Melatonin: Regulation of Viral Phase Separation and Epitranscriptomics in Post-Acute Sequelae of COVID-19
Abstract
1. Introduction
2. Viral Persistence May Modulate Innate Immune Response
3. SARS-CoV-2 Proteins Phase Separation Disrupt Host Biomolecular Condensates That Regulate Gene Expression and Interferon Immune Signaling
3.1. SARS-CoV-2 Evades Host Interferon Responses by Inhibition of the JAK-STAT Signaling Pathway in a Time-Sensitive Manner
3.2. The Effects of Melatonin Preactivation of the IFN Signaling Response Are Time- and Dose-Dependent
3.3. SARS-CoV-2 Molecular Condensates Are Viral Replication Factories That Enhance Immune Suppression and Evasion
3.4. Interactions between Viral Intrinsically Disordered Regions and Host Biomolecular Condensates Enhance Viral Replication by Exploiting Stress Responses
3.5. SARS-CoV-2 Nucleocapsid Enlists Nonstructural Protein 1 to Shut down Host mRNA Translation and Modulate Expression of IFN Genes
4. Melatonin Is an Ancient Molecule That Can Regulate Virus Phase Separation
4.1. ATP and RNA Controls N Protein Phase Separation in a Biphasic Manner
4.2. Elevated Extracellular ATP May Reduce Viral Replication
5. Melatonin Protects Mitochondria and ATP Production to Inhibit N Protein Phase Separation
5.1. Melatonin Rescues Mitochondrial Membrane Potential from SARS-CoV-2 Envelope Protein-Induced Depolarization
5.1.1. Membrane Depolarization Impairs Oxidative Phosphorylation and Cation Homeostasis
5.1.2. Viroporin Ion Channel Activities May Regulate Virus Phase Separation
5.2. Melatonin Attenuates Membrane Depolarization and Balances Ion Homeostasis by Antioxidant-Dependent and -Independent Mechanisms to Protect Mitochondria and Lymphocytes during Viral Infection and PASC
5.3. Melatonin Protects Mitochondria Cristae Morphology and ATP Production via Antioxidant-Dependent and -Independent Mechanisms
5.3.1. Melatonin Suppresses Aerobic Glycolysis to Enhance Oxidative Phosphorylation
5.3.2. Melatonin and Metabolites Preserve Cardiolipin Function in Cristae by Preventing Lipid Peroxidation Cascades
5.4. Melatonin Targets NLRP3 Inflammasomes via Cardiolipin and DDX3X
5.5. DDX3X Is a “Double-Edged Sword” That Mediates Host Antiviral Immunity and Viral Replication
5.6. N Protein Must Phase Separation to Target G3BP1 and Disassemble Stress Granules
5.7. The Formation of “Viral Factories” by N Protein LLPS Is Tuned by Phosphorylation
6. Melatonin Disrupts Formation of “Viral Factories” by Regulating GSK-3 Phosphorylation of N Protein Condensates
6.1. GSK-3 Phosphorylation of Gle1A Mediates Stress Granule Disassembly via Inhibition of DDX3X
6.2. Melatonin Inhibits GSK-3 Gene Expression and Promotes Phosphorylation to Deactivate GSK-3
7. Melatonin Regulates SARS-CoV-2-Mediated Crosstalk between the Epitranscriptome and Transcriptome via m6A Modifications and LINE1 Suppression
7.1. SARS-CoV-2 Derepression of LINE1 May Induce Genomic Instability That Exacerbates Disease Severity and Prolongs Recovery
7.1.1. Can SARS-CoV-2 Be Reverse-Transcribed to Form Viral-Host Chimeric Transcripts?
7.1.2. LINE1 Derepression and Global Hypomethylation May Be Associated with SARS-CoV-2-Mediated Pathologies
7.1.3. LINE1 Derepression and Global Hypomethylation Are Induced by Mitochondrial Dysfunction
7.2. Melatonin Suppresses LINE1 Derepression via Antioxidant-Dependent and -Independent Mechanisms
7.2.1. Oxidative Stress Activates LINE1 ORF1 Proteins to Associate with Stress Granules
7.2.2. Melatonin May Inhibit LINE1 Expression and Derepression via Regulation of ORF1 Protein Phase Separation
7.2.3. ORF1p Phase Separation Formation of Dynamic Condensates Is a Requisite for L1 Retrotransposition
7.2.4. Melatonin Enhances Complex I Functions, Reduces Oxidative Stress, and Regulates DNA Damage Response Elements to Restrain L1 Retrotransposition
7.3. m6A Modifications Regulate SARS-CoV-2-Mediated LINE1 Derepression
7.4. Viral Epitranscriptomics: The Hijacking of Host m6A for Viral Infection and Replication
7.5. Is m6A a Positive or Negative Regulator of SARS-CoV-2 Replication?
7.6. Melatonin Phosphorylation of GSK-3 Increases the m6A Demethylase FTO
7.7. SARS-CoV-2 Suppresses Innate Immune Responses by Hijacking DDXs to Enhance ALKBH5 and METTL3
7.8. G3BP1 Is Repelled by m6A METTL3 Modification, but Associates with YTHDF Proteins to Form Stress Granules
7.9. Melatonin Modulates the Expression of m6A METTL3 Methyltransferase in a Context-Dependent, Pleiotropic Manner
| m6A Modification Enzymes | Model/Description | Melatonin Doses | Melatonin’s Effects | Reference |
|---|---|---|---|---|
| METTL3/METT14 | Epididymal WAT/Alimentary obesity mouse model | 20 mg/kg IP injection × 14 days | Reduced transcription. | [861] |
| ALKBH5 | Epididymal WAT/Alimentary obesity mouse model | 20 mg/kg IP injection × 14 days | Reduced transcription. | [861] |
| FTO/YTHDF2 | Epididymal WAT/Alimentary obesity mouse model | 20 mg/kg IP injection × 14 days | Significantly increased transcriptions. | [861] |
| METTL3 | MSC-derived EV/SCI mouse model | 1 μmol/L for 48 h. | Reduced transcription | [862] |
| METTL3 | Long-term cultured ESCs | 10 μM × 90 days. | Maintained pluripotency of ESCs by significantly reducing METTL3 levels. | [863] |
| METTL3 | Mouse SSC Cr (VI)-induced m6A downregulation | 50 μM pretreatment | Restored METTL3 levels, attenuated m6A modification reduction. | [872] |
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 4-HNE | 4-hydroxynonenal |
| ALKBH5 | alpha-ketoglutarate-dependent dioxygenase alkB homolog 5 |
| Ca2+ | calcium |
| CL | cardiolipin |
| CNS | central nervous system |
| DNA | deoxyribonucleic acid |
| DMR | differentially methylated region |
| EBOV | Ebola virus |
| ER | endoplasmic reticulum |
| FTO | fat mass and obesity-associated protein |
| GSK | glycogen synthase kinase |
| HPI | hour post-infection |
| IB | inclusion body |
| IBM | inner boundary membrane |
| IDR | intrinsically disordered region |
| IFN | interferon |
| IMM | inner mitochondrial membrane |
| I.P. | intraperitoneal |
| ISG | interferon-stimulated gene |
| ISR | integrated stress response |
| JAK-STAT | Janus kinase-signal transducers and activators of transcription |
| K+ | potassium ion |
| LINE1, L1 | long interspersed nuclear element 1 |
| m6A | N6-methyladenosine |
| METTL3 | methyltransferase 3 |
| METTL14 | methyltransferase 14 |
| mPTP | mitochondrial permeability transition pore |
| mRNA | messenger RNA |
| NLRP3 | NLR pyrin domain containing 3 |
| Nrf2 | nuclear factor erythroid 2-related factor |
| Nsp1 | nonstructural protein 1 |
| PASC | post-acute sequelae of COVID-19 |
| PBMC | peripheral blood mononuclear cells |
| PI | post-infection |
| RdRp | RNA-dependent RNA polymerase |
| RBP | RNA-binding protein |
| RIRR | ROS-induced ROS release |
| RNA | ribonucleic acid |
| RNA-seq | RNA sequencing |
| RNP | ribonucleoprotein |
| ROS | reactive oxygen species |
| RSV | respiratory syncytial virus |
| RT | reverse transcriptase |
| RTE | retrotransposable element, retrotransposon |
| SG | stress granule |
| S/R | serine/arginine |
| TE | transposable element |
| VSV | vesicular stomatitis virus |
| YTHDF2 | YTH-domain family 2 |
| ZIKV | Zika virus |
References
- Rössler, A.; Riepler, L.; Bante, D.; von Laer, D.; Kimpel, J. SARS-CoV-2 Omicron Variant Neutralization in Serum from Vaccinated and Convalescent Persons. N. Engl. J. Med. 2022, 386, 698–700. [Google Scholar] [CrossRef] [PubMed]
- Quaglia, F.; Salladini, E.; Carraro, M.; Minervini, G.; Tosatto, S.C.E.; Le Mercier, P. SARS-CoV-2 Variants Preferentially Emerge at Intrinsically Disordered Protein Sites Helping Immune Evasion. FEBS J. 2022, 289, 4240–4250. [Google Scholar] [CrossRef] [PubMed]
- Lipsitch, M.; Krammer, F.; Regev-Yochay, G.; Lustig, Y.; Balicer, R.D. SARS-CoV-2 Breakthrough Infections in Vaccinated Individuals: Measurement, Causes and Impact. Nat. Rev. Immunol. 2022, 22, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Bergwerk, M.; Gonen, T.; Lustig, Y.; Amit, S.; Lipsitch, M.; Cohen, C.; Mandelboim, M.; Levin, E.G.; Rubin, C.; Indenbaum, V.; et al. COVID-19 Breakthrough Infections in Vaccinated Health Care Workers. N. Engl. J. Med. 2021, 385, 1474–1484. [Google Scholar] [CrossRef] [PubMed]
- Davis, H.E.; Assaf, G.S.; McCorkell, L.; Wei, H.; Low, R.J.; Re’em, Y.; Redfield, S.; Austin, J.P.; Akrami, A. Characterizing Long COVID in an International Cohort: 7 Months of Symptoms and Their Impact. EClinicalMedicine 2021, 38, 101019. [Google Scholar] [CrossRef]
- Hayes, L.D.; Ingram, J.; Sculthorpe, N.F. More Than 100 Persistent Symptoms of SARS-CoV-2 (Long COVID): A Scoping Review. Front. Med. 2021, 8, 750378. [Google Scholar] [CrossRef]
- Vehar, S.; Boushra, M.; Ntiamoah, P.; Biehl, M. Post-Acute Sequelae of SARS-CoV-2 Infection: Caring for the “Long-Haulers.”. Cleve. Clin. J. Med. 2021, 88, 267–272. [Google Scholar] [CrossRef]
- Mehandru, S.; Merad, M. Pathological Sequelae of Long-Haul COVID. Nat. Immunol. 2022, 23, 194–202. [Google Scholar] [CrossRef]
- Collns, F.S. NIH Launches New Initiative to Study “Long COVID.” National Institutes of Health (NIH). Available online: https://www.nih.gov/about-nih/who-we-are/nih-director/statements/nih-launches-new-initiative-study-long-covid (accessed on 4 March 2022).
- LongCovidSOS. The Impact of COVID Vaccination on Symptoms of Long Covid. An International Survey of 900 People with Lived Experience (May 2021). Patient Safety Learning—The Hub. Available online: https://www.pslhub.org/learn/coronavirus-covid19/data-and-statistics/the-impact-of-covid-vaccination-on-symptoms-of-long-covid-an-international-survey-of-900-people-with-lived-experience-may-2021-r4636/ (accessed on 16 March 2022).
- Blankson, J.N.; Persaud, D.; Siliciano, R.F. The Challenge of Viral Reservoirs in HIV-1 Infection. Annu. Rev. Med. 2002, 53, 557–593. [Google Scholar] [CrossRef]
- Wu, Y.; Guo, C.; Tang, L.; Hong, Z.; Zhou, J.; Dong, X.; Yin, H.; Xiao, Q.; Tang, Y.; Qu, X.; et al. Prolonged Presence of SARS-CoV-2 Viral RNA in Faecal Samples. Lancet Gastroenterol. Hepatol. 2020, 5, 434–435. [Google Scholar] [CrossRef]
- Mendes Correa, M.C.; Leal, F.E.; Villas Boas, L.S.; Witkin, S.S.; de Paula, A.; Tozetto Mendonza, T.R.; Ferreira, N.E.; Curty, G.; de Carvalho, P.S.; Buss, L.F.; et al. Prolonged Presence of Replication-Competent SARS-CoV-2 in Mildly Symptomatic Individuals: A Report of Two Cases. J. Med. Virol. 2021, 93, 5603–5607. [Google Scholar] [CrossRef] [PubMed]
- Hong, K.; Cao, W.; Liu, Z.; Lin, L.; Zhou, X.; Zeng, Y.; Wei, Y.; Chen, L.; Liu, X.; Han, Y.; et al. Prolonged Presence of Viral Nucleic Acid in Clinically Recovered COVID-19 Patients Was Not Associated with Effective Infectiousness. Emerg. Microbes Infect. 2020, 9, 2315–2321. [Google Scholar] [CrossRef] [PubMed]
- Gaebler, C.; Wang, Z.; Lorenzi, J.C.C.; Muecksch, F.; Finkin, S.; Tokuyama, M.; Cho, A.; Jankovic, M.; Schaefer-Babajew, D.; Oliveira, T.Y.; et al. Evolution of Antibody Immunity to SARS-CoV-2. Nature 2021, 591, 639–644. [Google Scholar] [CrossRef] [PubMed]
- Proal, A.D.; VanElzakker, M.B. Long COVID or Post-Acute Sequelae of COVID-19 (PASC): An Overview of Biological Factors That May Contribute to Persistent Symptoms. Front. Microbiol. 2021, 12, 698169. [Google Scholar] [CrossRef]
- Kalkeri, R.; Goebel, S.; Sharma, G.D. SARS-CoV-2 Shedding from Asymptomatic Patients: Contribution of Potential Extrapulmonary Tissue Reservoirs. Am. J. Trop. Med. Hyg. 2020, 103, 18–21. [Google Scholar] [CrossRef]
- Viszlayová, D.; Sojka, M.; Dobrodenková, S.; Szabó, S.; Bilec, O.; Turzová, M.; Ďurina, J.; Baloghová, B.; Borbély, Z.; Kršák, M. SARS-CoV-2 RNA in the Cerebrospinal Fluid of a Patient with Long COVID. Ther Adv. Infect. Dis. 2021, 8, 20499361211048572. [Google Scholar] [CrossRef]
- Eriksen, A.Z.; Møller, R.; Makovoz, B.; Uhl, S.A.; tenOever, B.R.; Blenkinsop, T.A. SARS-CoV-2 Infects Human Adult Donor Eyes and hESC-Derived Ocular Epithelium. Cell Stem Cell 2021, 28, 1205–1220.e7. [Google Scholar] [CrossRef]
- Zhou, L.; Xu, Z.; Castiglione, G.M.; Soiberman, U.S.; Eberhart, C.G.; Duh, E.J. ACE2 and TMPRSS2 Are Expressed on the Human Ocular Surface, Suggesting Susceptibility to SARS-CoV-2 Infection. Ocul. Surf. 2020, 18, 537–544. [Google Scholar] [CrossRef]
- Colavita, F.; Curiale, S.; Lapa, D.; Castilletti, C. Live and Replication-Competent SARS-CoV-2 in Ocular Fluids. JAMA Ophthalmol. 2021, 139, 1041. [Google Scholar] [CrossRef]
- Colavita, F.; Lapa, D.; Carletti, F.; Lalle, E.; Bordi, L.; Marsella, P.; Nicastri, E.; Bevilacqua, N.; Giancola, M.L.; Corpolongo, A.; et al. SARS-CoV-2 Isolation From Ocular Secretions of a Patient With COVID-19 in Italy With Prolonged Viral RNA Detection. Ann. Intern. Med. 2020, 173, 242–243. [Google Scholar] [CrossRef]
- Menuchin-Lasowski, Y.; Schreiber, A.; Lecanda, A.; Mecate-Zambrano, A.; Brunotte, L.; Psathaki, O.E.; Ludwig, S.; Rauen, T.; Schöler, H.R. SARS-CoV-2 Infects and Replicates in Photoreceptor and Retinal Ganglion Cells of Human Retinal Organoids. Stem Cell Rep. 2022, 17, 789–803. [Google Scholar] [CrossRef] [PubMed]
- de Melo, G.D.; Lazarini, F.; Levallois, S.; Hautefort, C.; Michel, V.; Larrous, F.; Verillaud, B.; Aparicio, C.; Wagner, S.; Gheusi, G.; et al. COVID-19-Related Anosmia Is Associated with Viral Persistence and Inflammation in Human Olfactory Epithelium and Brain Infection in Hamsters. Sci. Transl. Med. 2021, 13, eabf8396. [Google Scholar] [CrossRef] [PubMed]
- Hu, F.; Chen, F.; Ou, Z.; Fan, Q.; Tan, X.; Wang, Y.; Pan, Y.; Ke, B.; Li, L.; Guan, Y.; et al. A Compromised Specific Humoral Immune Response against the SARS-CoV-2 Receptor-Binding Domain Is Related to Viral Persistence and Periodic Shedding in the Gastrointestinal Tract. Cell. Mol. Immunol. 2020, 17, 1119–1125. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, A.; Zlitni, S.; Brooks, E.F.; Vance, S.E.; Dahlen, A.; Hedlin, H.; Park, R.M.; Han, A.; Schmidtke, D.T.; Verma, R.; et al. Gastrointestinal Symptoms and Fecal Shedding of SARS-CoV-2 RNA Suggest Prolonged Gastrointestinal Infection. Med 2022, 3, 371–387.e9. [Google Scholar] [CrossRef] [PubMed]
- Elgarhy, L.H.; Salem, M.L. Could Injured Skin Be a Reservoir for SARS-CoV-2 Virus Spread? Clin. Dermatol. 2020, 38, 762–763. [Google Scholar] [CrossRef]
- Ryan, P.M.; Caplice, N.M. Is Adipose Tissue a Reservoir for Viral Spread, Immune Activation, and Cytokine Amplification in Coronavirus Disease 2019? Obesity 2020, 28, 1191–1194. [Google Scholar] [CrossRef]
- Vibholm, L.K.; Nielsen, S.S.F.; Pahus, M.H.; Frattari, G.S.; Olesen, R.; Andersen, R.; Monrad, I.; Andersen, A.H.F.; Thomsen, M.M.; Konrad, C.V.; et al. SARS-CoV-2 Persistence Is Associated with Antigen-Specific CD8 T-Cell Responses. EBioMedicine 2021, 64, 103230. [Google Scholar] [CrossRef]
- Herrera, D.; Serrano, J.; Roldán, S.; Sanz, M. Is the Oral Cavity Relevant in SARS-CoV-2 Pandemic? Clin. Oral Investig. 2020, 24, 2925–2930. [Google Scholar] [CrossRef]
- Xu, J.; Li, Y.; Gan, F.; Du, Y.; Yao, Y. Salivary Glands: Potential Reservoirs for COVID-19 Asymptomatic Infection. J. Dent. Res. 2020, 99, 989. [Google Scholar] [CrossRef]
- Troeltzsch, M.; Berndt, R.; Troeltzsch, M. Is the Oral Cavity a Reservoir for Prolonged SARS-CoV-2 Shedding? Med. Hypotheses 2021, 146, 110419. [Google Scholar] [CrossRef]
- Badran, Z.; Gaudin, A.; Struillou, X.; Amador, G.; Soueidan, A. Periodontal Pockets: A Potential Reservoir for SARS-CoV-2? Med. Hypotheses 2020, 143, 109907. [Google Scholar] [CrossRef] [PubMed]
- Gupta, M.; Mahanty, S.; Greer, P.; Towner, J.S.; Shieh, W.-J.; Zaki, S.R.; Ahmed, R.; Rollin, P.E. Persistent Infection with Ebola Virus under Conditions of Partial Immunity. J. Virol. 2004, 78, 958–967. [Google Scholar] [CrossRef] [PubMed]
- Viola, M.V.; Scott, C.; Duffy, P.D. Persistent Measles Virus Infection in Vitro and in Man. Arthritis Rheum. 1978, 21 (Suppl. 5), S47–S51. [Google Scholar] [CrossRef] [PubMed]
- Riddell, M.A.; Moss, W.J.; Hauer, D.; Monze, M.; Griffin, D.E. Slow Clearance of Measles Virus RNA after Acute Infection. J. Clin. Virol. 2007, 39, 312–317. [Google Scholar] [CrossRef] [PubMed]
- Ireland, D.D.C.; Manangeeswaran, M.; Lewkowicz, A.P.; Engel, K.; Clark, S.M.; Laniyan, A.; Sykes, J.; Lee, H.-N.; McWilliams, I.L.; Kelley-Baker, L.; et al. Long-Term Persistence of Infectious Zika Virus: Inflammation and Behavioral Sequela in Mice. PLoS Pathog. 2020, 16, e1008689. [Google Scholar] [CrossRef] [PubMed]
- Desimmie, B.A.; Raru, Y.Y.; Awadh, H.M.; He, P.; Teka, S.; Willenburg, K.S. Insights into SARS-CoV-2 Persistence and Its Relevance. Viruses 2021, 13, 1025. [Google Scholar] [CrossRef]
- Caniego-Casas, T.; Martínez-García, L.; Alonso-Riaño, M.; Pizarro, D.; Carretero-Barrio, I.; Martínez-de-Castro, N.; Ruz-Caracuel, I.; de Pablo, R.; Saiz, A.; Royo, R.N.; et al. RNA SARS-CoV-2 Persistence in the Lung of Severe COVID-19 Patients: A Case Series of Autopsies. Front. Microbiol. 2022, 13, 824967. [Google Scholar] [CrossRef] [PubMed]
- Randall, R.E.; Griffin, D.E. Within Host RNA Virus Persistence: Mechanisms and Consequences. Curr. Opin. Virol. 2017, 23, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Regnery, R.L.; Johnson, K.M.; Kiley, M.P. Virion Nucleic Acid of Ebola Virus. J. Virol. 1980, 36, 465–469. [Google Scholar] [CrossRef]
- Thorson, A.E.; Deen, G.F.; Bernstein, K.T.; Liu, W.J.; Yamba, F.; Habib, N.; Sesay, F.R.; Gaillard, P.; Massaquoi, T.A.; McDonald, S.L.R.; et al. Persistence of Ebola Virus in Semen among Ebola Virus Disease Survivors in Sierra Leone: A Cohort Study of Frequency, Duration, and Risk Factors. PLoS Med. 2021, 18, e1003273. [Google Scholar] [CrossRef]
- Keita, A.K.; Koundouno, F.R.; Faye, M.; Düx, A.; Hinzmann, J.; Diallo, H.; Ayouba, A.; Le Marcis, F.; Soropogui, B.; Ifono, K.; et al. Resurgence of Ebola Virus in 2021 in Guinea Suggests a New Paradigm for Outbreaks. Nature 2021, 597, 539–543. [Google Scholar] [CrossRef] [PubMed]
- Oliveira Souto, I.; Alejo-Cancho, I.; Gascón Brustenga, J.; Peiró Mestres, A.; Muñoz Gutiérrez, J.; Martínez Yoldi, M.J. Persistence of Zika Virus in Semen 93 Days after the Onset of Symptoms. Enferm. Infecc. Microbiol. Clin. 2018, 36, 21–23. [Google Scholar] [CrossRef] [PubMed]
- Bhatnagar, J.; Rabeneck, D.B.; Martines, R.B.; Reagan-Steiner, S.; Ermias, Y.; Estetter, L.B.C.; Suzuki, T.; Ritter, J.; Keating, M.K.; Hale, G.; et al. Zika Virus RNA Replication and Persistence in Brain and Placental Tissue. Emerg. Infect. Dis. 2017, 23, 405–414. [Google Scholar] [CrossRef]
- de Noronha, L.; Zanluca, C.; Burger, M.; Suzukawa, A.A.; Azevedo, M.; Rebutini, P.Z.; Novadzki, I.M.; Tanabe, L.S.; Presibella, M.M.; Duarte Dos Santos, C.N. Zika Virus Infection at Different Pregnancy Stages: Anatomopathological Findings, Target Cells and Viral Persistence in Placental Tissues. Front. Microbiol. 2018, 9, 2266. [Google Scholar] [CrossRef]
- Adams Waldorf, K.M.; Nelson, B.R.; Stencel-Baerenwald, J.E.; Studholme, C.; Kapur, R.P.; Armistead, B.; Walker, C.L.; Merillat, S.; Vornhagen, J.; Tisoncik-Go, J.; et al. Congenital Zika Virus Infection as a Silent Pathology with Loss of Neurogenic Output in the Fetal Brain. Nat. Med. 2018, 24, 368–374. [Google Scholar] [CrossRef] [PubMed]
- White, M.K.; Wollebo, H.S.; David Beckham, J.; Tyler, K.L.; Khalili, K. Zika Virus: An Emergent Neuropathological Agent. Ann. Neurol. 2016, 80, 479–489. [Google Scholar] [CrossRef]
- Kristensson, K.; Norrby, E. Persistence of RNA Viruses in the Central Nervous System. Annu. Rev. Microbiol. 1986, 40, 159–184. [Google Scholar] [CrossRef]
- Guo, L.; Wang, G.; Wang, Y.; Zhang, Q.; Ren, L.; Gu, X.; Huang, T.; Zhong, J.; Wang, Y.; Wang, X.; et al. SARS-CoV-2-Specific Antibody and T-Cell Responses 1 Year after Infection in People Recovered from COVID-19: A Longitudinal Cohort Study. Lancet Microbe 2022, 3, e348–e356. [Google Scholar] [CrossRef]
- Rank, A.; Tzortzini, A.; Kling, E.; Schmid, C.; Claus, R.; Löll, E.; Burger, R.; Römmele, C.; Dhillon, C.; Müller, K.; et al. One Year after Mild COVID-19: The Majority of Patients Maintain Specific Immunity, But One in Four Still Suffer from Long-Term Symptoms. J. Clin. Med. Res. 2021, 10, 3305. [Google Scholar] [CrossRef]
- Thorne, L.G.; Bouhaddou, M.; Reuschl, A.-K.; Zuliani-Alvarez, L.; Polacco, B.; Pelin, A.; Batra, J.; Whelan, M.V.X.; Hosmillo, M.; Fossati, A.; et al. Evolution of Enhanced Innate Immune Evasion by SARS-CoV-2. Nature 2021, 602, 487–495. [Google Scholar] [CrossRef]
- Clark, S.A.; Clark, L.E.; Pan, J.; Coscia, A.; McKay, L.G.A.; Shankar, S.; Johnson, R.I.; Brusic, V.; Choudhary, M.C.; Regan, J.; et al. SARS-CoV-2 Evolution in an Immunocompromised Host Reveals Shared Neutralization Escape Mechanisms. Cell 2021, 184, 2605–2617.e18. [Google Scholar] [CrossRef] [PubMed]
- Min, Y.-Q.; Huang, M.; Sun, X.; Deng, F.; Wang, H.; Ning, Y.-J. Immune Evasion of SARS-CoV-2 from Interferon Antiviral System. Comput. Struct. Biotechnol. J. 2021, 19, 4217–4225. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.K.; Blanco, M.R.; Bruce, E.A.; Honson, D.D.; Chen, L.M.; Chow, A.; Bhat, P.; Ollikainen, N.; Quinodoz, S.A.; Loney, C.; et al. SARS-CoV-2 Disrupts Splicing, Translation, and Protein Trafficking to Suppress Host Defenses. Cell 2020, 183, 1325–1339.e21. [Google Scholar] [CrossRef] [PubMed]
- Kamitani, W.; Narayanan, K.; Huang, C.; Lokugamage, K.; Ikegami, T.; Ito, N.; Kubo, H.; Makino, S. Severe Acute Respiratory Syndrome Coronavirus nsp1 Protein Suppresses Host Gene Expression by Promoting Host mRNA Degradation. Proc. Natl. Acad. Sci. USA 2006, 103, 12885–12890. [Google Scholar] [CrossRef] [PubMed]
- Hyman, A.A.; Weber, C.A.; Jülicher, F. Liquid-Liquid Phase Separation in Biology. Annu. Rev. Cell Dev. Biol. 2014, 30, 39–58. [Google Scholar] [CrossRef]
- Gomes, E.; Shorter, J. The Molecular Language of Membraneless Organelles. J. Biol. Chem. 2019, 294, 7115–7127. [Google Scholar] [CrossRef]
- Feng, Z.; Chen, X.; Wu, X.; Zhang, M. Formation of Biological Condensates via Phase Separation: Characteristics, Analytical Methods, and Physiological Implications. J. Biol. Chem. 2019, 294, 14823–14835. [Google Scholar] [CrossRef]
- Ning, W.; Guo, Y.; Lin, S.; Mei, B.; Wu, Y.; Jiang, P.; Tan, X.; Zhang, W.; Chen, G.; Peng, D.; et al. DrLLPS: A Data Resource of Liquid–liquid Phase Separation in Eukaryotes. Nucleic Acids Res. 2019, 48, D288–D295. [Google Scholar] [CrossRef]
- Azaldegui, C.A.; Vecchiarelli, A.G.; Biteen, J.S. The Emergence of Phase Separation as an Organizing Principle in Bacteria. Biophys. J. 2021, 120, 1123–1138. [Google Scholar] [CrossRef]
- Salvador-Castell, M.; Demé, B.; Oger, P.; Peters, J. Lipid Phase Separation Induced by the Apolar Polyisoprenoid Squalane Demonstrates Its Role in Membrane Domain Formation in Archaeal Membranes. Langmuir 2020, 36, 7375–7382. [Google Scholar] [CrossRef]
- Hansma, H.G. Better than Membranes at the Origin of Life? Life 2017, 7, 28. [Google Scholar] [CrossRef] [PubMed]
- Banani, S.F.; Lee, H.O.; Hyman, A.A.; Rosen, M.K. Biomolecular Condensates: Organizers of Cellular Biochemistry. Nat. Rev. Mol. Cell Biol. 2017, 18, 285–298. [Google Scholar] [CrossRef]
- White, J.P.; Lloyd, R.E. Regulation of Stress Granules in Virus Systems. Trends Microbiol. 2012, 20, 175–183. [Google Scholar] [CrossRef] [PubMed]
- McCormick, C.; Khaperskyy, D.A. Translation Inhibition and Stress Granules in the Antiviral Immune Response. Nat. Rev. Immunol. 2017, 17, 647–660. [Google Scholar] [CrossRef] [PubMed]
- Buchan, J.R.; Parker, R. Eukaryotic Stress Granules: The Ins and Outs of Translation. Mol. Cell 2009, 36, 932–941. [Google Scholar] [CrossRef]
- Kedersha, N.; Ivanov, P.; Anderson, P. Stress Granules and Cell Signaling: More than Just a Passing Phase? Trends Biochem. Sci. 2013, 38, 494–506. [Google Scholar] [CrossRef]
- Riback, J.A.; Katanski, C.D.; Kear-Scott, J.L.; Pilipenko, E.V.; Rojek, A.E.; Sosnick, T.R.; Drummond, D.A. Stress-Triggered Phase Separation Is an Adaptive, Evolutionarily Tuned Response. Cell 2017, 168, 1028–1040.e19. [Google Scholar] [CrossRef]
- Mahboubi, H.; Stochaj, U. Cytoplasmic Stress Granules: Dynamic Modulators of Cell Signaling and Disease. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 884–895. [Google Scholar] [CrossRef]
- Baumann, K. mRNA Translation in Stress Granules Is Not Uncommon. Nat. Rev. Mol. Cell Biol. 2021, 22, 164. [Google Scholar] [CrossRef]
- Moon, S.L.; Morisaki, T.; Stasevich, T.J.; Parker, R. Coupling of Translation Quality Control and mRNA Targeting to Stress Granules. J. Cell Biol. 2020, 219, e202004120. [Google Scholar] [CrossRef]
- Xiao, Q.; McAtee, C.K.; Su, X. Phase Separation in Immune Signalling. Nat. Rev. Immunol. 2021, 22, 188–199. [Google Scholar] [CrossRef] [PubMed]
- Pakos-Zebrucka, K.; Koryga, I.; Mnich, K.; Ljujic, M.; Samali, A.; Gorman, A.M. The Integrated Stress Response. EMBO Rep. 2016, 17, 1374–1395. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhang, Z.; Li, Y.; Li, Y. The Regulation of Integrated Stress Response Signaling Pathway on Viral Infection and Viral Antagonism. Front. Microbiol. 2021, 12, 814635. [Google Scholar] [CrossRef]
- Gil, J.; Esteban, M. The Interferon-Induced Protein Kinase (PKR), Triggers Apoptosis through FADD-Mediated Activation of Caspase 8 in a Manner Independent of Fas and TNF-Alpha Receptors. Oncogene 2000, 19, 3665–3674. [Google Scholar] [CrossRef]
- Lee, S.B.; Rodríguez, D.; Rodríguez, J.R.; Esteban, M. The Apoptosis Pathway Triggered by the Interferon-Induced Protein Kinase PKR Requires the Third Basic Domain, Initiates Upstream of Bcl-2, and Involves ICE-like Proteases. Virology 1997, 231, 81–88. [Google Scholar] [CrossRef] [PubMed]
- Yoneyama, M.; Jogi, M.; Onomoto, K. Regulation of Antiviral Innate Immune Signaling by Stress-Induced RNA Granules. J. Biochem. 2016, 159, 279–286. [Google Scholar] [CrossRef]
- Miller, C.L. Stress Granules and Virus Replication. Future Virol. 2011, 6, 1329–1338. [Google Scholar] [CrossRef]
- Zheng, Z.-Q.; Wang, S.-Y.; Xu, Z.-S.; Fu, Y.-Z.; Wang, Y.-Y. SARS-CoV-2 Nucleocapsid Protein Impairs Stress Granule Formation to Promote Viral Replication. Cell Discov. 2021, 7, 38. [Google Scholar] [CrossRef]
- Ahlquist, P. Parallels among Positive-Strand RNA Viruses, Reverse-Transcribing Viruses and Double-Stranded RNA Viruses. Nat. Rev. Microbiol. 2006, 4, 371–382. [Google Scholar] [CrossRef]
- Guo, Y.; Hinchman, M.M.; Lewandrowski, M.; Cross, S.T.; Sutherland, D.M.; Welsh, O.L.; Dermody, T.S.; Parker, J.S.L. The Multi-Functional Reovirus σ3 Protein Is a Virulence Factor That Suppresses Stress Granule Formation and Is Associated with Myocardial Injury. PLoS Pathog. 2021, 17, e1009494. [Google Scholar] [CrossRef]
- van Leeuwen, W.; Rabouille, C. Cellular Stress Leads to the Formation of Membraneless Stress Assemblies in Eukaryotic Cells. Traffic 2019, 20, 623–638. [Google Scholar] [CrossRef] [PubMed]
- Anderson, P.; Kedersha, N. RNA Granules. J. Cell Biol. 2006, 172, 803–808. [Google Scholar] [CrossRef] [PubMed]
- Gaete-Argel, A.; Velásquez, F.; Márquez, C.L.; Rojas-Araya, B.; Bueno-Nieto, C.; Marín-Rojas, J.; Cuevas-Zúñiga, M.; Soto-Rifo, R.; Valiente-Echeverría, F. Tellurite Promotes Stress Granules and Nuclear SG-Like Assembly in Response to Oxidative Stress and DNA Damage. Front. Cell Dev. Biol. 2021, 9, 622057. [Google Scholar] [CrossRef] [PubMed]
