Selectivity Determinants of RHO GTPase Binding to IQGAPs
Abstract
1. Introduction
2. Results
2.1. IQGAP1/2 Selectively Bind CDC42 and RAC1-Like Members of the RHO Family
2.2. RAC2 Exhibited the Highest Affinity for IQGAP1
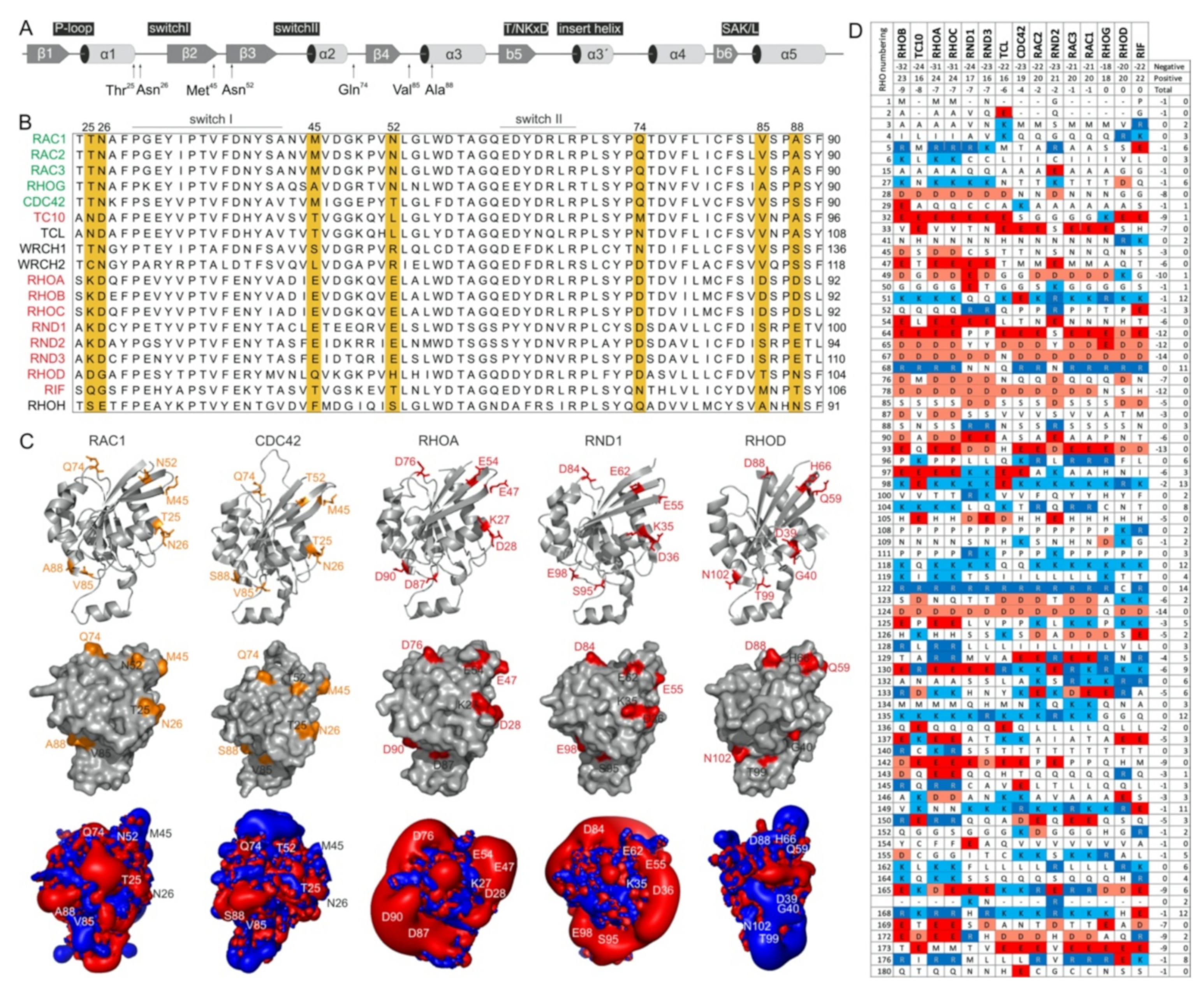
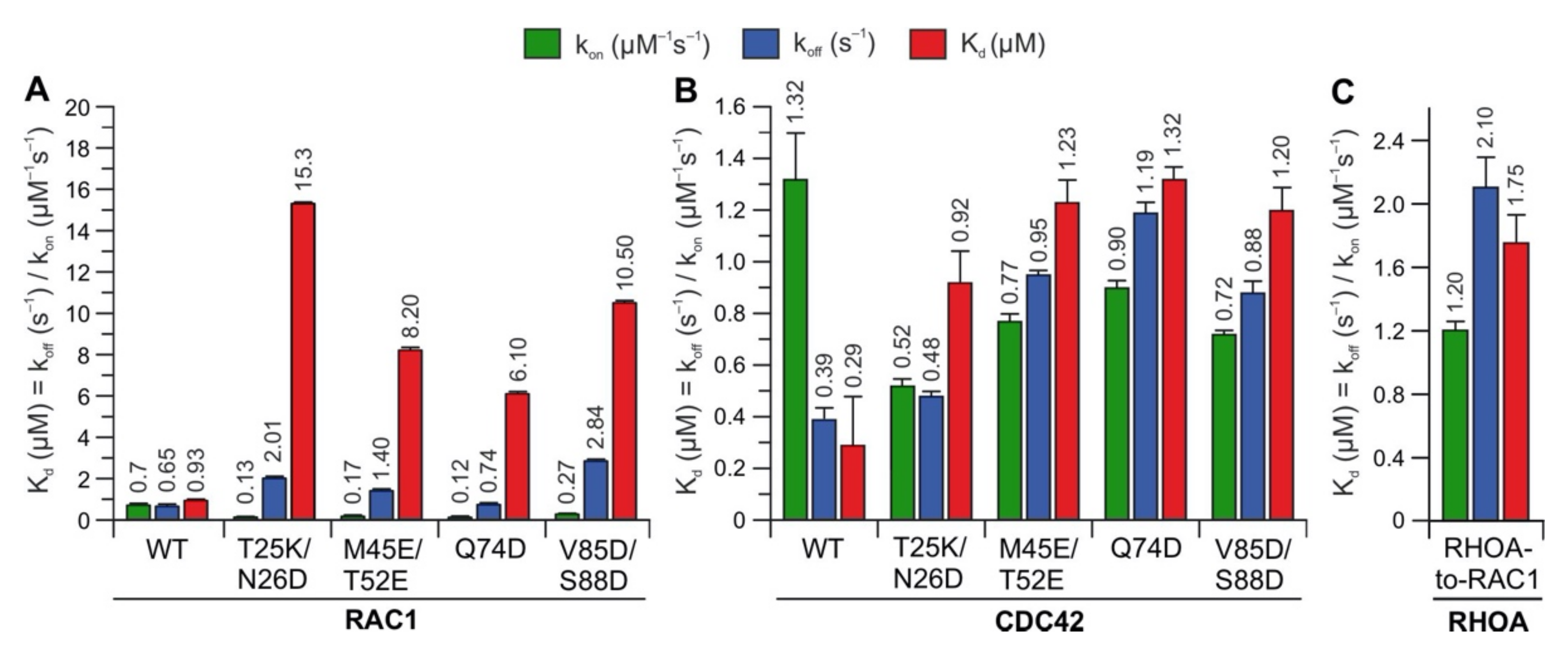
2.3. Potential Hotspots for IQGAP Binding Appear Outside the Switch Regions
2.4. PAK1, p50GAP, and DOCK2 Compete with IQGAP1 for Binding RAC1
2.5. IQGAP1 Binding Hotspots Significantly Vary among RHO GTPases
3. Discussion
4. Materials and Methods
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Abel, A.M.; Schuldt, K.M.; Rajasekaran, K.; Hwang, D.; Riese, M.J.; Rao, S.; Thakar, M.S.; Malarkannan, S. IQGAP1: Insights into the function of a molecular puppeteer. Mol. Immunol. 2015, 65, 336–349. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.; Anderson, R.A. IQGAP1 is a phosphoinositide effector and kinase scaffold. Adv. Biol. Regul. 2016, 60, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Hedman, A.C.; Smith, J.M.; Sacks, D.B. The biology of IQGAP proteins: Beyond the cytoskeleton. EMBO Rep. 2015, 16, 427–446. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.M.; Hedman, A.C.; Sacks, D.B. IQGAPs choreograph cellular signaling from the membrane to the nucleus. Trends Cell Biol. 2015, 25, 171–184. [Google Scholar] [CrossRef]
- Watanabe, T.; Wang, S.; Kaibuchi, K. IQGAPs as Key Regulators of Actin-cytoskeleton Dynamics Mini-review and Review. Cell Struct. Funct. 2015, 40, 69–77. [Google Scholar] [CrossRef]
- Tanos, B.E.; Yeaman, C.; Rodriguez-Boulan, E. An emerging role for IQGAP1 in tight junction control. Small GTPases 2018, 9, 375–383. [Google Scholar] [CrossRef]
- Nussinov, R.; Zhang, M.; Tsai, C.J.; Jang, H. Calmodulin and IQGAP1 activation of PI3Kα and Akt in KRAS, HRAS and NRAS-driven cancers. Biochim. Biophys. Acta Mol. Basis Dis. 2018, 1864, 2304–2314. [Google Scholar] [CrossRef]
