Genome-Wide Identification, Expansion Mechanism and Expression Profiling Analysis of GLABROUS1 Enhancer-Binding Protein (GeBP) Gene Family in Gramineae Crops
Abstract
:1. Introduction
2. Results
2.1. Identification, Phylogenetic Analysis, and Classification in the Gramineae Crops
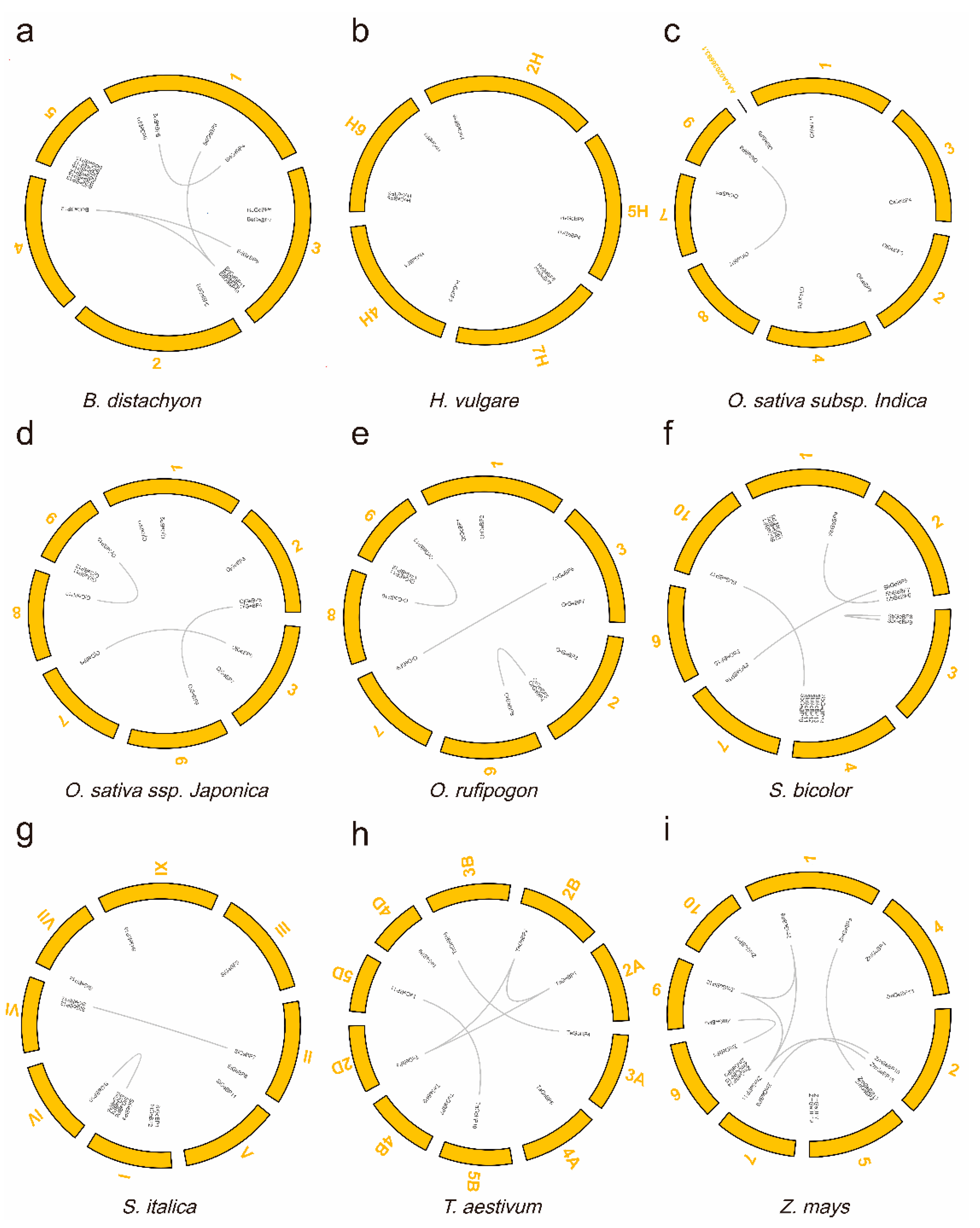
2.2. The Expansion and Evolutionary Pattern of the GeBP Genes
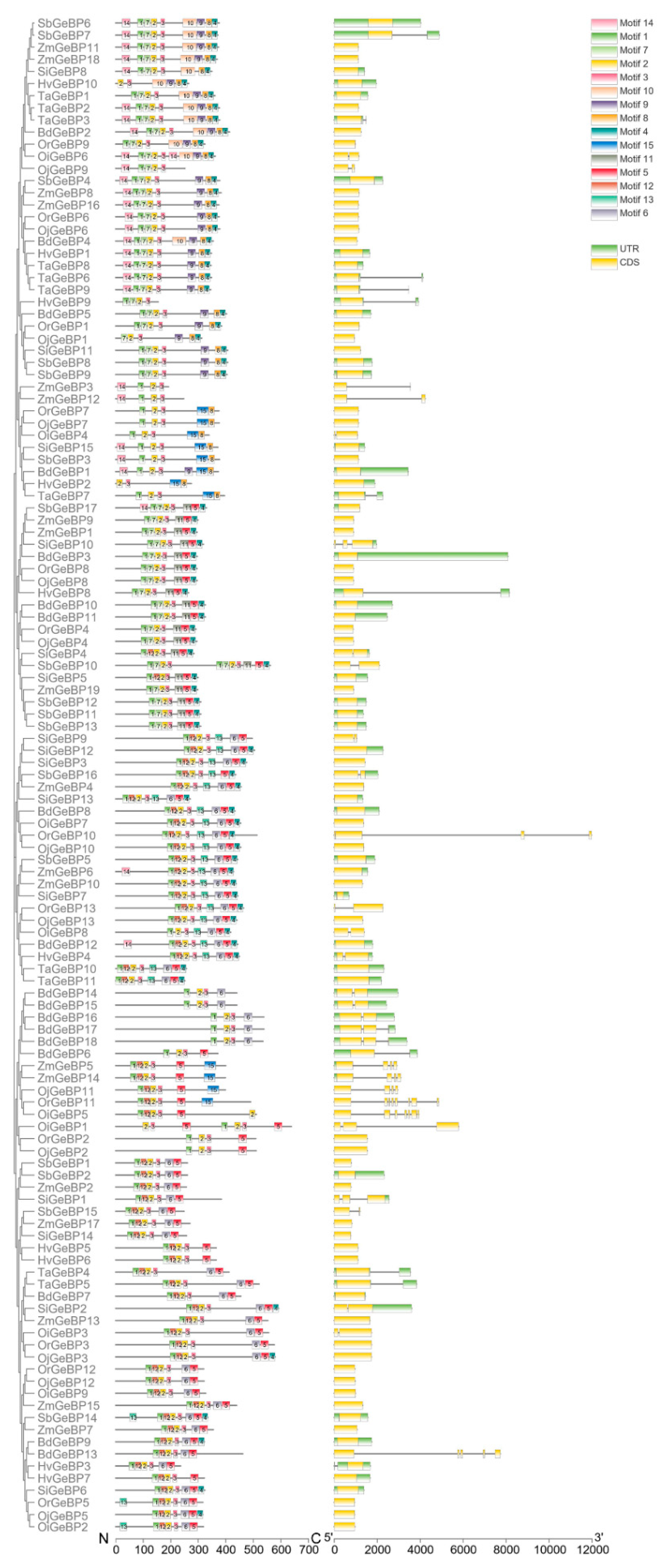
2.3. The Conserved Motif and Gene Structure Analysis
2.4. Expression Profiles of the GeBP Genes across Different Rice Tissues and Developmental Stages
2.5. Expression Profiles of the GeBP Genes in Rice under Various Hormonal Stresses
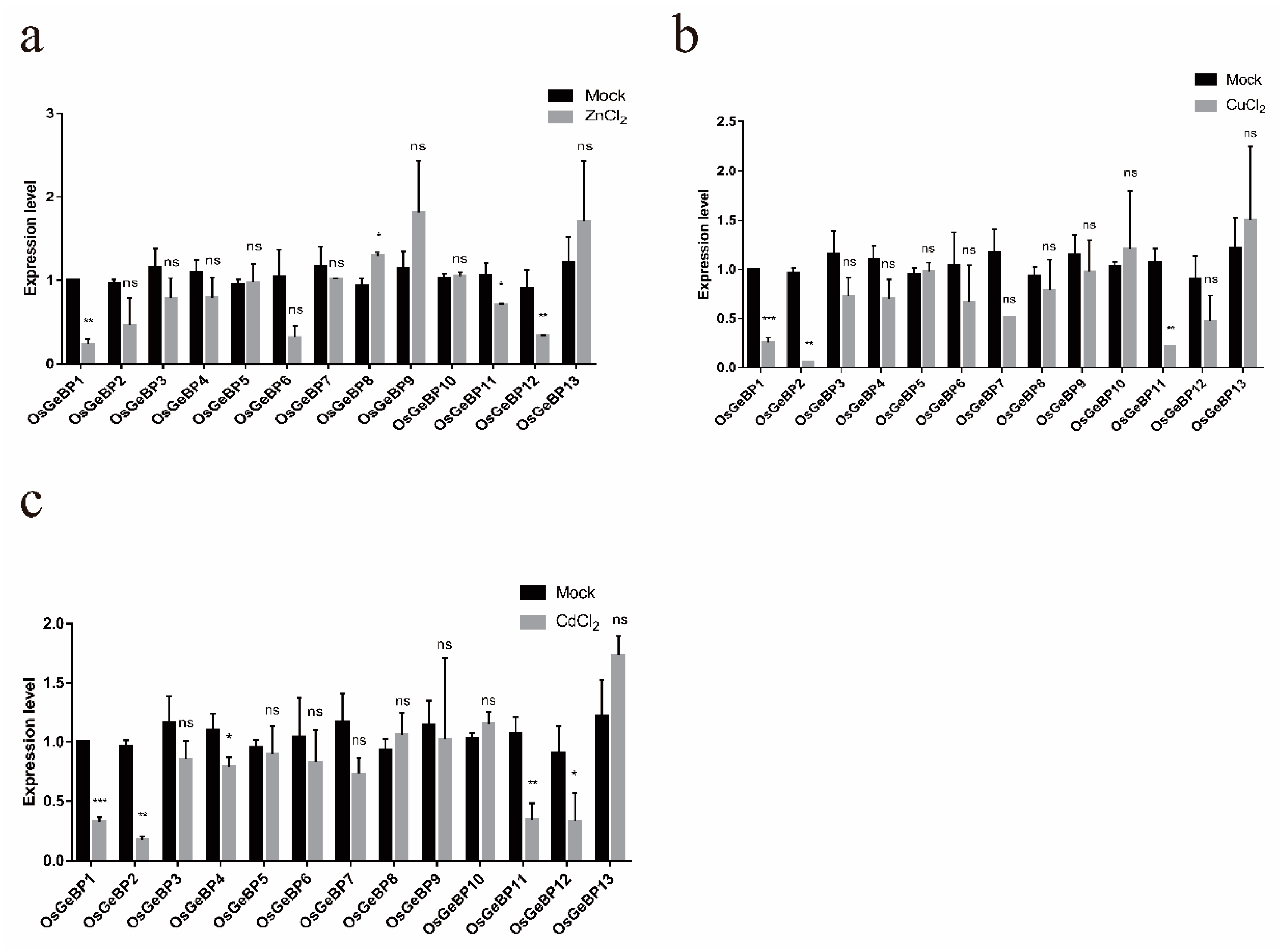
2.6. Expression Profiles of the GeBP Genes in Rice under Various Metal Ion Stresses
2.7. The Subcellular Localization of OsGeBP11 and OsGeBP12 in Rice
3. Materials and Methods
3.1. Plant Materials and Growth Conditions
3.2. Hormone and Metal Ion Stress Treatments
3.3. Genome-Wide Identification of the GeBP Genes in Gramineae Crops
3.4. Phylogenetic Relationship Analyses
3.5. Duplication Events, Orthologous Groups, Conserved Motifs, and Gene Structure Analyses
3.6. RNA Extraction, cDNA Synthesis, and Quantitative Real-Time PCR
3.7. Transient Expression of the Enhanced Green Fluorescent Protein (eGFP) Constructs in the Rice Protoplast
3.8. Statistical Analysis
4. Discussion
4.1. Characterization of the GeBP Genes in Gramineae Crops
4.2. Expression Profiles of the GeBP Genes in Gramineae Crops
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| GeBP | GLABROUS1 enhancer-binding protein |
| TF | Transcription factor |
| GA | Gibberellin |
| CK | Cytokinin |
| cDNA | Complementary DNA |
| NJ | Neighbor-joining |
| Ka/Ks | nonsynonymous (Ka) /Synonymous (Ks) substitution |
| qRT-PCR | Quantitative real-time PCR |
| Bd | Brachypodium distachyon |
| Hv | Hordeum vulgare |
| Oi | Oryza. sativa ssp.indica |
| Oj | Oryza. sativa ssp. japonica |
| Or | Oryza rufipogon |
| Sb | Sorghum bicolor |
| Si | Setaria italica |
| Ta | Triticum aestivum |
