The Roles of Cullins E3 Ubiquitin Ligases in the Lipid Biosynthesis of the Green Microalgae Chlamydomonas reinhardtii
Abstract
:1. Introduction
2. Results and Discussions
2.1. Cullins in C. Reinhardtii
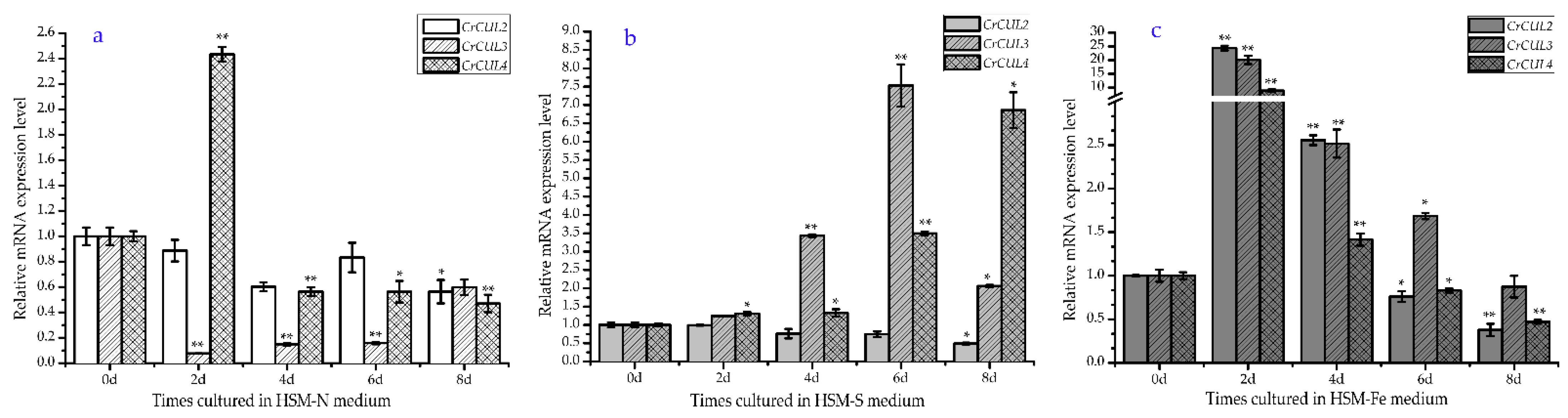
2.2. Expression Patterns of CrCUL Genes under Different Nutrition Deficiency Conditions
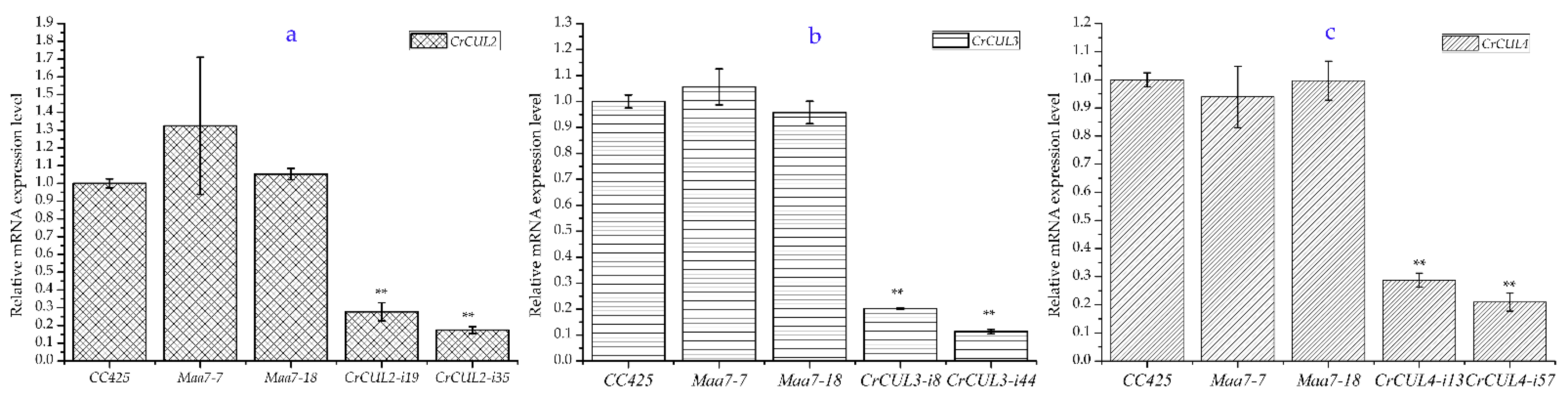
2.3. RNAi Expression of CrCULs in C. reinhardtii
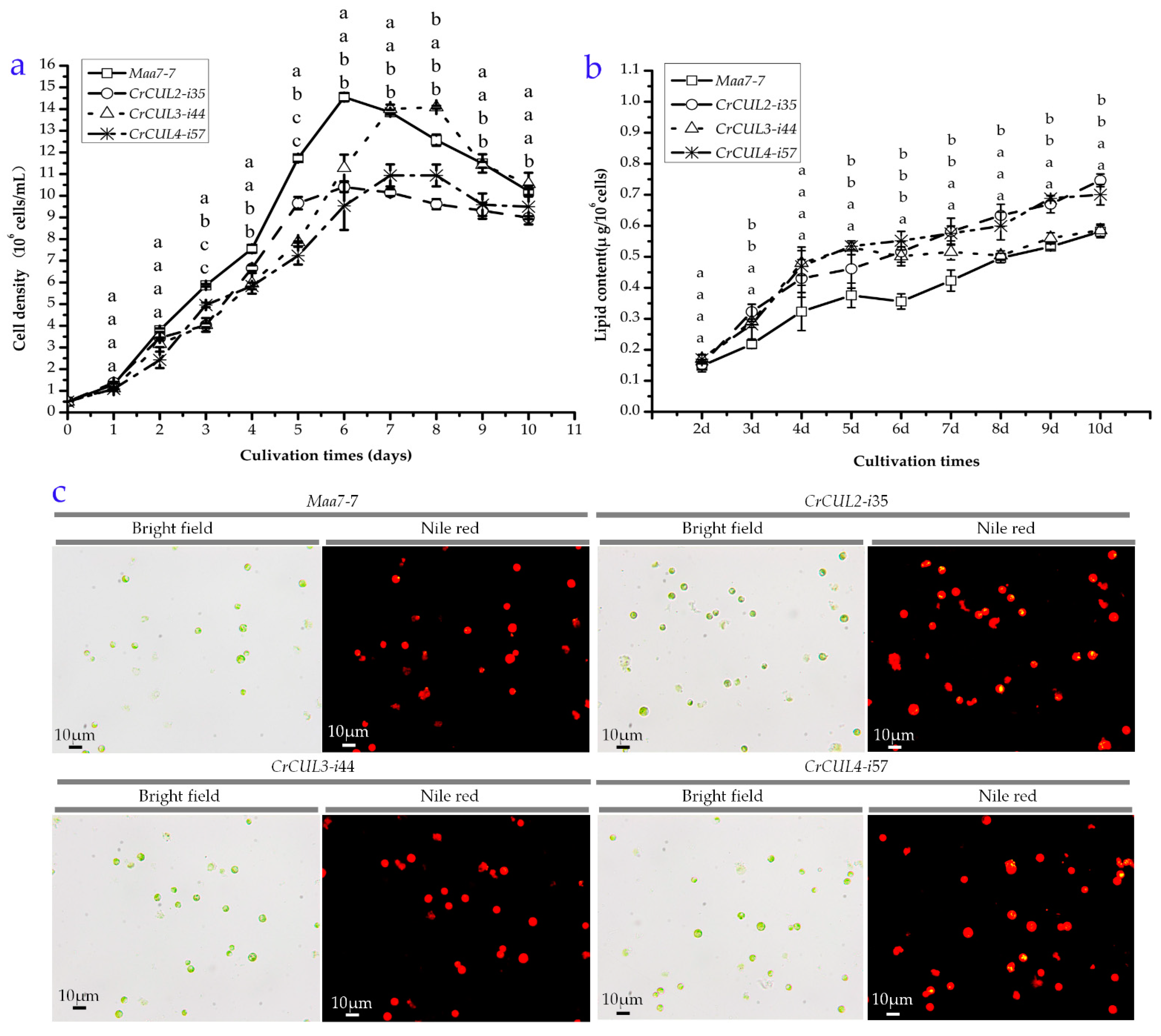
2.4. Growth Kinetics and Lipid Contents of CrCUL RNAi Lines
2.5. Fatty Acid Composition Analysis
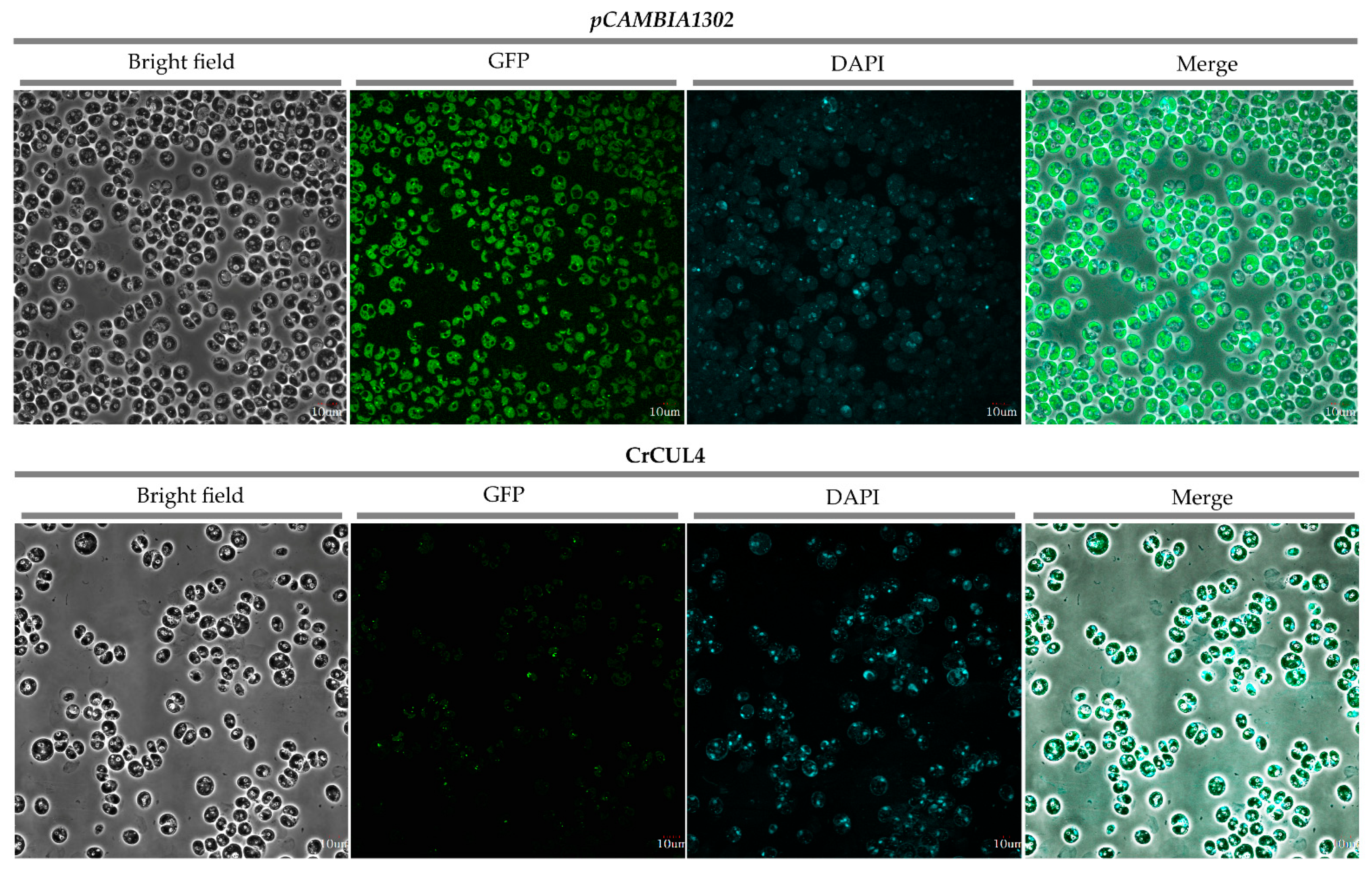
2.6. Subcellular Localization of CrCUL4
3. Materials and Methods
3.1. In Silico Identification and Sequence Analysis of CrCULs
3.2. C. reinhardtii Strain and Growth Conditions
3.3. Gene Expression Analysis of CrCULs
3.4. RNA Interference (RNAi) Vector Construction and Algae Transformation
3.5. Growth and Neutral Lipid Analysis
3.6. Fatty Acid Methyl Ester Profiling
3.7. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CRLs | Cullin-RING E3 ligases |
| CUL | Cullin |
| C. reinhardtii | Chlamydomonas reinhardtii |
| RNAi | RNA interference |
| TAG | Triacylglycerol |
| PDAT | Diacylglycerol acyltransferase |
| DGAT | Diacylglycerol acyltransferase |
| GPAT | Glycerol-3-phosphate acyltransferase |
| GPDH | Glycerol-3-Phosphate Dehydrogenases |
| Fat | Acyl-ACP thioesterase |
| ACCase | Acetyl-CoA carboxylase |
| PEPC | Phosphoenolpyruvate carboxylase |
| UPP | Ubiquitin proteasome pathway |
| E3s | E3 ubiquitin ligases |
| LD | Lipid droplets |
| SRS | Substrate-recognition subunit |
| SKP1 | S-phase kinase-associated protein 1 |
| SOCS/BC | Suppressor of cytokine signaling/elongin-BC |
| BTB | Broad complex, Tramtrack, Bric-a-brac |
| DDB1 | DNA damage-binding protein-1 |
| RBX1 | RING Box 1 |
| PARC | PARkin-like cytoplasmic protein |
| FA | Fatty acid |
| TAP | Tris Acetate Phosphate medium |