- White, C.D.; Brown, M.D.; Sacks, D.B. IQGAPs in cancer: A family of scaffold proteins underlying tumorigenesis. FEBS Lett. 2009, 583, 1817–1824. [Google Scholar] [CrossRef]
- Roy, M.; Li, Z.; Sacks, D.B. IQGAP1 Binds ERK2 and Modulates Its Activity. J. Biol. Chem. 2004, 279, 17329–17337. [Google Scholar] [CrossRef]
- Roy, M.; Li, Z.; Sacks, D.B. IQGAP1 Is a Scaffold for Mitogen-Activated Protein Kinase Signaling. Mol. Cell. Biol. 2005, 25, 7940–7952. [Google Scholar] [CrossRef]
- Ren, J.G.; Li, Z.; Sacks, D.B. IQGAP1 modulates activation of B-Raf. Proc. Natl. Acad. Sci. USA 2007, 104, 10465–10469. [Google Scholar] [CrossRef]
- Benseñor, L.B.; Kan, H.M.; Wang, N.; Wallrabe, H.; Davidson, L.A.; Cai, Y.; Schafer, D.A.; Bloom, G.S. IQGAP1 regulates cell motility by linking growth factor signaling to actin assembly. J. Cell Sci. 2007, 120, 658–669. [Google Scholar] [CrossRef]
- Le Clainche, C.; Schlaepfer, D.; Ferrari, A.; Klingauf, M.; Grohmanova, K.; Veligodskiy, A.; Didry, D.; Le, D.; Egile, C.; Carlier, M.F.; et al. IQGAP1 stimulates actin assembly through the N-wasp-Arp2/3 pathway. J. Biol. Chem. 2007, 282, 426–435. [Google Scholar] [CrossRef]
- Pelikan-Conchaudron, A.; Le Clainche, C.; Didry, D.; Carlier, M.F. The IQGAP1 protein is a calmodulin-regulated barbed end capper of actin filaments: Possible implications in its function in cell migration. J. Biol. Chem. 2011, 286, 35119–35128. [Google Scholar] [CrossRef]
- Usatyuk, P.V.; Gorshkova, I.A.; He, D.; Zhao, Y.; Kalari, S.K.; Garcia, J.G.N.; Natarajan, V. Phospholipase D-mediated Activation of IQGAP1 through Rac1 regulates hyperoxia-induced p47phox translocation and reactive oxygen species generation in lung endothelial cells. J. Biol. Chem. 2009, 284, 15339–15352. [Google Scholar] [CrossRef]
