The Rice CHD3/Mi-2 Chromatin Remodeling Factor Rolled Fine Striped Promotes Flowering Independent of Photoperiod
Abstract
:1. Introduction
2. Results
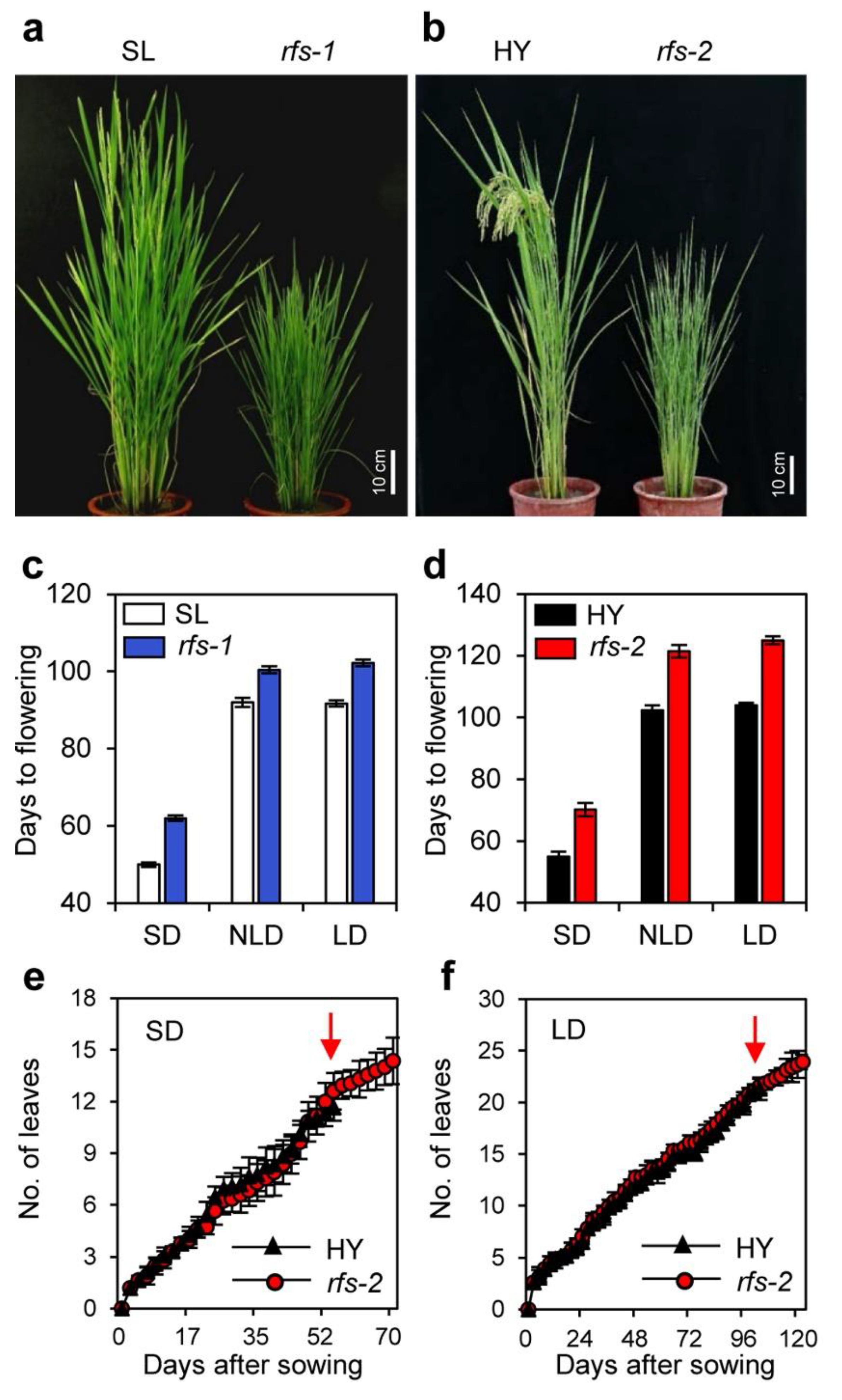
2.1. The rfs Mutants Exhibit a Late Flowering Phenotype Independent of the Photoperiod
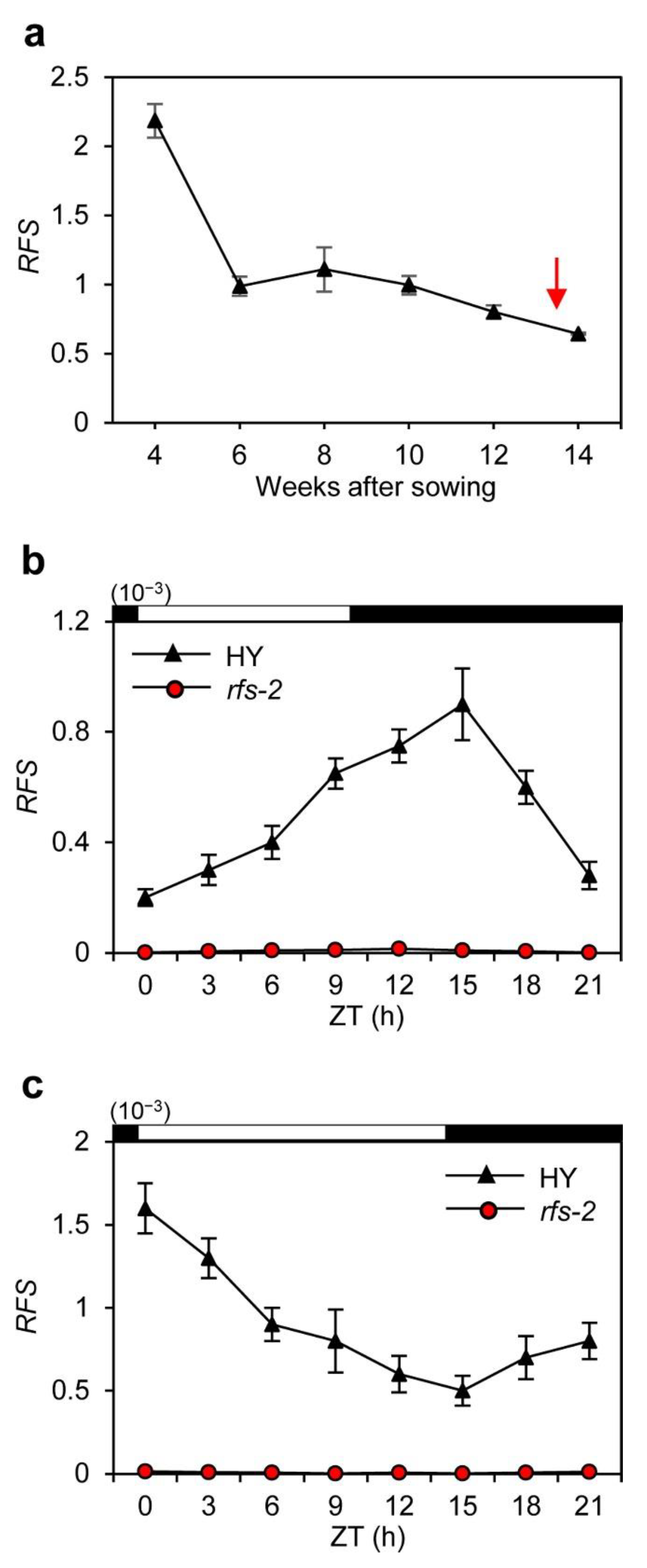
2.2. Expression Pattern of RFS
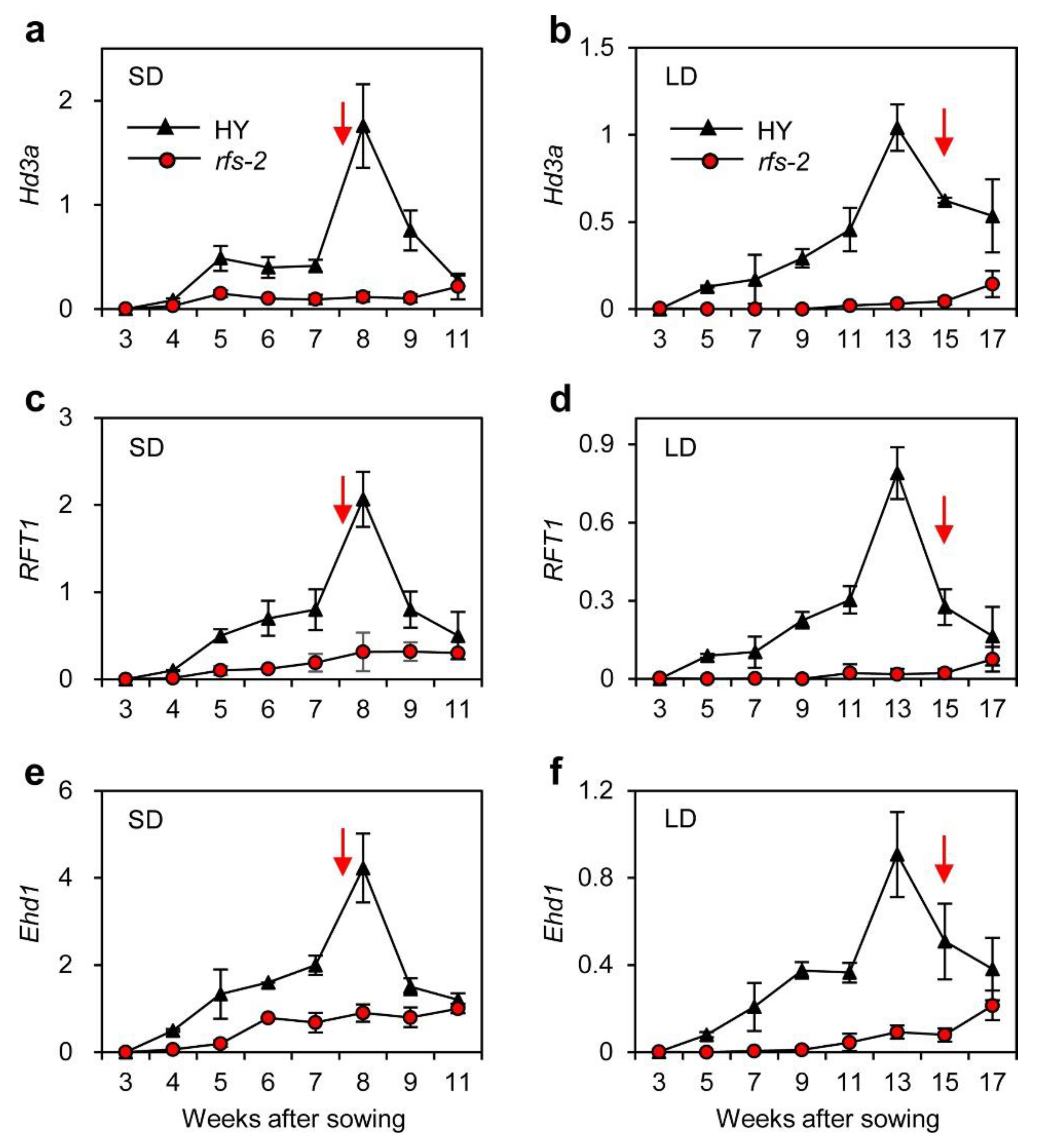
2.3. Expression Patterns of Hd3a and RFT1 in the rfs Mutants
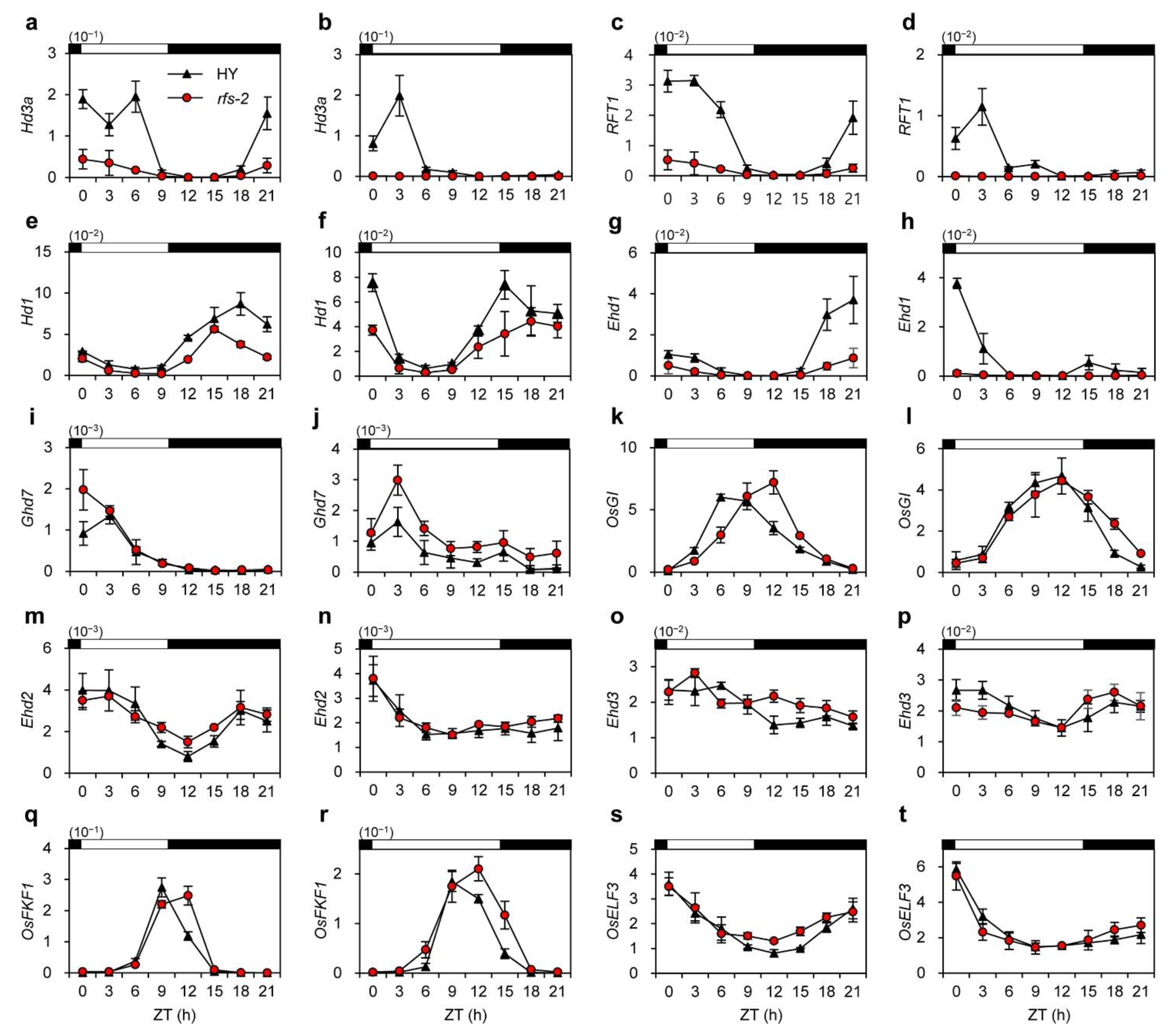
2.4. Expression Analysis of Flowering-Time Genes in the rfs Mutants
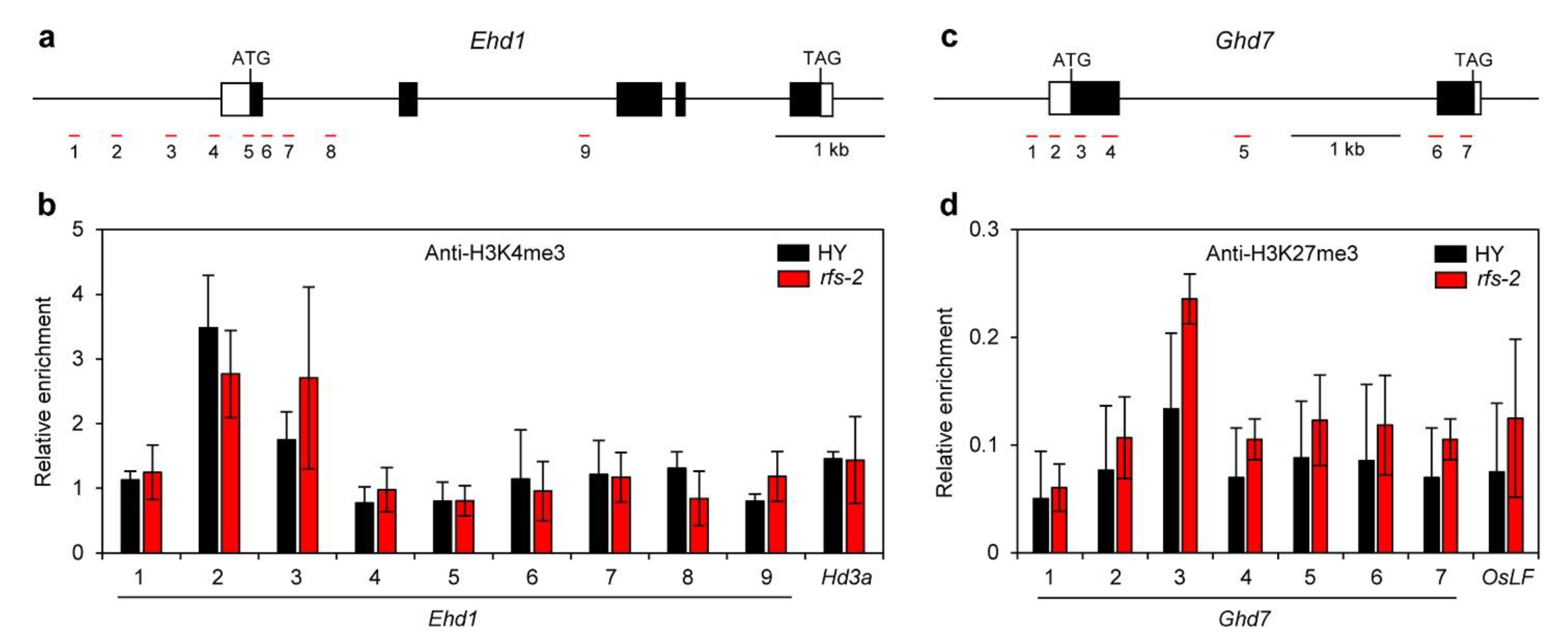
2.5. Histone Methylation Levels of Ehd1 and Ghd7 in the rfs Mutants
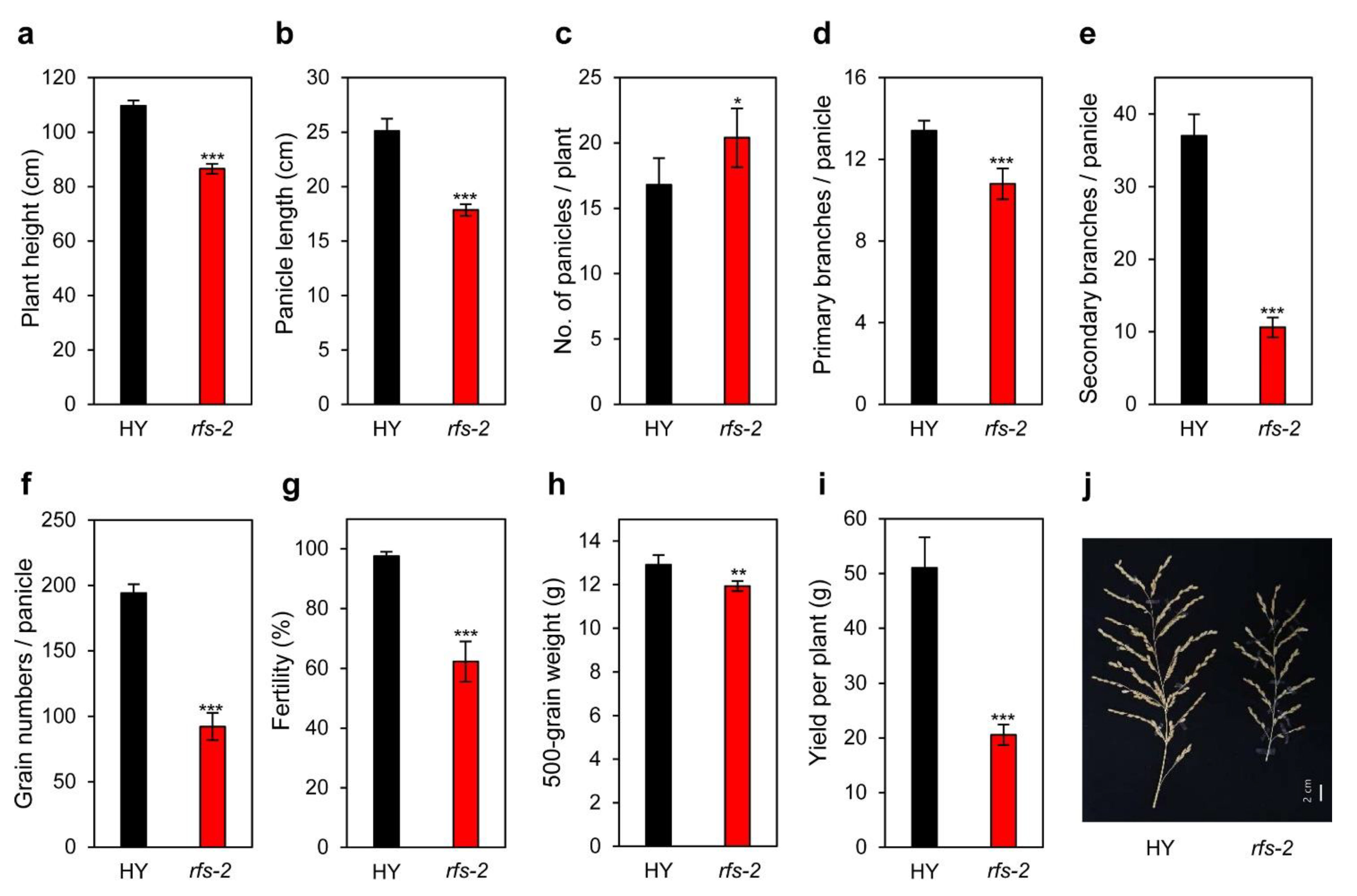
2.6. Agronomic Traits of the rfs Mutants
3. Discussion
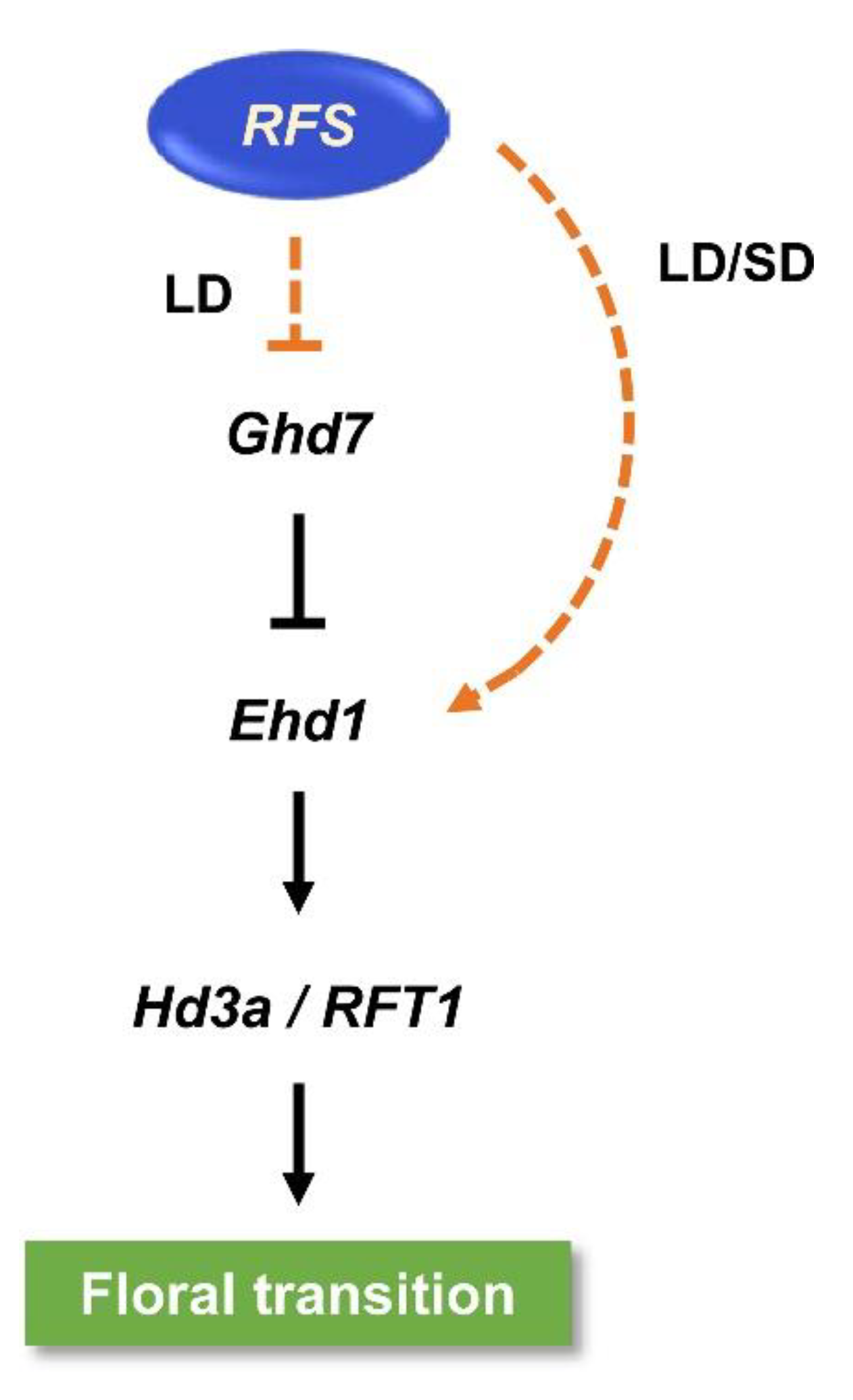
3.1. Regulatory Roles of RFS in Flowering-Time Pathways
3.2. RFS May Control Expression of Flowering-Time Genes via Histone Modification
3.3. RFS Is Involved in Inflorescence Development
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. RNA Extraction and Reverse Transcription-Quantitative PCR (RT-qPCR) Analysis
4.3. Chromatin Immunoprecipitation (ChIP) Assay
4.4. Measurement of Agronomic Traits
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| RFS | Rolled Fine Striped |
| SD | Short day |
| LD | Long day |
| NLD | Natural long day |
| H3K4me3 | Trimethylation of histone H3 lysine 4 |
| H3K27me3 | Trimethylation of histone H3 lysine 27 |
| SL | Seolak |
| HY | Hwayoung |
| ZT | Zeitgeber time |
| DAS | Days after sowing |
| RT-qPCR | Reverse transcription quantitative PCR |
| ChIP | Chromatin Immunoprecipitation |
References
