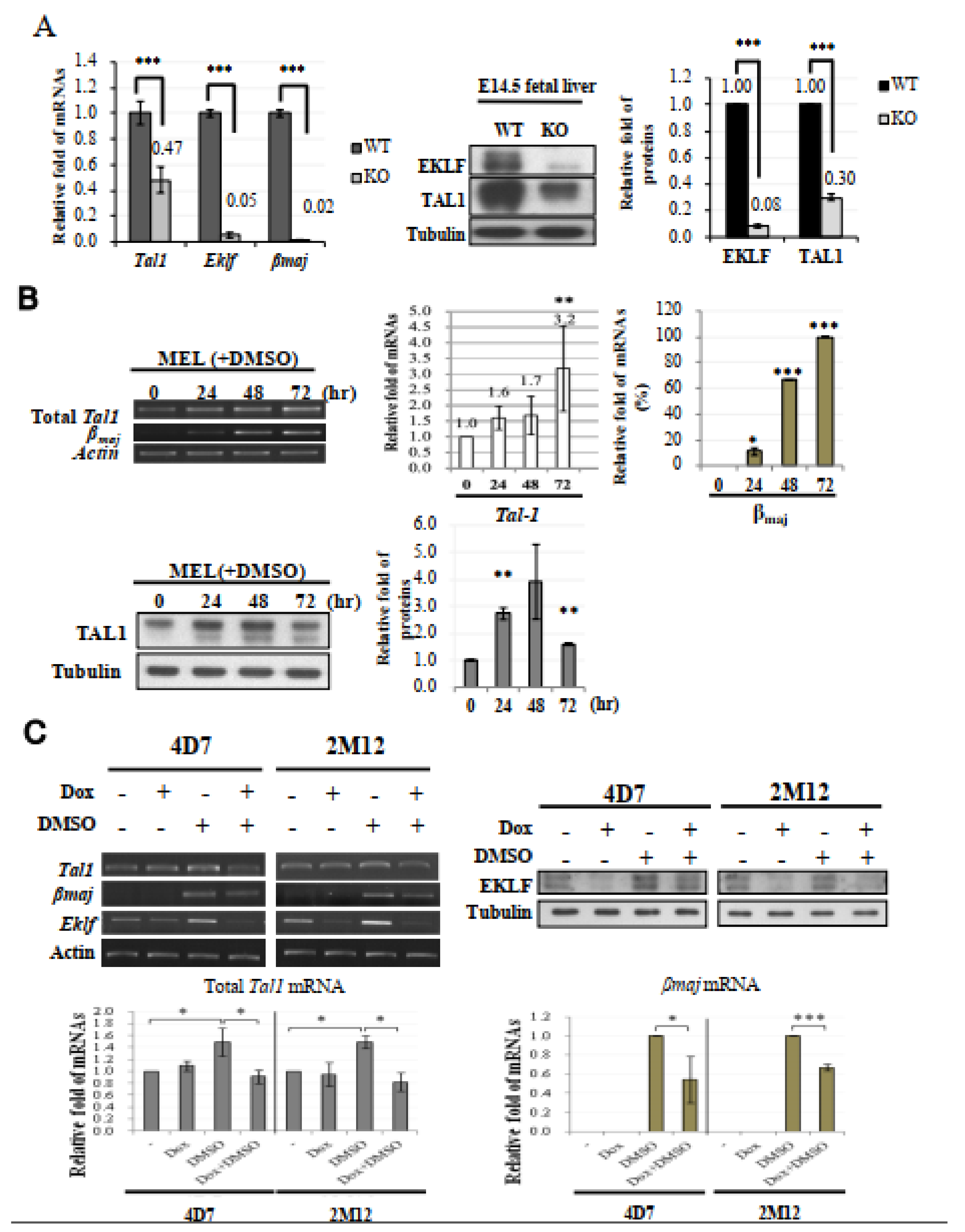
2.4. Likelihood of Tal1 Gene as a Regulatory Target of EKLF in E14.5 Fetal Liver Cells
2.5. EKLF As an Activator of Tal1 Gene Expression during Erythroid Differentiation
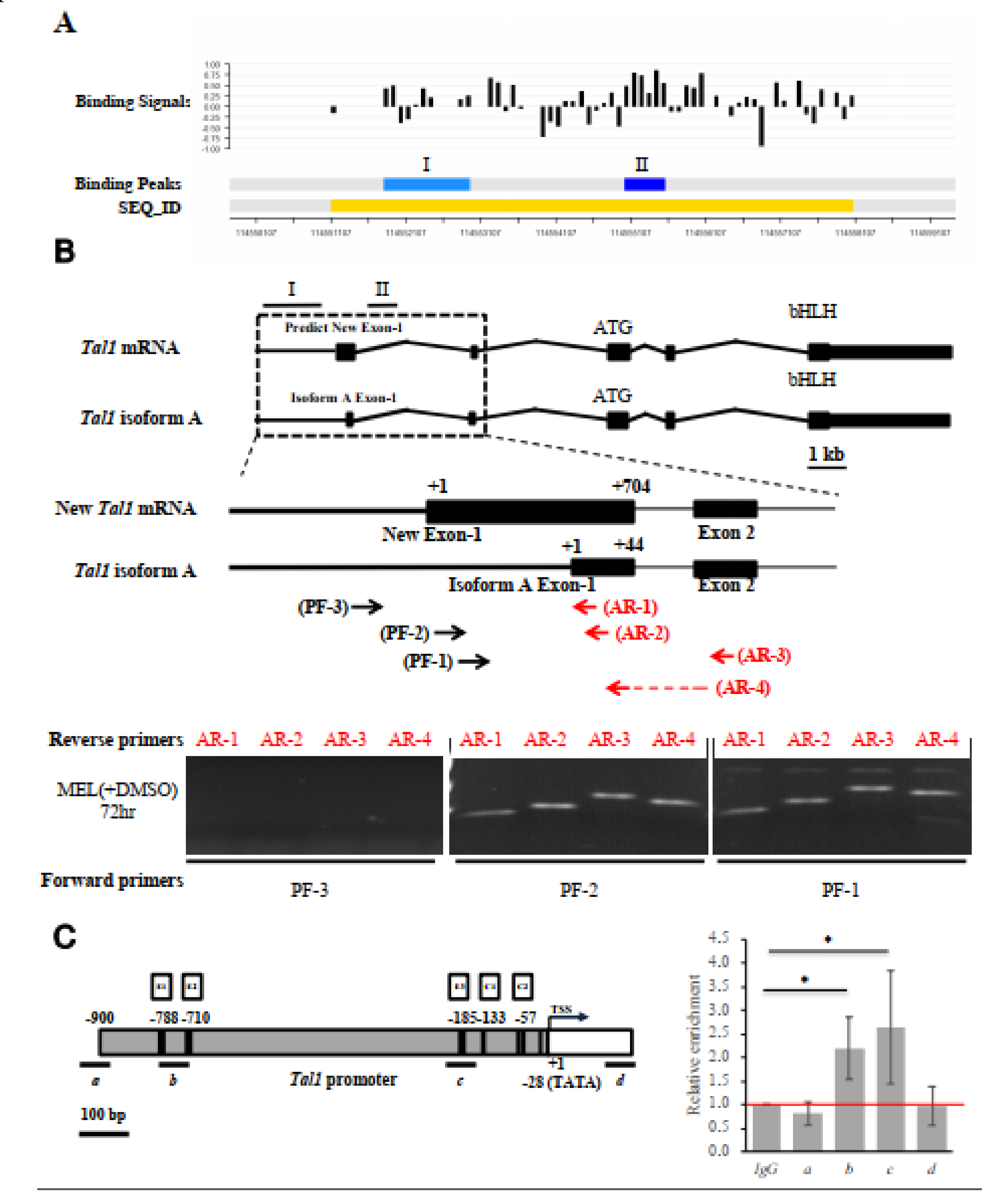
2.6. Binding In Vivo of EKLF to the Upstream Promoter of Tal1 Gene
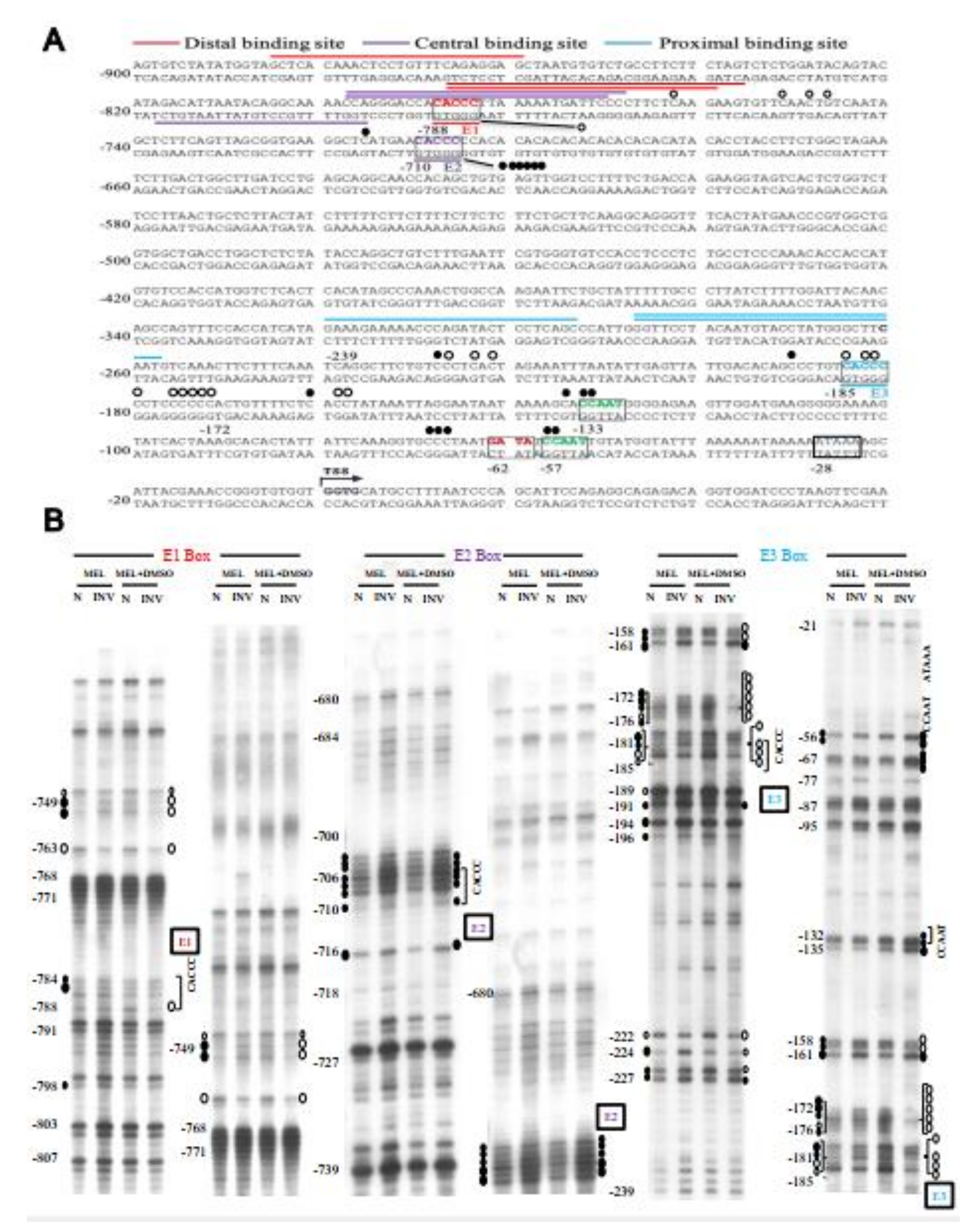
2.7. Binding In Vivo of EKLF to the Proximal CACCC Box of Tal1 Promoter-Genomic Footprinting Analysis
2.8. Requirement of the Proximal CACCC Motif for Transcriptional Activation of the Tal1 Promoter by EKLF
3. Discussion
4. Material and Methods
4.1. Generation of Eklf−/− Mice
4.2. Gene expression Profiling by Affymetrix Array Hybridization
4.3. Identification of Differentially Expressed Genes
4.4. Identification of EKLF-Bound Targets by Using NimbleGen ChIP-Chip Array Hybridization
4.5. Matching between Affymetrix Probes and NimbleGen ChIP-Chip Probes
4.6. Co-Occurrence of Binding Motifs and Relative Distance Distribution
4.7. Functional Enrichment Analysis
4.8. ChIP-qPCR
4.9. Plasmid Construction
4.10. Cell Culture, Differentiation, DNA Transfection, and Knockdown of Gene Expression
4.11. RNA Analysis
4.12. Western Blotting Analysis and Antibodies
4.13. Reporter Assay
4.14. In Vivo Genomic Footprinting
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Orkin, S.H.; Zon, L.I. Hematopoiesis: An Evolving Paradigm for Stem Cell Biology. Cell 2008, 132, 631–644. [Google Scholar] [CrossRef] [Green Version]
- Doré, L.C.; Crispino, J.D. Transcription factor networks in erythroid cell and megakaryocyte development. Blood 2011, 118, 231–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dzierzak, E.; Philipsen, S. Erythropoiesis: Development and Differentiation. Cold Spring Harb. Perspect. Med. 2013, 3, a011601. [Google Scholar] [CrossRef]
- Kerenyi, M.A.; Orkin, S.H. Networking erythropoiesis. J. Exp. Med. 2010, 207, 2537–2541. [Google Scholar] [CrossRef] [Green Version]
- Crispino, J.D.; Weiss, M.J. Erythro-megakaryocytic transcription factors associated with hereditary anemia. Blood 2014, 123, 3080–3088. [Google Scholar] [CrossRef] [Green Version]