| Zm | Zea mays |
| WGD | Whole genome duplication |
| MYA | Million years |
| OGs | Orthologous groups |
References
- Kim, T.-W.; Wang, Z.-Y. Brassinosteroid Signal Transduction from Receptor Kinases to Transcription Factors. Annu. Rev. Plant Biol. 2010, 61, 681–704. [Google Scholar] [CrossRef] [Green Version]
- Yoon, J.; Cho, L.-H.; Yang, W.; Pasriga, R.; Wu, Y.; Hong, W.-J.; Bureau, C.; Wi, S.J.; Zhang, T.; Wang, R.; et al. Homeobox transcription factor OsZHD2 promotes root meristem activity in rice by inducing ethylene biosynthesis. J. Exp. Bot. 2020, 71, 5348–5364. [Google Scholar] [CrossRef]
- Liu, C.; Ou, S.; Mao, B.; Tang, J.; Wang, W.; Wang, H.; Cao, S.; Schläppi, M.R.; Zhao, B.; Xiao, G.; et al. Early selection of bZIP73 facilitated adaptation of japonica rice to cold climates. Nat. Commun. 2018, 9, 1–12. [Google Scholar] [CrossRef]
- Hu, Y.; Han, X.; Yang, M.; Zhang, M.; Pan, J.; Yu, D. The Transcription Factor INDUCER OF CBF EXPRESSION1 Interacts with ABSCISIC ACID INSENSITIVE5 and DELLA Proteins to Fine-Tune Abscisic Acid Signaling during Seed Germination in Arabidopsis. Plant Cell 2019, 31, 1520–1538. [Google Scholar] [CrossRef]
- Jin, J.; Zhang, H.; Kong, L.; Gao, G.; Luo, J. PlantTFDB 3.0: A portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res. 2014, 42, D1182–D1187. [Google Scholar] [CrossRef] [Green Version]
- Riechmann, J.L.; Heard, J.; Martin, G.; Reuber, L.Z.-C.; Jiang, C.-Z.; Keddie, J.; Adam, L.; Pineda, O.; Ratcliffe, O.J.; Samaha, R.R.; et al. Arabidopsis Transcription Factors: Genome-Wide Comparative Analysis Among Eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar] [CrossRef] [PubMed]
- Curaba, J.; Herzog, M.; Vachon, G. GeBP, the first member of a new gene family inArabidopsis, encodes a nuclear protein with DNA-binding activity and is regulated byKNAT1. Plant J. 2003, 33, 305–317. [Google Scholar] [CrossRef] [PubMed]
- Hülskamp, M.; Miséra, S.; Jürgens, G. Genetic dissection of trichome cell development in Arabidopsis. Cell 1994, 76, 555–566. [Google Scholar] [CrossRef]
