Genome-Wide Identification of the Salvia miltiorrhiza SmCIPK Gene Family and Revealing the Salt Resistance Characteristic of SmCIPK13
Abstract
:1. Introduction
2. Result
2.1. Determination and Sequence Characterization of SmCIPKs Gene Family Members
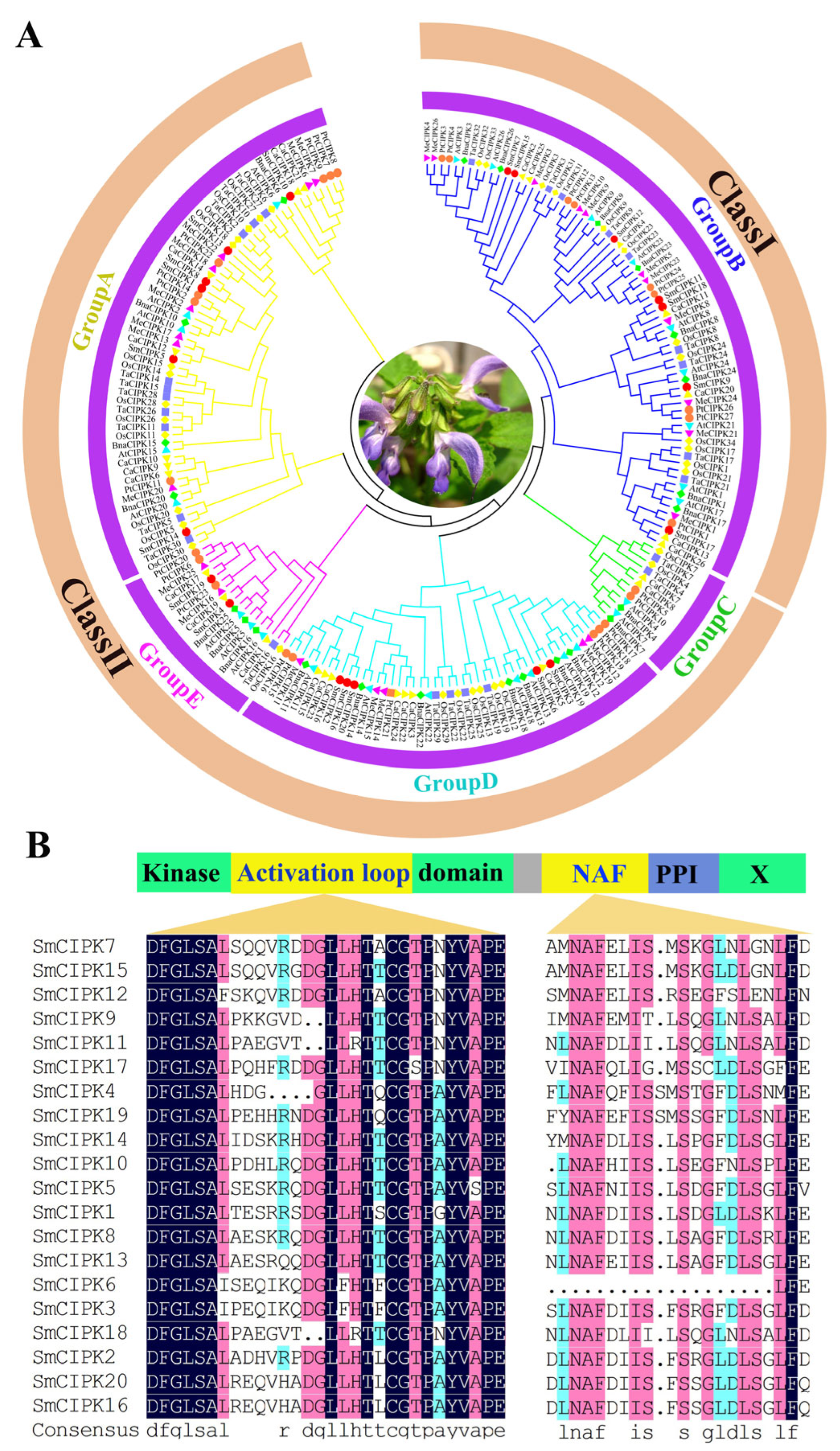
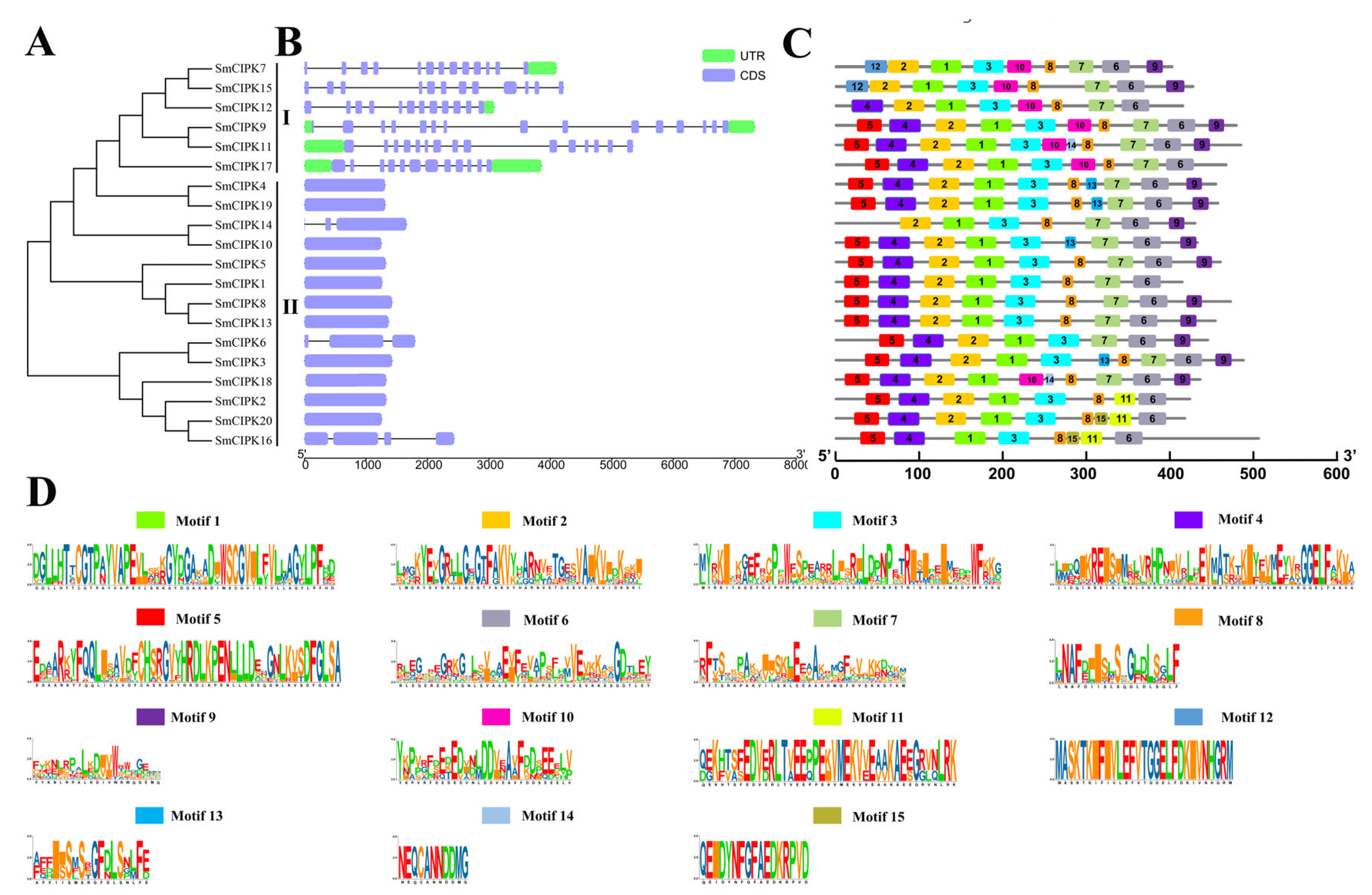
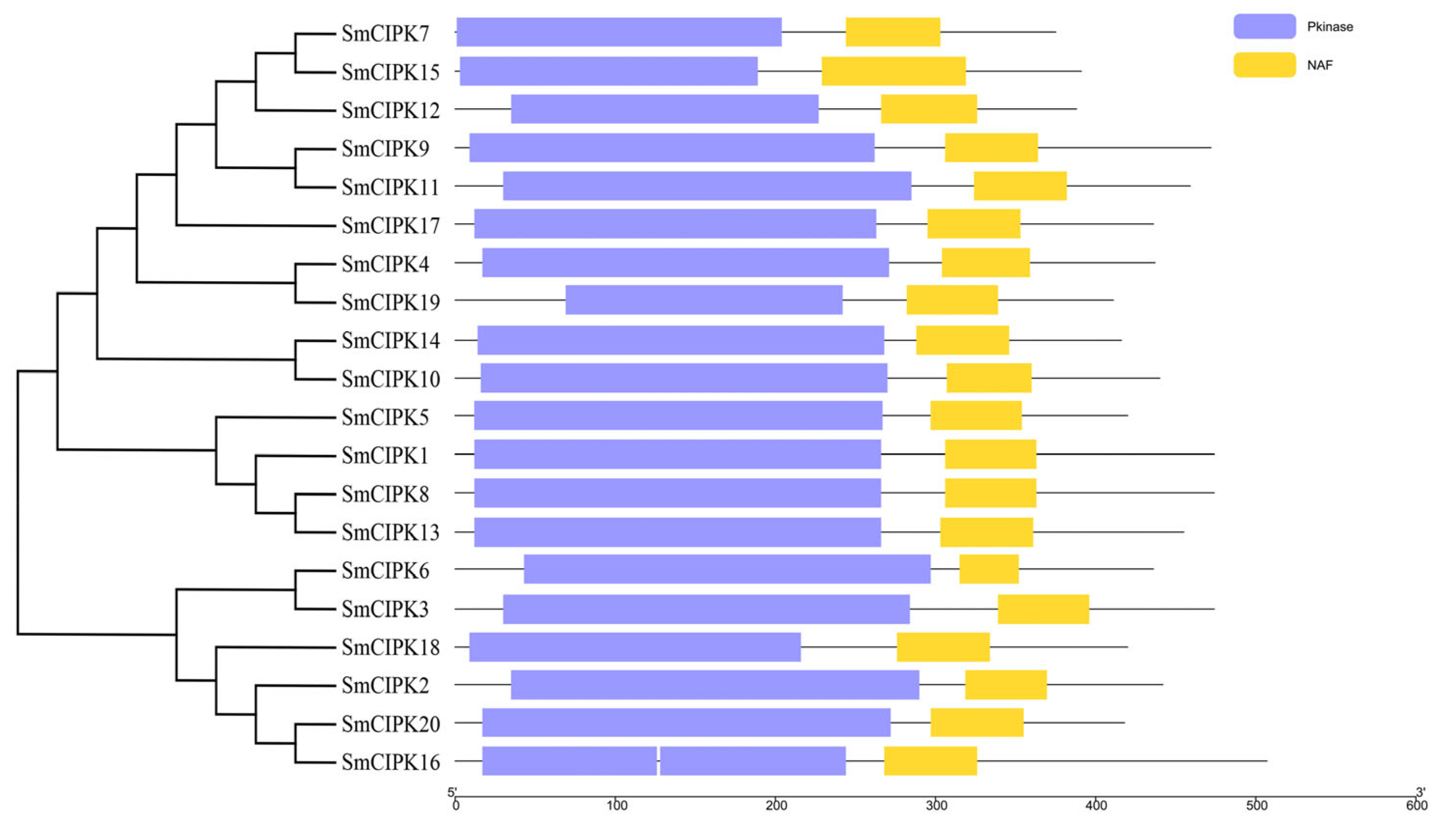
2.2. Phylogenetic and Sequence Analysis of SmCIPKs
2.3. Expression Profiles of SmCIPK Genes
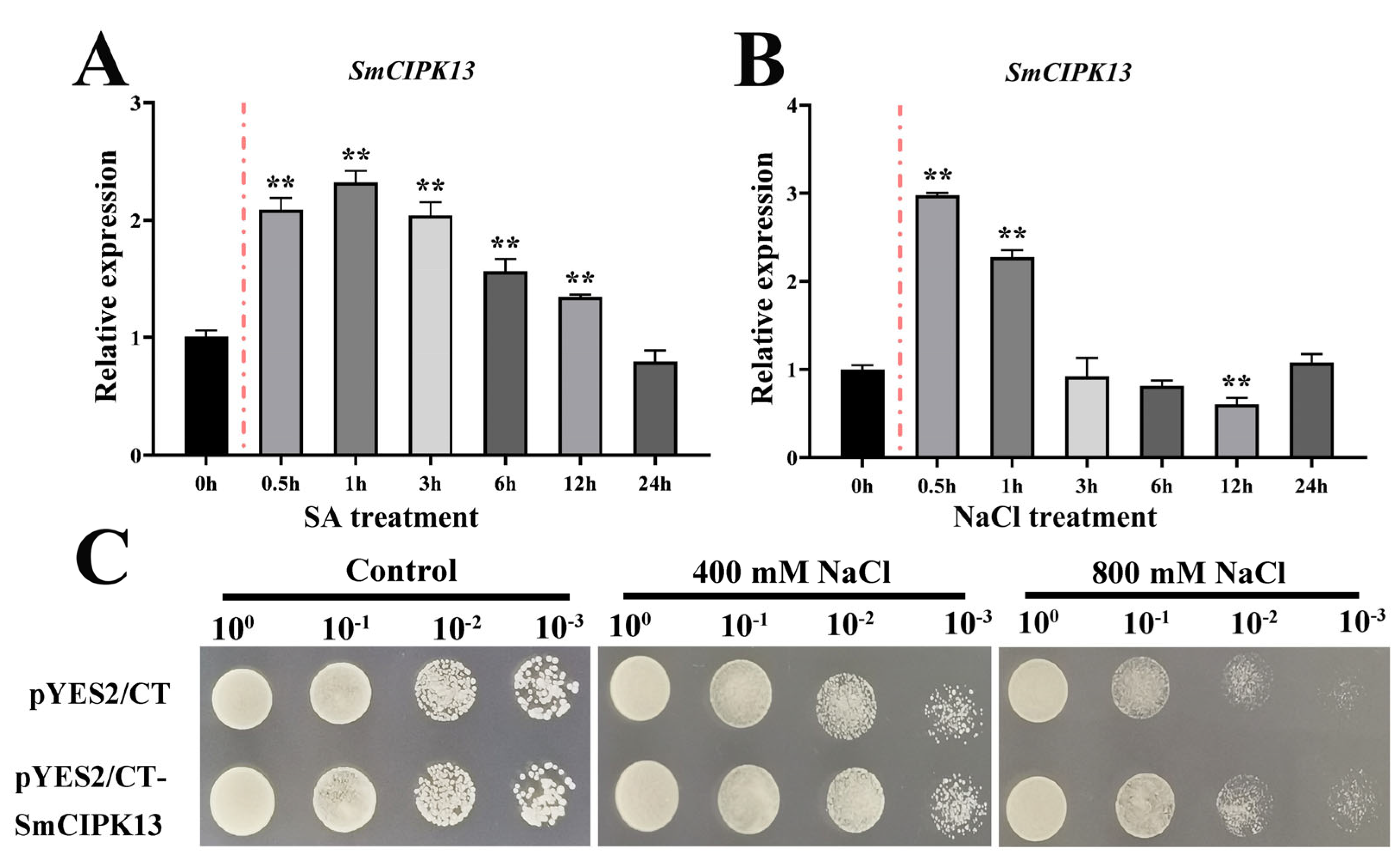
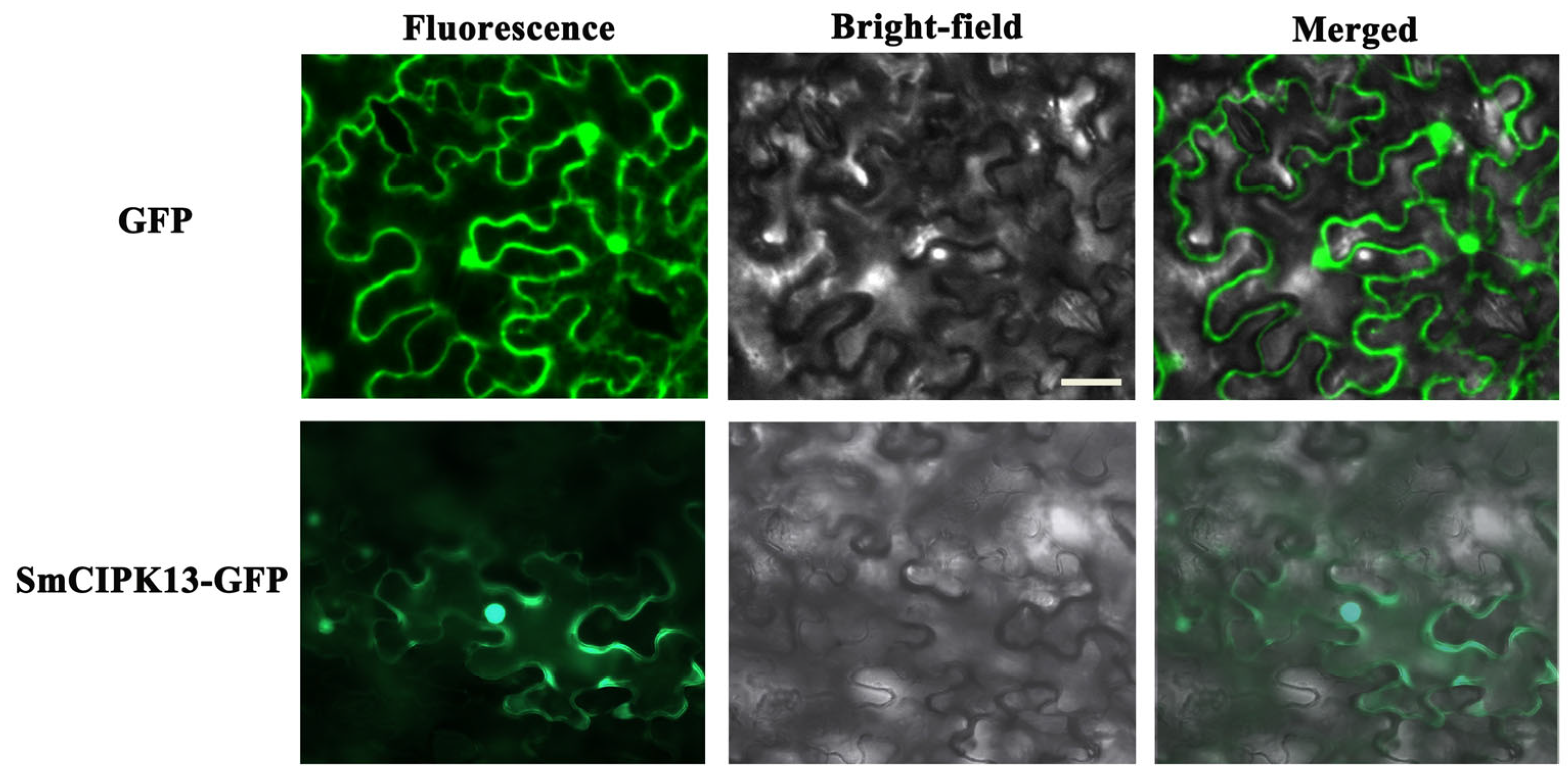
2.4. Localization of SmCIPK13 Protein and Analysis of Its Response to Abiotic Stresses
2.5. Expression and Overexpression of SmCIPK13 Promoted Salt Resistance in Yeast and Arabidopsis, Respectively
3. Discussion
4. Material and Methods
4.1. Plant Materials and Treatment
4.2. Identification and Characterization of Gene Family Members
4.3. Protein Sequence, Phylogenetic Tree Construction and Gene Structure Analysis
4.4. Analysis of SmCIPKs Expression Patterns in Different Tissues and under Different Treatments
4.5. Plasmid Construction, Transgenic Arabidopsis and Stress Treatment
4.6. Subcellular Localization of SmCIPK13 and Salt Stress Tolerance Assay in Yeast
4.7. Statistics Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Knight, H.; Knight, M.R. Abiotic stress signalling pathways: Specificity and cross-talk. Trends Plant Sci. 2001, 6, 262–267. [Google Scholar] [CrossRef]
- Xi, Y.; Liu, J.; Dong, C.; Cheng, Z.M. The CBL and CIPK Gene Family in Grapevine (Vitis vinifera): Genome-Wide Analysis and Expression Profiles in Response to Various Abiotic Stresses. Front Plant Sci. 2017, 8, 978. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luan, S.; Lan, W.; Chul Lee, S. Potassium nutrition, sodium toxicity, and calcium signaling: Connections through the CBL-CIPK network. Curr. Opin. Plant Biol. 2009, 12, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Batistič, O.; Kudla, J.J. Integration and channeling of calcium signaling through the CBL calcium sensor/CIPK protein kinase network. Planta 2004, 219, 915–924. [Google Scholar] [CrossRef]
