The Drosophila miR-959–962 Cluster Members Repress Toll Signaling to Regulate Antibacterial Defense during Bacterial Infection
Abstract
:1. Introduction
2. Results
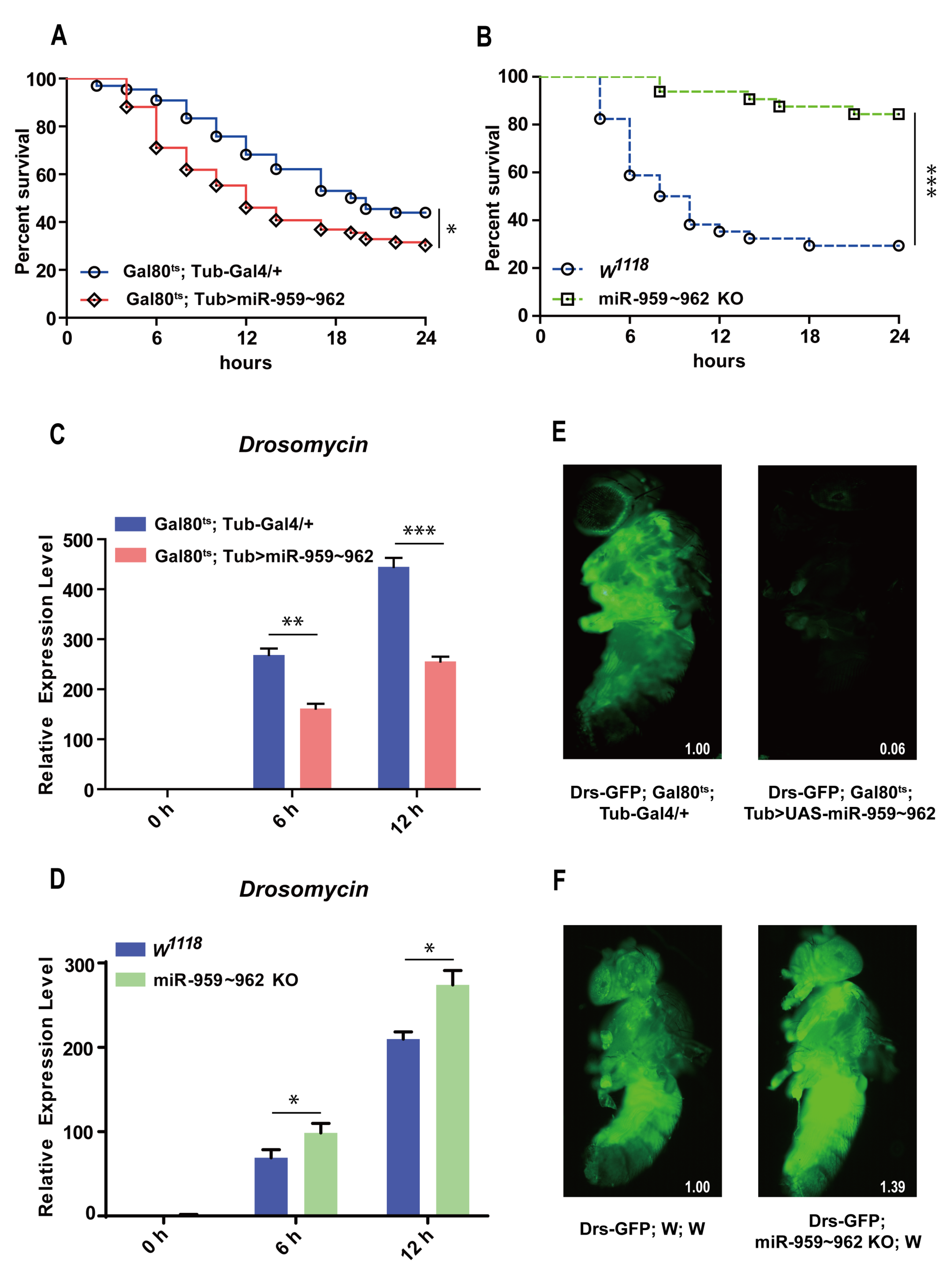
2.1. The miR-959–962 Cluster Could Negatively Regulate Drosophila Toll-Related Immune Response
2.2. Each Member from miR-959–962 Cluster Plays a Negative Regulatory Role in Drosophila Toll Pathway
2.3. The Immune-Related Genes Are Potentially Targeted by miRNA Members from miR-959–962 Cluster In Vitro
2.4. The miR-959–962 Member Simultaneously or Serperately Target Key Components of Toll Pathway (Tube, dl, and Toll) In Vivo
2.5. Dynamic Expression Patterns of miR-959–962 Cluster Members in Wild-Type Flies after M. luteus or PBS Infection
3. Discussion
4. Materials and Methods
4.1. Drosophila Stocks and Husbandry
4.2. Adult Immune Challenge
4.3. Quantitative RT–PCR Analysis
4.4. miRNA Targets Prediction
4.5. Recombinant Plasmids Generation
4.6. Cell Transfection and Luciferase Assays
4.7. Data Processing and Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| miRNA | microRNA |
| Drs | Drosomycin |
| dl | dorsal |
| 3′ UTR | 3′ untranslated region |
| imd | immune deficiency |
| AMP | antimicrobial peptide |
| M. luteus | Micrococcus luteus |
| E. faecalis | Enterococcus faecalis |
| Drs-GFP | Drosomycin–green fluorescent protein |
References
- Ayyar, K.K.; Reddy, K.V.R. MAPK and NF-kappaB signalling pathways regulate the expression of miRNA, let-7f in human endocervical epithelial cells. J. Cell. Biochem. 2018, 119, 4751–4759. [Google Scholar] [CrossRef] [PubMed]
- Parkin, J.; Cohen, B. An overview of the immune system. Lancet 2001, 357, 1777–1789. [Google Scholar] [CrossRef]
- Kounatidis, I.; Ligoxygakis, P. Drosophila as a model system to unravel the layers of innate immunity to infection. Open Biol. 2012, 2, 120075. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoffmann, J.A. The immune response of Drosophila. Nature 2003, 426, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Lemaitre, B.; Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 2007, 25, 697–743. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hultmark, D. Drosophila immunity: Paths and patterns. Curr. Opin. Immunol. 2003, 15, 12–19. [Google Scholar] [CrossRef]
- Valanne, S.; Wang, J.H.; Ramet, M. The Drosophila Toll signaling pathway. J. Immunol. 2011, 186, 649–656. [Google Scholar] [CrossRef] [Green Version]
- Michel, T.; Reichhart, J.M.; Hoffmann, J.A.; Royet, J. Drosophila Toll is activated by Gram-positive bacteria through a circulating peptidoglycan recognition protein. Nature 2001, 414, 756–759. [Google Scholar] [CrossRef]
- Bischoff, V.; Vignal, C.; Boneca, I.G.; Michel, T.; Hoffmann, J.A.; Royet, J. Function of the drosophila pattern-recognition receptor PGRP-SD in the detection of Gram-positive bacteria. Nat. Immunol. 2004, 5, 1175–1180. [Google Scholar] [CrossRef]
- Wang, L.; Weber, A.N.; Atilano, M.L.; Filipe, S.R.; Gay, N.J.; Ligoxygakis, P. Sensing of Gram-positive bacteria in Drosophila: GNBP1 is needed to process and present peptidoglycan to PGRP-SA. EMBO J. 2006, 25, 5005–5014. [Google Scholar] [CrossRef] [Green Version]
