Genome-Wide Characterization of Salt-Responsive miRNAs, circRNAs and Associated ceRNA Networks in Tomatoes
Abstract
:1. Introduction
2. Results
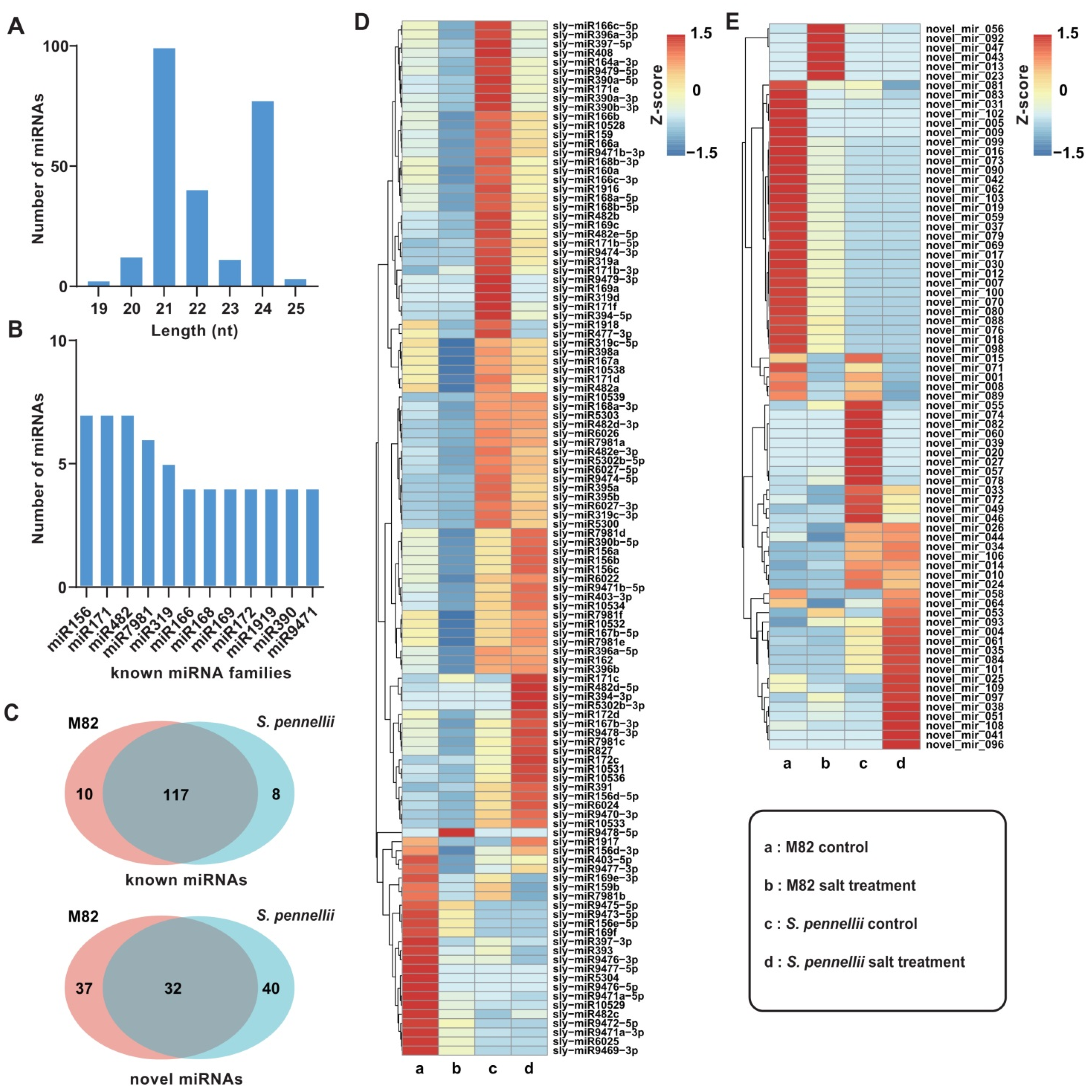
2.1. Small RNA Sequencing of Tomato Roots
2.2. Identification and Characterization of Salt-Responsive miRNAs in Tomatoes
2.3. Prediction and Functional Category of the Putative Target Genes of Salt-Responsive miRNAs
2.4. Validation of Salt-Responsive miRNAs
2.5. Identification and Characterization of Salt-Responsive mRNAs and CircRNAs
2.6. GO and KEGG Analysis of Differentially Expressed mRNAs
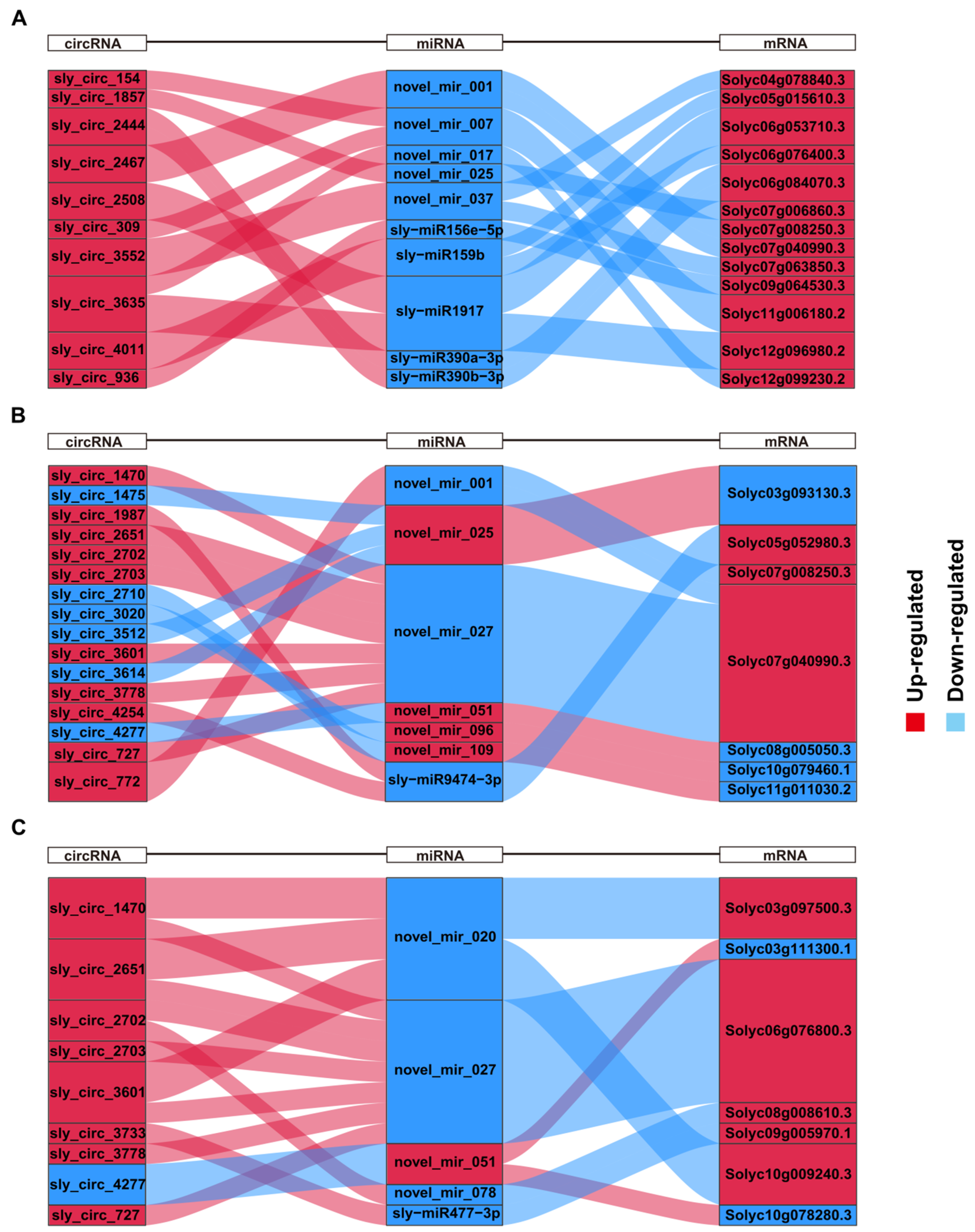
2.7. Construction of Salt-Responsive circRNA–miRNA–mRNA Networks
3. Discussion
3.1. Analysis of Salt-Responsive mRNAs in Tomatoes
3.2. miRNAs Play Crucial Roles in Tomatoes under Salt Stress
3.3. ceRNA Networks Could Shed New Light on the Regulatory Roles of ncRNAs
4. Materials and Methods
4.1. Plant Materials and Preparation
4.2. Small RNA Sequencing and Identification of DE-miRNAs
4.3. Prediction and Annotation of Potential Target Genes of DE-miRNAs
4.4. Validation of miRNAs by qRT-PCR
4.5. Strand-Specific RNA Sequencing and Identification of Differentially Expressed mRNAs and circRNAs
4.6. Construction of the circRNA–miRNA–mRNA Networks
4.7. GO and KEGG Pathway Analyses of mRNAs in the CeRNA Networks
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Francois, L.E.; Grieve, C.M.; Maas, E.V.; Lesch, S.M. Time of Salt Stress Affects Growth and Yield Components of Irrigated Wheat. Agron. J. 1994, 86, 100–107. [Google Scholar] [CrossRef]
- Ashraf, M.; O’leary, J. Responses of some newly developed salt-tolerant genotypes of spring wheat to salt stress: 1. Yield components and ion distribution. J. Agron. Crop. Sci. 1996, 176, 91–101. [Google Scholar] [CrossRef]
- Maas, E.V.; Grattan, S. Crop yields as affected by salinity. Agric. Drain. 1999, 38, 55–108. [Google Scholar]
- Iqbal, M.; Ashraf, M. Gibberellic acid mediated induction of salt tolerance in wheat plants: Growth, ionic partitioning, photosynthesis, yield and hormonal homeostasis. Environ. Exp. Bot. 2013, 86, 76–85. [Google Scholar] [CrossRef]
- Zörb, C.; Geilfus, C.M.; Dietz, K.J. Salinity and crop yield. Plant Biol. 2019, 21, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-K. Genetic Analysis of Plant Salt Tolerance Using Arabidopsis. Plant Physiol. 2000, 124, 941–948. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Parida, A.K.; Das, A.B. Salt tolerance and salinity effects on plants: A review. Ecotoxicol. Environ. Saf. 2005, 60, 324–349. [Google Scholar] [CrossRef] [PubMed]
- Cuartero, J.; Fernández-Muñoz, R. Tomato and salinity. Sci. Hortic. 1998, 78, 83–125. [Google Scholar] [CrossRef]
- Dehan, K.; Tal, M. Salt tolerance in the wild relatives of the cultivated tomato: Responses of Solanum pennellii to high salinity. Irrig. Sci. 1978, 1, 71–76. [Google Scholar] [CrossRef]
- Tal, M.; Shannon, M. Salt tolerance in the wild relatives of the cultivated tomato: Responses of Lycopersicon esculentum, L. cheesmanii, L. peruvianum, Solanum pennellii and F1 hybrids to high salinity. Funct. Plant Biol. 1983, 10, 109–117. [Google Scholar] [CrossRef]
- Martínez, J.P.; Antúnez, A.; Araya, H.; Pertuzé, R.; Fuentes, L.; Lizana, C.; Lutts, S. Salt stress differently affects growth, water status and antioxidant enzyme activities in Solanum lycopersicum and its wild relative Solanum chilense. Aust. J. Bot. 2014, 62, 359–368. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.-K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, J.; Wang, Y.; Wang, G. Salt stress-induced programmed cell death via Ca2+-mediated mitochondrial permeability transition in tobacco protoplasts. Plant Growth Regul. 2005, 45, 243–250. [Google Scholar] [CrossRef]
- Yuan, K.; Rashotte, A.M.; Wysocka-Diller, J.W. ABA and GA signaling pathways interact and regulate seed germination and seedling development under salt stress. Acta Physiol. Plant. 2010, 33, 261–271. [Google Scholar] [CrossRef]
- Duan, L.; Dietrich, D.; Ng, C.H.; Chan, P.M.Y.; Bhalerao, R.; Bennett, M.; Dinneny, J.R. Endodermal ABA Signaling Promotes Lateral Root Quiescence during Salt Stress in Arabidopsis Seedlings. Plant Cell 2013, 25, 324–341. [Google Scholar] [CrossRef] [Green Version]
- Shen, X.; Wang, Z.; Song, X.; Xu, J.; Jiang, C.; Zhao, Y.; Ma, C.; Zhang, H. Transcriptomic profiling revealed an important role of cell wall remodeling and ethylene signaling pathway during salt acclimation in Arabidopsis. Plant Mol. Biol. 2014, 86, 303–317. [Google Scholar] [CrossRef] [PubMed]
- Parre, E.; Ghars, M.A.; Leprince, A.-S.; Thiery, L.; Lefebvre, D.; Bordenave, M.; Richard, L.; Mazars, C.; Abdelly, C.; Savouré, A. Calcium Signaling via Phospholipase C Is Essential for Proline Accumulation upon Ionic But Not Nonionic Hyperosmotic Stresses in Arabidopsis. Plant Physiol. 2007, 144, 503–512. [Google Scholar] [CrossRef] [Green Version]
- Kovtun, Y.; Chiu, W.-L.; Tena, G.; Sheen, J. Functional analysis of oxidative stress-activated mitogen-activated protein kinase cascade in plants. Proc. Natl. Acad. Sci. USA 2000, 97, 2940–2945. [Google Scholar] [CrossRef] [Green Version]
- Kim, S.Y. The role of ABF family bZIP class transcription factors in stress response. Physiol. Plant. 2005, 126, 519–527. [Google Scholar] [CrossRef]
- Ashraf, M.; Akram, N.A.; Arteca, R.N.; Foolad, M.R. The Physiological, Biochemical and Molecular Roles of Brassinosteroids and Salicylic Acid in Plant Processes and Salt Tolerance. Crit. Rev. Plant Sci. 2010, 29, 162–190. [Google Scholar] [CrossRef]
- Quan, R.; Lin, H.; Mendoza, I.; Zhang, Y.; Cao, W.; Yang, Y.; Shang, M.; Chen, S.; Pardo, J.M.; Guo, Y. SCABP8/CBL10, a Putative Calcium Sensor, Interacts with the Protein Kinase SOS2 to Protect Arabidopsis Shoots from Salt Stress. Plant Cell 2007, 19, 1415–1431. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Q.; Chen, Z.-Z.; Zhou, X.-F.; Yin, H.-B.; Li, X.; Xin, X.-F.; Hong, X.-H.; Zhu, J.-K.; Gong, Z. Overexpression of SOS (Salt Overly Sensitive) Genes Increases Salt Tolerance in Transgenic Arabidopsis. Mol. Plant 2009, 2, 22–31. [Google Scholar] [CrossRef] [Green Version]
