Functional Characterisation of Three Glycine N-Acyltransferase Variants and the Effect on Glycine Conjugation to Benzoyl–CoA
Abstract
:1. Introduction
2. Results and Discussion
2.1. Level of Conservation of the GLYAT Gene
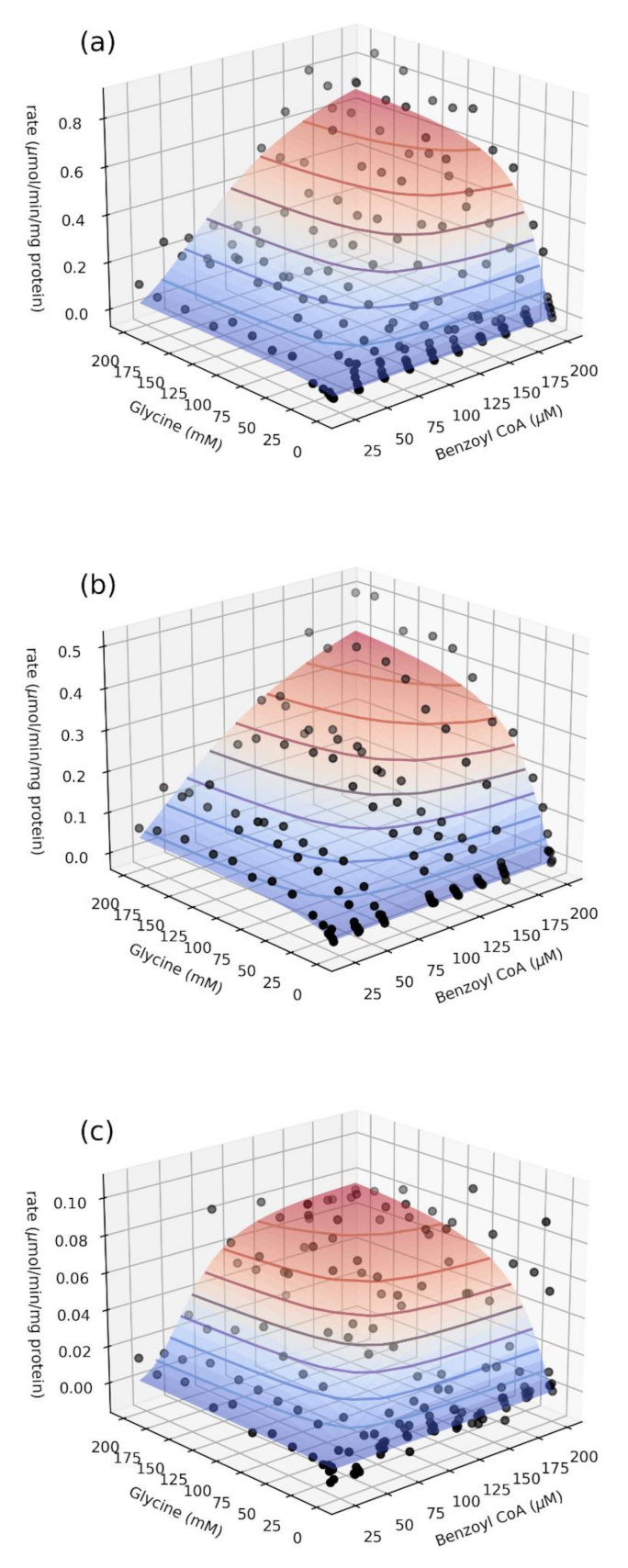
2.2. Relative Enzyme Activity and Catalytic Parameters
2.3. Comparison of GLYAT Kinetic Parameters to Literature Values
3. Materials and Methods
3.1. Missense Variants Identified in GLYAT Using gnomAD
3.2. Haplotype Data Obtained from Ensembl
3.3. Tajima’s Test of Neutrality
3.4. Phylogenetic Analyses
3.5. Expression and Nickel-Affinity Purification of the Recombinant GLYAT Haplotypes
3.6. Bi-Substrate (Benzoyl–CoA and Glycine) Kinetic Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- del Olmo, A.; Calzada, J.; Nunez, M. Benzoic acid and its derivatives as naturally occurring compounds in foods and as additives: Uses, exposure, and controversy. Crit. Rev. Food Sci. Nutr. 2017, 57, 3084–3103. [Google Scholar] [CrossRef] [PubMed]
- van der Sluis, R. Analyses of the genetic diversity and protein expression variation of the acyl: CoA medium-chain ligases, ACSM2A and ACSM2B. Mol. Genet. Genom. MGG 2018, 293, 1279–1292. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.; Isselbacher, K.J. The isolation and identification of N-isovalerylglycine from urine of patients with isovaleric acidemia. J. Biol. Chem. 1967, 242, 2966–2972. [Google Scholar] [CrossRef]
- Bartlett, K.; Gompertz, D. The specificity of glycine-N-acylase and acylglycine excretion in the organicacidaemias. Biochem. Med. 1974, 10, 15–23. [Google Scholar] [CrossRef]
- Killinberg, P.G.; Davidson, E.D.; Webster, L.T. Evidence for a medium-chain fatty acid: Coenzyme A ligase (adenosine monophosphate) that activates salicylate. Mol. Pharmacol. 1971, 7, 260–268. [Google Scholar]
- Knights, K.M. Role of hepatic fatty acid: Coenzyme A ligases in the metabolism of xenobiotic carboxylic acids. Clin. Exp. Pharmacol. Physiol. 1998, 25, 776–782. [Google Scholar] [CrossRef]
- Nandi, D.L.; Lucas, S.V.; Webster, L.T., Jr. Benzoyl-coenzyme A: Glycine N-acyltransferase and phenylacetyl-coenzyme A: Glycine N-acyltransferase from bovine liver mitochondria. Purification and characterization. J. Biol. Chem. 1979, 254, 7230–7237. [Google Scholar] [CrossRef]
- Schachter, D.; Taggart, J.V. Glycine N-acylase: Purification and properties. J. Biol. Chem. 1954, 208, 263–275. [Google Scholar] [CrossRef]
- Rechner, A.R.; Kuhnle, G.; Bremner, P.; Hubbard, G.P.; Moore, K.P.; Rice-Evans, C.A. The metabolic fate of dietary polyphenols in humans. Free Radic. Biol. Med. 2002, 33, 220–235. [Google Scholar] [CrossRef]
- Lemarie, F.; Beauchamp, E.; Legrand, P.; Rioux, V. Revisiting the metabolism and physiological functions of caprylic acid (C8:0) with special focus on ghrelin octanoylation. Biochimie 2016, 120, 40–48. [Google Scholar] [CrossRef]
