Subcellular Localization of Acyl-CoA: Lysophosphatidylethanolamine Acyltransferases (LPEATs) and the Effects of Knocking-Out and Overexpression of Their Genes on Autophagy Markers Level and Life Span of A. thaliana
Abstract
1. Introduction
2. Results
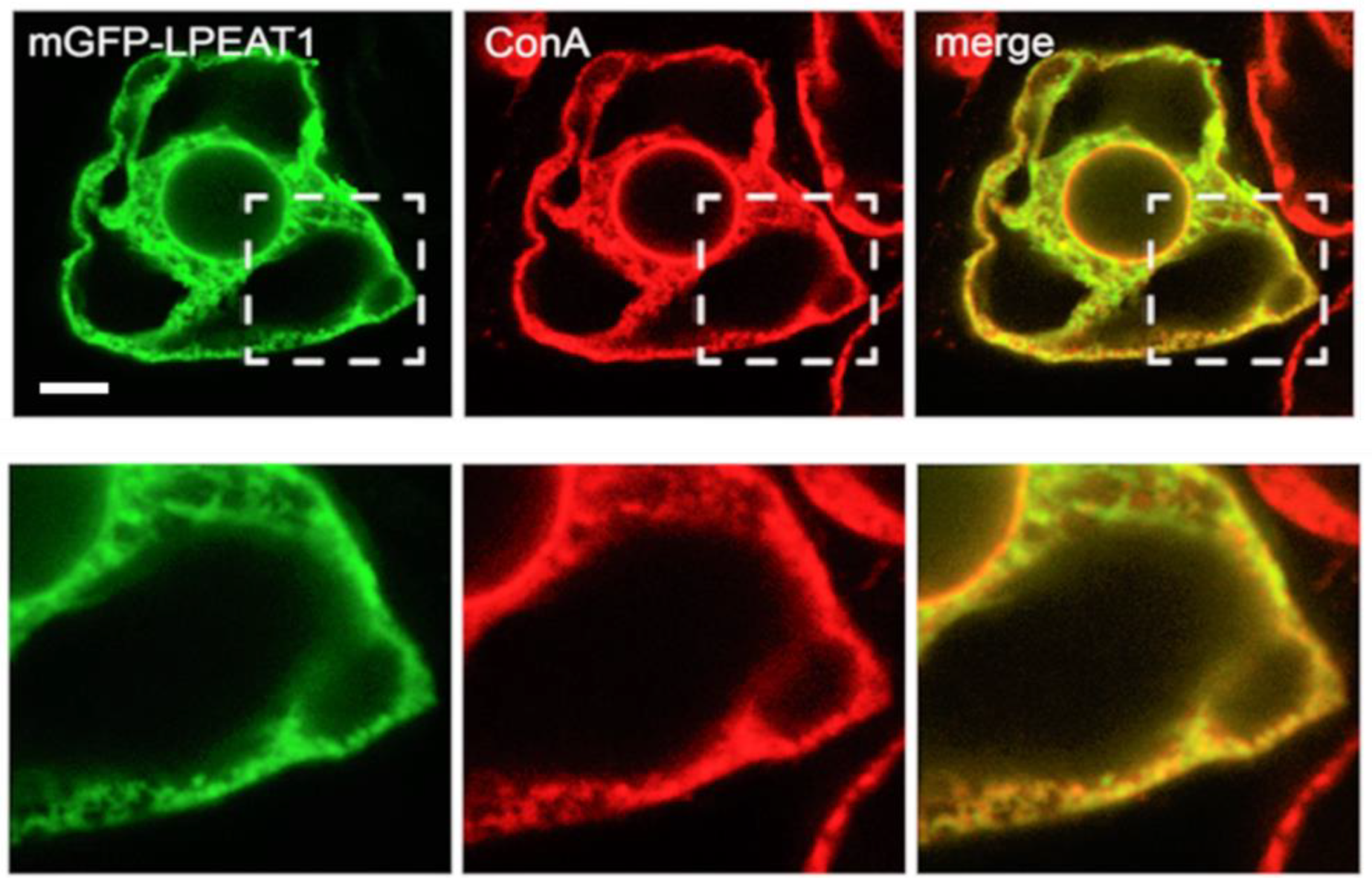
2.1. Subcellular Localisation of Arabidopsis LPEAT1 and LPEAT2
2.2. NBR1 Transcript Levels and NBR1 Protein Abundance in Rosette Leaves of Wild-Type and LPEAT Transgenic Arabidopsis Plants
2.3. ATG8a Transcript Levels and ATG8 Protein Abundance in Rosette Leaves of Wild-Type and LPEAT Transgenic Arabidopsis Plants
2.4. Relative Amount of Different Forms of ATG8 Proteins in Rosette Leaves of Wild Type Arabidopsis Plants and Plants with Knocked-Out or Overexpressed LPEAT
2.5. Ultrastructure of Cells of Wild Type Plants and Lpeat Mutants of A. thaliana
2.6. Lifespan of Wild-Type Arabidopsis Plants and Plants with Knocked-Out or Overexpressed LPEAT
3. Discussion
3.1. Subcellular Localisation of LPEAT1 and LPEAT2
3.2. Autophagy Markers Levels
3.3. Ultrastructure of Mesophyll Cells
3.4. Life Span of Tested Arabidopsis Lines
3.5. Conclusions
4. Methods
4.1. Plant Materials and Growth Conditions
4.2. Subcellular Localisation of LPEAT1 and LPEAT2 in BY-2 Cells
4.3. Determination of NBR1 and ATG8 Protein Levels in Leaves of A. thaliana
4.4. Measuring ATG8a and NBR1 Expression Levels in Leaves of A. thaliana by q-RT-PCR
4.5. Ultrastructural Analysis of Cells of Leaves of Wild Type Plants and Lpeat Mutants of A. thaliana
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ståhl, U.; Stalberg, K.; Stymne, S.; Ronne, H. A family of eukaryotic lysophospholipid acyltransferases with broad specificity. FEBS Lett. 2008, 582, 305–309. [Google Scholar] [CrossRef]
- Stalberg, K.; Ståhl, U.; Stymne, S.; Ohlrogge, J. Characterisation of two Arabidopsis thaliana acyltransferases with prefer-ence for lysophosphatidylethanolamine. BMC Plant Biol. 2009, 9, 1–8. [Google Scholar] [CrossRef]