- Sanyal, S.; Pandey, A.; Pandey, G. The CBL-CIPK signaling module in plants: A mechanistic perspective. Physiol. Plant. 2015, 155, 89–108. [Google Scholar] [CrossRef]
- Liu, J.X.; Ishitani, M.; Halfter, U.; Kim, C.S.; Zhu, J.K. The Arabidopsis thaliana SOS2 gene encodes a protein kinase that is required for salt tolerance. Proc. Natl. Acad. Sci. USA 2000, 97, 3730–3734. [Google Scholar] [CrossRef]
- Halfter, U.; Ishitani, M.; Zhu, J.K. The Arabidopsis SOS2 protein kinase physically interacts with and is activated by the calcium-binding protein SOS3. Proc. Natl. Acad. Sci. USA 2000, 97, 3735–3740. [Google Scholar] [CrossRef]
- Prodhan, M.Y.; Munemasa, S.; Nahar, M.N.; Nakamura, Y.; Murata, Y. Guard cell salicylic acid signaling is integrated into abscisic acid signaling via the Ca2+/CPK-dependent pathway. Plant Physiol. 2018, 178, 441–450. [Google Scholar] [CrossRef] [Green Version]
- Halford, N.G.; Grahame Hardie, D. SNF1-related protein kinases: Global regulators of carbon metabolism in plants? Plant Mol. Biol. 2004, 37, 735–748. [Google Scholar] [CrossRef]
- Hrabak, E.M.; Chan, C.W.M.; Gribskov, M.; Harper, J.F.; Choi, J.H.; Halford, N.G.; Kudla, J.; Luan, S.; Nimmo, H.G.; Sussman, M.R.; et al. The Arabidopsis CDPK-SnRK superfamily of protein kinases. Plant Physiol. 2003, 132, 666–680. [Google Scholar] [CrossRef] [Green Version]