- Gregus, Z.; Fekete, T.; Varga, F.; Klaassen, C.D. Dependence of glycine conjugation on availability of glycine: Role of the glycine cleavage system. Xenobiotica Fate Foreign Compd. Biol. Syst. 1993, 23, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Gregus, Z.; Fekete, T.; Varga, F.; Klaassen, C.D. Effect of valproic acid on glycine conjugation of benzoic acid. J. Pharmacol. Exp. Ther. 1993, 267, 1068–1075. [Google Scholar] [PubMed]
- Knights, K.M.; Miners, J.O. Amino Acid Conjugation: A Novel Route of Xenobiotic Carboxylic Acid Metabolism in Man, 1st ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2012. [Google Scholar]
- Knights, K.M.; Sykes, M.J.; Miners, J.O. Amino acid conjugation: Contribution to the metabolism and toxicity of xenobiotic carboxylic acids. Expert Opin. Drug Metab. Toxicol. 2007, 3, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Beyoglu, D.; Idle, J.R. The glycine deportation system and its pharmacological consequences. Pharmacol. Ther. 2012, 135, 151–167. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leth, T.; Christensen, T.; Larsen, I.K. Estimated intake of benzoic and sorbic acids in Denmark. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2010, 27, 783–792. [Google Scholar] [CrossRef] [PubMed]
- Mischek, D.; Krapfenbauer-Cermak, C. Exposure assessment of food preservatives (sulphites. benzoic and sorbic acid) in Austria. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2012, 29, 371–382. [Google Scholar] [CrossRef]
- Piper, P.W. Yeast superoxide dismutase mutants reveal a pro-oxidant action of weak organic acid food preservatives. Free Radic. Biol. Med. 1999, 27, 1219–1227. [Google Scholar] [CrossRef]
- Eigenmann, P.A.; Haenggeli, C.A. Food colourings. preservatives, and hyperactivity. Lancet 2007, 370, 1524–1525. [Google Scholar] [CrossRef]
- Piper, J.D.; Piper, P.W. Benzoate and Sorbate Salts: A Systematic Review of the Potential Hazards of These Invaluable Preservatives and the Expanding Spectrum of Clinical Uses for Sodium Benzoate. Compr. Rev. Food Sci. Food Saf. 2017, 16, 868–880. [Google Scholar] [CrossRef] [Green Version]
- Badenhorst, C.P.; van der Sluis, R.; Erasmus, E.; van Dijk, A.A. Glycine conjugation: Importance in metabolism. the role of glycine N-acyltransferase, and factors that influence interindividual variation. Expert Opin. Drug Metab. Toxicol. 2013, 9, 1139–1153. [Google Scholar] [CrossRef] [PubMed]
- Badenhorst, C.P.; Erasmus, E.; van der Sluis, R.; Nortje, C.; van Dijk, A.A. A new perspective on the importance of glycine conjugation in the metabolism of aromatic acids. Drug Metab. Rev. 2014, 46, 343–361. [Google Scholar] [CrossRef]
- Levy, G.; Yaffe, S.J. Clinical implications of salicylate-induced liver damage. Am. J. Dis. Child. 1975, 129, 1385–1386. [Google Scholar] [CrossRef] [PubMed]
- Zimmerman, H.J. Effects of aspirin and acetaminophen on the liver. Arch. Intern. Med. 1981, 141, 333–342. [Google Scholar] [CrossRef] [PubMed]