- Lands, W.E. Lipid metabolism. Annu. Rev. Biochem. 1965, 34, 313–346. [Google Scholar] [CrossRef]
- Jasieniecka-Gazarkiewicz, K.; Demski, K.; Lager, I.; Stymne, S.; Banaś, A. Possible role of different yeast and plant lysophospholipid:acyl-CoA acyltransferases (LPLAT) in acyl remodeling of phospholipids. LIPIDS 2016, 51, 15–32. [Google Scholar] [CrossRef]
- Jasieniecka-Gazarkiewicz, K.; Lager, I.; Carlsson, A.S.; Gutbrod, K.; Peisker, H.; Dörman, P.; Stymne, S.; Banaś, A. Acyl-CoA:lysophosphosphatidylethanolamine acyltransferase activity regulates growth of Arabidopsis. Plant Physiol. 2017, 174, 986–998. [Google Scholar] [CrossRef]
- Wang, L.; Shen, W.; Kazachkov, M.; Chen, Q.; Carlsson, A.S.; Stymne, S.; Weselake, R.J.; Zou, J. Metabolic interactions be-tween the Lands cycle and the Kennedy pathway of glycerolipid synthesis in Arabidopsis developing seeds. Plant Cell 2012, 24, 4652–4669. [Google Scholar] [CrossRef]
- Christie, W.W.; Han, X. Chapter 1—Lipids: Their structures and occurrence. In Lipid Analysis, 4th ed.; Christie, W.W., Han, X., Eds.; Woodhead Publishing: Sawston, UK, 2012; pp. 3–19. [Google Scholar]
- Kennedy, E.P.; Weiss, S.B. The function of cytidine coenzymes in the biosynthesis of phospholipides. J. Biol. Chem. 1956, 222, 193–214. [Google Scholar] [CrossRef]
- Klińska, S.; Jasieniecka-Gazarkiewicz, K.; Demski, K.; Banaś, A. Editing of phosphatidic acid and phosphatidylethanolamine by acyl-CoA: Lysophospholipid acyltransferases in developing Camelina sativa seeds. Planta 2020, 252, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Rockenfeller, P.; Koska, M.; Pietrocola, F.; Minois, N.; Knittelfelder, O.; Sica, V.; Frantz, J.; Carmona-Gutierrez, D.; Kroemer, G.; Madeo, F. Phosphatidylethanolamine positively regulates authophagy and longevity. Cell Death Differ. 2015, 23, 499–508. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Bassham, D.C. Autophagy: Pathways for Self-Eating in Plant Cells. Annu. Rev. Plant Biol. 2012, 63, 215–237. [Google Scholar] [CrossRef] [PubMed]