- Kolukisaoglu, U.; Weinl, S.; Blazevic, D.; Batisti, O.; Kudla, J. Calcium sensors and their interacting protein kinases: Genomics of the Arabidopsis and rice CBL-CIPK signaling networks. Plant Physiol. 2004, 134, 43–58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kimura, S.; Kawarazaki, T.; Nibori, H.; Michikawa, M.; Imai, A.; Kaya, H.; Kuchitsu, K. The CBL-interacting protein kinase CIPK26 is a novel interactor of Arabidopsis NADPH oxidase AtRbohF that negatively modulates its ROS-producing activity in a heterologous expression system. J. Biochem. 2013, 153, 191–195. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.L.; Halfter, U.; Ishitani, M.; Zhu, J.K. Molecular characterization of functional domains in the protein kinase SOS2 that is required for plant salt tolerance. Plant Cell 2001, 13, 1383–1400. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, R.-J.; Wang, C.; Li, K.; Luan, S. The CBL-CIPK calcium signaling network: Unified paradigm from 20 years of discoveries. Trends Plant Sci. 2020, 25, 604–617. [Google Scholar] [CrossRef]
- Johnson, L.N.; Noble, M.E.M.; Owen, D.J. Active and inactive protein kinases: Structural basis for regulation. Cell 1996, 85, 149–158. [Google Scholar] [CrossRef] [Green Version]
- Gong, D.; Guo, Y.; Jagendorf, A.T.; Zhu, J.K. Biochemical characterization of the Arabidopsis protein kinase SOS2 that functions in salt tolerance. Plant Physiol. 2002, 130, 256–264. [Google Scholar] [CrossRef] [Green Version]
- Albrecht, V.; Ritz, O.; Linder, S.; Harter, K.; Kudla, J. The NAF domain defines a novel protein protein interaction module conserved in Ca2+ regulated kinases. EMBO J. 2001, 20, 1051–1063. [Google Scholar] [CrossRef] [Green Version]
- Kim, K.N.; Cheong, Y.H.; Gupta, R.; Luan, S. Interaction specificity of Arabidopsis calcineurin B-like calcium sensors and their target kinases. Plant Physiol. 2000, 124, 1844–1853. [Google Scholar] [CrossRef] [Green Version]
- Ohta, M.; Guo, Y.; Halfter, U.; Zhu, J.K. A novel domain in the protein kinase SOS2 mediates interaction with the protein phosphatase 2C ABI2. Proc. Natl. Acad. Sci. USA 2003, 100, 11771–11776. [Google Scholar] [CrossRef] [Green Version]