- Komiya, R.; Ikegami, A.; Tamaki, S.; Yokoi, S.; Shimamoto, K. Hd3a and RFT1 are essential for flowering in rice. Development 2008, 135, 767–774. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsuji, H.; Taoka, K.; Shimamoto, K. Regulation of flowering in rice: Two florigen genes, a complex gene network, and natural variation. Curr. Opin. Plant Biol. 2011, 14, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Yano, M.; Katayose, Y.; Ashikari, M.; Yamanouchi, U.; Monna, L.; Fuse, T.; Baba, T.; Yamamoto, K.; Umehara, Y.; Nagamura, Y.; et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 2000, 12, 2473–2484. [Google Scholar] [PubMed] [Green Version]
- Doi, K.; Izawa, T.; Fuse, T.; Yamanouchi, U.; Kubo, T.; Shimatani, Z.; Yano, M.; Yoshimura, A. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev. 2004, 18, 926–936. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xue, W.; Xing, Y.; Weng, X.; Zhao, Y.; Tang, W.; Wang, L.; Zhou, H.; Yu, S.; Xu, C.; Li, X.; et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 2008, 40, 761–767. [Google Scholar] [CrossRef]
- Matsubara, K.; Yamanouchi, U.; Wang, Z.X.; Minobe, Y.; Izawa, T.; Yano, M. Ehd2, a rice ortholog of the maize INDETERMINATE1 gene, promotes flowering by up-regulating Ehd1. Plant Physiol. 2008, 148, 1425–1435. [Google Scholar] [CrossRef] [Green Version]
- Park, S.J.; Kim, S.L.; Lee, S.; Je, B.I.; Piao, H.L.; Park, S.H.; Kim, C.M.; Ryu, C.H.; Park, S.H.; Xuan, Y.H.; et al. Rice Indeterminate 1 (OsId1) is necessary for the expression of Ehd1 (Early heading date 1) regardless of photoperiod. Plant J. 2008, 56, 1018–1029. [Google Scholar] [CrossRef]
- Wu, C.; You, C.; Li, C.; Long, T.; Chen, G.; Byrne, M.E.; Zhang, Q. RID1, encoding a Cys2/His2-type zinc finger transcription factor, acts as a master switch from vegetative to floral development in rice. Proc. Natl. Acad. Sci. USA 2008, 105, 12915–12920. [Google Scholar] [CrossRef] [Green Version]
- Matsubara, K.; Yamanouchi, U.; Nonoue, Y.; Sugimoto, K.; Wang, Z.X.; Minobe, Y.; Yano, M. Ehd3, encoding a plant homeodomain finger-containing protein, is a critical promoter of rice flowering. Plant J. 2011, 66, 603–612. [Google Scholar] [CrossRef]
