Chemical Exposures Affect Innate Immune Response to SARS-CoV-2
Abstract
:1. Introduction
2. Results
2.1. Monocytes
2.1.1. Top CSSG for Monocytes
2.1.2. Genes Affected by Chemicals and COVID-19
2.2. NK Cell
2.2.1. Top CSSG for NK-Cells
2.2.2. Genes Affected by Chemicals and COVID-19
2.3. T-Cells
2.3.1. Top CSSG for T-Cells
2.3.2. Genes Affected by Chemicals and COVID-19
2.4. B-Cells
2.4.1. Top CSSG for B-Cells
2.4.2. Genes Affected by Chemicals and COVID-19
2.5. Dendritic Cells
2.5.1. Top CSSG for Dendritic Cell
2.5.2. Genes Affected by Chemicals and COVID-19
2.6. Top Sensitive Overlapping Genes in Multiple Cell Types
3. Discussion
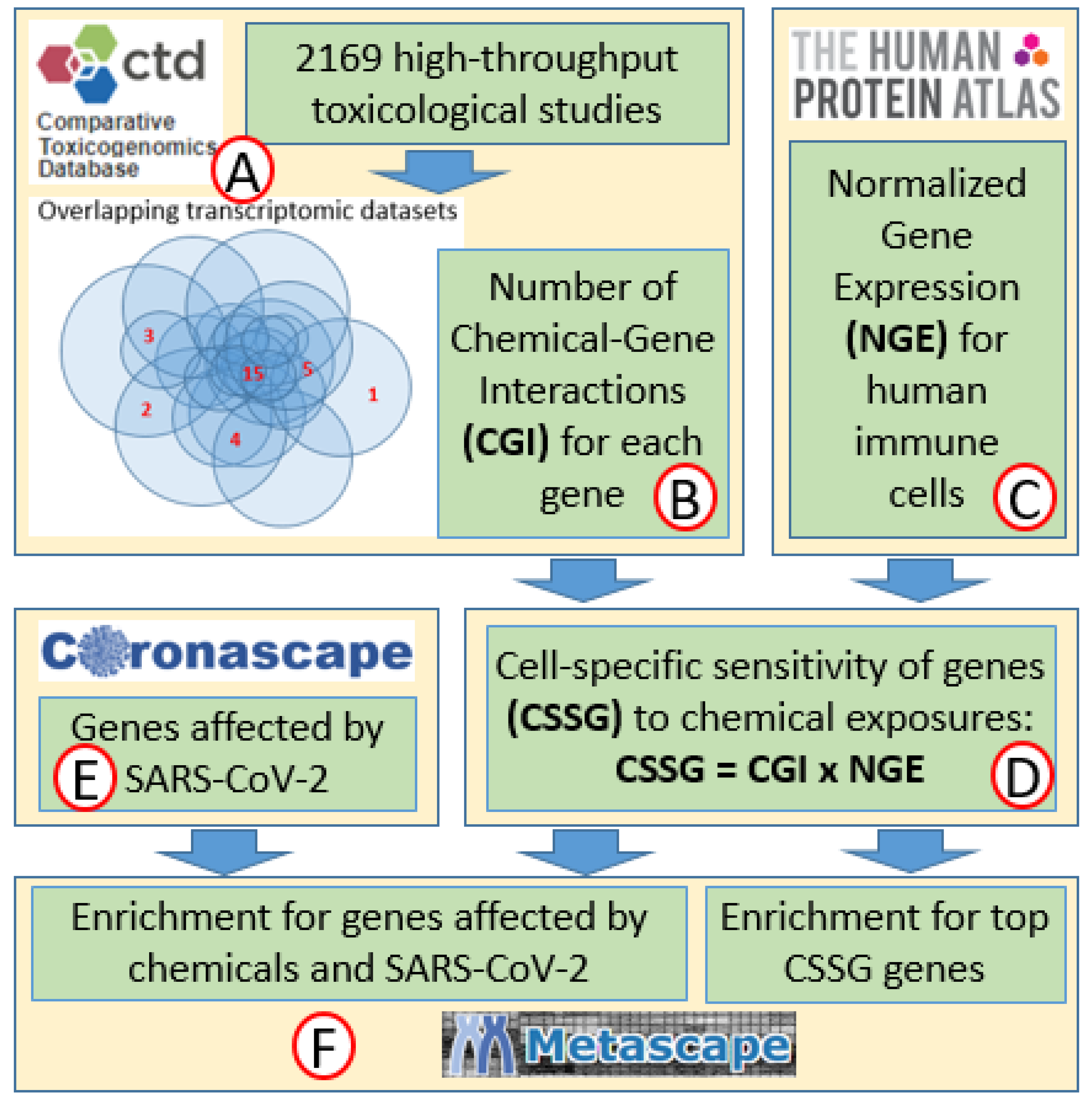
4. Materials and Methods
4.1. Sensitivity of Genes to Chemical Exposures
4.2. Sensitivities of Genes to Chemical Exposures Normalized for Expression in Different Cell Types
4.3. Genes Affected by SARS-CoV-2 in Human Immune Cells
4.4. Overlap of Top CSSG Genes with Genes Affected by COVID-19
4.5. Enrichment Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Morens, D.M.; Fauci, A.S. Emerging Pandemic Diseases: How We Got to COVID-19. Cell 2020, 182, 1077–1092. [Google Scholar] [CrossRef]
- World Health Organization. Coronavirus Disease (COVID-19). Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019 (accessed on 7 October 2021).
- Wu, X.; Nethery, R.C.; Sabath, M.B.; Braun, D.; Dominici, F. Exposure to air pollution and COVID-19 mortality in the United States. medRxiv 2020. [Google Scholar] [CrossRef]
- Ali, N.; Islam, F. The Effects of Air Pollution on COVID-19 Infection and Mortality—A Review on Recent Evidence. Front. Public Health 2020, 8, 580057. [Google Scholar] [CrossRef]
- Bashir, M.F.; Ma, B.J.; Bilal; Komal, B.; Bashir, M.A.; Farooq, T.H.; Iqbal, N.; Bashir, M. Correlation between environmental pollution indicators and COVID-19 pandemic: A brief study in Californian context. Environ. Res. 2020, 187, 109652. [Google Scholar] [CrossRef]
- Bilal; Bashir, M.F.; Benghoul, M.; Numan, U.; Shakoor, A.; Komal, B.; Bashir, M.A.; Bashir, M.; Tan, D. Environmental pollution and COVID-19 outbreak: Insights from Germany. Air Qual. Atmos. Health 2020, 13, 1385–1394. [Google Scholar] [CrossRef] [PubMed]
- Bourdrel, T.; Annesi-Maesano, I.; Alahmad, B.; Maesano, C.N.; Bind, M.-A. The impact of outdoor air pollution on COVID-19: A review of evidence from in vitro, animal, and human studies. Eur. Respir. Rev. 2021, 30, 200242. [Google Scholar] [CrossRef] [PubMed]
- Espejo, W.; Celis, J.E.; Chiang, G.; Bahamonde, P. Environment and COVID-19: Pollutants, impacts, dissemination, management and recommendations for facing future epidemic threats. Sci. Total Environ. 2020, 747, 141314. [Google Scholar] [CrossRef]