- Emara, M.M.; Fujimura, K.; Sciaranghella, D.; Ivanova, V.; Ivanov, P.; Anderson, P. Hydrogen Peroxide Induces Stress Granule Formation Independent of eIF2α Phosphorylation. Biochem. Biophys. Res. Commun. 2012, 423, 763–769. [Google Scholar] [CrossRef]
- Lian, X.J.; Gallouzi, I.-E. Oxidative Stress Increases the Number of Stress Granules in Senescent Cells and Triggers a Rapid Decrease in p21waf1/cip1 Translation. J. Biol. Chem. 2009, 284, 8877–8887. [Google Scholar] [CrossRef]
- Sathyanarayanan, U.; Musa, M.; Bou Dib, P.; Raimundo, N.; Milosevic, I.; Krisko, A. ATP Hydrolysis by Yeast Hsp104 Determines Protein Aggregate Dissolution and Size in Vivo. Nat. Commun. 2020, 11, 5226. [Google Scholar] [CrossRef]
- Reineke, L.C.; Cheema, S.A.; Dubrulle, J.; Neilson, J.R. Chronic Starvation Induces Noncanonical pro-Death Stress Granules. J. Cell Sci. 2018, 131, jcs220244. [Google Scholar] [CrossRef]
- Ying, S.; Khaperskyy, D.A. UV Damage Induces G3BP1-Dependent Stress Granule Formation That Is Not Driven by mTOR Inhibition-Mediated Translation Arrest. J. Cell Sci. 2020, 133, jcs248310. [Google Scholar] [CrossRef]
- Moutaoufik, M.T.; El Fatimy, R.; Nassour, H.; Gareau, C.; Lang, J.; Tanguay, R.M.; Mazroui, R.; Khandjian, E.W. UVC-Induced Stress Granules in Mammalian Cells. PLoS ONE 2014, 9, e112742. [Google Scholar] [CrossRef]
- Timalsina, S.; Arimoto-Matsuzaki, K.; Kitamura, M.; Xu, X.; Wenzhe, Q.; Ishigami-Yuasa, M.; Kagechika, H.; Hata, Y. Chemical Compounds That Suppress Hypoxia-Induced Stress Granule Formation Enhance Cancer Drug Sensitivity of Human Cervical Cancer HeLa Cells. J. Biochem. 2018, 164, 381–391. [Google Scholar] [CrossRef]
- van der Laan, A.M.A.; van Gemert, A.M.C.; Dirks, R.W.; Noordermeer, J.N.; Fradkin, L.G.; Tanke, H.J.; Jost, C.R. mRNA Cycles through Hypoxia-Induced Stress Granules in Live Drosophila Embryonic Muscles. Int. J. Dev. Biol. 2012, 56, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Namkoong, S.; Ho, A.; Woo, Y.M.; Kwak, H.; Lee, J.H. Systematic Characterization of Stress-Induced RNA Granulation. Mol. Cell 2018, 70, 175–187. [Google Scholar] [CrossRef] [PubMed]
- Tweedie, A.; Nissan, T. Hiding in Plain Sight: Formation and Function of Stress Granules During Microbial Infection of Mammalian Cells. Front. Mol. Biosci 2021, 8, 647884. [Google Scholar] [CrossRef] [PubMed]
- Child, J.R.; Chen, Q.; Reid, D.W.; Jagannathan, S.; Nicchitta, C.V. Recruitment of Endoplasmic Reticulum-Targeted and Cytosolic mRNAs into Membrane-Associated Stress Granules. RNA 2021, 27, 1241–1256. [Google Scholar] [CrossRef]
- Zhang, X.; Sridharan, S.; Zagoriy, I.; Oegema, C.E.; Ching, C.; Pflaesterer, T.; Fung, H.K.H.; Poser, I.; Mueller, C.W.; Hyman, A.A.; et al. Molecular Mechanisms of Stress-Induced Reactivation in Mumps Virus Condensates. bioRxiv 2022. [Google Scholar] [CrossRef]
- Emara, M.M.; Brinton, M.A. Interaction of TIA-1/TIAR with West Nile and Dengue Virus Products in Infected Cells Interferes with Stress Granule Formation and Processing Body Assembly. Proc. Natl. Acad. Sci. USA 2007, 104, 9041–9046. [Google Scholar] [CrossRef]
- Katoh, H.; Okamoto, T.; Fukuhara, T.; Kambara, H.; Morita, E.; Mori, Y.; Kamitani, W.; Matsuura, Y. Japanese Encephalitis Virus Core Protein Inhibits Stress Granule Formation through an Interaction with Caprin-1 and Facilitates Viral Propagation. J. Virol. 2013, 87, 489–502. [Google Scholar] [CrossRef]
- John, L.; Samuel, C.E. Induction of Stress Granules by Interferon and down-Regulation by the Cellular RNA Adenosine Deaminase ADAR1. Virology 2014, 454–455, 299–310. [Google Scholar] [CrossRef]
- Courtney, S.C.; Scherbik, S.V.; Stockman, B.M.; Brinton, M.A. West Nile Virus Infections Suppress Early Viral RNA Synthesis and Avoid Inducing the Cell Stress Granule Response. J. Virol. 2012, 86, 3647–3657. [Google Scholar] [CrossRef]
- Blázquez, A.-B.; Martín-Acebes, M.A.; Poderoso, T.; Saiz, J.-C. Relevance of Oxidative Stress in Inhibition of eIF2 Alpha Phosphorylation and Stress Granules Formation during Usutu Virus Infection. PLoS Negl. Trop. Dis. 2021, 15, e0009072. [Google Scholar] [CrossRef]
- Hou, S.; Kumar, A.; Xu, Z.; Airo, A.M.; Stryapunina, I.; Wong, C.P.; Branton, W.; Tchesnokov, E.; Götte, M.; Power, C.; et al. Zika Virus Hijacks Stress Granule Proteins and Modulates the Host Stress Response. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Sen, G.C. Viruses and Interferons. Annu. Rev. Microbiol. 2001, 55, 255–281. [Google Scholar] [CrossRef] [PubMed]
- Schultz, U.; Kaspers, B.; Staeheli, P. The Interferon System of Non-Mammalian Vertebrates. Dev. Comp. Immunol. 2004, 28, 499–508. [Google Scholar] [CrossRef] [PubMed]
- Randall, R.E.; Goodbourn, S. Interferons and Viruses: An Interplay between Induction, Signalling, Antiviral Responses and Virus Countermeasures. J. Gen. Virol. 2008, 89 Pt 1, 1–47. [Google Scholar] [CrossRef] [PubMed]
- Stark, G.R.; Darnell, J.E., Jr. The JAK-STAT Pathway at Twenty. Immunity 2012, 36, 503–514. [Google Scholar] [CrossRef]
- Liu, S.-Y.; Sanchez, D.J.; Aliyari, R.; Lu, S.; Cheng, G. Systematic Identification of Type I and Type II Interferon-Induced Antiviral Factors. Proc. Natl. Acad. Sci. USA 2012, 109, 4239–4244. [Google Scholar] [CrossRef]
- Katze, M.G.; He, Y.; Gale, M., Jr. Viruses and Interferon: A Fight for Supremacy. Nat. Rev. Immunol. 2002, 2, 675–687. [Google Scholar] [CrossRef]
- Lin, R.-J.; Chang, B.-L.; Yu, H.-P.; Liao, C.-L.; Lin, Y.-L. Blocking of Interferon-Induced Jak-Stat Signaling by Japanese Encephalitis Virus NS5 through a Protein Tyrosine Phosphatase-Mediated Mechanism. J. Virol. 2006, 80, 5908–5918. [Google Scholar] [CrossRef]
- Vazquez, C.; Swanson, S.E.; Negatu, S.G.; Dittmar, M.; Miller, J.; Ramage, H.R.; Cherry, S.; Jurado, K.A. SARS-CoV-2 Viral Proteins NSP1 and NSP13 Inhibit Interferon Activation through Distinct Mechanisms. PLoS ONE 2021, 16, e0253089. [Google Scholar] [CrossRef]
- Oh, S.J.; Shin, O.S. SARS-CoV-2 Nucleocapsid Protein Targets RIG-I-Like Receptor Pathways to Inhibit the Induction of Interferon Response. Cells 2021, 10, 530. [Google Scholar] [CrossRef]
- Guo, K.; Barrett, B.S.; Mickens, K.L.; Vladar, E.K.; Morrison, J.H.; Hasenkrug, K.J.; Poeschla, E.M.; Santiago, M.L. Interferon Resistance of Emerging SARS-CoV-2 Variants. bioRxiv 2021. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Zhou, Z.; Xiao, X.; Tian, Z.; Dong, X.; Wang, C.; Li, L.; Ren, L.; Lei, X.; Xiang, Z.; et al. SARS-CoV-2 nsp12 Attenuates Type I Interferon Production by Inhibiting IRF3 Nuclear Translocation. Cell Mol. Immunol. 2021, 18, 945–953. [Google Scholar] [CrossRef] [PubMed]
- Park, A.; Iwasaki, A. Type I and Type III Interferons—Induction, Signaling, Evasion, and Application to Combat COVID-19. Cell Host Microbe 2020, 27, 870–878. [Google Scholar] [CrossRef] [PubMed]
- Hadjadj, J.; Yatim, N.; Barnabei, L.; Corneau, A.; Boussier, J.; Smith, N.; Péré, H.; Charbit, B.; Bondet, V.; Chenevier-Gobeaux, C.; et al. Impaired Type I Interferon Activity and Inflammatory Responses in Severe COVID-19 Patients. Science 2020, 369, 718–724. [Google Scholar] [CrossRef]
- Matsuyama, T.; Kubli, S.P.; Yoshinaga, S.K.; Pfeffer, K.; Mak, T.W. An Aberrant STAT Pathway Is Central to COVID-19. Cell Death Differ. 2020, 27, 3209–3225. [Google Scholar] [CrossRef]
- Jafarzadeh, A.; Nemati, M.; Jafarzadeh, S. Contribution of STAT3 to the Pathogenesis of COVID-19. Microb. Pathog. 2021, 154, 104836. [Google Scholar] [CrossRef]
- Tan, L.Y.; Komarasamy, T.V.; Rmt Balasubramaniam, V. Hyperinflammatory Immune Response and COVID-19: A Double Edged Sword. Front. Immunol. 2021, 12, 742941. [Google Scholar] [CrossRef]
- Ravid, J.D.; Leiva, O.; Chitalia, V.C. Janus Kinase Signaling Pathway and Its Role in COVID-19 Inflammatory, Vascular, and Thrombotic Manifestations. Cells 2022, 11, 306. [Google Scholar] [CrossRef]
- Grant, A.H.; Estrada, A., 3rd; Ayala-Marin, Y.M.; Alvidrez-Camacho, A.Y.; Rodriguez, G.; Robles-Escajeda, E.; Cadena-Medina, D.A.; Rodriguez, A.C.; Kirken, R.A. The Many Faces of JAKs and STATs Within the COVID-19 Storm. Front. Immunol. 2021, 12, 690477. [Google Scholar] [CrossRef]
- Neubauer, A.; Johow, J.; Mack, E.; Burchert, A.; Meyn, D.; Kadlubiec, A.; Torje, I.; Wulf, H.; Vogelmeier, C.F.; Hoyer, J.; et al. The Janus-Kinase Inhibitor Ruxolitinib in SARS-CoV-2 Induced Acute Respiratory Distress Syndrome (ARDS). Leukemia 2021, 35, 2917–2923. [Google Scholar] [CrossRef]
- Yan, B.; Freiwald, T.; Chauss, D.; Wang, L.; West, E.; Mirabelli, C.; Zhang, C.J.; Nichols, E.-M.; Malik, N.; Gregory, R.; et al. SARS-CoV-2 Drives JAK1/2-Dependent Local Complement Hyperactivation. Sci. Immunol. 2021, 6, eabg0833. [Google Scholar] [CrossRef] [PubMed]
- Goletti, D.; Cantini, F. Baricitinib Therapy in Covid-19 Pneumonia—An Unmet Need Fulfilled. N. Engl. J. Med. 2021, 384, 867–869. [Google Scholar] [CrossRef] [PubMed]
- Guimarães, P.O.; Quirk, D.; Furtado, R.H.; Maia, L.N.; Saraiva, J.F.; Antunes, M.O.; Kalil Filho, R.; Junior, V.M.; Soeiro, A.M.; Tognon, A.P.; et al. Tofacitinib in Patients Hospitalized with Covid-19 Pneumonia. N. Engl. J. Med. 2021, 385, 406–415. [Google Scholar] [CrossRef] [PubMed]
- Satarker, S.; Tom, A.A.; Shaji, R.A.; Alosious, A.; Luvis, M.; Nampoothiri, M. JAK-STAT Pathway Inhibition and Their Implications in COVID-19 Therapy. Postgrad. Med. 2021, 133, 489–507. [Google Scholar] [CrossRef]
- Chen, D.-Y.; Khan, N.; Close, B.J.; Goel, R.K.; Blum, B.; Tavares, A.H.; Kenney, D.; Conway, H.L.; Ewoldt, J.K.; Chitalia, V.C.; et al. SARS-CoV-2 Disrupts Proximal Elements in the JAK-STAT Pathway. J. Virol. 2021, 95, e0086221. [Google Scholar] [CrossRef]
- Xia, H.; Cao, Z.; Xie, X.; Zhang, X.; Chen, J.Y.-C.; Wang, H.; Menachery, V.D.; Rajsbaum, R.; Shi, P.-Y. Evasion of Type I Interferon by SARS-CoV-2. Cell Rep. 2020, 33, 108234. [Google Scholar] [CrossRef]
- Yuen, C.-K.; Lam, J.-Y.; Wong, W.-M.; Mak, L.-F.; Wang, X.; Chu, H.; Cai, J.-P.; Jin, D.-Y.; To, K.K.-W.; Chan, J.F.-W.; et al. SARS-CoV-2 nsp13, nsp14, nsp15 and orf6 Function as Potent Interferon Antagonists. Emerg. Microbes Infect. 2020, 9, 1418–1428. [Google Scholar] [CrossRef]
- Rebendenne, A.; Valadão, A.L.C.; Tauziet, M.; Maarifi, G.; Bonaventure, B.; McKellar, J.; Planès, R.; Nisole, S.; Arnaud-Arnould, M.; Moncorgé, O.; et al. SARS-CoV-2 Triggers an MDA-5-Dependent Interferon Response Which Is Unable to Control Replication in Lung Epithelial Cells. J. Virol. 2021, 95, e02415-20. [Google Scholar] [CrossRef]
- Zandi, K.; Musall, K.; Oo, A.; Cao, D.; Liang, B.; Hassandarvish, P.; Lan, S.; Slack, R.L.; Kirby, K.A.; Bassit, L.; et al. Baicalein and Baicalin Inhibit SARS-CoV-2 RNA-Dependent-RNA Polymerase. Microorganisms 2021, 9, 893. [Google Scholar] [CrossRef]
- Savastano, A.; Ibáñez de Opakua, A.; Rankovic, M.; Zweckstetter, M. Nucleocapsid Protein of SARS-CoV-2 Phase Separates into RNA-Rich Polymerase-Containing Condensates. Nat. Commun. 2020, 11, 6041. [Google Scholar] [CrossRef]
- Tan, A.T.; Linster, M.; Tan, C.W.; Le Bert, N.; Chia, W.N.; Kunasegaran, K.; Zhuang, Y.; Tham, C.Y.L.; Chia, A.; Smith, G.J.D.; et al. Early Induction of Functional SARS-CoV-2-Specific T Cells Associates with Rapid Viral Clearance and Mild Disease in COVID-19 Patients. Cell Rep. 2021, 34, 108728. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, M.; Worlock, K.B.; Huang, N.; Lindeboom, R.G.H.; Butler, C.R.; Kumasaka, N.; Dominguez Conde, C.; Mamanova, L.; Bolt, L.; Richardson, L.; et al. Local and Systemic Responses to SARS-CoV-2 Infection in Children and Adults. Nature 2022, 602, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, C.A.; Clemmensen, A.; Sparrewath, C.; Tetens, M.M.; Krogfelt, K.A. Children Naturally Evading COVID-19—Why Children Differ from Adults. COVID 2022, 2, 369–378. [Google Scholar] [CrossRef]
- Speranza, E. Children Primed and Ready for SARS-CoV-2. Nat. Microbiol. 2021, 6, 1337–1338. [Google Scholar] [CrossRef]
- Loske, J.; Röhmel, J.; Lukassen, S.; Stricker, S.; Magalhães, V.G.; Liebig, J.; Chua, R.L.; Thürmann, L.; Messingschlager, M.; Seegebarth, A.; et al. Pre-Activated Antiviral Innate Immunity in the Upper Airways Controls Early SARS-CoV-2 Infection in Children. Nat. Biotechnol. 2021, 40, 319–324. [Google Scholar] [CrossRef]
- Kumar, A.; Ishida, R.; Strilets, T.; Cole, J.; Lopez-Orozco, J.; Fayad, N.; Felix-Lopez, A.; Elaish, M.; Evseev, D.; Magor, K.E.; et al. SARS-CoV-2 Nonstructural Protein 1 Inhibits the Interferon Response by Causing Depletion of Key Host Signaling Factors. J. Virol. 2021, 95, e0026621. [Google Scholar] [CrossRef] [PubMed]
- Waldhauser, F.; Weiszenbacher, G.; Tatzer, E.; Gisinger, B.; Waldhauser, M.; Schemper, M.; Frisch, H. Alterations in Nocturnal Serum Melatonin Levels in Humans with Growth and Aging. J. Clin. Endocrinol. Metab. 1988, 66, 648–652. [Google Scholar] [CrossRef] [PubMed]
- Bahrampour Juybari, K.; Pourhanifeh, M.H.; Hosseinzadeh, A.; Hemati, K.; Mehrzadi, S. Melatonin Potentials against Viral Infections Including COVID-19: Current Evidence and New Findings. Virus Res. 2020, 287, 198108. [Google Scholar] [CrossRef]
- Huang, S.-H.; Liao, C.-L.; Chen, S.-J.; Shi, L.-G.; Lin, L.; Chen, Y.-W.; Cheng, C.-P.; Sytwu, H.-K.; Shang, S.-T.; Lin, G.-J. Melatonin Possesses an Anti-Influenza Potential through Its Immune Modulatory Effect. J. Funct. Foods 2019, 58, 189–198. [Google Scholar] [CrossRef]
- Anderson, G.; Maes, M.; Markus, R.P.; Rodriguez, M. Ebola Virus: Melatonin as a Readily Available Treatment Option. J. Med. Virol. 2015, 87, 537–543. [Google Scholar] [CrossRef]
- Boga, J.A.; Coto-Montes, A.; Rosales-Corral, S.A.; Tan, D.-X.; Reiter, R.J. Beneficial Actions of Melatonin in the Management of Viral Infections: A New Use for This “Molecular Handyman”? Rev. Med. Virol. 2012, 22, 323–338. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, V.; Mohamed, M.; Kato, H. Melatonin in Bacterial and Viral Infections with Focus on Sepsis: A Review. Recent Pat. Endocr. Metab. Immune Drug Discov. 2012, 6, 30–39. [Google Scholar] [CrossRef] [PubMed]
- Ben-Nathan, D.; Maestroni, G.J.; Lustig, S.; Conti, A. Protective Effects of Melatonin in Mice Infected with Encephalitis Viruses. Arch. Virol. 1995, 140, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Wang, H.; Zhou, Q.; Li, Q.; Liu, N.; Li, Z.; Chen, C.; Deng, Y. Melatonin Ameliorates Axonal Hypomyelination of Periventricular White Matter by Transforming A1 to A2 Astrocyte via JAK2/STAT3 Pathway in Septic Neonatal Rats. J. Inflamm. Res. 2021, 14, 5919–5937. [Google Scholar] [CrossRef]
- Li, S.; Yang, S.; Sun, B.; Hang, C. Melatonin Attenuates Early Brain Injury after Subarachnoid Hemorrhage by the JAK-STAT Signaling Pathway. Int. J. Clin. Exp. Pathol. 2019, 12, 909–915. [Google Scholar]
- Raftery, N.; Stevenson, N.J. Advances in Anti-Viral Immune Defence: Revealing the Importance of the IFN JAK/STAT Pathway. Cell. Mol. Life Sci. 2017, 74, 2525–2535. [Google Scholar] [CrossRef]
- Lau, W.W.I.; Ng, J.K.Y.; Lee, M.M.K.; Chan, A.S.L.; Wong, Y.H. Interleukin-6 Autocrine Signaling Mediates Melatonin MT(1/2) Receptor-Induced STAT3 Tyr(705) Phosphorylation. J. Pineal Res. 2012, 52, 477–489. [Google Scholar] [CrossRef]
- Fitzgerald-Bocarsly, P. Human Natural Interferon-Alpha Producing Cells. Pharmacol. Ther. 1993, 60, 39–62. [Google Scholar] [CrossRef]
- Decker, P. Neutrophils and Interferon-α-Producing Cells: Who Produces Interferon in Lupus? Arthritis Res. Ther. 2011, 13, 118. [Google Scholar] [CrossRef]
- Peña, C.; Rincon, J.; Pedreanez, A.; Viera, N.; Mosquera, J. Chemotactic Effect of Melatonin on Leukocytes. J. Pineal Res. 2007, 43, 263–269. [Google Scholar] [CrossRef]
- Heinonen, S.; Rodriguez-Fernandez, R.; Diaz, A.; Oliva Rodriguez-Pastor, S.; Ramilo, O.; Mejias, A. Infant Immune Response to Respiratory Viral Infections. Immunol. Allergy Clin. N. Am. 2019, 39, 361–376. [Google Scholar] [CrossRef] [PubMed]
- Montiel, M.; Bonilla, E.; Valero, N.; Mosquera, J.; Espina, L.M.; Quiroz, Y.; Álvarez-Mon, M. Melatonin Decreases Brain Apoptosis, Oxidative Stress, and CD200 Expression and Increased Survival Rate in Mice Infected by Venezuelan Equine Encephalitis Virus. Antivir. Chem. Chemother. 2015, 24, 99–108. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Rico, M.; de la Muela, P.; Herrera-Morueco, J.J.; Geoffroy, P.A.; Limosin, F.; Hoertel, N.; AP-HP/Université de Paris/INSERM COVID-19 Research Collaboration/AP-HP COVID CDR Initiative/Entrepôt de Données de Santé AP-HP Consortium. Melatonin Does Not Reduce Mortality in Adult Hospitalized Patients with COVID-19: A Multicenter Retrospective Observational Study. J. Travel Med. 2022, 29, taab195. [Google Scholar] [CrossRef] [PubMed]
- Hasan, Z.T.; Atrakji, D.M.Q.Y.M.A.A.; Mehuaiden, D.A.K. The Effect of Melatonin on Thrombosis, Sepsis and Mortality Rate in COVID-19 Patients. Int. J. Infect. Dis. 2022, 114, 79–84. [Google Scholar] [CrossRef]
- Castillo, R.R.; Quizon, G.R.A.; Juco, M.J.M.; Roman, A.D.E.; de Leon, D.G.; Punzalan, F.E.R.; Guingon, R.B.L.; Morales, D.D.; Tan, D.-X.; Reiter, R.J. Melatonin as Adjuvant Treatment for Coronavirus Disease 2019 Pneumonia Patients Requiring Hospitalization (MAC-19 PRO): A Case Series. Melatonin Res. 2020, 3, 297–310. [Google Scholar] [CrossRef]
- Kamel, W.; Noerenberg, M.; Cerikan, B.; Chen, H.; Järvelin, A.I.; Kammoun, M.; Lee, J.Y.; Shuai, N.; Garcia-Moreno, M.; Andrejeva, A.; et al. Global Analysis of Protein-RNA Interactions in SARS-CoV-2-Infected Cells Reveals Key Regulators of Infection. Mol. Cell 2021, 81, 2851–2867.e7. [Google Scholar] [CrossRef]
- V’kovski, P.; Kratzel, A.; Steiner, S.; Stalder, H.; Thiel, V. Coronavirus Biology and Replication: Implications for SARS-CoV-2. Nat. Rev. Microbiol. 2021, 19, 155–170. [Google Scholar] [CrossRef]
- Khailany, R.A.; Safdar, M.; Ozaslan, M. Genomic Characterization of a Novel SARS-CoV-2. Gene Rep. 2020, 19, 100682. [Google Scholar] [CrossRef]
- Gerassimovich, Y.A.; Miladinovski-Bangall, S.J.; Bridges, K.M.; Boateng, L.; Ball, L.E.; Valafar, H.; Nag, A. Proximity-Dependent Biotinylation Detects Associations between SARS Coronavirus Nonstructural Protein 1 and Stress Granule-Associated Proteins. J. Biol. Chem. 2021, 297, 101399. [Google Scholar] [CrossRef]
- Yuan, S.; Peng, L.; Park, J.J.; Hu, Y.; Devarkar, S.C.; Dong, M.B.; Shen, Q.; Wu, S.; Chen, S.; Lomakin, I.B.; et al. Nonstructural Protein 1 of SARS-CoV-2 Is a Potent Pathogenicity Factor Redirecting Host Protein Synthesis Machinery toward Viral RNA. Mol. Cell 2020, 80, 1055–1066.e6. [Google Scholar] [CrossRef]
- Nakagawa, K.; Makino, S. Mechanisms of Coronavirus Nsp1-Mediated Control of Host and Viral Gene Expression. Cells 2021, 10, 300. [Google Scholar] [CrossRef] [PubMed]
- Thoms, M.; Buschauer, R.; Ameismeier, M.; Koepke, L.; Denk, T.; Hirschenberger, M.; Kratzat, H.; Hayn, M.; Mackens-Kiani, T.; Cheng, J.; et al. Structural Basis for Translational Shutdown and Immune Evasion by the Nsp1 Protein of SARS-CoV-2. Science 2020, 369, 1249–1255. [Google Scholar] [CrossRef] [PubMed]
- Liao, M.; Wu, J.; Dai, M.; Li, H.; Yan, N.; Yuan, R.; Pan, C. Rapid Detection of SARS-CoV-2, Replicating or Non-Replicating, Using RT-PCR. Int. J. Infect. Dis. 2021, 104, 471–473. [Google Scholar] [CrossRef] [PubMed]
- Lo, C.-Y.; Tsai, T.-L.; Lin, C.-N.; Lin, C.-H.; Wu, H.-Y. Interaction of Coronavirus Nucleocapsid Protein with the 5′- and 3’-Ends of the Coronavirus Genome Is Involved in Genome Circularization and Negative-Strand RNA Synthesis. FEBS J. 2019, 286, 3222–3239. [Google Scholar] [CrossRef]
- Wu, C.-H.; Chen, P.-J.; Yeh, S.-H. Nucleocapsid Phosphorylation and RNA Helicase DDX1 Recruitment Enables Coronavirus Transition from Discontinuous to Continuous Transcription. Cell Host Microbe 2014, 16, 462–472. [Google Scholar] [CrossRef]
- Spencer, K.-A.; Hiscox, J.A. Characterisation of the RNA Binding Properties of the Coronavirus Infectious Bronchitis Virus Nucleocapsid Protein Amino-Terminal Region. FEBS Lett. 2006, 580, 5993–5998. [Google Scholar] [CrossRef]
- Caruso, Í.P.; Sanches, K.; Da Poian, A.T.; Pinheiro, A.S.; Almeida, F.C.L. Dynamics of the SARS-CoV-2 Nucleoprotein N-Terminal Domain Triggers RNA Duplex Destabilization. Biophys. J. 2021, 120, 2814–2827. [Google Scholar] [CrossRef]
- Cubuk, J.; Alston, J.J.; Incicco, J.J.; Singh, S.; Stuchell-Brereton, M.D.; Ward, M.D.; Zimmerman, M.I.; Vithani, N.; Griffith, D.; Wagoner, J.A.; et al. The SARS-CoV-2 Nucleocapsid Protein Is Dynamic, Disordered, and Phase Separates with RNA. Nat. Commun. 2021, 12, 1936. [Google Scholar] [CrossRef]
- Zhao, H.; Wu, D.; Nguyen, A.; Li, Y.; Adão, R.C.; Valkov, E.; Patterson, G.H.; Piszczek, G.; Schuck, P. Energetic and Structural Features of SARS-CoV-2 N-Protein Co-Assemblies with Nucleic Acids. iScience 2021, 24, 102523. [Google Scholar] [CrossRef]
- Chen, H.; Cui, Y.; Han, X.; Hu, W.; Sun, M.; Zhang, Y.; Wang, P.-H.; Song, G.; Chen, W.; Lou, J. Liquid-Liquid Phase Separation by SARS-CoV-2 Nucleocapsid Protein and RNA. Cell Res. 2020, 30, 1143–1145. [Google Scholar] [CrossRef]
- Jack, A.; Ferro, L.S.; Trnka, M.J.; Wehri, E.; Nadgir, A.; Nguyenla, X.; Fox, D.; Costa, K.; Stanley, S.; Schaletzky, J.; et al. SARS-CoV-2 Nucleocapsid Protein Forms Condensates with Viral Genomic RNA. PLoS Biol. 2021, 19, e3001425. [Google Scholar] [CrossRef] [PubMed]
- Dolnik, O.; Gerresheim, G.K.; Biedenkopf, N. New Perspectives on the Biogenesis of Viral Inclusion Bodies in Negative-Sense RNA Virus Infections. Cells 2021, 10, 1460. [Google Scholar] [CrossRef] [PubMed]
- Perdikari, T.M.; Murthy, A.C.; Ryan, V.H.; Watters, S.; Naik, M.T.; Fawzi, N.L. SARS-CoV-2 Nucleocapsid Protein Phase-Separates with RNA and with Human hnRNPs. EMBO J. 2020, 39, e106478. [Google Scholar] [CrossRef] [PubMed]
- de Castro, I.F.; Volonté, L.; Risco, C. Virus Factories: Biogenesis and Structural Design. Cell. Microbiol. 2013, 15, 24–34. [Google Scholar] [CrossRef] [PubMed]
- Netherton, C.L.; Wileman, T. Virus Factories, Double Membrane Vesicles and Viroplasm Generated in Animal Cells. Curr. Opin. Virol. 2011, 1, 381–387. [Google Scholar] [CrossRef]
- Klein, S.; Cortese, M.; Winter, S.L.; Wachsmuth-Melm, M.; Neufeldt, C.J.; Cerikan, B.; Stanifer, M.L.; Boulant, S.; Bartenschlager, R.; Chlanda, P. SARS-CoV-2 Structure and Replication Characterized by in Situ Cryo-Electron Tomography. Nat. Commun. 2020, 11, 5885. [Google Scholar] [CrossRef]
- Wheeler, J.R.; Matheny, T.; Jain, S.; Abrisch, R.; Parker, R. Distinct Stages in Stress Granule Assembly and Disassembly. Elife 2016, 5, e18413. [Google Scholar] [CrossRef]
- Chatterjee, S.; Kan, Y.; Brzezinski, M.; Koynov, K.; Regy, R.M.; Murthy, A.C.; Burke, K.A.; Michels, J.J.; Mittal, J.; Fawzi, N.L.; et al. Reversible Kinetic Trapping of FUS Biomolecular Condensates. Adv. Sci. 2022, 9, e2104247. [Google Scholar] [CrossRef]
- Hallegger, M.; Chakrabarti, A.M.; Lee, F.C.Y.; Lee, B.L.; Amalietti, A.G.; Odeh, H.M.; Copley, K.E.; Rubien, J.D.; Portz, B.; Kuret, K.; et al. TDP-43 Condensation Properties Specify Its RNA-Binding and Regulatory Repertoire. Cell 2021, 184, 4680–4696. [Google Scholar] [CrossRef]
- Etibor, T.A.; Yamauchi, Y.; Amorim, M.J. Liquid Biomolecular Condensates and Viral Lifecycles: Review and Perspectives. Viruses 2021, 13, 366. [Google Scholar] [CrossRef]
- Su, J.M.; Wilson, M.Z.; Samuel, C.E.; Ma, D. Formation and Function of Liquid-Like Viral Factories in Negative-Sense Single-Stranded RNA Virus Infections. Viruses 2021, 13, 126. [Google Scholar] [CrossRef] [PubMed]
- Hoenen, T.; Shabman, R.S.; Groseth, A.; Herwig, A.; Weber, M.; Schudt, G.; Dolnik, O.; Basler, C.F.; Becker, S.; Feldmann, H. Inclusion Bodies Are a Site of Ebolavirus Replication. J. Virol. 2012, 86, 11779–11788. [Google Scholar] [CrossRef] [PubMed]
- Cifuentes-Muñoz, N.; Branttie, J.; Slaughter, K.B.; Dutch, R.E. Human Metapneumovirus Induces Formation of Inclusion Bodies for Efficient Genome Replication and Transcription. J. Virol. 2017, 91, e01282-17. [Google Scholar] [CrossRef] [PubMed]
- Alenquer, M.; Vale-Costa, S.; Etibor, T.A.; Ferreira, F.; Sousa, A.L.; Amorim, M.J. Influenza A Virus Ribonucleoproteins Form Liquid Organelles at Endoplasmic Reticulum Exit Sites. Nat. Commun. 2019, 10, 1629. [Google Scholar] [CrossRef]
- Zhou, Y.; Su, J.M.; Samuel, C.E.; Ma, D. Measles Virus Forms Inclusion Bodies with Properties of Liquid Organelles. J. Virol. 2019, 93, e00948-19. [Google Scholar] [CrossRef]
- Lahaye, X.; Vidy, A.; Pomier, C.; Obiang, L.; Harper, F.; Gaudin, Y.; Blondel, D. Functional Characterization of Negri Bodies (NBs) in Rabies Virus-Infected Cells: Evidence That NBs Are Sites of Viral Transcription and Replication. J. Virol. 2009, 83, 7948–7958. [Google Scholar] [CrossRef]
- Nikolic, J.; Le Bars, R.; Lama, Z.; Scrima, N.; Lagaudrière-Gesbert, C.; Gaudin, Y.; Blondel, D. Negri Bodies Are Viral Factories with Properties of Liquid Organelles. Nat. Commun. 2017, 8, 58. [Google Scholar] [CrossRef]
- Galloux, M.; Risso-Ballester, J.; Richard, C.-A.; Fix, J.; Rameix-Welti, M.-A.; Eléouët, J.-F. Minimal Elements Required for the Formation of Respiratory Syncytial Virus Cytoplasmic Inclusion Bodies In Vivo and In Vitro. MBio 2020, 11. [Google Scholar] [CrossRef]
- Rincheval, V.; Lelek, M.; Gault, E.; Bouillier, C.; Sitterlin, D.; Blouquit-Laye, S.; Galloux, M.; Zimmer, C.; Eleouet, J.-F.; Rameix-Welti, M.-A. Functional Organization of Cytoplasmic Inclusion Bodies in Cells Infected by Respiratory Syncytial Virus. Nat. Commun. 2017, 8, 563. [Google Scholar] [CrossRef]
- Tawar, R.G.; Duquerroy, S.; Vonrhein, C.; Varela, P.F.; Damier-Piolle, L.; Castagné, N.; MacLellan, K.; Bedouelle, H.; Bricogne, G.; Bhella, D.; et al. Crystal Structure of a Nucleocapsid-like Nucleoprotein-RNA Complex of Respiratory Syncytial Virus. Science 2009, 326, 1279–1283. [Google Scholar] [CrossRef]
- Heinrich, B.S.; Maliga, Z.; Stein, D.A.; Hyman, A.A.; Whelan, S.P.J. Phase Transitions Drive the Formation of Vesicular Stomatitis Virus Replication Compartments. MBio 2018, 9, e02290-17. [Google Scholar] [CrossRef]
- Heinrich, B.S.; Cureton, D.K.; Rahmeh, A.A.; Whelan, S.P.J. Protein Expression Redirects Vesicular Stomatitis Virus RNA Synthesis to Cytoplasmic Inclusions. PLoS Pathog. 2010, 6, e1000958. [Google Scholar] [CrossRef]
- Zhu, N.; Wang, W.; Liu, Z.; Liang, C.; Wang, W.; Ye, F.; Huang, B.; Zhao, L.; Wang, H.; Zhou, W.; et al. Morphogenesis and Cytopathic Effect of SARS-CoV-2 Infection in Human Airway Epithelial Cells. Nat. Commun. 2020, 11, 3910. [Google Scholar] [CrossRef] [PubMed]
- Miyake, T.; Farley, C.M.; Neubauer, B.E.; Beddow, T.P.; Hoenen, T.; Engel, D.A. Ebola Virus Inclusion Body Formation and RNA Synthesis Are Controlled by a Novel Domain of Nucleoprotein Interacting with VP35. J. Virol. 2020, 94, e02100-19. [Google Scholar] [CrossRef]
- Wendt, L.; Brandt, J.; Bodmer, B.S.; Reiche, S.; Schmidt, M.L.; Traeger, S.; Hoenen, T. The Ebola Virus Nucleoprotein Recruits the Nuclear RNA Export Factor NXF1 into Inclusion Bodies to Facilitate Viral Protein Expression. Cells 2020, 9, 187. [Google Scholar] [CrossRef]
- Nelson, E.V.; Schmidt, K.M.; Deflubé, L.R.; Doğanay, S.; Banadyga, L.; Olejnik, J.; Hume, A.J.; Ryabchikova, E.; Ebihara, H.; Kedersha, N.; et al. Ebola Virus Does Not Induce Stress Granule Formation during Infection and Sequesters Stress Granule Proteins within Viral Inclusions. J. Virol. 2016, 90, 7268–7284. [Google Scholar] [CrossRef] [PubMed]