- Minina, E.A.; Sanchez-Vera, V.; Moschou, P.N.; Suárez, M.F.; Sundberg, E.; Weih, M.; Bozhkov, P.V. Autophagy mediates caloric restriction-induced lifespan extension inArabidopsis. Aging Cell 2013, 12, 327–329. [Google Scholar] [CrossRef]
- Brandizzi, F.; Irons, S.; Kearns, A.; Hawes, C. BY-2 Cells: Culture and Transformation for Live Cell Imaging. Curr. Protoc. Cell Biol. 2003, 19, 1–17. [Google Scholar] [CrossRef]
- Sparkes, I.A.; Runions, J.; Kearns, A.; Hawes, C. Rapid, transient expression of fluorescent fusion proteins in tabacco plants and generation of stably transformed plants. Nat. Protoc. 2006, 1, 2019–2025. [Google Scholar] [CrossRef]
- Miao, Y.; Jiang, L. Transient expression of fluorescent fusion proteins in protoplasts of suspension cultured cells. Nat. Protoc. 2007, 2, 2348–2353. [Google Scholar] [CrossRef]
- Tartakoff, A.M.; Vassalli, P. Lectin-binding sites as markers of Golgi subcompartments: Proximal-to-distal maturation of oligosaccharides. J. Cell Biol. 1983, 97, 1243–1248. [Google Scholar] [CrossRef]
- Dunkley, T.P.J.; Hester, S.; Shadforth, I.P.; Runions, J.; Weimar, T.; Hanton, S.L.; Griffin, J.L.; Bessant, C.; Brandizzi, F.; Hawes, C.; et al. Mapping the Arabidopsis organelle proteome. Proc. Natl. Acad. Sci. USA 2006, 103, 6518–6523. [Google Scholar] [CrossRef] [PubMed]
- Nikolovski, N.; Rubtsov, D.; Segura, M.P.; Miles, G.P.; Stevens, T.J.; Dunkley, T.P.; Munro, S.; Lilley, K.S.; DuPree, P. Putative Glycosyltransferases and Other Plant Golgi Apparatus Proteins Are Revealed by LOPIT Proteomics. Plant Physiol. 2012, 160, 1037–1051. [Google Scholar] [CrossRef]
- SUBA. 2017. Available online: http://suba.live/ (accessed on 22 October 2020).
