The Analysis of the Human Megakaryocyte and Platelet Coding Transcriptome in Healthy and Diseased Subjects
Abstract
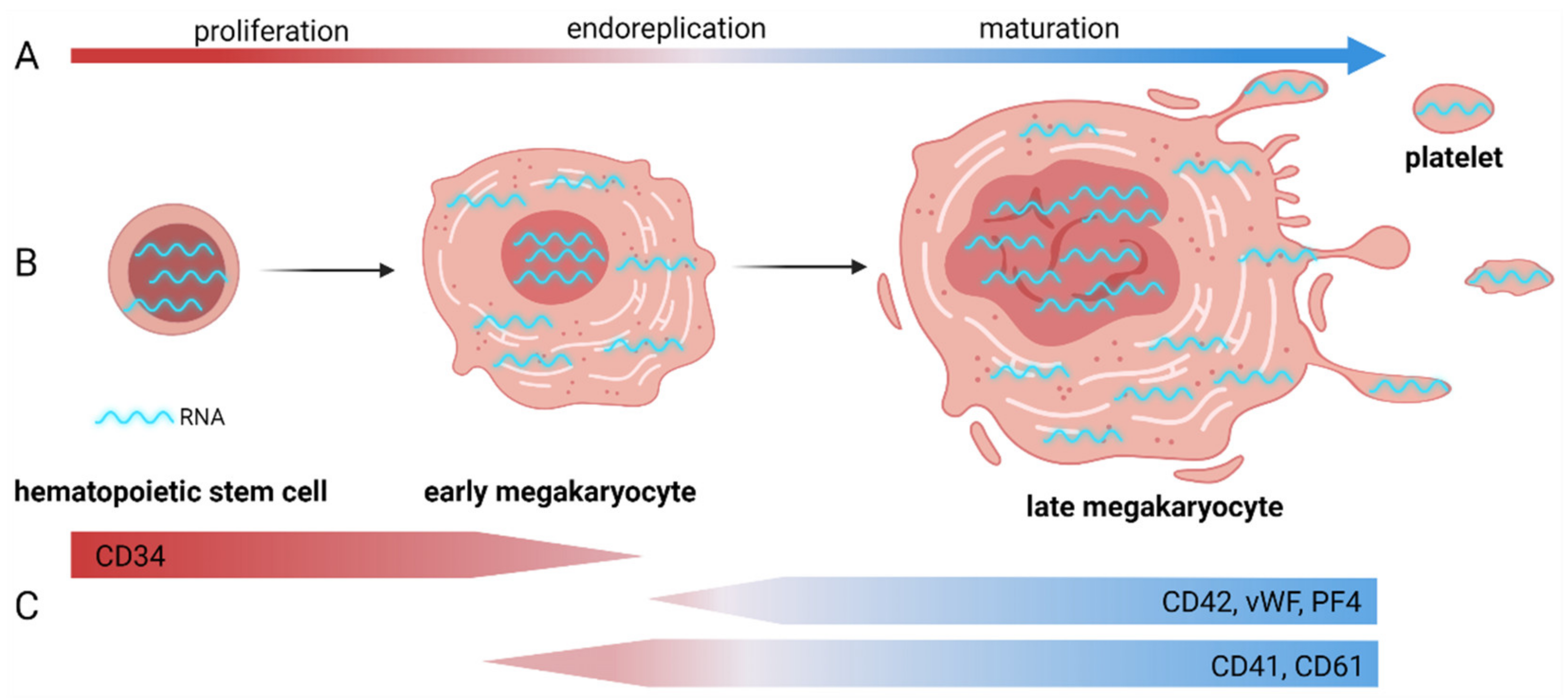
:1. Introduction to Megakaryopoiesis and Platelet Formation with a Focus on RNA
2. Bulk Transcriptomics of Healthy Megakaryocytes
3. Bulk Transcriptomics of Healthy Platelets
4. Single-Cell Transcriptomics of Healthy Megakaryocytes
| Reference | Cell Type(S) | Summary of Results | Technology |
|---|---|---|---|
| Choudry et al. (2021) [73] | MKs and HSCs | MKs in lower ploidy states highly express platelet-specific genes. As polyploidization increases and the cell prepares for thrombopoiesis, gene expression is redirected towards transcriptional programs involved in translation and posttranslational processing. Two MK-biased HSC subpopulations were also observed and shown to originate from the BM. Finally, BM MKs from patients with recent myocardial infarction showed a specific gene expression signature that supports the modulation of MK differentiation in this thrombotic state. | G&T-seq |
| Estevez et al. (2021) [98] | HSCs | The effect of germline monoallelic mutations in RUNX1, found in patients suffering from familial platelet disorder with a predisposition to myeloid malignancy, was studied by inserting the patient mutations into iPSC-derived hematopoietic progenitor cells (iHSCs) and performing scRNA-seq. There was found to be a marked deficiency of MK-biased iHSCs in mutated cultures, and gene sets that were upregulated included response to stress, regulation of signal transduction and immune signaling-related gene sets. An increased sensitivity to transforming growth factor β1 and an increase in the stress pathway through upregulation of c-jun N-terminal kinase-2 phosphorylation were observed. | 10X Chromium |
| Lawrence et al. (2022) [94] | In vitro differentiating cells from iPSCs and HSCs up to MKs | Analysis of iPSC-derived MK differentiation and transcriptomic comparison with primary hematopoietic stem and progenitor cells. The in vitro cells do not pass through states resembling HSCs or MPPs as seen in vivo, but the further differentiated MK progenitor cells do exhibit a very similar transcriptome to their in vivo counterparts. A surface marker panel is described for MK progenitors, allowing for selection from culture and for insights into this intermediary state. | 10X Chromium and smart-seq |
| Liu et al. (2021) [90] | MKs | Cellular heterogeneity within MKs was mapped, and an MK subpopulation with high enrichment of immune-associated genes was identified. The immune signature could be traced back to the progenitor stage, and two surface markers, CD148 and CD48, were identified. This type of MK can respond rapidly to immune stimuli both in vitro and in vivo, exhibiting high expression of immune receptors and mediators, which might act as immune-surveillance cells. | Smart-seq |
| Lu et al. (2018) [96] | MEPs, common myeloid progenitors, and MK and erythroid progenitors | MEPs have a distinct gene expression signature that represents a continuous transition state from common myeloid progenitor cells to MK and erythroid progenitor cells. | Fluidigm C1 |
| Psaila et al. (2017) [95] | CD34+ peripheral blood cells | Myelofibrosis causes an increased number of immature/low ploidy MKs with an altered transcriptome. Patient HSPCs have increased expression of MK-associated genes, including VWF and ITGA2B. Patient CD34+ progenitor cells showed increased expression of PF4 and TGFβ. | 10X Chromium |
| Psaila et al. (2020) [97] | HSPCs | MK-biased hematopoiesis in myelofibrosis was observed, with heterogeneous MKp showing a highly expressed fibrosis signature and an aberrant metabolic and inflammatory signature. Targeting the aberrant expression of surface G6B may selectively ablate the myelofibrosis HSPC clone. | 10X Chromium |
| Riemondy et al. (2019) [93] | Lymphocytes and MK mixture | The introduction of a method for resampling cell-type-wise, cell-wise or sample-wise from an existing complex scRNA library. | 10X Chromium |
| Sun et al. (2021) [91] | Human and mouse MKs | Three distinct MK subpopulations were observed to possess gene signatures related to platelet generation, HSC niche interaction or inflammatory response. The first type of MK was mostly found near blood vessels, and the second was near HSCs. The third type, containing a gene signature related to the inflammatory response, was lower in ploidy, consisted of 5% of MKs and was capable of engulfing and digesting bacteria and stimulating T cells in vitro. | Smart-seq |