- Quintero, F.J.; Martinez-Atienza, J.; Villalta, I.; Jiang, X.; Kim, W.-Y.; Ali, Z.; Fujii, H.; Mendoza, I.; Yun, D.-J.; Zhu, J.-K.; et al. Activation of the plasma membrane Na/H antiporter Salt-Overly-Sensitive 1 (SOS1) by phosphorylation of an auto-inhibitory C-terminal domain. Proc. Natl. Acad. Sci. USA 2011, 108, 2611–2616. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qiu, Q.-S.; Guo, Y.; Quintero, F.J.; Pardo, J.M.; Schumaker, K.S.; Zhu, J.-K. Regulation of Vacuolar Na+/H+ Exchange in Arabidopsis thaliana by the Salt-Overly-Sensitive (SOS) Pathway. J. Biol. Chem. 2004, 279, 207–215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quan, R.; Wang, J.; Yang, D.; Zhang, H.; Zhang, Z.; Huang, R. EIN3 and SOS2 synergistically modulate plant salt tolerance. Sci. Rep. 2017, 7, 44637. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.-K.; Liu, J.; Xiong, L. Genetic analysis of salt tolerance in Arabidopsis: Evidence for a critical role of potassium nutrition. Plant Cell 1998, 10, 1181–1191. [Google Scholar] [CrossRef] [Green Version]
- Sathee, L.; Sairam, R.K.; Chinnusamy, V.; Jha, S.K. Differential transcript abundance of salt overly sensitive (SOS) pathway genes is a determinant of salinity stress tolerance of wheat. Acta Physiol. Plant. 2015, 37, 169. [Google Scholar] [CrossRef]
- Qiu, Q.-S.; Guo, Y.; Dietrich, M.A.; Schumaker, K.S.; Zhu, J.K. Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc. Natl. Acad. Sci. USA 2002, 99, 8436–8441. [Google Scholar] [CrossRef] [Green Version]
- Ji, H.; Pardo, J.M.; Batelli, G.; Van Oosten, M.J.; Bressan, R.A.; Li, X. The Salt Overly Sensitive (SOS) Pathway: Established and Emerging Roles. Mol. Plant 2013, 6, 275–286. [Google Scholar] [CrossRef] [Green Version]
- Latz, A.; Mehlmer, N.; Zapf, S.; Mueller, T.D.; Wurzinger, B.; Pfister, B.; Csaszar, E.; Hedrich, R.; Teige, M.; Becker, D. Salt Stress Triggers Phosphorylation of the Arabidopsis Vacuolar K+ Channel TPK1 by Calcium-Dependent Protein Kinases (CDPKs). Mol. Plant 2013, 6, 1274–1289. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Liu, D.; Yang, B.; Liu, W.-Z.; Mu, B.; Song, H.; Chen, B.; Li, Y.; Ren, D.; Deng, H.; et al. Arabidopsis CPK6 positively regulates ABA signaling and drought tolerance through phosphorylating ABA-responsive element-binding factors. J. Exp. Bot. 2019, 71, 188–203. [Google Scholar] [CrossRef]
- Nguyen, N.H.; Jung, C.; Cheong, J.-J. Chromatin remodeling for the transcription of type 2C protein phosphatase genes in response to salt stress. Plant Physiol. Biochem. 2019, 141, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, T.; Fujita, Y.; Sayama, H.; Kidokoro, S.; Maruyama, K.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J. 2010, 61, 672–685. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Chen, S.; Dai, S.; Wang, R.; Li, N.; Shen, X.; Zhou, X.; Lu, C.; Zheng, X.; Hu, Z.; et al. NaCl-induced alternations of cellular and tissue ion fluxes in roots of salt-resistant and salt-sensitive poplar species. Plant Physiol. 2009, 149, 1141–1153. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, X.; Deng, L.; Li, J.; Zhou, X.; Li, N.; Zhang, D.; Lu, Y.; Wang, R.; Sun, J.; Lu, C.; et al. Effect of NaCl on leaf H+-ATPase and the relevance to salt tolerance in two contrasting poplar species. Trees 2010, 24, 597–607. [Google Scholar] [CrossRef] [Green Version]
- Sun, J.; Wang, M.-J.; Ding, M.-Q.; Deng, S.-R.; Liu, M.-Q.; Lu, C.-F.; Zhou, X.-Y.; Shen, X.; Zheng, X.-J.; Zhang, Z.-K.; et al. H2O2 and cytosolic Ca2+ signals triggered by the PM H+-coupled transport system mediate K+/Na+ homeostasis in NaCl-stressed Populus euphratica cells. Plant Cell Environ. 2010, 33, 943–958. [Google Scholar] [CrossRef]
- Sun, J.; Li, L.; Liu, M.; Wang, M.; Ding, M.; Deng, S.; Lu, C.; Zhou, X.; Shen, X.; Zheng, X.; et al. Hydrogen peroxide and nitric oxide mediate K+/Na+ homeostasis and antioxidant defense in NaCl-stressed callus cells of two contrasting poplars. Plant Cell Tissue Organ Cult. 2010, 103, 205–215. [Google Scholar] [CrossRef]
- Yu, Z.; Duan, X.; Luo, L.; Dai, S.; Ding, Z.; Xia, G. How Plant Hormones Mediate Salt Stress Responses. Trends Plant Sci. 2020, 25, 1117–1130. [Google Scholar] [CrossRef]
- Li, H.; Yu, M.; Du, X.; Wang, Z.; Wu, W.; Quintero, F.J.; Jin, X.; Li, H.; Wang, Y. NRT1.5/NPF7.3 functions as a proton-coupled H+/K+ antiporter for K+ loading into the xylem in Arabidopsis. Plant Cell Online 2017, 29, 2016–2026. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drechsler, N.; Zheng, Y.; Bohner, A.; Nobmann, B.; von Wirén, N.; Kunze, R.; Rausch, C. Nitrate-dependent control of shoot K homeostasis by the nitrate transporter1/peptide transporter family member NPF7.3/NRT1.5 and the stelar K+ outward rectifier SKOR in Arabidopsis. Plant Physiol. 2015, 169, 2832–2847. [Google Scholar]
- Xu, P.; Fang, S.; Chen, H.; Cai, W. The brassinosteroid-responsive xyloglucan endotransglucosylase/hydrolase 19 ( XTH19 ) and XTH23 genes are involved in lateral root development under salt stress in Arabidopsis. Plant J. 2020, 104, 59–75. [Google Scholar] [CrossRef] [PubMed]
- Jones-Rhoades, M.W.; Bartel, D.P. Computational Identification of Plant MicroRNAs and Their Targets, Including a Stress-Induced miRNA. Mol. Cell 2004, 14, 787–799. [Google Scholar] [CrossRef]
- Ding, D.; Zhang, L.; Wang, H.; Liu, Z.; Zhang, Z.; Zheng, Y. Differential expression of miRNAs in response to salt stress in maize roots. Ann. Bot. 2008, 103, 29–38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, B.; Ge, L.; Liang, R.; Li, W.; Ruan, K.; Lin, H.; Jin, Y. Members of miR-169 family are induced by high salinity and transiently inhibit the NF-YA transcription factor. BMC Mol. Biol. 2009, 10, 29. [Google Scholar] [CrossRef] [Green Version]
- Yin, Z.; Li, Y.; Yu, J.; Liu, Y.; Li, C.; Han, X.; Shen, F. Difference in miRNA expression profiles between two cotton cultivars with distinct salt sensitivity. Mol. Biol. Rep. 2011, 39, 4961–4970. [Google Scholar] [CrossRef]