- Lee, Y.S.; Jeong, D.H.; Lee, D.Y.; Yi, J.; Ryu, C.H.; Kim, S.L.; Jeong, H.J.; Choi, S.C.; Jin, P.; Yang, J.; et al. OsCOL4 is a constitutive flowering repressor upstream of Ehd1 and downstream of OsphyB. Plant J. 2010, 63, 18–30. [Google Scholar] [CrossRef]
- Niu, Y.X.; Bai, J.; Zheng, S.Z. The Regulation and Function of Histone Methylation. J. Plant Biol. 2018, 61, 347–357. [Google Scholar] [CrossRef]
- Liu, B.; Wei, G.; Shi, J.; Jin, J.; Shen, T.; Ni, T.; Shen, W.H.; Yu, Y.; Dong, A. SET DOMAIN GROUP 708, a histone H3 lysine 36-specific methyltransferase, controls flowering time in rice (Oryza sativa). New Phytol. 2016, 210, 577–588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, K.; Yu, Y.; Dong, A.; Shen, W.H. SET DOMAIN GROUP701 encodes a H3K4-methyltransferase and regulates multiple key processes of rice plant development. New Phytol. 2017, 215, 609–623. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, S.C.; Lee, S.; Kim, S.R.; Lee, Y.S.; Liu, C.; Cao, X.; An, G. Trithorax group protein Oryza sativa Trithorax1 controls flowering time in rice via interaction with early heading date3. Plant Physiol. 2014, 164, 1326–1337. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, P.; Wang, S.; Jiang, H.; Cheng, B.; Wu, K.; Ding, Y. The COMPASS-Like Complex Promotes Flowering and Panicle Branching in Rice. Plant Physiol. 2018, 176, 2761–2771. [Google Scholar] [CrossRef] [Green Version]
- Jiang, P.; Wang, S.; Zheng, H.; Li, H.; Zhang, F.; Su, Y.; Xu, Z.; Lin, H.; Qian, Q.; Ding, Y. SIP1 participates in regulation of flowering time in rice by recruiting OsTrx1 to Ehd1. New Phytol. 2018, 219, 422–435. [Google Scholar] [CrossRef] [Green Version]
- Sun, C.; Fang, J.; Zhao, T.; Xu, B.; Zhang, F.; Liu, L.; Tang, J.; Zhang, G.; Deng, X.; Chen, F.; et al. The histone methyltransferase SDG724 mediates H3K36me2/3 deposition at MADS50 and RFT1 and promotes flowering in rice. Plant Cell 2012, 24, 3235–3247. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.; Liu, Y.; Wang, B.; Luo, Q.; Shi, J.; Gan, J.; Shen, W.H.; Yu, Y.; Dong, A. The transcription factor OsSUF4 interacts with SDG725 in promoting H3K36me3 establishment. Nat. Commun. 2019, 10, 2999. [Google Scholar] [CrossRef]