- Frontelo, P.; Manwani, D.; Galdass, M.; Karsunky, H.; Lohmann, F.; Gallagher, P.G.; Bieker, J.J. Novel role for EKLF in megakaryocyte lineage commitment. Blood 2007, 110, 3871–3880. [Google Scholar] [CrossRef] [Green Version]
- Bouilloux, F.; Juban, G.; Cohet, N.; Buet, D.; Guyot, B.; Vainchenker, W.; Louache, F.; Morle, F. EKLF restricts megakaryocytic differentiation at the benefit of erythrocytic differentiation. Blood 2008, 112, 576–584. [Google Scholar] [CrossRef] [PubMed]
- Tallack, M.R.; Perkins, A.C. Megakaryocyte-erythroid lineage promiscuity in EKLF null mouse blood. Haematologica 2009, 95, 144–147. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siatecka, M.; Bieker, J.J. The multifunctional role of EKLF/KLF1 during erythropoiesis. Blood 2011, 118, 2044–2054. [Google Scholar] [CrossRef] [Green Version]
- Perkins, A.; Xu, X.; Higgs, D.-R.; The KLF1 Consensus Workgroup; Patrinos, G.-P.; Arnaud, L.; Bieker, J.-J.; Philipsen, S. “Krüppeling” erythropoiesis: An unexpected broad spectrum of human red blood cell disorders due to KLF1 variants unveiled by genomic sequencing. Blood 2016, 127, 1856–1862. [Google Scholar] [CrossRef] [Green Version]
- Hung, C.H.; Wang, K.Y.; Liou, Y.H.; Wang, J.P.; Huang, A.Y.S.; Lee, T.L.; Jiang, S.T.; Liao, N.S.; Shyu, Y.C.; Shen, J.C.K. Negative regulation of the differentiation of Flk2− CD34− LSK hematopoietic stem cell by EKLF/KLF1. Int. J. Mol. Sci. 2020, 21, 8448. [Google Scholar] [CrossRef]
- Miller, I.J.; Bieker, J.J. A novel, erythroid cell-specific murine transcription factor that binds to the CACCC element and is related to the Krüppel family of nuclear proteins. Mol. Cell. Biol. 1993, 13, 2776–2786. [Google Scholar] [CrossRef] [Green Version]
- Andrews, N.; Orkin, S.H. Transcriptional control of erythropoiesis. Curr. Opin. Hematol. 1994, 1, 119–124. [Google Scholar] [PubMed]
