Combined Pharmacophore and Grid-Independent Molecular Descriptors (GRIND) Analysis to Probe 3D Features of Inositol 1,4,5-Trisphosphate Receptor (IP3R) Inhibitors in Cancer
Abstract
:1. Introduction
2. Results
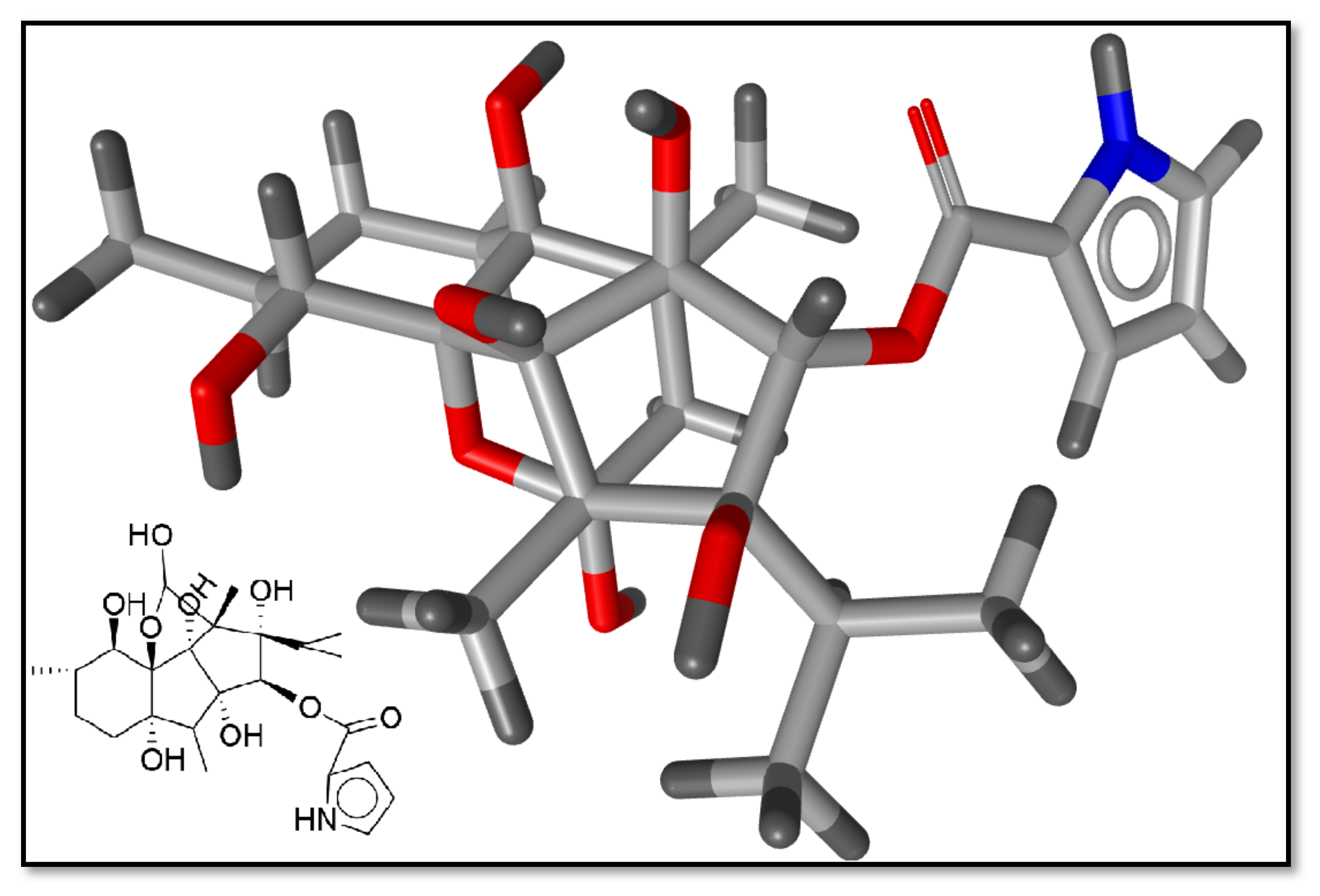
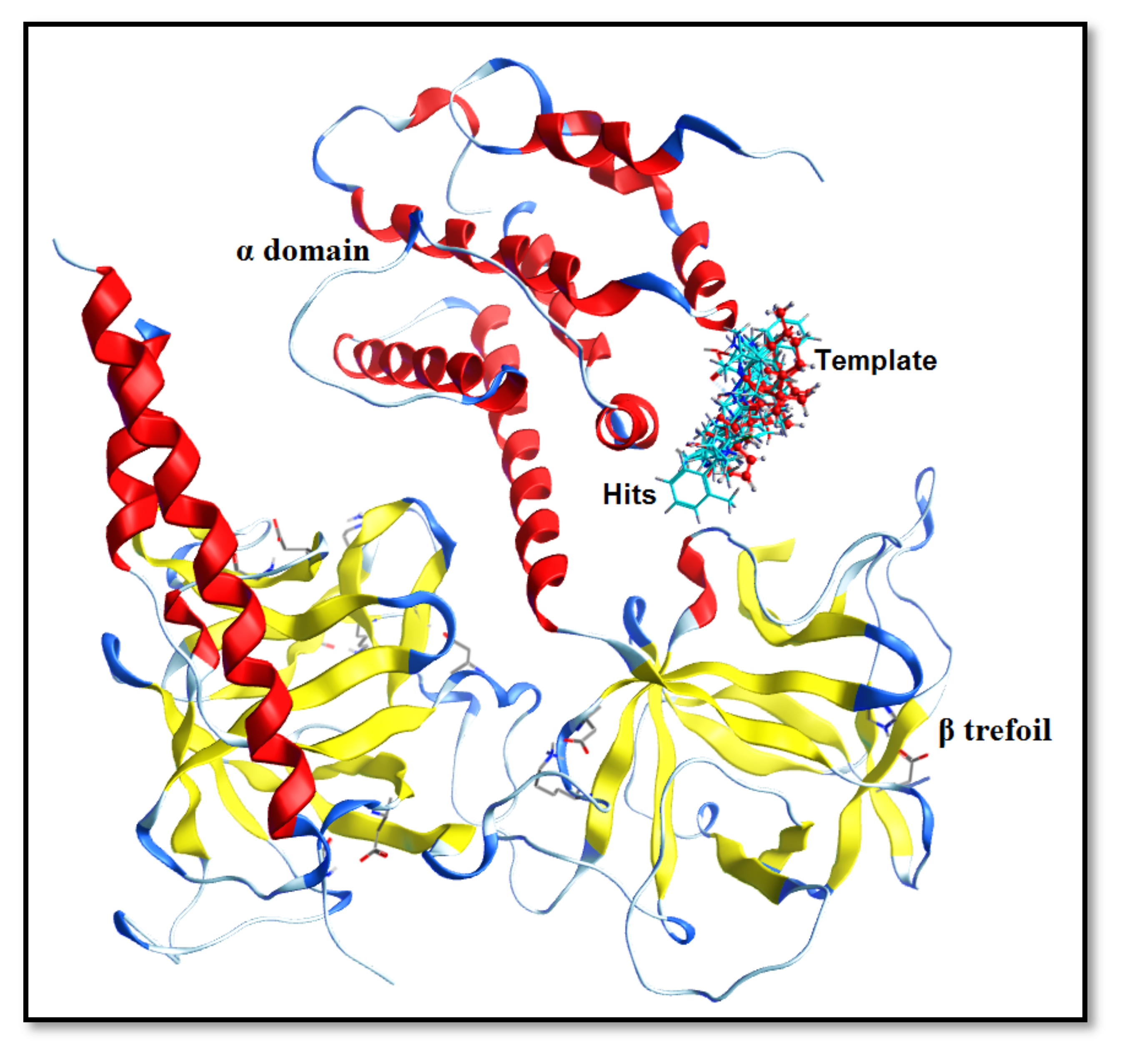
2.1. Preliminary Data Analysis and Template Selection
2.2. Pharmacophore Model Generation and Validation
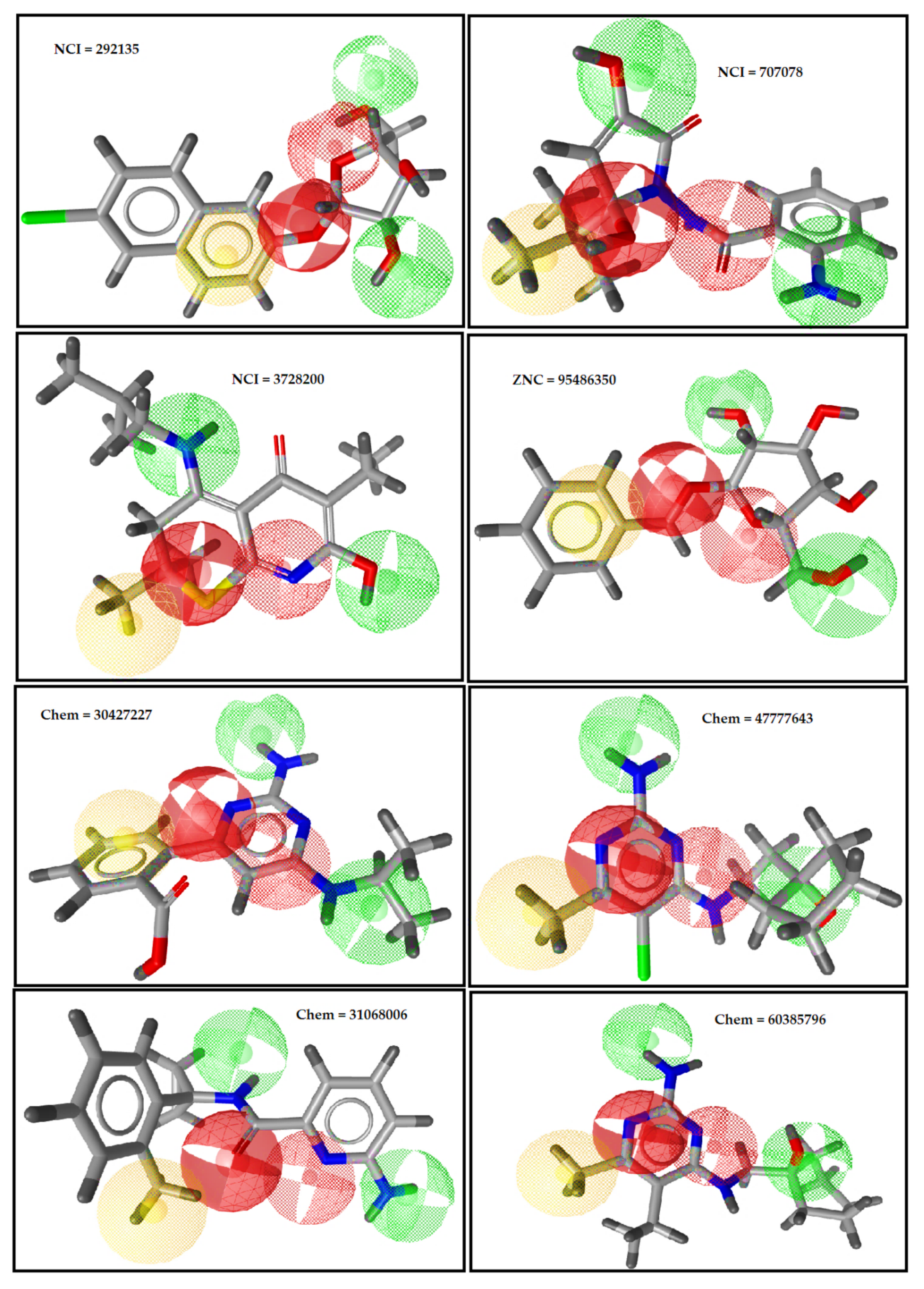
2.3. Pharmacophore-Based Virtual Screening
2.4. Molecular-Docking Simulation and PLIF Analysis
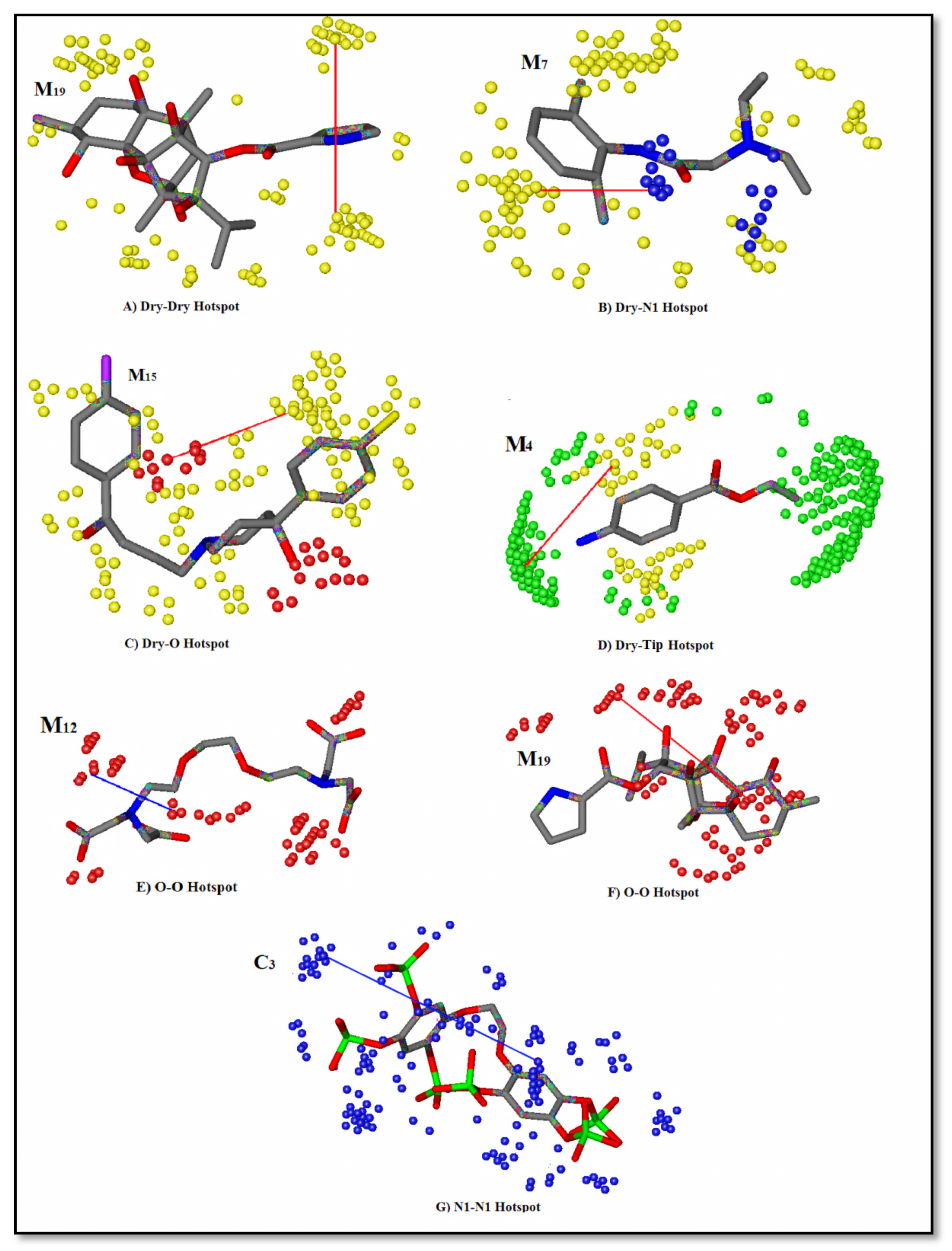
2.5. Grid-Independent Molecular Descriptor (GRIND) Analysis
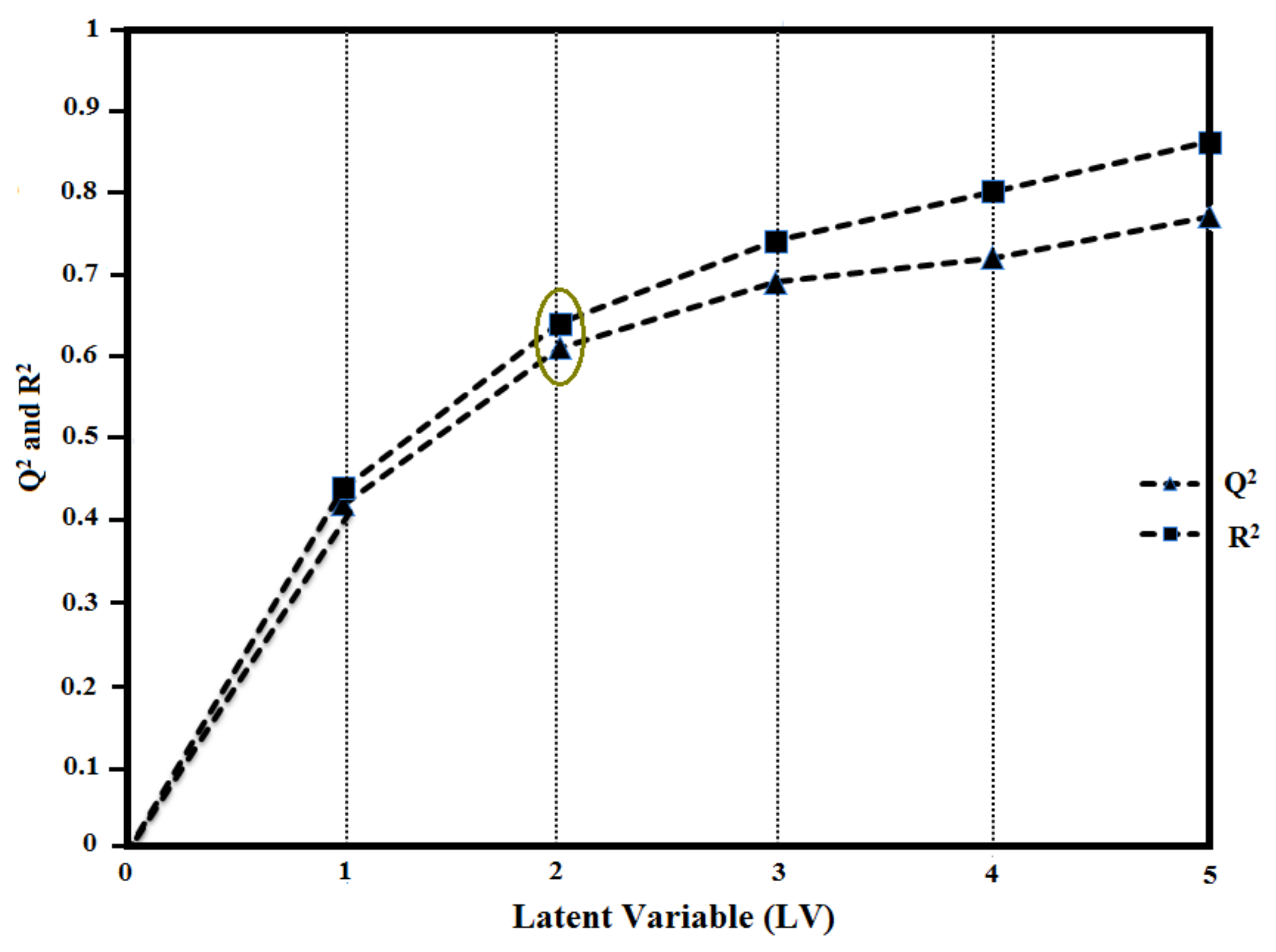
2.6. Validation of GRIND Model
3. Discussion
4. Materials and Methods
4.1. Ligand Dataset (Collection and Refinement)
4.2. Molecular-Docking Simulations
4.3. Template Selection Criteria for Pharmacophore Modeling
4.4. Pharmacophore Model Generation and Validation
4.5. Pharmacophore-Based Virtual Screening
4.6. Grid-Independent Molecular Descriptor (GRIND) Calculation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Shapovalov, G.; Ritaine, A.; Skryma, R.; Prevarskaya, N. Role of TRP ion channels in cancer and tumorigenesis. In Seminars in Immunopathology; Springer: Berlin/Heidelberg, Germany, 2016. [Google Scholar]
- Roderick, L.H.; Cook, S.J. Ca2+ signalling checkpoints in cancer: Remodelling Ca2+ for cancer cell proliferation and survival. Nat. Rev. Cancer 2008, 8, 361–375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bong, H.A.; Monteith, G.R. Calcium signaling and the therapeutic targeting of cancer cells. Biochim. Et Biophys. Acta Mol. Cell Res. 2018, 1865, 1786–1794. [Google Scholar] [CrossRef]
- Bittremieux, M.; Parys, J.B.; Pinton, P.; Bultynck, G. ER functions of oncogenes and tumor suppressors: Modulators of intracellular Ca2+ signaling. Biochim. Biophys. Acta Mol. Cell Res. 2016, 1863, 1364–1378. [Google Scholar] [CrossRef]
- Chen, Y.F.; Chen, Y.T.; Chiu, W.T.; Shen, M.R. Remodeling of calcium signaling in tumor progression. J. Biomed. Sci. 2013, 20, 23. [Google Scholar] [CrossRef] [Green Version]
- Xu, X.; Balk, S.P.; Isaacs, W.B.; Ma, J. Calcium signaling: An underlying link between cardiac disease and carcinogenesis. Cell Biosci. 2018, 8, 39. [Google Scholar] [CrossRef] [Green Version]
- Xu, M.; Seas, A.; Kiyani, M.; Ji, K.; Bell, S.Y.; Hannah, N. A temporal examination of calcium signaling in cancer-from tumorigenesis, to immune evasion, and metastasis. Cell Biosci. 2018, 8, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tarn, D.; Yu, C.J.; Lu, J.; Hartz, A.; Tamanoi, F.; Zink, J.I. In vitro delivery of calcium ions by nanogated mesoporous silica nanoparticles to induce cancer cellular apoptosis. Mol. Syst. Des. Eng. 2017, 2, 384–392. [Google Scholar] [CrossRef]
- Marchi, S.; Pinton, P. Alterations of calcium homeostasis in cancer cells. Curr. Opin. Pharmacol. 2016, 29, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Monteith, G.R.; Davis, F.M.; Roberts-Thomson, S.J. Calcium channels and pumps in cancer: Changes and consequences. J. Biol. Chem. 2012, 287, 31666–31673. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prevarskaya, N.; Ouadid-Ahidouch, H.; Skryma, R.; Shuba, Y. Remodelling of Ca2+ transport in cancer: How it contributes to cancer hallmarks? Philos. Trans. R. Soc. B Biol. Sci. 2014, 369, 20130097. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roberts-Thomson, S.J.; Chalmers, S.B.; Monteith, G.R. The calcium-signaling toolkit in cancer: Remodeling and targeting. Cold Spring Harb. Perspect. Biol. 2019, 11, a035204. [Google Scholar] [CrossRef] [Green Version]
- Boroughs, K.L.; DeBerardinis, R.J. Metabolic pathways promoting cancer cell survival and growth. Nat. Cell Biol. 2015, 17, 351–359. [Google Scholar] [CrossRef] [Green Version]
- Bultynck, G. Onco-IP3Rs feed cancerous cravings for mitochondrial Ca2+. Trends Biochem. Sci. 2016, 41, 390–393. [Google Scholar] [CrossRef] [PubMed]
- Mound, A.; Vautrin-Glabik, A.; Foulon, A.; Botia, B.; Hague, F.; Parys, J.B.; Ouadid-Ahidouch, H.; Rodat-Despoix, L. Downregulation of type 3 inositol (1, 4, 5)-trisphosphate receptor decreases breast cancer cell migration through an oscillatory Ca2+ signal. Oncotarget 2017, 8, 72324. [Google Scholar] [CrossRef] [PubMed]
- Shibao, K.; Fiedler, M.J.; Nagata, J.; Minagawa, N.; Hirata, K.; Nakayama, Y.; Iwakiri, Y.; Nathanson, M.H.; Yamaguchi, K. The type III inositol 1, 4, 5-trisphosphate receptor is associated with aggressiveness of colorectal carcinoma. Cell Calcium 2010, 48, 315–323. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singh, A.; Chagtoo, M.; Tiwari, S.; George, N.; Chakravarti, B.; Khan, S.; Lakshmi, S.; Godbole, M. Inhibition of Inositol 1, 4, 5-Trisphosphate Receptor Induce Breast Cancer Cell Death Through Deregulated Autophagy and Cellular Bioenergetics. J. Cell. Biochem. 2017, 118, 2333–2346. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.S.; Han, K.S.; Ku, B.M.; Lee, Y.K.; Hong, J.; Shin, H.Y.; Almonte, A.G.; Woo, D.H.; Brat, D.J.; Hwang, E.M. Caffeine-mediated inhibition of calcium release channel inositol 1, 4, 5-trisphosphate receptor subtype 3 blocks glioblastoma invasion and extends survival. Cancer Res. 2010, 70, 1173–1183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prole, L.D.; Taylor, C.W. Structure and function of IP3 receptors. Cold Spring Harb. Perspect. Biol. 2019, 11, a035063. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loncke, J.; Kerkhofs, M.; Kaasik, A.; Bezprozvanny, I.; Bultynck, G. Recent advances in understanding IP3R function with focus on ER-mitochondrial Ca2+ transfers. Curr. Opin. Physiol. 2020, 17, 80–88. [Google Scholar] [CrossRef]