- Babu, M.M. The Contribution of Intrinsically Disordered Regions to Protein Function, Cellular Complexity, and Human Disease. Biochem. Soc. Trans. 2016, 44, 1185–1200. [Google Scholar] [CrossRef]
- Owen, I.; Shewmaker, F. The Role of Post-Translational Modifications in the Phase Transitions of Intrinsically Disordered Proteins. Int. J. Mol. Sci. 2019, 20, 5501. [Google Scholar] [CrossRef] [PubMed]
- van der Lee, R.; Buljan, M.; Lang, B.; Weatheritt, R.J.; Daughdrill, G.W.; Dunker, A.K.; Fuxreiter, M.; Gough, J.; Gsponer, J.; Jones, D.T.; et al. Classification of Intrinsically Disordered Regions and Proteins. Chem. Rev. 2014, 114, 6589–6631. [Google Scholar] [CrossRef]
- Xue, B.; Dunker, A.K.; Uversky, V.N. Orderly Order in Protein Intrinsic Disorder Distribution: Disorder in 3500 Proteomes from Viruses and the Three Domains of Life. J. Biomol. Struct. Dyn. 2012, 30, 137–149. [Google Scholar] [CrossRef]
- Radivojac, P.; Obradovic, Z.; Smith, D.K.; Zhu, G.; Vucetic, S.; Brown, C.J.; Lawson, J.D.; Dunker, A.K. Protein Flexibility and Intrinsic Disorder. Protein Sci. 2004, 13, 71–80. [Google Scholar] [CrossRef] [PubMed]
- Dogan, J.; Gianni, S.; Jemth, P. The Binding Mechanisms of Intrinsically Disordered Proteins. Phys. Chem. Chem. Phys. 2014, 16, 6323–6331. [Google Scholar] [CrossRef] [PubMed]
- Morris, O.M.; Torpey, J.H.; Isaacson, R.L. Intrinsically Disordered Proteins: Modes of Binding with Emphasis on Disordered Domains. Open Biol. 2021, 11, 210222. [Google Scholar] [CrossRef] [PubMed]
- Saito, A.; Shofa, M.; Ode, H.; Yumiya, M.; Hirano, J.; Okamoto, T.; Yoshimura, S.H. How Do Flaviviruses Hijack Host Cell Functions by Phase Separation? Viruses 2021, 13, 1479. [Google Scholar] [CrossRef] [PubMed]
- Mishra, P.M.; Verma, N.C.; Rao, C.; Uversky, V.N.; Nandi, C.K. Chapter One—Intrinsically Disordered Proteins of Viruses: Involvement in the Mechanism of Cell Regulation and Pathogenesis. In Progress in Molecular Biology and Translational Science; Uversky, V.N., Ed.; Academic Press: Cambridge, MA, USA, 2020; Volume 174, pp. 1–78. [Google Scholar] [CrossRef]
- Tokunaga, M.; Miyamoto, Y.; Suzuki, T.; Otani, M.; Inuki, S.; Esaki, T.; Nagao, C.; Mizuguchi, K.; Ohno, H.; Yoneda, Y.; et al. Novel Anti-Flavivirus Drugs Targeting the Nucleolar Distribution of Core Protein. Virology 2020, 541, 41–51. [Google Scholar] [CrossRef]
- Fraser, J.E.; Rawlinson, S.M.; Heaton, S.M.; Jans, D.A. Dynamic Nucleolar Targeting of Dengue Virus Polymerase NS5 in Response to Extracellular pH. J. Virol. 2016, 90, 5797–5807. [Google Scholar] [CrossRef]
- Aminev, A.G.; Amineva, S.P.; Palmenberg, A.C. Encephalomyocarditis Viral Protein 2A Localizes to Nucleoli and Inhibits Cap-Dependent mRNA Translation. Virus Res. 2003, 95, 45–57. [Google Scholar] [CrossRef]
- Yang, X.; Hu, Z.; Fan, S.; Zhang, Q.; Zhong, Y.; Guo, D.; Qin, Y.; Chen, M. Picornavirus 2A Protease Regulates Stress Granule Formation to Facilitate Viral Translation. PLoS Pathog. 2018, 14, e1006901. [Google Scholar] [CrossRef]
- Wu, S.; Wang, Y.; Lin, L.; Si, X.; Wang, T.; Zhong, X.; Tong, L.; Luan, Y.; Chen, Y.; Li, X.; et al. Protease 2A Induces Stress Granule Formation during Coxsackievirus B3 and Enterovirus 71 Infections. Virol. J. 2014, 11, 192. [Google Scholar] [CrossRef]
- Liu, D.; Ndongwe, T.P.; Puray-Chavez, M.; Casey, M.C.; Izumi, T.; Pathak, V.K.; Tedbury, P.R.; Sarafianos, S.G. Effect of P-Body Component Mov10 on HCV Virus Production and Infectivity. FASEB J. 2020, 34, 9433–9449. [Google Scholar] [CrossRef]
- Dougherty, J.D.; White, J.P.; Lloyd, R.E. Poliovirus-Mediated Disruption of Cytoplasmic Processing Bodies. J. Virol. 2011, 85, 64–75. [Google Scholar] [CrossRef] [PubMed]
- Giri, R.; Kumar, D.; Sharma, N.; Uversky, V.N. Intrinsically Disordered Side of the Zika Virus Proteome. Front. Cell. Infect. Microbiol. 2016, 6, 144. [Google Scholar] [CrossRef] [PubMed]
- Wurm, T.; Chen, H.; Hodgson, T.; Britton, P.; Brooks, G.; Hiscox, J.A. Localization to the Nucleolus Is a Common Feature of Coronavirus Nucleoproteins, and the Protein May Disrupt Host Cell Division. J. Virol. 2001, 75, 9345–9356. [Google Scholar] [CrossRef] [PubMed]
- Tenchov, R.; Zhou, Q.A. Intrinsically Disordered Proteins: Perspective on COVID-19 Infection and Drug Discovery. ACS Infect. Dis. 2022, 8, 422–432. [Google Scholar] [CrossRef]
- Kumar, A.; Kumar, A.; Kumar, P.; Garg, N.; Giri, R. SARS-CoV-2 NSP1 C-Terminal (residues 131-180) Is an Intrinsically Disordered Region in Isolation. Curr. Res. Virol Sci 2021, 2, 100007. [Google Scholar] [CrossRef]
- Tomaszewski, T.; DeVries, R.S.; Dong, M.; Bhatia, G.; Norsworthy, M.D.; Zheng, X.; Caetano-Anollés, G. New Pathways of Mutational Change in SARS-CoV-2 Proteomes Involve Regions of Intrinsic Disorder Important for Virus Replication and Release. Evol. Bioinform. Online 2020, 16, 1176934320965149. [Google Scholar] [CrossRef]
- Martínez-Flores, D.; Zepeda-Cervantes, J.; Cruz-Reséndiz, A.; Aguirre-Sampieri, S.; Sampieri, A.; Vaca, L. SARS-CoV-2 Vaccines Based on the Spike Glycoprotein and Implications of New Viral Variants. Front. Immunol. 2021, 12, 701501. [Google Scholar] [CrossRef]
- Dangi, T.; Sanchez, S.; Park, M.; Class, J.; Richner, M.; Richner, J.M.; Penaloza-MacMaster, P. Nucleocapsid-Specific Humoral Responses Improve the Control of SARS-CoV-2. bioRxiv 2022. [Google Scholar] [CrossRef]
- Giri, R.; Bhardwaj, T.; Shegane, M.; Gehi, B.R.; Kumar, P.; Gadhave, K.; Oldfield, C.J.; Uversky, V.N. Understanding COVID-19 via Comparative Analysis of Dark Proteomes of SARS-CoV-2, Human SARS and Bat SARS-like Coronaviruses. Cell. Mol. Life Sci. 2021, 78, 1655–1688. [Google Scholar] [CrossRef]
- Tidu, A.; Janvier, A.; Schaeffer, L.; Sosnowski, P.; Kuhn, L.; Hammann, P.; Westhof, E.; Eriani, G.; Martin, F. The Viral Protein NSP1 Acts as a Ribosome Gatekeeper for Shutting down Host Translation and Fostering SARS-CoV-2 Translation. RNA 2021, 27, 253–264. [Google Scholar] [CrossRef]
- Schubert, K.; Karousis, E.D.; Jomaa, A.; Scaiola, A.; Echeverria, B.; Gurzeler, L.-A.; Leibundgut, M.; Thiel, V.; Mühlemann, O.; Ban, N. SARS-CoV-2 Nsp1 Binds the Ribosomal mRNA Channel to Inhibit Translation. Nat. Struct. Mol. Biol. 2020, 27, 959–966. [Google Scholar] [CrossRef] [PubMed]
- Lapointe, C.P.; Grosely, R.; Johnson, A.G.; Wang, J.; Fernández, I.S.; Puglisi, J.D. Dynamic Competition between SARS-CoV-2 NSP1 and mRNA on the Human Ribosome Inhibits Translation Initiation. Proc. Natl. Acad. Sci. USA 2021, 118, e2017715118. [Google Scholar] [CrossRef] [PubMed]
- Yuan, S.; Balaji, S.; Lomakin, I.B.; Xiong, Y. Coronavirus Nsp1: Immune Response Suppression and Protein Expression Inhibition. Front. Microbiol. 2021, 12, 752214. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Miorin, L.; Makio, T.; Dehghan, I.; Gao, S.; Xie, Y.; Zhong, H.; Esparza, M.; Kehrer, T.; Kumar, A.; et al. Nsp1 Protein of SARS-CoV-2 Disrupts the mRNA Export Machinery to Inhibit Host Gene Expression. Sci. Adv. 2021, 7, eabe7386. [Google Scholar] [CrossRef]
- Luo, L.; Li, Z.; Zhao, T.; Ju, X.; Ma, P.; Jin, B.; Zhou, Y.; He, S.; Huang, J.; Xu, X.; et al. SARS-CoV-2 Nucleocapsid Protein Phase Separates with G3BPs to Disassemble Stress Granules and Facilitate Viral Production. Sci. Bull. 2021, 66, 1194–1204. [Google Scholar] [CrossRef]
- Cai, T.; Yu, Z.; Wang, Z.; Liang, C.; Richard, S. Arginine Methylation of SARS-Cov-2 Nucleocapsid Protein Regulates RNA Binding, Its Ability to Suppress Stress Granule Formation, and Viral Replication. J. Biol. Chem. 2021, 297, 100821. [Google Scholar] [CrossRef]
- Nabeel-Shah, S.; Lee, H.; Ahmed, N.; Burke, G.L.; Farhangmehr, S.; Ashraf, K.; Pu, S.; Braunschweig, U.; Zhong, G.; Wei, H.; et al. SARS-CoV-2 Nucleocapsid Protein Binds Host mRNAs and Attenuates Stress Granules to Impair Host Stress Response. iScience 2022, 25, 103562. [Google Scholar] [CrossRef]
- Wu, J.; Liu, W.; Gong, P. A Structural Overview of RNA-Dependent RNA Polymerases from the Flaviviridae Family. Int. J. Mol. Sci. 2015, 16, 12943–12957. [Google Scholar] [CrossRef]
- Semerdzhiev, S.A.; Fakhree, M.A.A.; Segers-Nolten, I.; Blum, C.; Claessens, M.M.A.E. Interactions between SARS-CoV-2 N-Protein and α-Synuclein Accelerate Amyloid Formation. ACS Chem. Neurosci. 2022, 13, 143–150. [Google Scholar] [CrossRef]
- Peluso, M.J.; Deeks, S.G.; Mustapic, M.; Kapogiannis, D.; Henrich, T.J.; Lu, S.; Goldberg, S.A.; Hoh, R.; Chen, J.Y.; Martinez, E.O.; et al. SARS-CoV-2 and Mitochondrial Proteins in Neural-Derived Exosomes of COVID-19. Ann. Neurol. 2022, 91, 772–781. [Google Scholar] [CrossRef]
- Visvabharathy, L.; Hanson, B.; Orban, Z.; Lim, P.H.; Palacio, N.M.; Jain, R.; Clark, J.R.; Graham, E.L.; Liotta, E.M.; Penaloza-MacMaster, P.; et al. Neuro-COVID Long-Haulers Exhibit Broad Dysfunction in T Cell Memory Generation and Responses to Vaccination. medRxiv 2021. medRxiv:2021.08.08.21261763. [Google Scholar] [CrossRef]
- Miglis, M.G.; Seliger, J.; Shaik, R.; Gibbons, C.H. A Case Series of Cutaneous Phosphorylated α-Synuclein in Long-COVID POTS. Clin. Auton. Res. 2022, 32, 209–212. [Google Scholar] [CrossRef] [PubMed]
- Mendez, A.S.; Ly, M.; González-Sánchez, A.M.; Hartenian, E.; Ingolia, N.T.; Cate, J.H.; Glaunsinger, B.A. The N-Terminal Domain of SARS-CoV-2 nsp1 Plays Key Roles in Suppression of Cellular Gene Expression and Preservation of Viral Gene Expression. Cell Rep. 2021, 37, 109841. [Google Scholar] [CrossRef] [PubMed]
- Sakuraba, S.; Xie, Q.; Kasahara, K.; Iwakiri, J.; Kono, H. Extended Ensemble Simulations of a SARS-CoV-2 nsp1-5′-UTR Complex. PLoS Comput. Biol. 2022, 18, e1009804. [Google Scholar] [CrossRef]
- Lokugamage, K.G.; Narayanan, K.; Huang, C.; Makino, S. Severe Acute Respiratory Syndrome Coronavirus Protein nsp1 Is a Novel Eukaryotic Translation Inhibitor That Represses Multiple Steps of Translation Initiation. J. Virol. 2012, 86, 13598–13608. [Google Scholar] [CrossRef]
- Finkel, Y.; Gluck, A.; Nachshon, A.; Winkler, R.; Fisher, T.; Rozman, B.; Mizrahi, O.; Lubelsky, Y.; Zuckerman, B.; Slobodin, B.; et al. SARS-CoV-2 Uses a Multipronged Strategy to Impede Host Protein Synthesis. Nature 2021, 594, 240–245. [Google Scholar] [CrossRef]
- Shemesh, M.; Aktepe, T.E.; Deerain, J.M.; McAuley, J.L.; Audsley, M.D.; David, C.T.; Purcell, D.F.J.; Urin, V.; Hartmann, R.; Moseley, G.W.; et al. SARS-CoV-2 Suppresses IFNβ Production Mediated by NSP1, 5, 6, 15, ORF6 and ORF7b but Does Not Suppress the Effects of Added Interferon. PLoS Pathog. 2021, 17, e1009800. [Google Scholar] [CrossRef]
- Joyce, G.F. RNA Evolution and the Origins of Life. Nature 1989, 338, 217–224. [Google Scholar] [CrossRef]
- Jankowsky, E.; Harris, M.E. Specificity and Nonspecificity in RNA-Protein Interactions. Nat. Rev. Mol. Cell Biol. 2015, 16, 533–544. [Google Scholar] [CrossRef]
- Lin, Y.; Protter, D.S.W.; Rosen, M.K.; Parker, R. Formation and Maturation of Phase-Separated Liquid Droplets by RNA-Binding Proteins. Mol. Cell 2015, 60, 208–219. [Google Scholar] [CrossRef]
- Kumar, S.; Mohapatra, T. Deciphering Epitranscriptome: Modification of mRNA Bases Provides a New Perspective for Post-Transcriptional Regulation of Gene Expression. Front. Cell Dev. Biol. 2021, 9, 628415. [Google Scholar] [CrossRef] [PubMed]
- Tsai, K.; Cullen, B.R. Epigenetic and Epitranscriptomic Regulation of Viral Replication. Nat. Rev. Microbiol. 2020, 18, 559–570. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Maimaitiyiming, Y.; Wang, L.; Cheng, X.; Hsu, C.-H. Modulation of Phase Separation by RNA: A Glimpse on N6-Methyladenosine Modification. Front. Cell Dev. Biol. 2021, 9, 786454. [Google Scholar] [CrossRef] [PubMed]
- Baquero-Perez, B.; Geers, D.; Díez, J. From A to m6A: The Emerging Viral Epitranscriptome. Viruses 2021, 13, 1049. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, E.M.; Courtney, D.G.; Tsai, K.; Cullen, B.R. Viral Epitranscriptomics. J. Virol. 2017, 91. [Google Scholar] [CrossRef]
- Lu, M.; Zhang, Z.; Xue, M.; Zhao, B.S.; Harder, O.; Li, A.; Liang, X.; Gao, T.Z.; Xu, Y.; Zhou, J.; et al. N6-Methyladenosine Modification Enables Viral RNA to Escape Recognition by RNA Sensor RIG-I. Nat. Microbiol. 2020, 5, 584–598. [Google Scholar] [CrossRef]
- Thiel, V. Viral RNA in an m6A Disguise. Nat. Microbiol. 2020, 5, 531–532. [Google Scholar] [CrossRef]
- Li, N.; Hui, H.; Bray, B.; Gonzalez, G.M.; Zeller, M.; Anderson, K.G.; Knight, R.; Smith, D.; Wang, Y.; Carlin, A.F.; et al. METTL3 Regulates Viral m6A RNA Modification and Host Cell Innate Immune Responses during SARS-CoV-2 Infection. Cell Rep. 2021, 35, 109091. [Google Scholar] [CrossRef]
- Winkler, R.; Gillis, E.; Lasman, L.; Safra, M.; Geula, S.; Soyris, C.; Nachshon, A.; Tai-Schmiedel, J.; Friedman, N.; Le-Trilling, V.T.K.; et al. m6A Modification Controls the Innate Immune Response to Infection by Targeting Type I Interferons. Nat. Immunol. 2019, 20, 173–182. [Google Scholar] [CrossRef]
- Reiter, R.J.; Rosales-Corral, S.; Tan, D.X.; Jou, M.J.; Galano, A.; Xu, B. Melatonin as a Mitochondria-Targeted Antioxidant: One of Evolution’s Best Ideas. Cell. Mol. Life Sci. 2017, 74, 3863–3881. [Google Scholar] [CrossRef]
- Kang, K.; Lee, K.; Park, S.; Byeon, Y.; Back, K. Molecular Cloning of Rice Serotonin N-Acetyltransferase, the Penultimate Gene in Plant Melatonin Biosynthesis. J. Pineal Res. 2013, 55, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Byeon, Y.; Choi, G.-H.; Lee, H.Y.; Back, K. Melatonin Biosynthesis Requires N-Acetylserotonin Methyltransferase Activity of Caffeic Acid O-Methyltransferase in Rice. J. Exp. Bot. 2015, 66, 6917–6925. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Choi, G.-H.; Back, K. Functional Characterization of Serotonin N-Acetyltransferase in Archaeon Thermoplasma Volcanium. Antioxidants 2022, 11, 596. [Google Scholar] [CrossRef] [PubMed]
- Doolittle, W.F. Evolution: Two Domains of Life or Three? Curr. Biol. 2020, 30, R177–R179. [Google Scholar] [CrossRef]
- Lau, Y.; Oamen, H.P.; Caudron, F. Protein Phase Separation during Stress Adaptation and Cellular Memory. Cells 2020, 9, 1302. [Google Scholar] [CrossRef]
- Franzmann, T.M.; Alberti, S. Protein Phase Separation as a Stress Survival Strategy. Cold Spring Harb. Perspect. Biol. 2019, 11, a034058. [Google Scholar] [CrossRef]
- Manchester, L.C.; Poeggeler, B.; Alvares, F.L.; Ogden, G.B.; Reiter, R.J. Melatonin Immunoreactivity in the Photosynthetic Prokaryote Rhodospirillum Rubrum: Implications for an Ancient Antioxidant System. Cell. Mol. Biol. Res. 1995, 41, 391–395. [Google Scholar]
- Tan, D.-X.; Manchester, L.C.; Liu, X.; Rosales-Corral, S.A.; Acuna-Castroviejo, D.; Reiter, R.J. Mitochondria and Chloroplasts as the Original Sites of Melatonin Synthesis: A Hypothesis Related to Melatonin’s Primary Function and Evolution in Eukaryotes. J. Pineal Res. 2013, 54, 127–138. [Google Scholar] [CrossRef]
- Pattanayak, G.K.; Liao, Y.; Wallace, E.W.J.; Budnik, B.; Drummond, D.A.; Rust, M.J. Daily Cycles of Reversible Protein Condensation in Cyanobacteria. Cell Rep. 2020, 32, 108032. [Google Scholar] [CrossRef]
- Guilhas, B.; Walter, J.-C.; Rech, J.; David, G.; Walliser, N.O.; Palmeri, J.; Mathieu-Demaziere, C.; Parmeggiani, A.; Bouet, J.-Y.; Le Gall, A.; et al. ATP-Driven Separation of Liquid Phase Condensates in Bacteria. Mol. Cell 2020, 79, 293–303.e4. [Google Scholar] [CrossRef]
- Tan, D.-X.; Manchester, L.C.; Qin, L.; Reiter, R.J. Melatonin: A Mitochondrial Targeting Molecule Involving Mitochondrial Protection and Dynamics. Int. J. Mol. Sci. 2016, 17, 2124. [Google Scholar] [CrossRef] [PubMed]
- Martín, M.; Macías, M.; León, J.; Escames, G.; Khaldy, H.; Acuña-Castroviejo, D. Melatonin Increases the Activity of the Oxidative Phosphorylation Enzymes and the Production of ATP in Rat Brain and Liver Mitochondria. Int. J. Biochem. Cell Biol. 2002, 34, 348–357. [Google Scholar] [CrossRef]
- Tan, D.-X.; Zheng, X.; Kong, J.; Manchester, L.C.; Hardeland, R.; Kim, S.J.; Xu, X.; Reiter, R.J. Fundamental Issues Related to the Origin of Melatonin and Melatonin Isomers during Evolution: Relation to Their Biological Functions. Int. J. Mol. Sci. 2014, 15, 15858–15890. [Google Scholar] [CrossRef] [PubMed]
- Coon, S.L.; Klein, D.C. Evolution of Arylalkylamine N-Acetyltransferase: Emergence and Divergence. Mol. Cell. Endocrinol. 2006, 252, 2–10. [Google Scholar] [CrossRef] [PubMed]
- Patel, A.; Malinovska, L.; Saha, S.; Wang, J.; Alberti, S.; Krishnan, Y.; Hyman, A.A. ATP as a Biological Hydrotrope. Science 2017, 356, 753–756. [Google Scholar] [CrossRef]
- Snead, W.T.; Gladfelter, A.S. The Control Centers of Biomolecular Phase Separation: How Membrane Surfaces, PTMs, and Active Processes Regulate Condensation. Mol. Cell 2019, 76, 295–305. [Google Scholar] [CrossRef]
- Henninger, J.E.; Oksuz, O.; Shrinivas, K.; Sagi, I.; LeRoy, G.; Zheng, M.M.; Andrews, J.O.; Zamudio, A.V.; Lazaris, C.; Hannett, N.M.; et al. RNA-Mediated Feedback Control of Transcriptional Condensates. Cell 2021, 184, 207–225.e24. [Google Scholar] [CrossRef]
- Manchester, K.L. Free Energy ATP Hydrolysis and Phosphorylation Potential. Biochem. Educ. 1980, 8, 70–72. [Google Scholar] [CrossRef]
- Jain, S.; Wheeler, J.R.; Walters, R.W.; Agrawal, A.; Barsic, A.; Parker, R. ATPase-Modulated Stress Granules Contain a Diverse Proteome and Substructure. Cell 2016, 164, 487–498. [Google Scholar] [CrossRef]
- Aida, H.; Shigeta, Y.; Harada, R. The Role of ATP in Solubilizing RNA-Binding Protein Fused in Sarcoma. Proteins 2022, 90, 1606–1612. [Google Scholar] [CrossRef]
- Conn, G.L.; Gittis, A.G.; Lattman, E.E.; Misra, V.K.; Draper, D.E. A Compact RNA Tertiary Structure Contains a Buried Backbone-K+ Complex. J. Mol. Biol. 2002, 318, 963–973. [Google Scholar] [CrossRef]
- Garcia-Jove Navarro, M.; Kashida, S.; Chouaib, R.; Souquere, S.; Pierron, G.; Weil, D.; Gueroui, Z. RNA Is a Critical Element for the Sizing and the Composition of Phase-Separated RNA-Protein Condensates. Nat. Commun. 2019, 10, 3230. [Google Scholar] [CrossRef] [PubMed]
- Campos-Melo, D.; Hawley, Z.C.E.; Droppelmann, C.A.; Strong, M.J. The Integral Role of RNA in Stress Granule Formation and Function. Front. Cell Dev. Biol. 2021, 9, 621779. [Google Scholar] [CrossRef]
- Dang, M.; Li, Y.; Song, J. ATP Biphasically Modulates LLPS of SARS-CoV-2 Nucleocapsid Protein and Specifically Binds Its RNA-Binding Domain. Biochem. Biophys. Res. Commun. 2021, 541, 50–55. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Ye, Q.; Singh, D.; Cao, Y.; Diedrich, J.K.; Yates, J.R., 3rd; Villa, E.; Cleveland, D.W.; Corbett, K.D. The SARS-CoV-2 Nucleocapsid Phosphoprotein Forms Mutually Exclusive Condensates with RNA and the Membrane-Associated M Protein. Nat. Commun. 2021, 12, 502. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; He, H.; Wang, L.; Zhang, N.; Huang, H.; Xiong, Q.; Yan, Y.; Wu, N.; Ren, H.; Han, H.; et al. Virus-Triggered ATP Release Limits Viral Replication through Facilitating IFN-β Production in a P2X7-Dependent Manner. J. Immunol. 2017, 199, 1372–1381. [Google Scholar] [CrossRef]
- Kouzaki, H.; Iijima, K.; Kobayashi, T.; O’Grady, S.M.; Kita, H. The Danger Signal, Extracellular ATP, Is a Sensor for an Airborne Allergen and Triggers IL-33 Release and Innate Th2-Type Responses. J. Immunol. 2011, 186, 4375–4387. [Google Scholar] [CrossRef]
- Strauss, M.; Hofhaus, G.; Schröder, R.R.; Kühlbrandt, W. Dimer Ribbons of ATP Synthase Shape the Inner Mitochondrial Membrane. EMBO J. 2008, 27, 1154–1160. [Google Scholar] [CrossRef]
- Wang, T.; Chen, Z.; Wang, X.; Shyy, J.Y.-J.; Zhu, Y. Cholesterol Loading Increases the Translocation of ATP Synthase Beta Chain into Membrane Caveolae in Vascular Endothelial Cells. Biochim. Biophys. Acta 2006, 1761, 1182–1190. [Google Scholar] [CrossRef]
- Kim, B.-W.; Choo, H.-J.; Lee, J.-W.; Kim, J.-H.; Ko, Y.-G. Extracellular ATP Is Generated by ATP Synthase Complex in Adipocyte Lipid Rafts. Exp. Mol. Med. 2004, 36, 476–485. [Google Scholar] [CrossRef]
- Russo, C.; Raiden, S.; Algieri, S.; De Carli, N.; Davenport, C.; Sarli, M.; Bruera, M.J.; Seery, V.; Sananez, I.; Simaz, N.; et al. Extracellular ATP and Imbalance of CD4+ T Cell Compartment in Pediatric COVID-19. Front. Cell. Infect. Microbiol. 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Sant, A.J.; DiPiazza, A.T.; Nayak, J.L.; Rattan, A.; Richards, K.A. CD4 T Cells in Protection from Influenza Virus: Viral Antigen Specificity and Functional Potential. Immunol. Rev. 2018, 284, 91–105. [Google Scholar] [CrossRef] [PubMed]
- Iwamura, A.P.D.; Tavares da Silva, M.R.; Hümmelgen, A.L.; Soeiro Pereira, P.V.; Falcai, A.; Grumach, A.S.; Goudouris, E.; Neto, A.C.; Prando, C. Immunity and Inflammatory Biomarkers in COVID-19: A Systematic Review. Rev. Med. Virol. 2021, 31, e2199. [Google Scholar] [CrossRef] [PubMed]
- Loh, D.; Reiter, R.J. Melatonin: Regulation of Biomolecular Condensates in Neurodegenerative Disorders. Antioxidants 2021, 10, 1483. [Google Scholar] [CrossRef] [PubMed]
- Mehrzadi, S.; Karimi, M.Y.; Fatemi, A.; Reiter, R.J.; Hosseinzadeh, A. SARS-CoV-2 and Other Coronaviruses Negatively Influence Mitochondrial Quality Control: Beneficial Effects of Melatonin. Pharmacol. Ther. 2021, 224, 107825. [Google Scholar] [CrossRef]
- McBride, H.M.; Neuspiel, M.; Wasiak, S. Mitochondria: More than Just a Powerhouse. Curr. Biol. 2006, 16, R551–R560. [Google Scholar] [CrossRef]
- Brown, G.C. Control of Respiration and ATP Synthesis in Mammalian Mitochondria and Cells. Biochem. J. 1992, 284 Pt 1, 1–13. [Google Scholar] [CrossRef]
- Elesela, S.; Lukacs, N.W. Role of Mitochondria in Viral Infections. Life 2021, 11, 232. [Google Scholar] [CrossRef]
- Khan, M.; Syed, G.H.; Kim, S.-J.; Siddiqui, A. Mitochondrial Dynamics and Viral Infections: A Close Nexus. Biochim. Biophys. Acta 2015, 1853 Pt B, 2822–2833. [Google Scholar] [CrossRef]
- Kim, S.-J.; Khan, M.; Quan, J.; Till, A.; Subramani, S.; Siddiqui, A. Hepatitis B Virus Disrupts Mitochondrial Dynamics: Induces Fission and Mitophagy to Attenuate Apoptosis. PLoS Pathog. 2013, 9, e1003722. [Google Scholar] [CrossRef]
- Kim, S.-J.; Syed, G.H.; Khan, M.; Chiu, W.-W.; Sohail, M.A.; Gish, R.G.; Siddiqui, A. Hepatitis C Virus Triggers Mitochondrial Fission and Attenuates Apoptosis to Promote Viral Persistence. Proc. Natl. Acad. Sci. USA 2014, 111, 6413–6418. [Google Scholar] [CrossRef]
- Gatti, P.; Ilamathi, H.S.; Todkar, K.; Germain, M. Mitochondria Targeted Viral Replication and Survival Strategies-Prospective on SARS-CoV-2. Front. Pharmacol. 2020, 11, 578599. [Google Scholar] [CrossRef]
- Shang, C.; Liu, Z.; Zhu, Y.; Lu, J.; Ge, C.; Zhang, C.; Li, N.; Jin, N.; Li, Y.; Tian, M.; et al. SARS-CoV-2 Causes Mitochondrial Dysfunction and Mitophagy Impairment. Front. Microbiol. 2021, 12, 780768. [Google Scholar] [CrossRef]
- Wu, K.E.; Fazal, F.M.; Parker, K.R.; Zou, J.; Chang, H.Y. RNA-GPS Predicts SARS-CoV-2 RNA Residency to Host Mitochondria and Nucleolus. Cell Syst. 2020, 11, 102–108.e3. [Google Scholar] [CrossRef]
- Pliss, A.; Kuzmin, A.N.; Prasad, P.N.; Mahajan, S.D. Mitochondrial Dysfunction: A Prelude to Neuropathogenesis of SARS-CoV-2. ACS Chem. Neurosci. 2022, 13, 308–312. [Google Scholar] [CrossRef]
- Cortese, M.; Lee, J.-Y.; Cerikan, B.; Neufeldt, C.J.; Oorschot, V.M.J.; Köhrer, S.; Hennies, J.; Schieber, N.L.; Ronchi, P.; Mizzon, G.; et al. Integrative Imaging Reveals SARS-CoV-2-Induced Reshaping of Subcellular Morphologies. Cell Host Microbe 2020, 28, 853–866.e5. [Google Scholar] [CrossRef]
- Santos, A.F.; Póvoa, P.; Paixão, P.; Mendonça, A.; Taborda-Barata, L. Changes in Glycolytic Pathway in SARS-COV 2 Infection and Their Importance in Understanding the Severity of COVID-19. Front. Chem. 2021, 9, 685196. [Google Scholar] [CrossRef]
- Ma, K.; Wu, H.; Li, P.; Li, B. LC3-II May Mediate ATR-Induced Mitophagy in Dopaminergic Neurons through SQSTM1/p62 Pathway. Acta Biochim. Biophys. Sin. 2018, 50, 1047–1061. [Google Scholar] [CrossRef]
- Wang, S.; Zhao, Z.; Feng, X.; Cheng, Z.; Xiong, Z.; Wang, T.; Lin, J.; Zhang, M.; Hu, J.; Fan, Y.; et al. Melatonin Activates Parkin Translocation and Rescues the Impaired Mitophagy Activity of Diabetic Cardiomyopathy through Mst1 Inhibition. J. Cell. Mol. Med. 2018, 22, 5132–5144. [Google Scholar] [CrossRef]
- da Costa, L.S.; Outlioua, A.; Anginot, A.; Akarid, K.; Arnoult, D. RNA Viruses Promote Activation of the NLRP3 Inflammasome through Cytopathogenic Effect-Induced Potassium Efflux. Cell Death Dis. 2019, 10, 346. [Google Scholar] [CrossRef]
- Muñoz-Planillo, R.; Kuffa, P.; Martínez-Colón, G.; Smith, B.L.; Rajendiran, T.M.; Núñez, G. K+ Efflux Is the Common Trigger of NLRP3 Inflammasome Activation by Bacterial Toxins and Particulate Matter. Immunity 2013, 38, 1142–1153. [Google Scholar] [CrossRef]
- Ichinohe, T.; Pang, I.K.; Iwasaki, A. Influenza Virus Activates Inflammasomes via Its Intracellular M2 Ion Channel. Nat. Immunol. 2010, 11, 404–410. [Google Scholar] [CrossRef]
- de Zoete, M.R.; Palm, N.W.; Zhu, S.; Flavell, R.A. Inflammasomes. Cold Spring Harb. Perspect. Biol. 2014, 6, a016287. [Google Scholar] [CrossRef]
- DeDiego, M.L.; Nieto-Torres, J.L.; Jimenez-Guardeño, J.M.; Regla-Nava, J.A.; Castaño-Rodriguez, C.; Fernandez-Delgado, R.; Usera, F.; Enjuanes, L. Coronavirus Virulence Genes with Main Focus on SARS-CoV Envelope Gene. Virus Res. 2014, 194, 124–137. [Google Scholar] [CrossRef]
- Farag, N.S.; Breitinger, U.; Breitinger, H.G.; El Azizi, M.A. Viroporins and Inflammasomes: A Key to Understand Virus-Induced Inflammation. Int. J. Biochem. Cell Biol. 2020, 122, 105738. [Google Scholar] [CrossRef]
- McClenaghan, C.; Hanson, A.; Lee, S.-J.; Nichols, C.G. Coronavirus Proteins as Ion Channels: Current and Potential Research. Front. Immunol. 2020, 11, 573339. [Google Scholar] [CrossRef]
- Arya, R.; Kumari, S.; Pandey, B.; Mistry, H.; Bihani, S.C.; Das, A.; Prashar, V.; Gupta, G.D.; Panicker, L.; Kumar, M. Structural Insights into SARS-CoV-2 Proteins. J. Mol. Biol. 2021, 433, 166725. [Google Scholar] [CrossRef]
- Cao, Y.; Yang, R.; Lee, I.; Zhang, W.; Sun, J.; Wang, W.; Meng, X. Characterization of the SARS-CoV-2 E Protein: Sequence, Structure, Viroporin, and Inhibitors. Protein Sci. 2021, 30, 1114–1130. [Google Scholar] [CrossRef]
- Mandala, V.S.; McKay, M.J.; Shcherbakov, A.A.; Dregni, A.J.; Kolocouris, A.; Hong, M. Structure and Drug Binding of the SARS-CoV-2 Envelope Protein Transmembrane Domain in Lipid Bilayers. Nat. Struct. Mol. Biol. 2020, 27, 1202–1208. [Google Scholar] [CrossRef]
- Breitinger, U.; Ali, N.K.M.; Sticht, H.; Breitinger, H.-G. Inhibition of SARS CoV Envelope Protein by Flavonoids and Classical Viroporin Inhibitors. Front. Microbiol. 2021, 12, 692423. [Google Scholar] [CrossRef]
- Rizwan, T.; Kothidar, A.; Meghwani, H.; Sharma, V.; Shobhawat, R.; Saini, R.; Vaishnav, H.K.; Singh, V.; Pratap, M.; Sihag, H.; et al. Comparative Analysis of SARS-CoV-2 Envelope Viroporin Mutations from COVID-19 Deceased and Surviving Patients Revealed Implications on Its Ion-Channel Activities and Correlation with Patient Mortality. J. Biomol. Struct. Dyn. 2021, 1–16. [Google Scholar] [CrossRef]
- Cao, Y.; Yang, R.; Wang, W.; Lee, I.; Zhang, R.; Zhang, W.; Sun, J.; Xu, B.; Meng, X. Computational Study of the Ion and Water Permeation and Transport Mechanisms of the SARS-CoV-2 Pentameric E Protein Channel. Front. Mol. Biosci 2020, 7, 565797. [Google Scholar] [CrossRef]
