1. Introduction
Species within the
Fusarium genus represent some of the most devastating and important pathogens of many of the world’s agricultural crops. The
Fusarium oxysporum species complex (FOSC) includes isolates that can be pathogens, saprophytes and even endophytes [
1,
2]. The plant pathogenic lineages of
F. oxysporum have been recorded to cause vascular wilt in over 100 species of plants [
3,
4,
5]. However, an individual pathogenic isolate of
F. oxysporum is a specialist and typically causes disease in only one or two host species. As a consequence of this host-specificity, isolates of
F. oxysporum are classified into special groups called ‘
formae speciales’ (plural, ff. spp.; singular–
forma speciaies, f. sp.) according to their specific host [
1]. Many
formae speciales consist of multiple, genetically distinct clonal lineages or vegetatively compatible groups (VCGs) [
3,
6,
7].
Fusarium wilt of banana is one of the most notorious examples of disease caused by
F. oxysporum and is also known as Panama disease of banana [
8,
9]. The isolates that cause Panama disease in banana have historically been referred to as
F. oxysporum f. sp.
cubense (
Foc). However, recent taxonomic revisions within the FOSC have divided isolates of
Foc into eight different species [
10,
11]. The relationship between these newly described species and other methods of describing and classifying isolates of
Foc have been summarised in
Table 1. For clarity of communication and consistency, the synonym
Foc shall be used throughout this study to indicate all isolates and species within the FOSC that are pathogenic to bananas. This convention for the naming of isolates
F. oxysporum shall also be extended to the other
formae speciales that have also been recently epitypified [
11]. Isolates within
Foc are further classified into cultivar-specific races and VCGs (
Table 1) [
3,
6,
7,
12,
13]. Presently, tropical race 4 (TR4; VCG 01213/16) is of significant concern as it is highly pathogenic to all commercial cultivars and is rapidly spreading throughout the world’s banana producing regions [
14,
15,
16,
17,
18]. Phylogenetic analyses of the VCGs that comprise
Foc have demonstrated that isolates within this
forma specialis do not share a recent common ancestor and are commonly more closely related to isolates of
F. oxysporum that are non-pathogenic or pathogenic to other host species [
4,
5,
19,
20,
21]. It is hypothesised that the polyphyletic distribution of
Foc in the FOSC is due to either convergent evolution and/or the horizontal transmission of the genes conferring host-specific pathogenicity. One such class of pathogenicity genes includes effectors.
Effector genes encode for small proteins that are secreted by plant pathogens, including
F. oxysporum, during the infection of a host [
23]. These proteins are often critical for infection as they manipulate the defence response of a host. In
F. oxysporum, a family of effector genes termed secreted in xylem (
SIX) genes have been identified [
24]. Currently, 14
SIX genes have been experimentally identified in the tomato pathotype,
F. oxysporum f. sp.
lycopersici (
Fol) [
24,
25,
26,
27,
28]. Subsequent to their discovery,
SIX3,
SIX4,
SIX5 and
SIX6 have been experimentally verified to be required for full virulence of
F. oxysporum in its respective host [
24,
29,
30,
31,
32,
33]. Furthermore, studies in the tomato and cucurbits have demonstrated that the small, dispensable accessory chromosomes to which the
SIX genes are localised can be horizontally transferred from pathogenic lineages of
F. oxysporum to a genetically distinct, non-pathogenic isolate of
F. oxysporum [
33,
34]. In both the tomato and cucurbit pathosystems, the transfer of accessory chromosomes from a pathogenic donor into a non-pathogenic recipient was accompanied by a gain of pathogenicity towards the respective host. These experiments have been important for demonstrating the significant impact these accessory chromosomes and associated pathogenicity genes have on the phenotype of their host strain.
Although 14
SIX genes have been described in the
Fol lineages, not all 14
SIX genes are carried by other
formae speciales [
22,
30,
35,
36,
37,
38]. In fact, van Dam et al. demonstrated that the
formae speciales of
F. oxysporum each harbour a unique combination of effectors. These effector profiles can be used to distinguish the different host-specific lineages [
39]. Furthermore, variation in the
SIX genes has been particularly useful for distinguishing different races and clonal lineages that exist within a
forma specialis [
22,
26,
37,
39,
40]. The unique combination of effectors has been proposed to confer the host-specific pathogenicity exhibited by plant pathogens, such as
F. oxysporum. Several
SIX genes, including
SIX2 and
SIX6, have been previously reported in other species of
Fusarium, including
F. verticillioides,
F. proliferatum,
F. foetens,
F. hostae and
F. agapanthi [
36,
41,
42]. Homologues of
SIX1 and
SIX6 have also been described outside the Fusarium genus in
Colletotrichum orbiculare and
C. higginsianum, as well as
SIX1 in
Leptosphaeria maculans [
43,
44,
45].
