Integrated Analysis of Microarray, Small RNA, and Degradome Datasets Uncovers the Role of MicroRNAs in Temperature-Sensitive Genic Male Sterility in Wheat
Abstract
:1. Introduction
2. Results
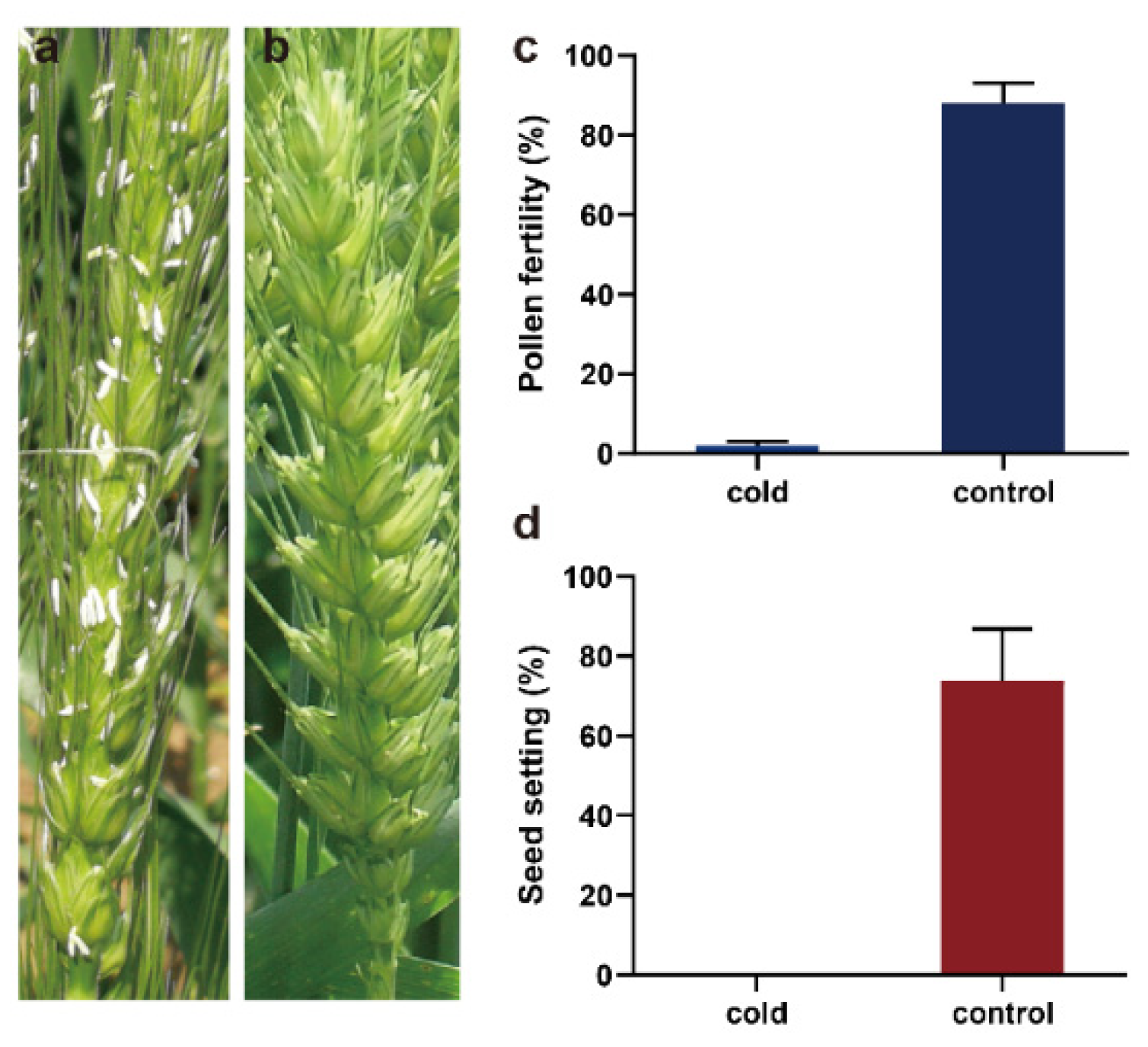
2.1. Pollen Development Was Defective in BS366 under Cold Conditions
2.2. Construction of the Co-Expression Network for All of the Expressed Genes
2.3. Expression and Function Analysis of Genes in the Blue and Brown Modules
2.4. The Differentially Expressed Genes between Cold and Control Conditions
2.5. Differential Expression of Genic Male Sterility Genes
2.6. Mapping of sRNA Reads to the Reference Genome
2.7. Differentially Expressed miRNA Identification, Target Prediction and Function Analysis
2.8. The Antagonistic miRNA–Target Pairs Identified during Three Stages
2.9. Identification of miRNA Target Genes via Degradome Sequencing
3. Discussion
3.1. The Potential Role of Conserved miRNAs in the Anther Development of Wheat
3.2. Impaired Cell Plate Formation during Meiosis
3.3. Suppression of Lipid Metabolism Was Responsible for Male Sterility in BS366
3.4. The Auxin Signaling Pathway Is Essential to Male Sterility in BS366
4. Materials and Methods
4.1. Plant Materials
4.2. Phenotypic Analysis of BS366
4.3. Data Acquisition
4.4. Bioinformatics Analysis
4.5. Weighted Gene Co-Expression Network Analysis
4.6. Sample Preparation, RNA Isolation, and Real-Time qRT–PCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Longin, C.F.; Muhleisen, J.; Maurer, H.P.; Zhang, H.; Gowda, M.; Reif, J.C. Hybrid breeding in autogamous cereals. Theor. Appl. Genet. 2012, 125, 1087–1096. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Zhang, D. Molecular Control of Male Fertility for Crop Hybrid Breeding. Trends Plant Sci. 2018, 23, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, J.; Chen, S.; Heng, Y.; Chen, Z.; Yang, J.; Zhou, K.; Pei, J.; He, H.; Deng, X.W.; et al. Poaceae-specific MS1 encodes a phospholipid-binding protein for male fertility in bread wheat. Proc. Natl. Acad. Sci. USA 2017, 114, 12614–12619. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tucker, E.J.; Baumann, U.; Kouidri, A.; Suchecki, R.; Baes, M.; Garcia, M.; Okada, T.; Dong, C.; Wu, Y.; Sandhu, A.; et al. Molecular identification of the wheat male fertility gene Ms1 and its prospects for hybrid breeding. Nat. Commun. 2017, 8, 869. [Google Scholar] [CrossRef]
- Xia, C.; Zhang, L.C.; Zou, C.; Gu, Y.Q.; Duan, J.L.; Zhao, G.Y.; Wu, J.J.; Liu, Y.; Fang, X.H.; Gao, L.F.; et al. A TRIM insertion in the promoter of Ms2 causes male sterility in wheat. Nat. Commun. 2017, 8, 15407. [Google Scholar] [CrossRef] [Green Version]