- Hauser, M.-T.; Harr, B.; Schlötterer, C. Trichome Distribution in Arabidopsis thaliana and its Close Relative Arabidopsis lyrata: Molecular Analysis of the Candidate Gene GLABROUS1. Mol. Biol. Evol. 2001, 18, 1754–1763. [Google Scholar] [CrossRef] [Green Version]
- Perazza, D.; Vachon, G.; Herzog, M. Gibberellins Promote Trichome Formation by Up-RegulatingGLABROUS1 in Arabidopsis1. Plant Physiol. 1998, 117, 375–383. [Google Scholar] [CrossRef] [Green Version]
- Dill, A.; Jung, H.-S.; Sun, T.-P. The DELLA motif is essential for gibberellin-induced degradation of RGA. Proc. Natl. Acad. Sci. USA 2001, 98, 14162–14167. [Google Scholar] [CrossRef] [Green Version]
- Ferreira, F.J.; Kieber, J.J. Cytokinin signaling. Curr. Opin. Plant Biol. 2005, 8, 518–525. [Google Scholar] [CrossRef] [PubMed]
- Hay, A.; Kaur, H.; Phillips, A.; Hedden, P.; Hake, S.; Tsiantis, M. The Gibberellin Pathway Mediates KNOTTED1-Type Homeobox Function in Plants with Different Body Plans. Curr. Biol. 2002, 12, 1557–1565. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Z.; An, L.; Sun, L.; Zhu, S.; Xi, W.; Broun, P.; Yu, H.; Gan, Y. Zinc Finger Protein5 Is Required for the Control of Trichome Initiation by Acting Upstream of Zinc Finger Protein8 in Arabidopsis. Plant Physiol. 2011, 157, 673–682. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Z.; An, L.; Sun, L.; Gan, Y. ZFP5encodes a functionally equivalentGISprotein to control trichome initiation. Plant Signal. Behav. 2012, 7, 28–30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jasinski, S.; Piazza, P.; Craft, J.; Hay, A.; Woolley, L.; Rieu, I.; Phillips, A.; Hedden, P.; Tsiantis, M. KNOX Action in Arabidopsis Is Mediated by Coordinate Regulation of Cytokinin and Gibberellin Activities. Curr. Biol. 2005, 15, 1560–1565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yanai, O.; Shani, E.; Dolezal, K.; Tarkowski, P.; Sablowski, R.; Sandberg, G.; Samach, A.; Ori, N. Arabidopsis KNOXI Proteins Activate Cytokinin Biosynthesis. Curr. Biol. 2005, 15, 1566–1571. [Google Scholar] [CrossRef] [Green Version]