| Wang et al. (2021) [92] | Human MKs | A comprehensive single-cell transcriptomic landscape of human MKs was constructed where MKs show cellular heterogeneity with distinct metabolic and cell cycle signatures. CD14+ MKs with immune characteristics were generated along a distinct trajectory, and THBS1 was identified as an early marker for MK-biased endothelial cells from human embryonic stem cells. | 10X Chromium |
5. Single-Cell Transcriptomics of Healthy Platelets
6. Platelet and Megakaryocyte Transcriptome to Decipher Inherited Platelet Disorders
7. Conclusions and Future Perspectives
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Akashi, K.; Traver, D.; Miyamoto, T.; Weissman, I.L. A clonogenic common myeloid progenitor that gives rise to all myeloid lineages. Nature 2000, 404, 193–197. [Google Scholar] [CrossRef] [PubMed]
- Woolthuis, C.M.; Park, C.Y. Hematopoietic stem/progenitor cell commitment to the megakaryocyte lineage. Blood 2016, 127, 1242–1248. [Google Scholar] [CrossRef] [Green Version]
- Donné, R.; Nader, M.B.; Desdouets, C. Cellular and Molecular Mechanisms Controlling Ploidy. Ref. Modul. Life Sci. 2018. [Google Scholar] [CrossRef]
- Michelson, A.; Cattaneo, M.; Frelinger, A.; Newman, P. Platelets; Academic Press: Cambridge, MA, USA, 2019. [Google Scholar]
- Davizon-Castillo, P.; Rowley, J.W.; Rondina, M.T. Megakaryocyte and Platelet Transcriptomics for Discoveries in Human Health and Disease. Arterioscler. Thromb. Vasc. Biol. 2020, 40, 1432–1440. [Google Scholar] [CrossRef] [PubMed]
- Eicke, D.; Baigger, A.; Schulze, K.; Latham, S.; Halloin, C.; Zweigerdt, R.; Guzman, C.A.; Blasczyk, R.; Figueiredo, C. Large-scale production of megakaryocytes in microcarrier-supported stirred suspension bioreactors. Sci. Rep. 2018, 8, 10146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, Q.; Shabrani, N.; Thon, J.N.; Huo, H.; Thiel, A.; Machlus, K.R.; Kim, K.; Brooks, J.; Li, F.; Luo, C.; et al. Scalable Generation of Universal Platelets from Human Induced Pluripotent Stem Cells. Stem Cell Rep. 2014, 3, 817–831. [Google Scholar] [CrossRef] [Green Version]
- Mills, J.A.; Paluru, P.; Weiss, M.J.; Gadue, P.; French, D.L. Hematopoietic Differentiation of Pluripotent Stem Cells in Culture. Methods Mol. Biol. 2014, 1185, 181–194. [Google Scholar] [CrossRef]
- Pick, M.; Azzola, L.; Osborne, E.; Stanley, E.G.; Elefanty, A.G. Generation of Megakaryocytic Progenitors from Human Embryonic Stem Cells in a Feeder- and Serum-Free Medium. PLoS ONE 2013, 8, e55530. [Google Scholar] [CrossRef] [Green Version]
- Elcheva, I.; Brok-Volchanskaya, V.; Kumar, A.; Liu, P.; Lee, J.-H.; Tong, L.; Vodyanik, M.; Swanson, S.; Stewart, R.; Kyba, M.; et al. Direct induction of haematoendothelial programs in human pluripotent stem cells by transcriptional regulators. Nat. Commun. 2014, 5, 4372. [Google Scholar] [CrossRef] [Green Version]
- Moreau, T.; Evans, A.L.; Vasquez, L.; Tijssen, M.R.; Yan, Y.; Trotter, M.W.; Howard, D.; Colzani, M.; Arumugam, M.; Wu, W.H.; et al. Large-scale production of megakaryocytes from human pluripotent stem cells by chemically defined forward programming. Nat. Commun. 2016, 7, 11208. [Google Scholar] [CrossRef]
- Chen, T.W.; Yao, C.L.; Chu, I.M.; Chuang, T.L.; Hsieh, T.B.; Hwang, S.M. Large generation of megakaryocytes from serum-free expanded human CD34+ cells. Biochem. Biophys. Res. Commun. 2009, 378, 112–117. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, S.; Takayama, N.; Hirata, S.; Seo, H.; Endo, H.; Ochi, K.; Fujita, K.-I.; Koike, T.; Harimoto, K.-I.; Dohda, T.; et al. Expandable Megakaryocyte Cell Lines Enable Clinically Applicable Generation of Platelets from Human Induced Pluripotent Stem Cells. Cell Stem Cell 2014, 14, 535–548. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wright, J.H. The histogenesis of the blood platelets. J. Morphol. 1910, 21, 263–278. [Google Scholar] [CrossRef]
- Kosaki, G. In vivo platelet production from mature megakaryocytes: Does platelet release occur via proplatelets? Int. J. Hematol. 2005, 81, 208–219. [Google Scholar] [CrossRef]
- Bender, M.; Thon, J.N.; Ehrlicher, A.J.; Wu, S.; Mazutis, L.; Deschmann, E.; Sola-Visner, M.; Italiano, J.E.; Hartwig, J.H. Microtubule sliding drives proplatelet elongation and is dependent on cytoplasmic dynein. Blood 2015, 125, 860–868. [Google Scholar] [CrossRef] [Green Version]
- Jiang, J.; Woulfe, D.S.; Papoutsakis, E.T. Shear enhances thrombopoiesis and formation of microparticles that induce megakaryocytic differentiation of stem cells. Blood 2014, 124, 2094–2103. [Google Scholar] [CrossRef]