- Sui, P.; Shi, J.; Gao, X.; Shen, W.H.; Dong, A. H3K36 methylation is involved in promoting rice flowering. Mol. Plant 2013, 6, 975–977. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Hu, J.; Qian, Q.; Xue, H.W. LC2 and OsVIL2 promote rice flowering by photoperiod-induced epigenetic silencing of OsLF. Mol. Plant 2013, 6, 514–527. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Lee, S.; Hang, R.; Kim, S.R.; Lee, Y.S.; Cao, X.; Amasino, R.; An, G. OsVIL2 functions with PRC2 to induce flowering by repressing OsLFL1 in rice. Plant J. 2013, 73, 566–578. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Zhou, C.; Zhao, Y.; Zhou, S.; Wang, W.; Zhou, D.X. The rice enhancer of zeste [E(z)] genes SDG711 and SDG718 are respectively involved in long day and short day signaling to mediate the accurate photoperiod control of flowering time. Front. Plant Sci. 2014, 5, 591. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fu, X.; Li, C.; Liang, Q.; Zhou, Y.; He, H.; Fan, L.M. CHD3 chromatin-remodeling factor PICKLE regulates floral transition partially via modulating LEAFY expression at the chromatin level in Arabidopsis. Sci. China Life Sci. 2016, 59, 516–528. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, J.; Oh, D.H.; Dassanayake, M.; Nguyen, K.T.; Ogas, J.; Choi, G.; Sun, T.P. Gibberellin Signaling Requires Chromatin Remodeler PICKLE to Promote Vegetative Growth and Phase Transitions. Plant Physiol. 2017, 173, 1463–1474. [Google Scholar] [CrossRef] [Green Version]
- Jing, Y.; Guo, Q.; Zha, P.; Lin, R. The chromatin-remodelling factor PICKLE interacts with CONSTANS to promote flowering in Arabidopsis. Plant Cell Environ. 2019, 42, 2495–2507. [Google Scholar] [CrossRef]