- Chevalier, F.; Perazza, D.; Laporte, F.; Le Hénanff, G.; Hornitschek, P.; Bonneville, J.-M.; Herzog, M.; Vachon, G. GeBP and GeBP-Like Proteins Are Noncanonical Leucine-Zipper Transcription Factors That Regulate Cytokinin Response in Arabidopsis. Plant Physiol. 2008, 146, 1142–1154. [Google Scholar] [CrossRef] [Green Version]
- Chien, J.C.; Sussex, I.M. Differential Regulation of Trichome Formation on the Adaxial and Abaxial Leaf Surfaces by Gibberellins and Photoperiod in Arabidopsis thaliana (L.) Heynh. Plant Physiol. 1996, 111, 1321–1328. [Google Scholar] [CrossRef] [Green Version]
- Ogas, J.; Cheng, J.-C.; Sung, Z.R.; Somerville, C. Cellular Differentiation Regulated by Gibberellin in theArabidopsis thaliana pickleMutant. Science 1997, 277, 91–94. [Google Scholar] [CrossRef]
- Khare, D.; Mitsuda, N.; Lee, S.; Song, W.; Hwang, D.; Ohme-Takagi, M.; Martinoia, E.; Lee, Y.; Hwang, J. Root avoidance of toxic metals requires the GeBP-LIKE 4 transcription factor in Arabidopsis thaliana. New Phytol. 2017, 213, 1257–1273. [Google Scholar] [CrossRef] [Green Version]
- Clemens, S. Toxic metal accumulation, responses to exposure and mechanisms of tolerance in plants. Biochimie 2006, 88, 1707–1719. [Google Scholar] [CrossRef]
- A Bowling, S.; Guo, A.; Cao, H.; Gordon, A.S.; Klessig, D.F.; Dong, X. A mutation in Arabidopsis that leads to constitutive expression of systemic acquired resistance. Plant Cell 1994, 6, 1845–1857. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brininstool, G.; Kasili, R.; Simmons, L.A.; Kirik, V.; Hülskamp, M.; Larkin, J.C. Constitutive Expressor of Pathogenesis-Related Genes5 affects cell wall biogenesis and trichome development. BMC Plant Biol. 2008, 8, 58. [Google Scholar] [CrossRef] [Green Version]
- Jing, H.-C.; Sturre, M.J.G.; Hille, J.; Dijkwel, P.P. Arabidopsis onset of leaf deathmutants identify a regulatory pathway controlling leaf senescence. Plant J. 2002, 32, 51–63. [Google Scholar] [CrossRef] [Green Version]