- Liao, Y.; Yuan, Q.; Torres, J.; Tam, J.P.; Liu, D.X. Biochemical and Functional Characterization of the Membrane Association and Membrane Permeabilizing Activity of the Severe Acute Respiratory Syndrome Coronavirus Envelope Protein. Virology 2006, 349, 264–275. [Google Scholar] [CrossRef]
- Pervushin, K.; Tan, E.; Parthasarathy, K.; Lin, X.; Jiang, F.L.; Yu, D.; Vararattanavech, A.; Soong, T.W.; Liu, D.X.; Torres, J. Structure and Inhibition of the SARS Coronavirus Envelope Protein Ion Channel. PLoS Pathog. 2009, 5, e1000511. [Google Scholar] [CrossRef]
- Mehregan, A.; Pérez-Conesa, S.; Zhuang, Y.; Elbahnsi, A.; Pasini, D.; Lindahl, E.; Howard, R.J.; Ulens, C.; Delemotte, L. Probing Effects of the SARS-CoV-2 E Protein on Membrane Curvature and Intracellular Calcium. Biochim. Biophys. Acta Biomembr. 2022, 1864, 183994. [Google Scholar] [CrossRef]
- Nardacci, R.; Colavita, F.; Castilletti, C.; Lapa, D.; Matusali, G.; Meschi, S.; Del Nonno, F.; Colombo, D.; Capobianchi, M.R.; Zumla, A.; et al. Evidences for Lipid Involvement in SARS-CoV-2 Cytopathogenesis. Cell Death Dis. 2021, 12, 263. [Google Scholar] [CrossRef]
- Wang, P.; Luo, R.; Zhang, M.; Wang, Y.; Song, T.; Tao, T.; Li, Z.; Jin, L.; Zheng, H.; Chen, W.; et al. A Cross-Talk between Epithelium and Endothelium Mediates Human Alveolar–capillary Injury during SARS-CoV-2 Infection. Cell Death Dis. 2020, 11, 1–17. [Google Scholar] [CrossRef]
- Mannella, C.A.; Pfeiffer, D.R.; Bradshaw, P.C.; Moraru, I.I.; Slepchenko, B.; Loew, L.M.; Hsieh, C.E.; Buttle, K.; Marko, M. Topology of the Mitochondrial Inner Membrane: Dynamics and Bioenergetic Implications. IUBMB Life 2001, 52, 93–100. [Google Scholar] [CrossRef]
- Zorova, L.D.; Popkov, V.A.; Plotnikov, E.Y.; Silachev, D.N.; Pevzner, I.B.; Jankauskas, S.S.; Babenko, V.A.; Zorov, S.D.; Balakireva, A.V.; Juhaszova, M.; et al. Mitochondrial Membrane Potential. Anal. Biochem. 2018, 552, 50–59. [Google Scholar] [CrossRef]
- Zick, M.; Rabl, R.; Reichert, A.S. Cristae Formation-Linking Ultrastructure and Function of Mitochondria. Biochim. Biophys. Acta 2009, 1793, 5–19. [Google Scholar] [CrossRef]
- Gilkerson, R.W.; Selker, J.M.L.; Capaldi, R.A. The Cristal Membrane of Mitochondria Is the Principal Site of Oxidative Phosphorylation. FEBS Lett. 2003, 546, 355–358. [Google Scholar] [CrossRef]
- Mitchell, P. Coupling of Phosphorylation to Electron and Hydrogen Transfer by a Chemi-Osmotic Type of Mechanism. Nature 1961, 191, 144–148. [Google Scholar] [CrossRef]
- Wolf, D.M.; Segawa, M.; Kondadi, A.K.; Anand, R.; Bailey, S.T.; Reichert, A.S.; van der Bliek, A.M.; Shackelford, D.B.; Liesa, M.; Shirihai, O.S. Individual Cristae within the Same Mitochondrion Display Different Membrane Potentials and Are Functionally Independent. EMBO J. 2019, 38, e101056. [Google Scholar] [CrossRef]
- Rieger, B.; Junge, W.; Busch, K.B. Lateral pH Gradient between OXPHOS Complex IV and F(0)F(1) ATP-Synthase in Folded Mitochondrial Membranes. Nat. Commun. 2014, 5, 3103. [Google Scholar] [CrossRef]
- Garcia, G.C.; Bartol, T.M.; Phan, S.; Bushong, E.A.; Perkins, G.; Sejnowski, T.J.; Ellisman, M.H.; Skupin, A. Mitochondrial Morphology Provides a Mechanism for Energy Buffering at Synapses. Sci. Rep. 2019, 9, 18306. [Google Scholar] [CrossRef]
- Scott, I.D.; Nicholls, D.G. Energy Transduction in Intact Synaptosomes. Influence of Plasma-Membrane Depolarization on the Respiration and Membrane Potential of Internal Mitochondria Determined in Situ. Biochem. J. 1980, 186, 21–33. [Google Scholar] [CrossRef]
- Mannella, C.A. Consequences of Folding the Mitochondrial Inner Membrane. Front. Physiol. 2020, 11, 536. [Google Scholar] [CrossRef]
- Gottlieb, E.; Armour, S.M.; Harris, M.H.; Thompson, C.B. Mitochondrial Membrane Potential Regulates Matrix Configuration and Cytochrome c Release during Apoptosis. Cell Death Differ. 2003, 10, 709–717. [Google Scholar] [CrossRef]
- Rasola, A.; Bernardi, P. Mitochondrial Permeability Transition in Ca(2+)-Dependent Apoptosis and Necrosis. Cell Calcium 2011, 50, 222–233. [Google Scholar] [CrossRef]
- Liesa, M. Why Does a Mitochondrion Need Its Individual Cristae to Be Functionally Autonomous? Mol. Cell Oncol. 2020, 7, 1705119. [Google Scholar] [CrossRef]
- Twig, G.; Elorza, A.; Molina, A.J.A.; Mohamed, H.; Wikstrom, J.D.; Walzer, G.; Stiles, L.; Haigh, S.E.; Katz, S.; Las, G.; et al. Fission and Selective Fusion Govern Mitochondrial Segregation and Elimination by Autophagy. EMBO J. 2008, 27, 433–446. [Google Scholar] [CrossRef] [PubMed]
- Twig, G.; Shirihai, O.S. The Interplay between Mitochondrial Dynamics and Mitophagy. Antioxid. Redox Signal. 2011, 14, 1939–1951. [Google Scholar] [CrossRef]
- Medini, H.; Zirman, A.; Mishmar, D. Immune System Cells from COVID-19 Patients Display Compromised Mitochondrial-Nuclear Expression Co-Regulation and Rewiring toward Glycolysis. iScience 2021, 24, 103471. [Google Scholar] [CrossRef]
- Wu, M.; Neilson, A.; Swift, A.L.; Moran, R.; Tamagnine, J.; Parslow, D.; Armistead, S.; Lemire, K.; Orrell, J.; Teich, J.; et al. Multiparameter Metabolic Analysis Reveals a Close Link between Attenuated Mitochondrial Bioenergetic Function and Enhanced Glycolysis Dependency in Human Tumor Cells. Am. J. Physiol. Cell Physiol. 2007, 292, C125–C136. [Google Scholar] [CrossRef]
- Codo, A.C.; Davanzo, G.G.; de Brito Monteiro, L.; de Souza, G.F.; Muraro, S.P.; Virgilio-da-Silva, J.V.; Prodonoff, J.S.; Carregari, V.C.; de Biagi Junior, C.A.O.; Crunfli, F.; et al. Elevated Glucose Levels Favor SARS-CoV-2 Infection and Monocyte Response through a HIF-1α/Glycolysis-Dependent Axis. Cell Metab. 2020, 32, 437–446.e5. [Google Scholar] [CrossRef]
- Aklima, J.; Onojima, T.; Kimura, S.; Umiuchi, K.; Shibata, T.; Kuraoka, Y.; Oie, Y.; Suganuma, Y.; Ohta, Y. Effects of Matrix pH on Spontaneous Transient Depolarization and Reactive Oxygen Species Production in Mitochondria. Front. Cell Dev. Biol. 2021, 9, 692776. [Google Scholar] [CrossRef]
- Moreno Davila, H. Molecular and Functional Diversity of Voltage-Gated Calcium Channels. Ann. N. Y. Acad. Sci. 1999, 868, 102–117. [Google Scholar] [CrossRef]
- Pitt, G.S.; Matsui, M.; Cao, C. Voltage-Gated Calcium Channels in Nonexcitable Tissues. Annu. Rev. Physiol. 2021, 83, 183–203. [Google Scholar] [CrossRef]
- Catterall, W.A. Voltage-Gated Calcium Channels. Cold Spring Harb. Perspect. Biol. 2011, 3, a003947. [Google Scholar] [CrossRef]
- Agirre, A.; Barco, A.; Carrasco, L.; Nieva, J.L. Viroporin-Mediated Membrane Permeabilization. Pore Formation by Nonstructural Poliovirus 2B Protein. J. Biol. Chem. 2002, 277, 40434–40441. [Google Scholar] [CrossRef]
- Firth, A.E.; Chung, B.Y.; Fleeton, M.N.; Atkins, J.F. Discovery of Frameshifting in Alphavirus 6K Resolves a 20-Year Enigma. Virol. J. 2008, 5, 108. [Google Scholar] [CrossRef] [PubMed]
- González, M.E. Vpu Protein: The Viroporin Encoded by HIV-1. Viruses 2015, 7, 4352–4368. [Google Scholar] [CrossRef] [PubMed]
- To, J.; Torres, J. Viroporins in the Influenza Virus. Cells 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Lescar, J.; Tam, J.P.; Liu, D.X. Expression of SARS-Coronavirus Envelope Protein in Escherichia Coli Cells Alters Membrane Permeability. Biochem. Biophys. Res. Commun. 2004, 325, 374–380. [Google Scholar] [CrossRef]
- Landstrom, A.P.; Dobrev, D.; Wehrens, X.H.T. Calcium Signaling and Cardiac Arrhythmias. Circ. Res. 2017, 120, 1969–1993. [Google Scholar] [CrossRef]
- Larsen, H.E.; Bardsley, E.N.; Lefkimmiatis, K.; Paterson, D.J. Dysregulation of Neuronal Ca2+ Channel Linked to Heightened Sympathetic Phenotype in Prohypertensive States. J. Neurosci. 2016, 36, 8562–8573. [Google Scholar] [CrossRef]
- Jamal, S.M.; Landers, D.B.; Hollenberg, S.M.; Turi, Z.G.; Glotzer, T.V.; Tancredi, J.; Parrillo, J.E. Prospective Evaluation of Autonomic Dysfunction in Post-Acute Sequela of COVID-19. J. Am. Coll. Cardiol. 2022. [Google Scholar] [CrossRef]
- Dani, M.; Dirksen, A.; Taraborrelli, P.; Torocastro, M.; Panagopoulos, D.; Sutton, R.; Lim, P.B. Autonomic Dysfunction in “Long COVID”: Rationale, Physiology and Management Strategies. Clin. Med. 2021, 21, e63–e67. [Google Scholar] [CrossRef]
- Papadopoulou, M.; Bakola, E.; Papapostolou, A.; Stefanou, M.-I.; Gaga, M.; Zouvelou, V.; Michopoulos, I.; Tsivgoulis, G. Autonomic Dysfunction in Long-COVID Syndrome: A Neurophysiological and Neurosonology Study. J. Neurol. 2022, 1–2. [Google Scholar] [CrossRef]
- Raj, S.R.; Arnold, A.C.; Barboi, A.; Claydon, V.E.; Limberg, J.K.; Lucci, V.-E.M.; Numan, M.; Peltier, A.; Snapper, H.; Vernino, S.; et al. Long-COVID Postural Tachycardia Syndrome: An American Autonomic Society Statement. Clin. Auton. Res. 2021, 31, 365–368. [Google Scholar] [CrossRef]
- Bisaccia, G.; Ricci, F.; Recce, V.; Serio, A.; Iannetti, G.; Chahal, A.A.; Ståhlberg, M.; Khanji, M.Y.; Fedorowski, A.; Gallina, S. Post-Acute Sequelae of COVID-19 and Cardiovascular Autonomic Dysfunction: What Do We Know? J. Cardiovasc. Dev. Dis 2021, 8, 156. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Cao, R.; Zhong, W. Host Calcium Channels and Pumps in Viral Infections. Cells 2019, 9, 94. [Google Scholar] [CrossRef] [PubMed]
- Hyser, J.M.; Estes, M.K. Pathophysiological Consequences of Calcium-Conducting Viroporins. Annu Rev. Virol. 2015, 2, 473–496. [Google Scholar] [CrossRef] [PubMed]
- Berktaş, B.M.; Gökçek, A.; Hoca, N.T.; Koyuncu, A. COVID-19 Illness and Treatment Decrease Bone Mineral Density of Surviving Hospitalized Patients. Eur. Rev. Med. Pharmacol. Sci. 2022, 26, 3046–3056. [Google Scholar] [CrossRef]
- Mandala, V.S.; Loftis, A.R.; Shcherbakov, A.A.; Pentelute, B.L.; Hong, M. Atomic Structures of Closed and Open Influenza B M2 Proton Channel Reveal the Conduction Mechanism. Nat. Struct. Mol. Biol. 2020, 27, 160–167. [Google Scholar] [CrossRef]
- Gargan, S.; Stevenson, N.J. Unravelling the Immunomodulatory Effects of Viral Ion Channels, towards the Treatment of Disease. Viruses 2021, 13, 2165. [Google Scholar] [CrossRef]
- Bohmwald, K.; Gálvez, N.M.S.; Andrade, C.A.; Mora, V.P.; Muñoz, J.T.; González, P.A.; Riedel, C.A.; Kalergis, A.M. Modulation of Adaptive Immunity and Viral Infections by Ion Channels. Front. Physiol. 2021, 12, 736681. [Google Scholar] [CrossRef]
- Feske, S.; Wulff, H.; Skolnik, E.Y. Ion Channels in Innate and Adaptive Immunity. Annu. Rev. Immunol. 2015, 33, 291–353. [Google Scholar] [CrossRef]
- Kaivola, J.; Nyman, T.A.; Matikainen, S. Inflammasomes and SARS-CoV-2 Infection. Viruses 2021, 13, 2513. [Google Scholar] [CrossRef]
- Campbell, G.R.; To, R.K.; Hanna, J.; Spector, S.A. SARS-CoV-2, SARS-CoV-1, and HIV-1 Derived ssRNA Sequences Activate the NLRP3 Inflammasome in Human Macrophages through a Non-Classical Pathway. iScience 2021, 24, 102295. [Google Scholar] [CrossRef]
- Causton, H.C. SARS-CoV2 Infection and the Importance of Potassium Balance. Front. Med. 2021, 8, 744697. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Akinyemi, I.A.; Chitre, S.A.; Loeb, J.C.; Lednicky, J.A.; McIntosh, M.T.; Bhaduri-McIntosh, S. SARS-CoV-2 Viroporin Encoded by ORF3a Triggers the NLRP3 Inflammatory Pathway. Virology 2022, 568, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, K.; Goyal, P.; Grewer, C. Mechanism and Potential Sites of Potassium Interaction with Glutamate Transporters. J. Gen. Physiol. 2020, 152, e202012577. [Google Scholar] [CrossRef] [PubMed]
- Kozlov, A.G.; Cheng, X.; Zhang, H.; Shinn, M.K.; Weiland, E.; Nguyen, B.; Shkel, I.A.; Zytkiewicz, E.; Finkelstein, I.J.; Record, M.T., Jr.; et al. How Glutamate Promotes Liquid-Liquid Phase Separation and DNA Binding Cooperativity of E. Coli SSB Protein. J. Mol. Biol. 2022, 434, 167562. [Google Scholar] [CrossRef]
- Rimmele, T.S.; Rocher, A.-B.; Wellbourne-Wood, J.; Chatton, J.-Y. Control of Glutamate Transport by Extracellular Potassium: Basis for a Negative Feedback on Synaptic Transmission. Cereb. Cortex 2017, 27, 3272–3283. [Google Scholar] [CrossRef][Green Version]
- Bharadwaj, S.; Singh, M.; Kirtipal, N.; Kang, S.G. SARS-CoV-2 and Glutamine: SARS-CoV-2 Triggered Pathogenesis via Metabolic Reprograming of Glutamine in Host Cells. Front. Mol. Biosci 2020, 7, 627842. [Google Scholar] [CrossRef]
- Wang, J.; Yang, G.; Wang, X.; Wen, Z.; Shuai, L.; Luo, J.; Wang, C.; Sun, Z.; Liu, R.; Ge, J.; et al. SARS-CoV-2 Uses Metabotropic Glutamate Receptor Subtype 2 as an Internalization Factor to Infect Cells. Cell Discov. 2021, 7, 119. [Google Scholar] [CrossRef]
- Díaz-Resendiz, K.J.G.; Benitez-Trinidad, A.B.; Covantes-Rosales, C.E.; Toledo-Ibarra, G.A.; Ortiz-Lazareno, P.C.; Girón-Pérez, D.A.; Bueno-Durán, A.Y.; Pérez-Díaz, D.A.; Barcelos-García, R.G.; Girón-Pérez, M.I. Loss of Mitochondrial Membrane Potential (ΔΨm ) in Leucocytes as Post-COVID-19 Sequelae. J. Leukoc. Biol. 2022, 112, 23–29. [Google Scholar] [CrossRef]
- Wong, R.S.M.; Wu, A.; To, K.F.; Lee, N.; Lam, C.W.K.; Wong, C.K.; Chan, P.K.S.; Ng, M.H.L.; Yu, L.M.; Hui, D.S.; et al. Haematological Manifestations in Patients with Severe Acute Respiratory Syndrome: Retrospective Analysis. BMJ 2003, 326, 1358–1362. [Google Scholar] [CrossRef]
- Zou, Z.-Y.; Ren, D.; Chen, R.-L.; Yu, B.-J.; Liu, Y.; Huang, J.-J.; Yang, Z.-J.; Zhou, Z.-P.; Feng, Y.-W.; Wu, M. Persistent Lymphopenia after Diagnosis of COVID-19 Predicts Acute Respiratory Distress Syndrome: A Retrospective Cohort Study. Eur. J. Inflam. 2021, 19, 20587392211036825. [Google Scholar] [CrossRef]
- Ghizlane, E.A.; Manal, M.; Abderrahim, E.K.; Abdelilah, E.; Mohammed, M.; Rajae, A.; Amine, B.M.; Houssam, B.; Naima, A.; Brahim, H. Lymphopenia in Covid-19: A Single Center Retrospective Study of 589 Cases. Ann. Med. Surg. 2021, 69, 102816. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; Gao, Y.; Wang, G.; Song, G.; Liu, S.; Sun, D.; Xu, Y.; Tian, Z. Functional Exhaustion of Antiviral Lymphocytes in COVID-19 Patients. Cell. Mol. Immunol. 2020, 17, 533–535. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Meng, M.; Kumar, R.; Wu, Y.; Huang, J.; Deng, Y.; Weng, Z.; Yang, L. Lymphopenia Is Associated with Severe Coronavirus Disease 2019 (COVID-19) Infections: A Systemic Review and Meta-Analysis. Int. J. Infect. Dis. 2020, 96, 131–135. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Wang, Q.; Zhang, D.; Ding, J.; Huang, Q.; Tang, Y.-Q.; Wang, Q.; Miao, H. Lymphopenia Predicts Disease Severity of COVID-19: A Descriptive and Predictive Study. Signal Transduct. Target. Ther. 2020, 5, 33. [Google Scholar] [CrossRef]
- Liu, J.; Li, S.; Liu, J.; Liang, B.; Wang, X.; Wang, H.; Li, W.; Tong, Q.; Yi, J.; Zhao, L.; et al. Longitudinal Characteristics of Lymphocyte Responses and Cytokine Profiles in the Peripheral Blood of SARS-CoV-2 Infected Patients. EBioMedicine 2020, 55, 102763. [Google Scholar] [CrossRef]
- Ledderose, C.; Bao, Y.; Lidicky, M.; Zipperle, J.; Li, L.; Strasser, K.; Shapiro, N.I.; Junger, W.G. Mitochondria Are Gate-Keepers of T Cell Function by Producing the ATP That Drives Purinergic Signaling. J. Biol. Chem. 2014, 289, 25936–25945. [Google Scholar] [CrossRef]
- Desdín-Micó, G.; Soto-Heredero, G.; Mittelbrunn, M. Mitochondrial Activity in T Cells. Mitochondrion 2018, 41, 51–57. [Google Scholar] [CrossRef]
- Feske, S.; Giltnane, J.; Dolmetsch, R.; Staudt, L.M.; Rao, A. Gene Regulation Mediated by Calcium Signals in T Lymphocytes. Nat. Immunol. 2001, 2, 316–324. [Google Scholar] [CrossRef]
- Campello, S.; Lacalle, R.A.; Bettella, M.; Mañes, S.; Scorrano, L.; Viola, A. Orchestration of Lymphocyte Chemotaxis by Mitochondrial Dynamics. J. Exp. Med. 2006, 203, 2879–2886. [Google Scholar] [CrossRef]
- Zhang, K.; Li, H.; Song, Z. Membrane Depolarization Activates the Mitochondrial Protease OMA1 by Stimulating Self-Cleavage. EMBO Rep. 2014, 15, 576–585. [Google Scholar] [CrossRef]
- Jahangir, A.; Ozcan, C.; Holmuhamedov, E.L.; Terzic, A. Increased Calcium Vulnerability of Senescent Cardiac Mitochondria: Protective Role for a Mitochondrial Potassium Channel Opener. Mech. Ageing Dev. 2001, 122, 1073–1086. [Google Scholar] [CrossRef]
- Glitsch, M.D.; Bakowski, D.; Parekh, A.B. Store-Operated Ca2+ Entry Depends on Mitochondrial Ca2+ Uptake. EMBO J. 2002, 21, 6744–6754. [Google Scholar] [CrossRef]
- Santos, J.H.; Hunakova, L.U.; Chen, Y.; Bortner, C.; Van Houten, B. Cell Sorting Experiments Link Persistent Mitochondrial DNA Damage with Loss of Mitochondrial Membrane Potential and Apoptotic Cell Death. J. Biol. Chem. 2003, 278, 1728–1734. [Google Scholar] [CrossRef]
- Hu, L.; Zhang, S.; Wen, H.; Liu, T.; Cai, J.; Du, D.; Zhu, D.; Chen, F.; Xia, C. Melatonin Decreases M1 Polarization via Attenuating Mitochondrial Oxidative Damage Depending on UCP2 Pathway in Prorenin-Treated Microglia. PLoS ONE 2019, 14, e0212138. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.-J.; Ji, D.-M.; Liu, Z.-B.; Wang, T.-J.; Xie, F.-F.; Zhang, Z.-G.; Wei, Z.-L.; Zhou, P.; Cao, Y.-X. Melatonin Maintains Mitochondrial Membrane Potential and Decreases Excessive Intracellular Ca2+ Levels in Immature Human Oocytes. Life Sci. 2019, 235, 116810. [Google Scholar] [CrossRef] [PubMed]
- Lançoni, R.; Celeghini, E.C.C.; Alves, M.B.R.; Lemes, K.M.; Gonella-Diaza, A.M.; Oliveira, L.Z.; de Arruda, R.P. Melatonin Added to Cryopreservation Extenders Improves the Mitochondrial Membrane Potential of Postthawed Equine Sperm. J. Equine Vet. Sci. 2018, 69, 78–83. [Google Scholar] [CrossRef]
- Kumari, S.; Dash, D. Melatonin Elevates Intracellular Free Calcium in Human Platelets by Inositol 1,4,5-Trisphosphate Independent Mechanism. FEBS Lett. 2011, 585, 2345–2351. [Google Scholar] [CrossRef]
- Pieri, C.; Recchioni, R.; Moroni, F.; Marcheselli, F.; Marra, M.; Marinoni, S.; Di Primio, R. Melatonin Regulates the Respiratory Burst of Human Neutrophils and Their Depolarization. J. Pineal Res. 1998, 24, 43–49. [Google Scholar] [CrossRef]
- Fischer, T.W.; Zmijewski, M.A.; Wortsman, J.; Slominski, A. Melatonin Maintains Mitochondrial Membrane Potential and Attenuates Activation of Initiator (casp-9) and Effector Caspases (casp-3/casp-7) and PARP in UVR-Exposed HaCaT Keratinocytes. J. Pineal Res. 2008, 44, 397–407. [Google Scholar] [CrossRef]
- NavaneethaKrishnan, S.; Rosales, J.L.; Lee, K.-Y. mPTP Opening Caused by Cdk5 Loss Is due to Increased Mitochondrial Ca2+ Uptake. Oncogene 2020, 39, 2797–2806. [Google Scholar] [CrossRef]
- Park, J.; Lee, J.; Choi, C. Mitochondrial Network Determines Intracellular ROS Dynamics and Sensitivity to Oxidative Stress through Switching Inter-Mitochondrial Messengers. PLoS ONE 2011, 6, e23211. [Google Scholar] [CrossRef] [PubMed]
- Niki, E. Lipid Peroxidation: Physiological Levels and Dual Biological Effects. Free Radic. Biol. Med. 2009, 47, 469–484. [Google Scholar] [CrossRef] [PubMed]
- Ayala, A.; Muñoz, M.F.; Argüelles, S. Lipid Peroxidation: Production, Metabolism, and Signaling Mechanisms of Malondialdehyde and 4-Hydroxy-2-Nonenal. Oxid. Med. Cell. Longev. 2014, 2014, 360438. [Google Scholar] [CrossRef] [PubMed]
- Žarković, N.; Orehovec, B.; Milković, L.; Baršić, B.; Tatzber, F.; Wonisch, W.; Tarle, M.; Kmet, M.; Mataić, A.; Jakovčević, A.; et al. Preliminary Findings on the Association of the Lipid Peroxidation Product 4-Hydroxynonenal with the Lethal Outcome of Aggressive COVID-19. Antioxidants 2021, 10, 1341. [Google Scholar] [CrossRef] [PubMed]
- Reiter, R.J.; Tan, D.-X.; Galano, A. Melatonin Reduces Lipid Peroxidation and Membrane Viscosity. Front. Physiol. 2014, 5, 377. [Google Scholar] [CrossRef]
- García, J.J.; López-Pingarrón, L.; Almeida-Souza, P.; Tres, A.; Escudero, P.; García-Gil, F.A.; Tan, D.-X.; Reiter, R.J.; Ramírez, J.M.; Bernal-Pérez, M. Protective Effects of Melatonin in Reducing Oxidative Stress and in Preserving the Fluidity of Biological Membranes: A Review. J. Pineal Res. 2014, 56, 225–237. [Google Scholar] [CrossRef]
- Petrosillo, G.; Moro, N.; Ruggiero, F.M.; Paradies, G. Melatonin Inhibits Cardiolipin Peroxidation in Mitochondria and Prevents the Mitochondrial Permeability Transition and Cytochrome c Release. Free Radic. Biol. Med. 2009, 47, 969–974. [Google Scholar] [CrossRef]
- Livrea, M.A.; Tesoriere, L.; D’Arpa, D.; Morreale, M. Reaction of Melatonin with Lipoperoxyl Radicals in Phospholipid Bilayers. Free Radic. Biol. Med. 1997, 23, 706–711. [Google Scholar] [CrossRef]
- Galano, A.; Reiter, R.J. Melatonin and Its Metabolites vs Oxidative Stress: From Individual Actions to Collective Protection. J. Pineal Res. 2018, 65, e12514. [Google Scholar] [CrossRef]
- Tan, D.X.; Manchester, L.C.; Reiter, R.J.; Plummer, B.F.; Limson, J.; Weintraub, S.T.; Qi, W. Melatonin Directly Scavenges Hydrogen Peroxide: A Potentially New Metabolic Pathway of Melatonin Biotransformation. Free Radic. Biol. Med. 2000, 29, 1177–1185. [Google Scholar] [CrossRef]
- Fischer, T.W.; Scholz, G.; Knöll, B.; Hipler, U.C.; Elsner, P. Melatonin Reduces UV-Induced Reactive Oxygen Species in a Dose-Dependent Manner in IL-3-Stimulated Leukocytes. J. Pineal Res. 2001, 31, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Reiter, R.J. Melatonin: Lowering the High Price of Free Radicals. News Physiol. Sci. 2000, 15, 246–250. [Google Scholar] [CrossRef] [PubMed]
- Tan, D.-X.; Manchester, L.C.; Terron, M.P.; Flores, L.J.; Reiter, R.J. One Molecule, Many Derivatives: A Never-Ending Interaction of Melatonin with Reactive Oxygen and Nitrogen Species? J. Pineal Res. 2007, 42, 28–42. [Google Scholar] [CrossRef] [PubMed]
- Vanecek, J.; Klein, D.C. Sodium-Dependent Effects of Melatonin on Membrane Potential of Neonatal Rat Pituitary Cells. Endocrinology 1992, 131, 939–946. [Google Scholar] [CrossRef] [PubMed]
- Bortner, C.D.; Gomez-Angelats, M.; Cidlowski, J.A. Plasma Membrane Depolarization without Repolarization Is an Early Molecular Event in Anti-Fas-Induced Apoptosis. J. Biol. Chem. 2001, 276, 4304–4314. [Google Scholar] [CrossRef] [PubMed]
- Ching, A.C.; Hughes, M.R.; Poon, A.M.; Pang, S.F. Melatonin Receptors and Melatonin Inhibition of Duck Salt Gland Secretion. Gen. Comp. Endocrinol. 1999, 116, 229–240. [Google Scholar] [CrossRef]
- Hughes, M.R.; Kitamura, N.; Bennett, D.C.; Gray, D.A.; Sharp, P.J.; Poon, A.M.S. Effect of Melatonin on Salt Gland and Kidney Function of Gulls, Larus Glaucescens. Gen. Comp. Endocrinol. 2007, 151, 300–307. [Google Scholar] [CrossRef]
- Farouk, S.; Al-Huqail, A.A. Sustainable Biochar And/or Melatonin Improve Salinity Tolerance in Borage Plants by Modulating Osmotic Adjustment, Antioxidants, and Ion Homeostasis. Plants 2022, 11, 765. [Google Scholar] [CrossRef]
- Jiang, C.; Cui, Q.; Feng, K.; Xu, D.; Li, C.; Zheng, Q. Melatonin Improves Antioxidant Capacity and Ion Homeostasis and Enhances Salt Tolerance in Maize Seedlings. Acta Physiol. Plant 2016, 38, 82. [Google Scholar] [CrossRef]
- Li, C.; Wang, P.; Wei, Z.; Liang, D.; Liu, C.; Yin, L.; Jia, D.; Fu, M.; Ma, F. The Mitigation Effects of Exogenous Melatonin on Salinity-Induced Stress in Malus Hupehensis. J. Pineal Res. 2012, 53, 298–306. [Google Scholar] [CrossRef]
- Chakravarty, S.; Rizvi, S.I. Circadian Modulation of Sodium-Potassium ATPase and Sodium—Proton Exchanger in Human Erythrocytes: In Vitro Effect of Melatonin. Cell. Mol. Biol. 2011, 57, 80–86. [Google Scholar] [PubMed]
- Loh, D.; Reiter, R.J. Melatonin: Regulation of Prion Protein Phase Separation in Cancer Multidrug Resistance. Molecules 2022, 27, 705. [Google Scholar] [CrossRef] [PubMed]
- Morth, J.P.; Pedersen, B.P.; Buch-Pedersen, M.J.; Andersen, J.P.; Vilsen, B.; Palmgren, M.G.; Nissen, P. A Structural Overview of the Plasma Membrane Na+,K+-ATPase and H+-ATPase Ion Pumps. Nat. Rev. Mol. Cell Biol. 2011, 12, 60–70. [Google Scholar] [CrossRef] [PubMed]
- Noel, J.; Roux, D.; Pouysségur, J. Differential Localization of Na+/H+ Exchanger Isoforms (NHE1 and NHE3) in Polarized Epithelial Cell Lines. J. Cell Sci. 1996, 109 Pt 5, 929–939. [Google Scholar] [CrossRef] [PubMed]
- Clausen, M.V.; Hilbers, F.; Poulsen, H. The Structure and Function of the Na,K-ATPase Isoforms in Health and Disease. Front. Physiol. 2017, 8, 371. [Google Scholar] [CrossRef]
- Grinstein, S.; Rotin, D.; Mason, M.J. Na+/H+ Exchange and Growth Factor-Induced Cytosolic pH Changes. Role in Cellular Proliferation. Biochim. Biophys. Acta 1989, 988, 73–97. [Google Scholar] [CrossRef]
- Cha, C.Y.; Oka, C.; Earm, Y.E.; Wakabayashi, S.; Noma, A. A Model of Na+/H+ Exchanger and Its Central Role in Regulation of pH and Na+ in Cardiac Myocytes. Biophys. J. 2009, 97, 2674–2683. [Google Scholar] [CrossRef]
- Alberti, S.; Gladfelter, A.; Mittag, T. Considerations and Challenges in Studying Liquid-Liquid Phase Separation and Biomolecular Condensates. Cell 2019, 176, 419–434. [Google Scholar] [CrossRef]
- Królicka, A.L.; Kruczkowska, A.; Krajewska, M.; Kusztal, M.A. Hyponatremia in Infectious Diseases-A Literature Review. Int. J. Environ. Res. Public Health 2020, 17. [Google Scholar] [CrossRef]
- Machado, R.R.G.; Glaser, T.; Araujo, D.B.; Petiz, L.L.; Oliveira, D.B.L.; Durigon, G.S.; Leal, A.L.; Pinho, J.R.R.; Ferreira, L.C.S.; Ulrich, H.; et al. Inhibition of Severe Acute Respiratory Syndrome Coronavirus 2 Replication by Hypertonic Saline Solution in Lung and Kidney Epithelial Cells. ACS Pharmacol. Transl. Sci. 2021, 4, 1514–1527. [Google Scholar] [CrossRef]
- Kühlbrandt, W. Biology, Structure and Mechanism of P-Type ATPases. Nat. Rev. Mol. Cell Biol. 2004, 5, 282–295. [Google Scholar] [CrossRef] [PubMed]
- Dalskov, S.-M.; Immerdal, L.; Niels-Christiansen, L.-L.; Hansen, G.H.; Schousboe, A.; Danielsen, E.M. Lipid Raft Localization of GABA A Receptor and Na+, K+-ATPase in Discrete Microdomain Clusters in Rat Cerebellar Granule Cells. Neurochem. Int. 2005, 46, 489–499. [Google Scholar] [CrossRef] [PubMed]
- Welker, P.; Geist, B.; Frühauf, J.-H.; Salanova, M.; Groneberg, D.A.; Krause, E.; Bachmann, S. Role of Lipid Rafts in Membrane Delivery of Renal Epithelial Na+-K+-ATPase, Thick Ascending Limb. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2007, 292, R1328–R1337. [Google Scholar] [CrossRef] [PubMed]
- Fujii, T.; Takahashi, Y.; Itomi, Y.; Fujita, K.; Morii, M.; Tabuchi, Y.; Asano, S.; Tsukada, K.; Takeguchi, N.; Sakai, H. K+-Cl- Cotransporter-3a Up-Regulates Na+,K+-ATPase in Lipid Rafts of Gastric Luminal Parietal Cells. J. Biol. Chem. 2008, 283, 6869–6877. [Google Scholar] [CrossRef] [PubMed]
- Pytel, E.; Olszewska-Banaszczyk, M.; Koter-Michalak, M.; Broncel, M. Increased Oxidative Stress and Decreased Membrane Fluidity in Erythrocytes of CAD Patients. Biochem. Cell Biol. 2013, 91, 315–318. [Google Scholar] [CrossRef] [PubMed]
- Padmavathi, P.; Reddy, V.D.; Maturu, P.; Varadacharyulu, N. Smoking-Induced Alterations in Platelet Membrane Fluidity and Na(+)/K(+)-ATPase Activity in Chronic Cigarette Smokers. J. Atheroscler. Thromb. 2010, 17, 619–627. [Google Scholar] [CrossRef]
- Yelinova, V.; Glazachev, Y.; Khramtsov, V.; Kudryashova, L.; Rykova, V.; Salganik, R. Studies of Human and Rat Blood under Oxidative Stress: Changes in Plasma Thiol Level, Antioxidant Enzyme Activity, Protein Carbonyl Content, and Fluidity of Erythrocyte Membrane. Biochem. Biophys. Res. Commun. 1996, 221, 300–303. [Google Scholar] [CrossRef]
- Sutherland, E.; Dixon, B.S.; Leffert, H.L.; Skally, H.; Zaccaro, L.; Simon, F.R. Biochemical Localization of Hepatic Surface-Membrane Na+,K+-ATPase Activity Depends on Membrane Lipid Fluidity. Proc. Natl. Acad. Sci. USA 1988, 85, 8673–8677. [Google Scholar] [CrossRef]