- Jing, Y.; Guo, Q.; Lin, R. The Chromatin-Remodeling Factor PICKLE Antagonizes Polycomb Repression of FT to Promote Flowering. Plant Physiol. 2019, 181, 656–668. [Google Scholar] [CrossRef]
- Sang, Q.; Pajoro, A.; Sun, H.; Song, B.; Yang, X.; Stolze, S.C.; Andrés, F.; Schneeberger, K.; Nakagami, H.; Coupland, G. Mutagenesis of a Quintuple Mutant Impaired in Environmental Responses Reveals Roles for CHROMATIN REMODELING4 in the Arabidopsis Floral Transition. Plant Cell 2020, 32, 1479–1500. [Google Scholar] [CrossRef] [Green Version]
- Ma, X.; Ma, J.; Zhai, H.; Xin, P.; Chu, J.; Qiao, Y.; Han, L. CHR729 Is a CHD3 Protein That Controls Seedling Development in Rice. PLoS ONE 2015, 10, e0138934. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Wang, D.; Gan, T.; Liu, L.; Long, W.; Wang, Y.; Niu, M.; Li, X.; Zheng, M.; Jiang, L.; et al. CRL6, a member of the CHD protein family, is required for crown root development in rice. Plant Physiol. Biochem. 2016, 105, 185–194. [Google Scholar] [CrossRef]
- Xu, J.; Wang, L.; Zhou, M.; Zeng, D.; Hu, J.; Zhu, L.; Ren, D.; Dong, G.; Gao, Z.; Guo, L.; et al. Narrow albino leaf 1 is allelic to CHR729, regulates leaf morphogenesis and development by affecting auxin metabolism in rice. Plant Growth Regul. 2017, 82, 175–186. [Google Scholar] [CrossRef]
- Hu, Y.; Liu, D.; Zhong, X.; Zhang, C.; Zhang, Q.; Zhou, D.X. CHD3 protein recognizes and regulates methylated histone H3 lysines 4 and 27 over a subset of targets in the rice genome. Proc. Natl. Acad. Sci. USA 2012, 109, 5773–5778. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cho, S.H.; Lee, C.H.; Gi, E.; Yim, Y.; Koh, H.J.; Kang, K.; Paek, N.C. The Rice Rolled Fine Striped (RFS) CHD3/Mi-2 Chromatin Remodeling Factor Epigenetically Regulates Genes Involved in Oxidative Stress Responses During Leaf Development. Front. Plant Sci. 2018, 9, 364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Itoh, J.I.; Hasegawa, A.; Kitano, H.; Nagato, Y.A. recessive heterochronic mutation, plastochron1, shortens the plastochron and elongates the vegetative phase in rice. Plant Cell 1998, 10, 1511–1522. [Google Scholar] [PubMed] [Green Version]