- Porcu, S.; Manchinu, M.F.; Marongiu, M.F.; Sogos, V.; Poddie, D.; Asunis, I.; Porcu, L.; Marini, M.G.; Moi, P.; Cao, A.; et al. Klf1 Affects DNase II-Alpha Expression in the Central Macrophage of a Fetal Liver Erythroblastic Island: A Non-Cell-Autonomous Role in Definitive Erythropoiesis. Mol. Cell. Biol. 2011, 31, 4144–4154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perkins, A.C.; Sharpe, A.H.; Orkin, S.H. Lethal β-thalassaemia in mice lacking the erythroid CACCC-transcription factor EKLF. Nature 1995, 375, 318–322. [Google Scholar] [CrossRef] [PubMed]
- Nuez, B.; Michalovich, D.; Bygrave, A.; Ploemacher, R.; Grosveld, F. Defective haematopoiesis in fetal liver resulting from inactivation of the EKLF gene. Nat. Cell Biol. 1995, 375, 316–318. [Google Scholar] [CrossRef] [PubMed]
- Drissen, R.; von Lindern, M.; Kolbus, A.; Driegen, S.; Steinlein, P.; Beug, H.; Grosveld, F.; Philipsen, S. The Erythroid Phenotype of EKLF-Null Mice: Defects in Hemoglobin Metabolism and Membrane Stability. Mol. Cell. Biol. 2005, 25, 5205–5214. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Bieker, J.J. Stage-Specific Repression by the EKLF Transcriptional Activator. Mol. Cell. Biol. 2004, 24, 10416–10424. [Google Scholar] [CrossRef] [Green Version]
- Shyu, Y.-C.; Wen, S.-C.; Lee, T.-L.; Chen, X.; Hsu, C.-T.; Chen, H.; Chen, R.-L.; Hwang, J.-L.; Shen, C.-K.J. Chromatin-binding in vivo of the erythroid kruppel-like factor, EKLF, in the murine globin loci. Cell Res. 2006, 16, 347–355. [Google Scholar] [CrossRef] [Green Version]
- Shyu, Y.C.; Lee, T.L.; Wen, S.C.; Chen, H.; Hsiao, W.Y.; Chen, X.; Hwang, J.; Shen, C.K. Subcellular transport of EKLF and switch-on of murine adult beta maj globin gene transcription. Mol. Cell. Biol. 2007, 27, 2309–2323. [Google Scholar] [CrossRef] [Green Version]
- Zhang, W.; Bieker, J.J. Acetylation and modulation of erythroid Kruppel-like factor (EKLF) activity by interaction with histone acetyltransferases. Proc. Natl. Acad. Sci. USA 1998, 95, 9855–9860. [Google Scholar] [CrossRef] [Green Version]
- Kadam, S.; McAlpine, G.S.; Phelan, M.L.; Kingston, R.E.; Jones, K.A.; Emerson, B.M. Functional selectivity of recombinant mammalian SWI/SNF subunits. Genes Dev. 2000, 14, 2441–2451. [Google Scholar] [CrossRef] [Green Version]
