Overexpression of an Osa-miR162a Derivative in Rice Confers Cross-Kingdom RNA Interference-Mediated Brown Planthopper Resistance without Perturbing Host Development
Abstract
:1. Introduction
2. Results
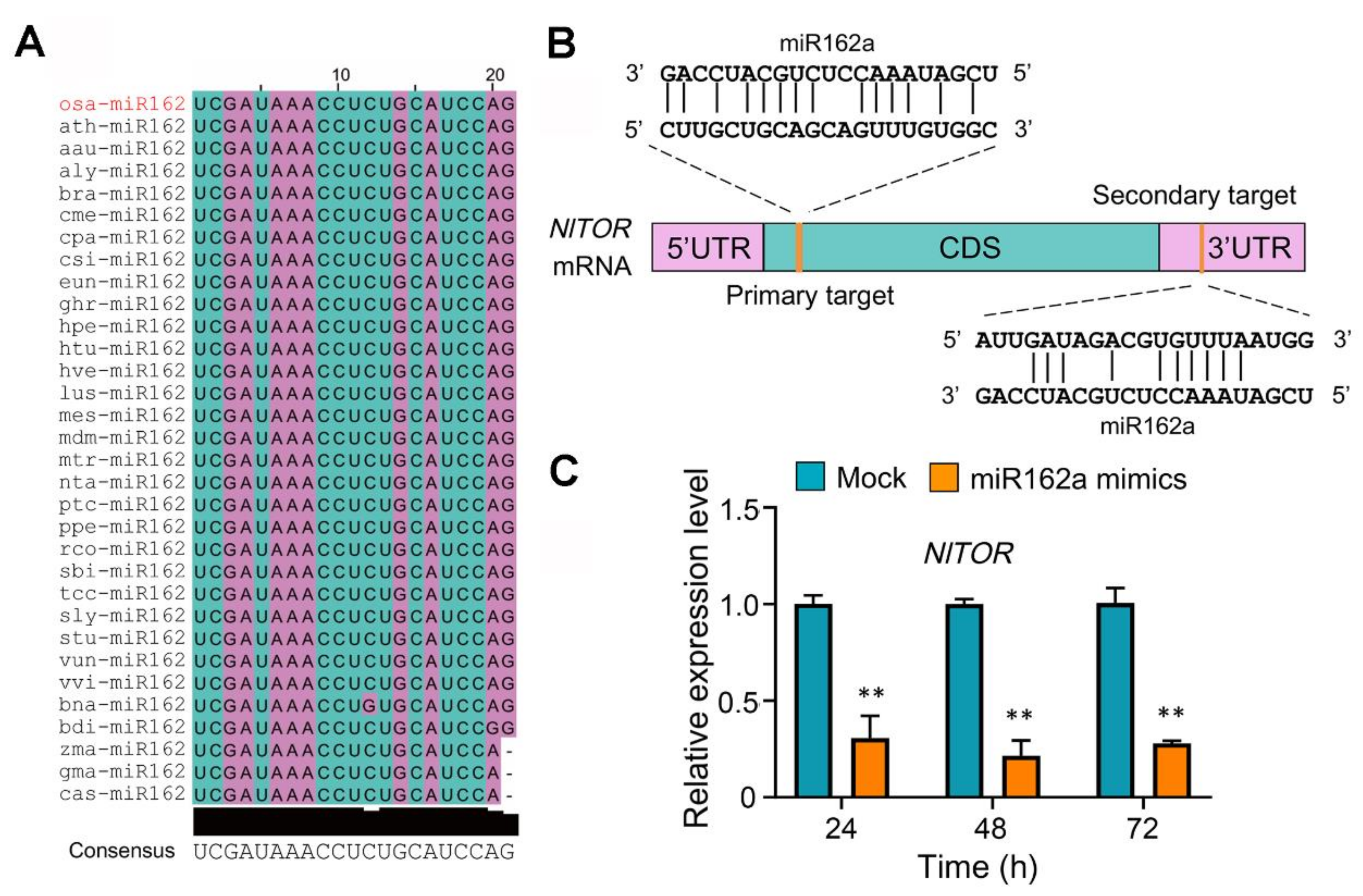
2.1. The NlTOR Gene of BPH Is a Target of Rice Osa-miR162a
2.2. Injected Osa-miR162a Mimics Reduce the Fecundity of BPH
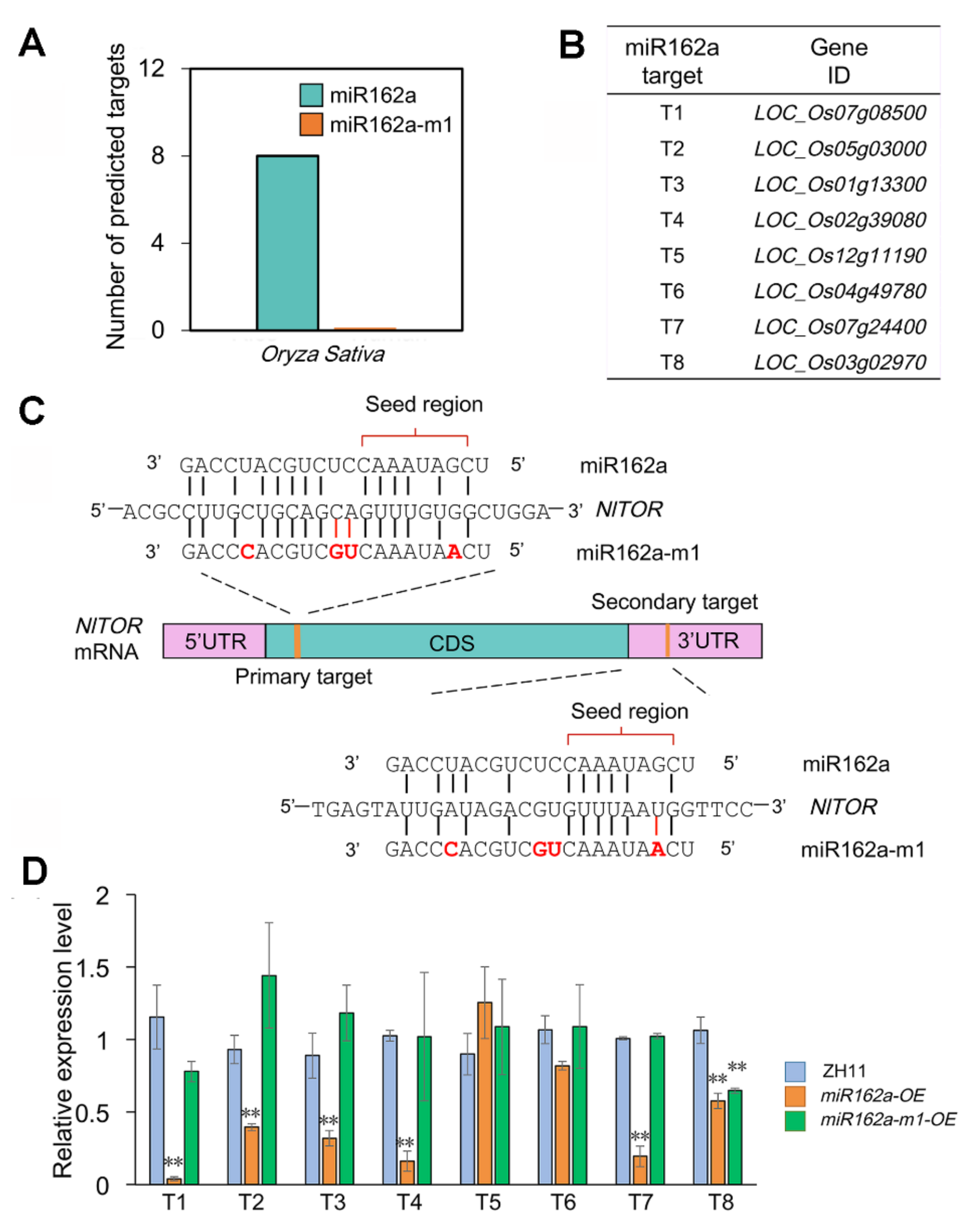
2.3. Osa-miR162a-m1 Exhibits Minimal Perturbation on Rice Endogenous Gene Expression
2.4. Rice Development Is Not Defected upon Osa-miR162a-m1 Overexpression
2.5. BPH Adults Fed with miR162a-m1-OE Plants Display Impaired Fecundity
3. Discussion
4. Materials and Methods
4.1. Plant and Insect Materials
4.2. Molecular Cloning and Transforming Rice Plants
4.3. Feeding BPH with Osa-miR162a Mimics
4.4. Injecting BPH with Osa-miR162a Mimics
4.5. Bioassay of the Fecundity
4.6. RT-qPCR or Stem-Loop RT-qPCR
4.7. Feeding BPH with miR162a-OE or miR162a-m1-OE Plants
4.8. BPH Host Choice Test
4.9. Bioinformatic Prediction of miRNA Target Genes
4.10. Statistical Analysis
5. Patents
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lou, Y.G. Basic research on the outbreak mechanism and sustainable management of rice planthoppers. Chin. J. Appl. Entomol. 2011, 48, 231–238. [Google Scholar]
- Ndiath, M.O. Insecticides and insecticide resistance. Methods Mol. Biol. 2019, 2013, 287–304. [Google Scholar]
- Vaughn, T.; Cavato, T.; Brar, G.; Coombe, T.; DeGooyer, T.; Ford, S.; Groth, M.; Howe, A.; Johnson, S.; Kolacz, K.; et al. A method of controlling corn rootworm feeding using a Bacillus thuringiensis protein expressed in transgenic maize. Crop Sci. 2005, 45, 931–938. [Google Scholar] [CrossRef]
- Tabashnik, B.E.; Gassmann, A.J.; Crowder, D.W.; Yves, C. Insect resistance to Bt crops: Evidence versus theory. Nat. Biotechnol. 2008, 26, 199–202. [Google Scholar] [CrossRef]
- Pallava, B. Hardy cotton-munching pests are latest blow to GM crops. Science 2010, 327, 1439. [Google Scholar]
- Sokol, N.S.; Xu, P.; Jan, Y.N.; Ambros, V. Drosophila let-7 microRNA is required for remodeling of the neuromusculature during metamorphosis. Genes Dev. 2008, 22, 1591–1596. [Google Scholar] [CrossRef] [Green Version]