- Junt, T.; Schulze, H.; Chen, Z.; Massberg, S.; Goerge, T.; Krueger, A.; Wagner, D.D.; Graf, T.; Italiano, J.E.; Shivdasani, R.A.; et al. Dynamic visualization of thrombopoiesis within bone marrow. Science 2007, 317, 1767–1770. [Google Scholar] [CrossRef] [Green Version]
- Lefrançais, E.; Ortiz-Muñoz, G.; Caudrillier, A.; Mallavia, B.; Liu, F.; Sayah, D.M.; Thornton, E.E.; Headley, M.B.; David, T.; Coughlin, S.R.; et al. The lung is a site of platelet biogenesis and a reservoir for haematopoietic progenitors. Nature 2017, 544, 105–109. [Google Scholar] [CrossRef]
- Thon, J.N.; Montalvo, A.; Patel-Hett, S.; Devine, M.T.; Richardson, J.L.; Ehrlicher, A.; Larson, M.K.; Hoffmeister, K.; Hartwig, J.H.; Italiano, J.E. Cytoskeletal mechanics of proplatelet maturation and platelet release. J. Cell Biol. 2010, 191, 861–874. [Google Scholar] [CrossRef] [Green Version]
- Boscher, J.; Guinard, I.; Eckly, A.; Lanza, F.; Léon, C. Blood platelet formation at a glance. J. Cell Sci. 2020, 133, jcs244731. [Google Scholar] [CrossRef]
- Bornert, A.; Boscher, J.; Pertuy, F.; Eckly, A.; Stegner, D.; Strassel, C.; Gachet, C.; Lanza, F.; Léon, C. Cytoskeletal-based mechanisms differently regulate in vivo and in vitro proplatelet formation. Haematologica 2021, 106, 1368–1380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kowata, S.; Isogai, S.; Murai, K.; Ito, S.; Tohyama, K.; Ema, M.; Hitomi, J.; Ishida, Y. Platelet demand modulates the type of intravascular protrusion of megakaryocytes in bone marrow. Thromb. Haemost. 2014, 112, 743–756. [Google Scholar] [CrossRef] [PubMed]
- Radley, J.; Scurfield, G. The mechanism of platelet release. Blood 1980, 56, 996–999. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nishimura, S.; Nagasaki, M.; Kunishima, S.; Sawaguchi, A.; Sakata, A.; Sakaguchi, H.; Ohmori, T.; Manabe, I.; Italiano, J.E.; Ryu, T.; et al. IL-1α induces thrombopoiesis through megakaryocyte rupture in response to acute platelet needs. J. Cell Biol. 2015, 209, 453–466. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van der Meijden, P.E.J.; Heemskerk, J.W.M. Platelet biology and functions: New concepts and clinical perspectives. Nat. Rev. Cardiol. 2018, 16, 166–179. [Google Scholar] [CrossRef]
- Wagner, R. Glycogen content of isolated white blood cells in glycogen storage disease. Am. J. Dis. Child. 1947, 73, 559–564. [Google Scholar] [CrossRef]
- Warshaw, A.L.; Laster, L.; Shulman, N.R. Protein Synthesis by Human Platelets from the Section on. J. Biol. Chem. 1967, 242, 2094–2097. [Google Scholar] [CrossRef]
- Bray, P.F.; McKenzie, S.E.; Edelstein, L.C.; Nagalla, S.; Delgrosso, K.; Ertel, A.; Kupper, J.; Jing, Y.; Londin, E.; Loher, P.; et al. The complex transcriptional landscape of the anucleate human platelet. BMC Genom. 2013, 14, 1. [Google Scholar] [CrossRef] [Green Version]
- Nassa, G.; Giurato, G.; Cimmino, G.; Rizzo, F.; Ravo, M.; Salvati, A.; Nyman, T.A.; Zhu, Y.; Vesterlund, M.; Lehtiö, J.; et al. Splicing of platelet resident pre-mRNAs upon activation by physiological stimuli results in functionally relevant proteome modifications. Sci. Rep. 2017, 8, 498. [Google Scholar] [CrossRef]
- Schubert, S.; Weyrich, A.S.; Rowley, J.W. A tour through the transcriptional landscape of platelets. Blood 2014, 124, 493–502. [Google Scholar] [CrossRef] [Green Version]
- Clancy, L.; Freedman, J.E. New paradigms in thrombosis: Novel mediators and biomarkers platelet RNA transfer. J. Thromb. Thrombolysis 2014, 37, 12–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Czajka, P.; Fitas, A.; Jakubik, D.; Eyileten, C.; Gasecka, A.; Wicik, Z.; Siller-Matula, J.M.; Filipiak, K.J.; Postula, M. MicroRNA as Potential Biomarkers of Platelet Function on Antiplatelet Therapy: A Review. Front. Physiol. 2021, 12, 498. [Google Scholar] [CrossRef] [PubMed]
- Dangwal, S.; Thum, T. MicroRNAs in platelet physiology and pathology. Hamostaseologie 2013, 33, 17–20. [Google Scholar] [CrossRef] [Green Version]
- Edelstein, L.C.; Mckenzie, S.E.; Shaw, C.; Holinstat, M.A.; Kunapuli, S.P.; Bray, P.F. MicroRNAs in platelet production and activation. J. Thromb. Haemost. 2013, 11 (Suppl. 1), 340–350. [Google Scholar] [CrossRef]
- Gidlöf, O.; van der Brug, M.; Öhman, J.; Gilje, P.; Olde, B.; Wahlestedt, C.; Erlinge, D. Platelets activated during myocardial infarction release functional miRNA, which can be taken up by endothelial cells and regulate ICAM1 expression. Blood 2013, 121, 3908–3917. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laffont, B.; Corduan, A.; Pĺe, H.; Duchez, A.C.; Cloutier, N.; Boilard, E.; Provost, P. Activated platelets can deliver mRNA regulatory Ago2•microRNA complexes to endothelial cells via microparticles. Blood 2013, 122, 253–261. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mayr, M.; Zampetaki, A.; Willeit, P.; Willeit, J.; Kiechl, S. MicroRNAs within the continuum of postgenomics biomarker discovery. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 206–214. [Google Scholar] [CrossRef] [Green Version]