- García, J.J.; Reiter, R.J.; Guerrero, J.M.; Escames, G.; Yu, B.P.; Oh, C.S.; Muñoz-Hoyos, A. Melatonin Prevents Changes in Microsomal Membrane Fluidity during Induced Lipid Peroxidation. FEBS Lett. 1997, 408, 297–300. [Google Scholar] [CrossRef]
- García, J.J.; Piñol-Ripoll, G.; Martínez-Ballarín, E.; Fuentes-Broto, L.; Miana-Mena, F.J.; Venegas, C.; Caballero, B.; Escames, G.; Coto-Montes, A.; Acuña-Castroviejo, D. Melatonin Reduces Membrane Rigidity and Oxidative Damage in the Brain of SAMP8 Mice. Neurobiol. Aging 2011, 32, 2045–2054. [Google Scholar] [CrossRef]
- Ochoa, J.J.; Vílchez, M.J.; Palacios, M.A.; García, J.J.; Reiter, R.J.; Muñoz-Hoyos, A. Melatonin Protects against Lipid Peroxidation and Membrane Rigidity in Erythrocytes from Patients Undergoing Cardiopulmonary Bypass Surgery. J. Pineal Res. 2003, 35, 104–108. [Google Scholar] [CrossRef]
- Bolmatov, D.; McClintic, W.T.; Taylor, G.; Stanley, C.B.; Do, C.; Collier, C.P.; Leonenko, Z.; Lavrentovich, M.O.; Katsaras, J. Deciphering Melatonin-Stabilized Phase Separation in Phospholipid Bilayers. Langmuir 2019, 35, 12236–12245. [Google Scholar] [CrossRef]
- Santamaria, A.; Batchu, K.C.; Matsarskaia, O.; Prévost, S.F.; Russo, D.; Natali, F.; Seydel, T.; Hoffmann, I.; Laux, V.; Haertlein, M.; et al. Strikingly Different Roles of SARS-CoV-2 Fusion Peptides Uncovered by Neutron Scattering. J. Am. Chem. Soc. 2022, 144, 2968–2979. [Google Scholar] [CrossRef]
- Deng, Y.; Angelova, A. Coronavirus-Induced Host Cubic Membranes and Lipid-Related Antiviral Therapies: A Focus on Bioactive Plasmalogens. Front. Cell Dev. Biol. 2021, 9, 630242. [Google Scholar] [CrossRef]
- Deng, Y.; Lee, E.L.-H.; Chong, K.; Almsherqi, Z.A. Evaluation of Radical Scavenging System in Amoeba Chaos Carolinense during Nutrient Deprivation. Interface Focus 2017, 7, 20160113. [Google Scholar] [CrossRef]
- Mannella, C.A. Structure and Dynamics of the Mitochondrial Inner Membrane Cristae. Biochim. Biophys. Acta 2006, 1763, 542–548. [Google Scholar] [CrossRef]
- Zheng, X.; Sun, Z.; Yu, L.; Shi, D.; Zhu, M.; Yao, H.; Li, L. Interactome Analysis of the Nucleocapsid Protein of SARS-CoV-2 Virus. Pathogens 2021, 10, 1155. [Google Scholar] [CrossRef]
- Yu, Q.; Guo, M.; Zeng, W.; Zeng, M.; Zhang, X.; Zhang, Y.; Zhang, W.; Jiang, X.; Yu, B. Interactions between NLRP3 Inflammasome and Glycolysis in Macrophages: New Insights into Chronic Inflammation Pathogenesis. Immun. Inflamm. Dis. 2022, 10, e581. [Google Scholar] [CrossRef]
- Ajaz, S.; McPhail, M.J.; Singh, K.K.; Mujib, S.; Trovato, F.M.; Napoli, S.; Agarwal, K. Mitochondrial Metabolic Manipulation by SARS-CoV-2 in Peripheral Blood Mononuclear Cells of Patients with COVID-19. Am. J. Physiol. Cell Physiol. 2021, 320, C57–C65. [Google Scholar] [CrossRef]
- Mookerjee, S.A.; Gerencser, A.A.; Nicholls, D.G.; Brand, M.D. Quantifying Intracellular Rates of Glycolytic and Oxidative ATP Production and Consumption Using Extracellular Flux Measurements. J. Biol. Chem. 2017, 292, 7189–7207. [Google Scholar] [CrossRef]
- Reiter, R.J.; Sharma, R.; Rosales-Corral, S. Anti-Warburg Effect of Melatonin: A Proposed Mechanism to Explain Its Inhibition of Multiple Diseases. Int. J. Mol. Sci. 2021, 22, 764. [Google Scholar] [CrossRef]
- Reiter, R.J.; Sharma, R.; Ma, Q.; Rosales-Corral, S.; Acuna-Castroviejo, D.; Escames, G. Inhibition of Mitochondrial Pyruvate Dehydrogenase Kinase: A Proposed Mechanism by Which Melatonin Causes Cancer Cells to Overcome Cytosolic Glycolysis, Reduce Tumor Biomass and Reverse Insensitivity to Chemotherapy. Melatonin Res. 2019, 2, 105–119. [Google Scholar] [CrossRef]
- Gray, L.R.; Tompkins, S.C.; Taylor, E.B. Regulation of Pyruvate Metabolism and Human Disease. Cell. Mol. Life Sci. 2014, 71, 2577–2604. [Google Scholar] [CrossRef]
- Chen, X.; Hao, B.; Li, D.; Reiter, R.J.; Bai, Y.; Abay, B.; Chen, G.; Lin, S.; Zheng, T.; Ren, Y.; et al. Melatonin Inhibits Lung Cancer Development by Reversing the Warburg Effect via Stimulating the SIRT3/PDH Axis. J. Pineal Res. 2021, e12755. [Google Scholar] [CrossRef]
- Go, G.; Yoon, Y.M.; Yoon, S.; Lee, G.; Lim, J.H.; Han, S.-Y.; Lee, S.H. Melatonin Protects Chronic Kidney Disease Mesenchymal Stem/stromal Cells against Accumulation of Methylglyoxal via Modulation of Hexokinase-2 Expression. Biomol. Ther. 2022, 30, 28. [Google Scholar] [CrossRef]
- Pérez-Torres, I.; Guarner-Lans, V.; Rubio-Ruiz, M.E. Reductive Stress in Inflammation-Associated Diseases and the Pro-Oxidant Effect of Antioxidant Agents. Int. J. Mol. Sci. 2017, 18, 2098. [Google Scholar] [CrossRef]
- Dawson, T.L.; Gores, G.J.; Nieminen, A.L.; Herman, B.; Lemasters, J.J. Mitochondria as a Source of Reactive Oxygen Species during Reductive Stress in Rat Hepatocytes. Am. J. Physiol. 1993, 264 Pt 1, C961–C967. [Google Scholar] [CrossRef]
- Hahn, A.; Parey, K.; Bublitz, M.; Mills, D.J.; Zickermann, V.; Vonck, J.; Kühlbrandt, W.; Meier, T. Structure of a Complete ATP Synthase Dimer Reveals the Molecular Basis of Inner Mitochondrial Membrane Morphology. Mol. Cell 2016, 63, 445–456. [Google Scholar] [CrossRef]
- Esparza-Perusquía, M.; Olvera-Sánchez, S.; Pardo, J.P.; Mendoza-Hernández, G.; Martínez, F.; Flores-Herrera, O. Structural and Kinetics Characterization of the F1F0-ATP Synthase Dimer. New Repercussion of Monomer-Monomer Contact. Biochim. Biophys. Acta Bioenerg. 2017, 1858, 975–981. [Google Scholar] [CrossRef]
- Davies, K.M.; Anselmi, C.; Wittig, I.; Faraldo-Gómez, J.D.; Kühlbrandt, W. Structure of the Yeast F1Fo-ATP Synthase Dimer and Its Role in Shaping the Mitochondrial Cristae. Proc. Natl. Acad. Sci. USA 2012, 109, 13602–13607. [Google Scholar] [CrossRef]
- Spikes, T.E.; Montgomery, M.G.; Walker, J.E. Interface Mobility between Monomers in Dimeric Bovine ATP Synthase Participates in the Ultrastructure of Inner Mitochondrial Membranes. Proc. Natl. Acad. Sci. USA 2021, 118. [Google Scholar] [CrossRef]
- Elías-Wolff, F.; Lindén, M.; Lyubartsev, A.P.; Brandt, E.G. Curvature Sensing by Cardiolipin in Simulated Buckled Membranes. Soft Matter 2019, 15, 792–802. [Google Scholar] [CrossRef]
- Ikon, N.; Ryan, R.O. Cardiolipin and Mitochondrial Cristae Organization. Biochim. Biophys. Acta Biomembr. 2017, 1859, 1156–1163. [Google Scholar] [CrossRef]
- Mileykovskaya, E.; Dowhan, W. Cardiolipin-Dependent Formation of Mitochondrial Respiratory Supercomplexes. Chem. Phys. Lipids 2014, 179, 42–48. [Google Scholar] [CrossRef]
- Pfeiffer, K.; Gohil, V.; Stuart, R.A.; Hunte, C.; Brandt, U.; Greenberg, M.L.; Schägger, H. Cardiolipin Stabilizes Respiratory Chain Supercomplexes. J. Biol. Chem. 2003, 278, 52873–52880. [Google Scholar] [CrossRef]
- Horvath, S.E.; Daum, G. Lipids of Mitochondria. Prog. Lipid Res. 2013, 52, 590–614. [Google Scholar] [CrossRef]
- Agrawal, A.; Ramachandran, R. Exploring the Links between Lipid Geometry and Mitochondrial Fission: Emerging Concepts. Mitochondrion 2019, 49, 305–313. [Google Scholar] [CrossRef]
- Parui, P.P.; Sarakar, Y.; Majumder, R.; Das, S.; Yang, H.; Yasuhara, K.; Hirota, S. Determination of Proton Concentration at Cardiolipin-Containing Membrane Interfaces and Its Relation with the Peroxidase Activity of Cytochrome C. Chem. Sci. 2019, 10, 9140–9151. [Google Scholar] [CrossRef]
- Haines, T.H.; Dencher, N.A. Cardiolipin: A Proton Trap for Oxidative Phosphorylation. FEBS Lett. 2002, 528, 35–39. [Google Scholar] [CrossRef]
- Afzal, N.; Lederer, W.J.; Jafri, M.S.; Mannella, C.A. Effect of Crista Morphology on Mitochondrial ATP Output: A Computational Study. Curr. Res. Physiol. 2021, 4, 163–176. [Google Scholar] [CrossRef]
- Vähäheikkilä, M.; Peltomaa, T.; Róg, T.; Vazdar, M.; Pöyry, S.; Vattulainen, I. How Cardiolipin Peroxidation Alters the Properties of the Inner Mitochondrial Membrane? Chem. Phys. Lipids 2018, 214, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Claypool, S.M. Cardiolipin, a Critical Determinant of Mitochondrial Carrier Protein Assembly and Function. Biochim. Biophys. Acta 2009, 1788, 2059–2068. [Google Scholar] [CrossRef]
- Paradies, G.; Paradies, V.; De Benedictis, V.; Ruggiero, F.M.; Petrosillo, G. Functional Role of Cardiolipin in Mitochondrial Bioenergetics. Biochim. Biophys. Acta 2014, 1837, 408–417. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y. Emerging Roles of Cardiolipin Remodeling in Mitochondrial Dysfunction Associated with Diabetes, Obesity, and Cardiovascular Diseases. J. Biomed. Res. 2010, 24, 6–15. [Google Scholar] [CrossRef]
- Chicco, A.J.; Sparagna, G.C. Role of Cardiolipin Alterations in Mitochondrial Dysfunction and Disease. Am. J. Physiol. Cell Physiol. 2007, 292, C33–C44. [Google Scholar] [CrossRef]
- Paradies, G.; Paradies, V.; Ruggiero, F.M.; Petrosillo, G. Mitochondrial Bioenergetics and Cardiolipin Alterations in Myocardial Ischemia-Reperfusion Injury: Implications for Pharmacological Cardioprotection. Am. J. Physiol. Heart Circ. Physiol. 2018, 315, H1341–H1352. [Google Scholar] [CrossRef]
- Dolinsky, V.W.; Cole, L.K.; Sparagna, G.C.; Hatch, G.M. Cardiac Mitochondrial Energy Metabolism in Heart Failure: Role of Cardiolipin and Sirtuins. Biochim. Biophys. Acta 2016, 1861, 1544–1554. [Google Scholar] [CrossRef]
- Han, X.; Yang, J.; Yang, K.; Zhao, Z.; Abendschein, D.R.; Gross, R.W. Alterations in Myocardial Cardiolipin Content and Composition Occur at the Very Earliest Stages of Diabetes: A Shotgun Lipidomics Study. Biochemistry 2007, 46, 6417–6428. [Google Scholar] [CrossRef]
- Schlame, M.; Ren, M. Barth Syndrome, a Human Disorder of Cardiolipin Metabolism. FEBS Lett. 2006, 580, 5450–5455. [Google Scholar] [CrossRef]
- Jiang, F.; Ryan, M.T.; Schlame, M.; Zhao, M.; Gu, Z.; Klingenberg, M.; Pfanner, N.; Greenberg, M.L. Absence of Cardiolipin in the crd1 Null Mutant Results in Decreased Mitochondrial Membrane Potential and Reduced Mitochondrial Function. J. Biol. Chem. 2000, 275, 22387–22394. [Google Scholar] [CrossRef]
- Ikonomidis, I.; Lekakis, J.; Vamvakou, G.; Loizou, S.; Revela, I.; Andreotti, F.; Kremastinos, D.T.; Nihoyannopoulos, P. IgA Anticardiolipin Antibody Is Associated with the Extent of Daily-Life Ischaemia in Patients with Chronic Coronary Artery Disease. Heart 2007, 93, 1412–1413. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Saleh, J.; Peyssonnaux, C.; Singh, K.K.; Edeas, M. Mitochondria and Microbiota Dysfunction in COVID-19 Pathogenesis. Mitochondrion 2020, 54, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Taha, M.; Samavati, L. Antiphospholipid Antibodies in COVID-19: A Meta-Analysis and Systematic Review. RMD Open 2021, 7, e001580. [Google Scholar] [CrossRef] [PubMed]
- Craig, W.Y.; Poulin, S.E.; Neveux, L.M.; Palomaki, G.E.; Dostal-Johnson, D.A.; Ledue, T.B.; Ritchie, R.F. Anti-Oxidized LDL Antibodies and Antiphospholipid Antibodies in Healthy Subjects: Relationship with Lipoprotein- and Oxidation-Related Analytes. J. Autoimmun. 1995, 8, 713–726. [Google Scholar] [CrossRef] [PubMed]
- Hasan Ali, O.; Bomze, D.; Risch, L.; Brugger, S.D.; Paprotny, M.; Weber, M.; Thiel, S.; Kern, L.; Albrich, W.C.; Kohler, P.; et al. Severe Coronavirus Disease 2019 (COVID-19) Is Associated With Elevated Serum Immunoglobulin (Ig) A and Antiphospholipid IgA Antibodies. Clin. Infect. Dis. 2021, 73, e2869–e2874. [Google Scholar] [CrossRef]
- Martín-Fernández, M.; Aller, R.; Heredia-Rodríguez, M.; Gómez-Sánchez, E.; Martínez-Paz, P.; Gonzalo-Benito, H.; Sánchez-de Prada, L.; Gorgojo, Ó.; Carnicero-Frutos, I.; Tamayo, E.; et al. Lipid Peroxidation as a Hallmark of Severity in COVID-19 Patients. Redox Biol. 2021, 48, 102181. [Google Scholar] [CrossRef]
- Petrosillo, G.; Di Venosa, N.; Pistolese, M.; Casanova, G.; Tiravanti, E.; Colantuono, G.; Federici, A.; Paradies, G.; Ruggiero, F.M. Protective Effect of Melatonin against Mitochondrial Dysfunction Associated with Cardiac Ischemia- Reperfusion: Role of Cardiolipin. FASEB J. 2006, 20, 269–276. [Google Scholar] [CrossRef]
- Römsing, S. Development and Validation of Bioanalytical Methods: Application to Melatonin and Selected Anti-Infective Drugs. Ph.D. Thesis, Acta Universitatis Upsaliensis, Uppsala, Sweden, 2010. [Google Scholar]
- Bongiorno, D.; Ceraulo, L.; Ferrugia, M.; Filizzola, F.; Giordano, C.; Ruggirello, A.; Liveri, V.T. H-NMR and FT-IR Study of the State of Melatonin Confined in Membrane Models: Location and Interactions of Melatonin in Water Free Lecithin and AOT Reversed Micelles. ARKIVOC 2004, 2004, 251–262. [Google Scholar] [CrossRef]
- Ceraulo, L.; Ferrugia, M.; Tesoriere, L.; Segreto, S.; Livrea, M.A.; Turco Liveri, V. Interactions of Melatonin with Membrane Models: Portioning of Melatonin in AOT and Lecithin Reversed Micelles. J. Pineal Res. 1999, 26, 108–112. [Google Scholar] [CrossRef]
- Galano, A.; Tan, D.X.; Reiter, R.J. Cyclic 3-Hydroxymelatonin, a Key Metabolite Enhancing the Peroxyl Radical Scavenging Activity of Melatonin. RSC Adv. 2014, 4, 5220. [Google Scholar] [CrossRef]
- Tan, D.X.; Manchester, L.C.; Reiter, R.J.; Plummer, B.F. Cyclic 3-Hydroxymelatonin: A Melatonin Metabolite Generated as a Result of Hydroxyl Radical Scavenging. Biol. Signals Recept. 1999, 8, 70–74. [Google Scholar] [CrossRef] [PubMed]
- Bielski, B.H.; Arudi, R.L.; Sutherland, M.W. A Study of the Reactivity of HO2/O2- with Unsaturated Fatty Acids. J. Biol. Chem. 1983, 258, 4759–4761. [Google Scholar] [CrossRef]
- Aikens, J.; Dix, T.A. Perhydroxyl Radical (HOO.) Initiated Lipid Peroxidation. The Role of Fatty Acid Hydroperoxides. J. Biol. Chem. 1991, 266, 15091–15098. [Google Scholar] [CrossRef]
- Ademowo, O.S.; Dias, H.K.I.; Burton, D.G.A.; Griffiths, H.R. Lipid (per) Oxidation in Mitochondria: An Emerging Target in the Ageing Process? Biogerontology 2017, 18, 859–879. [Google Scholar] [CrossRef] [PubMed]
- Repetto, M.; Semprine, J.; Boveris, A. Lipid Peroxidation: Chemical Mechanism, Biological Implications and Analytical Determination. In Lipid Peroxidation; Catala, A., Ed.; IntechOpen: Rijeka, Croatia, 2012. [Google Scholar] [CrossRef]
- Ito, M.; Yanagi, Y.; Ichinohe, T. Encephalomyocarditis Virus Viroporin 2B Activates NLRP3 Inflammasome. PLoS Pathog. 2012, 8, e1002857. [Google Scholar] [CrossRef] [PubMed]
- Toldo, S.; Bussani, R.; Nuzzi, V.; Bonaventura, A.; Mauro, A.G.; Cannatà, A.; Pillappa, R.; Sinagra, G.; Nana-Sinkam, P.; Sime, P.; et al. Inflammasome Formation in the Lungs of Patients with Fatal COVID-19. Inflamm. Res. 2021, 70, 7–10. [Google Scholar] [CrossRef]
- Arioz, B.I.; Tarakcioglu, E.; Olcum, M.; Genc, S. The Role of Melatonin on NLRP3 Inflammasome Activation in Diseases. Antioxidants 2021, 10, 1020. [Google Scholar] [CrossRef]
- Seoane, P.I.; Lee, B.; Hoyle, C.; Yu, S.; Lopez-Castejon, G.; Lowe, M.; Brough, D. The NLRP3-Inflammasome as a Sensor of Organelle Dysfunction. J. Cell Biol. 2020, 219, 12. [Google Scholar] [CrossRef]
- Samir, P.; Kanneganti, T.-D. DDX3X Sits at the Crossroads of Liquid-Liquid and Prionoid Phase Transitions Arbitrating Life and Death Cell Fate Decisions in Stressed Cells. DNA Cell Biol. 2020, 39, 1091–1095. [Google Scholar] [CrossRef]
- Franklin, B.S.; Bossaller, L.; De Nardo, D.; Ratter, J.M.; Stutz, A.; Engels, G.; Brenker, C.; Nordhoff, M.; Mirandola, S.R.; Al-Amoudi, A.; et al. The Adaptor ASC Has Extracellular and “Prionoid” Activities That Propagate Inflammation. Nat. Immunol. 2014, 15, 727–737. [Google Scholar] [CrossRef]
- Xia, S.; Chen, Z.; Shen, C.; Fu, T.-M. Higher-Order Assemblies in Immune Signaling: Supramolecular Complexes and Phase Separation. Protein Cell 2021, 12, 680–694. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Hara, H.; Núñez, G. Mechanism and Regulation of NLRP3 Inflammasome Activation. Trends Biochem. Sci. 2016, 41, 1012–1021. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Xu, W.; Zhou, R. NLRP3 Inflammasome Activation and Cell Death. Cell. Mol. Immunol. 2021, 18, 2114–2127. [Google Scholar] [CrossRef] [PubMed]
- Franchi, L.; Eigenbrod, T.; Muñoz-Planillo, R.; Nuñez, G. The Inflammasome: A Caspase-1-Activation Platform That Regulates Immune Responses and Disease Pathogenesis. Nat. Immunol. 2009, 10, 241–247. [Google Scholar] [CrossRef]
- Huang, Z.; Tyurina, Y.Y.; Jiang, J.; Tokarska-Schlattner, M.; Boissan, M.; Lacombe, M.-L.; Epand, R.; Schlattner, U.; Epand, R.M.; Kagan, V.E. Externalization of Cardiolipin as an “Eat-Me” Mitophageal Signal Is Facilitated by NDPK-D. Biophys. J. 2014, 106, 184a. [Google Scholar] [CrossRef]
- Elliott, E.I.; Miller, A.N.; Banoth, B.; Iyer, S.S.; Stotland, A.; Weiss, J.P.; Gottlieb, R.A.; Sutterwala, F.S.; Cassel, S.L. Cutting Edge: Mitochondrial Assembly of the NLRP3 Inflammasome Complex Is Initiated at Priming. J. Immunol. 2018, 200, 3047–3052. [Google Scholar] [CrossRef]
- Sefik, E.; Qu, R.; Junqueira, C.; Kaffe, E.; Mirza, H.; Zhao, J.; Brewer, J.R.; Han, A.; Steach, H.R.; Israelow, B.; et al. Inflammasome Activation in Infected Macrophages Drives COVID-19 Pathology. Nature 2022, 606, 585–593. [Google Scholar] [CrossRef]
- Yalcinkaya, M.; Liu, W.; Islam, M.N.; Kotini, A.G.; Gusarova, G.A.; Fidler, T.P.; Papapetrou, E.P.; Bhattacharya, J.; Wang, N.; Tall, A.R. Modulation of the NLRP3 Inflammasome by Sars-CoV-2 Envelope Protein. Sci. Rep. 2021, 11, 24432. [Google Scholar] [CrossRef]
- Freeman, T.L.; Swartz, T.H. Targeting the NLRP3 Inflammasome in Severe COVID-19. Front. Immunol. 2020, 11, 1518. [Google Scholar] [CrossRef]
- Zeng, J.; Xie, X.; Feng, X.-L.; Xu, L.; Han, J.-B.; Yu, D.; Zou, Q.-C.; Liu, Q.; Li, X.; Ma, G.; et al. Specific Inhibition of the NLRP3 Inflammasome Suppresses Immune Overactivation and Alleviates COVID-19 like Pathology in Mice. EBioMedicine 2022, 75, 103803. [Google Scholar] [CrossRef]
- Zhao, N.; Di, B.; Xu, L.-L. The NLRP3 Inflammasome and COVID-19: Activation, Pathogenesis and Therapeutic Strategies. Cytokine Growth Factor Rev. 2021, 61, 2–15. [Google Scholar] [CrossRef] [PubMed]
- Abais, J.M.; Xia, M.; Zhang, Y.; Boini, K.M.; Li, P.-L. Redox Regulation of NLRP3 Inflammasomes: ROS as Trigger or Effector? Antioxid. Redox Signal. 2015, 22, 1111–1129. [Google Scholar] [CrossRef] [PubMed]
- Mitroulis, I.; Skendros, P.; Ritis, K. Targeting IL-1beta in Disease; the Expanding Role of NLRP3 Inflammasome. Eur. J. Intern. Med. 2010, 21, 157–163. [Google Scholar] [CrossRef] [PubMed]
- Tőzsér, J.; Benkő, S. Natural Compounds as Regulators of NLRP3 Inflammasome-Mediated IL-1β Production. Mediat. Inflamm. 2016, 2016, 5460302. [Google Scholar] [CrossRef]
- Yang, D.; Elner, S.G.; Bian, Z.-M.; Till, G.O.; Petty, H.R.; Elner, V.M. Pro-Inflammatory Cytokines Increase Reactive Oxygen Species through Mitochondria and NADPH Oxidase in Cultured RPE Cells. Exp. Eye Res. 2007, 85, 462–472. [Google Scholar] [CrossRef]
- Yoo, H.G.; Shin, B.A.; Park, J.S.; Lee, K.H.; Chay, K.O.; Yang, S.Y.; Ahn, B.W.; Jung, Y.D. IL-1beta Induces MMP-9 via Reactive Oxygen Species and NF-kappaB in Murine Macrophage RAW 264.7 Cells. Biochem. Biophys. Res. Commun. 2002, 298, 251–256. [Google Scholar] [CrossRef]
- Ratajczak, M.Z.; Kucia, M. SARS-CoV-2 Infection and Overactivation of Nlrp3 Inflammasome as a Trigger of Cytokine “Storm” and Risk Factor for Damage of Hematopoietic Stem Cells. Leukemia 2020, 34, 1726–1729. [Google Scholar] [CrossRef]
- Somasekharan, S.P.; Gleave, M. SARS-CoV-2 Nucleocapsid Protein Interacts with Immunoregulators and Stress Granules and Phase Separates to Form Liquid Droplets. FEBS Lett. 2021, 595, 2872–2896. [Google Scholar] [CrossRef]
- Park, S.H.; Lee, S.G.; Kim, Y.; Song, K. Assignment of a Human Putative RNA Helicase Gene, DDX3, to Human X Chromosome Bands p11.3-->p11.23. Cytogenet. Cell Genet. 1998, 81, 178–179. [Google Scholar] [CrossRef]
- Vesuna, F.; Akhrymuk, I.; Smith, A.; Winnard, P.T.; Lin, S.-C.; Scharpf, R.; Kehn-Hall, K.; Raman, V. RK-33, a Small Molecule Inhibitor of Host RNA Helicase DDX3, Suppresses Multiple Variants of SARS-CoV-2. bioRxiv 2022. [Google Scholar] [CrossRef]
- Kumar, R.; Singh, N.; Abdin, M.Z.; Patel, A.H.; Medigeshi, G.R. Dengue Virus Capsid Interacts with DDX3X-A Potential Mechanism for Suppression of Antiviral Functions in Dengue Infection. Front. Cell. Infect. Microbiol. 2017, 7, 542. [Google Scholar] [CrossRef] [PubMed]
- Brai, A.; Riva, V.; Saladini, F.; Zamperini, C.; Trivisani, C.I.; Garbelli, A.; Pennisi, C.; Giannini, A.; Boccuto, A.; Bugli, F.; et al. DDX3X Inhibitors, an Effective Way to Overcome HIV-1 Resistance Targeting Host Proteins. Eur. J. Med. Chem. 2020, 200, 112319. [Google Scholar] [CrossRef] [PubMed]
- Yedavalli, V.S.R.K.; Neuveut, C.; Chi, Y.-H.; Kleiman, L.; Jeang, K.-T. Requirement of DDX3 DEAD Box RNA Helicase for HIV-1 Rev-RRE Export Function. Cell 2004, 119, 381–392. [Google Scholar] [CrossRef] [PubMed]
- Pène, V.; Li, Q.; Sodroski, C.; Hsu, C.-S.; Liang, T.J. Dynamic Interaction of Stress Granules, DDX3X, and IKK-α Mediates Multiple Functions in Hepatitis C Virus Infection. J. Virol. 2015, 89, 5462–5477. [Google Scholar] [CrossRef]
- Nelson, C.; Mrozowich, T.; Gemmill, D.L.; Park, S.M.; Patel, T.R. Human DDX3X Unwinds Japanese Encephalitis and Zika Viral 5′ Terminal Regions. Int. J. Mol. Sci. 2021, 22, 413. [Google Scholar] [CrossRef]
- Winnard, P.T., Jr.; Vesuna, F.; Raman, V. Targeting Host DEAD-Box RNA Helicase DDX3X for Treating Viral Infections. Antivir. Res. 2021, 185, 104994. [Google Scholar] [CrossRef]
- Saito, M.; Iestamantavicius, V.; Hess, D.; Matthias, P. Monitoring Acetylation of the RNA Helicase DDX3X, a Protein Critical for Formation of Stress Granules. In RNA Remodeling Proteins: Methods and Protocols; Boudvillain, M., Ed.; Springer: New York, NY, USA, 2021; pp. 217–234. [Google Scholar] [CrossRef]
- Samir, P.; Kesavardhana, S.; Patmore, D.M.; Gingras, S.; Malireddi, R.K.S.; Karki, R.; Guy, C.S.; Briard, B.; Place, D.E.; Bhattacharya, A.; et al. DDX3X Acts as a Live-or-Die Checkpoint in Stressed Cells by Regulating NLRP3 Inflammasome. Nature 2019, 573, 590–594. [Google Scholar] [CrossRef]
- Cui, B.C.; Sikirzhytski, V.; Aksenova, M.; Lucius, M.D.; Levon, G.H.; Mack, Z.T.; Pollack, C.; Odhiambo, D.; Broude, E.; Lizarraga, S.B.; et al. Pharmacological Inhibition of DEAD-Box RNA Helicase 3 Attenuates Stress Granule Assembly. Biochem. Pharmacol. 2020, 182, 114280. [Google Scholar] [CrossRef]
- Lage, S.L.; Amaral, E.P.; Hilligan, K.L.; Laidlaw, E.; Rupert, A.; Namasivayan, S.; Rocco, J.; Galindo, F.; Kellogg, A.; Kumar, P.; et al. Persistent Oxidative Stress and Inflammasome Activation in CD14highCD16- Monocytes From COVID-19 Patients. Front. Immunol. 2021, 12, 799558. [Google Scholar] [CrossRef]
- Mishra, S.R.; Mahapatra, K.K.; Behera, B.P.; Patra, S.; Bhol, C.S.; Panigrahi, D.P.; Praharaj, P.P.; Singh, A.; Patil, S.; Dhiman, R.; et al. Mitochondrial Dysfunction as a Driver of NLRP3 Inflammasome Activation and Its Modulation through Mitophagy for Potential Therapeutics. Int. J. Biochem. Cell Biol. 2021, 136, 106013. [Google Scholar] [CrossRef]
- Favero, G.; Franceschetti, L.; Bonomini, F.; Rodella, L.F.; Rezzani, R. Melatonin as an Anti-Inflammatory Agent Modulating Inflammasome Activation. Int. J. Endocrinol. 2017, 2017, 1835195. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, X.; Grailer, J.J.; Wang, N.; Wang, M.; Yao, J.; Zhong, R.; Gao, G.F.; Ward, P.A.; Tan, D.-X.; et al. Melatonin Alleviates Acute Lung Injury through Inhibiting the NLRP3 Inflammasome. J. Pineal Res. 2016, 60, 405–414. [Google Scholar] [CrossRef]
- Ma, S.; Chen, J.; Feng, J.; Zhang, R.; Fan, M.; Han, D.; Li, X.; Li, C.; Ren, J.; Wang, Y.; et al. Melatonin Ameliorates the Progression of Atherosclerosis via Mitophagy Activation and NLRP3 Inflammasome Inhibition. Oxid. Med. Cell. Longev. 2018, 2018, 9286458. [Google Scholar] [CrossRef] [PubMed]
- Squeglia, F.; Romano, M.; Ruggiero, A.; Maga, G.; Berisio, R. Host DDX Helicases as Possible SARS-CoV-2 Proviral Factors: A Structural Overview of Their Hijacking Through Multiple Viral Proteins. Front. Chem. 2020, 8, 602162. [Google Scholar] [CrossRef] [PubMed]
- Ciccosanti, F.; Di Rienzo, M.; Romagnoli, A.; Colavita, F.; Refolo, G.; Castilletti, C.; Agrati, C.; Brai, A.; Manetti, F.; Botta, L.; et al. Proteomic Analysis Identifies the RNA Helicase DDX3X as a Host Target against SARS-CoV-2 Infection. Antivir. Res. 2021, 190, 105064. [Google Scholar] [CrossRef]
- Hernández-Díaz, T.; Valiente-Echeverría, F.; Soto-Rifo, R. RNA Helicase DDX3: A Double-Edged Sword for Viral Replication and Immune Signaling. Microorganisms 2021, 9, 1206. [Google Scholar] [CrossRef] [PubMed]
- Valiente-Echeverría, F.; Hermoso, M.A.; Soto-Rifo, R. RNA Helicase DDX3: At the Crossroad of Viral Replication and Antiviral Immunity. Rev. Med. Virol. 2015, 25, 286–299. [Google Scholar] [CrossRef]
- Riva, V.; Maga, G. From the Magic Bullet to the Magic Target: Exploiting the Diverse Roles of DDX3X in Viral Infections and Tumorigenesis. Future Med. Chem. 2019, 11, 1357–1381. [Google Scholar] [CrossRef]
- Wang, W.; Jia, M.; Zhao, C.; Yu, Z.; Song, H.; Qin, Y.; Zhao, W. RNF39 Mediates K48-Linked Ubiquitination of DDX3X and Inhibits RLR-Dependent Antiviral Immunity. Sci. Adv. 2021, 7, eabe5877. [Google Scholar] [CrossRef]
- Soulat, D.; Bürckstümmer, T.; Westermayer, S.; Goncalves, A.; Bauch, A.; Stefanovic, A.; Hantschel, O.; Bennett, K.L.; Decker, T.; Superti-Furga, G. The DEAD-Box Helicase DDX3X Is a Critical Component of the TANK-Binding Kinase 1-Dependent Innate Immune Response. EMBO J. 2008, 27, 2135–2146. [Google Scholar] [CrossRef]
- Oshiumi, H.; Sakai, K.; Matsumoto, M.; Seya, T. DEAD/H BOX 3 (DDX3) Helicase Binds the RIG-I Adaptor IPS-1 to up-Regulate IFN-Beta-Inducing Potential. Eur. J. Immunol. 2010, 40, 940–948. [Google Scholar] [CrossRef] [PubMed]
- Oshiumi, H.; Kouwaki, T.; Seya, T. Accessory Factors of Cytoplasmic Viral RNA Sensors Required for Antiviral Innate Immune Response. Front. Immunol. 2016, 7, 200. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Dai, T.; Qin, Z.; Pan, T.; Chu, F.; Lou, L.; Zhang, L.; Yang, B.; Huang, H.; Lu, H.; et al. Targeting Liquid-Liquid Phase Separation of SARS-CoV-2 Nucleocapsid Protein Promotes Innate Antiviral Immunity by Elevating MAVS Activity. Nat. Cell Biol. 2021, 23, 718–732. [Google Scholar] [CrossRef]
- Hou, F.; Sun, L.; Zheng, H.; Skaug, B.; Jiang, Q.-X.; Chen, Z.J. MAVS Forms Functional Prion-like Aggregates to Activate and Propagate Antiviral Innate Immune Response. Cell 2011, 146, 448–461. [Google Scholar] [CrossRef]
- Protter, D.S.W.; Parker, R. Principles and Properties of Stress Granules. Trends Cell Biol. 2016, 26, 668–679. [Google Scholar] [CrossRef] [PubMed]
- Parker, F.; Maurier, F.; Delumeau, I.; Duchesne, M.; Faucher, D.; Debussche, L.; Dugue, A.; Schweighoffer, F.; Tocque, B. A Ras-GTPase-Activating Protein SH3-Domain-Binding Protein. Mol. Cell. Biol. 1996, 16, 2561–2569. [Google Scholar] [CrossRef] [PubMed]
- Yang, P.; Mathieu, C.; Kolaitis, R.-M.; Zhang, P.; Messing, J.; Yurtsever, U.; Yang, Z.; Wu, J.; Li, Y.; Pan, Q.; et al. G3BP1 Is a Tunable Switch That Triggers Phase Separation to Assemble Stress Granules. Cell 2020, 181, 325–345.e28. [Google Scholar] [CrossRef] [PubMed]
- Deater, M.; Tamhankar, M.; Lloyd, R.E. TDRD3 Is an Antiviral Restriction Factor That Promotes IFN Signaling with G3BP1. PLoS Pathog. 2022, 18, e1010249. [Google Scholar] [CrossRef]
- Yang, W.; Ru, Y.; Ren, J.; Bai, J.; Wei, J.; Fu, S.; Liu, X.; Li, D.; Zheng, H. G3BP1 Inhibits RNA Virus Replication by Positively Regulating RIG-I-Mediated Cellular Antiviral Response. Cell Death Dis. 2019, 10, 946. [Google Scholar] [CrossRef]