- Gottar, M.; Gobert, V.; Matskevich, A.A.; Reichhart, J.M.; Wang, C.; Butt, T.M.; Belvin, M.; Hoffmann, J.A.; Ferrandon, D. Dual detection of fungal infections in Drosophila via recognition of glucans and sensing of virulence factors. Cell 2006, 127, 1425–1437. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weber, A.N.; Delamasure, S.T.; Hoffmann, J.A.; Lelievre, E.; Gascan, H.; Ray, K.P.; Morse, M.A.; Imler, J.L.; Gay, N.J. Binding of the Drosophila cytokine Spatzle to Toll is direct and establishes signaling. Nat. Immunol. 2003, 4, 794–800. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Yagi, Y.; Tanji, T.; Zhou, S.; Ip, Y.T. Multimerization and interaction of Toll and Spatzle in Drosophila. Proc. Natl. Acad. Sci. USA 2004, 101, 9369–9374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morisato, D.; Anderson, K.V. The spatzle gene encodes a component of the extracellular signaling pathway establishing the dorsal-ventral pattern of the Drosophila embryo. Cell 1994, 76, 677–688. [Google Scholar] [CrossRef]
- Schneider, D.S.; Jin, Y.; Morisato, D.; Anderson, K.V. A processed form of the Spatzle protein defines dorsal-ventral polarity in the Drosophila embryo. Development 1994, 120, 1243–1250. [Google Scholar]
- Delamasure, S.T.; Bilak, H.; Capovilla, M.; Hoffmann, J.A.; Imler, J.L. Drosophila MyD88 is required for the response to fungal and Gram-positive bacterial infections. Nat. Immunol. 2002, 3, 91–97. [Google Scholar] [CrossRef]
- Horng, T.; Medzhitov, R. Drosophila MyD88 is an adapter in the Toll signaling pathway. Proc. Natl. Acad. Sci. USA 2001, 98, 12654–12658. [Google Scholar] [CrossRef] [Green Version]
- Xiao, T.; Towb, P.; Wasserman, S.A.; Sprang, S.R. Three-dimensional structure of a complex between the death domains of Pelle and Tube. Cell 1999, 99, 545–555. [Google Scholar] [CrossRef] [Green Version]
- Ip, Y.T.; Reach, M.; Engstrom, Y.; Kadalayil, L.; Cai, H.; Gonzalez-Crespo, S.; Tatei, K.; Levine, M. Dif, a dorsal-related gene that mediates an immune response in Drosophila. Cell 1993, 75, 753–763. [Google Scholar] [CrossRef]
- Lemaitre, B.; Meister, M.; Govind, S.; Georgel, P.; Steward, R.; Reichhart, J.M.; Hoffmann, J.A. Functional analysis and regulation of nuclear import of dorsal during the immune response in Drosophila. EMBO J. 1995, 14, 536–545. [Google Scholar] [CrossRef] [Green Version]
- Wu, L.P.; Anderson, K.V. Regulated nuclear import of Rel proteins in the Drosophila immune response. Nature 1998, 392, 93–97. [Google Scholar] [CrossRef] [PubMed]
- Lemaitre, B.; Reichhart, J.M.; Hoffmann, J.A. Drosophila host defense: Differential induction of antimicrobial peptide genes after infection by various classes of microorganisms. Proc. Natl. Acad. Sci. USA 1997, 94, 14614–14619. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hetru, C.; Hoffmann, J.A. NF-kappaB in the immune response of Drosophila. Cold Spring Harb. Perspect. Biol. 2009, 1, a000232. [Google Scholar] [CrossRef] [PubMed]