- Komiya, R.; Yokoi, S.; Shimamoto, K. A gene network for long-day flowering activates RFT1 encoding a mobile flowering signal in rice. Development 2009, 136, 3443–3450. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ryu, C.H.; Lee, S.; Cho, L.H.; Kim, S.L.; Lee, Y.S.; Choi, S.C.; Jeong, H.J.; Yi, J.; Park, S.J.; Han, C.D.; et al. OsMADS50 and OsMADS56 function antagonistically in regulating long day (LD)-dependent flowering in rice. Plant Cell Environ. 2009, 32, 1412–1427. [Google Scholar] [CrossRef]
- Lee, Y.S.; An, G. OsGI controls flowering time by modulating rhythmic flowering time regulators preferentially under short day in rice. J. Plant Biol. 2015, 58, 137–145. [Google Scholar] [CrossRef]
- Han, S.H.; Yoo, S.C.; Lee, B.D.; An, G.; Paek, N.C. Rice FLAVIN-BINDING, KELCH REPEAT, F-BOX 1 (OsFKF1) promotes flowering independent of photoperiod. Plant Cell Environ. 2015, 38, 2527–2540. [Google Scholar] [CrossRef]
- Pan, Y.H.; Gao, L.J.; Liang, Y.T.; Zhao, Y.; Liang, H.F.; Chen, W.W.; Yang, X.H.; Qing, D.J.; Gao, J.; Wu, H.; et al. OrMKK3 Influences Morphology and Grain Size in Rice. J. Plant Biol. 2021, 4, 1–14. [Google Scholar]
- Zhang, H.; Rider, S.D.; Henderson, J.T., Jr.; Fountain, M.; Chuang, K.; Kandachar, V.; Simons, A.; Edenberg, H.J.; Romero-Severson, J.; Muir, W.M.; et al. The CHD3 remodeler PICKLE promotes trimethylation of histone H3 lysine 27. J. Biol. Chem. 2008, 283, 22637–22648. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Bishop, B.; Ringenberg, W.; Muir, W.M.; Ogas, J. The CHD3 remodeler PICKLE associates with genes enriched for trimethylation of histone H3 lysine 27. Plant Physiol. 2012, 159, 418–432. [Google Scholar] [CrossRef] [Green Version]