- Nagalla, S.; Shaw, C.; Kong, X.; Kondkar, A.A.; Edelstein, L.C.; Ma, L.; Chen, J.; McKnight, G.S.; López, J.A.; Yang, L.; et al. Platelet microRNA-mRNA coexpression profiles correlate with platelet reactivity. Blood 2011, 117, 5189–5197. [Google Scholar] [CrossRef]
- Angénieux, C.; Maitre, B.; Eckly, A.; Lanza, F.; Gachet, C.; de la Salle, H. Time-Dependent Decay of mRNA and Ribosomal RNA during Platelet Aging and Its Correlation with Translation Activity. PLoS ONE 2016, 11, e31529. [Google Scholar] [CrossRef]
- Dölken, L.; Ruzsics, Z.; Rädle, B.; Friedel, C.C.; Zimmer, R.; Mages, J.; Hoffmann, R.; Dickinson, P.; Forster, T.; Ghazal, P.; et al. High-resolution gene expression profiling for simultaneous kinetic parameter analysis of RNA synthesis and decay. RNA 2008, 14, 1959–1972. [Google Scholar] [CrossRef] [Green Version]
- McManus, J.; Cheng, Z.; Vogel, C. Next-generation analysis of gene expression regulation—Comparing the roles of synthesis and degradation. Mol. BioSyst. 2015, 11, 2680–2689. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burkhart, J.M.; Vaudel, M.; Gambaryan, S.; Radau, S.; Walter, U.; Martens, L.; Geiger, J.; Sickmann, A.; Zahedi, R.P. The first comprehensive and quantitative analysis of human platelet protein composition allows the comparative analysis of structural and functional pathways. Blood 2012, 120, e73–e82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Houng, A.K.; Maggini, L.; Clement, C.Y.; Reed, G.L. Identification and structure of activated-platelet protein-1, a protein with RNA-binding domain motifs that is expressed by activated platelets. Eur. J. Biochem. 1997, 243, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Landry, P.; Plante, I.; Ouellet, D.L.; Perron, M.P.; Rousseau, G.; Provost, P. Existence of a microRNA pathway in anucleate platelets. Nat. Struct. Mol. Biol. 2009, 16, 961–966. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dittrich, M.; Birschmann, I.; Pfrang, J.; Herterich, S.; Smolenski, A.; Walter, U.; Dandekar, T. Analysis of SAGE data in human platelets: Features of the transcriptome in an anucleate cell. Thromb. Haemost. 2006, 95, 643–651. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Denis, M.M.; Tolley, N.D.; Bunting, M.; Schwertz, H.; Jiang, H.; Lindemann, S.; Yost, C.C.; Rubner, F.J.; Albertine, K.H.; Swoboda, K.J.; et al. Escaping the nuclear confines: Signal-dependent pre-mRNA splicing in anucleate platelets. Cell 2005, 122, 379–391. [Google Scholar] [CrossRef] [Green Version]
- Power, K.A.; McRedmond, J.P.; de Stefani, A.; Gallagher, W.M.; Gaora, P.Ó. High-Throughput Proteomics Detection of Novel Splice Isoforms in Human Platelets. PLoS ONE 2009, 4, e5001. [Google Scholar] [CrossRef] [Green Version]
- Djaffar, I.; Chen, Y.P.; Creminon, C.; Maclouf, J.; Cieutat, A.M.; Gayet, O.; Rosa, J.P. A new alternative transcript encodes a 60 kDa truncated form of integrin beta 3. Biochem. J. 1994, 300 Pt 1, 69–74. [Google Scholar] [CrossRef]
- Censarek, P.; Steger, G.; Paolini, C.; Hohlfeld, T.; Grosser, T.; Zimmermann, N.; Fleckenstein, D.; Schrör, K.; Weber, A.-A. Alternative splicing of platelet cyclooxygenase-2 mRNA in patients after coronary artery bypass grafting. Thromb. Haemost. 2007, 98, 1309–1315. [Google Scholar] [CrossRef]
- Sun, L.; Mao, G.; Kunapuli, S.P.; Dhanasekaran, D.N.; Rao, A.K. Alternative splice variants of phospholipase C-β2 are expressed in platelets: Effect on Gαq-dependent activation and localization. Platelets 2009, 18, 217–223. [Google Scholar] [CrossRef]
- Huang, Y.Q.; Li, J.J.; Karpatkin, S. Identification of a family of alternatively spliced mRNA species of angiopoietin-1. Blood 2000, 95, 1993–1999. [Google Scholar] [CrossRef]
- Best, M.G.; Vancura, A.; Wurdinger, T. Platelet RNA as a circulating biomarker trove for cancer diagnostics. J. Thromb. Haemost. 2017, 15, 1295–1306. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schwertz, H.; Tolley, N.D.; Foulks, J.; Denis, M.M.; Risenmay, B.W.; Buerke, M.; Tilley, R.E.; Rondina, M.T.; Harris, E.M.; Kraiss, L.W.; et al. Signal-dependent splicing of tissue factor pre-mRNA modulates the thrombogenicity of human platelets. J. Exp. Med. 2006, 203, 2433–2440. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Münzer, P.; Tolios, A.; Pelzl, L.; Schmid, E.; Schmidt, E.-M.; Walker, B.; Fröhlich, H.; Borst, O.; Gawaz, M.; Lang, F. Thrombin-sensitive expression of the store operated Ca(2+) channel Orai1 in platelets. Biochem. Biophys. Res. Commun. 2013, 436, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.D. Exploiting the Hidden Treasure of Detained Introns. Cancer Cell 2017, 32, 393–395. [Google Scholar] [CrossRef] [Green Version]
