Identification and Fine Mapping of the Candidate Gene Controlling Multi-Inflorescence in Brassica napus
Abstract
:1. Introduction
2. Results
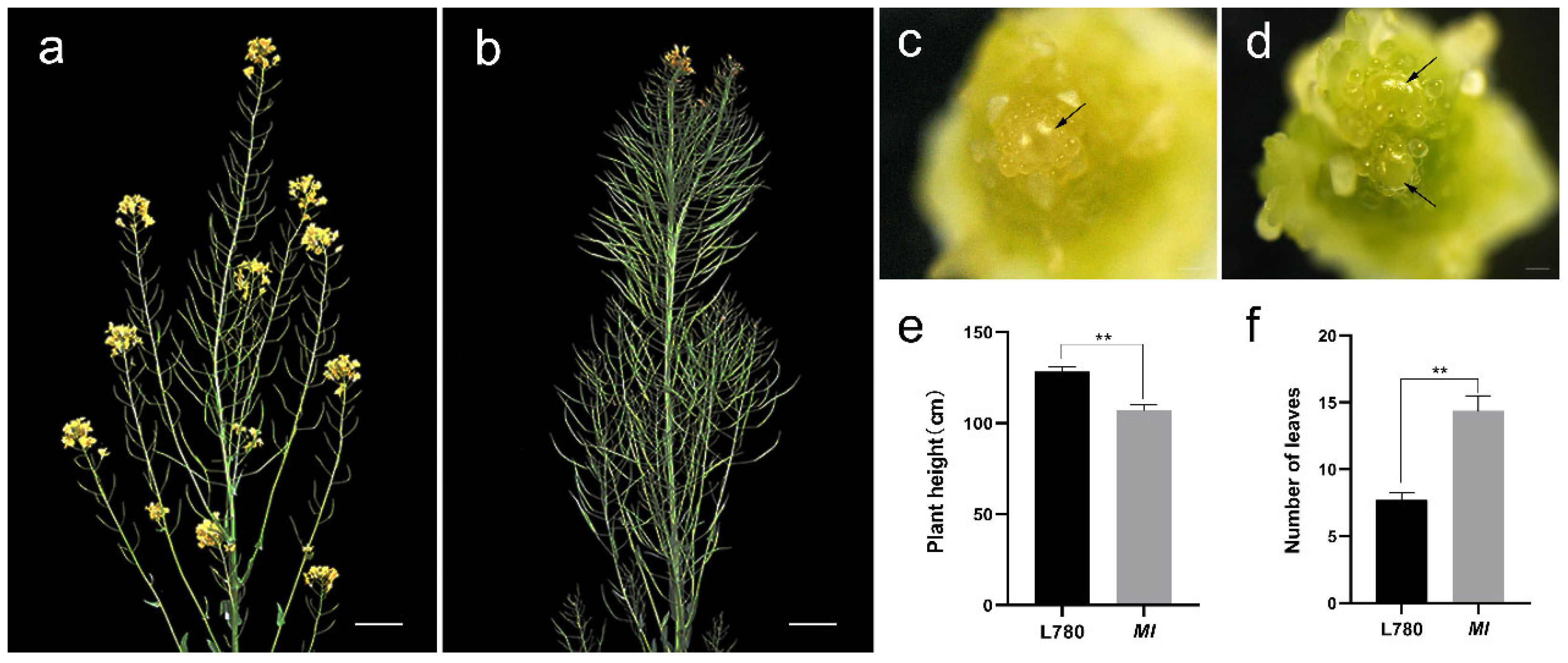
2.1. Morphological of Multi-Inflorescence and Possibility of Yield Increase
2.2. Fine Mapping of BnaMI Gene
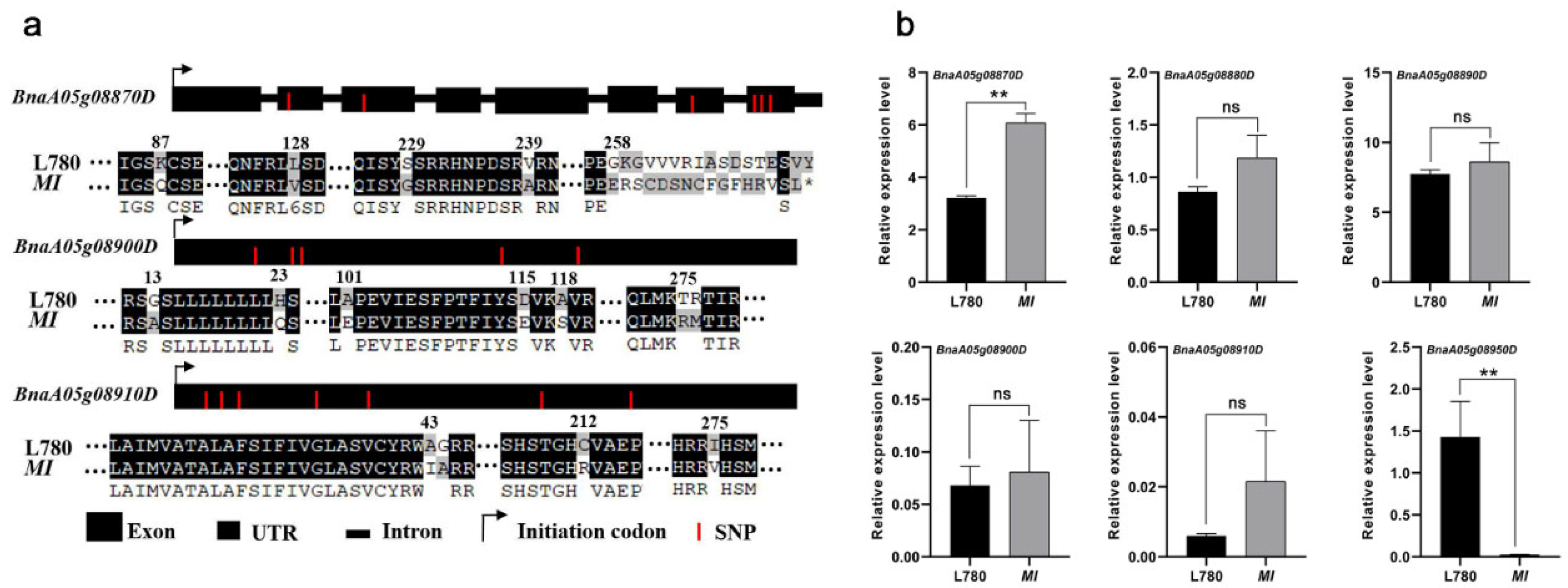
2.3. Candidate Gene Analysis in the Mapping Region
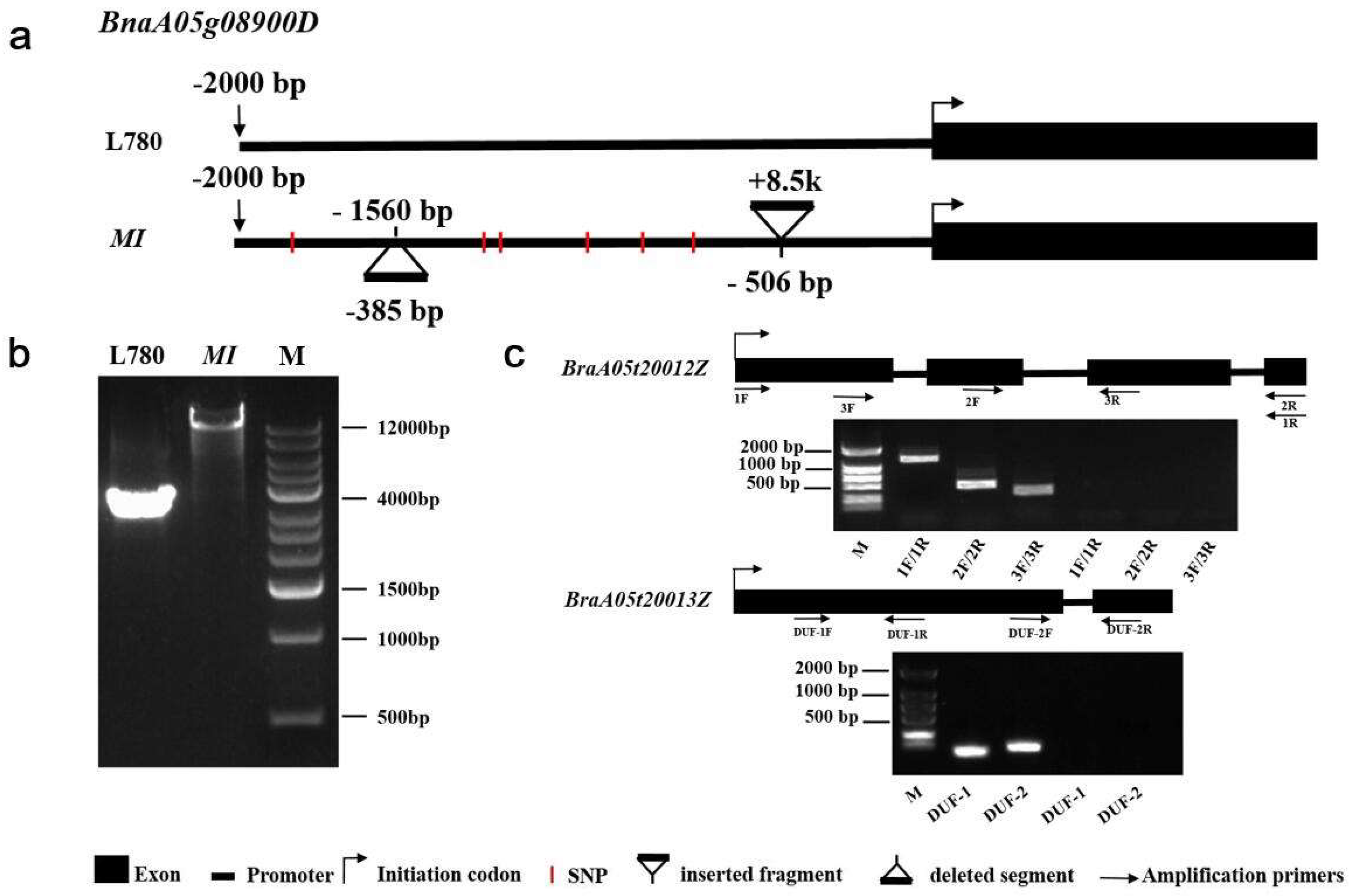
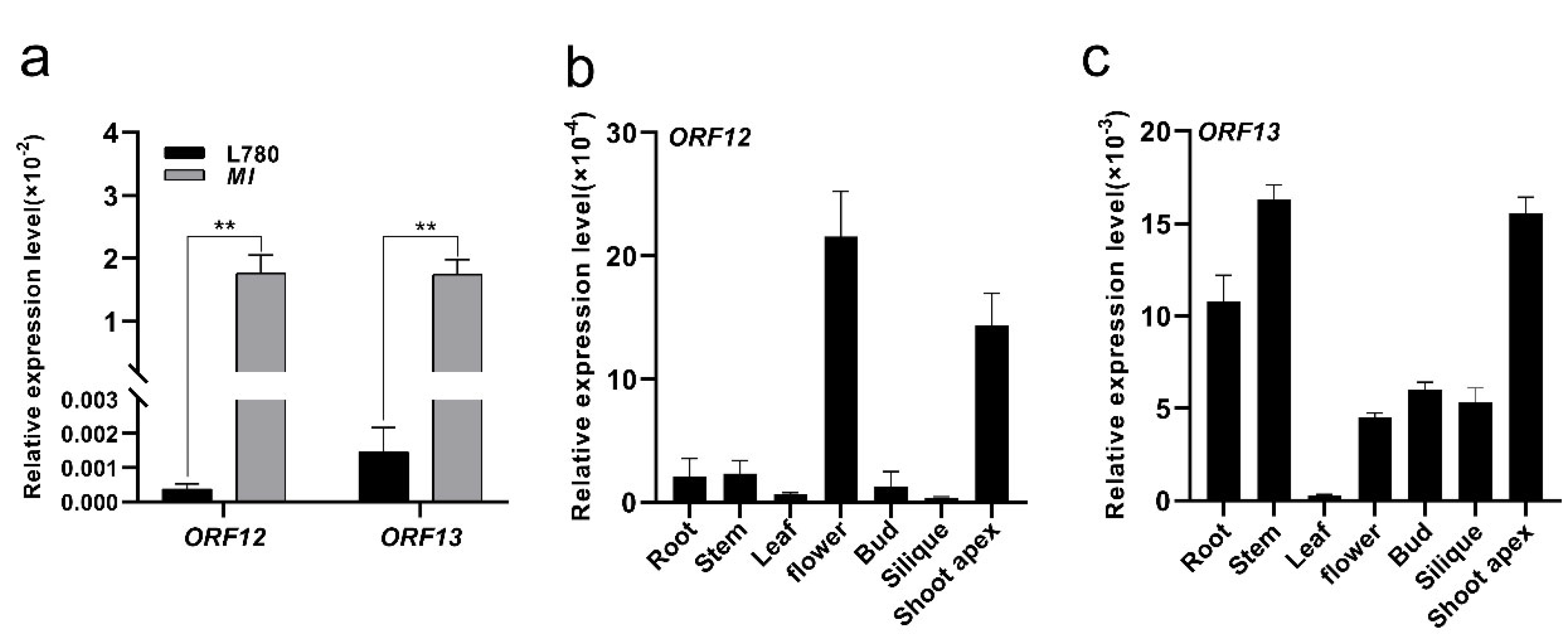
2.4. An Identified 8.5 kb Insertion by Comparative Sequencing
3. Discussion
4. Materials and Methods
4.1. Plant Materials, Population Development, and Trait Evaluation
4.2. Histological Analysis
4.3. Genomic DNA Extraction and Marker Development
4.4. Sequencing and functions prediction of candidate genes
4.5. RNA Extraction and Quantitative Real Time PCR (qRT−PCR)
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hu, Q.; Hua, W.; Yin, Y.; Zhang, X.K.; Liu, L.J.; Shi, J.Q.; Zhao, Y.G.; Qin, L.; Chen, C.; Wang, H.Z. Rapeseed research and production in China. Crop J. 2017, 5, 127–135. [Google Scholar] [CrossRef] [Green Version]
- Fattahi, F.; Fakheri, B.A.; Solouki, M.; Mollers, C.; Rezaizad, A. Mapping QTL controlling agronomic traits in a doubled haploid population of winter oilseed rape (Brassica napus L.). J. Genet. 2018, 97, 1389–1406. [Google Scholar] [CrossRef]
- Khush, G.S. Green revolution: The way forward. Nat. Rev. Genet. 2001, 2, 815–822. [Google Scholar] [CrossRef]
- Fu, D.; Zhou, Y. Progress and future development of hybrid rapeseed in China. Eng. Sci. 2013, 5, 13–18. [Google Scholar]
- Fu, T. Proceeding of oil crops scientific technological advantage. Chin. J. Oil Crop Sci. 2008, 30, 1–5. [Google Scholar]
- Reinhardt, D.; Kuhlemeier, C. Plant architecture. EMBO Rep. 2002, 3, 846–851. [Google Scholar] [CrossRef]