- Kaur, R.; Yuan, X.; Lu, M.L.; Balk, S.P. Increased PAK6 expression in prostate cancer and identification of PAK6 associated proteins. Prostate 2008, 68, 1510–1516. [Google Scholar] [CrossRef]
- Fukata, M.; Watanabe, T.; Noritake, J.; Nakagawa, M.; Yamaga, M.; Kuroda, S.; Matsuura, Y.; Iwamatsu, A.; Perez, F.; Kaibuchi, K. Rac1 and Cdc42 capture microtubules through IQGAP1 and CLIP-170. Cell 2002, 109, 873–885. [Google Scholar] [CrossRef]
- Watanabe, T.; Wang, S.; Noritake, J.; Sato, K.; Fukata, M.; Takefuji, M.; Nakagawa, M.; Izumi, N.; Akiyama, T.; Kaibuchi, K. Interaction with IQGAP1 links APC to Rac1, Cdc42, and actin filaments during cell polarization and migration. Dev. Cell 2004, 7, 871–883. [Google Scholar] [CrossRef]
- Mateer, S.C.; Morris, L.E.; Cromer, D.A.; Benseñor, L.B.; Bloom, G.S. Actin filament binding by a monomeric IQGAP1 fragment with a single calponin homology domain. Cell Motil. Cytoskeleton 2004, 58, 231–241. [Google Scholar] [CrossRef]
- Grohmanova, K.; Schlaepfer, D.; Hess, D.; Gutierrez, P.; Beck, M.; Kroschewski, R. Phosphorylation of IQGAP1 modulates its binding to Cdc42, revealing a new type of Rho-GTPase regulator. J. Biol. Chem. 2004, 279, 48495–48504. [Google Scholar] [CrossRef]
- Kurella, V.B.; Richard, J.M.; Parke, C.L.; LeCour, L.F.; Bellamy, H.D.; Worthylake, D.K. Crystal structure of the GTPase-activating protein-related domain from IQGAP1. J. Biol. Chem. 2009, 284, 14857–14865. [Google Scholar] [CrossRef]
- Li, Z.; McNulty, D.E.; Marler, K.J.M.; Lim, L.; Hall, C.; Annan, R.S.; Sacks, D.B. IQGAP1 promotes neurite outgrowth in a phosphorylation-dependent manner. J. Biol. Chem. 2005, 280, 13871–13878. [Google Scholar] [CrossRef]
- Nouri, K.; Fansa, E.K.; Amin, E.; Dvorsky, R.; Gremer, L.; Willbold, D.; Schmitt, L.; Timson, D.J.; Ahmadian, M.R. IQGAP1 interaction with RHO family proteins revisited kinetic and equilibrium evidence for multiple distinct binding sites. J. Biol. Chem. 2016, 291, 26364–26376. [Google Scholar] [CrossRef]