- Ni, F.; Qi, J.; Hao, Q.; Lyu, B.; Luo, M.C.; Wang, Y.; Chen, F.; Wang, S.; Zhang, C.; Epstein, L.; et al. Wheat Ms2 encodes for an orphan protein that confers male sterility in grass species. Nat. Commun. 2017, 8, 15121. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Guo, J.; Zhang, C.; Zheng, W.; Song, Y.; Wang, Y. Identification of Differentially Expressed miRNAs between a Wheat K-type Cytoplasmic Male Sterility Line and Its Near-Isogenic Restorer Line. Plant Cell Physiol. 2019, 60, 1604–1618. [Google Scholar] [CrossRef]
- Chen, L.; Heikkinen, L.; Wang, C.; Yang, Y.; Sun, H.; Wong, G. Trends in the development of miRNA bioinformatics tools. Brief Bioinform. 2019, 20, 1836–1852. [Google Scholar] [CrossRef] [Green Version]
- Ferreira e Silva, G.F.; Silva, E.M.; Azevedo Mda, S.; Guivin, M.A.; Ramiro, D.A.; Figueiredo, C.R.; Carrer, H.; Peres, L.E.; Nogueira, F.T. microRNA156-targeted SPL/SBP box transcription factors regulate tomato ovary and fruit development. Plant J. 2014, 78, 604–618. [Google Scholar] [CrossRef]
- Fan, Y.; Yang, J.; Mathioni, S.M.; Yu, J.; Shen, J.; Yang, X.; Wang, L.; Zhang, Q.; Cai, Z.; Xu, C.; et al. PMS1T, producing phased small-interfering RNAs, regulates photoperiod-sensitive male sterility in rice. Proc. Natl. Acad. Sci. USA 2016, 113, 15144–15149. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, Z.H.; Zhang, L.P.; Xu, C.G.; Yuan, S.H.; Zhang, F.T.; Zheng, Y.L.; Zhao, C.P. Uncovering Small RNA-Mediated Responses to Cold Stress in a Wheat Thermosensitive Genic Male-Sterile Line by Deep Sequencing. Plant Physiol. 2012, 159, 721–738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ding, Y.; Ma, Y.; Liu, N.; Xu, J.; Hu, Q.; Li, Y.; Wu, Y.; Xie, S.; Zhu, L.; Min, L.; et al. microRNAs involved in auxin signalling modulate male sterility under high-temperature stress in cotton (Gossypium hirsutum). Plant J. 2017, 91, 977–994. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, Y.; Guo, Z.; Li, L. Evolutionary conservation of microRNA regulatory programs in plant flower development. Dev. Biol. 2013, 380, 133–144. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ding, X.; Guo, J.; Zhang, Q.; Yu, L.; Zhao, T.; Yang, S. Heat-Responsive miRNAs Participate in the Regulation of Male Fertility Stability in Soybean CMS-Based F(1) under High Temperature Stress. Int. J. Mol. Sci. 2021, 22, 2446. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Peral, M.M.; Li, J.; Li, Y.; Allen, R.S.; Schnippenkoetter, W.; Ohms, S.; White, R.G.; Millar, A.A. The microRNA159-regulated GAMYB-like genes inhibit growth and promote programmed cell death in Arabidopsis. Plant Physiol. 2010, 154, 757–771. [Google Scholar] [CrossRef] [Green Version]
- Li, Z.; An, X.; Zhu, T.; Yan, T.; Wu, S.; Tian, Y.; Li, J.; Wan, X. Discovering and Constructing ceRNA-miRNA-Target Gene Regulatory Networks during Anther Development in Maize. Int. J. Mol. Sci. 2019, 20, 3480. [Google Scholar] [CrossRef] [Green Version]