- Su, W.H.; Ren, Y.J.; Wang, D.J.; Huang, L.; Fu, X.Q.; Ling, H.; Su, Y.C.; Huang, N.; Tang, H.; Xu, L.; et al. New insights into the evolution and functional divergence of the CIPK gene family in Saccharum. BMC Genom. 2020, 21, 868. [Google Scholar] [CrossRef]
- Cui, Y.P.; Su, Y.; Wang, J.J.; Jia, B.; Wu, M.; Pei, W.F.; Zhang, J.F.; Yu, J.W. Genome-Wide characterization and analysis of CIPK gene family in two cultivated allopolyploid cotton species: Sequence variation, association with seed oil content, and the role of GhCIPK6. Int. J. Mol. Sci. 2020, 21, 863. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheong, Y.H.; Pandey, G.K.; Grant, J.J.; Batistj, O.; Li, L.; Kim, B.G.; Lee, S.C.; Kudla, J.; Luan, S. Two calcineurin B-like calcium sensors, interacting with protein kinase CIPK23, regulate leaf transpiration and root potassium uptake in Arabidopsis. Plant J. 2007, 52, 223–239. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.G.; Waadt, R.; Cheong, Y.H.; Pandey, G.K.; Domínguez-Solís, J.R.; Schültke, S.; Lee, S.C.; Kudla, J.; Luan, S. The calcium sensor CBL10 mediates salt tolerance by regulating ion homeostasis in Arabidopsis. Plant J. 2007, 52, 473–484. [Google Scholar] [CrossRef] [PubMed]
- D’Angelo, C.; Weinl, S.; Batistic, O.; Pandey, G.; Cheong, Y.; Schültke, S.; Albrecht, V.; Ehlert, B.; Schulz, B.; Harter, K.; et al. Alternative complex formation of the Ca-regulated protein kinase CIPK1 controls abscisic acid-dependent and independent stress responses in Arabidopsis. Plant J. 2006, 48, 857–872. [Google Scholar] [CrossRef]
- Batistj, O.; Waadt, R.; Steinhorst, L.; Held, K.; Kudla, J. CBL-Mediated targeting of CIPKs facilitates the decoding of calcium signals emanating from distinct cellular stores. Plant J. 2010, 61, 211–222. [Google Scholar] [CrossRef]
