Transcriptional Analysis on Resistant and Susceptible Kiwifruit Genotypes Activating Different Plant-Immunity Processes against Pseudomonas syringae pv. actinidiae
Abstract
:1. Introduction
2. Results
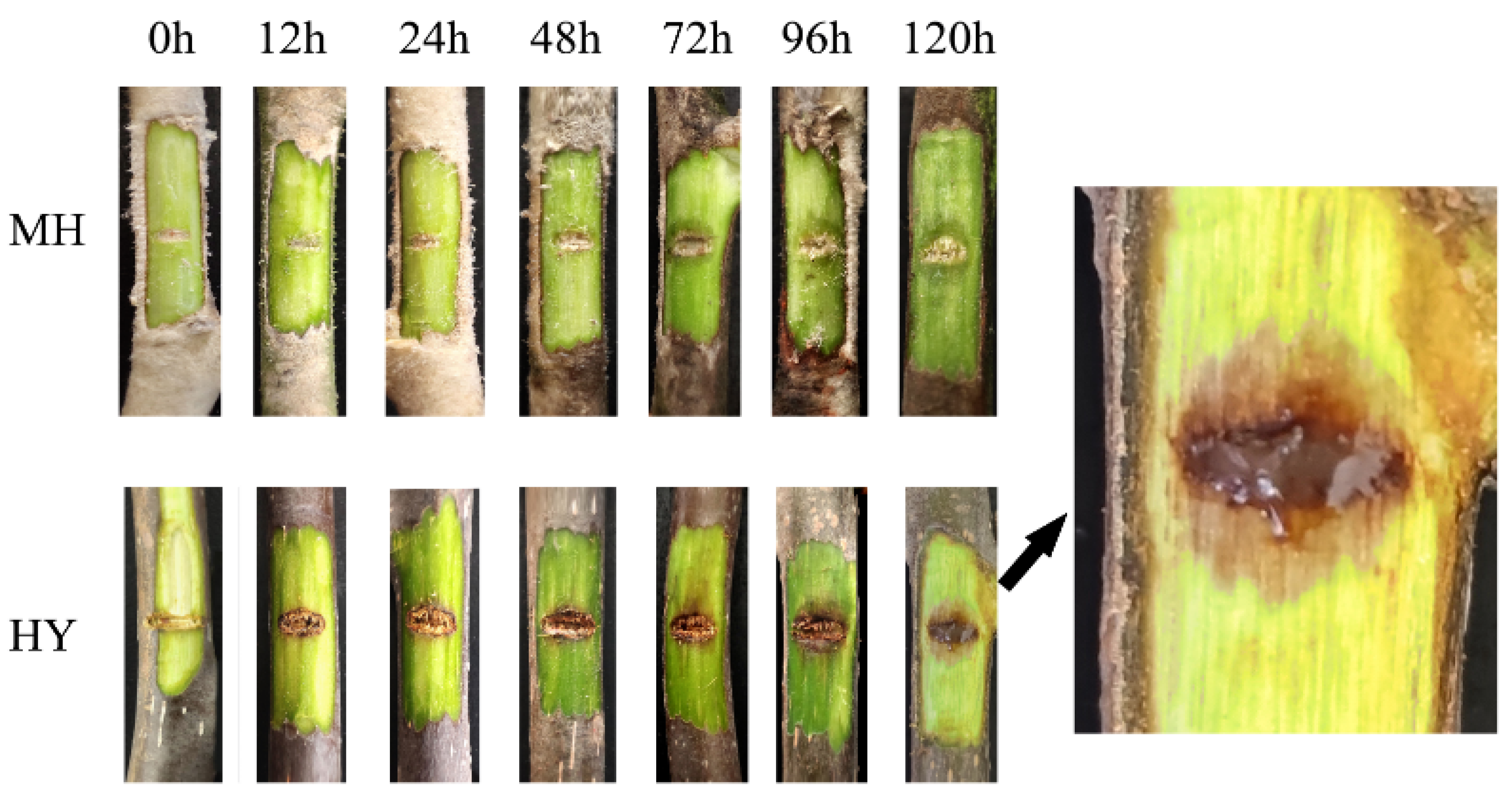
2.1. Symptoms of MH and HY Shoots Infected with Psa
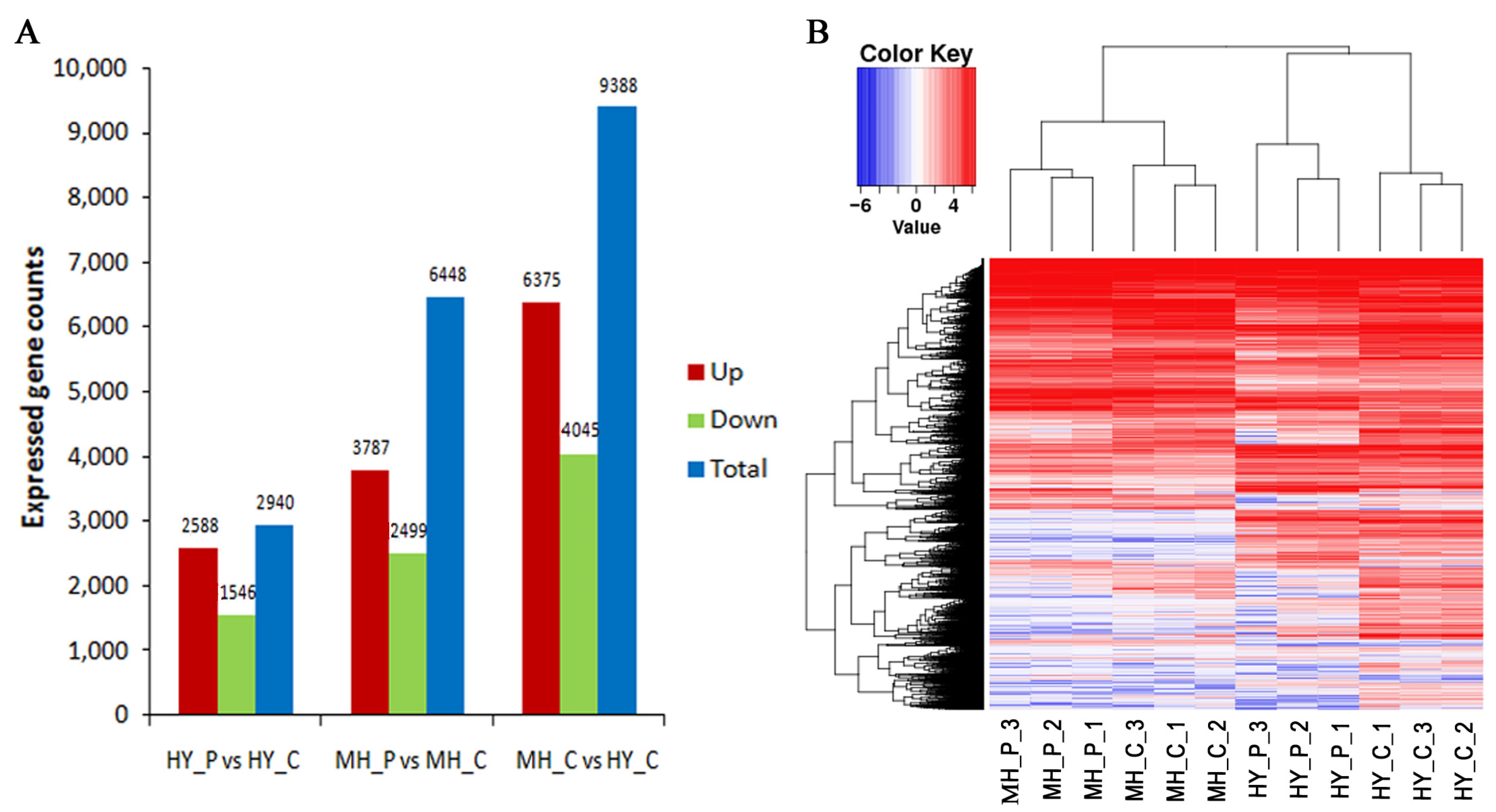
2.2. RNA-Seq Data Analysis and DEGs in Response to Psa Infection
2.3. Functional Classification of DEGs after Psa Inoculation
2.4. Pathway-Enrichment Analysis of DEGs after Psa Inoculation
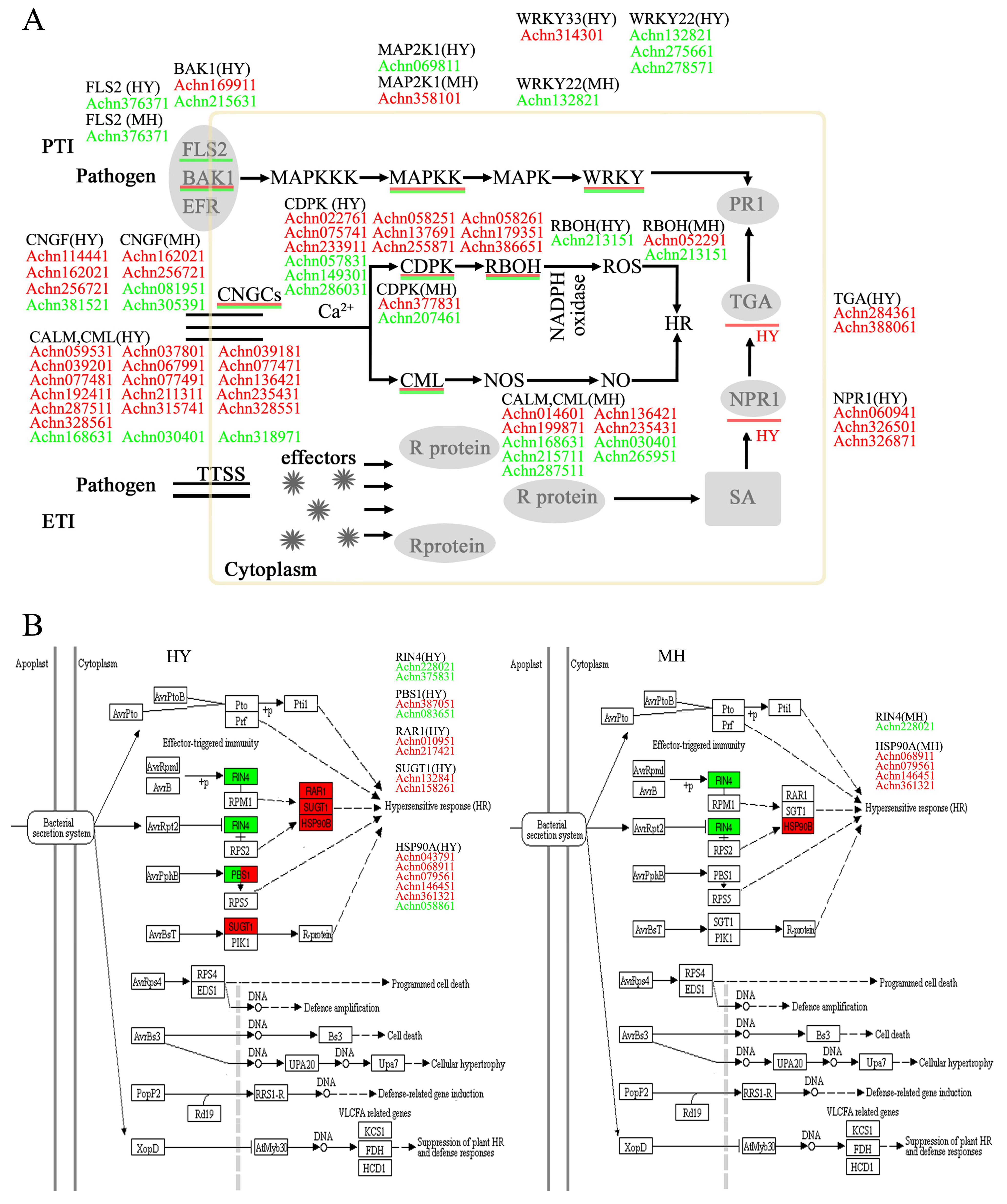
2.5. Analysis of Genes Related to Plant Innate Immune System
2.6. Regulation of Ca2+ Internal Flow and Reactive-Oxygen Species
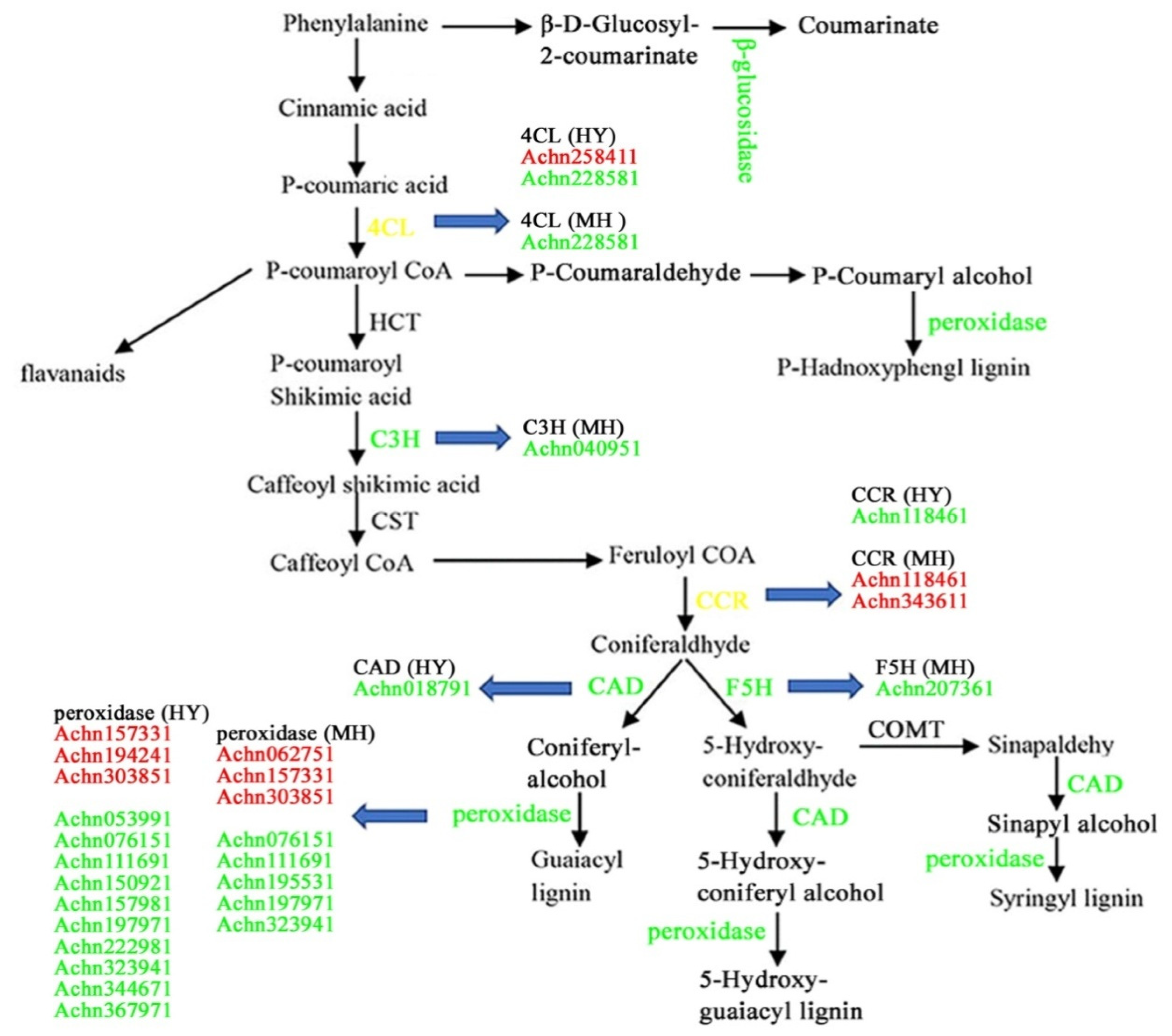
2.7. Analysis of DEGs Involved in Secondary Metabolism
2.8. Verification of Gene Expression by q-PCR
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Psa Inoculation
4.2. Total RNA Extraction, cDNA Library Construction, and Sequencing
4.3. Sequencing Read Mapping and Identification of DEGs
4.4. Gene Ontology and KEGG Pathway Analysis
4.5. Quantitative Real-Time PCR Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Serizawa, S.; Ichikawa, T.; Takikawa, Y.; Tsuyumu, S.; Goto, M. Occurrence of Bacterial Canker of Kiwifruit in Japan: Description of Symptoms, Isolation of the Pathogen and Screening of Bactericides. Ann. Phytopathol. Soc. Jpn. 1989, 55, 427–436. [Google Scholar] [CrossRef] [Green Version]
- Green, S.; Studholme, D.J.; Laue, B.E.; Dorati, F.; Lovell, H.; Arnold, D.; Cottrell, J.E.; Bridgett, S.; Blaxter, M.; Huitema, E.; et al. Comparative Genome Analysis Provides Insights into the Evolution and Adaptation of Pseudomonas syringae Pv. aesculi on Aesculus hippocastanum. PLoS ONE 2010, 5, e10224. [Google Scholar] [CrossRef] [PubMed]
- Carniel, E. The Yersinia High-Pathogenicity Island: An Iron-Uptake Island. Microbes Infect. 2001, 3, 561–569. [Google Scholar] [CrossRef]
- Raymond, K.N.; Dertz, E.A.; Kim, S.S. Enterobactin: An Archetype for Microbial Iron Transport. Proc. Natl. Acad. Sci. USA 2003, 100, 3584–3588. [Google Scholar] [CrossRef] [Green Version]
- Vanneste, J.L.; Yu, J.; Cornish, D.A.; Tanner, D.J.; Windner, R.; Chapman, J.R.; Taylor, R.K.; Mackay, J.F.; Dowlut, S. Identification, Virulence, and Distribution of Two Biovars of Pseudomonas syringae Pv. actinidiae in New Zealand. Plant Dis. 2013, 97, 708–719. [Google Scholar] [CrossRef] [Green Version]