- Wang, B.; Smith, S.M.; Li, J. Genetic Regulation of Shoot Architecture. Annu. Rev. Plant Biol. 2018, 69, 437–468. [Google Scholar] [CrossRef]
- Chatfield, S.P.; Stirnberg, P.; Forde, B.G.; Leyser, O. The hormonal regulation of axillary bud growth in Arabidopsis. Plant J. 2000, 24, 159–169. [Google Scholar] [CrossRef]
- Chickarmane, V.S.; Gordon, S.P.; Tarr, P.T.; Heisler, M.G.; Meyerowitz, E.M. Cytokinin signaling as a positional cue for patterning the apical-basal axis of the growing Arabidopsis shoot meristem. Proc. Natl. Acad. Sci. USA 2012, 109, 4002–4007. [Google Scholar] [CrossRef] [Green Version]
- Davis, E.L.; Rennie, P.; Steeves, T.A. Further Analytical and Experimental Studies on the Shoot Apex of Helianthus-Annuus—Variable Activity in the Central Zone. Can. J. Bot.-Rev. Can. Bot. 1979, 57, 971–980. [Google Scholar] [CrossRef]
- Lenhard, M.; Laux, T. Shoot meristem formation and maintenance. Curr. Opin. Plant Biol. 1999, 2, 44–50. [Google Scholar] [CrossRef]
- Fletcher, J.C. The CLV-WUS Stem Cell Signaling Pathway: A Roadmap to Crop Yield Optimization. Plants 2018, 7, 87. [Google Scholar] [CrossRef] [Green Version]
- Somssich, M.; Je, B.I.; Simon, R.; Jackson, D. CLAVATA-WUSCHEL signaling in the shoot meristem. Development 2016, 143, 3238–3248. [Google Scholar] [CrossRef] [Green Version]
- Rodríguez-Leal, D.; Lemmon, Z.H.; Man, J.; Bartlett, M.E.; Lippman, Z.B. Engineering quantitative trait variation for crop improvement by genome editing. Cell 2017, 171, 470–480.e8. [Google Scholar] [CrossRef] [Green Version]
- Yin, X.-J.; Volk, S.; Ljung, K.; Mehlmer, N.; Dolezal, K.; Ditengou, F.; Hanano, S.; Davis, S.J.; Schmelzer, E.; Sandberg, G.; et al. Ubiquitin lysine 63 chain–forming ligases regulate apical dominance in Arabidopsis. Plant Cell 2007, 19, 1898–1911. [Google Scholar] [CrossRef] [Green Version]