- Ma, X.; Li, Q.H.; Yu, Y.N.; Qiao, Y.M.; Haq, S.U.; Gong, Z.H. The CBl-CIPK pathway in plant response to stress signals. Int. J. Mol. Sci. 2020, 21, 5668. [Google Scholar] [CrossRef]
- Zhang, X.; Li, X.; Zhao, R.; Zhou, Y.; Jiao, Y. Evolutionary strategies drive a balance of the interacting gene products for the CBL and CIPK gene families. New Phytol. 2020, 226, 1506–1516. [Google Scholar] [CrossRef]
- Qiu, Q.S.; Guo, Y.; Dietrich, M.A.; Schumaker, K.S.; Zhu, J.K. Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc. Natl. Acad. Sci. USA 2002, 99, 8436–8441. [Google Scholar] [CrossRef] [Green Version]
- Shi, H.; Quintero, F.J.; Pardo, J.M.; Zhu, J.K. The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 2002, 14, 465–477. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Kim, B.-G.; Cheong, Y.H.; Pandey, G.K.; Luan, S. A Ca2+ signaling pathway regulates a K+ channel for low-K response in Arabidopsis. Proc. Natl. Acad. Sci. USA 2006, 103, 12625–12630. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.N.; Li, H.; Chen, L.Q.; Wang, Y.; Liu, L.L.; He, L.; Wu, W.H. A protein kinase, interacting with two calcineurin B-like proteins, regulates K+ transporter AKT1 in Arabidopsis. Cell 2006, 125, 1347–1360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fuglsang, A.T.; Guo, Y.; Cuin, T.A.; Qiu, Q.S.; Song, C.; Kristiansen, K.A.; Bych, K.; Schulz, A.; Shabala, S.; Schumaker, K.S.; et al. Arabidopsis protein kinase PKS5 Inhibits the plasma membrane H+-ATPase by preventing interaction with 14-3-3 protein. Plant Cell 2007, 19, 1617–1634. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ho, C.H.; Lin, S.; Hu, H.C.A.; Tsay, Y.F. CHL1 functions as a nitrate sensor in plants. Cell 2009, 138, 1184–1194. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Léran, S.; Edel, K.H.; Pervent, M.; Hashimoto, K.; Corratgé-Faillie, C.; Offenborn, J.N.; Tillard, P.; Gojon, A.; Kudla, J.; Lacombe, B. Nitrate sensing and uptake in Arabidopsis are enhanced by ABI2, a phosphatase inactivated by the stress hormone abscisic acid. Sci. Signal. 2015, 8, 43. [Google Scholar] [CrossRef]
- Tang, R.J.; Zhao, F.; Garcia, V.J.; Kleist, T.J.; Yang, L.; Zhang, H.X.; Luan, S. Tonoplast CBL-CIPK calcium signaling network regulates magnesium homeostasis in Arabidopsis. Proc. Natl. Acad. Sci. USA 2015, 112, 3134–3139. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.P.; Chen, L.M.; Liu, W.X.; Shen, L.K.; Wang, F.L.; Zhou, Y.; Zhang, Z.; Wu, W.H.; Wang, Y. AtKC1 and CIPK23 synergistically modulate AKT1-mediated low-potassium stress responses in Arabidopsis. Plant Physiol. 2016, 170, 2264–2277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drerup, M.M.; Schlücking, K.; Hashimoto, K.; Manishankar, P.; Steinhorst, L.; Kuchitsu, K.; Kudla, J. The Calcineurin B-like calcium sensors CBL1 and CBL9 together with their interacting protein kinase CIPK26 regulate the Arabidopsis NADPH oxidase RBOHF. Mol. Plant 2013, 62, 559–569. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.; Wang, C.; Xue, Y.; Liu, X.; Chen, S.; Song, C.; Yang, Y.; Guo, Y. Calcium-activated 14-3-3 proteins as a molecular switch in salt stress tolerance. Nat. Commun. 2019, 10, 1199. [Google Scholar] [CrossRef] [Green Version]
- Han, F.Q.; Zhang, X.L.; Yang, L.M.; Zhuang, M.; Zhang, Y.Y.; Liu, Y.; Li, Z.; Wang, Y.; Fang, Z.; Ji, J.; et al. Genome-Wide characterization and analysis of the anthocyanin biosynthetic genes in Brassica oleracea. Planta 2021, 254, 92. [Google Scholar] [CrossRef]
- Mistry, J.; Finn, R.D.; Eddy, S.R.; Bateman, A.; Punta, M. Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res. 2013, 41, e121. [Google Scholar] [CrossRef] [Green Version]