- Yu, J.; Meng, Z.; Liang, W.; Behera, S.; Kudla, J.; Tucker, M.R.; Luo, Z.; Chen, M.; Xu, D.; Zhao, G.; et al. A Rice Ca2+ Binding Protein Is Required for Tapetum Function and Pollen Formation. Plant Physiol. 2016, 172, 1772–1786. [Google Scholar] [CrossRef] [Green Version]
- Wu, M.F.; Tian, Q.; Reed, J.W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar] [CrossRef] [Green Version]
- Aukerman, M.J.; Sakai, H. Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell 2003, 15, 2730–2741. [Google Scholar] [CrossRef] [Green Version]
- Chen, X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 2004, 303, 2022–2025. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, Z.; Hu, X.; Cai, W.; Huang, W.; Zhou, X.; Luo, Q.; Yang, H.; Wang, J.; Huang, J. Arabidopsis miR171-targeted scarecrow-like proteins bind to GT cis-elements and mediate gibberellin-regulated chlorophyll biosynthesis under light conditions. PLoS Genet. 2014, 10, e1004519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hewezi, T.; Baum, T.J. Complex feedback regulations govern the expression of miRNA396 and its GRF target genes. Plant Signal. Behav. 2012, 7, 749–751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, L.; Sun, G.; Shi, C.; Sun, D. Transcriptome analysis reveals new microRNAs-mediated pathway involved in anther development in male sterile wheat. BMC Genom. 2018, 19, 333. [Google Scholar] [CrossRef] [Green Version]
- Wan, X.; Wu, S.; Li, Z.; Dong, Z.; An, X.; Ma, B.; Tian, Y.; Li, J. Maize Genic Male-Sterility Genes and Their Applications in Hybrid Breeding: Progress and Perspectives. Mol. Plant 2019, 12, 321–342. [Google Scholar] [CrossRef] [Green Version]
- Dong, X.; Hong, Z.; Sivaramakrishnan, M.; Mahfouz, M.; Verma, D.P. Callose synthase (CalS5) is required for exine formation during microgametogenesis and for pollen viability in Arabidopsis. Plant J. 2005, 42, 315–328. [Google Scholar] [CrossRef]
- Suzuki, T.; Narciso, J.O.; Zeng, W.; van de Meene, A.; Yasutomi, M.; Takemura, S.; Lampugnani, E.R.; Doblin, M.S.; Bacic, A.; Ishiguro, S. KNS4/UPEX1: A Type II Arabinogalactan β-(1,3)-Galactosyltransferase Required for Pollen Exine Development. Plant Physiol. 2017, 173, 183–205. [Google Scholar] [CrossRef] [Green Version]
- Sun, M.X.; Huang, X.Y.; Yang, J.; Guan, Y.F.; Yang, Z.N. Arabidopsis RPG1 is important for primexine deposition and functions redundantly with RPG2 for plant fertility at the late reproductive stage. Plant Reprod. 2013, 26, 83–91. [Google Scholar] [CrossRef]
- Addo-Quaye, C.; Miller, W.; Axtell, M.J. CleaveLand: A pipeline for using degradome data to find cleaved small RNA targets. Bioinformatics 2009, 25, 130–131. [Google Scholar] [CrossRef]