- Uemura, T.; Ueda, T.; Ohniwa, R.L.; Nakano, A.; Takeyasu, K.; Sato, M.H. Systematic analysis of SNARE molecules in Arabidopsis: Dissection of the post-Golgi network in plant cells. Cell Struct. Funct. 2004, 29, 49–65. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.J.; Sohn, E.J.; Lee, M.H.; Hwang, I. The Arabidopsis Rab5 homologs Rha1 and Ara7 localize to the prevacuolar compartment. Plant Cell Physiol. 2004, 45, 1211–1220. [Google Scholar] [CrossRef]
- Ueda, T.; Yamaguch, M.; Uchimiya, H.; Nakano, A. Ara6, a plant-unique novel type RabGTPase, functions in the endocytic pathway of Arabidopsis thaliana. EMBO J. 2001, 20, 4730–4741. [Google Scholar] [CrossRef] [PubMed]
- Scheuring, D.; Viotti, C.; Krüger, F.; Künzl, F.; Sturm, S.; Bubeck, J.; Hillmer, S.; Frigerio, L.; Robinson, D.G.; Pimpl, P.; et al. Multivesicular Bodies Mature from the Trans-Golgi Network/Early Endosome in Arabidopsis. Plant Cell 2011, 23, 3463–3481. [Google Scholar] [CrossRef]
- Zhuang, X.; Chung, K.P.; Luo, M.; Jiang, L. Autophagosome Biogenesis and the Endoplasmic Reticulum: A Plant Perspective. Trends Plant Sci. 2018, 23, 677–692. [Google Scholar] [CrossRef]
- Cui, Y.; He, Y.; Cao, W.; Gao, J.; Jiang, L. The Multivesicular Body and Autophagosome Pathways in Plants. Front. Plant Sci. 2018, 9, 1837. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, K.; Jikumaru, Y.; Kamiya, Y.; Kusano, M.; Consonni, C.; Panstruga, R.; Ohsumi, Y.; Shirasu, K. Autophagy nega-tively regulates cell death by controlling NPR1-dependent salicylic acid signaling during senescence and the innate immune response in Arabidopsis. Plant Cell 2009, 21, 2914–2927. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Zhuang, X. Using Microscopy Tools to Visualize Autophagosomal Structures in Plant Cells. In Advanced Structural Safety Studies; Humana Press: New York, NY, USA, 2017; Volume 1662, pp. 257–266. [Google Scholar]
- Suzuki, K.; Kirisako, T.; Kamada, Y.; Mizushima, N.; Noda, T.; Ohsumi, Y. The pre-autophagosomal structure organized by concerted functions of APG genes is essential for autophagosome formation. EMBO J. 2001, 20, 5971–5981. [Google Scholar] [CrossRef]
- Mizushima, N.; Yamamoto, A.; Matsui, M.; Yoshimori, T.; Ohsumi, Y. In Vivo Analysis of Autophagy in Response to Nutrient Starvation Using Transgenic Mice Expressing a Fluorescent Autophagosome Marker. Mol. Biol. Cell 2004, 15, 1101–1111. [Google Scholar] [CrossRef] [PubMed]
- Thompson, A.R.; Doelling, J.H.; Suttangkakul, A.; Vierstra, R.D. Autophagic Nutrient Recycling in Arabidopsis Directed by the ATG8 and ATG12 Conjugation Pathways. Plant Physiol. 2005, 138, 2097–2110. [Google Scholar] [CrossRef] [PubMed]
- Doelling, J.H.; Walker, J.M.; Friedman, E.M.; Thompson, A.R.; Viestra, R.D. The APG8/ 12-activating enzyme APG7 is required for proper nutrient recycling and senescence in Arabidopsis thaliana. J. Biol. Chem. 2002, 277, 33105–33114. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Zhang, P.; Zhu, R.; Fu, J.; Su, J.; Zheng, J.; Wang, Z.; Wang, D.; Gong, Q. Autophagy Is Rapidly Induced by Salt Stress and Is Required for Salt Tolerance in Arabidopsis. Front. Plant Sci. 2017, 8, 1459. [Google Scholar] [CrossRef] [PubMed]