- Schintler, L.; Kulkarni, R.; McNeely, C.L.; Haynes, K.E. Environmental and Occupational Exposure to Toxic Industrial Chemicals and COVID-19: An Exploratory Analysis of United States Counties; Social Science Research Network: Rochester, NY, USA, 2020. [Google Scholar]
- Li, D.; Sangion, A.; Li, L. Evaluating consumer exposure to disinfecting chemicals against coronavirus disease 2019 (COVID-19) and associated health risks. Environ. Int. 2020, 145, 106108. [Google Scholar] [CrossRef] [PubMed]
- Samara, F.; Badran, R.; Dalibalta, S. Are Disinfectants for the Prevention and Control of COVID-19 Safe? Health Secur. 2020, 18, 496–498. [Google Scholar] [CrossRef]
- Yari, S.; Moshammer, H.; Asadi, A.F.; Jarrahi, A.M. Side Effects of Using Disinfectants to Fight COVID-19. Asian Pac. J. Environ. Cancer 2020, 3, 9–13. [Google Scholar] [CrossRef]
- Grandjean, P.; Timmermann, C.A.G.; Kruse, M.; Nielsen, F.; Vinholt, P.J.; Boding, L.; Heilmann, C.; Mølbak, K. Severity of COVID-19 at elevated exposure to perfluorinated alkylates. PLoS ONE 2020, 15, e0244815. [Google Scholar] [CrossRef]
- Reyes, M.S.S.; Medina, P.M.B. Environmental pollutant exposure can exacerbate COVID-19 neurologic symptoms. Med. Hypotheses 2020, 144, 110136. [Google Scholar] [CrossRef]
- Suvorov, A.; Salemme, V.; McGaunn, J.; Poluyanoff, A.; Teffera, M.; Amir, S. Unbiased approach for the identification of molecular mechanisms sensitive to chemical exposures. Chemosphere 2021, 262, 128362. [Google Scholar] [CrossRef]
- Veraldi, A.; Costantini, A.S.; Bolejack, V.; Miligi, L.; Vineis, P.; van Loveren, H. Immunotoxic effects of chemicals: A matrix for occupational and environmental epidemiological studies. Am. J. Ind. Med. 2006, 49, 1046–1055. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chernis, N.; Masschelin, P.; Cox, A.R.; Hartig, S.M. Bisphenol AF promotes inflammation in human white adipocytes. Am. J. Physiol.-Cell Physiol. 2020, 318, C63–C72. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Hong, F.; Zhou, J.-L.; Zhang, Y.-Q. Lung inflammation caused by long-term exposure to titanium dioxide in mice involving in NF-κB signaling pathway. J. Biomed. Mater. Res. A 2017, 105, 720–727. [Google Scholar] [CrossRef] [PubMed]
- Negi, C.K.; Khan, S.; Dirven, H.; Bajard, L.; Bláha, L. Flame Retardants-Mediated Interferon Signaling in the Pathogenesis of Nonalcoholic Fatty Liver Disease. Int. J. Mol. Sci. 2021, 22, 4282. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Li, C.; Liu, W.; Jin, Y. Effect of exposure to volatile organic compounds (VOCs) on airway inflammatory response in mice. J. Toxicol. Sci. 2012, 37, 739–748. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lowery, S.A.; Sariol, A.; Perlman, S. Innate immune and inflammatory responses to SARS-CoV-2: Implications for COVID-19. Cell Host Microbe 2021, 29, 1052–1062. [Google Scholar] [CrossRef]