- Foskett, J.K. Inositol trisphosphate receptor Ca2+ release channels in neurological diseases. Pflügers Arch. -Eur. J. Physiol. 2010, 460, 481–494. [Google Scholar] [CrossRef] [Green Version]
- Furuichi, T.; Yoshikawa, S.; Miyawaki, A.; Wada, K.; Maeda, N.; Mikoshiba, K. Primary structure and functional expression of the inositol 1, 4, 5-trisphosphate-binding protein P 400. Nature 1989, 342, 32–38. [Google Scholar] [CrossRef]
- Mignery, G.A.; Newton, C.L.; Archer, B.T.; Südhof, T.C. Structure and expression of the rat inositol 1, 4, 5-trisphosphate receptor. J. Biol. Chem. 1990, 265, 12679–12685. [Google Scholar] [CrossRef]
- Serysheva, I.I. Toward a high-resolution structure of IP3R channel. Cell Calcium 2014, 56, 125–132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mikoshiba, K. IP3 receptor/Ca2+ channel: From discovery to new signaling concepts. J. Neurochem. 2007, 102, 1426–1446. [Google Scholar] [CrossRef] [PubMed]
- Hamada, K.; Miyatake, H.; Terauchi, A.; Mikoshiba, K. IP3-mediated gating mechanism of the IP3 receptor revealed by mutagenesis and X-ray crystallography. Proc. Natl. Acad. Sci. USA 2017, 114, 4661–4666. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Serysheva, I.I.; Bare, D.J.; Ludtke, S.J.; Kettlun, C.S.; Chiu, W.; Mignery, G.A. Structure of the type 1 inositol 1, 4, 5-trisphosphate receptor revealed by electron cryomicroscopy. J. Biol. Chem. 2003, 278, 21319–21322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoshikawa, S.; Tanimura, T.; Miyawaki, A.; Nakamura, M.; Yuzaki, M.; Furuichi, T.; Mikoshiba, K. Molecular cloning and characterization of the inositol 1, 4, 5-trisphosphate receptor in Drosophila melanogaster. J. Biol. Chem. 1992, 267, 16613–16619. [Google Scholar] [CrossRef]
- Uchida, K.; Miyauchi, H.; Furuichi, T.; Michikawa, T.; Mikoshiba, K. Critical regions for activation gating of the inositol 1, 4, 5-trisphosphate receptor. J. Biol. Chem. 2003, 278, 16551–16560. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Paknejad, N.; Hite, R.K. Structural basis for the regulation of inositol trisphosphate receptors by Ca2+ and IP3. Nat. Struct. Mol. Biol. 2018, 25, 660–668. [Google Scholar] [CrossRef]
- Azumaya, C.M.; Linton, E.A.; Risener, C.J.; Nakagawa, T.; Karakas, E. Cryo-EM structure of human type-3 inositol triphosphate receptor reveals the presence of a self-binding peptide that acts as an antagonist. J. Biol. Chem. 2020, 295, 1743–1753. [Google Scholar] [CrossRef] [PubMed]
- Cárdenas, C.; Miller, R.A.; Smith, I.; Bui, T.; Molgó, J.; Müller, M.; Vais, H.; Cheung, K.H.; Yang, J.; Parker, I. Essential regulation of cell bioenergetics by constitutive InsP3 receptor Ca2+ transfer to mitochondria. Cell 2010, 142, 270–283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cárdenas, C.; Müller, M.; McNeal, A.; Lovy, A.; Jaňa, F.; Bustos, G.; Urra, F.; Smith, N.; Molgó, J.; Diehl, J.A. Selective vulnerability of cancer cells by inhibition of Ca2+ transfer from endoplasmic reticulum to mitochondria. Cell Rep. 2016, 14, 2313–2324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tedeschi, G.; Nonnis, S.; Strumbo, B.; Cruciani, G.; Carosati, E.; Negri, A. On the catalytic role of the active site residue E121 of E. coli L-aspartate oxidase. Biochimie 2010, 92, 1335–1342. [Google Scholar] [CrossRef]
- Fontaine, F.; Pastor, M.; Zamora, I.; Sanz, F. Anchor−grind: Filling the gap between standard 3d qsar and the grid-independent descriptors. J. Med. Chem. 2005, 48, 2687–2694. [Google Scholar] [CrossRef] [PubMed]
- Cruciani, G.; Carosati, E.; De Boeck, B.; Ethirajulu, K.; Mackie, C.; Howe, T.; Vianello, R. MetaSite: Understanding metabolism in human cytochromes from the perspective of the chemist. J. Med. Chem. 2005, 48, 6970–6979. [Google Scholar] [CrossRef]
- Cruciani, G.; Pastor, M.; Guba, W. VolSurf: A new tool for the pharmacokinetic optimization of lead compounds. Eur. J. Pharm. Sci. 2000, 11, S29–S39. [Google Scholar] [CrossRef]
- Cases, M.; Briggs, K.; Steger, H.T.; Pognan, F.; Marc, P.; Kleinöder, T.; Schwab, C.H.; Pastor, M.; Wichard, J.; Sanz, F. The eTOX data-sharing project to advance in silico drug-induced toxicity prediction. Int. J. Mol. Sci. 2014, 15, 21136–21154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sciabola, S.; Stanton, R.V.; Mills, J.E.; Flocco, M.M.; Baroni, M.; Cruciani, G.; Perruccio, F.; Mason, J.S. High-throughput virtual screening of proteins using GRID molecular interaction fields. J. Chem. Inf. Model. 2010, 50, 155–169. [Google Scholar] [CrossRef] [PubMed]