- Chapman, J.R.; Taylor, R.K.; Weir, B.S.; Romberg, M.K.; Vanneste, J.L.; Luck, J.; Alexander, B.J.R. Phylogenetic Relationships among Global Populations of Pseudomonas syringae Pv. actinidiae. Phytopathology 2012, 102, 1034–1044. [Google Scholar] [CrossRef] [Green Version]
- Donati, I.; Cellini, A.; Buriani, G.; Mauri, S.; Kay, C.; Tacconi, G.; Spinelli, F. Pathways of Flower Infection and Pollen-Mediated Dispersion of Pseudomonas syringae Pv. actinidiae, the Causal Agent of Kiwifruit Bacterial Canker. Hortic. Res. 2018, 5, 56. [Google Scholar] [CrossRef] [Green Version]
- Vanneste, J.L. The Scientific, Economic, and Social Impacts of the New Zealand Outbreak of Bacterial Canker of Kiwifruit (Pseudomonas syringae Pv. actinidiae). Annu. Rev. Phytopathol. 2017, 55, 399. [Google Scholar] [CrossRef]
- Xin, X.-F.; Kvitko, B.; He, S.Y. Pseudomonas syringae: What It Takes to Be a Pathogen. Nat. Rev. Microbiol. 2018, 16, 316–328. [Google Scholar] [CrossRef]
- Everett, K.R.; Taylor, R.K.; Romberg, M.K.; Rees-George, J.; Fullerton, R.A.; Vanneste, J.L.; Manning, M.A. First Report of Pseudomonas Syringae Pv. Actinidiae Causing Kiwifruit Bacterial Canker in New Zealand. Australas. Plant Dis. Notes 2011, 6, 67–71. [Google Scholar] [CrossRef] [Green Version]
- Takikawa, Y.; Serizawa, S.; Ichikawa, T.; Tsuyumu, S.; Goto, M. Pseudomonas syringae Pv. actinidiae Pv. Nov.: The Causal Bacterium of Canker of Kiwifruit in Japan. Ann. Phytopathol. Soc. Jpn. 1989, 55, 437–444. [Google Scholar] [CrossRef] [Green Version]
- Gao, X.; Huang, Q.; Zhao, Z.; Han, Q.; Ke, X.; Qin, H.; Huang, L. Studies on the Infection, Colonization, and Movement of Pseudomonas syringae Pv. actinidiae in Kiwifruit Tissues Using a GFPuv-Labeled Strain. PLoS ONE 2016, 11, e0151169. [Google Scholar] [CrossRef] [PubMed]
- Thomma, B.P.H.J.; Nürnberger, T.; Joosten, M.H.A.J. Of PAMPs and Effectors: The Blurred PTI-ETI Dichotomy. Plant Cell 2011, 23, 4–15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Návarová, H.; Bernsdorff, F.; Döring, A.-C.; Zeier, J. Pipecolic Acid, an Endogenous Mediator of Defense Amplification and Priming, Is a Critical Regulator of Inducible Plant Immunity. Plant Cell 2012, 24, 5123–5141. [Google Scholar] [CrossRef] [Green Version]
- Bigeard, J.; Colcombet, J.; Hirt, H. Signaling Mechanisms in Pattern-Triggered Immunity (PTI). Mol. Plant 2015, 8, 521–539. [Google Scholar] [CrossRef] [Green Version]