- Szabo, P.A.; Dogra, P.; Gray, J.I.; Wells, S.B.; Connors, T.J.; Weisberg, S.P.; Krupska, I.; Matsumoto, R.; Poon, M.M.L.; Idzikowski, E.; et al. Longitudinal profiling of respiratory and systemic immune responses reveals myeloid cell-driven lung inflammation in severe COVID-19. Immunity 2021, 54, 797–814.e6. [Google Scholar] [CrossRef]
- Acharya, D.; Liu, G.; Gack, M.U. Dysregulation of type I interferon responses in COVID-19. Nat. Rev. Immunol. 2020, 20, 397–398. [Google Scholar] [CrossRef] [PubMed]
- Broman, N.; Rantasärkkä, K.; Feuth, T.; Valtonen, M.; Waris, M.; Hohenthal, U.; Rintala, E.; Karlsson, A.; Marttila, H.; Peltola, V.; et al. IL-6 and other biomarkers as predictors of severity in COVID-19. Ann. Med. 2021, 53, 410–412. [Google Scholar] [CrossRef] [PubMed]
- Mulchandani, R.; Lyngdoh, T.; Kakkar, A.K. Deciphering the COVID-19 cytokine storm: Systematic review and meta-analysis. Eur. J. Clin. Investig. 2021, 51, e13429. [Google Scholar] [CrossRef]
- Akamatsu, M.A.; de Castro, J.T.; Takano, C.Y.; Ho, P.L. Off balance: Interferons in COVID-19 lung infections. EBioMedicine 2021, 73, 103642. [Google Scholar] [CrossRef] [PubMed]
- Suvorov, A.; Salemme, V.; McGaunn, J.; Poluyanoff, A.; Amir, S. Data on chemical-gene interactions and biological categories enriched with genes sensitive to chemical exposures. Data Brief 2020, 33, 106398. [Google Scholar] [CrossRef]
- Ayoub Meo, S.; Adnan Abukhalaf, A.; Sami, W.; Hoang, T.D. Effect of environmental pollution PM2.5, carbon monoxide, and ozone on the incidence and mortality due to SARS-CoV-2 infection in London, United Kingdom. J. King Saud Univ.-Sci. 2021, 33, 101373. [Google Scholar] [CrossRef]
- Gupta, A.; Bherwani, H.; Gautam, S.; Anjum, S.; Musugu, K.; Kumar, N.; Anshul, A.; Kumar, R. Air pollution aggravating COVID-19 lethality? Exploration in Asian cities using statistical models. Environ. Dev. Sustain. 2021, 23, 6408–6417. [Google Scholar] [CrossRef]
- Linares, C.; Culqui, D.; Belda, F.; López-Bueno, J.A.; Luna, Y.; Sánchez-Martínez, G.; Hervella, B.; Díaz, J. Impact of environmental factors and Sahara dust intrusions on incidence and severity of COVID-19 disease in Spain. Effect in the first and second pandemic waves. Environ. Sci. Pollut. Res. 2021, 28, 51948–51960. [Google Scholar] [CrossRef]
- Haller, O.; Kochs, G.; Weber, F. Interferon, Mx, and viral countermeasures. Cytokine Growth Factor Rev. 2007, 18, 425–433. [Google Scholar] [CrossRef]