- Biswal, M.; Lu, J.; Song, J. SARS-CoV-2 Nucleocapsid Protein Targets a Conserved Surface Groove of the NTF2-like Domain of G3BP1. J. Mol. Biol. 2022, 434, 167516. [Google Scholar] [CrossRef]
- Wang, J.; Shi, C.; Xu, Q.; Yin, H. SARS-CoV-2 Nucleocapsid Protein Undergoes Liquid-Liquid Phase Separation into Stress Granules through Its N-Terminal Intrinsically Disordered Region. Cell Discov. 2021, 7, 5. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Deng, R.; Jiang, H.; Song, H.; Li, S.; Shen, Q.; Huang, W.; Nussinov, R.; Yu, J.; Zhang, J. The Mechanism of ATP-Dependent Allosteric Protection of Akt Kinase Phosphorylation. Structure 2015, 23, 1725–1734. [Google Scholar] [CrossRef] [PubMed]
- Carlson, C.R.; Asfaha, J.B.; Ghent, C.M.; Howard, C.J.; Hartooni, N.; Safari, M.; Frankel, A.D.; Morgan, D.O. Phosphoregulation of Phase Separation by the SARS-CoV-2 N Protein Suggests a Biophysical Basis for Its Dual Functions. Mol. Cell 2020, 80, 1092–1103.e4. [Google Scholar] [CrossRef] [PubMed]
- Lier, C.; Becker, S.; Biedenkopf, N. Dynamic Phosphorylation of Ebola Virus VP30 in NP-Induced Inclusion Bodies. Virology 2017, 512, 39–47. [Google Scholar] [CrossRef]
- Mühlberger, E.; Weik, M.; Volchkov, V.E.; Klenk, H.D.; Becker, S. Comparison of the Transcription and Replication Strategies of Marburg Virus and Ebola Virus by Using Artificial Replication Systems. J. Virol. 1999, 73, 2333–2342. [Google Scholar] [CrossRef]
- Nikolakaki, E.; Giannakouros, T. SR/RS Motifs as Critical Determinants of Coronavirus Life Cycle. Front. Mol. Biosci. 2020, 7, 219. [Google Scholar] [CrossRef]
- Zúñiga, S.; Cruz, J.L.G.; Sola, I.; Mateos-Gómez, P.A.; Palacio, L.; Enjuanes, L. Coronavirus Nucleocapsid Protein Facilitates Template Switching and Is Required for Efficient Transcription. J. Virol. 2010, 84, 2169–2175. [Google Scholar] [CrossRef]
- Kaidanovich-Beilin, O.; Woodgett, J.R. GSK-3: Functional Insights from Cell Biology and Animal Models. Front. Mol. Neurosci. 2011, 4, 40. [Google Scholar] [CrossRef]
- Hughes, K.; Nikolakaki, E.; Plyte, S.E.; Totty, N.F.; Woodgett, J.R. Modulation of the Glycogen Synthase Kinase-3 Family by Tyrosine Phosphorylation. EMBO J. 1993, 12, 803–808. [Google Scholar] [CrossRef]
- Lochhead, P.A.; Kinstrie, R.; Sibbet, G.; Rawjee, T.; Morrice, N.; Cleghon, V. A Chaperone-Dependent GSK3beta Transitional Intermediate Mediates Activation-Loop Autophosphorylation. Mol. Cell 2006, 24, 627–633. [Google Scholar] [CrossRef]
- Sutherland, C.; Cohen, P. The Alpha-Isoform of Glycogen Synthase Kinase-3 from Rabbit Skeletal Muscle Is Inactivated by p70 S6 Kinase or MAP Kinase-Activated Protein Kinase-1 in Vitro. FEBS Lett. 1994, 338, 37–42. [Google Scholar] [CrossRef]
- Sutherland, C.; Leighton, I.A.; Cohen, P. Inactivation of Glycogen Synthase Kinase-3 Beta by Phosphorylation: New Kinase Connections in Insulin and Growth-Factor Signalling. Biochem. J. 1993, 296 Pt 1, 15–19. [Google Scholar] [CrossRef] [PubMed]
- Sarhan, M.A.; Abdel-Hakeem, M.S.; Mason, A.L.; Tyrrell, D.L.; Houghton, M. Glycogen Synthase Kinase 3β Inhibitors Prevent Hepatitis C Virus Release/assembly through Perturbation of Lipid Metabolism. Sci. Rep. 2017, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Cuartas-López, A.M.; Gallego-Gómez, J.C. Glycogen Synthase Kinase 3ß Participates in Late Stages of Dengue Virus-2 Infection. Mem. Inst. Oswaldo Cruz 2020, 115, e190357. [Google Scholar] [CrossRef]
- Guendel, I.; Iordanskiy, S.; Van Duyne, R.; Kehn-Hall, K.; Saifuddin, M.; Das, R.; Jaworski, E.; Sampey, G.C.; Senina, S.; Shultz, L.; et al. Novel Neuroprotective GSK-3β Inhibitor Restricts Tat-Mediated HIV-1 Replication. J. Virol. 2014, 88, 1189–1208. [Google Scholar] [CrossRef]
- Marineau, A.; Khan, K.A.; Servant, M.J. Roles of GSK-3 and β-Catenin in Antiviral Innate Immune Sensing of Nucleic Acids. Cells 2020, 9, 897. [Google Scholar] [CrossRef]
- Alfhili, M.A.; Alsughayyir, J.; McCubrey, J.A.; Akula, S.M. GSK-3-Associated Signaling Is Crucial to Virus Infection of Cells. Biochim. Biophys. Acta Mol. Cell Res. 2020, 1867, 118767. [Google Scholar] [CrossRef]
- Yun, J.S.; Kim, N.H.; Song, H.; Cha, S.Y.; Hwang, K.H.; Lee, J.E.; Jeong, C.-H.; Song, S.H.; Kim, S.; Cho, E.S.; et al. Emergence of Glycogen Synthase Kinase-3 Interaction Domain Enhances Phosphorylation of SARS-CoV-2 Nucleocapsid Protein. bioRxiv 2022. [Google Scholar] [CrossRef]
- Kim, J.-M.; Rhee, J.E.; Yoo, M.; Kim, H.M.; Lee, N.-J.; Woo, S.H.; Jo, H.-J.; Kwon, D.; Lee, S.; Yoo, C.K.; et al. Increase in Viral Load in Patients With SARS-CoV-2 Delta Variant Infection in the Republic of Korea. Front. Microbiol. 2022, 13, 819745. [Google Scholar] [CrossRef]
- Butt, A.A.; Dargham, S.R.; Chemaitelly, H.; Al Khal, A.; Tang, P.; Hasan, M.R.; Coyle, P.V.; Thomas, A.G.; Borham, A.M.; Concepcion, E.G.; et al. Severity of Illness in Persons Infected With the SARS-CoV-2 Delta Variant vs Beta Variant in Qatar. JAMA Intern. Med. 2022, 182, 197–205. [Google Scholar] [CrossRef]
- Twohig, K.A.; Nyberg, T.; Zaidi, A.; Thelwall, S.; Sinnathamby, M.A.; Aliabadi, S.; Seaman, S.R.; Harris, R.J.; Hope, R.; Lopez-Bernal, J.; et al. Hospital Admission and Emergency Care Attendance Risk for SARS-CoV-2 Delta (B.1.617.2) Compared with Alpha (B.1.1.7) Variants of Concern: A Cohort Study. Lancet Infect. Dis. 2022, 22, 35–42. [Google Scholar] [CrossRef]
- Rudd, C.E. GSK-3 Inhibition as a Therapeutic Approach Against SARs CoV2: Dual Benefit of Inhibiting Viral Replication While Potentiating the Immune Response. Front. Immunol. 2020, 11, 1638. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Verma, A.; Garcia, G.; Ramage, H.; Myers, R.L.; Lucas, A.; Michaelson, J.J.; Coryell, W.; Kumar, A.; Charney, A.W.; et al. Targeting the Coronavirus Nucleocapsid Protein through GSK-3 Inhibition. Proc. Natl. Acad. Sci. USA 2021, 118, e2113401118. [Google Scholar] [CrossRef] [PubMed]
- Shapira, T.; Rens, C.; Pichler, V.; Rees, W.; Steiner, T.; Jean, F.; Winkler, D.; Sarai, I.; Pelech, S.; Av-Gay, Y. Inhibition of Glycogen Synthase Kinase-3-Beta (GSK3β) Blocks Nucleocapsid Phosphorylation and SARS-CoV-2 Replication. Res. Sq. 2022. [Google Scholar] [CrossRef]
- Jope, R.S.; Yuskaitis, C.J.; Beurel, E. Glycogen Synthase Kinase-3 (GSK3): Inflammation, Diseases, and Therapeutics. Neurochem. Res. 2007, 32, 577–595. [Google Scholar] [CrossRef]
- Dugo, L.; Collin, M.; Allen, D.A.; Patel, N.S.A.; Bauer, I.; Mervaala, E.M.A.; Louhelainen, M.; Foster, S.J.; Yaqoob, M.M.; Thiemermann, C. GSK-3beta Inhibitors Attenuate the Organ Injury/dysfunction Caused by Endotoxemia in the Rat. Crit. Care Med. 2005, 33, 1903–1912. [Google Scholar] [CrossRef]
- Hoeflich, K.P.; Luo, J.; Rubie, E.A.; Tsao, M.S.; Jin, O.; Woodgett, J.R. Requirement for Glycogen Synthase Kinase-3beta in Cell Survival and NF-kappaB Activation. Nature 2000, 406, 86–90. [Google Scholar] [CrossRef]
- Chen, X.; Liu, Y.; Zhu, J.; Lei, S.; Dong, Y.; Li, L.; Jiang, B.; Tan, L.; Wu, J.; Yu, S.; et al. GSK-3β Downregulates Nrf2 in Cultured Cortical Neurons and in a Rat Model of Cerebral Ischemia-Reperfusion. Sci. Rep. 2016, 6, 20196. [Google Scholar] [CrossRef]
- Lu, M.; Wang, P.; Qiao, Y.; Jiang, C.; Ge, Y.; Flickinger, B.; Malhotra, D.K.; Dworkin, L.D.; Liu, Z.; Gong, R. GSK3β-Mediated Keap1-Independent Regulation of Nrf2 Antioxidant Response: A Molecular Rheostat of Acute Kidney Injury to Chronic Kidney Disease Transition. Redox Biol. 2019, 26, 101275. [Google Scholar] [CrossRef]
- Culbreth, M.; Aschner, M. GSK-3β, a Double-Edged Sword in Nrf2 Regulation: Implications for Neurological Dysfunction and Disease. F1000Research 2018, 7, 1043. [Google Scholar] [CrossRef]
- Zhang, S.; Xin, F.; Zhang, X. The Compound Packaged in Virions Is the Key to Trigger Host Glycolysis Machinery for Virus Life Cycle in the Cytoplasm. iScience 2021, 24, 101915. [Google Scholar] [CrossRef] [PubMed]
- Hilliker, A. Analysis of RNA Helicases in P-Bodies and Stress Granules. Methods Enzymol. 2012, 511, 323–346. [Google Scholar] [CrossRef] [PubMed]
- de Nadal, E.; Ammerer, G.; Posas, F. Controlling Gene Expression in Response to Stress. Nat. Rev. Genet. 2011, 12, 833–845. [Google Scholar] [CrossRef] [PubMed]
- López-Maury, L.; Marguerat, S.; Bähler, J. Tuning Gene Expression to Changing Environments: From Rapid Responses to Evolutionary Adaptation. Nat. Rev. Genet. 2008, 9, 583–593. [Google Scholar] [CrossRef] [PubMed]
- Kedersha, N.; Anderson, P. Regulation of Translation by Stress Granules and Processing Bodies. Prog. Mol. Biol. Transl. Sci. 2009, 90, 155–185. [Google Scholar] [CrossRef] [PubMed]
- Montpetit, B.; Thomsen, N.D.; Helmke, K.J.; Seeliger, M.A.; Berger, J.M.; Weis, K. A Conserved Mechanism of DEAD-Box ATPase Activation by Nucleoporins and InsP6 in mRNA Export. Nature 2011, 472, 238–242. [Google Scholar] [CrossRef]
- Gray, S.; Cao, W.; Montpetit, B.; De La Cruz, E.M. The Nucleoporin Gle1 Activates DEAD-Box Protein 5 (Dbp5) by Promoting ATP Binding and Accelerating Rate Limiting Phosphate Release. Nucleic Acids Res. 2022, 50, 3998–4011. [Google Scholar] [CrossRef]
- Weirich, C.S.; Erzberger, J.P.; Flick, J.S.; Berger, J.M.; Thorner, J.; Weis, K. Activation of the DExD/H-Box Protein Dbp5 by the Nuclear-Pore Protein Gle1 and Its Coactivator InsP6 Is Required for mRNA Export. Nat. Cell Biol. 2006, 8, 668–676. [Google Scholar] [CrossRef]
- Aryanpur, P.P.; Regan, C.A.; Collins, J.M.; Mittelmeier, T.M.; Renner, D.M.; Vergara, A.M.; Brown, N.P.; Bolger, T.A. Gle1 Regulates RNA Binding of the DEAD-Box Helicase Ded1 in Its Complex Role in Translation Initiation. Mol. Cell. Biol. 2017, 37. [Google Scholar] [CrossRef]
- Glass, L.; Wente, S.R. Gle1 Mediates Stress Granule-Dependent Survival during Chemotoxic Stress. Adv. Biol. Regul. 2019, 71, 156–171. [Google Scholar] [CrossRef]
- Lin, D.H.; Correia, A.R.; Cai, S.W.; Huber, F.M.; Jette, C.A.; Hoelz, A. Structural and Functional Analysis of mRNA Export Regulation by the Nuclear Pore Complex. Nat. Commun. 2018, 9, 2319. [Google Scholar] [CrossRef] [PubMed]
- Aditi; Folkmann, A.W.; Wente, S.R. Cytoplasmic hGle1A Regulates Stress Granules by Modulation of Translation. Mol. Biol. Cell 2015, 26, 1476–1490. [Google Scholar] [CrossRef] [PubMed]
- Aditi; Mason, A.C.; Sharma, M.; Dawson, T.R.; Wente, S.R. MAPK- and Glycogen Synthase Kinase 3-Mediated Phosphorylation Regulates the DEAD-Box Protein Modulator Gle1 for Control of Stress Granule Dynamics. J. Biol. Chem. 2019, 294, 559–575. [Google Scholar] [CrossRef] [PubMed]
- Chang, L.; Karin, M. Mammalian MAP Kinase Signalling Cascades. Nature 2001, 410, 37–40. [Google Scholar] [CrossRef]
- Johnson, G.L.; Lapadat, R. Mitogen-Activated Protein Kinase Pathways Mediated by ERK, JNK, and p38 Protein Kinases. Science 2002, 298, 1911–1912. [Google Scholar] [CrossRef]
- Liu, L.; Xu, Y.; Reiter, R.J.; Pan, Y.; Chen, D.; Liu, Y.; Pu, X.; Jiang, L.; Li, Z. Inhibition of ERK1/2 Signaling Pathway Is Involved in Melatonin’s Antiproliferative Effect on Human MG-63 Osteosarcoma Cells. Cell. Physiol. Biochem. 2016, 39, 2297–2307. [Google Scholar] [CrossRef]
- Shin, I.-S.; Park, J.-W.; Shin, N.-R.; Jeon, C.-M.; Kwon, O.-K.; Lee, M.-Y.; Kim, H.-S.; Kim, J.-C.; Oh, S.-R.; Ahn, K.-S. Melatonin Inhibits MUC5AC Production via Suppression of MAPK Signaling in Human Airway Epithelial Cells. J. Pineal Res. 2014, 56, 398–407. [Google Scholar] [CrossRef]
- Esposito, E.; Genovese, T.; Caminiti, R.; Bramanti, P.; Meli, R.; Cuzzocrea, S. Melatonin Reduces Stress-Activated/mitogen-Activated Protein Kinases in Spinal Cord Injury. J. Pineal Res. 2009, 46, 79–86. [Google Scholar] [CrossRef]
- Das, R.; Balmik, A.A.; Chinnathambi, S. Melatonin Reduces GSK3β-Mediated Tau Phosphorylation, Enhances Nrf2 Nuclear Translocation and Anti-Inflammation. ASN Neuro 2020, 12, 1759091420981204. [Google Scholar] [CrossRef]
- Rhee, Y.-H.; Ahn, J.-C. Melatonin Attenuated Adipogenesis through Reduction of the CCAAT/enhancer Binding Protein Beta by Regulating the Glycogen Synthase 3 Beta in Human Mesenchymal Stem Cells. J. Physiol. Biochem. 2016, 72, 145–155. [Google Scholar] [CrossRef]
- Park, K.-H.; Kang, J.W.; Lee, E.-M.; Kim, J.S.; Rhee, Y.H.; Kim, M.; Jeong, S.J.; Park, Y.G.; Kim, S.H. Melatonin Promotes Osteoblastic Differentiation through the BMP/ERK/Wnt Signaling Pathways. J. Pineal Res. 2011, 51, 187–194. [Google Scholar] [CrossRef]
- Karim, R.; Tse, G.; Putti, T.; Scolyer, R.; Lee, S. The Significance of the Wnt Pathway in the Pathology of Human Cancers. Pathology 2004, 36, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.; Wu, X.; Wang, Y.; Ji, S.; Zhang, W.; Kang, J.; Li, J.; Fei, G. Melatonin Prevents LPS-Induced Epithelial-Mesenchymal Transition in Human Alveolar Epithelial Cells via the GSK-3β/Nrf2 Pathway. Biomed. Pharmacother. 2020, 132, 110827. [Google Scholar] [CrossRef] [PubMed]
- Hadj Ayed Tka, K.; Mahfoudh Boussaid, A.; Zaouali, M.A.; Kammoun, R.; Bejaoui, M.; Ghoul Mazgar, S.; Rosello Catafau, J.; Ben Abdennebi, H. Melatonin Modulates Endoplasmic Reticulum Stress and Akt/GSK3-Beta Signaling Pathway in a Rat Model of Renal Warm Ischemia Reperfusion. Anal. Cell. Pathol. 2015, 2015, 635172. [Google Scholar] [CrossRef] [PubMed]
- Cross, D.A.; Alessi, D.R.; Cohen, P.; Andjelkovich, M.; Hemmings, B.A. Inhibition of Glycogen Synthase Kinase-3 by Insulin Mediated by Protein Kinase B. Nature 1995, 378, 785–789. [Google Scholar] [CrossRef]
- Beitner-Johnson, D.; Rust, R.T.; Hsieh, T.C.; Millhorn, D.E. Hypoxia Activates Akt and Induces Phosphorylation of GSK-3 in PC12 Cells. Cell. Signal. 2001, 13, 23–27. [Google Scholar] [CrossRef]
- Perdomo, J.; Quintana, C.; González, I.; Hernández, I.; Rubio, S.; Loro, J.F.; Reiter, R.J.; Estévez, F.; Quintana, J. Melatonin Induces Melanogenesis in Human SK-MEL-1 Melanoma Cells Involving Glycogen Synthase Kinase-3 and Reactive Oxygen Species. Int. J. Mol. Sci. 2020, 21, 4970. [Google Scholar] [CrossRef]
- Sainz, R.M.; Mayo, J.C.; Rodriguez, C.; Tan, D.X.; Lopez-Burillo, S.; Reiter, R.J. Melatonin and Cell Death: Differential Actions on Apoptosis in Normal and Cancer Cells. Cell. Mol. Life Sci. 2003, 60, 1407–1426. [Google Scholar] [CrossRef]
- Reiter, R.J.; Tan, D.-X.; Fuentes-Broto, L. Melatonin: A Multitasking Molecule. Prog. Brain Res. 2010, 181, 127–151. [Google Scholar] [CrossRef]
- Zhang, H.-M.; Zhang, Y. Melatonin: A Well-Documented Antioxidant with Conditional pro-Oxidant Actions. J. Pineal Res. 2014, 57, 131–146. [Google Scholar] [CrossRef]
- Sagrillo-Fagundes, L.; Bienvenue-Pariseault, J.; Vaillancourt, C. Melatonin: The Smart Molecule That Differentially Modulates Autophagy in Tumor and Normal Placental Cells. PLoS ONE 2019, 14, e0202458. [Google Scholar] [CrossRef] [PubMed]
- Asadi, M.R.; Moslehian, M.S.; Sabaie, H.; Poornabi, M.; Ghasemi, E.; Hassani, M.; Hussen, B.M.; Taheri, M.; Rezazadeh, M. Stress Granules in the Anti-Cancer Medications Mechanism of Action: A Systematic Scoping Review. Front. Oncol. 2021, 11, 797549. [Google Scholar] [CrossRef] [PubMed]
- Lavalée, M.; Curdy, N.; Laurent, C.; Fournié, J.-J.; Franchini, D.-M. Cancer Cell Adaptability: Turning Ribonucleoprotein Granules into Targets. Trends Cancer Res. 2021, 7, 902–915. [Google Scholar] [CrossRef] [PubMed]
- Khong, A.; Ripin, N.; de Vasconcelos, L.M.; Spencer, S.; Parker, R. Stress Granules Promote Chemoresistance by Triggering Cellular Quiescence. bioRxiv 2022. [Google Scholar] [CrossRef]
- Fu, Y.; Dominissini, D.; Rechavi, G.; He, C. Gene Expression Regulation Mediated through Reversible m6A RNA Methylation. Nat. Rev. Genet. 2014, 15, 293–306. [Google Scholar] [CrossRef]
- Kan, R.L.; Chen, J.; Sallam, T. Crosstalk between Epitranscriptomic and Epigenetic Mechanisms in Gene Regulation. Trends Genet. 2022, 38, 182–193. [Google Scholar] [CrossRef]
- Wilkinson, E.; Cui, Y.-H.; He, Y.-Y. Context-Dependent Roles of RNA Modifications in Stress Responses and Diseases. Int. J. Mol. Sci. 2021, 22, 1949. [Google Scholar] [CrossRef]
- Lai, L.-C.; Kissinger, M.T.; Burke, P.V.; Kwast, K.E. Comparison of the Transcriptomic “Stress Response” Evoked by Antimycin A and Oxygen Deprivation in Saccharomyces Cerevisiae. BMC Genom. 2008, 9, 627. [Google Scholar] [CrossRef]
- Paramasivam, A.; Priyadharsini, J.V. Epigenetic Modifications of RNA and Their Implications in Antiviral Immunity. Epigenomics 2020, 12, 1673–1675. [Google Scholar] [CrossRef]
- Viswanathan, T.; Arya, S.; Chan, S.-H.; Qi, S.; Dai, N.; Misra, A.; Park, J.-G.; Oladunni, F.; Kovalskyy, D.; Hromas, R.A.; et al. Structural Basis of RNA Cap Modification by SARS-CoV-2. Nat. Commun. 2020, 11, 3718. [Google Scholar] [CrossRef]
- Manne, B.K.; Denorme, F.; Middleton, E.A.; Portier, I.; Rowley, J.W.; Stubben, C.; Petrey, A.C.; Tolley, N.D.; Guo, L.; Cody, M.; et al. Platelet Gene Expression and Function in Patients with COVID-19. Blood 2020, 136, 1317–1329. [Google Scholar] [CrossRef] [PubMed]
- Finlay, J.B.; Brann, D.H.; Hachem, R.A.; Jang, D.W.; Oliva, A.D.; Ko, T.; Gupta, R.; Wellford, S.A.; Ashley Moseman, E.; Jang, S.S.; et al. Persistent Post-COVID-19 Smell Loss Is Associated with Inflammatory Infiltration and Altered Olfactory Epithelial Gene Expression. bioRxiv 2022. [Google Scholar] [CrossRef]
- Bernardes, J.P.; Mishra, N.; Tran, F.; Bahmer, T.; Best, L.; Blase, J.I.; Bordoni, D.; Franzenburg, J.; Geisen, U.; Josephs-Spaulding, J.; et al. Longitudinal Multi-Omics Analyses Identify Responses of Megakaryocytes, Erythroid Cells, and Plasmablasts as Hallmarks of Severe COVID-19. Immunity 2020, 53, 1296–1314.e9. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Liu, X.-Z.; Tian, Q.; Fan, Y.-X.; Ye, Z.; Meng, T.-Q.; Wei, G.-H.; Xiong, C.-L.; Li, H.; He, X.; et al. Transcriptome and DNA Methylome Analysis of Peripheral Blood Samples Reveals Incomplete Restoration and Transposable Element Activation after 3-Month Recovery of COVID-19. medRxiv 2022. [Google Scholar] [CrossRef]
- Zhou, Y.; Yang, L.; Wang, H.; Chen, X.; Jiang, W.; Wang, Z.; Liu, S.; Liu, Y. Alterations in DNA Methylation Profiles in Cancellous Bone of Postmenopausal Women with Osteoporosis. FEBS Open Bio 2020, 10, 1516–1531. [Google Scholar] [CrossRef]
- Salvio, G.; Gianfelice, C.; Firmani, F.; Lunetti, S.; Balercia, G.; Giacchetti, G. Bone Metabolism in SARS-CoV-2 Disease: Possible Osteoimmunology and Gender Implications. Clin. Rev. Bone Miner. Metab. 2020, 18, 51–57. [Google Scholar] [CrossRef]
- Khosla, S. Minireview: The OPG/RANKL/RANK System. Endocrinology 2001, 142, 5050–5055. [Google Scholar] [CrossRef]
- Qiao, W.; Lau, H.E.; Xie, H.; Poon, V.K.-M.; Chan, C.C.-S.; Chu, H.; Yuan, S.; Yuen, T.T.-T.; Chik, K.K.-H.; Tsang, J.O.-L.; et al. SARS-CoV-2 Infection Induces Inflammatory Bone Loss in Golden Syrian Hamsters. Nat. Commun. 2022, 13, 1–16. [Google Scholar] [CrossRef]
- Mick, E.; Kamm, J.; Pisco, A.O.; Ratnasiri, K.; Babik, J.M.; Castañeda, G.; DeRisi, J.L.; Detweiler, A.M.; Hao, S.L.; Kangelaris, K.N.; et al. Upper Airway Gene Expression Reveals Suppressed Immune Responses to SARS-CoV-2 Compared with Other Respiratory Viruses. Nat. Commun. 2020, 11, 5854. [Google Scholar] [CrossRef]
- Mast, A.E.; Wolberg, A.S.; Gailani, D.; Garvin, M.R.; Alvarez, C.; Miller, J.I.; Aronow, B.; Jacobson, D. SARS-CoV-2 Suppresses Anticoagulant and Fibrinolytic Gene Expression in the Lung. Elife 2021, 10, e64330. [Google Scholar] [CrossRef]
- Salgado-Albarrán, M.; Navarro-Delgado, E.I.; Del Moral-Morales, A.; Alcaraz, N.; Baumbach, J.; González-Barrios, R.; Soto-Reyes, E. Comparative Transcriptome Analysis Reveals Key Epigenetic Targets in SARS-CoV-2 Infection. NPJ Syst. Biol. Appl. 2021, 7, 21. [Google Scholar] [CrossRef] [PubMed]
- Criscione, S.W.; Theodosakis, N.; Micevic, G.; Cornish, T.C.; Burns, K.H.; Neretti, N.; Rodić, N. Genome-Wide Characterization of Human L1 Antisense Promoter-Driven Transcripts. BMC Genom. 2016, 17, 463. [Google Scholar] [CrossRef] [PubMed]
- McKerrow, W.; Wang, X.; Mendez-Dorantes, C.; Mita, P.; Cao, S.; Grivainis, M.; Ding, L.; LaCava, J.; Burns, K.H.; Boeke, J.D.; et al. LINE-1 Expression in Cancer Correlates with p53 Mutation, Copy Number Alteration, and S Phase Checkpoint. Proc. Natl. Acad. Sci. USA 2022, 119, e2115999119. [Google Scholar] [CrossRef] [PubMed]
- Honda, T.; Nishikawa, Y.; Nishimura, K.; Teng, D.; Takemoto, K.; Ueda, K. Effects of Activation of the LINE-1 Antisense Promoter on the Growth of Cultured Cells. Sci. Rep. 2020, 10, 22136. [Google Scholar] [CrossRef]
- Hancks, D.C.; Kazazian, H.H., Jr. Roles for Retrotransposon Insertions in Human Disease. Mob. DNA 2016, 7, 9. [Google Scholar] [CrossRef]
- Beck, C.R.; Garcia-Perez, J.L.; Badge, R.M.; Moran, J.V. LINE-1 Elements in Structural Variation and Disease. Annu. Rev. Genom. Hum. Genet. 2011, 12, 187–215. [Google Scholar] [CrossRef] [PubMed]
- Martinez, J.; Simon, M.; Seluanov, A.; Gorbunova, V. 415 Line1 Derepression in Specific Retrotransposon Families in Aged Mice Leads to Cytosolic DNA and Increased Inflammation. J. Clin. Transl. Sci. 2022, 6, 81. [Google Scholar] [CrossRef]
- Chen, J.-M.; Stenson, P.D.; Cooper, D.N.; Férec, C. A Systematic Analysis of LINE-1 Endonuclease-Dependent Retrotranspositional Events Causing Human Genetic Disease. Hum. Genet. 2005, 117, 411–427. [Google Scholar] [CrossRef]
- Terry, D.M.; Devine, S.E. Aberrantly High Levels of Somatic LINE-1 Expression and Retrotransposition in Human Neurological Disorders. Front. Genet. 2019, 10, 1244. [Google Scholar] [CrossRef]
- Boeke, J.D.; Stoye, J.P. Retrotransposons, Endogenous Retroviruses, and the Evolution of Retroelements. In Retroviruses; Coffin, J.M., Hughes, S.H., Varmus, H.E., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2011. [Google Scholar]
- McClintock, B. Chromosome Organization and Genic Expression. Cold Spring Harb. Symp. Quant. Biol. 1951, 16, 13–47. [Google Scholar] [CrossRef]
- Ravindran, S. Barbara McClintock and the Discovery of Jumping Genes. Proc. Natl. Acad. Sci. USA 2012, 109, 20198–20199. [Google Scholar] [CrossRef] [PubMed]
- Penzkofer, T.; Jäger, M.; Figlerowicz, M.; Badge, R.; Mundlos, S.; Robinson, P.N.; Zemojtel, T. L1Base 2: More Retrotransposition-Active LINE-1s, More Mammalian Genomes. Nucleic Acids Res. 2017, 45, D68–D73. [Google Scholar] [CrossRef] [PubMed]
- Kemp, J.R.; Longworth, M.S. Crossing the LINE Toward Genomic Instability: LINE-1 Retrotransposition in Cancer. Front. Chem. 2015, 3, 68. [Google Scholar] [CrossRef] [PubMed]
- Daskalos, A.; Nikolaidis, G.; Xinarianos, G.; Savvari, P.; Cassidy, A.; Zakopoulou, R.; Kotsinas, A.; Gorgoulis, V.; Field, J.K.; Liloglou, T. Hypomethylation of Retrotransposable Elements Correlates with Genomic Instability in Non-Small Cell Lung Cancer. Int. J. Cancer 2009, 124, 81–87. [Google Scholar] [CrossRef]
- Gasior, S.L.; Wakeman, T.P.; Xu, B.; Deininger, P.L. The Human LINE-1 Retrotransposon Creates DNA Double-Strand Breaks. J. Mol. Biol. 2006, 357, 1383–1393. [Google Scholar] [CrossRef]
- Rodić, N.; Burns, K.H. Long Interspersed Element–1 (LINE-1): Passenger or Driver in Human Neoplasms? PLoS Genet. 2013, 9, e1003402. [Google Scholar] [CrossRef]
- Rangwala, S.H.; Zhang, L.; Kazazian, H.H., Jr. Many LINE1 Elements Contribute to the Transcriptome of Human Somatic Cells. Genome Biol. 2009, 10, R100. [Google Scholar] [CrossRef]
- Kubo, S.; Seleme, M.D.C.; Soifer, H.S.; Perez, J.L.G.; Moran, J.V.; Kazazian, H.H., Jr.; Kasahara, N. L1 Retrotransposition in Nondividing and Primary Human Somatic Cells. Proc. Natl. Acad. Sci. USA 2006, 103, 8036–8041. [Google Scholar] [CrossRef]
- Ardeljan, D.; Taylor, M.S.; Ting, D.T.; Burns, K.H. The Human Long Interspersed Element-1 Retrotransposon: An Emerging Biomarker of Neoplasia. Clin. Chem. 2017, 63, 816–822. [Google Scholar] [CrossRef]
- Swergold, G.D. Identification, Characterization, and Cell Specificity of a Human LINE-1 Promoter. Mol. Cell. Biol. 1990, 10, 6718–6729. [Google Scholar] [CrossRef]
- Kolosha, V.O.; Martin, S.L. High-Affinity, Non-Sequence-Specific RNA Binding by the Open Reading Frame 1 (ORF1) Protein from Long Interspersed Nuclear Element 1 (LINE-1). J. Biol. Chem. 2003, 278, 8112–8117. [Google Scholar] [CrossRef] [PubMed]
- Piskareva, O.; Ernst, C.; Higgins, N.; Schmatchenko, V. The Carboxy-Terminal Segment of the Human LINE-1 ORF2 Protein Is Involved in RNA Binding. FEBS Open Bio. 2013, 3, 433–437. [Google Scholar] [CrossRef] [PubMed]
- Sciamanna, I.; De Luca, C.; Spadafora, C. The Reverse Transcriptase Encoded by LINE-1 Retrotransposons in the Genesis, Progression, and Therapy of Cancer. Front. Chem. 2016, 4, 6. [Google Scholar] [CrossRef] [PubMed]
- Han, J.S.; Szak, S.T.; Boeke, J.D. Transcriptional Disruption by the L1 Retrotransposon and Implications for Mammalian Transcriptomes. Nature 2004, 429, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Poletti, V.; Mavilio, F. Interactions between Retroviruses and the Host Cell Genome. Mol. Ther. Methods Clin. Dev. 2018, 8, 31–41. [Google Scholar] [CrossRef] [PubMed]
- Geuking, M.B.; Weber, J.; Dewannieux, M.; Gorelik, E.; Heidmann, T.; Hengartner, H.; Zinkernagel, R.M.; Hangartner, L. Recombination of Retrotransposon and Exogenous RNA Virus Results in Nonretroviral cDNA Integration. Science 2009, 323, 393–396. [Google Scholar] [CrossRef]
- Yin, Y.; Liu, X.-Z.; He, X.; Zhou, L.-Q. Exogenous Coronavirus Interacts With Endogenous Retrotransposon in Human Cells. Front. Cell. Infect. Microbiol. 2021, 11, 609160. [Google Scholar] [CrossRef]
- Zhang, L.; Richards, A.; Barrasa, M.I.; Hughes, S.H.; Young, R.A.; Jaenisch, R. Reverse-Transcribed SARS-CoV-2 RNA Can Integrate into the Genome of Cultured Human Cells and Can Be Expressed in Patient-Derived Tissues. Proc. Natl. Acad. Sci. USA 2021, 118, e2105968118. [Google Scholar] [CrossRef]
- Kazachenka, A.; Kassiotis, G. SARS-CoV-2-Host Chimeric RNA-Sequencing Reads Do Not Necessarily Arise from Virus Integration into the Host DNA. Front. Microbiol. 2021, 12, 676693. [Google Scholar] [CrossRef]
- Yan, B.; Chakravorty, S.; Mirabelli, C.; Wang, L.; Trujillo-Ochoa, J.L.; Chauss, D.; Kumar, D.; Lionakis, M.S.; Olson, M.R.; Wobus, C.E.; et al. Host-Virus Chimeric Events in SARS-CoV-2-Infected Cells Are Infrequent and Artifactual. J. Virol. 2021, 95, e0029421. [Google Scholar] [CrossRef]
- Grandi, N.; Tramontano, E.; Berkhout, B. Integration of SARS-CoV-2 RNA in Infected Human Cells by Retrotransposons: An Unlikely Hypothesis and Old Viral Relationships. Retrovirology 2021, 18, 34. [Google Scholar] [CrossRef] [PubMed]
- Hirschenberger, M.; Hunszinger, V.; Sparrer, K.M.J. Implications of Innate Immunity in Post-Acute Sequelae of Non-Persistent Viral Infections. Cells 2021, 10, 2134. [Google Scholar] [CrossRef] [PubMed]
- Ryan, F.J.; Hope, C.M.; Masavuli, M.G.; Lynn, M.A.; Mekonnen, Z.A.; Yeow, A.E.L.; Garcia-Valtanen, P.; Al-Delfi, Z.; Gummow, J.; Ferguson, C.; et al. Long-Term Perturbation of the Peripheral Immune System Months after SARS-CoV-2 Infection. BMC Med. 2022, 20, 26. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, R.; Yu, J. New Understanding of the Relevant Role of LINE-1 Retrotransposition in Human Disease and Immune Modulation. Front. Cell Dev. Biol. 2020, 8, 657. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Li, W.; Wang, T.; Ran, D.; Davalos, V.; Planas-Serra, L.; Pujol, A.; Esteller, M.; Wang, X.; Yu, H. Accelerated Biological Aging in COVID-19 Patients. Nat. Commun. 2022, 13, 1–7. [Google Scholar] [CrossRef]