- Sila Ozdemir, E.; Jang, H.; Gursoy, A.; Keskin, O.; Li, Z.; Sacks, D.B.; Nussinov, R. Unraveling the molecular mechanism of interactions of the Rho GTPases Cdc42 and Rac1 with the scaffolding protein IQGAP2. J. Biol. Chem. 2018, 293, 3685–3699. [Google Scholar] [CrossRef]
- Wennerberg, K.; Der, C.J. Rho-family GTPases: It’s not only Rac and Rho (and I like it). J. Cell Sci. 2004, 117, 1301–1312. [Google Scholar] [CrossRef]
- Jaffe, A.B.; Hall, A. Rho GTPases: Biochemistry and biology. Annu. Rev. Cell Dev. Biol. 2005, 21, 247–269. [Google Scholar] [CrossRef]
- Haga, R.B.; Ridley, A.J. Rho GTPases: Regulation and roles in cancer cell biology. Small GTPases 2016, 7, 207–221. [Google Scholar] [CrossRef]
- Ahmadian, M.R.; Jaiswal, M.; Fansa, E.K.; Dvorsky, R. New insight into the molecular switch mechanism of human Rho family proteins: Shifting a paradigm. Biol. Chem. 2013, 394, 89–95. [Google Scholar] [CrossRef]
- Beljan, S.; Herak Bosnar, M.; Ćetković, H. Rho Family of Ras-Like GTPases in Early-Branching Animals. Cells 2020, 9, 2279. [Google Scholar] [CrossRef]
- Bishop, A.L.; Hall, A. Rho GTPases and their effector proteins. Biochem. J. 2000, 348, 241–255. [Google Scholar] [CrossRef]
- Dvorsky, R.; Ahmadian, M.R. Always look on the bright site of Rho: Structural implications for a conserved intermolecular interface. EMBO Rep. 2004, 5, 1130–1136. [Google Scholar] [CrossRef] [PubMed]
- Hall, A. Rho family GTPases. Biochem. Soc. Trans. 2012, 40, 1378–1382. [Google Scholar] [CrossRef] [PubMed]
- Elliott, S.F. Biochemical analysis of the interactions of IQGAP1 C-terminal domain with CDC42. World J. Biol. Chem. 2012, 3, 53. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, B.; Wang, Z.X.; Zheng, Y. Characterization of the interactions between the small GTPase Cdc42 and its GTPase-activating proteins and putative effectors: Comparison of kinetic properties of Cdc42 binding to the Cdc42-interactive domains. J. Biol. Chem. 1997, 272, 21999–22007. [Google Scholar] [CrossRef]