- Dercksen, M.; Duran, M.; Ijlst, L.; Mienie, L.J.; Reinecke, C.J.; Ruiter, J.P.; Waterham, H.R.; Wanders, R.J. Clinical variability of isovaleric acidemia in a genetically homogeneous population. J. Inherit. Metab. Dis. 2012, 35, 1021–1029. [Google Scholar] [CrossRef] [PubMed]
- Naglak, M.; Salvo, R.; Madsen, K.; Dembure, P.; Elsas, L. The Treatment of Isovaleric Acidemia with Glycine Supplement. Pediatric Res. 1988, 24, 9–13. [Google Scholar] [CrossRef] [Green Version]
- van der Sluis, R.; Badenhorst, C.P.; Erasmus, E.; van Dyk, E.; van der Westhuizen, F.H.; van Dijk, A.A. Conservation of the coding regions of the glycine N-acyltransferase gene further suggests that glycine conjugation is an essential detoxification pathway. Gene 2015, 571, 126–134. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, M.; Terai, K.; Kameda, N.; Matsumoto, A.; Kurokawa, Y.; Funase, Y.; Nishikawa, K.; Sugaya, N.; Hiruta, N.; Kishimoto, T. Designation of enzyme activity of glycine-N-acyltransferase family genes and depression of glycine-N-acyltransferase in human hepatocellular carcinoma. Biochem. Biophys. Res. Commun. 2012, 420, 901–906. [Google Scholar] [CrossRef]
- Wen, H.; Yang, H.J.; An, Y.J.; Kim, J.M.; Lee, D.H.; Jin, X.; Park, S.W.; Min, K.J.; Park, S. Enhanced phase II detoxification contributes to beneficial effects of dietary restriction as revealed by multi-platform metabolomics studies. Mol. Cell. Proteom. MCP 2013, 12, 575–586. [Google Scholar] [CrossRef] [Green Version]
- Mawal, Y.R.; Qureshi, I.A. Purification to homogeneity of mitochondrial acyl coa:glycine N-acyltransferase from human liver. Biochem. Biophys. Res. Commun. 1994, 205, 1373–1379. [Google Scholar] [CrossRef]
- van der Sluis, R.; Badenhorst, C.P.; van der Westhuizen, F.H.; van Dijk, A.A. Characterisation of the influence of genetic variations on the enzyme activity of a recombinant human glycine N-acyltransferase. Gene 2013, 515, 447–453. [Google Scholar] [CrossRef]
- Kelley, M.; Vessey, D. Characterization of the acyl-CoA: Amino acid N-acyltransferases from primate liver mitochondria. J. Biochem. Toxicol. 1994, 9, 153–158. [Google Scholar] [CrossRef] [PubMed]
- van der Westhuizen, F.H.; Pretorius, P.J.; Erasmus, E. The utilization of alanine. glutamic acid, and serine as amino acid substrates for glycine N-acyltransferase. J. Biochem. Mol. Toxicol. 2000, 14, 102–109. [Google Scholar] [CrossRef]
- Schulke, D.; Sass, J.O. Frequent sequence variants of human glycine N-acyltransferase (GLYAT) and inborn errors of metabolism. Biochimie 2021. [Google Scholar] [CrossRef] [PubMed]
- van der Sluis, R.; Ungerer, V.; Nortje, C.; van Dijk, A.A.; Erasmus, E. New insights into the catalytic mechanism of human glycine N-acyltransferase. J. Biochem. Mol. Toxicol. 2017, 31, e21963. [Google Scholar] [CrossRef] [PubMed]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alföldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef]
- Yates, A.; Akanni, W.; Amode, M.R.; Barrell, D.; Billis, K.; Carvalho-Silva, D.; Cummins, C.; Clapham, P.; Fitzgerald, S.; Gil, L.; et al. Ensembl 2016. Nucleic Acids Res. 2016, 44, D710–D716. [Google Scholar] [CrossRef] [PubMed]