- Gomez-Orte, E.; Belles, X. MicroRNA-dependent metamorphosis in hemimetabolan insects. Proc. Natl. Acad. Sci. USA 2009, 106, 21678–21682. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ling, L.; Ge, X.; Li, Z.; Zeng, B.; Xun, J.; Aslam, A.F.M.; Song, Q.; Shang, P.; Huang, Y.; Tan, A. MicroRNA Let-7 regulates molting and metamorphosis in the silkworm, Bombyx mori. Insect Biochem. Mol. Biol. 2014, 53, 13–21. [Google Scholar] [CrossRef]
- Belles, X. MicroRNAs and the evolution of insect metamorphosis. Annu. Rev. Entomol. 2017, 62, 111–125. [Google Scholar] [CrossRef] [Green Version]
- Wu, W.; Xiong, W.; Li, C.; Zhai, M.; Li, Y.; Ma, F.; Li, B. MicroRNA-dependent regulation of metamorphosis and identification of microRNAs in the red flour beetle, Tribolium castaneum. Genomics 2017, 109, 362–373. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Lu, K.; Zhou, Q. Dicer1 is crucial for the oocyte maturation of telotrophic ovary in Nilaparvata lugens (Stål) (Hemiptera: Geometroidea). Arch. Insect Biochem. Physiol. 2013, 84, 194–208. [Google Scholar] [CrossRef]
- Fu, X.; Li, T.; Chen, J.; Dong, Y.; Qiu, J.; Kang, K.; Zhang, W. Functional screen for microRNAs of Nilaparvata lugens reveals that targeting of glutamine synthase by miR-4868b regulates fecundity. J. Insect Physiol. 2015, 83, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Liang, Z.; Liang, Y.; Pang, R.; Zhang, W.Q. Conserved microRNAs miR-8-5p and miR-2a-3p modulate chitin biosynthesis in response to 20-hydroxyecdysone signaling in the brown planthopper, Nilaparvata lugens. Insect Biochem. Mol. Biol. 2013, 43, 839–848. [Google Scholar] [CrossRef]
- Li, T.; Chen, J.; Fan, X.; Chen, W.; Zhang, W.Q. MicroRNA and dsRNA targeting chitin synthase A reveal a great potential for pest management of the hemipteran insect Nilaparvata lugens. Pest Manag. Sci. 2017, 73, 1529–1537. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Li, T.; Pang, R.; Yue, X.Z.; Hu, J.; Zhang, W.Q. Genome-wide screening and functional analysis reveal that the specific microRNA nlu-miR-173 regulates molting by targeting Ftz-F1 in Nilaparvata lugens. Front. Physiol. 2018, 9, 1854. [Google Scholar] [CrossRef] [Green Version]
- Agrawal, A.; Rajamani, V.; Reddy, V.S.; Mukherjee, S.K.; Bhatnagar, R.K. Transgenic plants over-expressing insect-specific microRNA acquire insecticidal activity against Helicoverpa armigera: An alternative to Bt-toxin technology. Transgenic Res. 2015, 24, 1–11. [Google Scholar] [CrossRef]