- Laufs, P.; Dockx, J.; Kronenberger, J.; Traas, J. MGOUN1 and MGOUN2: Two genes required for primordium initiation at the shoot apical and floral meristems in Arabidopsis thaliana. Development 1998, 125, 1253–1260. [Google Scholar] [CrossRef]
- Xu, L.; Shen, W.H. Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr. Biol. 2008, 18, 1966–1971. [Google Scholar] [CrossRef] [Green Version]
- Cai, G.; Yang, Q.; Chen, H.; Yang, Q.; Zhang, C.; Fan, C.; Zhou, Y. Genetic dissection of plant architecture and yield-related traits in Brassica napus. Sci. Rep. 2016, 6, 21625. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Li, J.; Song, J.; Zhao, B.; Guo, C.; Wang, B.; Zhang, Q.; Wang, J.; King, G.J.; Liu, K. An auxin signaling gene BnaA3. IAA 7 contributes to improved plant architecture and yield heterosis in rapeseed. New Phytol. 2019, 222, 837–851. [Google Scholar] [CrossRef]
- Li, B.; Gao, J.; Chen, J.; Wang, Z.; Shen, W.; Yi, B.; Wen, J.; Ma, C.; Shen, J.; Fu, T.; et al. Identification and fine mapping of a major locus controlling branching in Brassica napus. Theor. Appl. Genet. 2020, 133, 771–783. [Google Scholar] [CrossRef]
- Li, B.; Wang, T.; Guo, Y.; Liu, X.; Deng, L.; Qu, L.; Li, M. Fine mapping of qDB.A03, a QTL for rapeseed branching, and identification of the candidate gene. Mol. Genet. Genom. 2022, 297, 699–710. [Google Scholar] [CrossRef]
- Ye, S.; Yan, L.; Ma, X.; Chen, Y.; Wu, L.; Ma, T.; Zhao, L.; Yi, B.; Ma, C.; Tu, J.; et al. Combined BSA-Seq Based Mapping and RNA-Seq Profiling Reveal Candidate Genes Associated with Plant Architecture in Brassica napus. Int. J. Mol. Sci. 2022, 23, 2472. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Q.; Cui, Y.; Liu, Z.; Chen, Z.; He, Y.; Mei, J.; Xiong, Q.; Li, X.; Qian, W. Genetic characterization and fine mapping for multi-inflorescence in Brassica napus L. Theor. Appl. Genet. 2018, 131, 2311–2319. [Google Scholar] [CrossRef]