- Perazza, D.; Laporte, F.; Balagué, C.; Chevalier, F.; Remo, S.; Bourge, M.; Larkin, J.; Herzog, M.; Vachon, G. GeBP/GPL Transcription Factors Regulate a Subset of CPR5-Dependent Processes. Plant Physiol. 2011, 157, 1232–1242. [Google Scholar] [CrossRef] [Green Version]
- García-Cano, E.; Hak, H.; Magori, S.; Lazarowitz, S.G.; Citovsky, V. The Agrobacterium F-Box Protein Effector VirF Destabilizes the Arabidopsis GLABROUS1 Enhancer/Binding Protein-Like Transcription Factor VFP4, a Transcriptional Activator of Defense Response Genes. Mol. Plant Microbe Interact. 2018, 31, 576–586. [Google Scholar] [CrossRef] [Green Version]
- Tian, Z.; Shen, G.; Yuan, G.; Song, K.; Lu, J.; Da, L. Effects of Echinochloa crusgalli and Cyperus difformis on yield and eco-economic thresholds of rice. J. Clean. Prod. 2020, 259, 120807. [Google Scholar] [CrossRef]
- Wang, W.; Mauleon, R.; Hu, Z.; Chebotarov, D.; Tai, S.; Wu, Z.; Li, M.; Zheng, T.; Fuentes, R.R.; Zhang, F.; et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 2018, 557, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Londo, J.; Chiang, Y.-C.; Hung, K.-H.; Chiang, T.-Y.; Schaal, B.A. Phylogeography of Asian wild rice, Oryza rufipogon, reveals multiple independent domestications of cultivated rice, Oryza sativa. Proc. Natl. Acad. Sci. USA 2006, 103, 9578–9583. [Google Scholar] [CrossRef] [Green Version]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [Green Version]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef]
- Kong, W.; Zhong, H.; Deng, X.; Gautam, M.; Gong, Z.; Zhang, Y.; Zhao, G.; Liu, C.; Li, Y. Evolutionary Analysis of GH3 Genes in Six Oryza Species/Subspecies and Their Expression under Salinity Stress in Oryza sativa ssp. japonica. Plants 2019, 8, 30. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Deng, X.; An, B.; Zhong, H.; Yang, J.; Kong, W.; Li, Y. A Novel Insight into Functional Divergence of the MST Gene Family in Rice Based on Comprehensive Expression Patterns. Genes 2019, 10, 239. [Google Scholar] [CrossRef] [Green Version]