- Ma, Y.Z.; Min, L.; Wang, M.J.; Wang, C.Z.; Zhao, Y.L.; Li, Y.Y.; Fang, Q.D.; Wu, Y.L.; Xie, S.; Ding, Y.H.; et al. Disrupted Genome Methylation in Response to High Temperature Has Distinct Affects on Microspore Abortion and Anther Indehiscence. Plant Cell 2018, 30, 1387–1403. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Alonso-Peral, M.; Wong, G.; Wang, M.B.; Millar, A.A. Ubiquitous miR159 repression of MYB33/65 in Arabidopsis rosettes is robust and is not perturbed by a wide range of stresses. BMC Plant Biol. 2016, 16, 179. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Millar, A.A.; Gubler, F. The Arabidopsis GAMYB-like genes, MYB33 and MYB65, are microRNA-regulated genes that redundantly facilitate anther development. Plant Cell 2005, 17, 705–721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Allen, R.S.; Li, J.; Stahle, M.I.; Dubroué, A.; Gubler, F.; Millar, A.A. Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc. Natl. Acad. Sci. USA 2007, 104, 16371–16376. [Google Scholar] [CrossRef] [Green Version]
- Hirokawa, N. Kinesin and dynein superfamily proteins and the mechanism of organelle transport. Science 1998, 279, 519–526. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, Y.R.; Giang, H.M.; Liu, B. A novel plant kinesin-related protein specifically associates with the phragmoplast organelles. Plant Cell 2001, 13, 2427–2439. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jürgens, G. Cytokinesis in higher plants. Annu. Rev. Plant Biol. 2005, 56, 281–299. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Zhang, L.; Yang, D.; Zhao, C.; Zheng, Y. Cold stress contributes to aberrant cytokinesis during male meiosis I in a wheat thermosensitive genic male sterile line. Plant Cell Environ. 2011, 34, 389–405. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.R.; Liu, B. The rise and fall of the phragmoplast microtubule array. Curr. Opin. Plant Biol. 2013, 16, 757–763. [Google Scholar] [CrossRef] [Green Version]
- Wan, X.; Wu, S.; Li, Z.; An, X.; Tian, Y. Lipid Metabolism: Critical Roles in Male Fertility and Other Aspects of Reproductive Development in Plants. Mol. Plant 2020, 13, 955–983. [Google Scholar] [CrossRef]
- Kunst, L.; Samuels, L. Plant cuticles shine: Advances in wax biosynthesis and export. Curr. Opin. Plant Biol. 2009, 12, 721–727. [Google Scholar] [CrossRef]
- Domínguez, E.; Heredia-Guerrero, J.A.; Heredia, A. Plant cutin genesis: Unanswered questions. Trends Plant Sci. 2015, 20, 551–558. [Google Scholar] [CrossRef] [PubMed]