- Kaffash Farkhad, N.; Reihani, H.; Sedaghat, A.; Moghadam, A.A.; Moghadam, A.B.; Tavakol-Afshari, J. Are mesenchymal stem cells able to manage cytokine storm in COVID-19 patients? A review of recent studies. Regen. Ther. 2021, 18, 152–160. [Google Scholar] [CrossRef]
- Miao, Y.; Fan, L.; Li, J.-Y. Potential Treatments for COVID-19 Related Cytokine Storm-Beyond Corticosteroids. Front. Immunol. 2020, 11, 1445. [Google Scholar] [CrossRef] [PubMed]
- Ruan, Q.; Yang, K.; Wang, W.; Jiang, L.; Song, J. Clinical predictors of mortality due to COVID-19 based on an analysis of data of 150 patients from Wuhan, China. Intensive Care Med. 2020, 46, 846–848. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, D.; Li, H.; Lu, X.-X.; Xiao, H.; Ren, J.; Zhang, F.-R.; Liu, Z.-S. Clinical features of severe pediatric patients with coronavirus disease 2019 in Wuhan: A single center’s observational study. World J. Pediatr. 2020, 16, 251–259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blanco-Melo, D.; Nilsson-Payant, B.E.; Liu, W.-C.; Uhl, S.; Hoagland, D.; Møller, R.; Jordan, T.X.; Oishi, K.; Panis, M.; Sachs, D.; et al. Imbalanced Host Response to SARS-CoV-2 Drives Development of COVID-19. Cell 2020, 181, 1036–1045.e9. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [Green Version]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Davis, A.P.; Grondin, C.J.; Johnson, R.J.; Sciaky, D.; McMorran, R.; Wiegers, J.; Wiegers, T.C.; Mattingly, C.J. The Comparative Toxicogenomics Database: Update 2019. Nucleic Acids Res. 2019, 47, D948–D954. [Google Scholar] [CrossRef]
- Uhlén, M.; Fagerberg, L.; Hallström, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, Å.; Kampf, C.; Sjöstedt, E.; Asplund, A.; et al. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef]
- Suvorov, A. Simple Method for Cutoff Point Identification In Descriptive High-Throughput Biological Studies. Preprints 2021. [Google Scholar] [CrossRef]
- Wilk, A.J.; Rustagi, A.; Zhao, N.Q.; Roque, J.; Martinez-Colon, G.J.; McKechnie, J.L.; Ivison, G.T.; Ranganath, T.; Vergara, R.; Hollis, T.; et al. A single-cell atlas of the peripheral immune response to severe COVID-19; Infectious Diseases (except HIV/AIDS). Nat. Med. 2020, 26, 1070–1076. [Google Scholar] [CrossRef]
- Zhang, J.-Y.; Wang, X.-M.; Xing, X.; Xu, Z.; Zhang, C.; Song, J.-W.; Fan, X.; Xia, P.; Fu, J.-L.; Wang, S.-Y.; et al. Single-cell landscape of immunological responses in patients with COVID-19. Nat. Immunol. 2020, 21, 1107–1118. [Google Scholar] [CrossRef] [PubMed]
- GEO Accession Viewer. Available online: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE148163 (accessed on 7 October 2021).
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Khodabakhshi, A.H.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 2019, 10, 1523. [Google Scholar] [CrossRef] [PubMed]