| HSM | Sueoka’s High Salt Medium |
| -N | Nitrogen deficiency |
| -S | Sulfur deficiency |
| -Fe | Iron deficiency |
| COP I | Coat Protein Complex I |
| WDR | WD40 repeat domain |
| IR | Tandem inverted repeat |
| GFP | Green fluorescent protein |
References
- Enamala, M.K.; Enamala, S.; Chavali, M.; Donepudi, J.; Yadavalli, R.; Kolapalli, B.; Aradhyula, T.V.; Velpuri, J.; Kuppam, C. Production of biofuels from microalgae—A review on cultivation, harvesting, lipid extraction, and numerous applications of microalgae. Renew. Sustain. Energy Rev. 2018, 94, 49–68. [Google Scholar] [CrossRef]
- Cakmak, T.; Angun, P.; Ozkan, A.D.; Cakmak, Z.; Olmez, T.T.; Tekinay, T. Nitrogen and sulfur deprivation differentiate lipid accumulation targets of Chlamydomonas reinhardtii. Bioengineered 2012, 3, 343–346. [Google Scholar] [CrossRef] [Green Version]
- Yoon, K.; Han, D.; Li, Y.; Sommerfeld, M.; Hu, Q. Phospholipid:diacylglycerol acyltransferase is a multifunctional enzyme involved in membrane lipid turnover and degradation while synthesizing triacylglycerol in the unicellular green microalga Chlamydomonas reinhardtii. Plant Cell 2012, 24, 3708–3724. [Google Scholar] [CrossRef] [Green Version]
- Driver, T.; Trivedi, D.K.; McIntosh, O.A.; Dean, A.P.; Goodacre, R.; Pittman, J.K. Two glycerol-3-phosphate dehydrogenases from Chlamydomonas have distinct roles in lipid metabolism. Plant Physiol. 2017, 174, 2083–2097. [Google Scholar] [CrossRef] [Green Version]
- Hung, C.H.; Ho, M.Y.; Kanehara, K.; Nakamura, Y. Functional study of diacylglycerol acyltransferase type 2 family in Chlamydomonas reinhardtii. FEBS Lett. 2013, 587, 2364–2370. [Google Scholar] [CrossRef] [Green Version]
- Herrera-Valencia, V.A.; Macario-Gonzalez, L.A.; Casais-Molina, M.L.; Beltran-Aguilar, A.G.; Peraza-Echeverria, S. In silico cloning and characterization of the glycerol-3-phosphate dehydrogenase (GPDH) gene family in the green microalga Chlamydomonas reinhardtii. Curr. Microbiol. 2012, 64, 477–485. [Google Scholar] [CrossRef]