- Jia, L.; Zhang, D.; Xiang, Z.; He, N. Nonfunctional ingestion of plant miRNAs in silkworm revealed by digital droplet PCR and transcriptome analysis. Sci. Rep. 2015, 5, 12290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, K.; Liu, M.; Fu, Z.; Zhou, Z.; Kong, Y.; Liang, H.; Lin, Z.; Luo, J.; Zheng, H.; Wan, P.; et al. Plant microRNAs in larval food regulate honeybee caste development. PLoS Genet. 2017, 13, e1006946. [Google Scholar] [CrossRef]
- Zhang, L.; Hou, D.X.; Chen, X.; Li, D.; Zhu, L.; Zhang, Y.; Li, J.; Bian, Z.; Liang, X.; Cai, X.; et al. Exogenous plant MIR168a specifically targets mammalian LDLRAP1: Evidence of cross-kingdom regulation by microRNA. Cell Res. 2012, 22, 107–126. [Google Scholar] [CrossRef]
- Wang, M.; Weiberg, A.; Lin, F.M.; Thomma, B.; Huang, H.D.; Jin, H.L. Bidirectional cross-kingdom RNAi and fungal uptake of external RNAs confer plant protection. Nat. Plants 2016, 2, 16151. [Google Scholar] [CrossRef]
- Zhang, T.; Zhao, Y.L.; Zhao, J.H.; Wang, S.; Jin, Y.; Chen, Z.Q.; Fang, Y.Y.; Hua, C.L.; Ding, S.W.; Guo, H.S. Cotton plants export microRNAs to inhibit virulence gene expression in a fungal pathogen. Nat. Plants 2016, 2, 16153. [Google Scholar] [CrossRef] [PubMed]
- Ge, Y.F.; Han, J.Y.; Zhou, G.X.; Xu, Y.; Ding, Y.; Shi, M.; Guo, C.; Wu, G. Silencing of miR156 confers enhanced resistance to brown planthopper in rice. Planta 2018, 248, 813–826. [Google Scholar] [CrossRef] [PubMed]
- Dai, Z.; Tan, J.; Zhou, C.; Yang, X.; Yang, F.; Zhang, S.; Sun, S.; Miao, X.; Shi, Z. The OsmiR396-OsGRF8-OsF3H-flavonoid pathway mediates resistance to the brown planthopper in rice (Oryza sativa). Plant Biotechnol. J. 2019, 17, 1657–1669. [Google Scholar] [CrossRef] [Green Version]
- Raikhel, A.S.; Dhadialla, T.S. Accumulation of yolk proteins in insect oocytes. Annu. Rev. Entomol. 1992, 37, 217–251. [Google Scholar] [CrossRef]
- Greenberg, S.M.; Spurgeon, D.W.; Sappington, T.W.; Sétamou, M. Size-dependent feeding and reproduction by Boll weevil (Coleoptera: Curculionidae). J. Econ. Entomol. 2005, 98, 749–756. [Google Scholar] [CrossRef] [Green Version]
- Tufail, M.; Naeemullah, M.; Elmogy, M.; Sharma, P.N.; Takeda, M.; Nakamura, C. Molecular cloning, transcriptional regulation, and differential expression profiling of vitellogenin in two wing-morphs of the brown planthopper, Nilaparvata lugens Stål (Hemiptera: Delphacidae). Insect Mol. Biol. 2010, 19, 787–798. [Google Scholar] [CrossRef]
- Roy, S.G.; Raikhel, A.S. Nutritional and hormonal regulation of the TOR effector 4E-binding protein (4E-BP) in the mosquito Aedes aegypti. FASEB J. 2012, 26, 1334–1342. [Google Scholar] [CrossRef] [Green Version]