- Zhang, W.; Kadam, S.; Emerson, B.M.; Bieker, J.J. Site-specific acetylation by p300 or CREB binding protein regulates erythroid Kruppel-like factor transcriptional activity via its interaction with the SWI-SNF complex. Mol. Cell. Biol. 2001, 21, 2413–2422. [Google Scholar] [CrossRef] [Green Version]
- Singleton, B.K.; Frayne, J.; Anstee, D.J. Blood group phenotypes resulting from mutations in erythroid transcription factors. Curr. Opin. Hematol. 2012, 19, 486–493. [Google Scholar] [CrossRef]
- Tallack, M.; Perkins, A. Three fingers on the switch: Krüppel-like factor 1 regulation of γ-globin to β-globin gene switching. Curr. Opin. Hematol. 2013, 20, 193–200. [Google Scholar] [CrossRef]
- Shyu, Y.C.; Lee, T.L.; Chen, X.; Hsu, P.H.; Wen, S.C.; Liaw, Y.W.; Lu, C.H.; Hsu, P.Y.; Lu, M.J.; Hwang, J.; et al. Tight regulation of a timed nuclear import wave of EKLF by PKCtheta and FOE during Pro-E to Baso-E transition. Dev. Cell 2014, 28, 409–422. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tallack, M.R.; Whitington, T.; Yuen, W.S.; Wainwright, E.N.; Keys, J.R.; Gardiner, B.B.; Nourbakhsh, E.; Cloonan, N.; Grimmond, S.M.; Bailey, T.L.; et al. A global role for KLF1 in erythropoiesis revealed by ChIP-seq in primary erythroid cells. Genome Res. 2010, 20, 1052–1063. [Google Scholar] [CrossRef] [Green Version]
- Pilon, A.M.; Ajay, S.S.; Kumar, S.A.; Steiner, L.A.; Cherukuri, P.F.; Wincovitch, S.; Anderson, S.M.; NISC Comparative Sequencing Center; Mullikin, J.C.; Gallagher, P.G.; et al. Genome-wide ChIP-Seq reveals a dramatic shift in the binding of the transcription factor erythroid Kruppel-like factor during erythrocyte differentiation. Blood 2011, 118, e139–e148. [Google Scholar] [CrossRef]