- Deng, X.D.; Gu, B.; Li, Y.J.; Hu, X.W.; Guo, J.C.; Fei, X.W. The roles of acyl-CoA: Diacylglycerol acyltransferase 2 genes in the biosynthesis of triacylglycerols by the green algae Chlamydomonas reinhardtii. Mol. Plant. 2012, 5, 945–947. [Google Scholar] [CrossRef] [Green Version]
- Russa, M.L.; Bogen, C.; Uhmeyer, A.; Doebbe, A.; Filippone, E.; Kruse, O.; Mussgnug, J.H. Functional analysis of three type-2 DGAT homologue genes for triacylglycerol production in the green microalga Chlamydomonas reinhardtii. J. Biotechnol. 2012, 162, 13–20. [Google Scholar] [CrossRef]
- Tan, K.; Lee, Y.K. Expression of the heterologous Dunaliella tertiolecta fatty acyl-ACP thioesterase leads to increased lipid production in Chlamydomonas reinhardtii. J. Biotechnol. 2017, 247, 60–67. [Google Scholar] [CrossRef]
- Kao, P.H.; Ng, I.S. CRISPRi mediated phosphoenolpyruvate carboxylase regulation to enhance the production of lipid in Chlamydomonas reinhardtii. Bioresour. Technol. 2017, 245, 1527–1537. [Google Scholar] [CrossRef]
- Chen, D.; Yuan, X.; Liang, L.; Liu, K.; Ye, H.; Liu, Z.; Liu, Y.; Huang, L.; He, W.; Chen, Y.; et al. Overexpression of acetyl-CoA carboxylase increases fatty acid production in the green alga Chlamydomonas reinhardtii. Biotechnol. Lett. 2019, 41, 1133–1145. [Google Scholar] [CrossRef]
- Miller, R.; Wu, G.; Deshpande, R.R.; Vieler, A.; Gartner, K.; Li, X.; Moellering, E.R.; Zauner, S.; Cornish, A.J.; Liu, B.; et al. Changes in transcript abundance in Chlamydomonas reinhardtii following nitrogen deprivation predict diversion of metabolism. Plant Physiol. 2010, 154, 1737–1752. [Google Scholar] [CrossRef] [Green Version]
- Ramanan, R.; Kim, B.H.; Cho, D.H.; Ko, S.R.; Oh, H.M.; Kim, H.S. Lipid droplet synthesis is limited by acetate availability in starchless mutant of Chlamydomonas reinhardtii. FEBS Lett. 2013, 587, 370–377. [Google Scholar] [CrossRef] [Green Version]
