Nuclear Functions of KaeA, a Subunit of the KEOPS Complex in Aspergillus nidulans
Abstract
:1. Introduction
2. Results
2.1. kaeA19 and kaeA25 Mutations Do Not Influence the Modification of A37 in tRNA
2.2. kaeA19 and kaeA25 Mutations Affect Expression of Many Genes
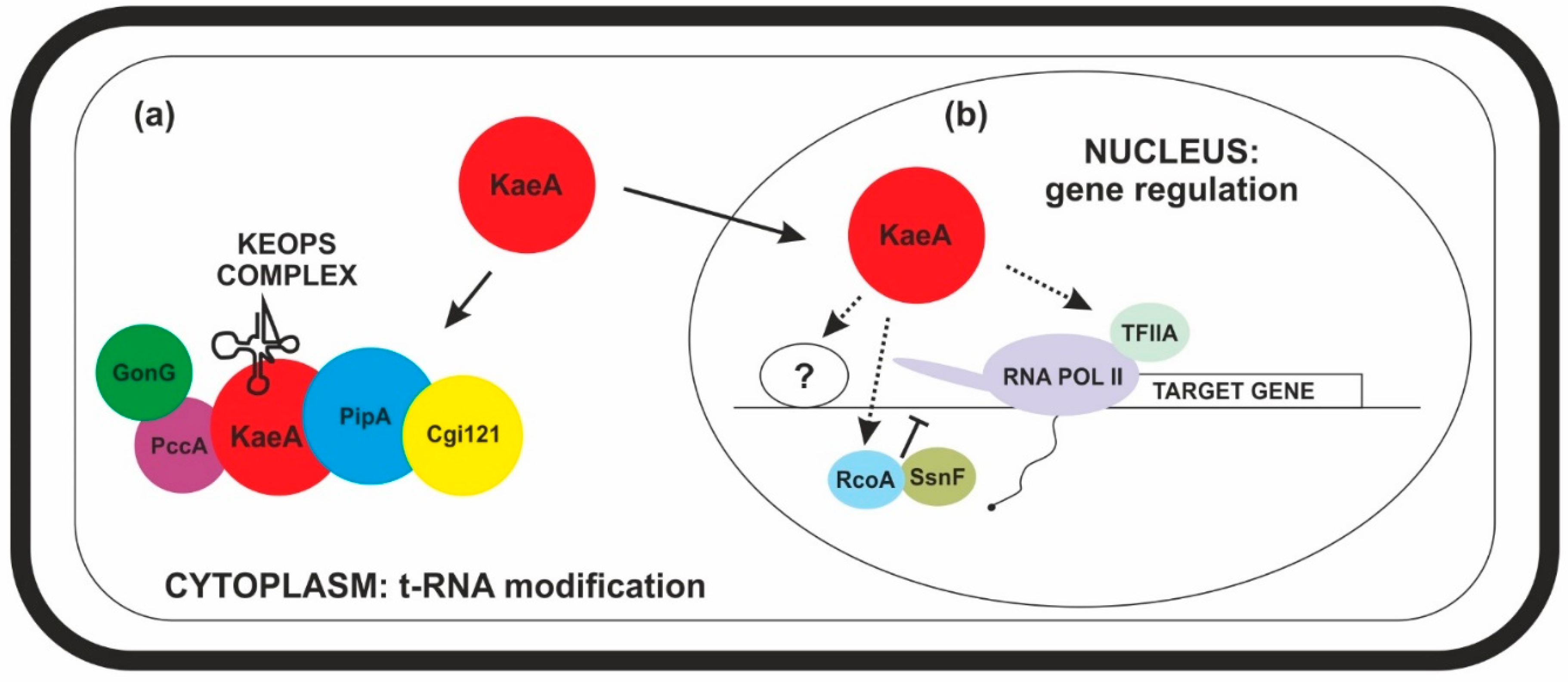
2.3. In Addition to Subunits of KEOPS Complex, KaeA Also Interacts with Other Proteins
2.4. KaeA Interacts with Chromatin
3. Discussion
4. Materials and Methods
4.1. A. nidulans and S. cerevisiae Strains, Growth Conditions, and Transformation
4.2. Transcriptomics
4.3. RT-qPCR Analysis
4.4. Plasmids for Aspergillus Transformation
4.4.1. Golden Gate Cloning
4.4.2. Overlap Extension PCR Cloning
4.5. Purification of Proteins Interacting with KaeA
4.6. Primer Extension
4.7. Western Analysis
4.8. ChIP-Seq Analysis
4.9. Bioinformatics Analysis of KaeA Protein
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Galperin, M.Y. Social Bacteria and Asocial Eukaryotes. Environ. Microbiol. 2008, 10, 281–288. [Google Scholar] [CrossRef] [PubMed]
- Facchin, S.; Lopreiato, R.; Stocchetto, S.; Arrigoni, G.; Cesaro, L.; Marin, O.; Carignani, G.; Pinna, L.A. Structure-Function Analysis of Yeast PiD261/Bud32, an Atypical Protein Kinase Essential for Normal Cell Life. Biochem. J. 2002, 364, 457–463. [Google Scholar] [CrossRef] [PubMed]
- Beenstock, J.; Sicheri, F. The Structural and Functional Workings of KEOPS. Nucleic Acids Res. 2021, 49, 10818–10834. [Google Scholar] [CrossRef] [PubMed]
- Oberto, J.; Breuil, N.; Hecker, A.; Farina, F.; Brochier-Armanet, C.; Culetto, E.; Forterre, P. Qri7/OSGEPL, the Mitochondrial Version of the Universal Kae1/YgjD Protein, Is Essential for Mitochondrial Genome Maintenance. Nucleic Acids Res. 2009, 37, 5343–5352. [Google Scholar] [CrossRef]