- Gaulton, A.; Bellis, L.; Bento, A.P.S.F.F.; Chambers, J.; Davies, M.; Hersey, A.; Light, Y.; McGlinchey, S.; Michalovich, D.; Al-Lazikani, B.; et al. ChEMBL: A large-scale bioactivity database for drug discovery. Nucl. Acids Res. 2012, 40, D1100–D1107. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.; Davis, R.J.; Nahorski, S.R.; Ballereau, S.; Spiess, B.; Potter, B.V. Synthesis, Calcium Mobilizing, and Physicochemical Properties of d-c hiro-Inositol 1, 3, 4, 6-Tetrakisphosphate, a Novel and Potent Ligand at the d-m yo-Inositol 1, 4, 5-Trisphosphate Receptor. J. Med. Chem. 1999, 42, 1991–1998. [Google Scholar] [CrossRef] [PubMed]
- Adelt, S.; Plettenburg, O.; Stricker, R.; Reiser, G.; Altenbach, H.J.; Vogel, G. Enzyme-assisted total synthesis of the optical antipodes d-myo-inositol 3, 4, 5-trisphosphate and d-myo-inositol 1, 5, 6-trisphosphate: Aspects of their structure−activity relationship to biologically active inositol phosphates. J. Med. Chem. 1999, 42, 1262–1273. [Google Scholar] [CrossRef] [PubMed]
- Podeschwa, M.; Plettenburg, O.; Brocke, J.; Block, O.; Adelt, S.; Altenbach, H.J. Stereoselective Synthesis of myo-, neo-, L-chiro, D-chiro, allo-, scyllo-, and epi-Inositol Systems via Conduritols Prepared from p-Benzoquinone. Eur. J. Org. Chem. 2003, 2003, 1958–1972. [Google Scholar] [CrossRef]
- Ta, T.A.; Feng, W.; Molinski, T.F.; Pessah, I.N. Hydroxylated xestospongins block IP3-induced Ca2+ release and sensitize Ca2+-induced Ca2+ release mediated by ryanodine receptors. Mol. Pharmacol. 2005. [Google Scholar] [CrossRef] [Green Version]
- Jaimovich Pérez, E.; Mattei, C.; Liberona, L.; José, C.; César, E.H.; Manuel, B.; Julien, D.; Cécile, L.; Dominique, M.J. Xestospongin B, a competitive inhibitor of IP3-mediated Ca2+ signalling in cultured rat myotubes, isolated myonuclei, and neuroblastoma (NG108-15) cells. FEBS Lett. 2005, 579, 2051–2057. [Google Scholar] [CrossRef]
- Gafni, J.; Munsch, J.A.; Lam, T.H.; Catlin, M.C.; Costa, L.G.; Molinski, T.F.; Pessah, I.N. Xestospongins: Potent membrane permeable blockers of the inositol 1, 4, 5-trisphosphate receptor. Neuron 1997, 19, 723–733. [Google Scholar] [CrossRef] [Green Version]
- Mills, S.J.; Luyten, T.; Erneux, C.; Parys, J.B.; Potter, B.V.L. Multivalent benzene polyphosphate derivatives are non-Ca2+-mobilizing Ins (1, 4, 5) P3 receptor antagonists. Messenger 2012, 1, 167–181. [Google Scholar] [CrossRef]
- Leeson, P.D.; Springthorpe, B. The influence of drug-like concepts on decision-making in medicinal chemistry. Nat. Rev. Drug Discov. 2007, 6, 881. [Google Scholar] [CrossRef] [PubMed]
- Ryckmans, T.; Edwards, M.P.; Horne, V.A.; Correia, A.M.; Owen, D.R.; Thompson, L.R.; Tran, I.; Tutt, M.F.; Young, T. Rapid assessment of a novel series of selective CB 2 agonists using parallel synthesis protocols: A lipophilic efficiency (LipE) analysis. Bioorganic Med. Chem. Lett. 2009, 19, 4406–4409. [Google Scholar] [CrossRef]
- Jabeen, I.; Pleban, K.; Rinner, U.; Chiba, P.; Ecker, G. Structure–activity relationships, ligand efficiency, and lipophilic efficiency profiles of benzophenone-type inhibitors of the multidrug transporter P-glycoprotein. J. Med. Chem. 2012, 55, 3261–3273. [Google Scholar] [CrossRef] [PubMed]
- Freeman-Cook, K.D.; Hoffman, R.L.; Johnson, T.W. Lipophilic efficiency: The most important efficiency metric in medicinal chemistry. Future Med. Chem. 2013, 5, 113–115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Podlipnik, C.; Solmajer, T.; Koller, J. Similarity of radial distribution function’s intervals. Match-Commun. Math. Comput. Chem. 2006, 56, 261–270. [Google Scholar]
- Lemmen, C.; Lengauer, T. Computational methods for the structural alignment of molecules. J. Comput. Aided Mol. Des. 2000, 14, 215–232. [Google Scholar] [CrossRef] [PubMed]
- Seidel, T.; Bryant, S.D.; Ibis, G.P.G.; Langer, T. 3D pharmacophore modeling techniques in computer-aided molecular design using LigandScout. Tutor. Chemoinform 2017, 281, 279–309. [Google Scholar]
- Güner, O.F. Pharmacophore Perception, Development, and Use in Drug Design; Internat University Line: La Jolla, CA, USA, 2000; Volume 2. [Google Scholar]
- Mannhold, R.; Kubinyi, H.; Folkers, G. Pharmacophores and Pharmacophore Searches; John Wiley & Sons: Hoboken, NJ, USA, 2006; Volume 32. [Google Scholar]
- Gurney, A.M.; Allam, M. Inhibition of calcium release from the sarcoplasmic reticulum of rabbit aorta by hydralazine. Br. J. Pharmacol. 1995, 114, 238–244. [Google Scholar] [CrossRef] [Green Version]
- Nagarkatti, N.; Deshpande, L.S.; DeLorenzo, R.J. Levetiracetam inhibits both ryanodine and IP3 receptor activated calcium induced calcium release in hippocampal neurons in culture. Neurosci. Lett. 2008, 436, 289–293. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seiler, S.M.; Arnold, A.J.; Stanton, H.C. Inhibitors of inositol trisphosphate-induced Ca2+ release from isolated platelet membrane vesicles. Biochem. Pharmacol. 1987, 36, 3331–3337. [Google Scholar] [CrossRef]