- Kong, W.; An, B.; Zhang, Y.; Yang, J.; Li, S.; Sun, T.; Li, Y. Sugar Transporter Proteins (STPs) in Gramineae Crops: Comparative Analysis, Phylogeny, Evolution, and Expression Profiling. Cells 2019, 8, 560. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, T.; Zeng, L.; Chen, X.; Rong, H.; Wu, J.; Batley, J.; Jiang, J.; Wang, Y. Genome-Wide Analysis of the Lateral Organ Boundaries Domain Gene Family in Brassica Napus. Genes 2020, 11, 280. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Wang, K.; Huang, W.; Liu, G.; Gao, Y.; Wang, J.; Huang, Q.; Ji, Y.; Qin, X.; Wan, L.; et al. The Rice Pentatricopeptide Repeat Protein RF5 Restores Fertility in Hong-Lian Cytoplasmic Male-Sterile Lines via a Complex with the Glycine-Rich Protein GRP162. Plant Cell 2012, 24, 109–122. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiao, H.; Zhang, Q.; Qin, X.; Xu, Y.; Ni, C.; Huang, J.; Zhu, L.; Zhong, F.; Liu, W.; Yao, G.; et al. Rice PPS1 encodes a DYW motif-containing pentatricopeptide repeat protein required for five consecutive RNA-editing sites of nad3 in mitochondria. New Phytol. 2018, 220, 878–892. [Google Scholar] [CrossRef] [Green Version]
- Qin, X.; Huang, Q.; Xiao, H.; Zhang, Q.; Ni, C.; Xu, Y.; Liu, G.; Yang, D.; Zhu, Y.; Hu, J. The rice DUF 1620-containing and WD 40-like repeat protein is required for the assembly of the restoration of fertility complex. New Phytol. 2016, 210, 934–945. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Swigoňová, Z.; Lai, J.; Ma, J.; Ramakrishna, W.; Llaca, V.; Bennetzen, J.L.; Messing, J. Close Split of Sorghum and Maize Genome Progenitors. Genome Res. 2004, 14, 1916–1923. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pickett, F.B.; Meeks-Wagner, D.R. Seeing double: Appreciating genetic redundancy. Plant Cell 1995, 7, 1347–1356. [Google Scholar] [CrossRef] [PubMed]
- Di Cristina, M.; Sessa, G.; Dolan, L.; Linstead, P.; Baima, S.; Ruberti, I.; Morelli, G. The Arabidopsis Athb-10 (GLABRA2) is an HD-Zip protein required for regulation of root hair development. Plant J. 1996, 10, 393–402. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Shen, J.; Zhang, L.; Qi, H.; Yang, L.; Wang, H.; Wang, J.; Wang, Y.; Du, H.; Tao, Z.; et al. Nuclear translocation of OsMFT1 that is impeded by OsFTIP1 promotes drought tolerance in rice. Mol. Plant 2021, 14, 1297–1311. [Google Scholar] [CrossRef]