In
Foc, homologues of
SIX1 and
SIX9 have been identified in all isolates of
Foc, while
SIX8 was only identified in the VCGs of race 4 of
Foc [
22,
35,
46]. The other
SIX genes identified in
Foc include
SIX2,
SIX6,
SIX7,
SIX10,
SIX13 and a pseudogeneised homologue of
SIX4 [
22,
35,
46]. Specific homologues of the
SIX genes were associated with the different races of
Foc. Furthermore, Czislowski et al. demonstrated evidence of horizontal transfer of the
SIX genes between the lineages of
Foc [
22]. Subsequently, it was hypothesised that the polyphyletic relationship of the
Foc lineages can be attributed to the horizontal transference of effectors and pathogenicity genes, including the
SIX genes [
22]. Although the distribution and diversity of the
SIX genes in the pathogenic lineages of
Foc has been studied, less is known about the
SIX genes in non-pathogenic lineages of
F. oxysporum associated with banana and other healthy hosts.
There has been a strong emphasis on understanding the distribution and diversity of effectors in the plant pathogenic forms of
F. oxysporum. Species of
Fusarium are commonly isolated from the tissue of healthy, asymptomatic host plants, including banana [
47,
48,
49,
50,
51]. For the purposes of this study, endophytically-associated non-pathogenic isolates are considered those that have been recovered from healthy, asymptotic host material. These isolates are to be considered non-pathogenic towards the host that they were recovered from—however, this should not automatically preclude them from possibly being pathogens of other hosts. The presence of effectors in non-pathogenic lineages of
F. oxysporum is not as well understood. This is in part due to the difficulty in classifying an isolate of
F. oxysporum as truly non-pathogenic. Although isolates of
F. oxysporum are common in soil environments and in healthy, asymptomatic hosts, the classification of these isolates as truly non-pathogenic is practically impossible, due to the number of hosts that must be screened in pathogenicity assays [
2]. The use of molecular methods to classify pathogenic lineages of
F. oxysporum is further complicated by the polyphyletic nature of many of the
formae speciales [
5]. Therefore, the development of a molecular method that is capable of distinguishing pathogenic and non-pathogenic lineages of
F. oxysporum is highly desirable. The majority of studies investigating the utility of the
SIX genes for identifying and distinguishing lineages of
F. oxysporum have predominantly focused on plant pathogenic
formae speciales. Few studies have sought to better understand the frequency of
SIX genes in isolates of
F. oxysporum recovered from the soil of different ecological environments.
Isolates of
F. oxysporum recovered from the rhizosphere or asymptomatic hosts have been shown to have a reduced number of effectors, including
SIX genes. Rocha et al. identified
SIX gene homologues in only 12% of
F. oxysporum isolate recovered from the soil of native environments [
52]. Inami et al. also screened isolates of
F. oxysporum recovered from the soil or tissue of wild tomato species for the presence of
SIX1,
SIX3 and
SIX4. However, they was unable to detect any of these genes in any of the isolates [
49]. Jelinski et al. found that the
SIX genes were more frequently identified in isolates of
F. oxysporum recovered from the soil of a tomato field known to be affected by
Fol [
53]. The isolates were screened for
SIX1–SIX7 and half of the isolates had the same
SIX gene profile as
Fol, while 3% of the isolates carried a combination of
SIX genes that was not typical of
Fol. Interestingly, one isolate that harboured the same
SIX genes as
Fol was non-pathogenic towards susceptible tomato in a subsequent pot trial. There are limited studies that have investigated whether
SIX genes are present in endophytic, non-pathogenic isolates of
F. oxysporum recovered from the internal tissue of healthy, asymptomatic hosts. It is also unclear if other
Fusarium species associated with the same ecological niche as
F. oxysporum also harbour
SIX genes. Furthermore, although
Fusarium species are commonly recovered from the tissue of banana hosts, it is unclear how frequently
SIX genes are associated with these isolates and if any of the
SIX genes in the endophytic isolates are similar to those found in the pathogenic lineages of
Foc.