- Allen, F.H. The Cambridge Structural Database: A quarter of a million crystal structures and rising. Acta Crystallogr. Sect. B Struct. Sci. 2002, 58, 380–388. [Google Scholar] [CrossRef]
- Milne, G.W.; George, W.A.; Nicklaus, J.S.; Marc, C.; Driscoll, J.S.; Wang, S.; Zaharevitz, D. National Cancer Institute drug information system 3D database. J. Chem. Inf. Comput. Sci. 1994, 34, 1219–1224. [Google Scholar] [CrossRef] [PubMed]
- Ihlenfeldt, W.-D.; Voigt, J.H.; Bienfait, B.; Oellien, F.; Nicklaus, M. Enhanced CACTVS browser of the Open NCI Database. J. Chem. Inf. Comput. Sci. 2002, 42, 46–57. [Google Scholar] [CrossRef]
- Irwin, J.J.; Sterling, T.; Mysinger, M.M.; Bolstad, E.S.; Coleman, R.G. ZINC: A free tool to discover chemistry for biology. J. Chem. Inf. Modeling 2012, 52, 1757–1768. [Google Scholar] [CrossRef] [PubMed]
- Guengerich, F.P. Cytochrome P450s and other enzymes in drug metabolism and toxicity. AAPS J. 2006, 8, E101–E111. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sushko, I.; Novotarskyi, S.; Körner, R.; Pandey, A.K.; Rupp, M.; Teetz, W.; Brandmaier, S.; Abdelaziz, A.; Prokopenko, V.V.; Tanchuk, V.Y.; et al. Online chemical modeling environment (OCHEM): Web platform for data storage, model development and publishing of chemical information. J. Comput. Aided Mol. Des. 2011, 25, 533–554. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Molecular Operating Environment (MOE). Chemical Computing Group Inc., 2019.01., 1010 Sherbooke St. West, Suite #910, Montreal, QC, Canada, 2015, Volume 49. Available online: www.chemcomp.com/index.htm (accessed on 17 September 2021).
- Ferguson, D.M.; Raber, D.J. A new approach to probing conformational space with molecular mechanics: Random incremental pulse search. J. Am. Chem. Soc. 1989, 111, 4371–4378. [Google Scholar] [CrossRef]
- Guth, B.D. Preclinical cardiovascular risk assessment in modern drug development. Toxicol. Sci. 2007, 97, 4–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Redfern, W.S.; Carlsson, L.; Davis, A.S.; Lynch, W.G.; MacKenzie, I.L.; Palethorpe, S.; Siegl, P.K.S.; Strang, I.; Sullivan, A.T.; Wallis, R. Relationships between preclinical cardiac electrophysiology, clinical QT interval prolongation and torsade de pointes for a broad range of drugs: Evidence for a provisional safety margin in drug development. Cardiovasc. Res. 2003, 58, 32–45. [Google Scholar] [CrossRef]
- Munawar, S.; Windley, M.J.; Tse, E.G.; Todd, M.H.; Hill, A.P.; Vandenberg, J.I.; Jabeen, I. Experimentally validated pharmacoinformatics approach to predict hERG inhibition potential of new chemical entities. Front. Pharmacol. 2018, 9, 1035. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W.L. Pymol: An open-source molecular graphics tool. CCP4 Newsl. Protein Crystallogr. 2002, 40, 82–92. [Google Scholar]
- Yoshikawa, F.; Morita, M.; Monkawa, T.; Michikawa, T.; Furuichi, T.; Mikoshiba, K. Mutational analysis of the ligand binding site of the inositol 1, 4, 5-trisphosphate receptor. J. Biol. Chem. 1996, 271, 18277–18284. [Google Scholar] [CrossRef] [Green Version]
- Bosanac, I.; Michikawa, T.; Mikoshiba, K.; Ikura, M. Structural insights into the regulatory mechanism of IP3 receptor. Biochim. Biophys. Acta Mol. Cell Res. 2004, 174, 89–102. [Google Scholar] [CrossRef] [Green Version]
- Ismatullah, H.; Jabeen, I.; Saeed, M.T. Biological Regulatory Network (BRN) Analysis and Molecular Docking Simulations to Probe the Modulation of IP3R Mediated Ca2+ Signaling in Cancer. Genes 2021, 12, 34. [Google Scholar] [CrossRef] [PubMed]
- Pentacle Version 1.0.7; Molecular Discovery Ltd.: Perugia, Italy, 2009; Available online: https://www.moldiscovery.com (accessed on 17 September 2021).
- Baroni, M.; Costantino, G.; Cruciani, G.; Riganelli, D.; Valigi, R.; Clementi, S. Generating optimal linear PLS estimations (GOLPE): An advanced chemometric tool for handling 3D-QSAR problems. Quant. Struct. Act. Relatsh. 1993, 12, 9–20. [Google Scholar] [CrossRef]
- Moro, S.; Bacilieri, M.; Ferrari, C.; Spalluto, G. Autocorrelation of molecular electrostatic potential surface properties combined with partial least squares analysis as alternative attractive tool to generate ligand-based 3D-QSARs. Curr. Drug Discov. Technol. 2005, 2, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Elisseeff, A.; Pontil, M. Leave-one-out error and stability of learning algorithms with applications. NATO Sci. Ser. Sub Ser. Iii Comput. Syst. Sci. 2003, 190, 111–130. [Google Scholar]
- Hawkins, D.M.; Basak, S.C.; Mills, D. Assessing model fit by cross-validation. J. Chem. Inf. Comput. Sci. 2003, 43, 579–586. [Google Scholar] [CrossRef]
- Veerasamy, R.; Rajak, H.; Jain, A.; Sivadasan, S.; Varghese, C.P.; Agrawal, R.K. Validation of QSAR models-strategies and importance. Int. J. Drug Des. Discov. 2011, 3, 511–519. [Google Scholar]