- Diepenbrock, W. Yield analysis of winter oilseed rape (Brassica napus L.): A review. Field Crops Res. 2000, 67, 35–49. [Google Scholar] [CrossRef]
- Zhao, W.; Chao, H.; Zhang, L.; Ta, N.; Zhao, Y.; Li, B.; Zhang, K.; Guan, Z.; Hou, D.; Chen, K.; et al. Integration of QTL Mapping and Gene Fishing Techniques to Dissect the Multi-Main Stem Trait in Rapeseed (Brassica napus L.). Front. Plant Sci. 2019, 10, 1152. [Google Scholar] [CrossRef]
- Zhu, K.; Zhang, W.; Sarwa, R.; Xu, S.; Li, K.; Yang, Y.; Li, Y.; Wang, Z.; Cao, J.; Li, Y.; et al. Proteomic analysis of a clavata-like phenotype mutant in Brassica napus. Genet. Mol. Biol. 2020, 43, e20190305. [Google Scholar] [CrossRef]
- Zhu, K.M.; Xu, S.; Li, K.X.; Chen, S.; Zafar, S.; Cao, W.; Wang, Z.; Ding, L.N.; Yang, Y.H.; Li, Y.M.; et al. Transcriptome analysis of the irregular shape of shoot apical meristem in dt (dou tou) mutant of Brassica napus L. Mol. Breed. 2019, 39, 39. [Google Scholar] [CrossRef] [Green Version]
- Lu, H.; Wu, H.; Wen, J.; Yi, B.; Ma, C.; Tu, J.; Fu, T.; Shen, J. Phenotypic characteristics and genetic analysis of multi-inflorescence trait in Brassica napus. Chin. J. Oil Crop Sci. 2019, 41, 850. [Google Scholar]
- Qin, M.; Yang, Y.; Cao, F.; Yang, C.; Li, C.; Cheng, H.; Yu, Y.; Li, D.; Zheng, H.; Peng, X.; et al. Impact of OsUreD Mutation on Rice Growth Phenotype and Nitrogen Nutritional Physiology. J. Agric. Sci. Technol. 2017, 19, 9. [Google Scholar]
- Sonoda, Y.; Yao, S.G.; Sako, K.; Sato, T.; Kato, W.; Ohto, M.A.; Ichikawa, T.; Matsui, M.; Yamaguchi, J.; Ikeda, A. SHA1, a novel RING finger protein, functions in shoot apical meristem maintenance in Arabidopsis. Plant J. 2007, 50, 586–596. [Google Scholar] [CrossRef]
- Rieu, I.; Laux, T. Signaling pathways maintaining stem cells at the plant shoot apex. Semin. Cell Dev. Biol. 2009, 20, 1083–1088. [Google Scholar] [CrossRef]