This study sought to ascertain if isolates of Fusarium associated with healthy, asymptomatic banana plants also carried any SIX gene homologues, and if so, was there any similarities to the SIX genes known to occur in pathogenic lineages of Foc. Similar to what has been observed for non-pathogenic lineages of the FOSC other Fusarium species, it was hypothesised that the SIX genes would be infrequently associated with isolates of endophytic, non-pathogenic Fusarium from the tissue of asymptomatic banana hosts. It was hypothesised that of the non-pathogenic, endophytic isolates identified to carry SIX genes, their SIX gene profile would be distinct from the SIX gene profiles of the pathogenic lineages of Foc.
3. Discussion
Plant pathogenic lineages of
F. oxysporum, such as the banana pathogen
Foc, represent a small fraction of the genetic diversity of the FOSC; many isolates exist in the rhizosphere as saprophytes whilst others can establish an endophytic, asymptomatic relationship with a host plant [
2]. The distribution, diversity and evolution of effectors in the pathogenic lineages within the FOSC has been well-studied. However, the presence of effectors, such as the
SIX genes, in putatively endophytic
F. oxysporum is poorly understood.
Initially, this study hypothesised that isolates of
F. oxysporum recovered from healthy, asymptomatic banana plants would carry no or very few
SIX genes. A major finding of this study was the high frequency of
SIX genes in endophytic isolates of
Fusarium associated with asymptomatic banana hosts. Similarly, Jelsinki et al. had also reported the occurrence of
SIX gene homologues in a high proportion of putatively endophytic isolates of
F. oxysporum recovered from tomato producing fields [
53]. These results are contradictory to Rocha et al. and Inami et al., which both reported a low or no incidence of
SIX genes in putatively endophytic or saprophytic isolates of
F. oxysporum [
49,
52]. The disparity between the results of these studies could be due to the land use and the environment from which the endophytes have been recovered. Both this study and Jelsinki et al. targeted endophytes in hosts being grown for agricultural purposes. In contrast, Rocha et al. and Inami et al. recovered putatively endophytic or saprophytic isolates of
Fusarium from natural ecosystems or wild, uncultivated hosts. In intensive agricultural systems, effectors (such as the
SIX proteins) could be important to the endophytic, as well as the pathogenic isolates of
Fusarium as they may facilitate the growth and survival of the fungus as an asymptomatic biotroph. For endophytic
Fusarium, the ability to better colonise and survive within their host may give the isolates a significant advantage of other fungi that are present in the same niche. Therefore, it is hypothesised that the high frequency of
SIX genes in the endophytes of
Fusarium may be the result of positive selection for endophytic isolates whose genomes include features, such as effectors that enable them to be better competitors in the ecological niche of a host plant, particularly in intensive agricultural systems.
There is increasing evidence that many mycorrhizal and endophytic species of fungi utilise effectors to successfully establish their mutualistic relationships with their hosts (reviewed by References [
56,
57]). The functions of three effectors in three mycorrhizal species have been well-studied with interesting results. The SP7 effector in the mutualist,
Rhizophagus irregularis, and the MiSSP7 effector in
Laccaria bicolour have both been shown to modulate plant hormone signalling pathways, thereby downregulating the host defence responses [
58,
59,
60,
61,
62]. The increasing number of sequenced genomes representing mycorrhizal and endophytic species have shown that hundreds of effector candidates are predicted to be encoded by these species [
63,
64,
65]. It is possible that similar to mycorrhizal species of fungi, the endophytic isolates of
F. oxysporum also utilise effectors to suppress a host’s defence responses to facilitate the establishment of an intimate endophytic relationship. However, the studies to date that have investigated the presence of
SIX genes in environmental isolates of
F. oxysporum have not established if the
SIX genes or other effectors are expressed or functional in these non-pathogenic interactions.