- Minina, E.A.; Moschou, P.N.; Vetukuri, R.R.; Sanchez-Vera, V.; Cardoso, C.; Liu, Q.; Elander, P.H.; Dalman, K.; Beganovic, M.; Yilmaz, J.L.; et al. Transcriptional stimulation of rate-limiting components of the autophagic pathway improves plant fitness. J. Exp. Bot. 2018, 69, 1415–1432. [Google Scholar] [CrossRef]
- Nakatogawa, H.; Ichimura, Y.; Ohsumi, Y. Atg8, a Ubiquitin-like Protein Required for Autophagosome Formation, Mediates Membrane Tethering and Hemifusion. Cell 2007, 130, 165–178. [Google Scholar] [CrossRef]
- Nath, S.; Dancourt, J.; Shteyn, V.; Puente, G.; Fong, W.M.; Nag, S.; Bewersdorf, J.; Yamamoto, A.; Antony, B.; Melia, T.J. Lipidation of LC3/GABARAP family of autophagy proteins relies on a membrane-curvature-sensing domain in Atg3. Nat. Cell Biol. 2014, 16, 415–424. [Google Scholar] [CrossRef]
- Le Bars, R.; Marion, J.; Le Borgne, R.; Satiat-Jeunemaitre, B.; Bianchi, M.W. ATG5 defines a phagophore domain connected to the endoplasmic reticulum during autophagosome formation in plants. Nat. Commun. 2014, 5, 1–10. [Google Scholar] [CrossRef]
- Johansen, T.; Lamark, T. Selective autophagy mediated by autophagic adapter proteins. Autophagy 2011, 7, 279–296. [Google Scholar] [CrossRef]
- Hanaoka, H.; Noda, T.; Shirano, Y.; Kato, Y.; Hayashi, H.; Shibata, D.; Tabata, S.; Ohsumi, Y. Leaf senescence and starvation-induced chlorosis are accelerated by the disruption of an Arabidopsis authophagy gene. Plant Physiol. 2020, 129, 1181–1193. [Google Scholar] [CrossRef]
- Shpilka, T.; Weidberg, H.; Pietrokovski, S.; Elazar, Z. Atg8: An autophagy-related ubiquitin-like protein family. Genome Biol. 2011, 12, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Izumi, M.; Ishida, H.; Nakamura, S.; Hidem, J. Entire Photodamaged Chloroplasts Are Transported to the Central Vacuole by Autophag. Plant Cell 2017, 29, 377–394. [Google Scholar] [CrossRef] [PubMed]
- Izumi, M.; Nakamura, S. Chloroplast Protein Turnover: The Influence of Extraplastidic Processes, Including Autophagy. Int. J. Mol. Sci. 2018, 19, 828. [Google Scholar] [CrossRef]
- Wang, M.; Li, X.; Luo, S.; Fan, B.; Zhu, C.; Chen, Z. Coordination and Crosstalk between Autophagosome and Multivesicular Body Pathways in Plant Stress Responses. Cells 2020, 9, 119. [Google Scholar] [CrossRef] [PubMed]
- Farag, K.M.; Palta, J.P. Improving postharvest keeping quality of vine-ripened tomato fruits with a natural lipid. HortScience 1991, 26, 162. [Google Scholar]
- Fara, K.M.; Palta, J.P. Use of lysophosphatidylethanolamine, a natural lipid, to retard tomato leaf and fruit senescence. Physiol. Plant 1993, 87, 515–521. [Google Scholar] [CrossRef]
- OÖzgen, M.; Park, S.; Palta, J.P. Mitigation of Ethylene-promoted Leaf Senescence by a Natural Lipid, Lysophosphatidylethanolamine. HortScience 2005, 40, 1166–1167. [Google Scholar] [CrossRef]
- Curtis, M.D.; Grossniklaus, U. A Gateway Cloning Vector Set for High-Throughput Functional Analysis of Genes in Planta. Plant Physiol. 2003, 133, 462–469. [Google Scholar] [CrossRef]
- Pyc, M.; Cai, Y.; Gidda, S.K.; Yurchenko, O.; Park, S.; Kretzschmar, F.K.; Ischebeck, T.; Valerius, O.; Braus, G.H.; Chapman, K.D.; et al. Arabidopsis lipid droplet-associated protein (LDAP)—Interacting protein (LDIP) influences lipid droplet size and neutral lipid homeostasis in both leaves and seeds. Plant J. 2017, 92, 1182–1201. [Google Scholar] [CrossRef]