- Smalle, J.; Vierstra, R.D. The ubiquitin 26S proteasome proteolytic pathway. Annu. Rev. Plant Biol. 2004, 55, 555–590. [Google Scholar] [CrossRef] [PubMed]
- Qi, L.; Heredia, J.E.; Altarejos, J.Y.; Screaton, R.; Goebel, N.; Niessen, S.; Macleod, I.X.; Liew, C.W.; Kulkarni, R.N.; Bain, J.; et al. TRB3 links the E3 ubiquitin ligase COP1 to lipid metabolism. Science 2006, 312, 1763–1766. [Google Scholar] [CrossRef] [PubMed]
- Thiele, C.; Spandl, J. Cell biology of lipid droplets. Curr. Opin. Cell. Biol. 2008, 20, 378–385. [Google Scholar] [CrossRef] [PubMed]
- Vierstra, Z.; Vierstra, R.D. The Cullin-RING ubiquitin-protein ligases. Annu. Rev. Plant Biol. 2011, 62, 299–334. [Google Scholar]
- Marín, I. Diversification of the cullin family. BMC Evol. Biol. 2009, 9, 267. [Google Scholar] [CrossRef] [Green Version]
- Roodbarkelari, F.; Bramsiepe, J.; Weinl, C.; Marquardt, S.; Novak, B.; Jakoby, M.J.; Lechner, E.; Genschik, P.; Schnittger, A. Cullin 4-ring finger-ligase plays a key role in the control of endoreplication cycles in Arabidopsis trichomes. Proc. Natl. Acad. Sci. USA 2010, 107, 15275–15280. [Google Scholar] [CrossRef] [Green Version]
- Dias, D.C.; Dolios, G.; Wang, R.; Pan, Z.Q. CUL7: A DOC domain-containing cullin selectively binds Skp1.Fbx29 to form an SCF-like complex. Proc. Natl. Acad. Sci. USA 2002, 99, 16601–16606. [Google Scholar] [CrossRef] [Green Version]
- Yin, C.Y.; Thilo, H.; Karl-Wilhelm, K. Mechanism of Cullin3 E3 Ubiquitin Ligase Dimerization. PLoS ONE 2012, 7, e41350. [Google Scholar]
- Dumbliauskas, E.; Lechner, E.; Jaciubek, M.; Berr, A.; Pazhouhandeh, M.; Alioua, M.; Cognat, V.; Brukhin, V.; Koncz, C.; Grossniklaus, U.; et al. The Arabidopsis CUL4-DDB1 complex interacts with MSI1 and is required to maintain MEDEA parental imprinting. EMBO J. 2011, 30, 731–743. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aihara, Y.; Fujimura-Kamada, K.; Yamasaki, T.; Minagawa, J. Algal photoprotection is regulated by the E3 ligase CUL4-DDB1DET1. Nat. Plants 2018, 5, 34–40. [Google Scholar] [CrossRef]