| NK Cells (649 Top CSSG) | T Cells (558 Top CSSG) | Monocytes (409 Top CSSG) | B Cells (408 Top CSSG) | Dendritic Cells (510 Top CSSG) | |
|---|---|---|---|---|---|
| 1 | mRNA catabolic process | Ribosome, cytoplasmic | Leukocyte degranulation | Ribosome, cytoplasmic | Regulation of expression of SLITs and ROBOs |
| 2 | Cytokine signaling in immune system | Cytokine signaling in immune system | Cellular responses to external stimuli | VEGFA-VEGFR2 signaling pathway | Leukocyte activation involved in immune response |
| 3 | VEGFA-VEGFR2 signaling pathway | VEGFA-VEGFR2 signaling pathway | Cytokine signaling in immune system | TRBP containing complex | Oxidation-reduction process |
| 4 | Oxidation-reduction process | Oxidation-reduction process | VEGFA-VEGFR2 signaling pathway | Cytokine signaling in immune system | VEGFA-VEGFR2 signaling pathway |
| 5 | Leukocyte activation involved in immune response | Positive regulation of cell death | Positive regulation of cell death | Nucleoside monophosphate metabolic process | Cytokine signaling in immune system |
| 6 | Ribonucleoprotein complex biogenesis | Apoptotic signaling pathway | Apoptotic signaling pathway | Ribosome assembly | TRBP containing complex |
| 7 | Protein processing in the endoplasmic reticulum | Leukocyte activation involved in immune response | Oxidation-reduction process | Regulation of translation | Apoptotic signaling pathway |
| 8 | Regulation of apoptotic signaling pathway | Ribonucleoprotein complex assembly | Nuclear receptors meta-pathway | Protein folding | Response to toxic substance |
| 9 | Protein folding | Regulation of cellular amide metabolic process | Cellular response to interleukin-12 | Apoptotic signaling pathway | Mitochondrion organization |
| 10 | Regulation of cellular amide metabolic process | Epstein–Barr virus infection | Hemostasis | Positive regulation of cell death | Positive regulation of cell death |
| NK Cells | T Cells | Monocytes | B Cells | Dendritic Cells | |
|---|---|---|---|---|---|
| 1 | STAT1 | TPT1 | S100A8 | MS4A1 | HLA-DQB1 |
| 2 | ISG15 | EEF1A1 | PLAC8 | CD74 | HSPA5 |
| 3 | ACTB | STAT1 | IFITM3 | JCHAIN; IGJ | ISG15 |
| 4 | GNLY | MX1 | PABPC1 | HSPA5 | PABPC1 |
| 5 | EEF2 | EEF2 | NAP1L1 | CALR | EEF2 |
| 6 | JAK1 | PABPC1 | S100A9 | HSP90B1 | TRIM22 |
| 7 | SP100 | EIF4B | EEF1A1 | PDIA6 | FKBP5 |
| 8 | PABPC1 | ISG15 | EEF2 | PDIA4 | PLSCR1 |
| 9 | TPT1 | IL7R | CD14 | CD79B | HSP90AB1 |
| 10 | ISG20 | PIM1 | IFI6 | CD79A | HSPA8 |
| NK Cells (24 Gene Overlap) | T Cells (55 Gene Overlap) | Monocytes (124 Gene Overlap) | B Cells (65 Gene Overlap) | Dendritic Cells (20 Gene Overlap) | |