- Lentaigne, C.; Greene, D.; Sivapalaratnam, S.; Favier, R.; Seyres, D.; Thys, C.; Grassi, L.; Mangles, S.; Sibson, K.; Stubbs, M.J.; et al. Germline mutations in the transcription factor IKZF5 cause thrombocytopenia. Blood 2019, 134, 2070–2081. [Google Scholar] [CrossRef]
- Psaila, B.; Mead, A.J. Single-cell approaches reveal novel cellular pathways for megakaryocyte and erythroid differentiation. Blood 2019, 133, 1427–1435. [Google Scholar] [CrossRef]
- Heazlewood, S.Y.; Williams, B.; Storan, M.J.; Nilsson, S.K. The Prospective Isolation of Viable, High Ploidy Megakaryocytes from Adult Murine Bone Marrow by Fluorescence Activated Cell Sorting. Methods Mol. Biol. 2013, 1035, 121–133. [Google Scholar] [CrossRef]
- Clien, L.; Kostadima, M.; Martens, J.H.A.; Canu, G.; Garcia, S.P.; Turro, E.; Downes, K.; Macaulay, I.C.; Bielczyk-Maczynska, E.; Coe, S.; et al. Transcriptional diversity during lineage commitment of human blood progenitors. Science 2014, 345, 1251033. [Google Scholar] [CrossRef] [Green Version]
- Cecchetti, L.; Tolley, N.D.; Michetti, N.; Bury, L.; Weyrich, A.S.; Gresele, P. Megakaryocytes differentially sort mRNAs for matrix metalloproteinases and their inhibitors into platelets: A mechanism for regulating synthetic events. Blood 2011, 118, 1903–1911. [Google Scholar] [CrossRef] [Green Version]
- Kammers, K.; Taub, M.A.; Rodriguez, B.; Yanek, L.R.; Ruczinski, I.; Martin, J.; Kanchan, K.; Battle, A.; Cheng, L.; Wang, Z.Z.; et al. Transcriptional profile of platelets and iPSC-derived megakaryocytes from whole-genome and RNA sequencing. Blood 2021, 137, 959–968. [Google Scholar] [CrossRef] [PubMed]
- De Kock, L.; Donck, F.V.; Thys, C.; Wijgaerts, A.; Eto, K.; Van Geet, C.; Freson, K. Combined transcriptome and proteome profiling of SRC kinase activity in healthy and E527K defective megakaryocytes. Haematologica 2021, 106, 3206. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.H.; Yao, C.; Bhan, A.; Schlaeger, T.; Keefe, J.; Rodriguez, B.A.; Hwang, S.-J.; Chen, M.-H.; Levy, D.; Johnson, A.D. Integrative genomic analysis reveals four protein biomarkers for platelet traits. Circ. Res. 2020, 127, 1182–1194. [Google Scholar] [CrossRef]
- Takaishi, K.; Takeuchi, M.; Tsukamoto, S.; Takayama, N.; Oshima, M.; Kimura, K.; Isshiki, Y.; Kayamori, K.; Hino, Y.; Oshima-Hasegawa, N.; et al. Suppressive effects of anagrelide on cell cycle progression and the maturation of megakaryocyte progenitor cell lines in human induced pluripotent stem cells. Haematologica 2020, 105, e216. [Google Scholar] [CrossRef]
- Conesa, A.; Madrigal, P.; Tarazona, S.; Gomez-Cabrero, D.; Cervera, A.; McPherson, A.; Szcześniak, M.W.; Gaffney, D.J.; Elo, L.L.; Zhang, X.; et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 2016, 17, 13. [Google Scholar] [CrossRef] [Green Version]
- Corchete, L.A.; Rojas, E.A.; Alonso-López, D.; de las Rivas, J.; Gutiérrez, N.C.; Burguillo, F.J. Systematic comparison and assessment of RNA-seq procedures for gene expression quantitative analysis. Sci. Rep. 2020, 10, 19737. [Google Scholar] [CrossRef] [PubMed]
- Geraci, F.; Saha, I.; Bianchini, M. Editorial: RNA-Seq Analysis: Methods, Applications and Challenges. Front. Genet. 2020, 11, 220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stark, R.; Grzelak, M.; Hadfield, J. RNA sequencing: The teenage years. Nat. Rev. Genet. 2019, 20, 631–656. [Google Scholar] [CrossRef]
- Londin, E.R.; Hatzimichael, E.; Loher, P.; Edelstein, L.; Shaw, C.; Delgrosso, K.; Fortina, P.; Bray, P.F.; E McKenzie, S.; Rigoutsos, I. The human platelet: Strong transcriptome correlations among individuals associate weakly with the platelet proteome. Biol. Direct 2014, 9, 3. [Google Scholar] [CrossRef] [Green Version]
- Londin, E.R.; Hatzimichael, E.; Loher, P.; Zhao, Y.; Jing, Y.; Chen, H.; Edelstein, L.C.; Nagalla, S.; Delgrosso, K.; Ertel, A.; et al. Beyond mRNAs and Mirnas: Unraveling the Full-Spectrum of the Normal Human Platelet Transcriptome through Next-Generation Sequencing. Blood 2012, 120, 3298. [Google Scholar] [CrossRef]