- Roy, K.; Mitra, I.; Kar, S.; Ojha, P.K.; Das, R.N.; Kabir, H. Comparative studies on some metrics for external validation of QSPR models. J. Chem. Inf. Modeling 2012, 52, 396–408. [Google Scholar] [CrossRef] [PubMed]
- Roy, P.P.; Roy, K. On some aspects of variable selection for partial least squares regression models. QSAR Comb. Sci. 2008, 27, 302–313. [Google Scholar] [CrossRef]
- Pratim, R.P.; Paul, S.; Mitra, I.; Roy, K. On two novel parameters for validation of predictive QSAR models. Molecules 2009, 14, 1660–1701. [Google Scholar] [CrossRef] [PubMed]
- Roy, K.; Kar, S.; Ambure, P. On a simple approach for determining applicability domain of QSAR models. Chemom. Intell. Lab. Syst. 2015, 145, 22–29. [Google Scholar] [CrossRef]
- Snedecor, G.; Cochran, W. Statistical Methods, 6th ed.; Oxford and IBH Publishing Co: New Delhi, India, 1967. [Google Scholar]
- Monteith, G.R.; Prevarskaya, N.; Roberts-Thomson, S.J. The calcium–cancer signalling nexus. Nat. Rev. Cancer 2017, 17, 367. [Google Scholar] [CrossRef] [Green Version]
- Cardenas, C.; Lovy, A.; Silva, P.E.; Urra, F.; Mizzoni, C.; Ahumada, C.U.; Bustos, G.; Jaňa, F.; Cruz, P.; Farias, P. Cancer cells with defective oxidative phosphorylation require endoplasmic reticulum–to–mitochondria Ca2+ transfer for survival. Sci. Signal. 2020, 13, 6. [Google Scholar] [CrossRef] [PubMed]
- Fan, G.; Baker, M.R.; Wang, Z.; Seryshev, A.B.; Ludtke, S.J.; Baker, M.L.; Serysheva, I.I. Cryo-EM reveals ligand induced allostery underlying InsP 3 R channel gating. Cell Res. 2018, 28, 1158–1170. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hotoda, H.; Murayama, K.; Miyamoto, S.; Iwata, Y.; Takahashi, M.; Kawase, Y.; Tanzawa, K.; Kaneko, M. Molecular recognition of adenophostin, a very potent Ca2+ inducer, at the D-myo-inositol 1, 4, 5-trisphosphate receptor. Biochemistry 1999, 38, 9234–9241. [Google Scholar] [CrossRef]
- Bosanac, I.; Alattia, J.R.; Mal, T.K.; Chan, J.; Talarico, S.; Tong, F.K.; Tong, K.I.; Yoshikawa, F.; Furuichi, T.; Iwai, M. Structure of the inositol 1, 4, 5-trisphosphate receptor binding core in complex with its ligand. Nature 2002, 420, 696. [Google Scholar] [CrossRef] [PubMed]
- Bosanac, I.; Yamazaki, H.; Matsuura, T.; Michikawa, T.; Mikoshiba, K.; Ikura, M. Crystal structure of the ligand binding suppressor domain of type 1 inositol 1, 4, 5-trisphosphate receptor. Mol. Cell 2005, 17, 193–203. [Google Scholar] [CrossRef]
- Foskett, J.K.; White, C.; Cheung, K.H.; Mak, D.D. Inositol trisphosphate receptor Ca2+ release channels. Physiol. Rev. 2007, 87, 593–658. [Google Scholar] [CrossRef] [Green Version]
- Marchant, J.S.; Beecroft, M.D.; Riley, A.M.; Jenkins, D.J.; Marwood, R.D.; Taylor, C.W.; Potter, B.V.L. Disaccharide polyphosphates based upon adenophostin A activate hepatic D-myo-inositol 1, 4, 5-trisphosphate receptors. Biochemistry 1997, 36, 12780–12790. [Google Scholar] [CrossRef] [PubMed]
- Glouchankova, L.; Krishna, U.M.; Potter, B.V.L.; Falck, J.R.; Bezprozvanny, I. Association of the inositol (1, 4, 5)-trisphosphate receptor ligand binding site with phosphatidylinositol (4, 5)-bisphosphate and adenophostin A. Mol. Cell Biol. Res. Commun. 2000, 3, 153–158. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Uchiyama, T.; Yoshikawa, F.; Hishida, A.; Furuichi, T.; Mikoshiba, K. A novel recombinant hyperaffinity inositol 1, 4, 5-trisphosphate (IP3) absorbent traps IP3, resulting in specific inhibition of IP3-mediated calcium signaling. J. Biol. Chem. 2002, 277, 8106–8113. [Google Scholar] [CrossRef] [Green Version]
- Chan, J.; Yamazaki, H.; Ishiyama, N.; Seo, M.D.; Mal, T.K.; Michikawa, T.; Mikoshiba, K.; Ikura, M. Structural studies of inositol 1, 4, 5-trisphosphate receptor coupling ligand binding to channel gating. J. Biol. Chem. 2010, 285, 36092–36099. [Google Scholar] [CrossRef] [Green Version]
- Lin, C.-C.; Baek, K.; Lu, Z. Apo and InsP 3-bound crystal structures of the ligand-binding domain of an InsP 3 receptor. Nat. Struct. Mol. Biol. 2011, 18, 1172. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Caron, G.; Ermondi, G. Influence of conformation on GRIND-based three-dimensional quantitative structure−activity relationship (3D-QSAR). J. Med. Chem. 2007, 50, 5039–5042. [Google Scholar] [CrossRef]
- Warren, G.L.; Andrews, C.W.; Capelli, A.M.; Clarke, B.; LaLonde, J.; Lambert, M.H.; Lindvall, M.; Nevins, N.; Semus, S.F.; Senger, S. A critical assessment of docking programs and scoring functions. J. Med. Chem. 2006, 49, 5912–5931. [Google Scholar] [CrossRef] [PubMed]
- Tropsha, A.; Gramatica, P.; Gombar, V.K. The importance of being earnest: Validation is the absolute essential for successful application and interpretation of QSPR models. QSAR Comb. Sci. 2003, 22, 69–77. [Google Scholar] [CrossRef]
- Adunyah, S.E.; Dean, W. Effects of sulfhydryl reagents and other inhibitors on Ca2+ transport and inositol trisphosphate-induced Ca2+ release from human platelet membranes. J. Biol. Chem. 1986, 261, 13071–13075. [Google Scholar] [CrossRef]