- Deb, S.; Sankaranarayanan, S.; Wewala, G.; Widdup, E.; Samuel, M.A. The S-Domain Receptor Kinase Arabidopsis Receptor Kinase2 and the U Box/Armadillo Repeat-Containing E3 Ubiquitin Ligase9 Module Mediates Lateral Root Development under Phosphate Starvation in Arabidopsis. Plant Physiol. 2014, 165, 1647–1656. [Google Scholar] [CrossRef]
- Fritze, C.E.; Anderson, T.R. Epitope tagging: General method for tracking recombinant proteins. Cellulases 2000, 327, 3–16. [Google Scholar] [CrossRef]
- Gidda, S.K.; Shockey, J.M.; Rothstein, S.J.; Dyer, J.M.; Mullen, R.T. Arabidopsis thaliana GPAT8 and GPAT9 are localized to the ER and possess distinct ER retrieval signals: Functional divergence of the dilysine ER retrieval motif in plant cells. Plant Physiol. Biochem. 2009, 47, 867–879. [Google Scholar] [CrossRef]
- Shockey, J.M.; Gidda, S.K.; Chapital, D.C.; Kuan, J.C.; Dhanoa, P.K.; Bland, J.M.; Rothstein, S.J.; Mullen, R.T.; Dyer, J.M. Tung Tree DGAT1 and DGAT2 have non redundant functions in triacylglycerol biosynthesis and are localized to different sub-domains of the endoplasmic reticulum. Plant Cell 2006, 18, 2294–2313. [Google Scholar] [CrossRef] [PubMed]
- Lingard, M.J.; Gidda, S.K.; Bingham, S.; Rothstein, S.J.; Mullen, R.T.; Trelease, R.N. Arabidopsis PEROXIN11c-e, FISSION1b, and DYNAMIN-RELATED PROTEIN3A cooperate in cell cycle-associated replication of peroxisomes. Plant Cell 2008, 20, 1567–1585. [Google Scholar] [CrossRef] [PubMed]
- Richardson, L.G.L.; Howard, A.S.M.; Khuu, N.; Gidda, S.K.; McCartney, A.; Morphy, B.J.; Mullen, R.T. Protein–Protein Interaction Network and Subcellular Localization of the Arabidopsis Thaliana ESCRT Machinery. Front. Plant Sci. 2011, 2, 20. [Google Scholar] [CrossRef] [PubMed]
- Marty, N.J.; Teresinski, H.J.; Hwang, Y.T.; Clendening, E.A.; Gidda, S.K.; Sliwinska, E.; Zhang, D.; Miernyk, J.A.; Brito, G.C.; Andrews, D.W.; et al. New insights into the targeting of subset of tail-anchored proteins to the outer mi-tochondrial membrane. Front. Plant Sci. 2014, 5, 426. [Google Scholar] [CrossRef]
- Belda-Palazon, B.; Rodriguez, L.; Fernandez, M.A.; Castillo, M.-C.; Anderson, E.M.; Gao, C.; Gonzalez-Guzman, M.; Peirats-Llobet, M.; Zhao, Q.; De Winne, N.; et al. FYVE1/FREE1 Interacts with the PYL4 ABA Receptor and Mediates Its Delivery to the Vacuolar Degradation Pathway. Plant Cell 2016, 28, 2291–2311. [Google Scholar] [CrossRef]
- Teresinski, H.J.; Gidda, S.K.; Nguyen, T.N.D.; Howard, N.J.M.; Porter, B.K.; Grimberg, N.; Smith, M.D.; Andrews, D.W.; Dyer, J.M.; Mullen, R.T. An RK/ST C-Terminal Motif is Required for Targeting of OEP7.2 and a Subset of Other Arabidopsis Tail-Anchored Proteins to the Plastid Outer Envelope Membrane. Plant Cell Physiol. 2018, 60, 516–537. [Google Scholar] [CrossRef] [PubMed]