- Myllymaki, H.; Valanne, S.; Ramet, M. The Drosophila imd signaling pathway. J. Immunol. 2014, 192, 3455–3462. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kambris, Z.; Brun, S.; Jang, I.H.; Nam, H.J.; Romeo, Y.; Takahashi, K.; Lee, W.J.; Ueda, R.; Lemaitre, B. Drosophila immunity: A large-scale in vivo RNAi screen identifies five serine proteases required for Toll activation. Curr. Biol. 2006, 16, 808–813. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chamy, L.E.; Leclerc, V.; Caldelari, I.; Reichhart, J.M. Sensing of ‘danger signals’ and pathogen-associated molecular patterns defines binary signaling pathways ‘upstream’ of Toll. Nat. Immunol. 2008, 9, 1165–1170. [Google Scholar] [CrossRef] [Green Version]
- Haghayeghi, A.; Sarac, A.; Czerniecki, S.; Grosshans, J.; Schock, F. Pellino enhances innate immunity in Drosophila. Mech. Dev. 2010, 127, 301–307. [Google Scholar] [CrossRef]
- Valanne, S.; Myllymaki, H.; Kallio, J.; Schmid, M.R.; Kleino, A.; Murumagi, A.; Airaksinen, L.; Kotipelto, T.; Kaustio, M.; Ulvila, J.; et al. Genome-wide RNA interference in Drosophila cells identifies G protein-coupled receptor kinase 2 as a conserved regulator of NF-kappaB signaling. J. Immunol. 2010, 184, 6188–6198. [Google Scholar] [CrossRef] [Green Version]
- Reed, D.E.; Huang, X.M.; Wohlschlegel, J.A.; Levine, M.S.; Senger, K. DEAF-1 regulates immunity gene expression in Drosophila. Proc. Natl. Acad. Sci. USA 2008, 105, 8351–8356. [Google Scholar] [CrossRef] [Green Version]
- Gordon, M.D.; Dionne, M.S.; Schneider, D.S.; Nusse, R. WntD is a feedback inhibitor of Dorsal/NF-kappaB in Drosophila development and immunity. Nature 2005, 437, 746–749. [Google Scholar] [CrossRef]
- Xiao, C.; Rajewsky, K. MicroRNA control in the immune system: Basic principles. Cell 2009, 136, 26–36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stark, A.; Brennecke, J.; Russell, R.B.; Cohen, S.M. Identification of Drosophila MicroRNA targets. PLoS Biol. 2003, 1, e60. [Google Scholar] [CrossRef] [PubMed]
- Ha, J.; Kim, H.; Yoon, Y.; Park, S. A method of extracting disease-related microRNAs through the propagation algorithm using the environmental factor based global miRNA network. BioMed Mater. Eng. 2015, 26 (Suppl. 1), S1763–S1772. [Google Scholar] [CrossRef] [Green Version]
- Lee, G.J.; Hyun, S. Multiple targets of the microRNA miR-8 contribute to immune homeostasis in Drosophila. Dev. Comp. Immunol. 2014, 45, 245–251. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Li, Y.; Shen, L.; Jin, P.; Chen, L.; Ma, F. miR-958 inhibits Toll signaling and Drosomycin expression via direct targeting of Toll and Dif in Drosophila melanogaster. Am. J. Physiol. Cell Physiol. 2017, 312, C103–C110. [Google Scholar] [CrossRef]
- Li, R.; Huang, Y.; Zhang, Q.; Zhou, H.; Jin, P.; Ma, F. The miR-317 functions as a negative regulator of Toll immune response and influences Drosophila survival. Dev. Comp. Immunol. 2019, 95, 19–27. [Google Scholar] [CrossRef]
- Li, Y.; Li, S.; Li, R.; Xu, J.; Jin, P.; Chen, L.; Ma, F. Genome-wide miRNA screening reveals miR-310 family members negatively regulate the immune response in Drosophila melanogaster via co-targeting Drosomycin. Dev. Comp. Immunol. 2017, 68, 34–45. [Google Scholar] [CrossRef]