- Hill, T.D.; Berggren, P.-O.; Boynton, A.L. Heparin inhibits inositol trisphosphate-induced calcium release from permeabilized rat liver cells. Biochem. Biophys. Res. Commun. 1987, 149, 897–901. [Google Scholar] [CrossRef]
- Föhr, K.; Wahl, Y.; Engling, R.; Kemmer, T.P.; Gratzl, M. Decavanadate displaces inositol 1, 4, 5-trisphosphate (IP3) from its receptor and inhibits IP3 induced Ca2+ release in permeabilized pancreatic acinar cells. Cell Calcium 1991, 12, 735–742. [Google Scholar] [CrossRef] [Green Version]
- Poitras, M.; Bernier, S.; Boulay, G.; Fournier, A.; Guillemette, G. Interaction of benzene 1, 2, 4-trisphosphate with inositol 1, 4, 5-trisphosphate receptor and metabolizing enzymes. Eur. J. Pharmacol. Mol. Pharmacol. 1993, 244, 203–210. [Google Scholar] [CrossRef]
- Sczekan, S.R.; Strumwasser, F. Antipsychotic drugs block IP3-dependent Ca2+-release from rat brain microsomes. Biol. Psychiatry 1996, 40, 497–502. [Google Scholar] [CrossRef]
- Snyder, S.H.; Axelrod, J.; Zweig, M. A sensitive and specific fluorescence assay for tissue serotonin. Biochem. Pharmacol. 1965, 14, 831–835. [Google Scholar] [CrossRef]
- McIntyre, J.C.; Sleight, R.G. Fluorescence assay for phospholipid membrane asymmetry. Biochemistry 1991, 30, 11819–11827. [Google Scholar] [CrossRef]
- Csizmadia, P. MarvinSketch and MarvinView: Molecule Applets for the World Wide Web. In Proceedings of the The 3rd International Electronic Conference on Synthetic Organic Chemistry, Budapest, Hungary, 1–30 September 1999. [Google Scholar] [CrossRef]
- Halgren, T.A. Merck molecular force field. II. MMFF94 van der Waals and electrostatic parameters for intermolecular interactions. J. Comput. Chem. 1996, 17, 520–552. [Google Scholar] [CrossRef]
- Gillet, V.J. Diversity selection algorithms. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2011, 1, 580–589. [Google Scholar] [CrossRef]
- Matter, H. Selecting optimally diverse compounds from structure databases: A validation study of two-dimensional and three-dimensional molecular descriptors. J. Med. Chem. 1997, 40, 1219–1229. [Google Scholar] [CrossRef] [PubMed]
- Schmuker, M.; Givehchi, A.; Schneider, G. Impact of different software implementations on the performance of the Maxmin method for diverse subset selection. Mol. Divers. 2004, 8, 421–425. [Google Scholar] [CrossRef] [PubMed]
- Mukhtar, S.; Kiani, Y.S.; Jabeen, I. Molecular docking simulations and GRID-independent molecular descriptor (GRIND) analysis to probe stereoselective interactions of CYP3A4 inhibitors. Med. Chem. Res. 2017, 26, 2322–2335. [Google Scholar] [CrossRef]
- Zafar, S.; Jabeen, I. GRID-independent molecular descriptor analysis and molecular docking studies to mimic the binding hypothesis of γ-aminobutyric acid transporter 1 (GAT1) inhibitors. PeerJ 2019, 7, e6283. [Google Scholar] [CrossRef] [PubMed]
- Munawar, S.; Vandenberg, J.I.; Jabeen, I. Molecular Docking Guided Grid-Independent Descriptor Analysis to Probe the Impact of Water Molecules on Conformational Changes of hERG Inhibitors in Drug Trapping Phenomenon. Int. J. Mol. Sci. 2019, 20, 3385. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Wang, L.-P.; McKiernan, K.A.; Gomes, J.; Beauchamp, K.A.; Head Gordon, T.; Rice, J.E.; Swope, W.C.; Martínez, T.J.; Pande, V.S. Building a more predictive protein force field: A systematic and reproducible route to AMBER-FB15. J. Phys. Chem. B 2017, 121, 4023–4039. [Google Scholar] [CrossRef]
- Nabuurs, S.B.; Wagener, M.; de Vlieg, J. A flexible approach to induced fit docking. J. Med. Chem. 2007, 50, 6507–6518. [Google Scholar] [CrossRef] [PubMed]
- Wilcox, R.A.; Primrose, W.U.; Nahorski, S.R.; Challiss, R.A.J. New developments in the molecular pharmacology of the myo-inositol 1, 4, 5-trisphosphate receptor. Trends Pharmacol. Sci. 1998, 19, 467–475. [Google Scholar] [CrossRef]
- Dobson, P.D.; Kell, D.B. Carrier-mediated cellular uptake of pharmaceutical drugs: An exception or the rule? Nat. Rev. Drug Discov. 2008, 7, 205. [Google Scholar] [CrossRef]
- Leo, A.J.; Hansch, C. Role of hydrophobic effects in mechanistic QSAR. Perspect. Drug Discov. Des. 1999, 17, 1–25. [Google Scholar] [CrossRef]
- Arnott, J.A.; Planey, S.L. The influence of lipophilicity in drug discovery and design. Expert Opin. Drug Discov. 2012, 7, 863–875. [Google Scholar] [CrossRef]
- Keserü, G.M.; Makara, G.M. The influence of lead discovery strategies on the properties of drug candidates. Nat. Rev. Drug Discov. 2009, 8, 203. [Google Scholar] [CrossRef]
- Mortenson, P.N.; Murray, C.W. Assessing the lipophilicity of fragments and early hits. J. Comput. Aided Mol. Des. 2011, 25, 663–667. [Google Scholar] [CrossRef]
- Hopkins, A.L.; Keserü, G.M.; Leeson, P.D.; Rees, D.C.; Reynolds, C.H. The role of ligand efficiency metrics in drug discovery. Nat. Rev. Drug Discov. 2014, 13, 105–121. [Google Scholar] [CrossRef]
- Wolber, G.; Dornhofer, A.A.; Langer, T. Efficient overlay of small organic molecules using 3D pharmacophores. J. Comput. Aided Mol. Des. 2006, 20, 773–788. [Google Scholar] [CrossRef] [PubMed]