- Trouvelot, S.; Héloir, M.-C.; Poinssot, B.; Gauthier, A.; Paris, F.; Guillier, C.; Combier, M.; Trdá, L.; Daire, X.; Adrian, M. Carbohydrates in Plant Immunity and Plant Protection: Roles and Potential Application as Foliar Sprays. Front. Plant Sci. 2014, 5, 592. [Google Scholar] [CrossRef] [Green Version]
- Fu, Z.Q.; Yan, S.; Saleh, A.; Wang, W.; Ruble, J.; Oka, N.; Mohan, R.; Spoel, S.H.; Tada, Y.; Zheng, N.; et al. NPR3 and NPR4 Are Receptors for the Immune Signal Salicylic Acid in Plants. Nature 2012, 486, 228–232. [Google Scholar] [CrossRef] [Green Version]
- Hartmann, M.; Kim, D.; Bernsdorff, F.; Ajami-Rashidi, Z.; Scholten, N.; Schreiber, S.; Zeier, T.; Schuck, S.; Reichel-Deland, V.; Zeier, J. Biochemical Principles and Functional Aspects of Pipecolic Acid Biosynthesis in Plant Immunity. Plant Physiol. 2017, 174, 124–153. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.; Liu, R.; Lim, G.-H.; de Lorenzo, L.; Yu, K.; Zhang, K.; Hunt, A.G.; Kachroo, A.; Kachroo, P. Pipecolic Acid Confers Systemic Immunity by Regulating Free Radicals. Sci. Adv. 2018, 4, eaar4509. [Google Scholar] [CrossRef] [Green Version]
- McCann, H.C.; Rikkerink, E.H.A.; Bertels, F.; Fiers, M.; Lu, A.; Rees-George, J.; Andersen, M.T.; Gleave, A.P.; Haubold, B.; Wohlers, M.W.; et al. Genomic Analysis of the Kiwifruit Pathogen Pseudomonas syringae Pv. actinidiae Provides Insight into the Origins of an Emergent Plant Disease. PLoS Pathog. 2013, 9, e1003503. [Google Scholar] [CrossRef]
- Wang, T.; Wang, G.; Jia, Z.-H.; Pan, D.-L.; Zhang, J.-Y.; Guo, Z.-R. Transcriptome Analysis of Kiwifruit in Response to Pseudomonas syringae Pv. actinidiae Infection. Int. J. Mol. Sci. 2018, 19, 373. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dorostkar, S.; Dadkhodaie, A.; Ebrahimie, E.; Heidari, B.; Ahmadi-Kordshooli, M. Comparative Transcriptome Analysis of Two Contrasting Resistant and Susceptible Aegilops Tauschii Accessions to Wheat Leaf Rust (Puccinia triticina) Using RNA-Sequencing. Sci. Rep. 2022, 12, 821. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.G.; Dangl, J.L. The Plant Immune System. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dodds, P.N.; Rathjen, J.P. Plant Immunity: Towards an Integrated View of Plant–Pathogen Interactions. Nat. Rev. Genet. 2010, 11, 539–548. [Google Scholar] [CrossRef] [PubMed]