| Seq_1 | Seq_2 | Ka | Ks | Ka/Ks | Duplication Type | Date (MYA) | Purifying Selection |
|---|---|---|---|---|---|---|---|
| BdGeBP10 | BdGeBP3 | 0.2840 | 0.5749 | 0.4940 | WGD or segmental duplication | 15.61 | YES |
| BdGeBP12 | BdGeBP10 | 0.8732 | 1.1050 | 0.7902 | WGD or segmental duplication | 47.98 | YES |
| BdGeBP12 | BdGeBP8 | 0.3481 | 0.7242 | 0.4807 | WGD or segmental duplication | 19.13 | YES |
| BdGeBP2 | BdGeBP4 | 0.2667 | 0.5949 | 0.4483 | WGD or segmental duplication | 14.65 | YES |
| OiGeBP7 | OiGeBP8 | 0.3827 | 0.7380 | 0.5185 | WGD or segmental duplication | 21.03 | YES |
| OjGeBP4 | OjGeBP8 | 0.3516 | 0.5702 | 0.6166 | WGD or segmental duplication | 19.32 | YES |
| OjGeBP6 | OjGeBP9 | 0.2728 | 0.5102 | 0.5346 | WGD or segmental duplication | 14.99 | YES |
| OjGeBP10 | OjGeBP13 | 0.3549 | 0.6963 | 0.5097 | WGD or segmental duplication | 19.50 | YES |
| OrGeBP6 | OrGeBP9 | 0.2853 | 0.6970 | 0.4093 | WGD or segmental duplication | 15.67 | YES |
| OrGeBP4 | OrGeBP8 | 0.3498 | 0.5667 | 0.6172 | WGD or segmental duplication | 19.22 | YES |
| OrGeBP10 | OrGeBP13 | 0.3429 | 0.7032 | 0.4876 | WGD or segmental duplication | 18.84 | YES |
| SbGeBP17 | SbGeBP10 | 0.4878 | 1.0186 | 0.4789 | WGD or segmental duplication | 26.80 | YES |
| SbGeBP4 | SbGeBP6 | 0.3067 | 0.8060 | 0.3806 | WGD or segmental duplication | 16.85 | YES |
| SbGeBP5 | SbGeBP16 | 0.3966 | 0.8534 | 0.4647 | WGD or segmental duplication | 21.79 | YES |
| SbGeBP8 | SbGeBP9 | 0.0045 | 0.0359 | 0.1244 | WGD or segmental duplication | 0.25 | YES |
| SiGeBP10 | SiGeBP4 | 0.4122 | 0.6411 | 0.6430 | WGD or segmental duplication | 22.65 | YES |
| SiGeBP12 | SiGeBP7 | 0.4039 | 0.7101 | 0.5687 | WGD or segmental duplication | 22.19 | YES |
| TaGeBP10 | TaGeBP11 | 0.0035 | 0.0843 | 0.0410 | WGD or segmental duplication | 0.19 | YES |
| TaGeBP1 | TaGeBP2 | 0.0739 | 0.1727 | 0.4280 | WGD or segmental duplication | 4.06 | YES |