- Wolber, G.; Langer, T. LigandScout: 3-D pharmacophores derived from protein-bound ligands and their use as virtual screening filters. J. Chem. Inf. Modeling 2005, 45, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Poli, G.; Seidel, T.; Langer, T. Conformational Sampling of Small Molecules With iCon: Performance Assessment in Comparison With OMEGA. Front. Chem. 2018, 6, 229. [Google Scholar] [CrossRef]
- Hodgkin, E.; Richards, W. Molecular similarity. Chem. Ber. 1988, 24, 1141. [Google Scholar]
- Durán, Á.; Pastor, M. An Advanced Tool for Computing and Handling GRid-INdependent. Descriptors; User Manual Version; 2011; Volume 1. [Google Scholar]
- Gasteiger, J.; Rudolph, C.; Sadowski, J. Automatic generation of 3D-atomic coordinates for organic molecules. Tetrahedron Comput. Methodol. 1990, 3, 537–547. [Google Scholar] [CrossRef]
- Artese, A.; Cross, S.; Costa, G.; Distinto, S.; Parrotta, L.; Alcaro, S.; Ortuso, F.; Cruciani, G. Molecular interaction fields in drug discovery: Recent advances and future perspectives. Wiley Interdisciplinary Reviews. Comput. Mol. Sci. 2013, 3, 594–613. [Google Scholar] [CrossRef]
- Mannhold, R.; Kubinyi, H.; Folkers, G. Molecular Interaction Fields: Applications in Drug Discovery and ADME Prediction; John Wiley & Sons: Hoboken, NJ, USA, 2006; Volume 27. [Google Scholar]
- Shu, Z.; Davies, G. Calculation of the Lennard-Jones nm potential energy parameters for metals. Physica Status Solidi. A Appl. Res. 1983, 78, 595–605. [Google Scholar]
- Hajigeorgiou, P.G. An extended Lennard-Jones potential energy function for diatomic molecules: Application to ground electronic states. J. Mol. Spectrosc. 2010, 263, 101–110. [Google Scholar] [CrossRef]
- Duran, A.; Martínez, G.C.; Pastor, M. Development and validation of AMANDA, a new algorithm for selecting highly relevant regions in molecular interaction fields. J. Chem. Inf. Modeling 2008, 48, 1813–1823. [Google Scholar] [CrossRef] [Green Version]
- Durán, A.Á. Development of high-performance algorithms for a new generation of versatile molecular descriptors. In The Pentacle Software; Universitat Pompeu Fabra: Barcelona, Spain, 2010; Available online: http://hdl.handle.net/10803/7201 (accessed on 17 September 2021).





| Inositol Phosphate (IP) (Class A) |  | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Comp. No. | R1 | R2 | R3 | R4 | R5 | R6 | Conformation | Key Name | IC50 (µM) | logP | clogP | pIC50 | LipE | Ref. | |
| A1 | PO3−2 | PO3−2 | OH | PO3−2 | PO3−2 | OH | R,S,S,S,S,S | DL-Ins(1,2,4,5)P4 | 0.03 | −7.5 | −7.2 | 1.6 | 14.8 | [41] | |
| A2 | PO3−2 | PO3−2 | OH | PO3−2 | PO3−2 | OH | S,S,S,R,R,R | scyllo-Ins(1,2,4,5)P4 | 0.02 | −7.5 | −7.2 | 1.8 | 15.1 | [42] | |
| A3 | PO3−2 | PO3−2 | OH | PO3−2 | OH | OH | S,S,R,R,R,R | DL-scyllo-Ins(1,2,4)P3 | 0.05 | −6.4 | −5.7 | 1.3 | 13.1 | [41] | |
| A4 | PO3−2 | OH | PO3−2 | PO3−2 | PO3−2 | OH | R,S,S,S,S,S | Ins(1,3,4,5)P4 | 0.01 | −7.5 | −6.5 | 2.5 | 15.1 | [42] | |
| A5 | PO3−2 | OH | PO3−2 | PO3−2 | OH | PO3−2 | R,S,R,S,S,R | D-chiro-Ins(1,3,4,6)P4 | 0.17 | −7.5 | −6.7 | 0.7 | 13.4 | [42] | |
| A6 | PO3−2 | OH | OH | PO3−2 | PO3−2 | PO3−2 | R,S,S,R,R,S | Ins(1,4,5,6)P4 | 0.43 | −7.7 | −8.5 | 0.2 | 14.9 | [41] | |
| A7 | PO3−2 | OH | OH | PO3−2 | PO3−2 | OH | R,R,S,R,R,S | Ins(1,4,5)P3 | 3.01 | −6.4 | −5.8 | 2.2 | 14.1 | [42] | |
| A8 | PO3−2 | OH | OH | OH | PO3−2 | PO3−2 | R,R,S,R,R,S | Ins(1,5,6)P3 | 0.04 | −6.2 | −5.8 | 0.4 | 13.1 | [42] | |
| A9 | OH | OH | PO3−2 | PO3−2 | PO3−2 | PO3−2 | S,R,R,S,R,S | Ins(3,4,5,6)P4 | 0.62 | −7.7 | −7.2 | 1.3 | 13.4 | [41] | |
| A10 | OH | OH | PO3−2 | PO3−2 | PO3−2 | OH | S,S,R,R,S,S | Ins(3,4,5)P3 | 0.01 | −6.6 | −5.7 | 1.9 | 13.9 | [41] | |
| A11 | OH | OH | OH | PO3−2 | PO3−2 | PO3−2 | R,S,S,S,R,S | Ins(4,5,6)P3 | 93.0 | −6.9 | −5.8 | −1.3 | 9.8 | [43] | |
| A12 | OH | OH | OH | PO3−2 | PO3−2 | OH | R,R,S,S,R,S | Ins(4, 5)P2 | 20.0 | −5.5 | −4.3 | −0.5 | 9.1 | [43] | |
| Xestospongins (Xe) (Class B) |  | ||||||||||||||
| Comp. No. | R1 | R4 | R5 | R8 | Conformation | Key Name | IC50 (µM) | logP | clogP | pIC50 | LipE | Ref. | |||
| B1 | OH | --- | OH | --- | R,R,S,R,R,S | Araguspongine C | 6.60 | 5.7 | 4.7 | 5.2 | 0.5 | [44] | |||
| B2 | OH | --- | --- | CH3 | S,S,R,S,R,R,R | Xestospongin B | 5.01 | 6.8 | 7.2 | 5.3 | −1.9 | [45] | |||
| B3 | OH | --- | --- | --- | S,S,R,R,S,R | Demethylated Xestospongin B | 5.86 | 6.5 | 6.8 | 5.2 | −1.5 | [46] | |||
| B4 | --- | OH | --- | --- | S,S,R,R,S,S,R | 7-(OH)-XeA | 6.40 | 6.3 | 6.8 | 5.2 | −1.5 | [44] | |||
| B5 | --- | --- | --- | --- | S,S,R,S,S,R | Xestospongin A | 2.53 | 7.3 | 8.1 | 5.6 | −2.4 | [46] | |||
| B6 | --- | --- | --- | --- | R,S,R,R,S,R | Araguspongine B | 0.65 | 7.3 | 8.0 | 6.2 | −1.8 | [46] | |||
| Benzene Phosphate Derivatives (Class C) |  | ||||||||||||||
| Comp. No. | R2 | R2′ | R3′ | R4 | R4′ | R5 | R5′ | R6 | Key Name | IC50 (µM) | logP | clogP | pIC50 | LipE | Ref. |
| C1 | PO3−2 | --- | PO3−2 | PO3−2 | --- | ---- | PO3−2 | PO3−2 | BiPh(2,3′,4,5′,6)P5 | 0.42 | −1.2 | −4.2 | 6.3 | 14.9 | [47] |