- Hu, B.H.; Jin, J.; Guo, A.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bailey, T.L.; Bodén, M.; Buske, F.A.; Frith, M.C.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME Suite: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools-an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Li, Q.; Zhao, H.X.; Wang, X.L.; Kang, J.Y.; Lv, B.B.; Dong, Q.X.; Li, C.L.; Chen, H.; Wu, Q. Tartary buckwheat transcription factor FtbZIP5, regulated by FtSnRK2.6, can improve salt/drought resistance in transgenic Arabidopsis. Int. J. Mol. Sci. 2020, 21, 1123. [Google Scholar] [CrossRef] [Green Version]
- Li, Q.; Wu, Q.; Wang, A.H.; Lv, B.B.; Dong, Q.X.; Yao, Y.J.; Wu, Q.; Zhao, H.X.; Li, C.L.; Chen, H.; et al. Tartary buckwheat transcription factor FtbZIP83 improves the drought/salt tolerance of Arabidopsis via an ABA-mediated pathway. Plant Physiol. Biochem. 2019, 144, 312–323. [Google Scholar] [CrossRef]


| Gene Name | Gene ID | CDS | AA | pI | Mw (Da) | Subcellular Localization |
|---|---|---|---|---|---|---|
| SmCIPK1 | SMil_00000703 | 1263 | 420 | 9.29 | 47,270.22 | Cytoplasm |
| SmCIPK2 | SMil_00000704 | 1329 | 442 | 8.81 | 49,427.94 | Cytoplasm |
| SmCIPK3 | SMil_00000853 | 1425 | 474 | 8.56 | 53,056.17 | Cytoplasm |
| SmCIPK4 | SMil_00001122 | 1311 | 436 | 9.28 | 49,667.43 | Cytoplasm |
| SmCIPK5 | SMil_00003013 | 1323 | 440 | 9.02 | 49,944.58 | Cytoplasm Nucleus |
| SmCIPK6 | SMil_00003083 | 1311 | 436 | 9.25 | 49,042.18 | Cytoplasm |
| SmCIPK7 | SMil_00003987 | 1128 | 375 | 5.99 | 42,844.14 | Cytoplasm |
| SmCIPK8 | SMil_00004681 | 1425 | 474 | 8.88 | 53,641.64 | Cytoplasm Nucleus |
| SmCIPK9 | SMil_00008243 | 1380 | 459 | 8.97 | 51,920.95 | Centrosome Cytoplasm Nucleus |
| SmCIPK10 | SMil_00009044 | 1251 | 416 | 9.07 | 46,960.19 | Cytoplasm |