An alternative explanation for the increased frequency of
SIX genes identified in the endophytic isolates compared to other studies is due to the primers used in this study. The universal primers developed in this study to amplify all
SIX gene homologues that were known at the time of this could account for the increased frequency of
SIX gene detection in isolates of
Fusarium. The other studies, such as Inami et al., Rocha et al. and Jelinski et al., utilised primers that were commonly designed to amplify
SIX genes from specific
formae speciales [
49,
52,
53]. While the use of PCR worked well for our purposes, it is an approach that is biased towards identifying known orthologues of previously identified
SIX genes. The most unbiased approach for effector discovery and analysis would be whole genome sequencing. Currently, the majority of genomes of
F. oxysporum that have been sequenced represent pathogenic lineages, and only two genomes of putatively non-pathogenic isolates of
F. oxysporum are publicly available. The two sequenced genomes of non-pathogenic
F. oxysporum, Fo47 and MN14, both show a reduced accessory genome [
38,
39]. These isolates were also shown by van Dam and Rep to have reduced numbers of a transposable element class termed mimps, which are associated with the promoter region of the
SIX genes and other predicted effector candidates of
F. oxysporum [
28,
42]. Interestingly, the genomes of
Foc also shared similar hallmarks similar to the non-pathogenic isolates, including a small accessory genome and reduced number of identified mimps [
38,
39,
42].
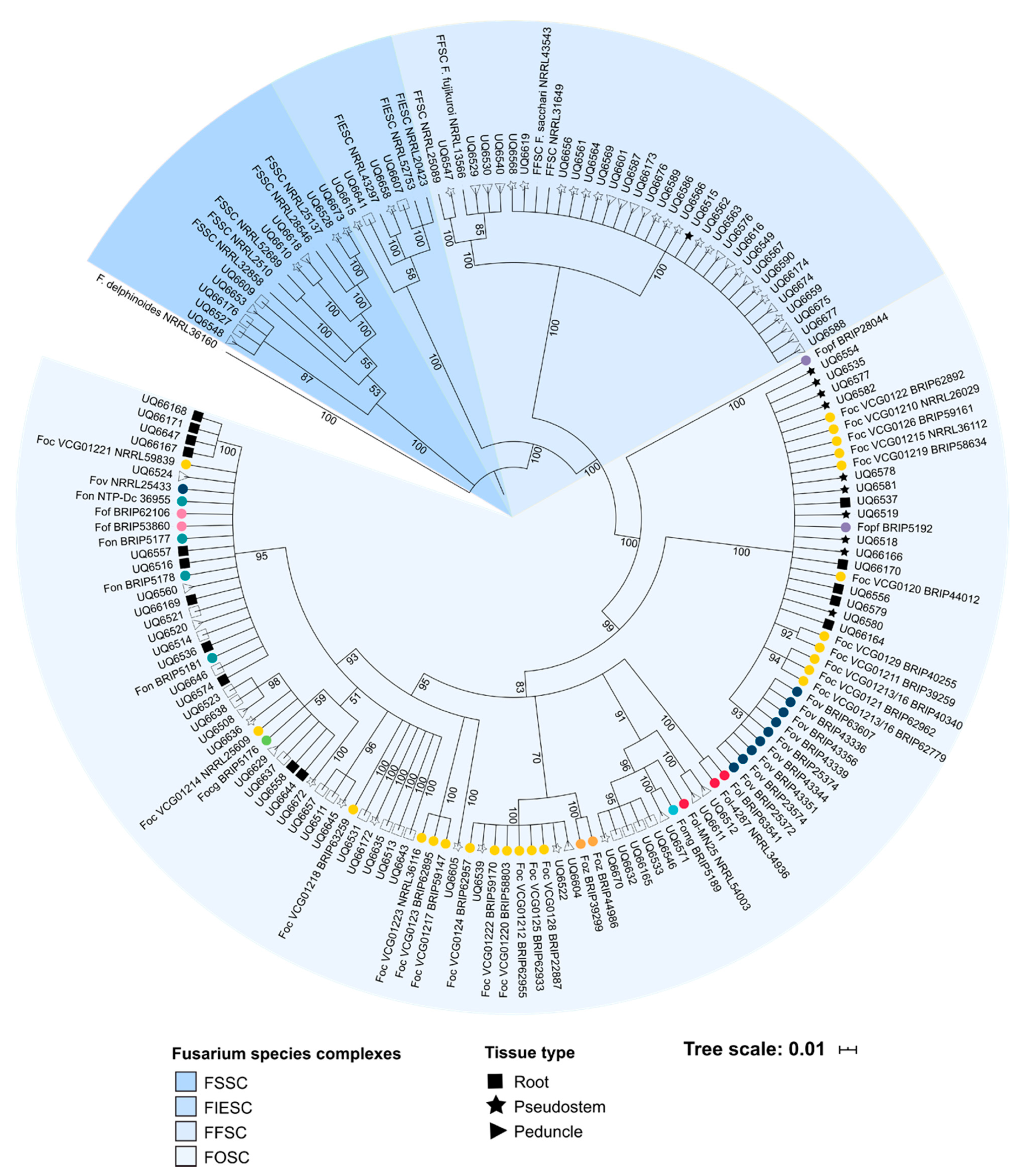
The cluster analysis of this study was also important for demonstrating that the SIX gene profiles of the putatively endophytic isolates were not the same as the SIX gene profiles of pathogenic formae speciales of F. oxysporum that were assessed in this study. Following the identification of the SIX genes in putatively endophytic isolates of Fusarium, it was hypothesised that endophytes from healthy, asymptomatic bananas should not have the same SIX gene profile as the pathogenic lineages of Foc. A hierarchical cluster analysis supported this hypothesis, and based on their SIX gene profiles, the majority of the endophytic isolates clustered separately from the pathogenic lineages of F. oxysporum. Several isolates of F. oxysporum recovered from Mullumbimby shared the same SIX gene profile as the race 1 VCGs of Foc, and some of the isolates recovered from Redlands clustered with the SR4 VCGs of Foc. These isolates also had an identical EF1-α sequence to the respective pathogenic VCGs of Foc. In both of these instances, the plants from which these isolates were recovered displayed both external and internal symptoms of Fusarium wilt at the time of sampling. Based on the SIX gene profiles and sequencing of the EF1-α locus, the race of Foc that was causing disease at both sites was identified, thus, demonstrating the utility of the SIX gene presence/absence typing.