- Ramsay, M. Africa: Continent of genome contrasts with implications for biomedical research and health. FEBS Lett. 2012, 586, 2813–2819. [Google Scholar] [CrossRef] [Green Version]
- Altshuler, D.M.; Gibbs, R.A.; Peltonen, L.; Altshuler, D.M.; Gibbs, R.A.; Peltonen, L.; Dermitzakis, E.; Schaffner, S.F.; Yu, F.; Peltonen, L.; et al. Integrating common and rare genetic variation in diverse human populations. Nature 2010, 467, 52–58. [Google Scholar]
- Buchanan, C.C.; Torstenson, E.S.; Bush, W.S.; Ritchie, M.D. A comparison of cataloged variation between International HapMap Consortium and 1000 Genomes Project data. J. Am. Med. Inform. Assoc. JAMIA 2012, 19, 289–294. [Google Scholar] [CrossRef] [Green Version]
- Retshabile, G.; Mlotshwa, B.C.; Williams, L.; Mwesigwa, S.; Mboowa, G.; Huang, Z.; Rustagi, N.; Swaminathan, S.; Katagirya, E.; Kyobe, S.; et al. Whole-Exome Sequencing Reveals Uncaptured Variation and Distinct Ancestry in the Southern African Population of Botswana. Am. J. Hum. Genet. 2018, 102, 731–743. [Google Scholar] [CrossRef] [Green Version]
- Vatsiou, A.I.; Bazin, E.; Gaggiotti, O.E. Changes in selective pressures associated with human population expansion may explain metabolic and immune related pathways enriched for signatures of positive selection. BMC Genom. 2016, 17, 504. [Google Scholar] [CrossRef] [Green Version]
- Sabbagh, A.; Darlu, P.; Crouau-Roy, B.; Poloni, E.S. Arylamine N-acetyltransferase 2 (NAT2) genetic diversity and traditional subsistence: A worldwide population survey. PLoS ONE 2011, 6, e18507. [Google Scholar] [CrossRef] [Green Version]
- Millburn, P. Factors Affecting Glucuronidation in vivo. Biochem. Soc. Trans. 1974, 2, 1182–1186. [Google Scholar] [CrossRef] [Green Version]
- Capel, I.D.; French, M.R.; Millburn, P.; Smith, R.L.; Williams, R.T. The fate of (14C) phenol in various species. Xenobiotica 1972, 2, 25–34. [Google Scholar] [CrossRef]
- Court, M.H.; Greenblatt, D.J. Molecular genetic basis for deficient acetaminophen glucuronidation by cats: UGT1A6 is a pseudogene. and evidence for reduced diversity of expressed hepatic UGT1A isoforms. Pharmacogenetics 2000, 10, 355–369. [Google Scholar] [CrossRef]
- Court, M.H.; Greenblatt, D.J. Molecular basis for deficient acetaminophen glucuronidation in cats. An interspecies comparison of enzyme kinetics in liver microsomes. Biochem. Pharmacol. 1997, 53, 1041–1047. [Google Scholar] [CrossRef]
- Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; Abecasis, G.R. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar]
- Cardenas, C.L.L.; Bourgine, J.; Cauffiez, C.; Allorge, D.; Lo-Guidice, J.M.; Broly, F.; Chevalier, D. Genetic polymorphisms of Glycine N-acyltransferase (GLYAT) in a French Caucasian population. Xenobiotica 2010, 40, 853–861. [Google Scholar] [CrossRef] [PubMed]