- Hansen, I.A.; Attardo, G.M.; Park, J.H.; Peng, Q.; Raikhel, A.S. Target of rapamycin-mediated amino acid signaling in mosquito anautogeny. Proc. Natl. Acad. Sci. USA 2004, 101, 10626–10631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qiu, J.; He, Y.; Zhang, J.; Kang, K.; Li, T.; Zhang, W. Discovery and functional identification of fecundity-related genes in the brown planthopper by large-scale RNA interference. Insect Mol. Biol. 2016, 25, 724–733. [Google Scholar] [CrossRef]
- Gulia-Nuss, M.; Robertson, A.E.; Brown, M.R.; Strand, M.R. Insulin-like peptides and the target of rapamycin pathway coordinately regulate blood digestion and egg maturation in the mosquito Aedes aegypti. PLoS ONE 2011, 6, e20401. [Google Scholar]
- Lu, K.; Chen, X.; Liu, W.T.; Zhou, Q. TOR pathway-mediated juvenile hormone synthesis regulates nutrient-dependent female reproduction in Nilaparvata lugens (Stål). Int. J. Mol. Sci. 2016, 17, 438. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Li, K.; Cai, W.; Zhao, J.; Zou, Y.; Hua, H. Knockdown of TOR causing ovarian diapause in a genetically stable brachypterous strain of Nilaparvata lugens. Arch. Insect Biochem. Physiol. 2017, 95, e21400. [Google Scholar] [CrossRef]
- Zhuo, J.C.; Xue, J.; Lu, J.B.; Huang, H.; Xu, H.; Zhang, C. Effect of RNAi-mediated knockdown of NlTOR gene on fertility of male Nilaparvata lugens. J. Insect Physiol. 2017, 98, 149–159. [Google Scholar] [CrossRef] [PubMed]
- Park, J.H.; Attardo, G.M.; Hansen, I.A.; Raikhel, A.S. GATA factor translation is the final downstream step in the amino acid/target-of-rapamycin-mediated vitellogenin gene expression in the anautogenous mosquito Aedes aegypti. J. Biol. Chem. 2006, 281, 11167–11176. [Google Scholar] [CrossRef] [Green Version]
- Tatebe, H.; Shiozaki, K. Evolutionary conservation of the components in the TOR signaling pathways. Biomolecules 2017, 7, 77. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Varkonyi-Gasic, E.; Wu, R.; Wood, M.; Walton, E.; Hellens, R.P. Protocol: A highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods 2007, 3, 12. [Google Scholar] [CrossRef] [Green Version]