The diagnostic potential of the
SIX genes in
Foc has also been utilised by Carvalhais et al. in the design and development of a molecular assay capable of distinguishing between the various races and vegetative compatibility groups (VCGs) of
Foc [
66]. Typically, molecular diagnostics are developed to be able to rapidly identify the intended target organism and return a binary ‘yes/no’ answer. This approach to molecular diagnostics is favoured for pathogens, such as TR4, due to the ability to design an assay that is sensitive, specific and fast. However, these methods of developing diagnostics have limitations, such as requiring an extensive prior understanding of the target pathogen. As a result, this approach to molecular diagnostics is often not effective and identifying or characterising novel pathotypes or even novel
formae speciales. The future of molecular-based diagnostics of the
F. oxysporum species complex lies in the ability to predict an isolate’s pathogenic potential from its genomic information. The diagnostic utility of the
SIX genes and effectors more broadly has the potential to provide a novel means of pathogen identification and characterisation in the
F. oxysporum species complex. Developing an improved understanding of effector presence/absence and evolution in the genetically diverse range of non-pathogenic and endophytic isolates of
Fusarium will be critical to the future utility of an effector-based multi-locus sequencing typing of
Fusarium.
In conclusion, this study has demonstrated that many of the Fusarium species isolated from healthy, asymptomatic banana tissue carry one or more SIX genes. Furthermore, the SIX gene profiles of isolates that are pathogenic to bananas have a complement of SIX genes that is distinct from their non-pathogenic counterparts. The results of this study could indicate that endophytic isolates of Fusarium, including F. oxysporum, could indicate that putative endophytes acquire and/or retain effectors to better colonise hosts and become more competitive in the ecological niche of a host banana plant. Although there may be a selective advantage for isolates that carry effectors, the endophytes may not be unable to cause disease in banana as they lack the full complement of molecular machinery required for virulence. Currently, the means by which putative endophytes acquire SIX genes is unknown and requires further investigation. While horizontal chromosome transfer has been reported in pathogenic lineages of Foc, it is not known whether SIX genes are also acquired by endophytes via HCT. This study is an important demonstration that endophytic Fusarium could be potential reservoirs of effector genes and could contribute to the evolution of novel pathotypes of F. oxysporum. For this reason, the endophytic and putatively non-pathogenic lineages of F. oxysporum continue to be of significant interest and warrant further research.