- Asai, T.; Tena, G.; Plotnikova, J.; Willmann, M.R.; Chiu, W.-L.; Gomez-Gomez, L.; Boller, T.; Ausubel, F.M.; Sheen, J. MAP Kinase Signalling Cascade in Arabidopsis Innate Immunity. Nature 2002, 415, 977–983. [Google Scholar] [CrossRef]
- Tena, G.; Asai, T.; Chiu, W.-L.; Sheen, J. Plant Mitogen-Activated Protein Kinase Signaling Cascades. Curr. Opin. Plant Biol. 2001, 4, 392–400. [Google Scholar] [CrossRef]
- Mao, G.; Meng, X.; Liu, Y.; Zheng, Z.; Chen, Z.; Zhang, S. Phosphorylation of a WRKY Transcription Factor by Two Pathogen-Responsive MAPKs Drives Phytoalexin Biosynthesis in Arabidopsis. Plant Cell 2011, 23, 1639–1653. [Google Scholar] [CrossRef] [Green Version]
- Lam, E.; Kato, N.; Lawton, M. Programmed Cell Death, Mitochondria and the Plant Hypersensitive Response. Nature 2001, 411, 848–853. [Google Scholar] [CrossRef]
- Coll, N.S.; Epple, P.; Dangl, J.L. Programmed Cell Death in the Plant Immune System. Cell Death Differ. 2011, 18, 1247–1256. [Google Scholar] [CrossRef] [Green Version]
- Chan, C.W.M.; Schorrak, L.M.; Smith, R.K.; Bent, A.F.; Sussman, M.R. A Cyclic Nucleotide-Gated Ion Channel, CNGC2, Is Crucial for Plant Development and Adaptation to Calcium Stress. Plant Physiol. 2003, 132, 728–731. [Google Scholar] [CrossRef] [Green Version]
- Kaplan, B.; Sherman, T.; Fromm, H. Cyclic Nucleotide-Gated Channels in Plants. FEBS Lett. 2007, 581, 2237–2246. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, W.; Smigel, A.; Tsai, Y.-C.; Braam, J.; Berkowitz, G.A. Innate Immunity Signaling: Cytosolic Ca2+ Elevation Is Linked to Downstream Nitric Oxide Generation through the Action of Calmodulin or a Calmodulin-Like Protein. Plant Physiol. 2008, 148, 818–828. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kobayashi, M.; Ohura, I.; Kawakita, K.; Yokota, N.; Fujiwara, M.; Shimamoto, K.; Doke, N.; Yoshioka, H. Calcium-Dependent Protein Kinases Regulate the Production of Reactive Oxygen Species by Potato NADPH Oxidase. Plant Cell 2007, 19, 1065–1080. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dubiella, U.; Seybold, H.; Durian, G.; Komander, E.; Lassig, R.; Witte, C.-P.; Schulze, W.X.; Romeis, T. Calcium-Dependent Protein Kinase/NADPH Oxidase Activation Circuit Is Required for Rapid Defense Signal Propagation. Proc. Natl. Acad. Sci. USA 2013, 110, 8744–8749. [Google Scholar] [CrossRef] [Green Version]
- Torres, M.A.; Jones, J.D.G.; Dangl, J.L. Reactive Oxygen Species Signaling in Response to Pathogens. Plant Physiol. 2006, 141, 373–378. [Google Scholar] [CrossRef] [Green Version]