- Zhang, S.; Zhou, J.; Han, S.; Yang, W.; Li, W.; Wei, H.; Li, X.; Qi, L. Four abiotic stress-induced miRNA families differentially regulated in the embryogenic and non-embryogenic callus tissues of Larix leptolepis. Biochem. Biophys. Res. Commun. 2010, 398, 355–360. [Google Scholar] [CrossRef]
- Zou, C.; Chen, A.; Xiao, L.; Muller, H.M.; Ache, P.; Haberer, G.; Zhang, M.; Jia, W.; Deng, P.; Huang, R.; et al. A high-quality genome assembly of quinoa provides insights into the molecular basis of salt bladder-based salinity tolerance and the exceptional nutritional value. Cell Res. 2017, 27, 1327–1340. [Google Scholar] [CrossRef]
- Zhao, B.; Liang, R.; Ge, L.; Li, W.; Xiao, H.; Lin, H.; Ruan, K.; Jin, Y. Identification of drought-induced microRNAs in rice. Biochem. Biophys. Res. Commun. 2007, 354, 585–590. [Google Scholar] [CrossRef]
- Zhou, L.; Liu, Y.; Liu, Z.; Kong, D.; Duan, M.; Luo, L. Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J. Exp. Bot. 2010, 61, 4157–4168. [Google Scholar] [CrossRef] [PubMed]
- Kantar, M.; Lucas, S.J.; Budak, H. miRNA expression patterns of Triticum dicoccoides in response to shock drought stress. Planta 2010, 233, 471–484. [Google Scholar] [CrossRef]
- Frazier, T.P.; Sun, G.; Burklew, C.E.; Zhang, B. Salt and Drought Stresses Induce the Aberrant Expression of microRNA Genes in Tobacco. Mol. Biotechnol. 2011, 49, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Chen, L.; Zhao, M.; Tian, Q.; Zhang, W.-H. Identification of drought-responsive microRNAs in Medicago truncatula by genome-wide high-throughput sequencing. BMC Genom. 2011, 12, 367. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Zhang, H.; Srivastava, A.K.; Pan, Y.; Bai, J.; Fang, J.; Shi, H.; Zhu, J.K. Knockdown of rice microRNA166 confers drought resistance by causing leaf rolling and altering stem xylem development. Plant Physiol. 2018, 176, 2082–2094. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Wang, Y.; Gao, X.; Liu, C.; Gai, S. Identification and characterization of microRNAs in tree peony during chilling induced dormancy release by high-throughput sequencing. Sci. Rep. 2018, 8, 4537. [Google Scholar] [CrossRef] [Green Version]
- Zhang, S.; Wang, Y.; Li, K.; Zou, Y.; Chen, L.; Li, X. Identification of Cold-Responsive miRNAs and Their Target Genes in Nitrogen-Fixing Nodules of Soybean. Int. J. Mol. Sci. 2014, 15, 13596–13614. [Google Scholar] [CrossRef]
- Yu, X.; Wang, H.; Lu, Y.; De Ruiter, M.; Cariaso, M.; Prins, M.; Van Tunen, A.; He, Y. Identification of conserved and novel microRNAs that are responsive to heat stress in Brassica rapa. J. Exp. Bot. 2011, 63, 1025–1038. [Google Scholar] [CrossRef] [PubMed]
- Kruszka, K.; Pacak, A.; Swida-Barteczka, A.; Nuc, P.; Alaba, S.; Wroblewska, Z.; Karlowski, W.; Jarmolowski, A.; Szweykowska-Kulinska, Z. Transcriptionally and post-transcriptionally regulated microRNAs in heat stress response in barley. J. Exp. Bot. 2014, 65, 6123–6135. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.; Ren, Y.; Zhang, Y.; Xu, J.; Sun, F.; Zhang, Z.; Wang, Y. Genome-wide identification and expression analysis of heat-responsive and novel microRNAs in Populus tomentosa. Gene 2012, 504, 160–165. [Google Scholar] [CrossRef]
- Sunkar, R.; Zhu, J.-K. Novel and Stress-Regulated MicroRNAs and Other Small RNAs from Arabidopsis. Plant Cell 2004, 16, 2001–2019. [Google Scholar] [CrossRef] [Green Version]
- Liu, H.-H.; Tian, X.; Li, Y.-J.; Wu, C.-A.; Zheng, C.-C. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 2008, 14, 836–843. [Google Scholar] [CrossRef] [Green Version]
- Jagadeeswaran, G.; Saini, A.; Sunkar, R. Biotic and abiotic stress down-regulate miR398 expression in Arabidopsis. Planta 2009, 229, 1009–1014. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Xie, K.; Xiong, L. Global expression profiling of rice microRNAs by one-tube stem-loop reverse transcription quantitative PCR revealed important roles of microRNAs in abiotic stress responses. Mol. Genet. Genom. 2010, 284, 477–488. [Google Scholar] [CrossRef] [PubMed]
- Jian, X.; Zhang, L.; Li, G.; Wang, X.; Cao, X.; Fang, X.; Chen, F. Identification of novel stress-regulated microRNAs from Oryza sativa L. Genomics 2010, 95, 47–55. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, B.; Duan, H.; Li, J.; Deng, X.W.; Yin, W.; Xia, X. Global identification of miRNAs and targets in Populus euphratica under salt stress. Plant Mol. Biol. 2013, 81, 525–539. [Google Scholar] [CrossRef]
- Zhuang, Y.; Zhou, X.-H.; Liu, J. Conserved miRNAs and Their Response to Salt Stress in Wild Eggplant Solanum linnaeanum Roots. Int. J. Mol. Sci. 2014, 15, 839–849. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Liu, D.; Chen, D.; Cheng, Y.; Zhang, X.; Song, L.; Hu, M.; Dong, J.; Shen, F. MicroRNA414c affects salt tolerance of cotton by regulating reactive oxygen species metabolism under salinity stress. RNA Biol. 2019, 16, 362–375. [Google Scholar] [CrossRef] [Green Version]
- Bai, Q.; Wang, X.; Chen, X.; Shi, G.; Liu, Z.; Guo, C.; Xiao, K. Wheat miRNA TaemiR408 Acts as an Essential Mediator in Plant Tolerance to Pi Deprivation and Salt Stress via Modulating Stress-Associated Physiological Processes. Front. Plant Sci. 2018, 9, 499. [Google Scholar] [CrossRef]
- Ye, C.; Chen, L.; Liu, C.; Zhu, Q.; Fan, L. Widespread noncoding circular RNA s in plants. New Phytol. 2015, 208, 88–95. [Google Scholar] [CrossRef]
- Sablok, G.; Zhao, H.; Sun, X. Plant Circular RNAs (circRNAs): Transcriptional Regulation Beyond miRNAs in Plants. Mol. Plant 2016, 9, 192–194. [Google Scholar] [CrossRef] [Green Version]
- Ashwal-Fluss, R.; Meyer, M.; Pamudurti, N.R.; Ivanov, A.; Bartok, O.; Hanan, M.; Evantal, N.; Memczak, S.; Rajewsky, N.; Kadener, S. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 2014, 56, 55–66. [Google Scholar] [CrossRef] [Green Version]
- Hansen, T.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef] [PubMed]