- Zhang, B.; Chernoff, J.; Zheng, Y. Interaction of Rac1 with GTPase-activating proteins and putative effectors. A comparison with Cdc42 and RhoA. J. Biol. Chem. 1998, 273, 8776–8782. [Google Scholar] [CrossRef]
- Owen, D.; Campbell, L.J.; Littlefield, K.; Evetts, K.A.; Li, Z.; Sacks, D.B.; Lowe, P.N.; Mott, H.R. The IQGAP1-Rac1 and IQGAP1-Cdc42 interactions: Interfaces differ between the complexes. J. Biol. Chem. 2008, 283, 1692–1704. [Google Scholar] [CrossRef]
- Mataraza, J.M.; Briggs, M.W.; Li, Z.; Frank, R.; Sacks, D.B. Identification and characterization of the Cdc42-binding site of IQGAP1. Biochem. Biophys. Res. Commun. 2003, 305, 315–321. [Google Scholar] [CrossRef]
- Li, R.; Debreceni, B.; Jia, B.; Gao, Y.; Tigyi, G.; Zheng, Y. Localization of the PAK1-, WASP-, and IQGAP1-specifying regions of Cdc42. J. Biol. Chem. 1999, 274, 29648–29654. [Google Scholar] [CrossRef]
- Lam, M.T.; Coppola, S.; Krumbach, O.H.F.; Prencipe, G.; Insalaco, A.; Cifaldi, C.; Brigida, I.; Zara, E.; Scala, S.; Di Cesare, S.; et al. A novel disorder involving dyshematopoiesis, inflammation, and HLH due to aberrant CDC42 function. J. Exp. Med. 2019, 216, 2778–2799. [Google Scholar] [CrossRef]
- Nouri, K.; Timson, D.J.; Ahmadian, M.R. New model for the interaction of IQGAP1 with CDC42 and RAC1. Small GTPases 2020, 11, 16–22. [Google Scholar] [CrossRef]
- LeCour, L.; Boyapati, V.K.; Liu, J.; Li, Z.; Sacks, D.B.; Worthylake, D.K. The Structural Basis for Cdc42-Induced Dimerization of IQGAPs. Structure 2016, 24, 1499–1508. [Google Scholar] [CrossRef]
- Identification of a Human RasGAP-Related Protein Containing Calmodulin-Binding Motifs. Available online: https://pubmed.ncbi.nlm.nih.gov/8051149/ (accessed on 26 February 2021).