- Li, X.; Ma, X.; Wang, H.; Zhu, Y.; Liu, X.; Li, T.; Zheng, Y.P.; Zhao, J.Q.; Zhang, J.W.; Huang, Y.Y.; et al. Osa-miR162a fine-tunes rice resistance to Magnaporthe oryzae and yield. Rice 2020, 13, 38. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Cao, X.L.; Zhu, Y.; Yang, X.M.; Zhang, K.N.; Xiao, Z.Y.; Wang, H.; Zhao, J.H.; Zhang, L.L.; Li, G.B.; et al. Osa-miR398b boosts H2O2 production and rice blast disease-resistance via multiple superoxide dismutases. New Phytol. 2019, 222, 1507–1522. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Li, Y.; Chern, M.; Zhu, Y.; Zhang, L.; Lu, J.; Li, X.P.; Dang, W.Q.; Ma, X.C.; Yang, Z.R.; et al. Suppression of rice miR168 improves yield, flowering time and immunity. Nat. Plants 2021, 7, 129–136. [Google Scholar] [CrossRef]
- Yu, B.; Yang, Z.; Li, J.; Minakhina, S.; Yang, M.; Padgett, R.W.; Steward, R.; Chen, X. Methylation as a crucial step in plant microRNA biogenesis. Science 2005, 307, 932–935. [Google Scholar] [CrossRef] [Green Version]
- Jung, M.; Schaefer, A.; Steiner, I.; Kempkensteffen, C.; Stephan, C.; Erbersdobler, A.; Jung, K. Robust MicroRNA stability in degraded RNA preparations from human tissue and cell samples. Clin. Chem. 2010, 56, 998–1006. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, Z.; Li, X.; Liu, J.; Dong, L.; Chen, Q.; Liu, J.; Kong, H.; Zhang, Q.; Qi, X.; Hou, D.; et al. Honeysuckle-encoded atypical microRNA2911 directly targets influenza A viruses. Cell Res. 2015, 25, 39–49. [Google Scholar] [CrossRef]
- Chin, A.R.; Fong, M.Y.; Somlo, G.; Wu, J.; Swiderski, P.; Wu, X.; Swiderski, P.; Wu, X.; Wang, S.E. Cross-kingdom inhibition of breast cancer growth by plant miR159. Cell Res. 2016, 26, 217–228. [Google Scholar] [CrossRef] [Green Version]
- Brodersen, P.; Sakvarelidze-Achard, L.; Bruun-Rasmussen, M.; Dunoyer, P.; Yamamoto, Y.Y.; Sieburth, L.; Voinnet, O. Widespread translational inhibition by plant miRNAs and siRNAs. Science 2008, 320, 1185–1190. [Google Scholar] [CrossRef] [PubMed]
- Brousse, C.; Liu, Q.; Beauclair, L.; Deremetz, A.; Axtell, M.J.; Bouche, N. A non-canonical plant microRNA target site. Nucleic Acids Res. 2014, 42, 5270–5279. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seok, H.; Ham, J.; Jang, E.S.; Chi, S.W. MicroRNA target recognition: Insights from transcriptome-wide non-canonical interactions. Mol. Cells 2016, 39, 375–381. [Google Scholar]