- Kulcheski, F.; Christoff, A.P.; Margis, R. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J. Biotechnol. 2016, 238, 42–51. [Google Scholar] [CrossRef]
- Li, Q.-F.; Zhang, Y.-C.; Chen, Y.-Q.; Yu, Y. Circular RNAs roll into the regulatory network of plants. Biochem. Biophys. Res. Commun. 2017, 488, 382–386. [Google Scholar] [CrossRef] [PubMed]
- Chu, Q.; Bai, P.; Zhu, X.; Zhang, X.; Mao, L.; Zhu, Q.H.; Fan, L.; Ye, C.Y. Characteristics of plant circular RNAs. Brief. Bioinform. 2020, 21, 135–143. [Google Scholar] [CrossRef]
- Lu, T.; Cui, L.; Zhou, Y.; Zhu, C.; Fan, D.; Gong, H.; Zhao, Q.; Zhou, C.; Zhao, Y.; Lu, D.; et al. Transcriptome-wide investigation of circular RNAs in rice. RNA 2015, 21, 2076–2087. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, B.; Hao, Z.; Zhu, Y.; Zhang, H.; Li, G. Genome-wide identification and functional analysis of circRNAs in Zea mays. PLoS ONE 2018, 13, e0202375. [Google Scholar] [CrossRef] [Green Version]
- Zuo, J.; Wang, Q.; Zhu, B.; Luo, Y.; Gao, L. Deciphering the roles of circRNAs on chilling injury in tomato. Biochem. Biophys. Res. Commun. 2016, 479, 132–138. [Google Scholar] [CrossRef]
- Wang, Y.; Yang, M.; Wei, S.; Qin, F.; Zhao, H.; Suo, B. Identification of Circular RNAs and Their Targets in Leaves of Triticum aestivum L. under Dehydration Stress. Front. Plant Sci. 2017, 7, 2024. [Google Scholar] [CrossRef] [Green Version]
- Zhao, W.; Cheng, Y.; Zhang, C.; You, Q.; Shen, X.; Guo, W.; Jiao, Y. Genome-wide identification and characterization of circular RNAs by high throughput sequencing in soybean. Sci. Rep. 2017, 7, 5636. [Google Scholar] [CrossRef] [Green Version]
- Zhao, T.; Wang, L.; Baoliang, Z.; Xueying, G.; Guan, X.; Zhou, B. Characterization of conserved circular RNA in polyploid Gossypium species and their ancestors. FEBS Lett. 2017, 591, 3660–3669. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Lin, J.; Wang, H.; Li, X.; Yang, Q.; Li, H.; Chang, Y. Identification and characterization of circRNAs in Pyrus betulifolia Bunge under drought stress. PLoS ONE 2018, 13, e0200692. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.-X.; Jia, J.-H.; Yang, L.; Xia, Y.C.; Zhang, H.L.; Jia, J.B.; Zhou, R.; Nie, P.Y.; Yin, J.L.; Ma, D.F.; et al. Identification of cucumber circular RNAs responsive to salt stress. BMC Plant Biol. 2019, 19, 164. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, P.; Fan, Y.; Sun, X.; Chen, L.; Terzaghi, W.; Bucher, E.; Li, L.; Dai, M. A large-scale circular RNA profiling reveals universal molecular mechanisms responsive to drought stress in maize and Arabidopsis. Plant J. 2019, 98, 697–713. [Google Scholar] [CrossRef]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The Rosetta Stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, Y.; Zhang, H.; Fan, M.; He, Y.; Guo, P. Genome-wide identification of long non-coding RNAs and circular RNAs reveal their ceRNA networks in response to cucumber green mottle mosaic virus infection in watermelon. Arch. Virol. 2020, 165, 1177–1190. [Google Scholar] [CrossRef]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak-Wolf, A.; Maier, L.; Mackowiak, S.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Guo, Z.-J.; Chen, X.-J.; Wu, X.-L.; Ling, J.-Q.; Xu, P. Overexpression of the AP2/EREBP transcription factor OPBP1 enhances disease resistance and salt tolerance in tobacco. Plant Mol. Biol. 2004, 55, 607–618. [Google Scholar] [CrossRef]
- Zhang, S.; Li, X.; Fan, S.; Zhou, L.; Wang, Y. Overexpression of HcSCL13, a Halostachys caspica GRAS transcription factor, enhances plant growth and salt stress tolerance in transgenic Arabidopsis. Plant Physiol. Biochem. 2020, 151, 243–254. [Google Scholar] [CrossRef]
- Ma, H.-S.; Liang, D.; Shuai, P.; Xia, X.-L.; Yin, W.-L. The salt- and drought-inducible poplar GRAS protein SCL7 confers salt and drought tolerance in Arabidopsis thaliana. J. Exp. Bot. 2010, 61, 4011–4019. [Google Scholar] [CrossRef] [Green Version]
- Wu, J.; Jiang, Y.; Liang, Y.; Chen, L.; Chen, W.; Cheng, B. Expression of the maize MYB transcription factor ZmMYB3R enhances drought and salt stress tolerance in transgenic plants. Plant Physiol. Biochem. 2019, 137, 179–188. [Google Scholar] [CrossRef]
- Wei, Q.; Luo, Q.; Wang, R.; Zhang, F.; He, Y.; Zhang, Y.; Qiu, D.; Li, K.; Chang, J.; Yang, G.; et al. A Wheat R2R3-type MYB Transcription Factor TaODORANT1 Positively Regulates Drought and Salt Stress Responses in Transgenic Tobacco Plants. Front. Plant Sci. 2017, 8, 1374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, H.-L.; Ma, N.-N.; Meng, X.; Zhang, S.; Wang, J.-R.; Chai, S.; Meng, Q.-W. A novel tomato MYC-type ICE1-like transcription factor, SlICE1a, confers cold, osmotic and salt tolerance in transgenic tobacco. Plant Physiol. Biochem. 2013, 73, 309–320. [Google Scholar] [CrossRef] [PubMed]
- Mao, K.; Dong, Q.; Li, C.; Liu, C.; Ma, F. Genome Wide Identification and Characterization of Apple bHLH Transcription Factors and Expression Analysis in Response to Drought and Salt Stress. Front. Plant Sci. 2017, 8, 480. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zang, D.; Wang, J.; Zhang, X.; Liu, Z.; Wang, Y. Arabidopsis heat shock transcription factor HSFA7b positively mediates salt stress tolerance by binding to an E-box-like motif to regulate gene expression. J. Exp. Bot. 2019, 70, 5355–5374. [Google Scholar] [CrossRef]
- Cai, R.; Dai, W.; Zhang, C.; Wang, Y.; Wu, M.; Zhao, Y.; Ma, Q.; Xiang, Y.; Cheng, B. The maize WRKY transcription factor ZmWRKY17 negatively regulates salt stress tolerance in transgenic Arabidopsis plants. Planta 2017, 246, 1215–1231. [Google Scholar] [CrossRef]
- Yu, Y.; Wang, N.; Hu, R.; Xiang, F. Genome-wide identification of soybean WRKY transcription factors in response to salt stress. SpringerPlus 2016, 5, 920. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, Z.; Wang, C.; Xue, F.; Zhang, H.; Ji, W. Wheat NAC transcription factor TaNAC29 is involved in response to salt stress. Plant Physiol. Biochem. 2015, 96, 356–363. [Google Scholar] [CrossRef]