- Zhang, B.; Zheng, Y. Negative regulation of Rho family GTPases Cdc42 and Rac2 by homodimer formation. J. Biol. Chem. 1998, 273, 25728–25733. [Google Scholar] [CrossRef]
- Jaiswal, M.; Gremer, L.; Dvorsky, R.; Haeusler, L.C.; Cirstea, I.C.; Uhlenbrock, K.; Ahmadian, M.R. Mechanistic insights into specificity, activity, and regulatory elements of the regulator of G-protein signaling (RGS)-containing Rho-specific guanine nucleotide exchange factors (GEFs) p115, PDZ-RhoGEF (PRG), and leukemia-associated RhoGEF (LARG). J. Biol. Chem. 2011, 286, 18202–18212. [Google Scholar] [CrossRef]
- Martinelli, S.; Krumbach, O.H.F.; Pantaleoni, F.; Coppola, S.; Amin, E.; Pannone, L.; Nouri, K.; Farina, L.; Dvorsky, R.; Lepri, F.; et al. Functional Dysregulation of CDC42 Causes Diverse Developmental Phenotypes. Am. J. Hum. Genet. 2018, 102, 309–320. [Google Scholar] [CrossRef]
- Haeusler, L.C.; Blumenstein, L.; Stege, P.; Dvorsky, R.; Ahmadian, M.R. Comparative functional analysis of the Rac GTPases. FEBS Lett. 2003, 555, 556–560. [Google Scholar] [CrossRef]
- Bhattacharya, M.; Sundaram, A.; Kudo, M.; Farmer, J.; Ganesan, P.; Khalifeh-Soltani, A.; Arjomandi, M.; Atabai, K.; Huang, X.; Sheppard, D. IQGAP1-dependent scaffold suppresses RhoA and inhibits airway smooth muscle contraction. J. Clin. Investig. 2014, 124, 4895–4898. [Google Scholar] [CrossRef]
- Casteel, D.E.; Turner, S.; Schwappacher, R.; Rangaswami, H.; Su-Yuo, J.; Zhuang, S.; Boss, G.R.; Pilz, R.B. Rho Isoform-specific Interaction with IQGAP1 promotes breast cancer cell proliferation and migration. J. Biol. Chem. 2012, 287, 38367–38378. [Google Scholar] [CrossRef]
- Fiegen, D.; Blumenstein, L.; Stege, P.; Vetter, I.R.; Ahmadian, M.R. Crystal structure of Rnd3/RhoE: Functional implications. FEBS Lett. 2002, 525, 100–104. [Google Scholar] [CrossRef]
- Fansa, E.K.; Dvorsky, R.; Zhang, S.C.; Fiegen, D.; Ahmadian, M.R. Interaction characteristics of Plexin-B1 with Rho family proteins. Biochem. Biophys. Res. Commun. 2013, 434, 785–790. [Google Scholar] [CrossRef]
- Lapouge, K.; Smith, S.J.M.; Walker, P.A.; Gamblin, S.J.; Smerdon, S.J.; Rittinger, K. Structure of the TPR domain of p67(phox) in complex with Rac·GTP. Mol. Cell 2000, 6, 899–907. [Google Scholar] [CrossRef]
- Ahmadian, M.R.; Hoffmann, U.; Goody, R.S.; Wittinghofer, A. Individual rate constants for the interaction of Ras proteins with GTPase-activating proteins determined by fluorescence spectroscopy. Biochemistry 1997, 36, 4535–4541. [Google Scholar] [CrossRef] [PubMed]
- D’Andrea, L.D.; Regan, L. TPR proteins: The versatile helix. Trends Biochem. Sci. 2003, 28, 655–662. [Google Scholar] [CrossRef] [PubMed]
- Diebold, B.A.; Bokoch, G.M. Molecular basis for Rac2 regulation of phagocyte NADPH oxidase. Nat. Immunol. 2001, 2, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, G.R.; Cerione, R.A. Rac inserts its way into the immune response. Nat. Immunol. 2001, 2, 194–196. [Google Scholar] [CrossRef]
- Dvorsky, R.; Blumenstein, L.; Vetter, I.R.; Ahmadian, M.R. Structural Insights into the Interaction of ROCKI with the Switch Regions of RhoA. J. Biol. Chem. 2004, 279, 7098–7104. [Google Scholar] [CrossRef]
- Sinha, N.; Smith-Gill, S. Electrostatics in Protein Binding and Function. Curr. Protein Pept. Sci. 2005, 3, 601–614. [Google Scholar] [CrossRef]
- Hemsath, L.; Dvorsky, R.; Fiegen, D.; Carlier, M.F.; Ahmadian, M.R. An electrostatic steering mechanism of Cdc42 recognition by Wiskott-Aldrich syndrome proteins. Mol. Cell 2005, 20, 313–324. [Google Scholar] [CrossRef]
- Vascon, F.; Gasparotto, M.; Giacomello, M.; Cendron, L.; Bergantino, E.; Filippini, F.; Righetto, I. Protein electrostatics: From computational and structural analysis to discovery of functional fingerprints and biotechnological design. Comput. Struct. Biotechnol. J. 2020, 18, 1774–1789. [Google Scholar] [CrossRef]