- Wander, P.L.; Lowy, E.; Beste, L.A.; Tulloch-Palomino, L.; Korpak, A.; Peterson, A.C.; Kahn, S.E.; Boyko, E.J. The Incidence of Diabetes Among 2,777,768 Veterans With and Without Recent SARS-CoV-2 Infection. Diabetes Care 2022, 45, 782–788. [Google Scholar] [CrossRef] [PubMed]
- Rubino, F.; Amiel, S.A.; Zimmet, P.; Alberti, G.; Bornstein, S.; Eckel, R.H.; Mingrone, G.; Boehm, B.; Cooper, M.E.; Chai, Z.; et al. New-Onset Diabetes in COVID-19. N. Engl. J. Med. 2020, 383, 789–790. [Google Scholar] [CrossRef]
- Barrett, C.E.; Koyama, A.K.; Alvarez, P.; Chow, W.; Lundeen, E.A.; Perrine, C.G.; Pavkov, M.E.; Rolka, D.B.; Wiltz, J.L.; Bull-Otterson, L.; et al. Risk for Newly Diagnosed Diabetes >30 Days After SARS-CoV-2 Infection Among Persons Aged <18 Years-United States, March 1, 2020-June 28, 2021. MMWR Morb. Mortal. Wkly. Rep. 2022, 71, 59–65. [Google Scholar] [CrossRef]
- Khunti, K.; Del Prato, S.; Mathieu, C.; Kahn, S.E.; Gabbay, R.A.; Buse, J.B. COVID-19, Hyperglycemia, and New-Onset Diabetes. Diabetes Care 2021, 44, 2645–2655. [Google Scholar] [CrossRef]
- Turcot, V.; Tchernof, A.; Deshaies, Y.; Pérusse, L.; Bélisle, A.; Marceau, S.; Biron, S.; Lescelleur, O.; Biertho, L.; Vohl, M.-C. LINE-1 Methylation in Visceral Adipose Tissue of Severely Obese Individuals Is Associated with Metabolic Syndrome Status and Related Phenotypes. Clin. Epigenetics 2012, 4, 10. [Google Scholar] [CrossRef]
- Garcia-Lacarte, M.; Milagro, F.I.; Zulet, M.A.; Martinez, J.A.; Mansego, M.L. LINE-1 Methylation Levels, a Biomarker of Weight Loss in Obese Subjects, Are Influenced by Dietary Antioxidant Capacity. Redox Rep. 2016, 21, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Martín-Núñez, G.M.; Rubio-Martín, E.; Cabrera-Mulero, R.; Rojo-Martínez, G.; Olveira, G.; Valdés, S.; Soriguer, F.; Castaño, L.; Morcillo, S. Type 2 Diabetes Mellitus in Relation to Global LINE-1 DNA Methylation in Peripheral Blood: A Cohort Study. Epigenetics 2014, 9, 1322–1328. [Google Scholar] [CrossRef]
- Min, B.; Jeon, K.; Park, J.S.; Kang, Y.-K. Demethylation and Derepression of Genomic Retroelements in the Skeletal Muscles of Aged Mice. Aging Cell 2019, 18, e13042. [Google Scholar] [CrossRef] [PubMed]
- Johnson, A.A.; Akman, K.; Calimport, S.R.G.; Wuttke, D.; Stolzing, A.; de Magalhães, J.P. The Role of DNA Methylation in Aging, Rejuvenation, and Age-Related Disease. Rejuvenation Res. 2012, 15, 483–494. [Google Scholar] [CrossRef] [PubMed]
- Horvath, S.; Raj, K. DNA Methylation-Based Biomarkers and the Epigenetic Clock Theory of Ageing. Nat. Rev. Genet. 2018, 19, 371–384. [Google Scholar] [CrossRef] [PubMed]
- Jintaridth, P.; Mutirangura, A. Distinctive Patterns of Age-Dependent Hypomethylation in Interspersed Repetitive Sequences. Physiol. Genom. 2010, 41, 194–200. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Lan, Y.; Pandey, R.R.; Homolka, D.; Berger, S.L.; Pillai, R.S.; Bartolomei, M.S.; Wang, P.J. TEX15 Associates with MILI and Silences Transposable Elements in Male Germ Cells. Genes Dev. 2020, 34, 745–750. [Google Scholar] [CrossRef]
- De Cecco, M.; Criscione, S.W.; Peterson, A.L.; Neretti, N.; Sedivy, J.M.; Kreiling, J.A. Transposable Elements Become Active and Mobile in the Genomes of Aging Mammalian Somatic Tissues. Aging 2013, 5, 867–883. [Google Scholar] [CrossRef]
- St Laurent, G., 3rd; Hammell, N.; McCaffrey, T.A. A LINE-1 Component to Human Aging: Do LINE Elements Exact a Longevity Cost for Evolutionary Advantage? Mech. Ageing Dev. 2010, 131, 299–305. [Google Scholar] [CrossRef]
- De Cecco, M.; Ito, T.; Petrashen, A.P.; Elias, A.E.; Skvir, N.J.; Criscione, S.W.; Caligiana, A.; Brocculi, G.; Adney, E.M.; Boeke, J.D.; et al. L1 Drives IFN in Senescent Cells and Promotes Age-Associated Inflammation. Nature 2019, 566, 73–78. [Google Scholar] [CrossRef]
- Ohka, F.; Natsume, A.; Motomura, K.; Kishida, Y.; Kondo, Y.; Abe, T.; Nakasu, Y.; Namba, H.; Wakai, K.; Fukui, T.; et al. The Global DNA Methylation Surrogate LINE-1 Methylation Is Correlated with MGMT Promoter Methylation and Is a Better Prognostic Factor for Glioma. PLoS ONE 2011, 6, e23332. [Google Scholar] [CrossRef] [PubMed]
- Chalitchagorn, K.; Shuangshoti, S.; Hourpai, N.; Kongruttanachok, N.; Tangkijvanich, P.; Thong-ngam, D.; Voravud, N.; Sriuranpong, V.; Mutirangura, A. Distinctive Pattern of LINE-1 Methylation Level in Normal Tissues and the Association with Carcinogenesis. Oncogene 2004, 23, 8841–8846. [Google Scholar] [CrossRef]
- Figueiredo, J.C.; Grau, M.V.; Wallace, K.; Levine, A.J.; Shen, L.; Hamdan, R.; Chen, X.; Bresalier, R.S.; McKeown-Eyssen, G.; Haile, R.W.; et al. Global DNA Hypomethylation (LINE-1) in the Normal Colon and Lifestyle Characteristics and Dietary and Genetic Factors. Cancer Epidemiol. Biomark. Prev. 2009, 18, 1041–1049. [Google Scholar] [CrossRef]
- Barchitta, M.; Quattrocchi, A.; Maugeri, A.; Vinciguerra, M.; Agodi, A. LINE-1 Hypomethylation in Blood and Tissue Samples as an Epigenetic Marker for Cancer Risk: A Systematic Review and Meta-Analysis. PLoS ONE 2014, 9, e109478. [Google Scholar] [CrossRef]
- Baba, Y.; Murata, A.; Watanabe, M.; Baba, H. Clinical Implications of the LINE-1 Methylation Levels in Patients with Gastrointestinal Cancer. Surg. Today 2014, 44, 1807–1816. [Google Scholar] [CrossRef] [PubMed]
- Caliri, A.W.; Caceres, A.; Tommasi, S.; Besaratinia, A. Hypomethylation of LINE-1 Repeat Elements and Global Loss of DNA Hydroxymethylation in Vapers and Smokers. Epigenetics 2020, 15, 816–829. [Google Scholar] [CrossRef] [PubMed]
- Moghimi, N.; Di Napoli, M.; Biller, J.; Siegler, J.E.; Shekhar, R.; McCullough, L.D.; Harkins, M.S.; Hong, E.; Alaouieh, D.A.; Mansueto, G.; et al. The Neurological Manifestations of Post-Acute Sequelae of SARS-CoV-2 Infection. Curr. Neurol. Neurosci. Rep. 2021, 21, 44. [Google Scholar] [CrossRef]
- Ramakrishnan, R.K.; Kashour, T.; Hamid, Q.; Halwani, R.; Tleyjeh, I.M. Unraveling the Mystery Surrounding Post-Acute Sequelae of COVID-19. Front. Immunol. 2021, 12, 686029. [Google Scholar] [CrossRef]
- Carfì, A.; Bernabei, R.; Landi, F.; Gemelli Against COVID-19 Post-Acute Care Study Group. Persistent Symptoms in Patients After Acute COVID-19. JAMA 2020, 324, 603–605. [Google Scholar] [CrossRef]
- Petersen, S.E.; Friedrich, M.G.; Leiner, T.; Elias, M.D.; Ferreira, V.M.; Fenski, M.; Flamm, S.D.; Fogel, M.; Garg, R.; Halushka, M.K.; et al. Cardiovascular Magnetic Resonance for Patients With COVID-19. JACC Cardiovasc. Imaging 2022, 15, 685–699. [Google Scholar] [CrossRef]
- Mitrani, R.D.; Dabas, N.; Goldberger, J.J. COVID-19 Cardiac Injury: Implications for Long-Term Surveillance and Outcomes in Survivors. Heart Rhythm 2020, 17, 1984–1990. [Google Scholar] [CrossRef] [PubMed]
- Giustino, G.; Croft, L.B.; Stefanini, G.G.; Bragato, R.; Silbiger, J.J.; Vicenzi, M.; Danilov, T.; Kukar, N.; Shaban, N.; Kini, A.; et al. Characterization of Myocardial Injury in Patients With COVID-19. J. Am. Coll. Cardiol. 2020, 76, 2043–2055. [Google Scholar] [CrossRef] [PubMed]
- Salabei, J.K.; Asnake, Z.T.; Ismail, Z.H.; Charles, K.; Stanger, G.-T.; Abdullahi, A.H.; Abraham, A.T.; Okonoboh, P. COVID-19 and the Cardiovascular System: An Update. Am. J. Med. Sci. 2022, 364, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Caro-Codón, J.; Rey, J.R.; Buño, A.; Iniesta, A.M.; Rosillo, S.O.; Castrejon-Castrejon, S.; Merino, C.; Marco, I.; Martinez, L.A.; Garcia-Veas, J.M.; et al. Characterization of Myocardial Injury in a Cohort of Patients with SARS-CoV-2 Infection. Med. Clin. 2021, 157, 274–280. [Google Scholar] [CrossRef]
- Siripanthong, B.; Asatryan, B.; Hanff, T.C.; Chatha, S.R.; Khanji, M.Y.; Ricci, F.; Muser, D.; Ferrari, V.A.; Nazarian, S.; Santangeli, P.; et al. The Pathogenesis and Long-Term Consequences of COVID-19 Cardiac Injury. JACC Basic Transl. Sci. 2022, 7, 294–308. [Google Scholar] [CrossRef]
- Loh, D. The Potential of Melatonin in the Prevention and Attenuation of Oxidative Hemolysis and Myocardial Injury from cd147 SARS-CoV-2 Spike Protein Receptor Binding. Melatonin Res. 2020, 3, 380–416. [Google Scholar] [CrossRef]
- Italia, L.; Tomasoni, D.; Bisegna, S.; Pancaldi, E.; Stretti, L.; Adamo, M.; Metra, M. COVID-19 and Heart Failure: From Epidemiology During the Pandemic to Myocardial Injury, Myocarditis, and Heart Failure Sequelae. Front. Cardiovasc. Med. 2021, 8, 713560. [Google Scholar] [CrossRef]
- Babapoor-Farrokhran, S.; Gill, D.; Walker, J.; Rasekhi, R.T.; Bozorgnia, B.; Amanullah, A. Myocardial Injury and COVID-19: Possible Mechanisms. Life Sci. 2020, 253, 117723. [Google Scholar] [CrossRef]
- Lindner, D.; Fitzek, A.; Bräuninger, H.; Aleshcheva, G.; Edler, C.; Meissner, K.; Scherschel, K.; Kirchhof, P.; Escher, F.; Schultheiss, H.-P.; et al. Association of Cardiac Infection With SARS-CoV-2 in Confirmed COVID-19 Autopsy Cases. JAMA Cardiol 2020, 5, 1281–1285. [Google Scholar] [CrossRef]
- Dal Ferro, M.; Bussani, R.; Paldino, A.; Nuzzi, V.; Collesi, C.; Zentilin, L.; Schneider, E.; Correa, R.; Silvestri, F.; Zacchigna, S.; et al. SARS-CoV-2, Myocardial Injury and Inflammation: Insights from a Large Clinical and Autopsy Study. Clin. Res. Cardiol. 2021, 110, 1822–1831. [Google Scholar] [CrossRef]
- Wei, L.; Liu, S.; Su, Z.; Cheng, R.; Bai, X.; Li, X. LINE-1 Hypomethylation Is Associated with the Risk of Coronary Heart Disease in Chinese Population. Arq. Bras. Cardiol. 2014, 102, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Cash, H.L.; McGarvey, S.T.; Houseman, E.A.; Marsit, C.J.; Hawley, N.L.; Lambert-Messerlian, G.M.; Viali, S.; Tuitele, J.; Kelsey, K.T. Cardiovascular Disease Risk Factors and DNA Methylation at the LINE-1 Repeat Region in Peripheral Blood from Samoan Islanders. Epigenetics 2011, 6, 1257–1264. [Google Scholar] [CrossRef] [PubMed]
- Guarrera, S.; Fiorito, G.; Onland-Moret, N.C.; Russo, A.; Agnoli, C.; Allione, A.; Di Gaetano, C.; Mattiello, A.; Ricceri, F.; Chiodini, P.; et al. Gene-Specific DNA Methylation Profiles and LINE-1 Hypomethylation Are Associated with Myocardial Infarction Risk. Clin. Epigenetics 2015, 7, 133. [Google Scholar] [CrossRef]
- Gao, C.; Howard-Quijano, K.; Rau, C.; Takamiya, T.; Song, Y.; Shivkumar, K.; Wang, Y.; Mahajan, A. Inflammatory and Apoptotic Remodeling in Autonomic Nervous System Following Myocardial Infarction. PLoS ONE 2017, 12, e0177750. [Google Scholar] [CrossRef]
- Mihalovic, M.; Tousek, P. Myocardial Injury after Stroke. J. Clin. Med. Res. 2021, 11, 2. [Google Scholar] [CrossRef]
- Sheng, W.; Wang, H.; Ma, X.; Qian, Y.; Zhang, P.; Wu, Y.; Zheng, F.; Chen, L.; Huang, G.; Ma, D. LINE-1 Methylation Status and Its Association with Tetralogy of Fallot in Infants. BMC Med. Genom. 2012, 5, 20. [Google Scholar] [CrossRef] [PubMed]
- Roncato, G.; Cardoso, B.; Spilimbergo, F.; Meyer, G.; Rigatto, K. Right Ventricular Hypertrophy or Dilatation in the Pulmonary Arterial Hypertension: Is There a Difference in the Autonomic Imbalance? Eur. Respir. J. 2013, 42 (Suppl. 57), 2652. [Google Scholar]
- Szekely, Y.; Lichter, Y.; Taieb, P.; Banai, A.; Hochstadt, A.; Merdler, I.; Gal Oz, A.; Rothschild, E.; Baruch, G.; Peri, Y.; et al. Spectrum of Cardiac Manifestations in COVID-19: A Systematic Echocardiographic Study. Circulation 2020, 142, 342–353. [Google Scholar] [CrossRef]
- Puntmann, V.O.; Carerj, M.L.; Wieters, I.; Fahim, M.; Arendt, C.; Hoffmann, J.; Shchendrygina, A.; Escher, F.; Vasa-Nicotera, M.; Zeiher, A.M.; et al. Outcomes of Cardiovascular Magnetic Resonance Imaging in Patients Recently Recovered From Coronavirus Disease 2019 (COVID-19). JAMA Cardiol. 2020, 5, 1265–1273. [Google Scholar] [CrossRef]
- Kohlrausch, F.B.; Berteli, T.S.; Wang, F.; Navarro, P.A.; Keefe, D.L. Control of LINE-1 Expression Maintains Genome Integrity in Germline and Early Embryo Development. Reprod. Sci. 2022, 29, 328–340. [Google Scholar] [CrossRef]
- Castañeda, J.; Genzor, P.; Bortvin, A. piRNAs, Transposon Silencing, and Germline Genome Integrity. Mutat. Res. 2011, 714, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Pattamadilok, J.; Huapai, N.; Rattanatanyong, P.; Vasurattana, A.; Triratanachat, S.; Tresukosol, D.; Mutirangura, A. LINE- 1 Hypomethylation Level as a Potential Prognostic Factor for Epithelial Ovarian Cancer. Int. J. Gynecol. Cancer 2008, 18. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, S.; Suzuki, H.; Niinuma, T.; Shimizu, H.; Nojima, M.; Iwaki, H.; Nobuoka, T.; Nishida, T.; Miyazaki, Y.; Takamaru, H.; et al. A Novel Correlation between LINE-1 Hypomethylation and the Malignancy of Gastrointestinal Stromal Tumors. Clin. Cancer Res. 2010, 16, 5114–5123. [Google Scholar] [CrossRef] [PubMed]
- Whongsiri, P.; Goering, W.; Lautwein, T.; Hader, C.; Niegisch, G.; Köhrer, K.; Hoffmann, M.J.; Schulz, W.A. Many Different LINE-1 Retroelements Are Activated in Bladder Cancer. Int. J. Mol. Sci. 2020, 21, 9433. [Google Scholar] [CrossRef]
- Thomas, C.A.; Paquola, A.C.M.; Muotri, A.R. LINE-1 Retrotransposition in the Nervous System. Annu. Rev. Cell Dev. Biol. 2012, 28, 555–573. [Google Scholar] [CrossRef] [PubMed]
- Upton, K.R.; Gerhardt, D.J.; Jesuadian, J.S.; Richardson, S.R.; Sánchez-Luque, F.J.; Bodea, G.O.; Ewing, A.D.; Salvador-Palomeque, C.; van der Knaap, M.S.; Brennan, P.M.; et al. Ubiquitous L1 Mosaicism in Hippocampal Neurons. Cell 2015, 161, 228–239. [Google Scholar] [CrossRef]
- Singer, T.; McConnell, M.J.; Marchetto, M.C.N.; Coufal, N.G.; Gage, F.H. LINE-1 Retrotransposons: Mediators of Somatic Variation in Neuronal Genomes? Trends Neurosci. 2010, 33, 345–354. [Google Scholar] [CrossRef]
- Coufal, N.G.; Garcia-Perez, J.L.; Peng, G.E.; Yeo, G.W.; Mu, Y.; Lovci, M.T.; Morell, M.; O’Shea, K.S.; Moran, J.V.; Gage, F.H. L1 Retrotransposition in Human Neural Progenitor Cells. Nature 2009, 460, 1127–1131. [Google Scholar] [CrossRef]
- Baillie, J.K.; Barnett, M.W.; Upton, K.R.; Gerhardt, D.J.; Richmond, T.A.; De Sapio, F.; Brennan, P.M.; Rizzu, P.; Smith, S.; Fell, M.; et al. Somatic Retrotransposition Alters the Genetic Landscape of the Human Brain. Nature 2011, 479, 534–537. [Google Scholar] [CrossRef]
- Zhao, B.; Wu, Q.; Ye, A.Y.; Guo, J.; Zheng, X.; Yang, X.; Yan, L.; Liu, Q.-R.; Hyde, T.M.; Wei, L.; et al. Somatic LINE-1 Retrotransposition in Cortical Neurons and Non-Brain Tissues of Rett Patients and Healthy Individuals. PLoS Genet. 2019, 15, e1008043. [Google Scholar] [CrossRef]
- Benitez-Guijarro, M.; Lopez-Ruiz, C.; Tarnauskaitė, Ž.; Murina, O.; Mian Mohammad, M.; Williams, T.C.; Fluteau, A.; Sanchez, L.; Vilar-Astasio, R.; Garcia-Canadas, M.; et al. RNase H2, Mutated in Aicardi-Goutières Syndrome, Promotes LINE-1 Retrotransposition. EMBO J. 2018, 37, e98506. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Du, J.; Goodier, J.L.; Hou, J.; Kang, J.; Kazazian, H.H., Jr.; Zhao, K.; Yu, X.-F. Aicardi-Goutières Syndrome Protein TREX1 Suppresses L1 and Maintains Genome Integrity through Exonuclease-Independent ORF1p Depletion. Nucleic Acids Res. 2017, 45, 4619–4631. [Google Scholar] [CrossRef]
- Coufal, N.G.; Garcia-Perez, J.L.; Peng, G.E.; Marchetto, M.C.N.; Muotri, A.R.; Mu, Y.; Carson, C.T.; Macia, A.; Moran, J.V.; Gage, F.H. Ataxia Telangiectasia Mutated (ATM) Modulates Long Interspersed Element-1 (L1) Retrotransposition in Human Neural Stem Cells. Proc. Natl. Acad. Sci. USA 2011, 108, 20382–20387. [Google Scholar] [CrossRef]
- Shpyleva, S.; Melnyk, S.; Pavliv, O.; Pogribny, I.; Jill James, S. Overexpression of LINE-1 Retrotransposons in Autism Brain. Mol. Neurobiol. 2018, 55, 1740–1749. [Google Scholar] [CrossRef] [PubMed]
- Stathopoulos, S.; Gaujoux, R.; Lindeque, Z.; Mahony, C.; Van Der Colff, R.; Van Der Westhuizen, F.; O’Ryan, C. DNA Methylation Associated with Mitochondrial Dysfunction in a South African Autism Spectrum Disorder Cohort. Autism Res. 2020, 13, 1079–1093. [Google Scholar] [CrossRef] [PubMed]
- Bundo, M.; Toyoshima, M.; Okada, Y.; Akamatsu, W.; Ueda, J.; Nemoto-Miyauchi, T.; Sunaga, F.; Toritsuka, M.; Ikawa, D.; Kakita, A.; et al. Increased l1 Retrotransposition in the Neuronal Genome in Schizophrenia. Neuron 2014, 81, 306–313. [Google Scholar] [CrossRef]
- Pereira, G.C.; Sanchez, L.; Schaughency, P.M.; Rubio-Roldán, A.; Choi, J.A.; Planet, E.; Batra, R.; Turelli, P.; Trono, D.; Ostrow, L.W.; et al. Properties of LINE-1 Proteins and Repeat Element Expression in the Context of Amyotrophic Lateral Sclerosis. Mob. DNA 2018, 9, 35. [Google Scholar] [CrossRef]
- Ravel-Godreuil, C.; Znaidi, R.; Bonnifet, T.; Joshi, R.L.; Fuchs, J. Transposable Elements as New Players in Neurodegenerative Diseases. FEBS Lett. 2021, 595, 2733–2755. [Google Scholar] [CrossRef]
- Peze-Heidsieck, E.; Bonnifet, T.; Znaidi, R.; Ravel-Godreuil, C.; Massiani-Beaudoin, O.; Joshi, R.L.; Fuchs, J. Retrotransposons as a Source of DNA Damage in Neurodegeneration. Front. Aging Neurosci. 2021, 13, 786897. [Google Scholar] [CrossRef]
- Baeken, M.W.; Moosmann, B.; Hajieva, P. Retrotransposon Activation by Distressed Mitochondria in Neurons. Biochem. Biophys. Res. Commun. 2020, 525, 570–575. [Google Scholar] [CrossRef]
- Delgado-Roche, L.; Mesta, F. Oxidative Stress as Key Player in Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) Infection. Arch. Med. Res. 2020, 51, 384–387. [Google Scholar] [CrossRef] [PubMed]
- Whongsiri, P.; Pimratana, C.; Wijitsettakul, U.; Jindatip, D.; Sanpavat, A.; Schulz, W.A.; Hoffmann, M.J.; Goering, W.; Boonla, C. LINE-1 ORF1 Protein Is Up-Regulated by Reactive Oxygen Species and Associated with Bladder Urothelial Carcinoma Progression. Cancer Genom. Proteom. 2018, 15, 143–151. [Google Scholar] [CrossRef]
- Patchsung, M.; Boonla, C.; Amnattrakul, P.; Dissayabutra, T.; Mutirangura, A.; Tosukhowong, P. Long Interspersed Nuclear Element-1 Hypomethylation and Oxidative Stress: Correlation and Bladder Cancer Diagnostic Potential. PLoS ONE 2012, 7, e37009. [Google Scholar] [CrossRef]
- Shaukat, A.; Hanif, S.; Shaukat, I.; Shukat, R.; Rajput, S.A.; Jiang, K.; Akhtar, M.; Yang, Y.; Guo, S.; Shaukat, I.; et al. Upregulated-Gene Expression of pro-Inflammatory Cytokines, Oxidative Stress and Apoptotic Markers through Inflammatory, Oxidative and Apoptosis Mediated Signaling Pathways in Bovine Pneumonia. Microb. Pathog. 2021, 155, 104935. [Google Scholar] [CrossRef] [PubMed]
- Kozlov, E.M.; Ivanova, E.; Grechko, A.V.; Wu, W.-K.; Starodubova, A.V.; Orekhov, A.N. Involvement of Oxidative Stress and the Innate Immune System in SARS-CoV-2 Infection. Diseases 2021, 9, 17. [Google Scholar] [CrossRef] [PubMed]
- Saheb Sharif-Askari, N.; Saheb Sharif-Askari, F.; Mdkhana, B.; Hussain Alsayed, H.A.; Alsafar, H.; Alrais, Z.F.; Hamid, Q.; Halwani, R. Upregulation of Oxidative Stress Gene Markers during SARS-COV-2 Viral Infection. Free Radic. Biol. Med. 2021, 172, 688–698. [Google Scholar] [CrossRef] [PubMed]
- Aratani, Y. Myeloperoxidase: Its Role for Host Defense, Inflammation, and Neutrophil Function. Arch. Biochem. Biophys. 2018, 640, 47–52. [Google Scholar] [CrossRef]
- Bjarnason, I. The Use of Fecal Calprotectin in Inflammatory Bowel Disease. Gastroenterol. Hepatol. 2017, 13, 53–56. [Google Scholar]
- Rando, O.J.; Verstrepen, K.J. Timescales of Genetic and Epigenetic Inheritance. Cell 2007, 128, 655–668. [Google Scholar] [CrossRef]
- Galindo-González, L.; Mhiri, C.; Deyholos, M.K.; Grandbastien, M.-A. LTR-Retrotransposons in Plants: Engines of Evolution. Gene 2017, 626, 14–25. [Google Scholar] [CrossRef]
- Kumar, A.; Bennetzen, J.L. Plant Retrotransposons. Annu. Rev. Genet. 1999, 33, 479–532. [Google Scholar] [CrossRef] [PubMed]
- Tharp, M.E.; Malki, S.; Bortvin, A. Maximizing the Ovarian Reserve in Mice by Evading LINE-1 Genotoxicity. Nat. Commun. 2020, 11, 330. [Google Scholar] [CrossRef] [PubMed]
- Giorgi, G.; Marcantonio, P.; Del Re, B. LINE-1 Retrotransposition in Human Neuroblastoma Cells Is Affected by Oxidative Stress. Cell Tissue Res. 2011, 346, 383–391. [Google Scholar] [CrossRef] [PubMed]
- Goodier, J.L.; Zhang, L.; Vetter, M.R.; Kazazian, H.H., Jr. LINE-1 ORF1 Protein Localizes in Stress Granules with Other RNA-Binding Proteins, Including Components of RNA Interference RNA-Induced Silencing Complex. Mol. Cell. Biol. 2007, 27, 6469–6483. [Google Scholar] [CrossRef]
- Mita, P.; Wudzinska, A.; Sun, X.; Andrade, J.; Nayak, S.; Kahler, D.J.; Badri, S.; LaCava, J.; Ueberheide, B.; Yun, C.Y.; et al. LINE-1 Protein Localization and Functional Dynamics during the Cell Cycle. Elife 2018, 7, e30058. [Google Scholar] [CrossRef]
- deHaro, D.; Kines, K.J.; Sokolowski, M.; Dauchy, R.T.; Streva, V.A.; Hill, S.M.; Hanifin, J.P.; Brainard, G.C.; Blask, D.E.; Belancio, V.P. Regulation of L1 Expression and Retrotransposition by Melatonin and Its Receptor: Implications for Cancer Risk Associated with Light Exposure at Night. Nucleic Acids Res. 2014, 42, 7694–7707. [Google Scholar] [CrossRef]
- Belancio, V.P. LINE-1 Activity as Molecular Basis for Genomic Instability Associated with Light Exposure at Night. Mob. Genet. Elem. 2015, 5, 1–5. [Google Scholar] [CrossRef]
- Cook, P.R.; Jones, C.E.; Furano, A.V. Phosphorylation of ORF1p Is Required for L1 Retrotransposition. Proc. Natl. Acad. Sci. USA 2015, 112, 4298–4303. [Google Scholar] [CrossRef]
- Furano, A.V.; Cook, P.R. The Challenge of ORF1p Phosphorylation: Effects on L1 Activity and Its Host. Mob. Genet. Elem. 2016, 6, e1119927. [Google Scholar] [CrossRef]
- Newton, J.C.; Naik, M.T.; Li, G.Y.; Murphy, E.L.; Fawzi, N.L.; Sedivy, J.M.; Jogl, G. Phase Separation of the LINE-1 ORF1 Protein Is Mediated by the N-Terminus and Coiled-Coil Domain. Biophys. J. 2021, 120, 2181–2191. [Google Scholar] [CrossRef]
- Sil, S.; Boeke, J.D.; Holt, L.J. Condensation of LINE-1 Is Required for Retrotransposition. bioRxiv 2022. [Google Scholar] [CrossRef]
- Thakar, J.H.; Wrogemann, K.; Blanchaer, M.C. Effect of Ruthenium Red on Oxidative Phosphorylation and the Calcium and Magnesium Content of Skeletal Muscle Mitochondria of Normal and BIO 14.6 Dystrophic Hamsters. Biochim. Biophys. Acta 1973, 314, 8–14. [Google Scholar] [CrossRef]
- Martín, M.; Macías, M.; Escames, G.; Reiter, R.J.; Agapito, M.T.; Ortiz, G.G.; Acuña-Castroviejo, D. Melatonin-Induced Increased Activity of the Respiratory Chain Complexes I and IV Can Prevent Mitochondrial Damage Induced by Ruthenium Red in Vivo. J. Pineal Res. 2000, 28, 242–248. [Google Scholar] [CrossRef]
- Velasco, I.; Tapia, R. Alterations of Intracellular Calcium Homeostasis and Mitochondrial Function Are Involved in Ruthenium Red Neurotoxicity in Primary Cortical Cultures. J. Neurosci. Res. 2000, 60, 543–551. [Google Scholar] [CrossRef]
- Hariharakrishnan, J.; Satpute, R.M.; Prasad, G.B.K.S.; Bhattacharya, R. Oxidative Stress Mediated Cytotoxicity of Cyanide in LLC-MK2 Cells and Its Attenuation by Alpha-Ketoglutarate and N-Acetyl Cysteine. Toxicol. Lett. 2009, 185, 132–141. [Google Scholar] [CrossRef]
- Kulasinghe, A.; Liu, N.; Tan, C.W.; Monkman, J.; Sinclair, J.E.; Bhuva, D.D.; Godbolt, D.; Pan, L.; Nam, A.; Sadeghirad, H.; et al. Transcriptomic Profiling of Cardiac Tissues from SARS-CoV-2 Patients Identifies DNA Damage. medRxiv 2022. [Google Scholar] [CrossRef]
- Jiang, H.; Mei, Y.-F. SARS-CoV-2 Spike Impairs DNA Damage Repair and Inhibits V(D)J Recombination In Vitro. Viruses 2021, 13, 2056. [Google Scholar] [CrossRef]
- Wu, J.; Lu, L.-Y.; Yu, X. The Role of BRCA1 in DNA Damage Response. Protein Cell 2010, 1, 117–123. [Google Scholar] [CrossRef]
- Mita, P.; Sun, X.; Fenyö, D.; Kahler, D.J.; Li, D.; Agmon, N.; Wudzinska, A.; Keegan, S.; Bader, J.S.; Yun, C.; et al. BRCA1 and S Phase DNA Repair Pathways Restrict LINE-1 Retrotransposition in Human Cells. Nat. Struct. Mol. Biol. 2020, 27, 179–191. [Google Scholar] [CrossRef]
- Kastan, M.B.; Bartek, J. Cell-Cycle Checkpoints and Cancer. Nature 2004, 432, 316–323. [Google Scholar] [CrossRef] [PubMed]
- Bartek, J.; Lukas, C.; Lukas, J. Checking on DNA Damage in S Phase. Nat. Rev. Mol. Cell Biol. 2004, 5, 792–804. [Google Scholar] [CrossRef] [PubMed]
- Das-Bradoo, S.; Ricke, R.M.; Bielinsky, A.-K. Interaction between PCNA and Diubiquitinated Mcm10 Is Essential for Cell Growth in Budding Yeast. Mol. Cell. Biol. 2006, 26, 4806–4817. [Google Scholar] [CrossRef] [PubMed]
- Girgert, R.; Hanf, V.; Emons, G.; Gründker, C. Membrane-Bound Melatonin Receptor MT1 down-Regulates Estrogen Responsive Genes in Breast Cancer Cells. J. Pineal Res. 2009, 47, 23–31. [Google Scholar] [CrossRef]
- Girgert, R.; Hanf, V.; Emons, G.; Gründker, C. Signal Transduction of the Melatonin Receptor MT1 Is Disrupted in Breast Cancer Cells by Electromagnetic Fields. Bioelectromagnetics 2010, 31, 237–245. [Google Scholar] [CrossRef]
- Bracci, M.; Ciarapica, V.; Zabaleta, M.E.; Tartaglione, M.F.; Pirozzi, S.; Giuliani, L.; Piva, F.; Valentino, M.; Ledda, C.; Rapisarda, V.; et al. BRCA1 and BRCA2 Gene Expression: Diurnal Variability and Influence of Shift Work. Cancers 2019, 11, 1146. [Google Scholar] [CrossRef]
- Gooley, J.J.; Chamberlain, K.; Smith, K.A.; Khalsa, S.B.S.; Rajaratnam, S.M.W.; Van Reen, E.; Zeitzer, J.M.; Czeisler, C.A.; Lockley, S.W. Exposure to Room Light before Bedtime Suppresses Melatonin Onset and Shortens Melatonin Duration in Humans. J. Clin. Endocrinol. Metab. 2011, 96, E463–E472. [Google Scholar] [CrossRef]
- Vriend, J.; Reiter, R.J. Breast Cancer Cells: Modulation by Melatonin and the Ubiquitin-Proteasome System—A Review. Mol. Cell. Endocrinol. 2015, 417, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Hill, S.M.; Frasch, T.; Xiang, S.; Yuan, L.; Duplessis, T.; Mao, L. Molecular Mechanisms of Melatonin Anticancer Effects. Integr. Cancer Ther. 2009, 8, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Fischer, T.W.; Slominski, A.; Zmijewski, M.A.; Reiter, R.J.; Paus, R. Melatonin as a Major Skin Protectant: From Free Radical Scavenging to DNA Damage Repair. Exp. Dermatol. 2008, 17, 713–730. [Google Scholar] [CrossRef] [PubMed]
- Santoro, R.; Marani, M.; Blandino, G.; Muti, P.; Strano, S. Melatonin Triggers p53Ser Phosphorylation and Prevents DNA Damage Accumulation. Oncogene 2012, 31, 2931–2942. [Google Scholar] [CrossRef]
- Liu, R.; Fu, A.; Hoffman, A.E.; Zheng, T.; Zhu, Y. Melatonin Enhances DNA Repair Capacity Possibly by Affecting Genes Involved in DNA Damage Responsive Pathways. BMC Cell Biol. 2013, 14, 1–8. [Google Scholar] [CrossRef]
- Majidinia, M.; Sadeghpour, A.; Mehrzadi, S.; Reiter, R.J.; Khatami, N.; Yousefi, B. Melatonin: A Pleiotropic Molecule That Modulates DNA Damage Response and Repair Pathways. J. Pineal Res. 2017, 63, e12416. [Google Scholar] [CrossRef]
- Mir, S.M.; Aliarab, A.; Goodarzi, G.; Shirzad, M.; Jafari, S.M.; Qujeq, D.; Samavarchi Tehrani, S.; Asadi, J. Melatonin: A Smart Molecule in the DNA Repair System. Cell Biochem. Funct. 2022, 40, 4–16. [Google Scholar] [CrossRef]
- Alonso-González, C.; González, A.; Martínez-Campa, C.; Gómez-Arozamena, J.; Cos, S. Melatonin Sensitizes Human Breast Cancer Cells to Ionizing Radiation by Downregulating Proteins Involved in Double-Strand DNA Break Repair. J. Pineal Res. 2015, 58, 189–197. [Google Scholar] [CrossRef]
- Watkins, W.S.; Feusier, J.E.; Thomas, J.; Goubert, C.; Mallick, S.; Jorde, L.B. The Simons Genome Diversity Project: A Global Analysis of Mobile Element Diversity. Genome Biol. Evol. 2020, 12, 779–794. [Google Scholar] [CrossRef]
- Kazazian, H.H., Jr.; Goodier, J.L. LINE Drive. Retrotransposition and Genome Instability. Cell 2002, 110, 277–280. [Google Scholar] [CrossRef]