- Guo, X.; Chen, G.; Cui, B.; Gao, Q.; Guo, J.-E.; Li, A.; Zhang, L.; Hu, Z. Solanum lycopersicum agamous-like MADS-box protein AGL15-like gene, SlMBP11, confers salt stress tolerance. Mol. Breed. 2016, 36, 125. [Google Scholar] [CrossRef]
- lhan, E.; Büyük, İ.; İnal, B. Transcriptome–Scale characterization of salt responsive bean TCP transcription factors. Gene 2018, 642, 64–73. [Google Scholar]
- Yiguo, S.; Shouyi, C. Molecular mechanism of plant responses to salt stress. Yi Chuan= Hered. 2001, 23, 365–369. [Google Scholar]
- Guohua, L. Research Progress in the Mechanism of Plant Resistance to Salt Stress. J. Anhui Agric. Sci. 2006, 34, 6111. [Google Scholar]
- Cheong, M.S.; Yun, D.-J. Salt-stress signaling. J. Plant Biol. 2007, 50, 148–155. [Google Scholar] [CrossRef]
- Yun, L. Research Advances on the Molecular Mechanism of Plant Salt-tolerance. J. Anhui Agric. Sci. 2010, 12, 6087–6089. [Google Scholar]
- Fariduddin, Q.; Zaid, A.; Mohammad, F. Plant Growth Regulators and Salt Stress: Mechanism of Tolerance Trade-Off. In Salt Stress, Microbes, and Plant Interactions: Causes and Solution; Springer: Berlin/Heidelberg, Germany, 2019; pp. 91–111. [Google Scholar]
- Lu, Y.; Liu, P.; Xu, Z.; Zhang, R.; Liu, L.; Li, L.; Chen, M.; Ye, X.; Chen, Y.; Ma, Y.; et al. Overexpression of W6 gene increases salt tolerance in transgenic tobacco plants. Acta Agron. Sin. 2008, 34, 984–990. [Google Scholar] [CrossRef]
- Kim, J.H.; Nguyen, N.H.; Jeong, C.Y.; Nguyen, N.T.; Hong, S.W.; Lee, H. Loss of the R2R3 MYB, AtMyb73, causes hyper-induction of the SOS1 and SOS3 genes in response to high salinity in Arabidopsis. J. Plant Physiol. 2013, 170, 1461–1465. [Google Scholar] [CrossRef]
- Rao, S.S.; El-Habbak, M.H.; Havens, W.M.; Singh, A.; Zheng, D.; Vaughn, L.; Haudenshield, J.S.; Hartman, G.L.; Korban, S.S.; Ghabrial, S.A. Overexpression of GmCaM4 in soybean enhances resistance to pathogens and tolerance to salt stress. Mol. Plant Pathol. 2014, 15, 145–160. [Google Scholar] [CrossRef]
- Wang, X.; Hou, C.; Zheng, K.; Li, Q.; Chen, S.; Wang, S. Overexpression of ERF96, a small ethylene response factor gene, enhances salt tolerance in Arabidopsis. Biol. Plant. 2017, 61, 693–701. [Google Scholar] [CrossRef]
- Zhang, A.; Liu, Y.; Wang, F.; Li, T.; Chen, Z.; Kong, D.; Bi, J.; Zhang, F.; Luo, X.; Wang, J.; et al. Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol. Breed. 2019, 39, 47. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.-R.; Liang, J.-H.; Wang, G.-F.; Sun, M.-X.; Peng, F.-T.; Xiao, Y.-S. Overexpression of PpSnRK1α in tomato enhanced salt tolerance by regulating ABA signaling pathway and reactive oxygen metabolism. BMC Plant Biol. 2020, 20, 128. [Google Scholar] [CrossRef]
- Jing, X.; Sun, Y.; Xiang, M.; Qian, Z.; Lang, T.; Zhao, R.; Shen, X.; Chen, S. Overexpression of KcTrxf in tobacco enhances salt tolerance through the regulation of ROS homeostasis under NaCl stress. J. Beijing For. Univ. 2015, 37, 17–26. [Google Scholar]
- Jamoussi, R.J.; Elabbassi, M.M.; Ben Jouira, H.; Hanana, M.; Zoghlami, N.; Ghorbel, A.; Mliki, A. Physiological responses of transgenic tobacco plants expressing the dehydration-responsive RD22 gene of Vitis vinifera to salt stress. Turk. J. Bot. 2014, 38, 268–280. [Google Scholar] [CrossRef]
- Xu, Z.-S.; Xiong, T.-F.; Ni, Z.-Y.; Chen, X.-P.; Chen, M.; Li, L.-C.; Gao, D.-Y.; Yu, X.-D.; Liu, P.; Ma, Y.-Z. Isolation and identification of two genes encoding leucine-rich repeat (LRR) proteins differentially responsive to pathogen attack and salt stress in tobacco. Plant Sci. 2009, 176, 38–45. [Google Scholar] [CrossRef]
- de Lorenzo, L.; Merchan, F.; Laporte, P.; Thompson, R.; Clarke, J.; Sousa, C.; Crespi, M. A novel plant leucine-rich repeat receptor kinase regulates the response of Medicago truncatula roots to salt stress. Plant Cell 2009, 21, 668–680. [Google Scholar] [CrossRef] [Green Version]
- Patankar, H.V.; Al-Harrasi, I.; Al Kharusi, L.; Jana, G.A.; Al-Yahyai, R.; Sunkar, R.; Yaish, M.W. Overexpression of a Metallothionein 2A Gene from Date Palm Confers Abiotic Stress Tolerance to Yeast and Arabidopsis thaliana. Int. J. Mol. Sci. 2019, 20, 2871. [Google Scholar] [CrossRef] [Green Version]
- Zhao, G.; Yu, H.; Liu, M.; Lu, Y.; Ouyang, B. Identification of salt-stress responsive microRNAs from Solanum lycopersicum and Solanum pimpinellifolium. Plant Growth Regul. 2017, 83, 129–140. [Google Scholar] [CrossRef]
- Zhou, J.; Liu, M.; Jiang, J.; Qiao, G.; Lin, S.; Li, H.; Xie, L.; Zhuo, R. Expression profile of miRNAs in Populus cathayana L. and Salix matsudana Koidz under salt stress. Mol. Biol. Rep. 2012, 39, 8645–8654. [Google Scholar] [CrossRef]
- Sailaja, B.; Voleti, S.; Subrahmanyam, D.; Sarla, N.; Prasanth, V.V.; Bhadana, V.; Mangrauthia, S. Prediction and Expression Analysis of miRNAs Associated with Heat Stress in Oryza sativa. Rice Sci. 2014, 21, 3–12. [Google Scholar] [CrossRef]
- Vashisht, I.; Mishra, P.; Pal, T.; Chanumolu, S.; Singh, T.R.; Chauhan, R.S. Mining NGS transcriptomes for miRNAs and dissecting their role in regulating growth, development, and secondary metabolites production in different organs of a medicinal herb, Picrorhiza kurroa. Planta 2015, 241, 1255–1268. [Google Scholar] [CrossRef]
- Kumar, G.; Singh, A.; Vinutha, T.; Kumar, S.; Kumar, R.R.; Praveen, S. Possible role of miRNAs and their targets, in modulating leaf morphology and plant growth during leaf curl virus infection in tomato. Indian J. Plant Physiol. 2017, 22, 608–615. [Google Scholar] [CrossRef]
- Curaba, J.; Singh, M.; Bhalla, P.L. miRNAs in the crosstalk between phytohormone signalling pathways. J. Exp. Bot. 2014, 65, 1425–1438. [Google Scholar] [CrossRef]
- De-La-Peña, C.; Nic-Can, G.I.; Avilez-Montalvo, J.; Cetz-Chel, J.E.; Loyola-Vargas, V.M. The Role of MiRNAs in Auxin Signaling and Regulation During Plant Development. In Plant Epigenetics; Springer: Cham, Switzerland, 2017; pp. 23–48. [Google Scholar] [CrossRef]
- Prigge, M.; Clark, S.E. Evolution of the class III HD-Zip gene family in land plants. Evol. Dev. 2006, 8, 350–361. [Google Scholar] [CrossRef]