| C2 | PO3−2 | PO3−2 | --- | PO3−2 | PO3−2 | PO3−2 | PO3−2 | --- | BiPh(2,2′4,4′,5,5′)P6 | 0.19 | −2.8 | −6.1 | 6.7 | 17.2 | [47] |
| C3 | PO3−2 | PO3−2 | --- | PO3−2 | PO3−2 | PO3−2 | PO3−2 | --- | 1,2,4-Dimer Biph(2,2′,4,4′,5,5′)P6 | 0.38 | −3.9 | −8.2 | 6.4 | 14.7 | [47] |
| Model No. | Pharmacophore Model (Template) | Model Score | Model Distance | Model Statistics | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1. |  | 0.68 * | Hyd | HBA1 | HBA2 | HBD1 | HBD2 | TP: TN: FP: FN: MCC: | 87% 72% 06% 03% 0.76 | |
| Hyd | 0 | |||||||||
| HBA1 | 2.62 | 0 | ||||||||
| HBA2 | 4.79 | 2.61 | 0 | |||||||
| HBD1 | 5.56 | 3.64 | 4.57 | 0 | ||||||
| HBD2 | 7.68 | 5.58 | 3.11 | 6.97 | 0 | |||||
| 2. |  | 0.67 | Hyd | HBA1 | HBD1 | HBD2 | HBD3 | TP: TN: FP: FN: MCC: | 51% 70% 14% 18% 0.26 | |
| Hyd | 0 | |||||||||
| HBA1 | 2.48 | 0 | ||||||||
| HBD1 | 3.46 | 4.17 | 0 | |||||||
| HBD2 | 5.56 | 3.63 | 6.33 | 0 | ||||||
| HBD3 | 7.43 | 5.58 | 7.8 | 7.01 | 0 | |||||
| 3. |  | 0.66 | Hyd | HBA | HBD1 | HBD2 | HBD3 | TP: TN: FP: FN: MCC: | 72% 29% 12% 33% 0.02 | |
| Hyd | 0 | |||||||||
| HBA | 3.95 | 0 | ||||||||
| HBD1 | 3.97 | 3.87 | 0 | |||||||
| HBD2 | 7.09 | 4.13 | 2.86 | 0 | ||||||
| HBD3 | 7.29 | 3.41 | 7.01 | 2.62 | 0 | |||||
| 4. |  | 0.65 | Hyd | HBA | HBD1 | HBD2 | Hyd | TP: TN: FP: FN: MCC: | 49% 71% 14% 27% 0.23 | |
| Hyd | 0 | |||||||||
| HBA | 2.32 | 0 | ||||||||
| HBD1 | 3.19 | 1.62 | 0 | |||||||
| HBD2 | 7.69 | 6.91 | 4.57 | 0 | ||||||
| Hyd | 6.22 | 4.41 | 3.17 | 2.04 | 0 | |||||
| 5. |  | 0.64 | Hyd | HBA | HBD1 | HBD2 | HBD3 | TP: TN: FP: FN: MCC: | 54% 57% 28% 27% 0.13 | |
| Hyd | 0 | |||||||||
| HBA | 2.32 | 0 | ||||||||
| HBD1 | 4.56 | 3.01 | 0 | |||||||
| HBD2 | 2.92 | 1.05 | 3.61 | 0 | ||||||
| HBD3 | 7.06 | 5.09 | 7.53 | 5.28 | 0 | |||||
| 6. |  | 0.63 | Hyd | HBA1 | HBA2 | HBD1 | HBD2 | TP: TN: FP: FN: MCC: | 60% 29% 57% 45% −0.07 | |
| Hyd | 0 | |||||||||
| HBA1 | 4.32 | 0 | ||||||||
| HBA2 | 4.46 | 2.21 | 0 | |||||||
| HBD1 | 6.87 | 3.07 | 5.73 | 0 | ||||||
| HBD2 | 4.42 | 6.05 | 5.04 | 9.61 | 0 | |||||
| 7. |  | 0.62 | Hyd | HBA | HBD1 | HBD2 | HBD3 | TP: TN: FP: FN: MCC: | 63% 71% 14% 42% 0.32 | |
| Hyd | 0 | |||||||||
| HBA | 2.49 | 0 | ||||||||
| HBD1 | 4.06 | 2.07 | 0 | |||||||
| HBD2 | 5.08 | 2.8 | 2.38 | 0 | ||||||
| HBD3 | 6.1 | 6.48 | 8.87 | 6.56 | 0 | |||||
| 8. |  | 0.61 | Hyd | HBA1 | HBA2 | HBD | TP: TN: FP: FN: MCC: | 55% 57% 42% 48% 0.08 | ||
| Hyd | 0 | |||||||||
| HBA1 | 4.28 | 0 | ||||||||
| HBA2 | 4.26 | 2.8 | 0 | |||||||
| HBD | 7.08 | 6.94 | 5.42 | 0 | ||||||
| 9. |  | 0.60 | HBA1 | HBA2 | HBA3 | HBD1 | HBD2 | TP: TN: FP: FN: MCC: | 58% 28% 57% 48% −0.09 | |
| HBA1 | 0 | |||||||||
| HBA2 | 2.52 | 0 | ||||||||
| HBA3 | 2.05 | 2.07 | 0 | |||||||
| HBD1 | 4.65 | 2.28 | 4.06 | 0 | ||||||
| HBD2 | 6.9 | 7.96 | 5.75 | 8.96 | 0 | |||||
| 10. |  | 0.60 | HBA1 | HBA2 | HBD1 | HBD2 | TP: TN: FP: FN: MCC: | 51% 42% 40% 54% −0.01 | ||
| HBA1 | 0 | |||||||||
| HBA2 | 3.26 | 0 | ||||||||
| HBD1 | 3.65 | 6.06 | 0 | |||||||
| HBD2 | 6.96 | 6.09 | 6.33 | 0 | ||||||
| Conformational Method | Fractional Factorial Design (FFD) Cycle | Comments FFD2 (LV2) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Complete | FFD1 | FFD2 | ||||||||
| Q2 LOO | R2 | SDEP | Q2 LOO | R2 | SDEP | Q2 LOO | R2 | SDEP | ||
| Energy Minimized | 0.07 | 0.93 | 2.8 | 0.12 | 0.93 | 2.7 | 0.23 | 0.94 | 2.5 | Inconsistent for auto- and cross-GRID variables |
| Standard 3D | 0.59 | 0.68 | 3.5 | 0.15 | 0.56 | 3.5 | 0.05 | 0.53 | 3.5 | Inconsistent for auto- and cross-GRID variables |
| Induced Fit Docked | 0.61 | 0.64 | 1.1 | 0.68 | 0.71 | 1.0 | *0.70 | 0.72 | 0.9 | Consistent for Dry-Dry, Dry-O, Dry-N1, and Dry-Tip correlogram (Figure 3) |
| Pharmacophore (Ligand-Based) | GRIND (Correlogram) | |||
|---|---|---|---|---|
| Pharmacophore Variables | Distances | GRIND Variables | Features at VRS | Distance |
| Hydro-HBA Hydro-HBD HBD-HBD | 4.79 Å 5.56 Å 6.97 Å | Dry-N1 Dry-O O-O | Hyd-HBD Hyd-HBA HBA-HBA | 7.6–8 Å 6.8–7.2 Å 10.4–10.8 Å |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ismatullah, H.; Jabeen, I. Combined Pharmacophore and Grid-Independent Molecular Descriptors (GRIND) Analysis to Probe 3D Features of Inositol 1,4,5-Trisphosphate Receptor (IP3R) Inhibitors in Cancer. Int. J. Mol. Sci. 2021, 22, 12993. https://doi.org/10.3390/ijms222312993
Ismatullah H, Jabeen I. Combined Pharmacophore and Grid-Independent Molecular Descriptors (GRIND) Analysis to Probe 3D Features of Inositol 1,4,5-Trisphosphate Receptor (IP3R) Inhibitors in Cancer. International Journal of Molecular Sciences. 2021; 22(23):12993. https://doi.org/10.3390/ijms222312993
Chicago/Turabian StyleIsmatullah, Humaira, and Ishrat Jabeen. 2021. "Combined Pharmacophore and Grid-Independent Molecular Descriptors (GRIND) Analysis to Probe 3D Features of Inositol 1,4,5-Trisphosphate Receptor (IP3R) Inhibitors in Cancer" International Journal of Molecular Sciences 22, no. 23: 12993. https://doi.org/10.3390/ijms222312993