- Yaacob, S.; Jamil, R.Z.R.; Suah, F.B.M. Sporopollenin based materials as a versatile choice for the detoxification of environmental pollutants-A review. Int. J. Biol. Macromol. 2022, 207, 990–1004. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Yu, S.I.; Jung, S.H.; Lee, B.H.; Suh, M.C. The F-Box Protein SAGL1 and ECERIFERUM3 Regulate Cuticular Wax Biosynthesis in Response to Changes in Humidity in Arabidopsis. Plant Cell 2019, 31, 2223–2240. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bu, R.F.; Wang, R.H.; Wei, Q.C.; Hu, H.Y.; Sun, H.L.; Song, P.W.; Yu, Y.G.; Liu, Q.L.; Zheng, Z.C.; Li, T.; et al. Silencing of glycerol-3-phosphate acyltransferase 6 (GPAT6) gene using a newly established virus induced gene silencing (VIGS) system in cucumber alleviates autotoxicity mimicked by cinnamic acid (CA). Plant Soil 2019, 438, 329–346. [Google Scholar] [CrossRef]
- Chang, Z.; Chen, Z.; Yan, W.; Xie, G.; Lu, J.; Wang, N.; Lu, Q.; Yao, N.; Yang, G.; Xia, J.; et al. An ABC transporter, OsABCG26, is required for anther cuticle and pollen exine formation and pollen-pistil interactions in rice. Plant Sci. 2016, 253, 21–30. [Google Scholar] [CrossRef] [Green Version]
- Vanneste, S.; Friml, J. Auxin: A trigger for change in plant development. Cell 2009, 136, 1005–1016. [Google Scholar] [CrossRef]
- Sundberg, E.; Østergaard, L. Distinct and dynamic auxin activities during reproductive development. Cold Spring Harb. Perspect. Biol. 2009, 1, a001628. [Google Scholar] [CrossRef] [Green Version]
- Uchida, N.; Takahashi, K.; Iwasaki, R.; Yamada, R.; Yoshimura, M.; Endo, T.A.; Kimura, S.; Zhang, H.; Nomoto, M.; Tada, Y.; et al. Chemical hijacking of auxin signaling with an engineered auxin-TIR1 pair. Nat. Chem. Biol. 2018, 14, 299–305. [Google Scholar] [CrossRef]
- Okushima, Y.; Overvoorde, P.J.; Arima, K.; Alonso, J.M.; Chan, A.; Chang, C.; Ecker, J.R.; Hughes, B.; Lui, A.; Nguyen, D.; et al. Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: Unique and overlapping functions of ARF7 and ARF19. Plant Cell 2005, 17, 444–463. [Google Scholar] [CrossRef] [Green Version]
- Sun, G. MicroRNAs and their diverse functions in plants. Plant Mol. Biol. 2012, 80, 17–36. [Google Scholar] [CrossRef]
- Cecchetti, V.; Altamura, M.M.; Falasca, G.; Costantino, P.; Cardarelli, M. Auxin regulates Arabidopsis anther dehiscence, pollen maturation, and filament elongation. Plant Cell 2008, 20, 1760–1774. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, X.F.; Wang, B.; Feng, Y.F.; Xue, J.S.; Qian, X.X.; Liu, S.Q.; Zhou, J.; Yu, Y.H.; Yang, N.Y.; Xu, P.; et al. AUXIN RESPONSE FACTOR17 Directly Regulates MYB108 for Anther Dehiscence. Plant Physiol. 2019, 181, 645–655. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mallory, A.C.; Bartel, D.P.; Bartel, B. MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 2005, 17, 1360–1375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, J.; Tian, L.; Sun, M.X.; Huang, X.Y.; Zhu, J.; Guan, Y.F.; Jia, Q.S.; Yang, Z.N. AUXIN RESPONSE FACTOR17 is essential for pollen wall pattern formation in Arabidopsis. Plant Physiol. 2013, 162, 720–731. [Google Scholar] [CrossRef] [Green Version]