- Ron, M.; Ulitsky, I. Context-Specific Effects of Sequence Elements on Subcellular Localization of Linear and Circular RNAs. Nat. Commun. 2022, 13, 2481. [Google Scholar] [CrossRef]
- Xiong, F.; Wang, R.; Lee, J.-H.; Li, S.; Chen, S.-F.; Liao, Z.; Hasani, L.A.; Nguyen, P.T.; Zhu, X.; Krakowiak, J.; et al. RNA m6A Modification Orchestrates a LINE-1-Host Interaction That Facilitates Retrotransposition and Contributes to Long Gene Vulnerability. Cell Res. 2021, 31, 861–885. [Google Scholar] [CrossRef]
- Hwang, S.-Y.; Jung, H.; Mun, S.; Lee, S.; Park, K.; Baek, S.C.; Moon, H.C.; Kim, H.; Kim, B.; Choi, Y.; et al. L1 Retrotransposons Exploit RNA m6A Modification as an Evolutionary Driving Force. Nat. Commun. 2021, 12, 880. [Google Scholar] [CrossRef]
- Selmi, T.; Lanzuolo, C. Driving Chromatin Organisation through N6-Methyladenosine Modification of RNA: What Do We Know and What Lies Ahead? Genes 2022, 13, 340. [Google Scholar] [CrossRef]
- Liao, S.; Sun, H.; Xu, C. YTH Domain: A Family of N6-Methyladenosine (m6A) Readers. Genom. Proteom. Bioinform. 2018, 16, 99–107. [Google Scholar] [CrossRef]
- Chen, C.; Liu, W.; Guo, J.; Liu, Y.; Liu, X.; Liu, J.; Dou, X.; Le, R.; Huang, Y.; Li, C.; et al. Nuclear m6A Reader YTHDC1 Regulates the Scaffold Function of LINE1 RNA in Mouse ESCs and Early Embryos. Protein Cell 2021, 12, 455–474. [Google Scholar] [CrossRef]
- He, C.; Lan, F. RNA m6A Meets Transposable Elements and Chromatin. Protein Cell 2021, 12, 906–910. [Google Scholar] [CrossRef]
- Boccaletto, P.; Machnicka, M.A.; Purta, E.; Piatkowski, P.; Baginski, B.; Wirecki, T.K.; de Crécy-Lagard, V.; Ross, R.; Limbach, P.A.; Kotter, A.; et al. MODOMICS: A Database of RNA Modification Pathways. 2017 Update. Nucleic Acids Res. 2018, 46, D303–D307. [Google Scholar] [CrossRef]
- Machnicka, M.A.; Milanowska, K.; Osman Oglou, O.; Purta, E.; Kurkowska, M.; Olchowik, A.; Januszewski, W.; Kalinowski, S.; Dunin-Horkawicz, S.; Rother, K.M.; et al. MODOMICS: A Database of RNA Modification Pathways—2013 Update. Nucleic Acids Res. 2013, 41, D262–D267. [Google Scholar] [CrossRef]
- Meyer, K.D.; Jaffrey, S.R. The Dynamic Epitranscriptome: N6-Methyladenosine and Gene Expression Control. Nat. Rev. Mol. Cell Biol. 2014, 15, 313–326. [Google Scholar] [CrossRef]
- Liu, N.; Pan, T. N6-Methyladenosine–encoded Epitranscriptomics. Nat. Struct. Mol. Biol. 2016, 23, 98–102. [Google Scholar] [CrossRef]
- Xiong, X.; Yi, C.; Peng, J. Epitranscriptomics: Toward A Better Understanding of RNA Modifications. Genom. Proteom. Bioinform. 2017, 15, 147–153. [Google Scholar] [CrossRef]
- Seo, K.W.; Kleiner, R.E. Mechanisms of Epitranscriptomic Gene Regulation. Biopolymers 2021, 112, e23403. [Google Scholar] [CrossRef]
- Gonzales-van Horn, S.R.; Sarnow, P. Making the Mark: The Role of Adenosine Modifications in the Life Cycle of RNA Viruses. Cell Host Microbe 2017, 21, 661–669. [Google Scholar] [CrossRef]
- Tan, B.; Gao, S.-J. RNA Epitranscriptomics: Regulation of Infection of RNA and DNA Viruses by N6 -Methyladenosine (m6 A). Rev. Med. Virol. 2018, 28, e1983. [Google Scholar] [CrossRef]
- Malone, B.; Urakova, N.; Snijder, E.J.; Campbell, E.A. Structures and Functions of Coronavirus Replication-Transcription Complexes and Their Relevance for SARS-CoV-2 Drug Design. Nat. Rev. Mol. Cell Biol. 2022, 23, 21–39. [Google Scholar] [CrossRef]
- Williams, G.D.; Gokhale, N.S.; Horner, S.M. Regulation of Viral Infection by the RNA Modification N6-Methyladenosine. Annu. Rev. Virol. 2019, 6, 235–253. [Google Scholar] [CrossRef]
- Netzband, R.; Pager, C.T. Epitranscriptomic Marks: Emerging Modulators of RNA Virus Gene Expression. Wiley Interdiscip. Rev. RNA 2020, 11, e1576. [Google Scholar] [CrossRef]
- Stamatopoulou, V.; Zaravinos, A. Epitranscriptomics Markers Regulate the Infection by RNA Viruses. In Epitranscriptomics; Jurga, S., Barciszewski, J., Eds.; Springer International Publishing: Cham, Switzerland, 2021; pp. 141–163. [Google Scholar] [CrossRef]
- Xue, M.; Zhao, B.S.; Zhang, Z.; Lu, M.; Harder, O.; Chen, P.; Lu, Z.; Li, A.; Ma, Y.; Xu, Y.; et al. Viral N6-Methyladenosine Upregulates Replication and Pathogenesis of Human Respiratory Syncytial Virus. Nat. Commun. 2019, 10, 4595. [Google Scholar] [CrossRef]
- Courtney, D.G.; Kennedy, E.M.; Dumm, R.E.; Bogerd, H.P.; Tsai, K.; Heaton, N.S.; Cullen, B.R. Epitranscriptomic Enhancement of Influenza A Virus Gene Expression and Replication. Cell Host Microbe 2017, 22, 377–386.e5. [Google Scholar] [CrossRef]
- Kennedy, E.M.; Bogerd, H.P.; Kornepati, A.V.R.; Kang, D.; Ghoshal, D.; Marshall, J.B.; Poling, B.C.; Tsai, K.; Gokhale, N.S.; Horner, S.M.; et al. Posttranscriptional m(6)A Editing of HIV-1 mRNAs Enhances Viral Gene Expression. Cell Host Microbe 2016, 19, 675–685. [Google Scholar] [CrossRef]
- Lichinchi, G.; Gao, S.; Saletore, Y.; Gonzalez, G.M.; Bansal, V.; Wang, Y.; Mason, C.E.; Rana, T.M. Dynamics of the Human and Viral m(6)A RNA Methylomes during HIV-1 Infection of T Cells. Nat. Microbiol 2016, 1, 16011. [Google Scholar] [CrossRef]
- Lichinchi, G.; Zhao, B.S.; Wu, Y.; Lu, Z.; Qin, Y.; He, C.; Rana, T.M. Dynamics of Human and Viral RNA Methylation during Zika Virus Infection. Cell Host Microbe 2016, 20, 666–673. [Google Scholar] [CrossRef]
- Wang, P.; Doxtader, K.A.; Nam, Y. Structural Basis for Cooperative Function of Mettl3 and Mettl14 Methyltransferases. Mol. Cell 2016, 63, 306–317. [Google Scholar] [CrossRef]
- Wang, J.; Wang, J.; Gu, Q.; Ma, Y.; Yang, Y.; Zhu, J.; Zhang, Q.A. The Biological Function of m6A Demethylase ALKBH5 and Its Role in Human Disease. Cancer Cell Int. 2020, 20, 347. [Google Scholar] [CrossRef]
- Rajecka, V.; Skalicky, T.; Vanacova, S. The Role of RNA Adenosine Demethylases in the Control of Gene Expression. Biochim. Biophys. Acta Gene Regul. Mech. 2019, 1862, 343–355. [Google Scholar] [CrossRef]
- Zou, S.; Toh, J.D.W.; Wong, K.H.Q.; Gao, Y.-G.; Hong, W.; Woon, E.C.Y. N(6)-Methyladenosine: A Conformational Marker That Regulates the Substrate Specificity of Human Demethylases FTO and ALKBH5. Sci. Rep. 2016, 6, 25677. [Google Scholar] [CrossRef]
- Liu, J.E.; Xu, Y.-P.; Li, K.; Ye, Q.; Zhou, H.-Y.; Sun, H.; Li, X.; Yu, L.; Deng, Y.-Q.; Li, R.-T.; et al. The m6A Methylome of SARS-CoV-2 in Host Cells. Cell Res. 2021, 31, 404–414. [Google Scholar] [CrossRef]
- Zaccara, S.; Jaffrey, S.R. A Unified Model for the Function of YTHDF Proteins in Regulating m6A-Modified mRNA. Cell 2020, 181, 1582–1595.e18. [Google Scholar] [CrossRef]
- Wang, X.; Lu, Z.; Gomez, A.; Hon, G.C.; Yue, Y.; Han, D.; Fu, Y.; Parisien, M.; Dai, Q.; Jia, G.; et al. N6-Methyladenosine-Dependent Regulation of Messenger RNA Stability. Nature 2014, 505, 117–120. [Google Scholar] [CrossRef]
- Qiu, X.; Hua, X.; Li, Q.; Zhou, Q.; Chen, J. m6A Regulator-Mediated Methylation Modification Patterns and Characteristics of Immunity in Blood Leukocytes of COVID-19 Patients. Front. Immunol. 2021, 12, 774776. [Google Scholar] [CrossRef]
- An, S.; Xie, Z.; Liao, Y.; Jiang, J.; Dong, W.; Yin, F.; Li, W.-X.; Ye, L.; Lin, J.; Liang, H. Systematic Analysis of Clinical Relevance and Molecular Characterization of m6A in COVID-19 Patients. Genes Dis. 2022, 9, 1170–1173. [Google Scholar] [CrossRef]
- Govorkova, E.A.; Murti, G.; Meignier, B.; de Taisne, C.; Webster, R.G. African Green Monkey Kidney (Vero) Cells Provide an Alternative Host Cell System for Influenza A and B Viruses. J. Virol. 1996, 70, 5519–5524. [Google Scholar] [CrossRef]
- Kumar, R.; Khandelwal, N.; Chander, Y.; Nagori, H.; Verma, A.; Barua, A.; Godara, B.; Pal, Y.; Gulati, B.R.; Tripathi, B.N.; et al. S-Adenosylmethionine-Dependent Methyltransferase Inhibitor DZNep Blocks Transcription and Translation of SARS-CoV-2 Genome with a Low Tendency to Select for Drug-Resistant Viral Variants. Antivir. Res. 2022, 197, 105232. [Google Scholar] [CrossRef]
- Relier, S.; Rivals, E.; David, A. The Multifaceted Functions of the Fat Mass and Obesity-Associated Protein (FTO) in Normal and Cancer Cells. RNA Biol. 2022, 19, 132–142. [Google Scholar] [CrossRef]
- Zannella, C.; Rinaldi, L.; Boccia, G.; Chianese, A.; Sasso, F.C.; De Caro, F.; Franci, G.; Galdiero, M. Regulation of m6A Methylation as a New Therapeutic Option against COVID-19. Pharmaceuticals 2021, 14, 1135. [Google Scholar] [CrossRef]
- Zhang, X.; Hao, H.; Ma, L.; Zhang, Y.; Hu, X.; Chen, Z.; Liu, D.; Yuan, J.; Hu, Z.; Guan, W. Methyltransferase-like 3 Modulates Severe Acute Respiratory Syndrome Coronavirus-2 RNA N6-Methyladenosine Modification and Replication. MBio 2021, 12, e0106721. [Google Scholar] [CrossRef]
- Meng, Y.; Zhang, Q.; Wang, K.; Zhang, X.; Yang, R.; Bi, K.; Chen, W.; Diao, H. RBM15-Mediated N6-Methyladenosine Modification Affects COVID-19 Severity by Regulating the Expression of Multitarget Genes. Cell Death Dis. 2021, 12, 732. [Google Scholar] [CrossRef]
- Burgess, H.M.; Depledge, D.P.; Thompson, L.; Srinivas, K.P.; Grande, R.C.; Vink, E.I.; Abebe, J.S.; Blackaby, W.P.; Hendrick, A.; Albertella, M.R.; et al. Targeting the m6A RNA Modification Pathway Blocks SARS-CoV-2 and HCoV-OC43 Replication. Genes Dev. 2021, 35, 1005–1019. [Google Scholar] [CrossRef]
- Li, X.; Jiang, Y.; Sun, X.; Wu, Y.; Chen, Z. METTL3 Is Required for Maintaining β-Cell Function. Metabolism 2021, 116, 154702. [Google Scholar] [CrossRef]
- Wang, J.; Ishfaq, M.; Xu, L.; Xia, C.; Chen, C.; Li, J. METTL3/m6A/miRNA-873-5p Attenuated Oxidative Stress and Apoptosis in Colistin-Induced Kidney Injury by Modulating Keap1/Nrf2 Pathway. Front. Pharmacol. 2019, 10, 517. [Google Scholar] [CrossRef]
- Gullberg, R.C.; Jordan Steel, J.; Moon, S.L.; Soltani, E.; Geiss, B.J. Oxidative Stress Influences Positive Strand RNA Virus Genome Synthesis and Capping. Virology 2015, 475, 219–229. [Google Scholar] [CrossRef]
- Chen, B.; Ye, F.; Yu, L.; Jia, G.; Huang, X.; Zhang, X.; Peng, S.; Chen, K.; Wang, M.; Gong, S.; et al. Development of Cell-Active N6-Methyladenosine RNA Demethylase FTO Inhibitor. J. Am. Chem. Soc. 2012, 134, 17963–17971. [Google Scholar] [CrossRef]
- Zhou, L.-L.; Xu, H.; Huang, Y.; Yang, C.-G. Targeting the RNA Demethylase FTO for Cancer Therapy. RSC Chem. Biol. 2021, 2, 1352–1369. [Google Scholar] [CrossRef]
- Li, G.-M.; Chen, J.-R.; Zhang, H.-Q.; Cao, X.-Y.; Sun, C.; Peng, F.; Yin, Y.-P.; Lin, Z.; Yu, L.; Chen, Y.; et al. Update on Pharmacological Activities, Security, and Pharmacokinetics of Rhein. Evid. Based Complement. Alternat. Med. 2021, 2021, 4582412. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.-W.; Su, Y.; Sheng, J.-T.; Gu, L.-M.; Zhao, Y.; Chen, X.-X.; Chen, C.; Li, W.-Z.; Li, K.-S.; Dai, J.-P. Anti-Influenza A Virus Activity of Rhein through Regulating Oxidative Stress, TLR4, Akt, MAPK, and NF-κB Signal Pathways. PLoS ONE 2018, 13, e0191793. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Wei, X.; Jian, Z.; Zhang, X. METTL3 Expression Is Associated with Glycolysis Metabolism and Sensitivity to Glycolytic Stress in Hepatocellular Carcinoma. Cancer Med. 2020, 9, 2859–2867. [Google Scholar] [CrossRef] [PubMed]
- Ardestani, A.; Azizi, Z. Targeting Glucose Metabolism for Treatment of COVID-19. Signal Transduct Target. Ther. 2021, 6, 112. [Google Scholar] [CrossRef]
- Zheng, Y.; Li, R.; Liu, S. Immunoregulation with mTOR Inhibitors to Prevent COVID-19 Severity: A Novel Intervention Strategy beyond Vaccines and Specific Antiviral Medicines. J. Med. Virol. 2020, 92, 1495–1500. [Google Scholar] [CrossRef]
- Faulds, K.J.; Egelston, J.N.; Sedivy, L.J.; Mitchell, M.K.; Garimella, S.; Kozlowski, H.; D’Alessandro, A.; Hansen, K.C.; Balsbaugh, J.L.; Phiel, C.J. Glycogen Synthase Kinase-3 (GSK-3) Activity Regulates mRNA Methylation in Mouse Embryonic Stem Cells. J. Biol. Chem. 2018, 293, 10731–10743. [Google Scholar] [CrossRef]
- Gerken, T.; Girard, C.A.; Tung, Y.-C.L.; Webby, C.J.; Saudek, V.; Hewitson, K.S.; Yeo, G.S.H.; McDonough, M.A.; Cunliffe, S.; McNeill, L.A.; et al. The Obesity-Associated FTO Gene Encodes a 2-Oxoglutarate-Dependent Nucleic Acid Demethylase. Science 2007, 318, 1469–1472. [Google Scholar] [CrossRef]
- Sanchez-Pulido, L.; Andrade-Navarro, M.A. The FTO (fat Mass and Obesity Associated) Gene Codes for a Novel Member of the Non-Heme Dioxygenase Superfamily. BMC Biochem. 2007, 8, 23. [Google Scholar] [CrossRef]
- Wen, C.; Lan, M.; Tan, X.; Wang, X.; Zheng, Z.; Lv, M.; Zhao, X.; Luo, H.; Liu, Y.; Wei, P.; et al. GSK3β Exacerbates Myocardial Ischemia/Reperfusion Injury by Inhibiting Myc. Oxid. Med. Cell. Longev. 2022, 2022, 2588891. [Google Scholar] [CrossRef]
- Pelengaris, S.; Khan, M.; Evan, G. C-MYC: More than Just a Matter of Life and Death. Nat. Rev. Cancer 2002, 2, 764–776. [Google Scholar] [CrossRef]
- Rocak, S.; Linder, P. DEAD-Box Proteins: The Driving Forces behind RNA Metabolism. Nat. Rev. Mol. Cell Biol. 2004, 5, 232–241. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.; Chen, G.; Jia, H.; He, X.; Jing, Z. DDX5 RNA Helicases: Emerging Roles in Viral Infection. Int. J. Mol. Sci. 2018, 19, 1122. [Google Scholar] [CrossRef] [PubMed]
- Goh, P.-Y.; Tan, Y.-J.; Lim, S.P.; Tan, Y.H.; Lim, S.G.; Fuller-Pace, F.; Hong, W. Cellular RNA Helicase p68 Relocalization and Interaction with the Hepatitis C Virus (HCV) NS5B Protein and the Potential Role of p68 in HCV RNA Replication. J. Virol. 2004, 78, 5288–5298. [Google Scholar] [CrossRef]
- Zhou, X.; Luo, J.; Mills, L.; Wu, S.; Pan, T.; Geng, G.; Zhang, J.; Luo, H.; Liu, C.; Zhang, H. DDX5 Facilitates HIV-1 Replication as a Cellular Co-Factor of Rev. PLoS ONE 2013, 8, e65040. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Ge, L.-L.; Li, P.-P.; Wang, Y.; Sun, M.-X.; Huang, L.; Ishag, H.; Di, D.-D.; Shen, Z.-Q.; Fan, W.-X.; et al. The DEAD-Box RNA Helicase DDX5 Acts as a Positive Regulator of Japanese Encephalitis Virus Replication by Binding to Viral 3′ UTR. Antivir. Res. 2013, 100, 487–499. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.-Y.; Chen, W.-N.; Poon, K.-M.V.; Zheng, B.-J.; Lin, X.; Wang, Y.-X.; Wen, Y.-M. Interaction between SARS-CoV Helicase and a Multifunctional Cellular Protein (Ddx5) Revealed by Yeast and Mammalian Cell Two-Hybrid Systems. Arch. Virol. 2009, 154, 507–512. [Google Scholar] [CrossRef]
- Zheng, G.; Dahl, J.A.; Niu, Y.; Fedorcsak, P.; Huang, C.-M.; Li, C.J.; Vågbø, C.B.; Shi, Y.; Wang, W.-L.; Song, S.-H.; et al. ALKBH5 Is a Mammalian RNA Demethylase That Impacts RNA Metabolism and Mouse Fertility. Mol. Cell 2013, 49, 18–29. [Google Scholar] [CrossRef]
- Zheng, Q.; Hou, J.; Zhou, Y.; Li, Z.; Cao, X. The RNA Helicase DDX46 Inhibits Innate Immunity by Entrapping m6A-Demethylated Antiviral Transcripts in the Nucleus. Nat. Immunol. 2017, 18, 1094–1103. [Google Scholar] [CrossRef]
- Zhou, J.; Zhang, X.; Hu, J.; Qu, R.; Yu, Z.; Xu, H.; Chen, H.; Yan, L.; Ding, C.; Zou, Q.; et al. m6A Demethylase ALKBH5 Controls CD4+ T Cell Pathogenicity and Promotes Autoimmunity. Sci. Adv. 2021, 7, eabg0470. [Google Scholar] [CrossRef]
- Soto-Rifo, R.; Ohlmann, T. The Role of the DEAD-Box RNA Helicase DDX3 in mRNA Metabolism. Wiley Interdiscip. Rev. RNA 2013, 4, 369–385. [Google Scholar] [CrossRef]
- Li, Q.; Zhang, P.; Zhang, C.; Wang, Y.; Wan, R.; Yang, Y.; Guo, X.; Huo, R.; Lin, M.; Zhou, Z.; et al. DDX3X Regulates Cell Survival and Cell Cycle during Mouse Early Embryonic Development. J. Biomed. Res. 2014, 28, 282–291. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Feng, T.; Pan, W.; Shi, X.; Dai, J. DEAD-Box RNA Helicase DDX3X Inhibits DENV Replication via Regulating Type One Interferon Pathway. Biochem. Biophys. Res. Commun. 2015, 456, 327–332. [Google Scholar] [CrossRef] [PubMed]
- Shriwas, O.; Priyadarshini, M.; Samal, S.K.; Rath, R.; Panda, S.; Das Majumdar, S.K.; Muduly, D.K.; Botlagunta, M.; Dash, R. DDX3 Modulates Cisplatin Resistance in OSCC through ALKBH5-Mediated m6A-Demethylation of FOXM1 and NANOG. Apoptosis 2020, 25, 233–246. [Google Scholar] [CrossRef]
- Shah, A.; Rashid, F.; Awan, H.M.; Hu, S.; Wang, X.; Chen, L.; Shan, G. The DEAD-Box RNA Helicase DDX3 Interacts with m6A RNA Demethylase ALKBH5. Stem Cells Int. 2017, 2017, 8596135. [Google Scholar] [CrossRef] [PubMed]
- Lou, X.; Wang, J.-J.; Wei, Y.-Q.; Sun, J.-J. Emerging Role of RNA Modification N6-Methyladenosine in Immune Evasion. Cell Death Dis. 2021, 12, 300. [Google Scholar] [CrossRef] [PubMed]
- Ji, D.; Juhas, M.; Tsang, C.M.; Kwok, C.K.; Li, Y.; Zhang, Y. Discovery of G-Quadruplex-Forming Sequences in SARS-CoV-2. Brief. Bioinform. 2021, 22, 1150–1160. [Google Scholar] [CrossRef]
- Xu, J.; Cai, Y.; Ma, Z.; Jiang, B.; Liu, W.; Cheng, J.; Guo, N.; Wang, Z.; Sealy, J.E.; Song, C.; et al. The RNA Helicase DDX5 Promotes Viral Infection via Regulating N6-Methyladenosine Levels on the DHX58 and NFκB Transcripts to Dampen Antiviral Innate Immunity. PLoS Pathog. 2021, 17, e1009530. [Google Scholar] [CrossRef]
- Zhao, W.; Wang, Z.; Chen, W.; Chen, M.; Han, J.; Yin, X.; Hu, X.; Wang, S.; Zan, J.; Zheng, L. SARS-CoV-2 Spike Protein Alleviates Atherosclerosis by Suppressing Macrophage Lipid Uptake through Regulating R-Loop Formation on MSR1 mRNA. Clin. Transl. Med. 2021, 11, e391. [Google Scholar] [CrossRef]
- Sidibé, H.; Dubinski, A.; Vande Velde, C. The Multi-Functional RNA-Binding Protein G3BP1 and Its Potential Implication in Neurodegenerative Disease. J. Neurochem. 2021, 157, 944–962. [Google Scholar] [CrossRef]
- Lee, A.K.; Klein, J.; Fon Tacer, K.; Lord, T.; Oatley, M.J.; Oatley, J.M.; Porter, S.N.; Pruett-Miller, S.M.; Tikhonova, E.B.; Karamyshev, A.L.; et al. Translational Repression of G3BP in Cancer and Germ Cells Suppresses Stress Granules and Enhances Stress Tolerance. Mol. Cell 2020, 79, 645–659.e9. [Google Scholar] [CrossRef]
- Montero, H.; Trujillo-Alonso, V. Stress Granules in the Viral Replication Cycle. Viruses 2011, 3, 2328–2338. [Google Scholar] [CrossRef] [PubMed]
- Onomoto, K.; Jogi, M.; Yoo, J.-S.; Narita, R.; Morimoto, S.; Takemura, A.; Sambhara, S.; Kawaguchi, A.; Osari, S.; Nagata, K.; et al. Critical Role of an Antiviral Stress Granule Containing RIG-I and PKR in Viral Detection and Innate Immunity. PLoS ONE 2012, 7, e43031. [Google Scholar] [CrossRef]
- Onomoto, K.; Yoneyama, M.; Fung, G.; Kato, H.; Fujita, T. Antiviral Innate Immunity and Stress Granule Responses. Trends Immunol. 2014, 35, 420–428. [Google Scholar] [CrossRef] [PubMed]
- Edupuganti, R.R.; Geiger, S.; Lindeboom, R.G.H.; Shi, H.; Hsu, P.J.; Lu, Z.; Wang, S.-Y.; Baltissen, M.P.A.; Jansen, P.W.T.C.; Rossa, M.; et al. N6-Methyladenosine (m6A) Recruits and Repels Proteins to Regulate mRNA Homeostasis. Nat. Struct. Mol. Biol. 2017, 24, 870–878. [Google Scholar] [CrossRef]
- Laver, J.D.; Ly, J.; Winn, A.K.; Karaiskakis, A.; Lin, S.; Nie, K.; Benic, G.; Jaberi-Lashkari, N.; Cao, W.X.; Khademi, A.; et al. The RNA-Binding Protein Rasputin/G3BP Enhances the Stability and Translation of Its Target mRNAs. Cell Rep. 2020, 30, 3353–3367.e7. [Google Scholar] [CrossRef] [PubMed]
- Ries, R.J.; Zaccara, S.; Klein, P.; Olarerin-George, A.; Namkoong, S.; Pickering, B.F.; Patil, D.P.; Kwak, H.; Lee, J.H.; Jaffrey, S.R. m6A Enhances the Phase Separation Potential of mRNA. Nature 2019, 571, 424–428. [Google Scholar] [CrossRef]
- Reineke, L.C.; Lloyd, R.E. The Stress Granule Protein G3BP1 Recruits Protein Kinase R to Promote Multiple Innate Immune Antiviral Responses. J. Virol. 2015, 89, 2575–2589. [Google Scholar] [CrossRef]
- Tsai, W.-C.; Gayatri, S.; Reineke, L.C.; Sbardella, G.; Bedford, M.T.; Lloyd, R.E. Arginine Demethylation of G3BP1 Promotes Stress Granule Assembly. J. Biol. Chem. 2016, 291, 22671–22685. [Google Scholar] [CrossRef]
- Fu, Y.; Zhuang, X. m6A-Binding YTHDF Proteins Promote Stress Granule Formation. Nat. Chem. Biol. 2020, 16, 955–963. [Google Scholar] [CrossRef]
- Anders, M.; Chelysheva, I.; Goebel, I.; Trenkner, T.; Zhou, J.; Mao, Y.; Verzini, S.; Qian, S.-B.; Ignatova, Z. Dynamic m6A Methylation Facilitates mRNA Triaging to Stress Granules. Life Sci. Alliance 2018, 1, e201800113. [Google Scholar] [CrossRef]
- Wu, J.; Frazier, K.; Zhang, J.; Gan, Z.; Wang, T.; Zhong, X. Emerging Role of m6 A RNA Methylation in Nutritional Physiology and Metabolism. Obes. Rev. 2020, 21, e12942. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.; Cheng, C.; Song, X.; Yang, L.; Deng, D.; Du, Z.; Chen, X.; Zou, T.; Qiao, L.; Li, N.; et al. Acupoint Combination Effect of Shenmen (HT 7) and Sanyinjiao (SP 6) in Treating Insomnia: Study Protocol for a Randomized Controlled Trial. Trials 2020, 21, 261. [Google Scholar] [CrossRef] [PubMed]
- Rong, B.; Feng, R.; Liu, C.; Wu, Q.; Sun, C. Reduced Delivery of Epididymal Adipocyte-Derived Exosomal Resistin Is Essential for Melatonin Ameliorating Hepatic Steatosis in Mice. J. Pineal Res. 2019, 66, e12561. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Tang, P.; Wang, J.; Ye, W.; Ge, X.; Rong, Y.; Ji, C.; Wang, Z.; Bai, J.; Fan, J.; et al. Extracellular Vesicles Derived from Melatonin-Preconditioned Mesenchymal Stem Cells Containing USP29 Repair Traumatic Spinal Cord Injury by Stabilizing NRF2. J. Pineal Res. 2021, 71, e12769. [Google Scholar] [CrossRef]
- Yang, L.; Liu, X.; Song, L.; Su, G.; Di, A.; Bai, C.; Wei, Z.; Li, G. Melatonin Restores the Pluripotency of Long-Term-Cultured Embryonic Stem Cells through Melatonin Receptor-Dependent m6A RNA Regulation. J. Pineal Res. 2020, 69, e12669. [Google Scholar] [CrossRef]
- Qi, S.; Fang, Z.; Wang, D.; Menendez, P.; Yao, K.; Ji, J. Concise Review: Induced Pluripotency by Defined Factors: Prey of Oxidative Stress. Stem. Cells 2015, 33, 1371–1376. [Google Scholar] [CrossRef]
- Lee, J.; Cho, Y.S.; Jung, H.; Choi, I. Pharmacological Regulation of Oxidative Stress in Stem Cells. Oxid. Med. Cell. Longev. 2018, 2018, 4081890. [Google Scholar] [CrossRef]
- Yang, B.; Chen, Q. Cross-Talk between Oxidative Stress and m6A RNA Methylation in Cancer. Oxid. Med. Cell. Longev. 2021, 2021, 6545728. [Google Scholar] [CrossRef]
- Chen, N.; Tang, J.; Su, Q.; Chou, W.-C.; Zheng, F.; Guo, Z.; Yu, G.; Shao, W.; Li, H.; Wu, S. Paraquat-Induced Oxidative Stress Regulates N6-Methyladenosine (m6A) Modification of Circular RNAs. Environ. Pollut. 2021, 290, 117816. [Google Scholar] [CrossRef]
- Su, Q.; Chen, N.; Tang, J.; Wang, J.; Chou, W.-C.; Zheng, F.; Shao, W.; Yu, G.; Cai, P.; Guo, Z.; et al. Paraquat-Induced Oxidative Stress Regulates N6-Methyladenosine (m6A) Modification of Long Noncoding RNAs in Neuro-2a Cells. Ecotoxicol. Environ. Saf. 2022, 237, 113503. [Google Scholar] [CrossRef]
- Tang, J.; Su, Q.; Guo, Z.; Zhou, J.; Zheng, F.; Yu, G.; Shao, W.; Hu, H.; Wu, S.; Li, H. N6-methyladenosine(m6A) Demethylase FTO Regulates Cellular Apoptosis Following Cobalt-Induced Oxidative Stress. Environ. Pollut. 2022, 297, 118749. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.-B.; Shi, G.-X.; Liu, T.; Li, B.; Jiang, S.-D.; Zheng, X.-F.; Jiang, L.-S. Oxidative Stress Aggravates Apoptosis of Nucleus Pulposus Cells through m6A Modification of MAT2A Pre-mRNA by METTL16. Oxid. Med. Cell. Longev. 2022, 2022, 4036274. [Google Scholar] [CrossRef] [PubMed]
- Bagchi, D.; Bagchi, M.; Stohs, S.J. Chromium (VI)-Induced Oxidative Stress, Apoptotic Cell Death and Modulation of p53 Tumor Suppressor Gene. Mol. Cell. Biochem. 2001, 222, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.; Li, T.; Yang, M.; Su, L.; Zhu, Z.; Zhao, S.; Zeng, W.; Zheng, Y. Melatonin Attenuates Chromium (VI)-Induced Spermatogonial Stem Cell/Progenitor Mitophagy by Restoration of METTL3-Mediated RNA N6-Methyladenosine Modification. Front. Cell Dev. Biol 2021, 9, 684398. [Google Scholar] [CrossRef]
- Sánchez-Sánchez, A.M.; Martín, V.; García-Santos, G.; Rodríguez-Blanco, J.; Casado-Zapico, S.; Suarez-Garnacho, S.; Antolín, I.; Rodriguez, C. Intracellular Redox State as Determinant for Melatonin Antiproliferative vs Cytotoxic Effects in Cancer Cells. Free Radic. Res. 2011, 45, 1333–1341. [Google Scholar] [CrossRef] [PubMed]
- Arnao, M.B.; Hernández-Ruiz, J. Melatonin and Reactive Oxygen and Nitrogen Species: A Model for the Plant Redox Network. Melatonin Res. 2019, 2, 152–168. [Google Scholar] [CrossRef]
- Li, H.; Chang, J.; Chen, H.; Wang, Z.; Gu, X.; Wei, C.; Zhang, Y.; Ma, J.; Yang, J.; Zhang, X. Exogenous Melatonin Confers Salt Stress Tolerance to Watermelon by Improving Photosynthesis and Redox Homeostasis. Front. Plant Sci. 2017, 8, 295. [Google Scholar] [CrossRef]
- Miller, E.; Morel, A.; Saso, L.; Saluk, J. Melatonin Redox Activity. Its Potential Clinical Applications in Neurodegenerative Disorders. Curr. Top. Med. Chem. 2015, 15, 163–169. [Google Scholar] [CrossRef]
- Osseni, R.A.; Rat, P.; Bogdan, A.; Warnet, J.M.; Touitou, Y. Evidence of Prooxidant and Antioxidant Action of Melatonin on Human Liver Cell Line HepG2. Life Sci. 2000, 68, 387–399. [Google Scholar] [CrossRef]
- Wölfler, A.; Caluba, H.C.; Abuja, P.M.; Dohr, G.; Schauenstein, K.; Liebmann, P.M. Prooxidant Activity of Melatonin Promotes Fas-Induced Cell Death in Human Leukemic Jurkat Cells. FEBS Lett. 2001, 502, 127–131. [Google Scholar] [CrossRef]
- Choutka, J.; Jansari, V.; Hornig, M.; Iwasaki, A. Unexplained Post-Acute Infection Syndromes. Nat. Med. 2022, 28, 911–923. [Google Scholar] [CrossRef] [PubMed]
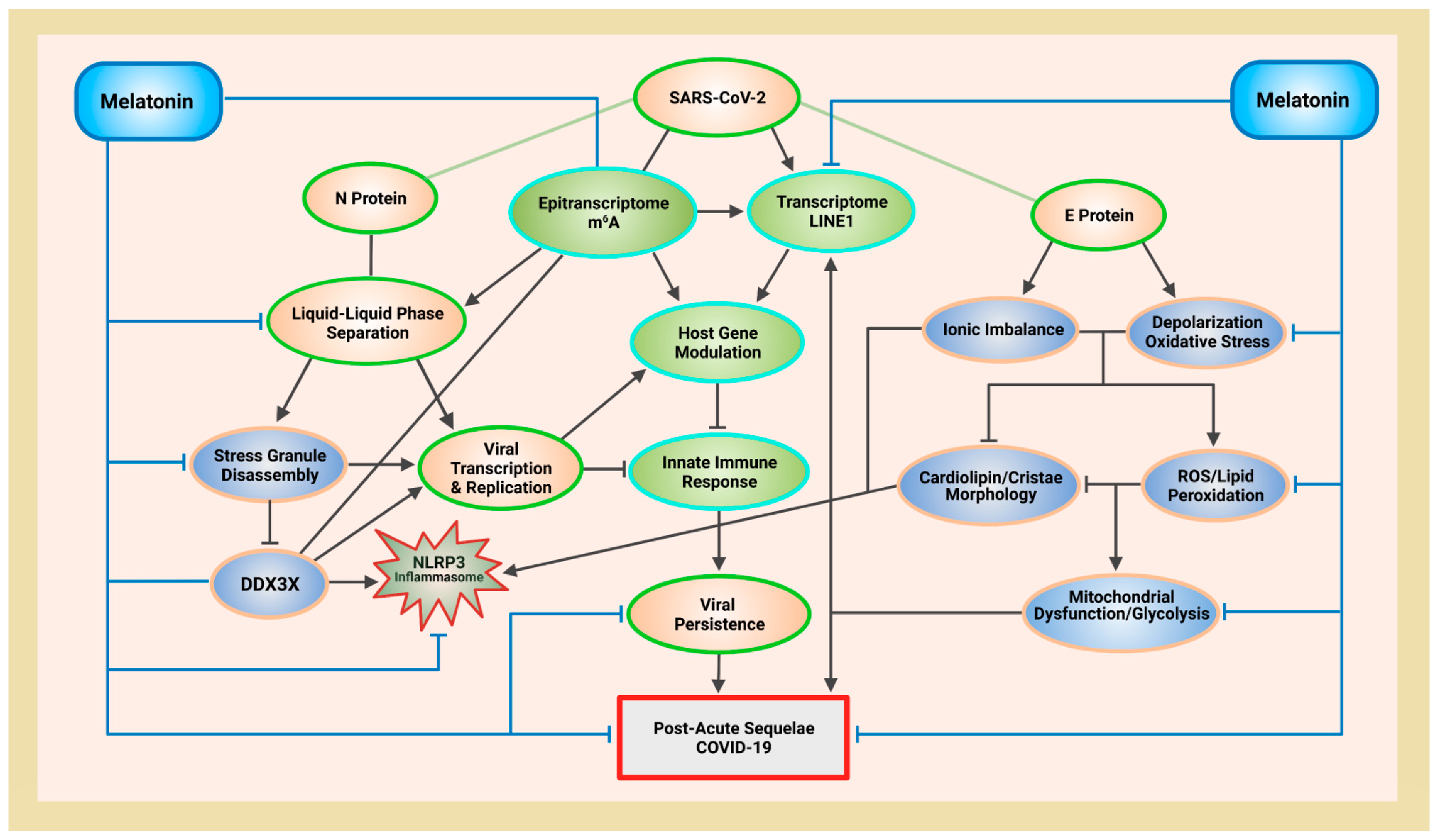
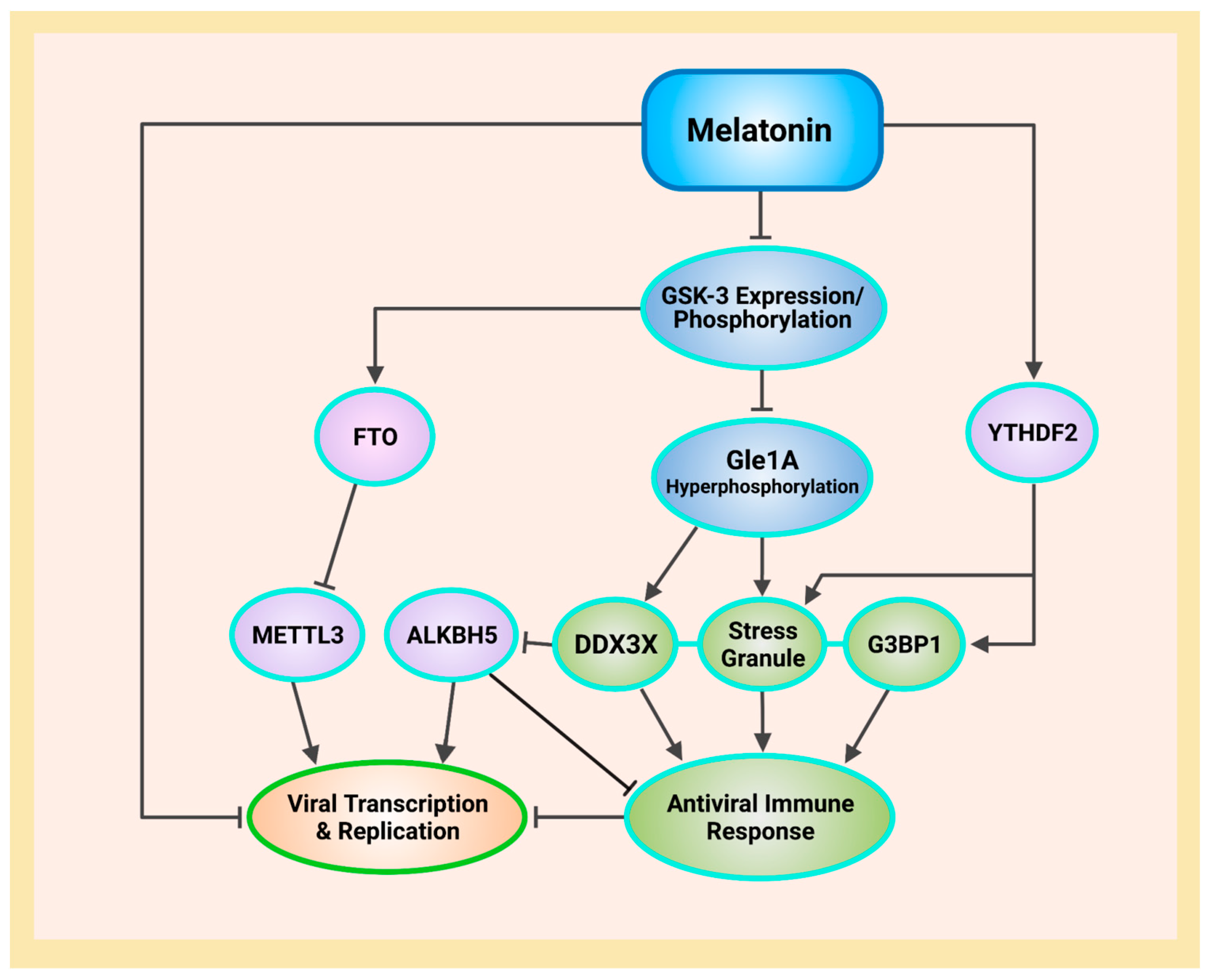
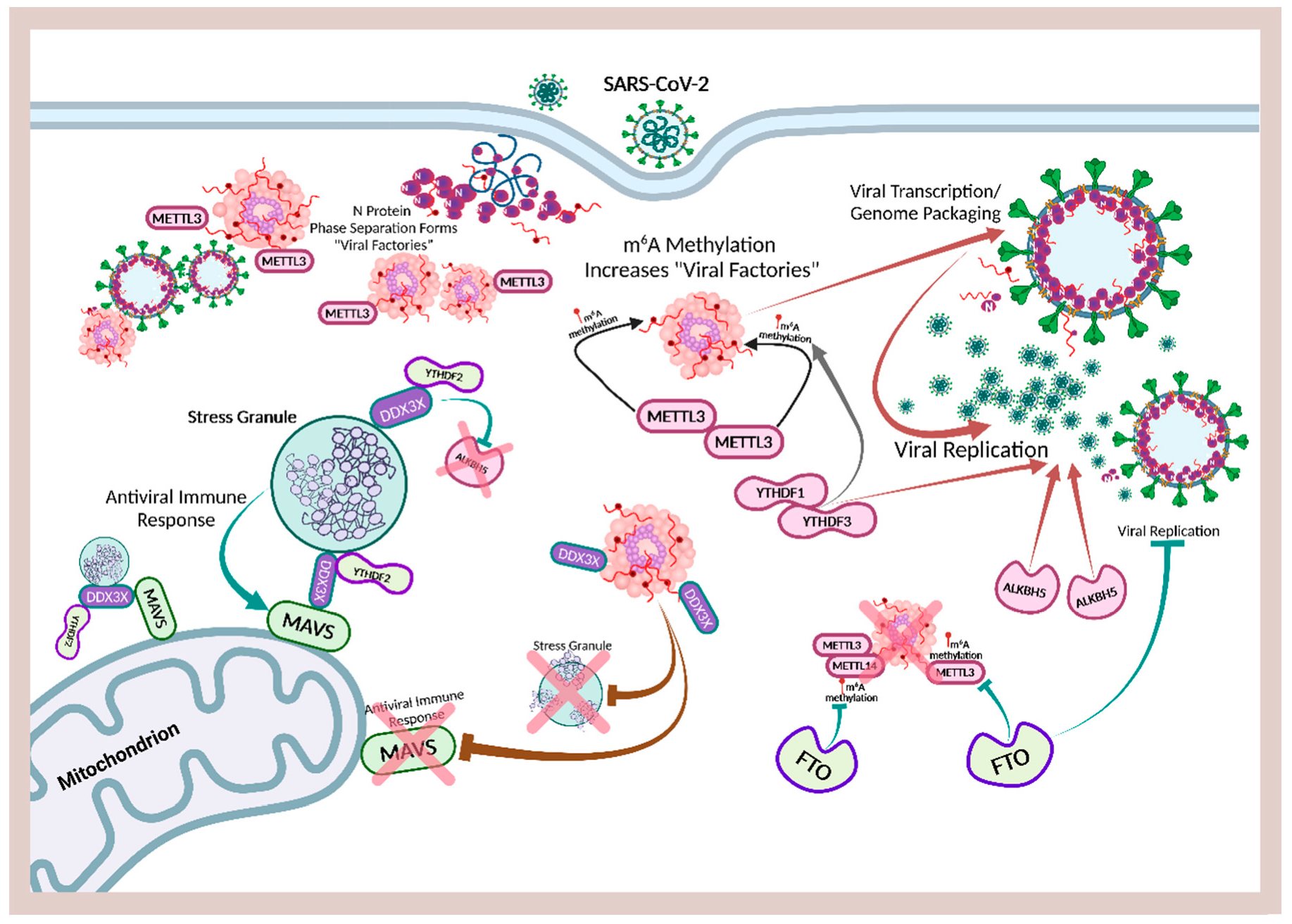
| m6A Modification Enzymes | Cell Line/Extraction Time | Method | Effects | Reference |
|---|---|---|---|---|
| METTL3 | Caco-2/24 hpi | Knockdown | Decreased replication | [251] |
| METTL3/METTL14 | Huh7/72 hpi | Knockdown | Increased replication | [802] |
| YTHDF2 | Huh7/72 hpi | Knockdown | Increased replication | [802] |
| ALKHB5 | Huh7/72 hpi | Knockdown | Decreased replication | [802] |
| FTO | Vero/14 hpi | Knockdown | Demethylase inhibitor dose-dependent interference with viral lifecyle with complete blockage of infection at highest dose. | [810] |
| METTL3 | Vero/24 hpi | Knockdown | Decreased viral titer, N protein copy number and expression | [811] |
| METTL3 | Vero/24 hpi | Overexpression | Elevated m6A modification | [811] |
| FTO | Vero/24 hpi | Knockdown | Increased viral titer, N protein copy number and expression | [811] |
| RBM15 | HuT 78/24 hpi | Knockdown | Inhibited inflammatory gene expression, lymphocyte apoptosis | [812] |
| METTL3 | A549+ACE/48 hpi | Knockdown | Reduced replication, synthesis of viral RNA and N protein | [813] |
| YTHDF1/YTHDF3 | A549+ACE/48 hpi | Knockdown | Reduced replication, synthesis of viral RNA and N protein | [813] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Loh, D.; Reiter, R.J. Melatonin: Regulation of Viral Phase Separation and Epitranscriptomics in Post-Acute Sequelae of COVID-19. Int. J. Mol. Sci. 2022, 23, 8122. https://doi.org/10.3390/ijms23158122
Loh D, Reiter RJ. Melatonin: Regulation of Viral Phase Separation and Epitranscriptomics in Post-Acute Sequelae of COVID-19. International Journal of Molecular Sciences. 2022; 23(15):8122. https://doi.org/10.3390/ijms23158122
Chicago/Turabian StyleLoh, Doris, and Russel J. Reiter. 2022. "Melatonin: Regulation of Viral Phase Separation and Epitranscriptomics in Post-Acute Sequelae of COVID-19" International Journal of Molecular Sciences 23, no. 15: 8122. https://doi.org/10.3390/ijms23158122
APA StyleLoh, D., & Reiter, R. J. (2022). Melatonin: Regulation of Viral Phase Separation and Epitranscriptomics in Post-Acute Sequelae of COVID-19. International Journal of Molecular Sciences, 23(15), 8122. https://doi.org/10.3390/ijms23158122