- Jing, Y.; Zhang, D.; Wang, X.; Tang, W.; Wang, W.; Huai, J.; Xu, G.; Chen, D.; Li, Y.; Lin, R. Arabidopsis chromatin remodeling factor PICKLE interacts with transcription factor HY5 to regulate hypocotyl cell elongation. Plant Cell 2013, 25, 242–256. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carter, B.; Bishop, B.; Ho, K.K.; Huang, R.; Jia, W.; Zhang, H.; Pascuzzi, P.E.; Deal, R.B.; Ogas, J. The Chromatin Remodelers PKL and PIE1 Act in an Epigenetic Pathway That Determines H3K27me3 Homeostasis in Arabidopsis. Plant Cell 2018, 30, 1337–1352. [Google Scholar] [CrossRef] [PubMed]
- Guo, T.; Wang, D.; Fang, J.; Zhao, J.; Yuan, S.; Xiao, L.; Li, X. Mutations in the Rice OsCHR4 Gene, Encoding a CHD3 Family Chromatin Remodeler, Induce Narrow and Rolled Leaves with Increased Cuticular Wax. Int. J. Mol. Sci. 2019, 20, 2567. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tong, J.K.; Hassig, C.A.; Schnitzler, G.R.; Kingston, R.E.; Schreiber, S.L. Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature 1998, 395, 917–921. [Google Scholar] [CrossRef]
- Xue, Y.; Wong, J.; Moreno, G.T.; Young, M.K.; Côté, J.; Wang, W. NURD, a novel complex with both ATP-dependent chromatin-remodeling and histone deacetylase activities. Mol. Cell 1998, 2, 851–861. [Google Scholar] [CrossRef]
- Yu, Y.; Liang, Z.; Song, X.; Fu, W.; Xu, J.; Lei, Y.; Yuan, L.; Ruan, J.; Chen, C.; Fu, W.; et al. BRAHMA-interacting proteins BRIP1 and BRIP2 are core subunits of Arabidopsis SWI/SNF complexes. Nat. Plants 2020, 6, 996–1007. [Google Scholar] [CrossRef]
- Tan, L.M.; Liu, R.; Gu, B.W.; Zhang, C.J.; Luo, J.; Guo, J.; Wang, Y.; Chen, L.; Du, X.; Li, S.; et al. Dual Recognition of H3K4me3 and DNA by the ISWI Component ARID5 Regulates the Floral Transition in Arabidopsis. Plant Cell 2020, 32, 2178–2195. [Google Scholar] [CrossRef]
- Ho, K.K.; Zhang, H.; Golden, B.L.; Ogas, J. PICKLE is a CHD subfamily II ATP-dependent chromatin remodeling factor. Biochim. Biophys. Acta 2013, 1829, 199–210. [Google Scholar] [CrossRef] [Green Version]