- Kissopoulou, A.; Jonasson, J.; Lindahl, T.L.; Osman, A. Next generation sequencing analysis of human platelet PolyA+ mRNAs and rRNA-depleted total RNA. PLoS ONE 2013, 8, e81809. [Google Scholar] [CrossRef] [PubMed]
- Supernat, A.; Popęda, M.; Pastuszak, K.; Best, M.G.; Grešner, P.; Veld, S.I.; Siek, B.; Bednarz-Knoll, N.; Rondina, M.T.; Stokowy, T.; et al. Transcriptomic landscape of blood platelets in healthy donors. Sci. Rep. 2021, 11, 15679. [Google Scholar] [CrossRef]
- Rondina, M.T.; Voora, D.; Simon, L.M.; Schwertz, H.; Harper, J.F.; Lee, O.; Bhatlekar, S.C.; Li, Q.; Eustes, A.S.; Montenont, E.; et al. Longitudinal RNA-seq analysis of the repeatability of gene expression and splicing in human platelets identifies a platelet SELP splice QTL. Circ. Res. 2020, 126, 501–516. [Google Scholar] [CrossRef] [PubMed]
- Clancy, L.; Beaulieu, L.M.; Tanriverdi, K.; Freedman, J.E. The role of RNA uptake in platelet heterogeneity. Thromb. Haemost. 2017, 117, 948–961. [Google Scholar] [CrossRef] [PubMed]
- Hille, L.; Lenz, M.; Vlachos, A.; Grüning, B.; Hein, L.; Neumann, F.; Nührenberg, T.G.; Trenk, D. Ultrastructural, transcriptional, and functional differences between human reticulated and non-reticulated platelets. J. Thromb. Haemost. 2020, 18, 2034–2046. [Google Scholar] [CrossRef]
- Picelli, S.; Björklund, Å.K.; Faridani, O.R.; Sagasser, S.; Winberg, G.; Sandberg, R. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 2013, 10, 1096–1100. [Google Scholar] [CrossRef]
- Picelli, S.; Faridani, O.R.; Björklund, Å.K.; Winberg, G.; Sagasser, S.; Sandberg, R. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 2013, 9, 171–181. [Google Scholar] [CrossRef]
- De Rop, F.; Ismail, J.N.; González-Blas, C.B.; Hulselmans, G.J.; Flerin, C.C.; Janssens, J.; Theunis, K.; Christiaens, V.M.; Wouters, J.; Marcassa, G.; et al. HyDrop enables droplet based single-cell ATAC-seq and single-cell RNA-seq using dissolvable hydrogel beads. eLife 2022, 11, e73971. [Google Scholar] [CrossRef]
- Cao, J.; Packer, J.S.; Ramani, V.; Cusanovich, D.A.; Huynh, C.; Daza, R.; Qiu, X.; Lee, C.; Furlan, S.N.; Steemers, F.J.; et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 2017, 357, 661–667. [Google Scholar] [CrossRef] [Green Version]
- Cao, J.; Cusanovich, D.A.; Ramani, V.; Aghamirzaie, D.; Pliner, H.A.; Hill, A.J.; Daza, R.M.; McFaline-Figueroa, J.L.; Packer, J.S.; Christiansen, L.; et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 2018, 361, 1380–1385. [Google Scholar] [CrossRef] [Green Version]
- Rosenberg, A.B.; Roco, C.M.; Muscat, R.A.; Kuchina, A.; Sample, P.; Yao, Z.; Graybuck, L.T.; Peeler, D.J.; Mukherjee, S.; Chen, W.; et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 2018, 360, 176–182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jovic, D.; Liang, X.; Zeng, H.; Lin, L.; Xu, F.; Luo, Y.; Correspondence, Y.; Luo, D.; Jovic, L. Single-cell RNA sequencing technologies and applications: A brief overview. Clin. Transl. Med. 2022, 12, e694. [Google Scholar] [CrossRef] [PubMed]
- Mereu, E.; Lafzi, A.; Moutinho, C.; Ziegenhain, C.; McCarthy, D.J.; Álvarez-Varela, A.; Batlle, E.; Sagar, G.D.; Lau, J.K.; Boutet, S.C.; et al. Benchmarking single-cell RNA-sequencing protocols for cell atlas projects. Nat. Biotechnol. 2020, 38, 747–755. [Google Scholar] [CrossRef] [PubMed]
- Adil, A.; Kumar, V.; Jan, A.T.; Asger, M. Single-Cell Transcriptomics: Current Methods and Challenges in Data Acquisition and Analysis. Front. Neurosci. 2021, 15, 398. [Google Scholar] [CrossRef] [PubMed]
- Andrews, T.S.; Kiselev, V.Y.; McCarthy, D.; Hemberg, M. Tutorial: Guidelines for the computational analysis of single-cell RNA sequencing data. Nat. Protoc. 2020, 16, 1–9. [Google Scholar] [CrossRef]
- Chen, G.; Ning, B.; Shi, T. Single-cell RNA-seq technologies and related computational data analysis. Front. Genet. 2019, 10, 317. [Google Scholar] [CrossRef]
- Luecken, M.D.; Theis, F.J. Current best practices in single-cell RNA-seq analysis: A tutorial. Mol. Syst. Biol. 2019, 15, e8746. [Google Scholar] [CrossRef]
- Choudry, F.A.; Bagger, F.O.; Macaulay, I.C.; Farrow, S.; Burden, F.; Kempster, C.; McKinney, H.; Olsen, L.R.; Huang, N.; Downes, K.; et al. Transcriptional characterization of human megakaryocyte polyploidization and lineage commitment. J. Thromb. Haemost. 2021, 19, 1236–1249. [Google Scholar] [CrossRef]
- Liu, C.; Wu, D.; Xia, M.; Li, M.; Sun, Z.; Shen, B.; Liu, Y.; Jiang, E.; Wang, H.; Su, P.; et al. Characterization of Cellular Heterogeneity and an Immune Subpopulation of Human Megakaryocytes. Adv. Sci. 2021, 8, 2100921. [Google Scholar] [CrossRef]