- Li, S.; Xu, J.; Sun, L.; Li, R.; Jin, P.; Ma, F. Drosophila miR-964 modulates Toll signaling pathway in response to bacterial infection. Dev. Comp. Immunol. 2017, 77, 252–258. [Google Scholar] [CrossRef]
- Li, R.; Zhou, H.; Jia, C.; Jin, P.; Ma, F. Drosophila Myc restores immune homeostasis of Imd pathway via activating miR-277 to inhibit imd/Tab2. PLoS Genet. 2020, 16, e1008989. [Google Scholar] [CrossRef]
- Choi, I.K.; Hyun, S. Conserved microRNA miR-8 in fat body regulates innate immune homeostasis in Drosophila. Dev. Comp. Immunol. 2012, 37, 50–54. [Google Scholar] [CrossRef]
- Chen, C.Z.; Schaffert, S.; Fragoso, R.; Loh, C. Regulation of immune responses and tolerance: The microRNA perspective. Immunol. Rev. 2013, 253, 112–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, W.; Julie Li, Y.S.; Huang, H.D.; Shyy, J.Y.; Chien, S. microRNA: A master regulator of cellular processes for bioengineering systems. Annu. Rev. Biomed. Eng. 2010, 12, 1–27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Zhang, R.; Su, B. Diversity and evolution of MicroRNA gene clusters. Sci. China C Life Sci. 2009, 52, 261–266. [Google Scholar] [CrossRef]
- Cullen, B.R. Transcription and processing of human microRNA precursors. Mol. Cell 2004, 16, 861–865. [Google Scholar] [CrossRef]
- Baskerville, S.; Bartel, D.P. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 2005, 11, 241–247. [Google Scholar] [CrossRef] [Green Version]
- Vodala, S.; Pescatore, S.; Rodriguez, J.; Buescher, M.; Chen, Y.W.; Weng, R.; Cohen, S.M.; Rosbash, M. The oscillating miRNA 959-964 cluster impacts Drosophila feeding time and other circadian outputs. Cell Metab. 2012, 16, 601–612. [Google Scholar] [CrossRef] [Green Version]
- Neyen, C.; Bretscher, A.J.; Binggeli, O.; Lemaitre, B. Methods to study Drosophila immunity. Methods 2014, 68, 116–128. [Google Scholar] [CrossRef] [Green Version]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [Green Version]
- Betel, D.; Wilson, M.; Gabow, A.; Marks, D.S.; Sander, C. The microRNA.org resource: Targets and expression. Nucleic Acids Res. 2008, 36, D149–D153. [Google Scholar] [CrossRef] [Green Version]
- Enright, A.J.; John, B.; Gaul, U.; Tuschl, T.; Sander, C.; Marks, D.S. MicroRNA targets in Drosophila. Genome Biol. 2003, 5, R1. [Google Scholar] [CrossRef] [PubMed] [Green Version]







Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, R.; Yao, X.; Zhou, H.; Jin, P.; Ma, F. The Drosophila miR-959–962 Cluster Members Repress Toll Signaling to Regulate Antibacterial Defense during Bacterial Infection. Int. J. Mol. Sci. 2021, 22, 886. https://doi.org/10.3390/ijms22020886
Li R, Yao X, Zhou H, Jin P, Ma F. The Drosophila miR-959–962 Cluster Members Repress Toll Signaling to Regulate Antibacterial Defense during Bacterial Infection. International Journal of Molecular Sciences. 2021; 22(2):886. https://doi.org/10.3390/ijms22020886
Chicago/Turabian StyleLi, Ruimin, Xiaolong Yao, Hongjian Zhou, Ping Jin, and Fei Ma. 2021. "The Drosophila miR-959–962 Cluster Members Repress Toll Signaling to Regulate Antibacterial Defense during Bacterial Infection" International Journal of Molecular Sciences 22, no. 2: 886. https://doi.org/10.3390/ijms22020886