- Torres, M.A.; Dangl, J.L. Functions of the Respiratory Burst Oxidase in Biotic Interactions, Abiotic Stress and Development. Curr. Opin. Plant Biol. 2005, 8, 397–403. [Google Scholar] [CrossRef]
- Slatnar, A.; Mikulič-Petkovšek, M.; Veberič, R.; Štampar, F. Research on the Involment of Phenoloics in the Defence of Horticultural Plants. Acta Agric. Slov. 2016, 107, 183. [Google Scholar] [CrossRef] [Green Version]
- Zhao, J.; Dixon, R.A. The ‘Ins’ and ‘Outs’ of Flavonoid Transport. Trends Plant Sci. 2010, 15, 72–80. [Google Scholar] [CrossRef] [Green Version]
- Petrussa, E.; Braidot, E.; Zancani, M.; Peresson, C.; Bertolini, A.; Patui, S.; Vianello, A. Plant Flavonoids—Biosynthesis, Transport and Involvement in Stress Responses. Int. J. Mol. Sci. 2013, 14, 14950–14973. [Google Scholar] [CrossRef]
- Gu, Y.-Q.; Wildermuth, M.C.; Chakravarthy, S.; Loh, Y.-T.; Yang, C.; He, X.; Han, Y.; Martin, G.B. Tomato Transcription Factors Pti4, Pti5, and Pti6 Activate Defense Responses When Expressed in Arabidopsis. Plant Cell 2002, 14, 817–831. [Google Scholar] [CrossRef] [Green Version]
- Sun, M.; Qiu, L.; Liu, Y.; Zhang, H.; Zhang, Y.; Qin, Y.; Mao, Y.; Zhou, M.; Du, X.; Qin, Z.; et al. Pto Interaction Proteins: Critical Regulators in Plant Development and Stress Response. Front. Plant Sci. 2022, 13, 774229. [Google Scholar] [CrossRef] [PubMed]
- Naoumkina, M.A.; Zhao, Q.; Gallego-Giraldo, L.; Dai, X.; Zhao, P.X.; Dixon, R.A. Genome-Wide Analysis of Phenylpropanoid Defence Pathways: Phenylpropanoid Defence Pathways. Mol. Plant Pathol. 2010, 11, 829–846. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Dixon, R.A. Transcriptional Networks for Lignin Biosynthesis: More Complex than We Thought? Trends Plant Sci. 2011, 16, 227–233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, L.; Zhu, L.; Tu, L.; Liu, L.; Yuan, D.; Jin, L.; Long, L.; Zhang, X. Lignin Metabolism Has a Central Role in the Resistance of Cotton to the Wilt Fungus Verticillium Dahliae as Revealed by RNA-Seq-Dependent Transcriptional Analysis and Histochemistry. J. Exp. Bot. 2011, 62, 5607–5621. [Google Scholar] [CrossRef] [Green Version]
- Glazebrook, J. Contrasting Mechanisms of Defense against Biotrophic and Necrotrophic Pathogens. Annu. Rev. Phytopathol. 2005, 43, 227. [Google Scholar] [CrossRef] [PubMed]
- Loake, G.; Grant, M. Salicylic Acid in Plant Defence—The Players and Protagonists. Curr. Opin. Plant Biol. 2007, 10, 466–472. [Google Scholar] [CrossRef]