- Sun, S.; Jin, C.; Si, J.; Lei, Y.; Chen, K.; Cui, Y.; Liu, Z.; Liu, J.; Zhao, M.; Zhang, X.; et al. Single-cell analysis of ploidy and the transcriptome reveals functional and spatial divergency in murine megakaryopoiesis. Blood 2021, 138, 1211–1224. [Google Scholar] [CrossRef]
- Wang, H.; He, J.; Xu, C.; Chen, X.; Yang, H.; Shi, S.; Liu, C.; Zeng, Y.; Wu, D.; Bai, Z.; et al. Decoding Human Megakaryocyte Development. Cell Stem Cell 2021, 28, 535–549.e8. [Google Scholar] [CrossRef] [PubMed]
- Riemondy, K.A.; Ransom, M.; Alderman, C.; Gillen, A.E.; Fu, R.; Finlay-Schultz, J.; Kirkpatrick, G.D.; Di Paola, J.; Kabos, P.; Sartorius, C.A.; et al. Recovery and analysis of transcriptome subsets from pooled single-cell RNA-seq libraries. Nucleic Acids Res. 2019, 47, e20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lawrence, M.; Shahsavari, A.; Bornelöv, S.; Moreau, T.; McDonald, R.; Vallance, T.M.; Kania, K.; Paramor, M.; Baye, J.; Perrin, M.; et al. Developmental biology mapping the biogenesis of forward programmed megakaryocytes from induced pluripotent stem cells. Sci. Adv. 2022, 8, eabj8618. [Google Scholar] [CrossRef]
- Psaila, B.; Heuston, E.F.; Milojkovic, D.; Lichtenberg, J.; Anderson, S.M.; Hoffman, R.; Roberts, I.; Mead, A.J.; Bodine, D.M.; Barkas, N.; et al. A Single-Cell Approach to Unraveling Abnormal Megakaryocyte Differentiation and Function in Myelofibrosis. Blood 2017, 130, 490. [Google Scholar] [CrossRef]
- Lu, Y.C.; Sanada, C.; Xavier-Ferrucio, J.; Wang, L.; Zhang, P.-X.; Grimes, H.L.; Venkatasubramanian, M.; Chetal, K.; Aronow, B.; Salomonis, N.; et al. The Molecular Signature of Megakaryocyte-Erythroid Progenitors Reveals a Role for the Cell Cycle in Fate Specification. Cell Rep. 2018, 25, 2083–2093.e4. [Google Scholar] [CrossRef] [Green Version]
- Psaila, B.; Wang, G.; Rodriguez-Meira, A.; Li, R.; Heuston, E.F.; Murphy, L.; Yee, D.; Hitchcock, I.S.; Sousos, N.; O’Sullivan, J.; et al. Single-Cell Analyses Reveal Megakaryocyte-Biased Hematopoiesis in Myelofibrosis and Identify Mutant Clone-Specific Targets. Mol. Cell 2020, 78, 477–492.e8. [Google Scholar] [CrossRef]
- Estevez, B.; Borst, S.; Jarocha, D.J.; Sudunagunta, V.S.; Gonzalez, M.; Garifallou, J.; Hakonarson, H.; Gao, P.; Tan, K.; Liu, P.P.; et al. RUNX-1 haploinsufficiency causes a marked deficiency of megakaryocyte-biased hematopoietic progenitor cells. Blood 2021, 137, 2662–2675. [Google Scholar] [CrossRef]
- Tolhurst, G.; Carter, R.N.; Miller, N.; Mahaut-Smith, M.P. Purification of Native Bone Marrow Megakaryocytes for Studies of Gene Expression. Methods Mol. Biol. 2012, 788, 259–273. [Google Scholar] [CrossRef]
- Jia, E.; Shi, H.; Wang, Y.; Zhou, Y.; Liu, Z.; Pan, M.; Bai, Y.; Zhao, X.; Ge, Q. Optimization of library preparation based on SMART for ultralow RNA-seq in mice brain tissues. BMC Genom. 2021, 22, 809. [Google Scholar] [CrossRef]
- Lesyk, G.; Jurasz, P. Advances in Platelet Subpopulation Research. Front. Cardiovasc. Med. 2019, 6, 138. [Google Scholar] [CrossRef] [Green Version]
- Lentaigne, C.; Freson, K.; Laffan, M.A.; Turro, E.; Ouwehand, W.H. Inherited platelet disorders: Toward DNA-based diagnosis. Blood 2016, 127, 2814–2823. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Megy, K.; Downes, K.; Simeoni, I.; Bury, L.; Morales, J.; Mapeta, R.; Bellissimo, D.B.; Bray, P.F.; Goodeve, A.C.; Gresele, P.; et al. Curated disease-causing genes for bleeding, thrombotic, and platelet disorders: Communication from the SSC of the ISTH. J. Thromb. Haemost. 2019, 17, 1253–1260. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sivapalaratnam, S.; Collins, J.; Gomez, K. Diagnosis of inherited bleeding disorders in the genomic era. Br. J. Haematol. 2017, 179, 363–376. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kahr, W.H.A.; Hinckley, J.; Li, L.; Schwertz, H.; Christensen, H.; Rowley, J.W.; Pluthero, F.; Urban, D.; Fabbro, S.; Nixon, B.; et al. Mutations in NBEAL2, encoding a BEACH protein, cause gray platelet syndrome. Nat. Genet. 2011, 43, 738–740. [Google Scholar] [CrossRef]
- Sims, M.C.; Mayer, L.; Collins, J.H.; Bariana, T.K.; Megy, K.; Lavenu-Bombled, C.; Seyres, D.; Kollipara, L.; Burden, F.S.; Greene, D.; et al. Novel manifestations of immune dysregulation and granule defects in gray platelet syndrome. Blood 2020, 136, 1956–1967. [Google Scholar] [CrossRef]
- Noetzli, L.; Lo, R.; Sherick, A.L.; Callaghan, M.; Noris, P.; Savoia, A.; Rajpurkar, M.; Jones, K.; Gowan, K.; Balduini, C.L.; et al. Germline mutations in ETV6 are associated with thrombocytopenia, red cell macrocytosis and predisposition to lymphoblastic leukemia. Nat. Genet. 2015, 47, 535–538. [Google Scholar] [CrossRef] [Green Version]