- Sarikas, A.; Hartmann, T.; Pan, Z.Q. The cullin protein family. Genome Biol. 2011, 12, 220. [Google Scholar] [CrossRef] [Green Version]
- Hannah, J.; Zhou, P. Distinct and overlapping functions of the cullin E3 ligase scaffolding proteins CUL4A and CUL4B. Gene 2015, 573, 33–45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dieterle, M.; Thomann, A.; Renou, J.P.; Parmentier, Y.; Cognat, V.; Lemonnier, G.; Muller, R.; Shen, W.H.; Kretsch, T.; Genschik, P. Molecular and functional characterization of Arabidopsis Cullin 3A. Plant J. 2005, 41, 386–399. [Google Scholar] [CrossRef] [Green Version]
- Fischer, E.S.; Scrima, A.; Bohm, K.; Matsumoto, S.; Lingaraju, G.M.; Faty, M.; Yasuda, T.; Cavadini, S.; Wakasugi, M.; Hanaoka, F.; et al. The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 2011, 147, 1024–1039. [Google Scholar] [CrossRef] [Green Version]
- Lv, H.; Qu, G.; Qi, X.; Lu, L.; Tian, C.; Ma, Y. Transcriptome analysis of Chlamydomonas reinhardtii during the process of lipid accumulation. Genomics 2013, 101, 229–237. [Google Scholar] [CrossRef] [Green Version]
- Wase, N.; Black, P.N.; Stanley, B.A.; DiRusso, C.C. Integrated quantitative analysis of nitrogen stress response in Chlamydomonas reinhardtii using metabolite and protein profiling. J. Proteome Res. 2014, 13, 1373–1396. [Google Scholar] [CrossRef]
- Roberts, D.; Pedmale, U.V.; Morrow, J.; Sachdev, S.; Lechner, E.; Tang, X.; Zheng, N.; Hannink, M.; Genschik, P.; Liscum, E. Modulation of phototropic responsiveness in Arabidopsis through ubiquitination of phototropin 1 by the CUL3-Ring E3 ubiquitin ligase CRL3NPH3. Plant Cell 2011, 23, 3627–3640. [Google Scholar] [CrossRef] [Green Version]