- Tallack, M.R.; Magor, G.W.; Dartigues, B.; Sun, L.; Huang, S.; Fittock, J.M.; Fry, S.V.; Glazov, E.A.; Bailey, T.L.; Perkins, A.C. Novel roles for KLF1 in erythropoiesis revealed by mRNA-seq. Genome Res. 2012, 22, 2385–2398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Robb, L.; Lyons, I.; Li, R.; Hartley, L.; Kontgen, F.; Harvey, R.; Metcalf, D.; Begley, C.G. Absence of yolk sac hematopoiesis from mice with a targeted disruption of the scl gene. Proc. Natl. Acad. Sci. USA 1995, 92, 7075–7079. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shivdasani, R.A.; Mayer, E.L.; Orkin, S.H. Absence of blood formation in mice lacking the T-cell leukaemia oncoprotein tal-1/SCL. Nat. Cell Biol. 1995, 373, 432–434. [Google Scholar] [CrossRef] [PubMed]
- Aplan, P.D.; Nakahara, K.; Orkin, S.H.; Kirsch, I.R. The SCL gene product: A positive regulator of erythroid differentiation. EMBO J. 1992, 11, 4073–4081. [Google Scholar] [CrossRef]
- Hall, M.A.; Curtis, D.; Metcalf, D.; Elefanty, A.; Sourris, K.; Robb, L.; Göthert, J.; Jane, S.M.; Begley, C.G. The critical regulator of embryonic hematopoiesis, SCL, is vital in the adult for megakaryopoiesis, erythropoiesis, and lineage choice in CFU-S12. Proc. Natl. Acad. Sci. USA 2003, 100, 992–997. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mikkola, H.K.A.; Klintman, J.; Yang, H.; Hock, H.; Schlaeger, T.M.; Fujiwara, Y.; Orkin, S.H. Haematopoietic stem cells retain long-term repopulating activity and multipotency in the absence of stem-cell leukaemia SCL/tal-1 gene. Nat. Cell Biol. 2003, 421, 547–551. [Google Scholar] [CrossRef] [PubMed]
- Weiss, M.J.; Orkin, S.H. Transcription factor GATA-1 permits survival and maturation of erythroid precursors by preventing apoptosis. Proc. Natl. Acad. Sci. USA 1995, 92, 9623–9627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, M.; Riva, L.; Xie, H.; Schindler, Y.; Moran, T.B.; Cheng, Y.; Yu, D.; Hardison, R.; Weiss, M.; Orkin, S.H.; et al. Insights into GATA-1-Mediated Gene Activation versus Repression via Genome-wide Chromatin Occupancy Analysis. Mol. Cell 2009, 36, 682–695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimizu, R.; Yamamoto, M. Contribution of GATA1 dysfunction to multi-step leukemogenesis. Cancer Sci. 2012, 103, 2039–2044. [Google Scholar] [CrossRef]