- Belamkar, V.; Weeks, N.T.; Bharti, A.K.; Farmer, A.D.; Graham, M.A.; Cannon, S.B. Comprehensive characterization and RNA-Seq profiling of the HD-Zip transcription factor family in soybean (Glycine max) during dehydration and salt stress. BMC Genom. 2014, 15, 950. [Google Scholar] [CrossRef] [Green Version]
- He, F.; Xu, C.; Fu, X.; Shen, Y.; Guo, L.; Leng, M.; Luo, K. The MicroRNA390/TRANS-ACTING SHORT INTERFERING RNA3 Module Mediates Lateral Root Growth under Salt Stress via the Auxin Pathway. Plant Physiol. 2018, 177, 775–791. [Google Scholar] [CrossRef] [Green Version]
- Wen, F.L.; Yue, Y.; He, T.F.; Gao, X.M.; Zhou, Z.S.; Long, X.H. Identification of miR390-TAS3-ARF pathway in response to salt stress in Helianthus tuberosus L. Gene 2020, 738, 144460. [Google Scholar] [CrossRef]
- Dong, Y.; Liu, X.; Yao, N.; Zhao, L.; Li, H. Expression of safflower miR397a gene and its role in LAC2 regulation. J. Northwest Agric. For. Univ. (Nat. Sci. Ed.) 2016, 44, 173–180. [Google Scholar]
- Zhao, X.; Pang, M.; Zhao, Q.; Ren, Y.; Hao, Y.; You, C. Cloning and expression analysis of tomato LeLAC miR397 gene. Acta Hortic. Sin. 2015, 42, 1285–1298. [Google Scholar]
- Chen, L.; Luan, Y.; Zhai, J. Sp-miR396a-5p acts as a stress-responsive genes regulator by conferring tolerance to abiotic stresses and susceptibility to Phytophthora nicotianae infection in transgenic tobacco. Plant Cell Rep. 2015, 34, 2013–2025. [Google Scholar] [CrossRef]
- Barciszewska-Pacak, M.; Milanowska, K.; Knop, K.; Bielewicz, D.; Nuc, P.; Plewka, P.; Pacak, A.M.; Vazquez, F.; Karlowski, W.; Jarmolowski, A.; et al. Arabidopsis microRNA expression regulation in a wide range of abiotic stress responses. Front. Plant Sci. 2015, 6, 410. [Google Scholar] [CrossRef]
- Song, Z.; Zhang, L.; Wang, Y.; Li, H.; Li, S.; Zhao, H.; Zhang, H. Constitutive Expression of miR408 Improves Biomass and Seed Yield in Arabidopsis. Front. Plant Sci. 2018, 8, 2114. [Google Scholar] [CrossRef]
- Macovei, A.; Tuteja, N. microRNAs targeting DEAD-box helicases are involved in salinity stress response in rice (Oryza sativa L.). BMC Plant Biol. 2012, 12, 183. [Google Scholar] [CrossRef] [Green Version]
- Cao, C.; Long, R.; Zhang, T.; Kang, J.; Wang, Z.; Wang, P.; Sun, H.; Yu, J.; Yang, Q. Genome-Wide Identification of microRNAs in Response to Salt/Alkali Stress in Medicago truncatula through High-Throughput Sequencing. Int. J. Mol. Sci. 2018, 19, 4076. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.; Xu, L.; Wang, Y.; Keyun, Z.; Zhu, X.; Luo, X.; Gong, Y.; Wang, R.; Limera, C.; Zhang, K.; et al. Identification of novel and salt-responsive miRNAs to explore miRNA-mediated regulatory network of salt stress response in radish (Raphanus sativus L.). BMC Genom. 2015, 16, 197. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.; Zhu, P.; Kang, H.; Liu, L.; Cao, Q.; Sun, J.; Dong, T.; Zhu, M.; Li, Z.; Xu, T. High-throughput deep sequencing reveals the important role that microRNAs play in the salt response in sweet potato (Ipomoea batatas L.). BMC Genom. 2020, 21, 164. [Google Scholar] [CrossRef] [Green Version]
- Eren, H.; Pekmezci, M.; Okay, S.; Türktaş, M.; Inal, B.; Ilhan, E.; Atak, M.; Erayman, M.; Unver, T. Hexaploid wheat (Triticum aestivum) root miRNome analysis in response to salt stress. Ann. Appl. Biol. 2015, 167, 208–216. [Google Scholar] [CrossRef]
- Hao, Y.-J.; Wei, W.; Song, Q.-X.; Chen, H.-W.; Zhang, Y.-Q.; Wang, F.; Zou, H.-F.; Lei, G.; Tian, A.-G.; Zhang, W.-K.; et al. Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants. Plant J. 2011, 68, 302–313. [Google Scholar] [CrossRef]
- Candar-Cakir, B.; Arican, E.; Zhang, B. Small RNA and degradome deep sequencing reveals drought-and tissue-specific micrornas and their important roles in drought-sensitive and drought-tolerant tomato genotypes. Plant Biotechnol. J. 2016, 14, 1727–1746. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naidoo, G.; Naidoo, Y. Effects of salinity and nitrogen on growth, ion relations and proline accumulation in Triglochin bulbosa. Wetl. Ecol. Manag. 2001, 9, 491–497. [Google Scholar] [CrossRef]
- Ding, X.; Tian, C.; Zhang, S.; Song, J.; Zhang, F.; Mi, G.; Feng, G. Effects of NO3−-N on the growth and salinity tolerance of Tamarix laxa Willd. Plant Soil 2009, 331, 57–67. [Google Scholar] [CrossRef]
- Yuan, J.F.; Feng, G.; Ma, H.Y.; Tian, C.Y. Effect of nitrate on root development and nitrogen uptake of Suaeda physophora under NaCl salinity. Pedosphere 2010, 20, 536–544. [Google Scholar] [CrossRef]
- Song, J.; Ding, X.; Feng, G.; Zhang, F. Nutritional and osmotic roles of nitrate in a euhalophyte and a xerophyte in saline conditions. New Phytol. 2006, 171, 357–366. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- Sunkar, R.; Chinnusamy, V.; Zhu, J.; Zhu, J.-K. Small RNAs as big players in plant abiotic stress responses and nutrient deprivation. Trends Plant Sci. 2007, 12, 301–309. [Google Scholar] [CrossRef]
- Hwang, E.-W.; Shin, S.-J.; Yu, B.-K.; Byun, M.-O.; Kwon, H.-B. miR171 Family Members are Involved in Drought Response in Solanum tuberosum. J. Plant Biol. 2010, 54, 43–48. [Google Scholar] [CrossRef]
- Shi, X.; Jiang, F.; Wen, J.; Cui, S.; Zhou, Y.; Wu, Z. MicroRNA319 family members play an important role in Solanum habrochaites and S. lycopersicum responses to chilling and heat stresses. Biol. Plant. 2019, 63, 200–209. [Google Scholar] [CrossRef]
- Zhou, M.; Li, D.; Li, Z.; Hu, Q.; Yang, C.; Zhu, L.; Luo, H. Constitutive Expression of a miR319 Gene Alters Plant Development and Enhances Salt and Drought Tolerance in Transgenic Creeping Bentgrass. Plant Physiol. 2013, 161, 1375–1391. [Google Scholar] [CrossRef] [Green Version]
- Zhou, M.; Luo, H. Role of microRNA319 in creeping bentgrass salinity and drought stress response. Plant Signal. Behav. 2014, 9, 1375–1391. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Li, D.; Yan, J.; Wang, K.; Luo, H.; Zhang, W. MiR319-mediated ethylene biosynthesis, signalling and salt stress response in switchgrass. Plant Biotechnol. J. 2019, 17, 2370–2383. [Google Scholar] [CrossRef]