- Christians, M.J.; Rottier, A.; Wiersma, C. Light regulates the RUBylation levels of individual Cullin proteins in Arabidopsis thaliana. Plant Mol. Biol. Rep. 2018, 36, 123–134. [Google Scholar] [CrossRef] [Green Version]
- Devadasu, E.; Chinthapalli, D.K.; Chouhan, N.; Madireddi, S.K.; Rasineni, G.K.; Sripadi, P.; Subramanyam, R. Changes in the photosynthetic apparatus and lipid droplet formation in Chlamydomonas reinhardtii under iron deficiency. Photosynth. Res. 2019, 139, 253–266. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, S.I.; Castruita, M.; Malasarn, D.; Urzica, E.; Erde, J.; Page, M.D.; Yamasaki, H.; Casero, D.; Pellegrini, M.; Merchant, S.S.; et al. The proteome of copper, iron, zinc, and manganese micronutrient deficiency in Chlamydomonas reinhardtii. Mol. Cell. Proteom. 2013, 12, 65–86. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beller, M.; Sztalryd, C.; Southall, N.; Ming, B.; Oliver, B. CopI complex is a regulator of lipid homeostasis. PLoS Biol. 2008, 6, e292. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rohr, J.; Sarkar, N.; Balenger, S.; Jeong, B.R.; Cerutti, H. Tandem inverted repeat system for selection of effective transgenic RNAi strains in Chlamydomonas. Plant J. 2004, 40, 611–621. [Google Scholar] [CrossRef] [Green Version]
- Teixeira, L.K.; Reed, S.I. Ubiquitin ligases and cell cycle control. Annu. Rev. Biochem. 2013, 82, 387–414. [Google Scholar] [CrossRef]
- Feng, H.; Zhong, W.; Punkosdy, G. CUL-2 is required for the G1-to-S-phase transition and mitotic chromosome condensation in Caenorhabditis elegans. Nat. Cell Biol. 1999, 1, 486–492. [Google Scholar] [CrossRef] [PubMed]
- Dubiel, D.; Bintig, W.; Kähne, T.; Dubiel, W.; Naumann, M. Cul3 neddylation is crucial for gradual lipid droplet formation during adipogenesis. BBA Mol. Cell Res. 2017, 1864, 1405–1412. [Google Scholar] [CrossRef]
- Chen, L.; Lee, J.H.; Weber, H.; Tohge, T.; Witt, S.; Roje, S.; Fernie, A.R.; Hellmann, H. Arabidopsis BPM proteins function as substrate adaptors to a cullin3-based E3 ligase to affect fatty acid metabolism in plants. Plant Cell 2013, 25, 2253–2264. [Google Scholar] [CrossRef] [Green Version]