- Lakehal, A.; Chaabouni, S.; Cavel, E.; Le Hir, R.; Ranjan, A.; Raneshan, Z.; Novák, O.; Păcurar, D.I.; Perrone, I.; Jobert, F.; et al. A Molecular Framework for the Control of Adventitious Rooting by TIR1/AFB2-Aux/IAA-Dependent Auxin Signaling in Arabidopsis. Mol. Plant 2019, 12, 1499–1514. [Google Scholar] [CrossRef]
- Liu, Y.J.; Li, D.; Gong, J.; Wang, Y.B.; Chen, Z.B.; Pang, B.S.; Chen, X.C.; Gao, J.G.; Yang, W.B.; Zhang, F.T.; et al. Comparative transcriptome and DNA methylation analysis in temperature-sensitive genic male sterile wheat BS366. BMC Genom. 2021, 22, 911. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Li, R.; Li, Y.; Kristiansen, K.; Wang, J. SOAP: Short oligonucleotide alignment program. Bioinformatics 2008, 24, 713–714. [Google Scholar] [CrossRef] [Green Version]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 2011, 39, D152–D157. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.K.; Feng, Z.X.; Wang, X.; Wang, X.W.; Zhang, X.G. DEGseq: An R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Ji, Y.; Chen, P.; Chen, J.; Pennerman, K.K.; Liang, X.; Yan, H.; Zhou, S.; Feng, G.; Wang, C.; Yin, G.; et al. Combinations of Small RNA, RNA, and Degradome Sequencing Uncovers the Expression Pattern of microRNA−mRNA Pairs Adapting to Drought Stress in Leaf and Root of Dactylis glomerata L. Int. J. Mol. Sci. 2018, 19, 3114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]












Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Li, D.; Zhang, S.; Zhang, L.; Gong, J.; Li, Y.; Chen, J.; Zhang, F.; Liao, X.; Chen, Z.; et al. Integrated Analysis of Microarray, Small RNA, and Degradome Datasets Uncovers the Role of MicroRNAs in Temperature-Sensitive Genic Male Sterility in Wheat. Int. J. Mol. Sci. 2022, 23, 8057. https://doi.org/10.3390/ijms23158057
Liu Y, Li D, Zhang S, Zhang L, Gong J, Li Y, Chen J, Zhang F, Liao X, Chen Z, et al. Integrated Analysis of Microarray, Small RNA, and Degradome Datasets Uncovers the Role of MicroRNAs in Temperature-Sensitive Genic Male Sterility in Wheat. International Journal of Molecular Sciences. 2022; 23(15):8057. https://doi.org/10.3390/ijms23158057
Chicago/Turabian StyleLiu, Yongjie, Dan Li, Shengquan Zhang, Liping Zhang, Jie Gong, Yanhong Li, Jiamin Chen, Fengting Zhang, Xiangzheng Liao, Zhaobo Chen, and et al. 2022. "Integrated Analysis of Microarray, Small RNA, and Degradome Datasets Uncovers the Role of MicroRNAs in Temperature-Sensitive Genic Male Sterility in Wheat" International Journal of Molecular Sciences 23, no. 15: 8057. https://doi.org/10.3390/ijms23158057