- Métrauxs, J.-P. Systemic Acquired Resistance and Salicylic Acid: Current State of Knowledge. Eur. J. Plant Pathol. 2001, 107, 13–18. [Google Scholar] [CrossRef]
- Zhang, Y.; Fan, W.; Kinkema, M.; Li, X.; Dong, X. Interaction of NPR1 with Basic Leucine Zipper Protein Transcription Factors That Bind Sequences Required for Salicylic Acid Induction of the PR-1 Gene. Proc. Natl. Acad. Sci. USA 1999, 96, 6523–6528. [Google Scholar] [CrossRef] [Green Version]
- Lebel, E.; Heifetz, P.; Thorne, L.; Uknes, S.; Ryals, J.; Ward, E. Functional Analysis of Regulatory Sequences ControllingPR-1 Gene Expression in Arabidopsis. Plant J. 1998, 16, 223–233. [Google Scholar] [CrossRef]
- Kunkel, B.N.; Brooks, D.M. Cross Talk between Signaling Pathways in Pathogen Defense. Curr. Opin. Plant Biol. 2002, 5, 325–331. [Google Scholar] [CrossRef]
- Li, J.; Brader, G.; Palva, E.T. The WRKY70 Transcription Factor: A Node of Convergence for Jasmonate-Mediated and Salicylate-Mediated Signals in Plant Defense. Plant Cell 2004, 16, 319–331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yasuda, M.; Ishikawa, A.; Jikumaru, Y.; Seki, M.; Umezawa, T.; Asami, T.; Maruyama-Nakashita, A.; Kudo, T.; Shinozaki, K.; Yoshida, S.; et al. Antagonistic Interaction between Systemic Acquired Resistance and the Abscisic Acid–Mediated Abiotic Stress Response in Arabidopsis. Plant Cell 2008, 20, 1678–1692. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Z.; Agnew, J.L.; Cohen, J.D.; He, P.; Shan, L.; Sheen, J.; Kunkel, B.N. Pseudomonas syringae Type III Effector AvrRpt2 Alters Arabidopsis thaliana Auxin Physiology. Proc. Natl. Acad. Sci. USA 2007, 104, 20131–20136. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, D.; Pajerowska-Mukhtar, K.; Culler, A.H.; Dong, X. Salicylic Acid Inhibits Pathogen Growth in Plants through Repression of the Auxin Signaling Pathway. Curr. Biol. 2007, 17, 1784–1790. [Google Scholar] [CrossRef] [Green Version]
- Choi, J.; Huh, S.U.; Kojima, M.; Sakakibara, H.; Paek, K.-H.; Hwang, I. The Cytokinin-Activated Transcription Factor ARR2 Promotes Plant Immunity via TGA3/NPR1-Dependent Salicylic Acid Signaling in Arabidopsis. Dev. Cell 2010, 19, 284–295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakashita, H.; Yasuda, M.; Nitta, T.; Asami, T.; Fujioka, S.; Arai, Y.; Sekimata, K.; Takatsuto, S.; Yamaguchi, I.; Yoshida, S. Brassinosteroid Functions in a Broad Range of Disease Resistance in Tobacco and Rice: Steroid Hormone-Mediated Disease Resistance. Plant J. 2003, 33, 887–898. [Google Scholar] [CrossRef] [Green Version]