| TaGeBP1 | TaGeBP3 | 0.0758 | 0.1790 | 0.4233 | WGD or segmental duplication | 4.16 | YES |
| TaGeBP2 | TaGeBP3 | 0.0107 | 0.0813 | 0.1319 | WGD or segmental duplication | 0.59 | YES |
| TaGeBP4 | TaGeBP5 | 0.0993 | 0.1508 | 0.6587 | WGD or segmental duplication | 5.46 | YES |
| ZmGeBP10 | ZmGeBP6 | 0.0599 | 0.2385 | 0.2510 | WGD or segmental duplication | 3.29 | YES |
| ZmGeBP11 | ZmGeBP16 | 0.3334 | 0.6935 | 0.4808 | WGD or segmental duplication | 18.32 | YES |
| ZmGeBP11 | ZmGeBP8 | 0.3210 | 0.8094 | 0.3966 | WGD or segmental duplication | 17.64 | YES |
| ZmGeBP12 | ZmGeBP3 | 0.0340 | 0.1367 | 0.2487 | WGD or segmental duplication | 1.87 | YES |
| ZmGeBP18 | ZmGeBP11 | 0.0306 | 0.1401 | 0.2183 | WGD or segmental duplication | 1.68 | YES |
| ZmGeBP1 | ZmGeBP9 | 0.0418 | 0.1690 | 0.2474 | WGD or segmental duplication | 2.30 | YES |
| ZmGeBP8 | ZmGeBP16 | 0.0377 | 0.1678 | 0.2248 | WGD or segmental duplication | 2.07 | YES |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, J.; Zhang, Q.; He, Y.; Liu, W.; Xu, Y.; Liu, K.; Xian, F.; Li, J.; Hu, J. Genome-Wide Identification, Expansion Mechanism and Expression Profiling Analysis of GLABROUS1 Enhancer-Binding Protein (GeBP) Gene Family in Gramineae Crops. Int. J. Mol. Sci. 2021, 22, 8758. https://doi.org/10.3390/ijms22168758
Huang J, Zhang Q, He Y, Liu W, Xu Y, Liu K, Xian F, Li J, Hu J. Genome-Wide Identification, Expansion Mechanism and Expression Profiling Analysis of GLABROUS1 Enhancer-Binding Protein (GeBP) Gene Family in Gramineae Crops. International Journal of Molecular Sciences. 2021; 22(16):8758. https://doi.org/10.3390/ijms22168758
Chicago/Turabian StyleHuang, Jishuai, Qiannan Zhang, Yurong He, Wei Liu, Yanghong Xu, Kejia Liu, Fengjun Xian, Junde Li, and Jun Hu. 2021. "Genome-Wide Identification, Expansion Mechanism and Expression Profiling Analysis of GLABROUS1 Enhancer-Binding Protein (GeBP) Gene Family in Gramineae Crops" International Journal of Molecular Sciences 22, no. 16: 8758. https://doi.org/10.3390/ijms22168758