- Sim, N.L.; Kumar, P.; Hu, J.; Henikoff, S.; Schneider, G.; Ng, P.C. SIFT web server: Predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 2012, 40, W452–W457. [Google Scholar] [CrossRef] [PubMed]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A method and server for predicting damaging missense mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Sibley, C.G.; Ahlquist, J.E. The phylogeny of the hominoid primates. as indicated by DNA-DNA hybridization. J. Mol. Evol. 1984, 20, 2–15. [Google Scholar] [CrossRef]
- Lovejoy, C.O. The origin of man. Science 1981, 211, 341–350. [Google Scholar] [CrossRef]
- McGrew, W.C. In search of the last common ancestor: New findings on wild chimpanzees. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 2010, 365, 3267–3276. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood. evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [Green Version]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. CABIOS 1992, 8, 275–282. [Google Scholar] [CrossRef]
- Taylor, D.J.; Piel, W.H. An assessment of accuracy. error, and conflict with support values from genome-scale phylogenetic data. Mol. Biol. Evol. 2004, 21, 1534–1537. [Google Scholar] [CrossRef] [Green Version]
- Felsenstein, J. Confidence Limits on Phylogenies: An Approach Using the Bootstrap. Evol. Int. J. Org. Evol. 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Dyda, F.; Klein, D.C.; Hickman, A.B. GCN5-related N-acetyltransferases: A structural overview. Annu. Rev. Biophys. Biomol. Struct. 2000, 29, 81–103. [Google Scholar] [CrossRef] [PubMed]
- Vetting, M.W.; de Carvalho, L.P.S.; Yu, M.; Hegde, S.S.; Magnet, S.; Roderick, S.L.; Blanchard, J.S. Structure and functions of the GNAT superfamily of acetyltransferases. Arch. Biochem. Biophys. 2005, 433, 212–226. [Google Scholar] [CrossRef]
- Ferdinand, W. The interpretation of non-hyperbolic rate curves for two-substrate enzymes. A possible mechanism for phosphofructokinase. Biochem. J. 1966, 98, 278–283. [Google Scholar] [CrossRef] [Green Version]
- Whittington, A.C.; Larion, M.; Bowler, J.M.; Ramsey, K.M.; Brüschweiler, R.; Miller, B.G. Dual allosteric activation mechanisms in monomeric human glucokinase. Proc. Natl. Acad. Sci. USA 2015, 112, 11553–11558. [Google Scholar] [CrossRef] [Green Version]
- Hilser, V.J.; Anderson, J.A.; Motlagh, H.N. Allostery vs. “allokairy”. Proc. Natl. Acad. Sci. USA 2015, 112, 11430–11431. [Google Scholar] [CrossRef] [Green Version]
- Fersht, A. Structure and Mechanism in Protein Science: A Guide to Enzyme Catalysis and Protein Folding; Hadler, G.L., Ed.; W. H. Freeman and Company: New York, NY, USA, 1999. [Google Scholar]
- Ebrecht, A.C.; Solamen, L.; Hill, B.L.; Iglesias, A.A.; Olsen, K.W.; Ballicora, M.A. Allosteric Control of Substrate Specificity of the Escherichia coli ADP-Glucose Pyrophosphorylase. Front. Chem. 2017, 5, 41. [Google Scholar] [CrossRef]
- Cornish-Bowden, A. Enzyme specificity: Its meaning in the general case. J. Theor. Biol. 1984, 108, 451–457. [Google Scholar] [CrossRef]