- Han, G.C.; Vinayachandran, V.; Bataille, A.R.; Park, B.; Chan-Salis, K.Y.; Keller, C.A.; Long, M.; Mahony, S.; Hardison, R.; Pugh, B.F. Genome-Wide Organization of GATA1 and TAL1 Determined at High Resolution. Mol. Cell. Biol. 2016, 36, 157–172. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kassouf, M.T.; Hughes, J.R.; Taylor, S.; McGowan, S.J.; Soneji, S.; Green, A.L.; Vyas, P.; Porcher, C. Genome-wide identification of TAL1’s functional targets: Insights into its mechanisms of action in primary erythroid cells. Genome Res. 2010, 20, 1064–1083. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anderson, K.P.; Crable, S.C.; Lingrel, J.B. Multiple Proteins Binding to a GATA-E Box-GATA Motif Regulate the Erythroid Krüppel-like Factor (EKLF) Gene. J. Biol. Chem. 1998, 273, 14347–14354. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pilon, A.M.; Arcasoy, M.O.; Dressman, H.K.; Vayda, S.E.; Maksimova, Y.D.; Sangerman, J.I.; Gallagher, P.G.; Bodine, D.M. Failure of Terminal Erythroid Differentiation in EKLF-Deficient Mice Is Associated with Cell Cycle Perturbation and Reduced Expression of E2F2. Mol. Cell. Biol. 2008, 28, 7394–7401. [Google Scholar] [CrossRef] [Green Version]
- Gregory, R.C.; Taxman, D.J.; Seshasayee, D.; Kensinger, M.H.; Bieker, J.J.; Wojchowski, D.M. Functional interaction of GATA1 with erythroid Krüppel-like factor and Sp1 at defined erythroid promoters. Blood 1996, 87, 1793–1801. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wadman, I.A.; Osada, H.; Grütz, G.G.; Agulnick, A.D.; Westphal, H.; Forster, A.; Rabbitts, T. The LIM-only protein Lmo2 is a bridging molecule assembling an erythroid, DNA-binding complex which includes the TAL1, E47, GATA-1 and Ldb1/NLI proteins. EMBO J. 1997, 16, 3145–3157. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, R.; Ohneda, K.; Yamamoto, M.; Philipsen, S. GATA1 Function, a Paradigm for Transcription Factors in Hematopoiesis. Mol. Cell. Biol. 2005, 25, 1215–1227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, L.; Freudenberg, J.; Cui, K.; Dale, R.; Song, S.