- Bivona, T.G.; Quatela, S.E.; Bodemann, B.O.; Ahearn, I.M.; Soskis, M.J.; Mor, A.; Miura, J.; Wiener, H.H.; Wright, L.; Saba, S.G.; et al. PKC regulates a farnesyl-electrostatic switch on K-Ras that promotes its association with Bcl-XL on mitochondria and induces apoptosis. Mol. Cell 2006, 21, 481–493. [Google Scholar] [CrossRef]
- Zhou, Y.; Hancock, J.F. A novel prenyl-polybasic domain code determines lipid-binding specificity of the K-Ras membrane anchor. Small GTPases 2020, 11, 220–224. [Google Scholar] [CrossRef]
- Connolly, A.; Gagnon, E. Electrostatic interactions: From immune receptor assembly to signaling. Immunol. Rev. 2019, 291, 26–43. [Google Scholar] [CrossRef]
- McLaughlin, S.; Murray, D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature 2005, 438, 605–611. [Google Scholar] [CrossRef]
- Jaiswal, M.; Dvorsky, R.; Ahmadian, M.R. Deciphering the molecular and functional basis of Dbl family proteins: A novel systematic approach toward classification of selective activation of the Rho family proteins. J. Biol. Chem. 2013, 288, 4486–4500. [Google Scholar] [CrossRef]
- Amin, E.; Jaiswal, M.; Derewenda, U.; Reis, K.; Nouri, K.; Koessmeier, K.T.; Aspenström, P.; Somlyo, A.V.; Dvorsky, R.; Ahmadian, M.R. Deciphering the molecular and functional basis of RHOGAP family proteins: A systematic approach toward selective inactivation of RHO family proteins. J. Biol. Chem. 2016, 291, 20353–20371. [Google Scholar] [CrossRef]
- Wennerberg, K.; Forget, M.A.; Ellerbroek, S.M.; Arthur, W.T.; Burridge, K.; Settleman, J.; Der Channing, C.; Hansen, S.H. Rnd proteins function as RhoA antagonists by activating p190 RhoGAP. Curr. Biol. 2003, 13, 1106–1115. [Google Scholar] [CrossRef]
- Chardin, P. Function and regulation of Rnd proteins. Nat. Rev. Mol. Cell Biol. 2006, 7, 54–62. [Google Scholar] [CrossRef]
- Swart-Mataraza, J.M.; Li, Z.; Sacks, D.B. IQGAP1 is a component of Cdc42 signaling to the cytoskeleton. J. Biol. Chem. 2002, 277, 24753–24763. [Google Scholar] [CrossRef]
- Joshi, S.; Singh, A.R.; Zulcic, M.; Bao, L.; Messer, K.; Ideker, T.; Dutkowski, J.; Durden, D.L. Rac2 controls tumor growth, metastasis and M1-M2 macrophage differentiation in vivo. PLoS ONE 2014, 9, 95893. [Google Scholar] [CrossRef]
- Okuyama, Y.; Nagashima, H.; Ushio-Fukai, M.; Croft, M.; Ishii, N.; So, T. IQGAP1 restrains T-cell cosignaling mediated by OX40. FASEB J. 2020, 34, 540–554. [Google Scholar] [CrossRef]
- Kulkarni, K.; Yang, J.; Zhang, Z.; Barford, D. Multiple factors confer specific Cdc42 and Rac protein activation by dedicator of cytokinesis (DOCK) nucleotide exchange factors. J. Biol. Chem. 2011, 286, 25341–25351. [Google Scholar] [CrossRef]
- Hemsath, L.; Ahmadian, M.R. Fluorescence approaches for monitoring interactions of Rho GTPases with nucleotides, regulators, and effectors. Methods 2005, 37, 173–182. [Google Scholar] [CrossRef]
- Jaiswal, M.; Dubey, B.N.; Koessmeier, K.T.; Gremer, L.; Ahmadian, M.R. Biochemical assays to characterize rho GTPases. Methods Mol. Biol. 2012, 827, 37–58. [Google Scholar] [CrossRef]
- Eberth, A.; Ahmadian, M.R. In vitro GEF and GAP assays. Curr. Protoc. Cell Biol. 2009, 43, 14.9.1–14.9.25. [Google Scholar]
- Fiegen, D.; Haeusler, L.C.; Blumenstein, L.; Herbrand, U.; Dvorsky, R.; Vetter, I.R.; Ahmadian, M.R. Alternative Splicing of Rac1 Generates Rac1b, a Self-activating GTPase. J. Biol. Chem. 2004, 279, 4743–4749. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. Nucl. Acid Symp. Ser. 1999, 41, 95–98. Available online: https://www.scirp.org/(S(lz5mqp453edsnp55rrgjct55))/reference/ReferencesPapers.aspx?ReferenceID=1383440 (accessed on 22 February 2021).