- Heremans, J.; Garcia-Perez, J.E.; Turro, E.; Schlenner, S.M.; Casteels, I.; Collin, R.; de Zegher, F.; Greene, D.; Humblet-Baron, S.; Lesage, S.; et al. Abnormal differentiation of B cells and megakaryocytes in patients with Roifman syndrome. J. Allergy Clin. Immunol. 2018, 142, 630–646. [Google Scholar] [CrossRef] [Green Version]
- Brechtmann, F.; Mertes, C.; Matusevičiūtė, A.; Yépez, V.A.; Avsec, Ž.; Herzog, M.; Bader, D.M.; Prokisch, H.; Gagneur, J. OUTRIDER: A Statistical Method for Detecting Aberrantly Expressed Genes in RNA Sequencing Data. Am. J. Hum. Genet. 2018, 103, 907. [Google Scholar] [CrossRef] [Green Version]
- Mertes, C.; Scheller, I.F.; Yépez, V.A.; Çelik, M.H.; Liang, Y.; Kremer, L.S.; Gusic, M.; Prokisch, H.; Gagneur, J. Detection of aberrant splicing events in RNA-seq data using FRASER. Nat. Commun. 2021, 12, 529. [Google Scholar] [CrossRef]
- Yépez, V.A.; Mertes, C.; Müller, M.F.; Klaproth-Andrade, D.; Wachutka, L.; Frésard, L.; Gusic, M.; Scheller, I.F.; Goldberg, P.F.; Prokisch, H.; et al. Detection of aberrant gene expression events in RNA sequencing data. Nat. Protoc. 2021, 16, 1276–1296. [Google Scholar] [CrossRef]
- Macaulay, I.C.; Haerty, W.; Kumar, P.; I Li, Y.; Hu, T.X.; Teng, M.J.; Goolam, M.; Saurat, N.; Coupland, P.; Shirley, L.M.; et al. G&T-seq: Parallel sequencing of single-cell genomes and transcriptomes. Nat. Methods 2015, 12, 519–522. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Zhang, B.; LaFave, L.M.; Earl, A.S.; Chiang, Z.; Hu, Y.; Ding, J.; Brack, A.; Kartha, V.K.; Tay, T.; et al. Chromatin Potential Identified by Shared Single-Cell Profiling of RNA and Chromatin. Cell 2020, 183, 1103–1116.e20. [Google Scholar] [CrossRef] [PubMed]
- Swanson, E.; Lord, C.; Reading, J.; Heubeck, A.T.; Genge, P.C.; Thomson, Z.; DA Weiss, M.; Li, X.-J.; Savage, A.K.; Green, R.R.; et al. Simultaneous trimodal single-cell measurement of transcripts, epitopes, and chromatin accessibility using tea-seq. eLife 2021, 10, e63632. [Google Scholar] [CrossRef]
- Leon-Mimila, P.; Wang, J.; Huertas-Vazquez, A. Relevance of Multi-Omics Studies in Cardiovascular Diseases. Front. Cardiovasc. Med. 2019, 6, 91. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turro, E.; Astle, W.J.; Megy, K.; Gräf, S.; Greene, D.; Shamardina, O.; Allen, H.L.; Sanchis-Juan, A.; Frontini, M.; Thys, C.; et al. Whole-genome sequencing of patients with rare diseases in a national health system. Nature 2020, 583, 96–102. [Google Scholar] [CrossRef]
- Greene, D.; Richardson, S.; Turro, E. A Fast Association Test for Identifying Pathogenic Variants Involved in Rare Diseases. Am. J. Hum. Genet. 2017, 101, 104–114. [Google Scholar] [CrossRef] [Green Version]
- Jaganathan, K.; Panagiotopoulou, S.K.; McRae, J.F.; Darbandi, S.F.; Knowles, D.; Li, Y.I.; Kosmicki, J.A.; Arbelaez, J.; Cui, W.; Schwartz, G.B.; et al. Predicting Splicing from Primary Sequence with Deep Learning. Cell 2018, 176, 535–548.e24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murdock, D.R.; Dai, H.; Burrage, L.C.; Rosenfeld, J.A.; Ketkar, S.; Müller, M.F.; Yépez, V.A.; Gagneur, J.; Liu, P.; Chen, S.; et al. Transcriptome-directed analysis for Mendelian disease diagnosis overcomes limitations of conventional genomic testing. J. Clin. Investig. 2021, 131, e141500. [Google Scholar] [CrossRef] [PubMed]

| Plate-Based | Droplet-Based |
|---|---|
| Higher sensitivity | Higher cell throughput |
| Better for large and fragile cells | Lower labor intensity |
| Cheaper setup cost | Cheaper cost per cell |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Wispelaere, K.; Freson, K. The Analysis of the Human Megakaryocyte and Platelet Coding Transcriptome in Healthy and Diseased Subjects. Int. J. Mol. Sci. 2022, 23, 7647. https://doi.org/10.3390/ijms23147647
De Wispelaere K, Freson K. The Analysis of the Human Megakaryocyte and Platelet Coding Transcriptome in Healthy and Diseased Subjects. International Journal of Molecular Sciences. 2022; 23(14):7647. https://doi.org/10.3390/ijms23147647
Chicago/Turabian StyleDe Wispelaere, Koenraad, and Kathleen Freson. 2022. "The Analysis of the Human Megakaryocyte and Platelet Coding Transcriptome in Healthy and Diseased Subjects" International Journal of Molecular Sciences 23, no. 14: 7647. https://doi.org/10.3390/ijms23147647
APA StyleDe Wispelaere, K., & Freson, K. (2022). The Analysis of the Human Megakaryocyte and Platelet Coding Transcriptome in Healthy and Diseased Subjects. International Journal of Molecular Sciences, 23(14), 7647. https://doi.org/10.3390/ijms23147647