- Hanamata, S.; Kurusu, T.; Okada, M.; Suda, A.; Kawamura, K.; Tsukada, E.; Kuchitsu, K. In vivo imaging and quantitative monitoring of autophagic flux in tobacco BY-2 cells. Plant Signal. Behav. 2013, 8, e22510. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, X.; Wang, H.; Lam, S.K.; Gao, C.; Wang, X.; Cai, Y.; Jiang, L. A BAR-domain protein SH3P2, which binds to phos-phatidylinositol 3-phosphate and ATG8, regulates autophagosome formation in Arabidopsis. Plant Cell 2013, 25, 4596–4615. [Google Scholar] [CrossRef] [PubMed]
- Klionsky, D.J.; Abdelmohse, K.; Abe, A.; Abedin, M.J.; Abeliovich, H.; Acevedo, A.A.; Adachi, H.; Adams, M.; Adams, P.D.; Deli, K.; et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 2016, 12, 1–222. [Google Scholar] [CrossRef]
- Marion, J.; Le Bars, R.; Besse, L.; Batoko, H.; Satiat-Jeunemaitre, B. Multiscale and multimodal approaches to study autopha-gy in model plants. Cells 2018, 7, 5. [Google Scholar] [CrossRef]
- Gidda, S.K.; Park, S.; Pyc, M.; Yurchenko, O.; Cai, Y.; Wu, P.; Andrews, D.W.; Chapman, K.D.; Dyer, J.M.; Mullen, R.T. Lipid Droplet-Associated Proteins (LDAPs) Are Required for the Dynamic Regulation of Neutral Lipid Compartmentation in Plant Cells. Plant Physiol. 2016, 170, 2052–2071. [Google Scholar] [CrossRef]
- Niedojadło, J.; Górska-Brylass, A. New type of snRNP containing nuclear bodies in plant cells. Biol. Cell 2003, 95, 303–310. [Google Scholar] [CrossRef]










Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jasieniecka-Gazarkiewicz, K.; Demski, K.; Gidda, S.K.; Klińska, S.; Niedojadło, J.; Lager, I.; Carlsson, A.S.; Minina, E.A.; Mullen, R.T.; Bozhkov, P.V.; et al. Subcellular Localization of Acyl-CoA: Lysophosphatidylethanolamine Acyltransferases (LPEATs) and the Effects of Knocking-Out and Overexpression of Their Genes on Autophagy Markers Level and Life Span of A. thaliana. Int. J. Mol. Sci. 2021, 22, 3006. https://doi.org/10.3390/ijms22063006
Jasieniecka-Gazarkiewicz K, Demski K, Gidda SK, Klińska S, Niedojadło J, Lager I, Carlsson AS, Minina EA, Mullen RT, Bozhkov PV, et al. Subcellular Localization of Acyl-CoA: Lysophosphatidylethanolamine Acyltransferases (LPEATs) and the Effects of Knocking-Out and Overexpression of Their Genes on Autophagy Markers Level and Life Span of A. thaliana. International Journal of Molecular Sciences. 2021; 22(6):3006. https://doi.org/10.3390/ijms22063006
Chicago/Turabian StyleJasieniecka-Gazarkiewicz, Katarzyna, Kamil Demski, Satinder K. Gidda, Sylwia Klińska, Janusz Niedojadło, Ida Lager, Anders S. Carlsson, Elena A. Minina, Robert T. Mullen, Peter V. Bozhkov, and et al. 2021. "Subcellular Localization of Acyl-CoA: Lysophosphatidylethanolamine Acyltransferases (LPEATs) and the Effects of Knocking-Out and Overexpression of Their Genes on Autophagy Markers Level and Life Span of A. thaliana" International Journal of Molecular Sciences 22, no. 6: 3006. https://doi.org/10.3390/ijms22063006
APA StyleJasieniecka-Gazarkiewicz, K., Demski, K., Gidda, S. K., Klińska, S., Niedojadło, J., Lager, I., Carlsson, A. S., Minina, E. A., Mullen, R. T., Bozhkov, P. V., Stymne, S., & Banaś, A. (2021). Subcellular Localization of Acyl-CoA: Lysophosphatidylethanolamine Acyltransferases (LPEATs) and the Effects of Knocking-Out and Overexpression of Their Genes on Autophagy Markers Level and Life Span of A. thaliana. International Journal of Molecular Sciences, 22(6), 3006. https://doi.org/10.3390/ijms22063006