- Carles, C.C.; Fletcher, J.C. Shoot apical meristem maintenance: The art of a dynamic balance. Trends Plant Sci. 2003, 8, 394–401. [Google Scholar] [CrossRef]
- Clark, S.E.; Williams, R.W.; Meyerowitz, E.M. The CLAVATA1gene encodes a putative receptor kinase that controls shoot and floral meristem size in Arabidopsis. Cell 1997, 89, 575–585. [Google Scholar] [CrossRef] [Green Version]
- Jeong, S.; Trotochaud, A.E.; Clark, S.E. The Arabidopsis CLAVATA2 gene encodes a receptor-like protein required for the stability of the CLAVATA1 receptor-like kinase. Plant Cell 1999, 11, 1925–1934. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Y.; Wagner, D. Plant Inflorescence Architecture: The Formation, Activity, and Fate of Axillary Meristems. Cold Spring Harb. Perspect. Biol. 2020, 12, a034652. [Google Scholar] [CrossRef]
- Lee, Z.H.; Hirakawa, T.; Yamaguchi, N.; Ito, T. The roles of plant hormones and their interactions with regulatory genes in determining meristem activity. Int. J. Mol. Sci. 2019, 20, 4065. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kurakawa, T.; Ueda, N.; Maekawa, M.; Kobayashi, K.; Kojima, M.; Nagato, Y.; Sakakibara, H.; Kyozuka, J.J.N. Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature 2007, 445, 652–655. [Google Scholar] [CrossRef]
- Landrein, B.; Formosa-Jordan, P.; Malivert, A.; Schuster, C.; Melnyk, C.W.; Yang, W.B.; Turnbull, C.; Meyerowitz, E.M.; Locke, J.C.W.; Jonsson, H. Nitrate modulates stem cell dynamics in Arabidopsis shoot meristems through cytokinins. Proc. Natl. Acad. Sci. USA 2018, 115, 1382–1387. [Google Scholar] [CrossRef] [Green Version]
- Vidaurre, D.P.; Ploense, S.; Krogan, N.T.; Berleth, T. AMP1 and MP antagonistically regulate embryo and meristem development in Arabidopsis. Development 2007, 134, 2561–2567. [Google Scholar] [CrossRef] [Green Version]
- Denay, G.; Chahtane, H.; Tichtinsky, G.; Parcy, F. A flower is born: An update on Arabidopsis floral meristem formation. Curr. Opin. Plant Biol. 2017, 35, 15–22. [Google Scholar] [CrossRef]