- Cornish-Bowden, A.; Cárdenas, M.L. Specificity of non-Michaelis-Menten enzymes: Necessary information for analyzing metabolic pathways. J. Phys. Chem. B 2010, 114, 16209–16213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quick, A.J. The conjugation of benzoic acid in man. J. Biol. Chem. 1931, 92, 65–85. [Google Scholar] [CrossRef]
- Levy, G. Pharmacokinetics of salicylate elimination in man. J. Pharm. Sci. 1965, 54, 959–967. [Google Scholar] [CrossRef] [PubMed]
- Schachter, D. The chemical estimation of acyl glucuronides and its application to studies on the metabolism of benzoate and salicylate in man. J. Clin. Investig. 1957, 36, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Campbell, L.; Wilson, H.K.; Samuel, A.M.; Gompertz, D. Interactions of m-xylene and aspirin metabolism in man. Br. J. Ind. Med. 1988, 45, 127–132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nortje, C.; van der Sluis, R.; van Dijk, A.A.; Erasmus, E. The Use of p-Aminobenzoic Acid as a Probe Substance for the Targeted Profiling of Glycine Conjugation. J. Biochem. Mol. Toxicol. 2016, 30, 136–147. [Google Scholar] [CrossRef]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2018, 46, D754–D761. [Google Scholar] [CrossRef]
- Simonsen, K.L.; Churchill, G.A.; Aquadro, C.F. Properties of statistical tests of neutrality for DNA polymorphism data. Genetics 1995, 141, 413–429. [Google Scholar] [CrossRef]
- Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 1989, 123, 585–595. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.X.; Li, W.H. Statistical tests of neutrality of mutations. Genetics 1993, 133, 693–709. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [Green Version]
- Kolvraa, S.; Gregersen, N. Acyl-CoA:glycine N-acyltransferase: Organelle localization and affinity toward straight- and branched-chained acyl-CoA esters in rat liver. Biochem. Med. Metab. Biol. 1986, 36, 98–105. [Google Scholar] [CrossRef]
- Kluyver, T.; Ragan-Kelley, B.; Pérez, F.; Granger, B.E.; Bussonnier, M.; Frederic, J.; Kelley, K.; Hamrick, J.B.; Grout, J.; Corlay, S.; et al. Jupyter Notebooks—A publishing format for reproducible computational workflows. In Positioning and Power in Academic Publishing: Players, Agents and Agendas; Loizides, F., Scmidt, B., Eds.; IOS Press: Amsterdam, The Netherlands, 2016; pp. 87–90. [Google Scholar]
- van der Walt, S.; Colbert, S.C.; Varoquaux, G. The NumPy Array: A Structure for Efficient Numerical Computation. Comput. Sci. Eng. 2011, 13, 22–30. [Google Scholar] [CrossRef] [Green Version]
- McKinney, W. Data Structures for Statistical Computing in Python. In Proceedings of the 9th Python in Science Conference, Austin, TX, USA, 28 June–3 July 2010. [Google Scholar]
- Hunter, J.D. Matplotlib: A 2D Graphics Environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Newville, M.; Otten, R.; Nelson, A.; Ingargiola, A.; Stensitzki, T.; Allan, D.; Fox, A.; Carter, F.; Michał; Pustakhod, D.; et al. lmfit/lmfit-py 1.0.2 (Version 1.0.2).Zenodo. 2021. Available online: https://zenodo.org/record/4516651 (accessed on 1 January 2020).