-H.; Dean, A.; Zhao, K.; Jothi, R.; Love, P.E. Ldb1-nucleated transcription complexes function as primary mediators of global erythroid gene activation. Blood 2013, 121, 4575–4585. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bosè, F.; Fugazza, C.; Casalgrandi, M.; Capelli, A.; Cunningham, J.M.; Zhao, Q.; Jane, S.M.; Ottolenghi, S.; Ronchi, A. Functional Interaction of CP2 with GATA-1 in the Regulation of Erythroid Promoters. Mol. Cell. Biol. 2006, 26, 3942–3954. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Crossley, M.; Tsang, A.; Bieker, J.; Orkin, S. Regulation of the erythroid Kruppel-like factor (EKLF) gene promoter by the erythroid transcription factor GATA-1. J. Biol. Chem. 1994, 269, 15440–15444. [Google Scholar] [CrossRef]
- Lecointe, N.; Bernard, O.; Naert, K.; Joulin, V.; Larsen, C.J.; Romeo, P.-H.; Mathieu-Mahul, D. GATA-and SP1-binding sites are required for the full activity of the tissue-specific promoter of the tal-1 gene. Oncogene 1994, 9, 2623–2632. [Google Scholar]
- Cheng, Y.; Wu, W.; Kumar, S.A.; Yu, D.; Deng, W.; Tripic, T.; King, D.C.; Chen, K.-B.; Zhang, Y.; Drautz, D.; et al. Erythroid GATA1 function revealed by genome-wide analysis of transcription factor occupancy, histone modifications, and mRNA expression. Genome Res. 2009, 19, 2172–2184. [Google Scholar] [CrossRef] [Green Version]
- Soler, E.; Andrieu-Soler, C.; De Boer, E.; Bryne, J.C.; Thongjuea, S.; Stadhouders, R.; Palstra, R.-J.; Stevens, M.; Kockx, C.; van Ijcken, W.; et al. The genome-wide dynamics of the binding of Ldb1 complexes during erythroid differentiation. Genes Dev. 2010, 24, 277–289. [Google Scholar] [CrossRef] [Green Version]
- Pop, R.; Shearstone, J.R.; Shen, Q.; Liu, Y.; Hallstrom, K.; Koulnis, M.; Gribnau, J.; Socolovsky, M. A key commitment step in erythropoiesis is synchronized with the cell cycle clock through mutual inhibition between PU.1 and S-phase progression. PLoS Biol. 2010, 8, e1000484. [Google Scholar] [CrossRef] [Green Version]
- Daftari, P.; Gvava, N.R.; Shen, C.K.J. Distinction between AP1 and NF-E2 factor-binding at specific chromtin regions in mammalian cells. Oncogene 1999, 18, 5482–5486. [Google Scholar] [CrossRef] [Green Version]