- Weigel, D.; Alvarez, J.; Smyth, D.R.; Yanofsky, M.F.; Meyerowitz, E.M. LEAFY controls floral meristem identity in Arabidopsis. Cell 1992, 69, 843–859. [Google Scholar] [CrossRef] [Green Version]
- Li, W.; Zhou, Y.; Liu, X.; Yu, P.; Cohen, J.D.; Meyerowitz, E.M. LEAFY controls auxin response pathways in floral primordium formation. Sci. Signal. 2013, 6, ra23. [Google Scholar] [CrossRef] [Green Version]
- Yamaguchi, N.; Wu, M.F.; Winter, C.M.; Berns, M.C.; Nole-Wilson, S.; Yamaguchi, A.; Coupland, G.; Krizek, B.A.; Wagner, D. A molecular framework for auxin-mediated initiation of flower primordia. Dev. Cell 2013, 24, 271–282. [Google Scholar] [CrossRef] [Green Version]
- Xiao, H.; Tang, J.; Li, Y.; Wang, W.; Li, X.; Jin, L.; Xie, R.; Luo, H.; Zhao, X.; Meng, Z.; et al. STAMENLESS 1, encoding a single C2H2 zinc finger protein, regulates floral organ identity in rice. Plant J. 2009, 59, 789–801. [Google Scholar] [CrossRef]
- Witte, C.P.; Rosso, M.G.; Romeis, T. Identification of three urease accessory proteins that are required for urease activation in Arabidopsis. Plant Physiol. 2005, 139, 1155–1162. [Google Scholar] [CrossRef] [Green Version]
- Bennetzen, J.L. Transposable element contributions to plant gene and genome evolution. Plant Mol. Biol. 2000, 42, 251–269. [Google Scholar] [CrossRef]
- Studer, A.; Zhao, Q.; Ross-Ibarra, J.; Doebley, J. Identification of a functional transposon insertion in the maize domestication gene tb1. Nat. Genet. 2011, 43, 1160–1163. [Google Scholar] [CrossRef] [Green Version]
- Xiao, H.; Jiang, N.; Schaffner, E.; Stockinger, E.J.; van der Knaap, E. A retrotransposon-mediated gene duplication underlies morphological variation of tomato fruit. Science 2008, 319, 1527–1530. [Google Scholar] [CrossRef]