- Nebert, D.; McKinnon, R.; Puga, A. Human Drug-Metabolizing Enzyme Polymorphisms: Effects on Risk of Toxicity and Cancer. DNA Cell Biol. 1996, 15, 273–280. [Google Scholar] [CrossRef]
- Wallig, M.A. Glucuronidation and susceptibility to chemical carcinogenesis. Toxicol. Sci. 2004, 78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lees, H.J.; Swann, J.R.; Wilson, I.D.; Nicholson, J.K.; Holmes, E. Hippurate: The Natural History of a Mammalian-Microbial Cometabolite. J. Proteome Res. 2013, 12, 1527–1546. [Google Scholar] [CrossRef] [PubMed]
- Tfouni, S.A.; Toledo, M.C. Estimates of the mean per capita daily intake of benzoic and sorbic acids in Brazil. Food Addit. Contam. 2002, 19, 647–654. [Google Scholar] [CrossRef] [PubMed]




| Population | Exomes | Genomes | Total |
|---|---|---|---|
| African/African American | 8 128 | 4 359 | 12,487 |
| Latino | 17,296 | 424 | 17,720 |
| Ashkenazi Jewish | 5 040 | 145 | 5 185 |
| East Asian | 9 197 | 780 | 9 977 |
| Finnish | 10,824 | 1 738 | 12,562 |
| Non-Finnish European | 56,885 | 7 718 | 64,603 |
| South Asian | 15,308 | # | 15,308 |
| Other* | 3 070 | 544 | 3 614 |
| Female | 57,787 | 6 967 | 64,754 |
| Male | 67,961 | 8 741 | 76,702 |
| Total | 125,748 | 15,708 | 138,632 |
| Gene | Number of Sequences (m) | Number of Segregating Sites (S) | Nucleotide Diversity (π) | Tajima Test Statistic (D) |
|---|---|---|---|---|
| GLYAT | 25 | 21 | 0.007748 | −2.13 |
| Haplotype | Vf µmol min−1 mg protein−1 | kcat s−1 | s0.5,gly mM | hgly | s0.5,benz µM | hbenz |
|---|---|---|---|---|---|---|
| 156Asn > Ser | 0.85 ± 0.06 | 0.48 ± 0.03 | 23 ± 2 | 1.6 ± 0.1 | 97 ± 3 | 2.1 ± 0.1 |
| 17Ser > Thr,156Asn > Ser | 0.62 ± 0.02 | 0.35 ± 0.01 | 29 ± 3 | 1.3 ± 0.1 | 118 ± 7 | 1.5 ± 0.1 |
| 156Asn > Ser,199Arg > Cys | 0.083 ± 0.005 | 0.047 ± 0.003 | 30 ± 3 | 1.4 ± 0.1 | 61 ± 3 | 3.5 ± 0.5 |
| Parameters | Values | Recombinant Variant/Isolated from Liver | Reference |
|---|---|---|---|
| KMapp (benzoyl-CoA) (µM) | 13 | Purified GLYAT from human liver | [33] |
| 28 ± 5 | Purified recombinant 17Ser > Thr variant | [31] | |
| 38 ± 4 | Purified recombinant 156Asn > Ser variant | [31] | |
| s0.5 | 61 ± 3 | Purified recombinant 156Asn > Ser,199Arg > Cys variant | This study (bi-substrate Hill) |
| KMapp (benzoyl-CoA) (µM) | 67 ± 5 | Partially purified GLYAT from human liver | [32] |
| 79 ± 38 | Purified recombinant wildtype | [34] | |
| 88 ± 66 | Purified recombinant 156Asn > Ser variant | [34] | |
| s0.5 | 97 ± 3 | Purified recombinant 156Asn > Ser variant | This study (bi-substrate Hill) |