- Trapnell, C.; Pachter, L.; Salzberg, S.L. TopHat: Discovering Splice Junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript Assembly and Quantification by RNA-Seq Reveals Unannotated Transcripts and Isoform Switching during Cell Differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Feng, Z.; Wang, X.; Wang, X.; Zhang, X. DEGseq: An R Package for Identifying Differentially Expressed Genes from RNA-Seq Data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential Expression Analysis for Sequence Count Data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [Green Version]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene Ontology Analysis for RNA-Seq: Accounting for Selection Bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Samples | Raw Reads | Clean Reads | Mapped Reads | Uniquely Mapped | Uniquely Mapped Rate |
|---|---|---|---|---|---|
| HY_C_1 | 53173060 | 47659800 | 42445216 | 41754675 | 87.61% |
| HY_C_2 | 50688944 | 44697010 | 39824177 | 38533070 | 86.21% |
| HY_C_3 | 50321846 | 45046286 | 39975928 | 39282108 | 87.20% |
| HY_P_1 | 52583390 | 47199212 | 40516165 | 39783757 | 84.29% |
| HY_P_2 | 47920524 | 42513064 | 36786553 | 35602846 | 83.75% |
| HY_P_3 | 52116128 | 46209178 | 41318821 | 40630090 | 87.93% |
| MH_C_1 | 47134410 | 41928396 | 16033373 | 14770334 | 35.23% |
| MH_C_2 | 51174634 | 45465026 | 17584600 | 16821959 | 37.00% |
| MH_C_3 | 53840934 | 47798112 | 18719524 | 17294295 | 36.18% |
| MH_P_1 | 51884426 | 45765796 | 17705514 | 17016783 | 37.18% |
| MH_P_2 | 50151192 | 44200726 | 17146800 | 15883761 | 35.94% |
| MH_P_3 | 53392744 | 47376418 | 18078332 | 16653103 | 35.15% |
| Description | DEGs of HY | Description | DEGs of MH | ||||
|---|---|---|---|---|---|---|---|
| Up | Down | All | Up | Down | All | ||
| Metabolic pathways | 153 | 232 | 385 | Metabolic pathways | 116 | 178 | 294 |
| Biosynthesis of secondary metabolites | 78 | 141 | 219 | Biosynthesis of secondary metabolites | 11 | 96 | 107 |
| Plant hormone signal transduction | 36 | 46 | 82 | Starch and sucrose metabolism | 30 | 39 | 69 |
| Microbial metabolism in diverse environments | 41 | 38 | 79 | Antigen processing and presentation | 58 | 7 | 65 |
| Plant–pathogen interaction | 40 | 20 | 60 | Microbial metabolism in diverse environments | 5 | 33 | 38 |