- Kansanen, E.; Jyrkkanen, H.K.; Levonen, A.L. Activation of stress signaling pathways by electrophilic oxidized and nitrated lipids. Free Radic. Biol. Med. 2012, 52, 973–982. [Google Scholar] [CrossRef]
- Zhang, Y.; Feng, S.; Chen, F.; Chen, H.; Wang, J.; Mccall, C.; Xiong, Y.; Deng, X.W. Arabidopsis DDB1-CUL4 associated factor1 forms a nuclear E3 ubiquitin ligase with DDB1 and CUL4 that is involved in multiple plant developmental processes. Plant Cell 2008, 20, 1437–1455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Faull, S.V.; Lau, A.M.C.; Martens, C.; Ahdash, Z.; Hansen, K.; Yebenes, H.; Schmidt, C.; Beuron, F.; Cronin, N.B.; Morris, E.P.; et al. Structural basis of Cullin 2 RING E3 ligase regulation by the COP9 signalosome. Nat. Commun. 2019, 10, 3814. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kindle, K.L. High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc. Natl. Acad. Sci. USA 1990, 87, 1228–1232. [Google Scholar] [CrossRef] [Green Version]
- Luo, Q.; Song, W.; Li, Y.; Wang, C.; Hu, Z. Flagella-Associated WDR-Containing Protein CrFAP89 Regulates Growth and Lipid Accumulation in Chlamydomonas reinhardtii. Front. Plant Sci. 2018, 9, 691. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Gene Name | Chromosome Position | Introns | ORF | Deduced Polypeptide | ||||
|---|---|---|---|---|---|---|---|---|
| Amino Acids | MW (kDa) | pI | GRAVY | Transmembrane Helices | ||||
| CrCUL2 | 17:4895502…4903556F | 19 | 2244 | 747 | 87.6 | 6.98 | −0.214 | 3 |
| CrCUL3 | 7:1516277…1525175F | 18 | 2211 | 736 | 84.8 | 8.14 | −0.473 | 0 |
| CrCUL4 | 12:3957943…3963740F | 14 | 2301 | 766 | 84.8 | 8.49 | −0.576 | 1 |
| FA | Maa7-7 | CrCUL2-i35 | CrCUL3-i44 | CrCUL4-i57 |
|---|---|---|---|---|
| C12:0 | 0.04 ± 0.01 | 0.04 ± 0.00 | 0.03 ± 0.00 | 0.04 ± 0.00 |