| 118 ± 7 | Purified recombinant 17Ser > Thr,156Asn > Ser variant | This study (bi-substrate Hill) | |
| KMapp (benzoyl-CoA) (µM) | 139 ± 85 | Purified recombinant L61 variant | [34] |
| 209 | Purified recombinant 156Asn > Ser variant | [28] | |
| 57900 * | Purified GLYAT from human liver | [30] | |
| KMapp (glycine) (mM) | 6.4 | Purified GLYAT from human liver | [33] |
| 6.5 ± 1 | Partially purified GLYAT from human liver | [32] | |
| s0.5 | 23 ± 2 | Purified recombinant 156Asn > Ser variant | This study (bi-substrate Hill) |
| KMapp (glycine) (mM) | 26.6 | Purified recombinant 156Asn > Ser variant | [28] |
| s0.5 | 29 ± 3 | Purified recombinant 17Ser > Thr,156Asn > Ser variant | This study (bi-substrate Hill) |
| 30 ± 3 | Purified recombinant 156Asn > Ser199Arg > Cys variant | This study (bi-substrate Hill) | |
| Vmax (nmol/min/mg) | 83 ± 5 | Purified recombinant 156Asn > Ser199Arg > Cys variant | This study (bi-substrate Hill) |
| 543 ± 21 | Purified GLYAT from human liver | [33] | |
| 620 ± 20 | Purified recombinant 17Ser > Thr,156Asn > Ser variant | This study (bi-substrate Hill) | |
| 665 ± 40 | Purified recombinant 17Ser > Thr variant | [31] | |
| 807 | Recombinant 156Asn > Ser variant | [28] | |
| 850 ± 60 | Purified recombinant 156Asn > Ser variant | This study (bi-substrate Hill) | |
| 1230 | Purified recombinant 156Asn > Ser variant | [31] | |
| 17,100 # | Purified GLYAT from human liver | [30] | |
| 121,000 + | Purified recombinant L61 variant | [34] | |
| 490,000 + | Purified recombinant wildtype | [34] | |
| 1,359,000 + | Purified recombinant 156Asn > Ser variant | [34] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rohwer, J.M.; Schutte, C.; van der Sluis, R. Functional Characterisation of Three Glycine N-Acyltransferase Variants and the Effect on Glycine Conjugation to Benzoyl–CoA. Int. J. Mol. Sci. 2021, 22, 3129. https://doi.org/10.3390/ijms22063129
Rohwer JM, Schutte C, van der Sluis R. Functional Characterisation of Three Glycine N-Acyltransferase Variants and the Effect on Glycine Conjugation to Benzoyl–CoA. International Journal of Molecular Sciences. 2021; 22(6):3129. https://doi.org/10.3390/ijms22063129
Chicago/Turabian StyleRohwer, Johann M., Chantelle Schutte, and Rencia van der Sluis. 2021. "Functional Characterisation of Three Glycine N-Acyltransferase Variants and the Effect on Glycine Conjugation to Benzoyl–CoA" International Journal of Molecular Sciences 22, no. 6: 3129. https://doi.org/10.3390/ijms22063129