- Mueller, P.R.; Wold, B. In vivo footprinting of a muscle specific enhancer by ligation mediated PCR. Science 1989, 246, 780–786. [Google Scholar] [CrossRef] [PubMed]
- Pfeifer, G.; Steigerwald, S.; Mueller, P.; Wold, B.; Riggs, A. Genomic sequencing and methylation analysis by ligation mediated PCR. Science 1989, 246, 810–813. [Google Scholar] [CrossRef]
- Zhang, Q.; Reddy, P.M.S.; Yu, C.-Y.; Bastiani, C.; Higgs, D.; Stamatoyannopoulos, G.; Papayannopoulou, T.; Shen, C.K.J. Transcriptional activation of human zeta 2 globin promoter by the alpha globin regulatory element (HS-40): Functional role of specific nuclear factor-DNA complexes. Mol. Cell. Biol. 1993, 13, 2298–2308. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wen, S.-C.; Roder, K.; Hu, K.-Y.; Rombel, I.; Gavva, N.R.; Daftari, P.; Kuo, Y.-Y.; Wang, C.; Shen, C.-K.J. Loading of DNA-binding factors to an erythroid enhancer. Mol. Cell. Biol. 2000, 20, 1993–2003. [Google Scholar] [CrossRef] [PubMed] [Green Version]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hung, C.-H.; Lee, T.-L.; Huang, A.Y.-S.; Yang, K.-C.; Shyu, Y.-C.; Wen, S.-C.; Lu, M.-J.; Yuan, S.; Shen, C.-K.J. A Positive Regulatory Feedback Loop between EKLF/KLF1 and TAL1/SCL Sustaining the Erythropoiesis. Int. J. Mol. Sci. 2021, 22, 8024. https://doi.org/10.3390/ijms22158024
Hung C-H, Lee T-L, Huang AY-S, Yang K-C, Shyu Y-C, Wen S-C, Lu M-J, Yuan S, Shen C-KJ. A Positive Regulatory Feedback Loop between EKLF/KLF1 and TAL1/SCL Sustaining the Erythropoiesis. International Journal of Molecular Sciences. 2021; 22(15):8024. https://doi.org/10.3390/ijms22158024
Chicago/Turabian StyleHung, Chun-Hao, Tung-Liang Lee, Anna Yu-Szu Huang, Kang-Chung Yang, Yu-Chiau Shyu, Shau-Ching Wen, Mu-Jie Lu, Shinsheng Yuan, and Che-Kun James Shen. 2021. "A Positive Regulatory Feedback Loop between EKLF/KLF1 and TAL1/SCL Sustaining the Erythropoiesis" International Journal of Molecular Sciences 22, no. 15: 8024. https://doi.org/10.3390/ijms22158024