| SmCIPK11 | SMil_00010169 | 1419 | 472 | 5.69 | 54,055.99 | Centrosome Cytoplasm Nucleus |
| SmCIPK12 | SMil_00012715 | 1167 | 388 | 8.15 | 43,862.71 | Cytoplasm |
| SmCIPK13 | SMil_00014175 | 1368 | 455 | 9.14 | 51,091.17 | Cytoplasm |
| SmCIPK14 | SMil_00021346 | 1236 | 411 | 6.51 | 46,210.19 | Cytoplasm |
| SmCIPK15 | SMil_00025018 | 1176 | 391 | 5.52 | 44,490.61 | Cytoplasm |
| SmCIPK16 | SMil_00025262 | 1524 | 507 | 9.56 | 57,920.08 | Centrosome Cytoplasm Nucleus |
| SmCIPK17 | SMil_00026858 | 1380 | 459 | 6.01 | 51,419.66 | Cytoplasm |
| SmCIPK18 | SMil_00026916 | 1263 | 193 | 6.21 | 47,926.96 | Cytoplasm |
| SmCIPK19 | SMil_00027660 | 1314 | 437 | 9.04 | 49,288.96 | Cytoplasm |
| SmCIPK20 | SMil_00028555 | 1257 | 418 | 8.92 | 47,519.09 | Cytoplasm |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, S.; Li, Q. Genome-Wide Identification of the Salvia miltiorrhiza SmCIPK Gene Family and Revealing the Salt Resistance Characteristic of SmCIPK13. Int. J. Mol. Sci. 2022, 23, 6861. https://doi.org/10.3390/ijms23126861
Wang S, Li Q. Genome-Wide Identification of the Salvia miltiorrhiza SmCIPK Gene Family and Revealing the Salt Resistance Characteristic of SmCIPK13. International Journal of Molecular Sciences. 2022; 23(12):6861. https://doi.org/10.3390/ijms23126861
Chicago/Turabian StyleWang, Shuang, and Qi Li. 2022. "Genome-Wide Identification of the Salvia miltiorrhiza SmCIPK Gene Family and Revealing the Salt Resistance Characteristic of SmCIPK13" International Journal of Molecular Sciences 23, no. 12: 6861. https://doi.org/10.3390/ijms23126861
APA StyleWang, S., & Li, Q. (2022). Genome-Wide Identification of the Salvia miltiorrhiza SmCIPK Gene Family and Revealing the Salt Resistance Characteristic of SmCIPK13. International Journal of Molecular Sciences, 23(12), 6861. https://doi.org/10.3390/ijms23126861