- Webb, B.; Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinforma. 2016, 2016, 5.6.1–5.6.37. [Google Scholar] [CrossRef]
- DeLano, W.L. The PyMOL Molecular Graphics System; Delano Scientific: San Carlos, CA, USA, 2002; Available online: https://www.scirp.org/(S(vtj3fa45qm1ean45vvffcz55))/reference/ReferencesPapers.aspx?ReferenceID=1958992 (accessed on 22 February 2021).
- Holst, M.; Baker, N.; Wang, F. Adaptive multilevel finite element solution of the Poisson-Boltzmann equation I. Algorithms and examples. J. Comput. Chem. 2001, 22, 475. [Google Scholar] [CrossRef]
- Baker, N.A.; Sept, D.; Joseph, S.; Holst, M.J.; McCammon, J.A. Electrostatics of nanosystems: Application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 2001, 98, 10037–10041. [Google Scholar] [CrossRef]
- Lorensen, W.E.; Cline, H.E. Marching Cubes: A High Resolution 3D Surface Construction Algorithm. In Proceedings of the 14th Annual Conference on Computer Graphics and Interactive Techniques, SIGGRAPH 1987, Anaheim, CA, USA, 27–31 July 1987; Association for Computing Machinery, Inc.: New York, NY, USA, 1987; Volume 21, pp. 163–169. [Google Scholar]
- Merritt, E.A.; Bacon, D.J. Raster3D: Photorealistic Molecular Graphics. Methods Enzymol. 1997, 277, 505–524. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mosaddeghzadeh, N.; Nouri, K.; Krumbach, O.H.F.; Amin, E.; Dvorsky, R.; Ahmadian, M.R. Selectivity Determinants of RHO GTPase Binding to IQGAPs. Int. J. Mol. Sci. 2021, 22, 12596. https://doi.org/10.3390/ijms222212596
Mosaddeghzadeh N, Nouri K, Krumbach OHF, Amin E, Dvorsky R, Ahmadian MR. Selectivity Determinants of RHO GTPase Binding to IQGAPs. International Journal of Molecular Sciences. 2021; 22(22):12596. https://doi.org/10.3390/ijms222212596
Chicago/Turabian StyleMosaddeghzadeh, Niloufar, Kazem Nouri, Oliver H. F. Krumbach, Ehsan Amin, Radovan Dvorsky, and Mohammad R. Ahmadian. 2021. "Selectivity Determinants of RHO GTPase Binding to IQGAPs" International Journal of Molecular Sciences 22, no. 22: 12596. https://doi.org/10.3390/ijms222212596
APA StyleMosaddeghzadeh, N., Nouri, K., Krumbach, O. H. F., Amin, E., Dvorsky, R., & Ahmadian, M. R. (2021). Selectivity Determinants of RHO GTPase Binding to IQGAPs. International Journal of Molecular Sciences, 22(22), 12596. https://doi.org/10.3390/ijms222212596