- Song, J.B.; Gao, S.; Sun, D.; Li, H.; Shu, X.X.; Yang, Z.M. miR394 and LCR are involved in Arabidopsis salt and drought stress responses in an abscisic acid-dependent manner. BMC Plant Biol. 2013, 13, 210. [Google Scholar] [CrossRef] [Green Version]
- Tian, X.; Song, L.; Wang, Y.; Jin, W.; Tong, F.; Wu, F. miR394 Acts as a Negative Regulator of Arabidopsis Resistance to B. cinerea Infection by Targeting LCR. Front. Plant Sci. 2018, 9, 903. [Google Scholar] [CrossRef] [Green Version]
- Zsigmond, L.; Rigó, G.; Szarka, A.; Székely, G.; Ötvös, K.; Darula, Z.; Medzihradszky, K.F.; Koncz, C.; Koncz, Z.; Szabados, L. Arabidopsis PPR40 Connects Abiotic Stress Responses to Mitochondrial Electron Transport. Plant Physiol. 2008, 146, 1721–1737. [Google Scholar] [CrossRef] [Green Version]
- Qu, D.; Yan, F.; Meng, R.; Jiang, X.; Yang, H.; Gao, Z.; Dong, Y.; Yang, Y.; Zhao, Z. Identification of MicroRNAs and Their Targets Associated with Fruit-Bagging and Subsequent Sunlight Re-exposure in the “Granny Smith” Apple Exocarp Using High-Throughput Sequencing. Front. Plant Sci. 2016, 7, 27. [Google Scholar] [CrossRef] [Green Version]
- Su, Y.; Xu, C.; Liu, Y.; Hu, Y.; Wu, H. Circular RNA hsa_circ_0001649 inhibits hepatocellular carcinoma progression via multiple miRNAs sponge. Aging 2019, 11, 3362–3375. [Google Scholar] [CrossRef]
- O’Malley, D.M.; Whetten, R.; Bao, W.; Chen, C.-L.; Sederoff, R.R. The role of of laccase in lignification. Plant J. 1993, 4, 751–757. [Google Scholar] [CrossRef]
- Gavnholt, B.; Larsen, K. Molecular biology of plant laccases in relation to lignin formation. Physiol. Plant. 2002, 116, 273–280. [Google Scholar] [CrossRef]
- Hu, Q.; Min, L.; Yang, X.; Jin, S.; Zhang, L.; Li, Y.; Ma, Y.; Qi, X.; Li, D.; Liu, H.; et al. Laccase GhLac1 Modulates Broad-Spectrum Biotic Stress Tolerance via Manipulating Phenylpropanoid Pathway and Jasmonic Acid Synthesis. Plant Physiol. 2017, 176, 1808–1823. [Google Scholar] [CrossRef] [Green Version]
- Jeon, J.R.; Baldrian, P.; Murugesan, K.; Chang, Y.S. Laccase-catalysed oxidations of naturally occurring phenols: From in vivo biosynthetic pathways to green synthetic applications. Microb. Biotechnol. 2012, 5, 318–332. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franke, R.; Schreiber, L. Suberin—A biopolyester forming apoplastic plant interfaces. Curr. Opin. Plant Biol. 2007, 10, 252–259. [Google Scholar] [CrossRef]
- Höfer, R.; Briesen, I.; Beck, M.; Pinot, F.; Schreiber, L.; Franke, R. The Arabidopsis cytochrome P450 CYP86A1 encodes a fatty acid ω-hydroxylase involved in suberin monomer biosynthesis. J. Exp. Bot. 2008, 59, 2347–2360. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Beisson, F.; Koo, A.; Molina, I.; Pollard, M.; Ohlrogge, J. Identification of acyltransferases required for cutin biosynthesis and production of cutin with suberin-like monomers. Proc. Natl. Acad. Sci. USA 2007, 104, 18339–18344. [Google Scholar] [CrossRef] [Green Version]
- Compagnon, V.; Diehl, P.; Benveniste, I.; Meyer, D.; Schaller, H.; Schreiber, L.; Franke, R.; Pinot, F. CYP86B1 is required for very long chain ω-hydroxyacid and α, ω-dicarboxylic acid synthesis in root and seed suberin polyester. Plant Physiol. 2009, 150, 1831–1843. [Google Scholar] [CrossRef] [Green Version]
- Wang, P.; Wang, C.-M.; Gao, L.; Cui, Y.-N.; Yang, H.-L.; De Silva, N.D.G.; Ma, Q.; Bao, A.-K.; Flowers, T.J.; Rowland, O.; et al. Aliphatic suberin confers salt tolerance to Arabidopsis by limiting Na+ influx, K+ efflux and water backflow. Plant Soil 2020, 448, 603–620. [Google Scholar] [CrossRef]
- Bruckhoff, V.; Haroth, S.; Feussner, K.; König, S.; Brodhun, F.; Feussner, I. Functional Characterization of CYP94-Genes and Identification of a Novel Jasmonate Catabolite in Flowers. PLoS ONE 2016, 11, e0159875. [Google Scholar] [CrossRef] [Green Version]
- Kurotani, K.; Hayashi, K.; Hatanaka, S.; Toda, Y.; Ogawa, D.; Ichikawa, H.; Ishimaru, Y.; Tashita, R.; Suzuki, T.; Ueda, M.; et al. Elevated levels of CYP94 family gene expression alleviate the jasmonate response and enhance salt tolerance in rice. Plant Cell Physiol. 2015, 56, 779–789. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koo, A.; Cooke, T.F.; Howe, G.A. Cytochrome P450 CYP94B3 mediates catabolism and inactivation of the plant hormone jasmonoyl-L-isoleucine. Proc. Natl. Acad. Sci. USA 2011, 108, 9298–9303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Erice, G.; Ruíz-Lozano, J.M.; Zamarreño, Á.M.; García-Mina, J.M.; Aroca, R. Transcriptomic analysis reveals the importance of JA-Ile turnover in the response of Arabidopsis plants to plant growth promoting rhizobacteria and salinity. Environ. Exp. Bot. 2017, 143, 10–19. [Google Scholar] [CrossRef] [Green Version]
- Pavlović, I.; Pěnčík, A.; Novák, O.; Vujčić, V.; Brkanac, S.R.; Lepeduš, H.; Strnad, M.; Salopek-Sondi, B. Short-term salt stress in Brassica rapa seedlings causes alterations in auxin metabolism. Plant Physiol. Biochem. 2018, 125, 74–84. [Google Scholar] [CrossRef]
- Karlova, R.; van Haarst, J.; Maliepaard, C.; van de Geest, H.; Bovy, A.G.; Lammers, M.; Angenent, G.C.; de Maagd, R.A. Identification of microRNA targets in tomato fruit development using high-throughput sequencing and degradome analysis. J. Exp. Bot. 2013, 64, 1863–1878. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Zou, W.; Xiao, Y.; Cheng, L.; Liu, Y.; Gao, S.; Shi, Z.; Jiang, Y.; Qi, M.; Xu, T.; et al. MicroRNA1917 targets CTR4 splice variants to regulate ethylene responses in tomato. J. Exp. Bot. 2018, 69, 1011–1025. [Google Scholar] [CrossRef] [Green Version]
- Zhao, H.; Yin, C.; Ma, B.; Chen, S.; Zhang, J. Ethylene signaling in rice and Arabidopsis: New regulators and mechanisms. J. Integr. Plant Biol. 2020, 63, 102–125. [Google Scholar] [CrossRef]
- Ohta, M.; Guo, Y.; Halfter, U.; Zhu, J.-K. A novel domain in the protein kinase SOS2 mediates interaction with the protein phosphatase 2C ABI2. Proc. Natl. Acad. Sci. USA 2003, 100, 11771–11776. [Google Scholar] [CrossRef] [Green Version]