- Song, X.; Cao, X. Transposon-mediated epigenetic regulation contributes to phenotypic diversity and environmental adaptation in rice. Curr. Opin. Plant Biol. 2017, 36, 111–118. [Google Scholar] [CrossRef]
- Jia, H.; Li, M.; Li, W.; Liu, L.; Jian, Y.; Yang, Z.; Shen, X.; Ning, Q.; Du, Y.; Zhao, R. A serine/threonine protein kinase encoding gene KERNEL NUMBER PER ROW6 regulates maize grain yield. Nat. Commun. 2020, 11, 988. [Google Scholar] [CrossRef] [Green Version]
- Shi, L.; Song, J.; Guo, C.; Wang, B.; Guan, Z.; Yang, P.; Chen, X.; Zhang, Q.; King, G.J.; Wang, J.; et al. A CACTA-like transposable element in the upstream region of BnaA9.CYP78A9 acts as an enhancer to increase silique length and seed weight in rapeseed. Plant J. 2019, 98, 524–539. [Google Scholar] [CrossRef]
- Song, J.M.; Guan, Z.; Hu, J.; Guo, C.; Yang, Z.; Wang, S.; Liu, D.; Wang, B.; Lu, S.; Zhou, R.; et al. Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat. Plants 2020, 6, 34–45. [Google Scholar] [CrossRef]
- Rousseau-Gueutin, M.; Belser, C.; Da Silva, C.; Richard, G.; Istace, B.; Cruaud, C.; Falentin, C.; Boideau, F.; Boutte, J.; Delourme, R.; et al. Long-read assembly of the Brassica napus reference genome Darmor-bzh. Gigascience 2020, 9, giaa137. [Google Scholar] [CrossRef]
- Tang, S.; Zhao, H.; Lu, S.P.; Yu, L.Q.; Zhang, G.F.; Zhang, Y.T.; Yang, Q.Y.; Zhou, Y.M.; Wang, X.M.; Ma, W.; et al. Genome- and transcriptome-wide association studies provide insights into the genetic basis of natural variation of seed oil content in Brassica napus. Mol. Plant 2021, 14, 470–487. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, D.; Yu, H.; Lin, B.; Fu, Y.; Hua, S. Floral Initiation in Response to Planting Date Reveals the Key Role of Floral Meristem Differentiation Prior to Budding in Canola (Brassica napus L.). Front. Plant Sci. 2016, 7, 1369. [Google Scholar] [CrossRef] [Green Version]