- Li, J.-F.; Chung, H.; Niu, Y.; Bush, J.; McCormack, M.; Sheen, J. Comprehensive protein-based artificial microRNA screens for effective gene silencing in plants. Plant Cell 2013, 25, 1507–1522. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Reichel, M.; Millar, A.A. Determinants beyond both complementarity and cleavage govern MicroR159 efficacy in Arabidopsis. PLoS Genet. 2014, 10, e1004232. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Li, P.; Li, X.; Liu, C.; Cao, S.; Chu, C.; Cao, X. Loss of function of OsDLC1 affects microRNA accumulation and causes developmental defects in rice. Plant Physiol. 2005, 139, 296–305. [Google Scholar] [CrossRef] [Green Version]
- Hiei, Y.; Ohta, S.; Komari, T.; Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 1994, 6, 271–282. [Google Scholar] [CrossRef] [Green Version]
- Zhang, N.; Zhang, D.; Shen, W.; Li, J.-F. Engineered artificial microRNA precursors facilitate cloning and gene silencing in Arabidopsis and Rice. Int. J. Mol. Sci. 2019, 20, 5620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krüger, J.; Rehmsmeier, M. RNAhybrid: MicroRNA target prediction easy, fast and flexible. Nucleic Acids Res. 2006, 34, W451–W454. [Google Scholar] [CrossRef]
- Enright, A.J.; John, B.; Gaul, U.; Tuschl, T.; Sander, C.; Marks, D.S. MicroRNA targets in Drosophila. Genome Biol. 2003, 5, R1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schwab, R.; Ossowski, S.; Riester, M.; Warthmann, N.; Weigel, D. Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 2006, 18, 1121–1133. [Google Scholar] [CrossRef] [PubMed] [Green Version]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shen, W.; Cao, S.; Liu, J.; Zhang, W.; Chen, J.; Li, J.-F. Overexpression of an Osa-miR162a Derivative in Rice Confers Cross-Kingdom RNA Interference-Mediated Brown Planthopper Resistance without Perturbing Host Development. Int. J. Mol. Sci. 2021, 22, 12652. https://doi.org/10.3390/ijms222312652
Shen W, Cao S, Liu J, Zhang W, Chen J, Li J-F. Overexpression of an Osa-miR162a Derivative in Rice Confers Cross-Kingdom RNA Interference-Mediated Brown Planthopper Resistance without Perturbing Host Development. International Journal of Molecular Sciences. 2021; 22(23):12652. https://doi.org/10.3390/ijms222312652
Chicago/Turabian StyleShen, Wenzhong, Shanni Cao, Jinhui Liu, Wenqing Zhang, Jie Chen, and Jian-Feng Li. 2021. "Overexpression of an Osa-miR162a Derivative in Rice Confers Cross-Kingdom RNA Interference-Mediated Brown Planthopper Resistance without Perturbing Host Development" International Journal of Molecular Sciences 22, no. 23: 12652. https://doi.org/10.3390/ijms222312652