| C14:0 | 0.04 ± 0.01 | 0.12 ± 0.11 | 0.04 ± 0.00 | 0.05 ± 0.01 |
| C16:0 | 12.25 ± 0.65 a | 14.68 ± 0.89 b | 11.28 ± 1.60 a | 16.77 ± 1.47 c |
| C16:1 | 3.95 ± 0.11 | 3.45 ± 0.63 | 2.01 ± 0.68 | 4.55 ± 2.21 |
| C16:2 | 1.21 ± 0.12 a | 1.15 ± 0.20 b | 2.00 ± 0.03 a | 1.35 ± 0.19 a |
| C16:3 | 1.08 ± 0.36 | 0.80 ± 0.06 | 1.29 ± 0.11 | 1.35 ± 0.27 |
| C16:4 | 4.68 ± 0.19 | 6.35 ± 2.53 | 7.66 ± 0.85 | 7.37 ± 2.04 |
| C18:0 | 1.40 ± 0.12 a | 1.64 ± 0.09 b | 1.27 ± 0.09 a | 1.83 ± 0.15 b |
| C18:1n9t | 4.10 ± 0.40 | 4.77 ± 0.51 | 3.12 ± 1.13 | 4.44 ± 2.90 |
| C18:1n9c | 5.50 ± 0.54 | 5.88 ± 0.89 | 4.39 ± 0.56 | 6.85 ± 2.37 |
| C18:2 | 5.74 ± 0.23 | 6.27 ± 0.52 | 7.45 ± 0.98 | 7.31 ± 1.59 |
| C18:3 (5,9,12) | 6.30 ± 0.32 a | 6.47 ± 0.25 a | 5.56 ± 0.74 a | 9.65 ± 0.51 b |
| C18:3n3 | 8.20 ± 0.63 a | 9.67 ± 2.35 a | 10.81 ± 1.36 a | 12.64 ± 3.16 b |
| C20:0 | 0.42 ± 0.03 a | 0.50 ± 0.10 a | 0.55 ± 0.03 a | 0.63 ± 0.15 b |
| C20:1 | 0.26 ± 0.03 a | 0.47 ± 0.31 a | 0.60 ± 0.03 b | 0.44 ± 0.16 a |
| Total FA | 55.19 ± 0.92 a | 62.24 ± 2.78 b | 58.04 ± 2.11 a | 75.30 ± 2.00 c |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, Q.; Zou, X.; Wang, C.; Li, Y.; Hu, Z. The Roles of Cullins E3 Ubiquitin Ligases in the Lipid Biosynthesis of the Green Microalgae Chlamydomonas reinhardtii. Int. J. Mol. Sci. 2021, 22, 4695. https://doi.org/10.3390/ijms22094695
Luo Q, Zou X, Wang C, Li Y, Hu Z. The Roles of Cullins E3 Ubiquitin Ligases in the Lipid Biosynthesis of the Green Microalgae Chlamydomonas reinhardtii. International Journal of Molecular Sciences. 2021; 22(9):4695. https://doi.org/10.3390/ijms22094695
Chicago/Turabian StyleLuo, Qiulan, Xianghui Zou, Chaogang Wang, Yajun Li, and Zhangli Hu. 2021. "The Roles of Cullins E3 Ubiquitin Ligases in the Lipid Biosynthesis of the Green Microalgae Chlamydomonas reinhardtii" International Journal of Molecular Sciences 22, no. 9: 4695. https://doi.org/10.3390/ijms22094695