- Smaczniak, C.; Immink, R.G.; Muiño, J.M.; Blanvillain, R.; Busscher, M.; Busscher-Lange, J.; Dinh, Q.D.; Liu, S.; Westphal, A.H.; Boeren, S.; et al. Characterization of MADS-domain transcription factor complexes in Arabidopsis flower development. Proc. Natl. Acad. Sci. USA 2012, 109, 1560–1565. [Google Scholar] [CrossRef] [Green Version]
- Ikeda, K.; Ito, M.; Nagasawa, N.; Kyozuka, J.; Nagato, Y. Rice ABERRANT PANICLE ORGANIZATION 1, encoding an F-box protein, regulates meristem fate. Plant J. 2007, 51, 1030–1040. [Google Scholar] [CrossRef]
- Haring, M.; Offermann, S.; Danker, T.; Horst, I.; Peterhansel, C.; Stam, M. Chromatin immunoprecipitation: Optimization, quantitative analysis and data normalization. Plant Methods 2007, 3, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yoon, H.; Shim, Y.; Yoo, S.-C.; Kang, K.; Paek, N.-C. The Rice CHD3/Mi-2 Chromatin Remodeling Factor Rolled Fine Striped Promotes Flowering Independent of Photoperiod. Int. J. Mol. Sci. 2021, 22, 1303. https://doi.org/10.3390/ijms22031303
Yoon H, Shim Y, Yoo S-C, Kang K, Paek N-C. The Rice CHD3/Mi-2 Chromatin Remodeling Factor Rolled Fine Striped Promotes Flowering Independent of Photoperiod. International Journal of Molecular Sciences. 2021; 22(3):1303. https://doi.org/10.3390/ijms22031303
Chicago/Turabian StyleYoon, Hyeryung, Yejin Shim, Soo-Cheul Yoo, Kiyoon Kang, and Nam-Chon Paek. 2021. "The Rice CHD3/Mi-2 Chromatin Remodeling Factor Rolled Fine Striped Promotes Flowering Independent of Photoperiod" International Journal of Molecular Sciences 22, no. 3: 1303. https://doi.org/10.3390/ijms22031303
APA StyleYoon, H., Shim, Y., Yoo, S. -C., Kang, K., & Paek, N. -C. (2021). The Rice CHD3/Mi-2 Chromatin Remodeling Factor Rolled Fine Striped Promotes Flowering Independent of Photoperiod. International Journal of Molecular Sciences, 22(3), 1303. https://doi.org/10.3390/ijms22031303