|---|---|---|---|---|---|
| 1 | Regulation of multi-organism process | Cytokine signaling in immune system | Neutrophil degranulation | Regulation of multi-organism process | Regulation of viral life cycle |
| 2 | Interferon signaling | Chaperone-mediated autophagy | Cytokine signaling in immune system | Protein folding | Antigen processing and presentation |
| 3 | T cell mediated cytotoxicity | Translation factors | Regulation of multi-organism process | VEGFA-VEGFR2 signaling pathway | P2X7 receptor signaling complex |
| 4 | PID IL12 2PATHWAY | Regulation of multi-organism process | Defense response to other organisms | Translation factors | Viral entry into host cell |
| 5 | Protein methylation | Antigen processing and presentation | Activation of immune response | Regulation of myeloid cell differentiation | Response to virus |
| 6 | Antigen processing and presentation | Regulation of hemopoiesis | Apoptotic signaling pathway | B cell activation | Translation factors |
| 7 | Translation factors | I-kappaB kinase/NF-kappaB signaling | Response to inorganic substance | Cytokine signaling in immune system | Response to interferon-gamma |
| 8 | Negative regulation of binding | Epstein–Barr virus infection | Regulation of cytokine production | Chaperone-mediated protein folding | Negative regulation of the cellular component organization |
| 9 | Homotypic cell–cell adhesion | Response to interferon-gamma | Pertussis | Translation | RNA degradation |
| 10 | Allograft rejection | H2AX complex, isolated from cells without IR exposure | Antigen processing and presentation | Interaction with symbiont | Negative regulation of intrinsic apoptotic signaling pathway |
| Cell Type | Number of Datasets | Genes Affected by COVID-19 in More than One Dataset | Sources |
|---|---|---|---|
| Dendritic cells | 14 | 53 | [42] |
| NK cells | 19 | 81 | [42,43] |
| B-cells | 23 | 248 | [42,43,44] |
| Monocytes | 32 | 278 | [42,43] |
| T-cells | 35 | 116 | [42,43] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arowolo, O.; Pobezinsky, L.; Suvorov, A. Chemical Exposures Affect Innate Immune Response to SARS-CoV-2. Int. J. Mol. Sci. 2021, 22, 12474. https://doi.org/10.3390/ijms222212474
Arowolo O, Pobezinsky L, Suvorov A. Chemical Exposures Affect Innate Immune Response to SARS-CoV-2. International Journal of Molecular Sciences. 2021; 22(22):12474. https://doi.org/10.3390/ijms222212474
Chicago/Turabian StyleArowolo, Olatunbosun, Leonid Pobezinsky, and Alexander Suvorov. 2021. "Chemical Exposures Affect Innate Immune Response to SARS-CoV-2" International Journal of Molecular Sciences 22, no. 22: 12474. https://doi.org/10.3390/ijms222212474