| Biosynthesis of amino acids | 15 | 34 | 49 | Plant hormone-signal transduction | 9 | 27 | 36 |
| Starch and sucrose metabolism | 13 | 33 | 46 | Amino sugar and nucleotide sugar metabolism | 9 | 24 | 33 |
| Protein processing in endoplasmic reticulum | 32 | 10 | 42 | Biosynthesis of unsaturated fatty acids | 26 | 6 | 32 |
| Amino sugar and nucleotide sugar metabolism | 6 | 31 | 37 | Phagosome | 7 | 19 | 26 |
| Glycolysis/Gluconeogenesis | 14 | 19 | 33 | Biosynthesis of amino acids | 1 | 24 | 25 |
| Cell cycle | 5 | 26 | 31 | Galactose metabolism | 16 | 8 | 24 |
| Peroxisome | 20 | 10 | 30 | Purine metabolism | 4 | 20 | 24 |
| Pentose and glucuronate interconversions | 6 | 24 | 30 | Pentose and glucuronate interconversions | 8 | 15 | 23 |
| Purine metabolism | 6 | 23 | 29 | Glycolysis/gluconeogenesis | 5 | 18 | 23 |
| Cysteine and methionine metabolism | 7 | 21 | 28 | Protein processing in endoplasmic reticulum | 2 | 20 | 22 |
| Pyrimidine metabolism | 9 | 18 | 27 | Photosynthesis | 7 | 11 | 18 |
| Phenylalanine metabolism | 9 | 15 | 24 | Gap junction | 11 | 6 | 17 |
| Glutathione metabolism | 18 | 5 | 23 | Estrogen-signaling pathway | 15 | 1 | 16 |
| Epstein-Barr virus infection | 12 | 10 | 22 | Cell cycle | 2 | 14 | 16 |
| Phenylpropanoid biosynthesis | 6 | 15 | 21 | Plant–pathogen interaction | 1 | 15 | 16 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qin, X.; Zhang, M.; Li, Q.; Chen, D.; Sun, L.; Qi, X.; Cao, K.; Fang, J. Transcriptional Analysis on Resistant and Susceptible Kiwifruit Genotypes Activating Different Plant-Immunity Processes against Pseudomonas syringae pv. actinidiae. Int. J. Mol. Sci. 2022, 23, 7643. https://doi.org/10.3390/ijms23147643
Qin X, Zhang M, Li Q, Chen D, Sun L, Qi X, Cao K, Fang J. Transcriptional Analysis on Resistant and Susceptible Kiwifruit Genotypes Activating Different Plant-Immunity Processes against Pseudomonas syringae pv. actinidiae. International Journal of Molecular Sciences. 2022; 23(14):7643. https://doi.org/10.3390/ijms23147643
Chicago/Turabian StyleQin, Xiaobo, Min Zhang, Qiaohong Li, Dalei Chen, Leiming Sun, Xiujuan Qi, Ke Cao, and Jinbao Fang. 2022. "Transcriptional Analysis on Resistant and Susceptible Kiwifruit Genotypes Activating Different Plant-Immunity Processes against Pseudomonas syringae pv. actinidiae" International Journal of Molecular Sciences 23, no. 14: 7643. https://doi.org/10.3390/ijms23147643