- Li, G.; Meng, X.; Wang, R.; Mao, G.; Han, L.; Liu, Y.; Zhang, S. Dual-Level Regulation of ACC Synthase Activity by MPK3/MPK6 Cascade and Its Downstream WRKY Transcription Factor during Ethylene Induction in Arabidopsis. PLoS Genet. 2012, 8, e1002767. [Google Scholar] [CrossRef]
- Achard, P.; Cheng, H.; De Grauwe, L.; Decat, J.; Schoutteten, H.; Moritz, T.; Van Der Straeten, D.; Peng, J.; Harberd, N.P. Integration of plant responses to environmentally activated phytohormonal signals. Science 2006, 311, 91–94. [Google Scholar] [CrossRef]
- Chichkova, S.; Arellano, J.; Vance, C.P.; Hernández, G. Transgenic tobacco plants that overexpress alfalfa NADH-glutamate synthase have higher carbon and nitrogen content. J. Exp. Bot. 2001, 52, 2079–2087. [Google Scholar] [CrossRef]
- Abdel-Ghany, S.E.; Pilon, M. MicroRNA-mediated systemic down-regulation of copper protein expression in response to low copper availability in Arabidopsis. J. Biol. Chem. 2008, 283, 15932–15945. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trevisan, S.; Nonis, A.; Begheldo, M.; Manoli, A.; Palme, K.; Caporale, G.; Ruperti, B.; Quaggiotti, S. Expression and tissue-specific localization of nitrate-responsive miRNAs in roots of maize seedlings. Plant Cell Environ. 2012, 35, 1137–1155. [Google Scholar] [CrossRef]
- Lancien, M.; Martin, M.; Hsieh, M.H.; Leustek, T.; Goodman, H.; Coruzzi, G.M. Arabidopsis glt1-T mutant defines a role for NADH-GOGAT in the non-photorespiratory ammonium assimilatory pathway. Plant J. 2002, 29, 347–358. [Google Scholar] [CrossRef] [PubMed]
- Shang, Y.; Yan, L.; Liu, Z.-Q.; Cao, Z.; Mei, C.; Xin, Q.; Wu, F.-Q.; Wang, X.-F.; Du, S.-Y.; Jiang, T.; et al. The Mg-Chelatase H Subunit ofArabidopsisAntagonizes a Group of WRKY Transcription Repressors to Relieve ABA-Responsive Genes of Inhibition. Plant Cell 2010, 22, 1909–1935. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, T.; Zhang, X.-F.; Wang, X.-F.; Zhang, D.-P. Arabidopsis 3-ketoacyl-CoA thiolase-2 (KAT2), an enzyme of fatty acid β-oxidation, is involved in ABA signal transduction. Plant Cell Physiol. 2011, 52, 528–538. [Google Scholar] [CrossRef] [Green Version]
- Schilling, M. miRDeep2, Version 0.1.3; GitHub: Berlin, Germany, 2019.
- Robinson, M.D. edgeR, Version 3.24; Bioinformatics: Sydney, Australia, 2010.
- Premier Biosoft. Primer5, Version 0.11.3; PRIMER-E: Plymouth, UK, 2006.
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Wang, Z.; Wang, B.; Wang, J.; Xu, R.; Yang, T.; Huang, S.; Yu, Q.; Wang, H.; Gao, J. Identification and Characterization of Long Non-Coding RNA in Tomato Roots under Salt Stress. bioRxiv 2021, 11, 468377. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. Fastp, version 0.20.0; GitHub: Shenzhen, China, 2019.
- Kim, D.; Paggi, J.; Park, C.; Bennett, C.; Salzberg, S. HISAT2, Version 2.1.0; Nature Methods: Dallas, TX, USA, 2015.
- Consortium, T.G. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 2012, 485, 635. [Google Scholar] [CrossRef] [Green Version]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.; Rinn, J.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Love, M.; Huber, W.; Anders, S. DESeq2, Version 1.24.0; Bioinformatics: Boston, MA, USA, 2014.
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.; Wang, J.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape, Version 3.7.2; Genome Research: Seattle, WA, USA, 2003.
- Brunson, J. ggalluvial, Version 0.11.3; The Journal of Open Source Software: Minneapolis, MN, USA, 2020.





| Samples 1 | Raw Reads | Clean Reads | Length (18–30 nt) | Q20 (%) | Q30 (%) | GC (%) |
|---|---|---|---|---|---|---|
| MCK1 | 19,164,692 | 18,960,794 | 16,466,635 | 99.38 | 98.35 | 51 |
| MCK2 | 18,813,751 | 18,678,235 | 13,675,379 | 99.59 | 98.72 | 52 |
| PCK1 | 18,479,610 | 18,396,518 | 13,819,122 | 99.56 | 98.64 | 55 |
| PCK2 | 20,555,858 | 20,428,981 | 17,783,906 | 99.59 | 98.70 | 54 |
| MT1 | 21,729,061 | 21,590,130 | 15,863,323 | 99.55 | 98.62 | 50 |
| MT2 | 26,172,258 | 25,926,977 | 21,979,146 | 99.50 | 98.52 | 49 |
| PT1 | 19,849,787 | 19,700,143 | 15,616,279 | 99.57 | 98.65 | 52 |
| PT2 | 19,518,318 | 19,345,616 | 13,681,724 | 99.58 | 98.69 | 51 |
| Samples 1 | Read Number | Base Number | GC Content (%) | Q30 (%) |
|---|---|---|---|---|
| M82-CK1 | 136,735,222 | 10,287,608,907 | 45 | 89.42 |
| M82-CK2 | 122,505,086 | 9,237,784,418 | 45 | 87.07 |
| M82-CK3 | 127,746,338 | 9,622,066,421 | 45 | 88.25 |
| P-CK1 | 123,116,402 | 9,272,390,810 | 44 | 89.71 |
| P-CK2 | 132,255,878 | 9,958,938,470 | 43 | 88.65 |
| P-CK3 | 128,380,664 | 9,668,085,519 | 43 | 89.25 |
| M82-T1 | 151,887,684 | 11,430,890,506 | 45 | 89.05 |
| M82-T2 | 147,518,380 | 11,100,219,513 | 47 | 88.41 |
| M82-T3 | 150,787,186 | 11,347,268,749 | 46 | 88.80 |
| P-T1 | 110,834,866 | 8,292,641,368 | 43 | 89.71 |
| P-T2 | 160,240,518 | 11,989,101,650 | 43 | 86.94 |
| P-T3 | 148,489,644 | 11,109,924,975 | 43 | 88.08 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Z.; Li, N.; Yu, Q.; Wang, H. Genome-Wide Characterization of Salt-Responsive miRNAs, circRNAs and Associated ceRNA Networks in Tomatoes. Int. J. Mol. Sci. 2021, 22, 12238. https://doi.org/10.3390/ijms222212238
Wang Z, Li N, Yu Q, Wang H. Genome-Wide Characterization of Salt-Responsive miRNAs, circRNAs and Associated ceRNA Networks in Tomatoes. International Journal of Molecular Sciences. 2021; 22(22):12238. https://doi.org/10.3390/ijms222212238
Chicago/Turabian StyleWang, Zhongyu, Ning Li, Qinghui Yu, and Huan Wang. 2021. "Genome-Wide Characterization of Salt-Responsive miRNAs, circRNAs and Associated ceRNA Networks in Tomatoes" International Journal of Molecular Sciences 22, no. 22: 12238. https://doi.org/10.3390/ijms222212238
APA StyleWang, Z., Li, N., Yu, Q., & Wang, H. (2021). Genome-Wide Characterization of Salt-Responsive miRNAs, circRNAs and Associated ceRNA Networks in Tomatoes. International Journal of Molecular Sciences, 22(22), 12238. https://doi.org/10.3390/ijms222212238