- Doyle, J. DNA protocols for plants. In Molecular Techniques in Taxonomy; Springer: Berlin/Heidelberg, Germany, 1991; pp. 283–293. [Google Scholar]
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.; Tang, H.; Wang, X.; Chiquet, J.; Belcram, H.; Tong, C.; Samans, B.; et al. Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 2014, 345, 950–953. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]


| Gene Number | Arabidopsis Homologous Gene Number | Annotations Information |
|---|---|---|
| BnaA05g08870D | AT2G35035.1 | urease accessory protein D |
| BnaA05g08880D | AT2G35030.1 | Pentatricopeptide repeat (PPR) superfamily protein |
| BnaA05g08890D | AT2G35020.1 | N-acetylglucosamine-1-phosphate uridylyltransferase 2 |
| BnaA05g08900D | AT2G35000.1 | RING/U-box superfamily protein |
| BnaA05g08910D | AT2G34990.1 | RING/U-box superfamily protein |
| BnaA05g08950D | AT2G34930.1 | disease resistance family protein/LRR family protein |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, H.; Wu, H.; Zhu, G.; Yin, C.; Zhao, L.; Wen, J.; Yi, B.; Ma, C.; Tu, J.; Fu, T.; et al. Identification and Fine Mapping of the Candidate Gene Controlling Multi-Inflorescence in Brassica napus. Int. J. Mol. Sci. 2022, 23, 7244. https://doi.org/10.3390/ijms23137244
Lu H, Wu H, Zhu G, Yin C, Zhao L, Wen J, Yi B, Ma C, Tu J, Fu T, et al. Identification and Fine Mapping of the Candidate Gene Controlling Multi-Inflorescence in Brassica napus. International Journal of Molecular Sciences. 2022; 23(13):7244. https://doi.org/10.3390/ijms23137244
Chicago/Turabian StyleLu, Hongchen, Hanfei Wu, Guangfeng Zhu, Caijun Yin, Lun Zhao, Jing Wen, Bin Yi, Chaozhi Ma, Jinxing Tu, Tingdong Fu, and et al. 2022. "Identification and Fine Mapping of the Candidate Gene Controlling Multi-Inflorescence in Brassica napus" International Journal of Molecular Sciences 23, no. 13: 7244. https://doi.org/10.3390/ijms23137244