- Daugeron, M.C.; Lenstra, T.L.; Frizzarin, M.; el Yacoubi, B.; Liu, X.; Baudin-Baillieu, A.; Lijnzaad, P.; Decourty, L.; Saveanu, C.; Jacquier, A.; et al. Gcn4 Misregulation Reveals a Direct Role for the Evolutionary Conserved EKC/KEOPS in the t 6A Modifi-cation of TRNAs. Nucleic Acids Res. 2011, 39, 6148–6160. [Google Scholar] [CrossRef]
- el Yacoubi, B.; Hatin, I.; Deutsch, C.; Kahveci, T.; Rousset, J.P.; Iwata-Reuyl, D.; G. Murzin, A.; de Crécy-Lagard, V. A Role for the Universal Kae1/Qri7/YgjD (COG0533) Family in TRNA Modification. EMBO J. 2011, 30, 882–893. [Google Scholar] [CrossRef]
- Srinivasan, M.; Mehta, P.; Yu, Y.; Prugar, E.; Koonin, E.V.; Karzai, A.W.; Sternglanz, R. The Highly Conserved KEOPS/EKC Complex Is Essential for a Universal TRNA Modification, T6A. EMBO J. 2011, 30, 873–881. [Google Scholar] [CrossRef]
- Perrochia, L.; Crozat, E.; Hecker, A.; Zhang, W.; Bareille, J.; Collinet, B.; van Tilbeurgh, H.; Forterre, P.; Basta, T. In Vitro Biosynthesis of a Universal T6A TRNA Modification in Archaea and Eukarya. Nucleic Acids Res. 2013, 41, 1953–1964. [Google Scholar] [CrossRef]
- Beenstock, J.; Ona, S.M.; Porat, J.; Orlicky, S.; Wan, L.C.K.; Ceccarelli, D.F.; Maisonneuve, P.; Szilard, R.K.; Yin, Z.; Setiaputra, D.; et al. A Substrate Binding Model for the KEOPS TRNA Modifying Complex. Nat. Commun. 2020, 11, 6233. [Google Scholar] [CrossRef]
- De Crécy-Lagard, V.; Marck, C.; Brochier-Armanet, C.; Grosjean, H. Comparative RNomics and Modomics in Mollicutes: Prediction of Gene Function and Evolutionary Implications. IUBMB Life 2007, 59, 634–658. [Google Scholar] [CrossRef]
- Yarian, C.; Townsend, H.; Czestkowski, W.; Sochacka, E.; Malkiewicz, A.J.; Guenther, R.; Miskiewicz, A.; Agris, P.F. Accurate Translation of the Genetic Code Depends on TRNA Modified Nucleosides. J. Biol. Chem. 2002, 277, 16391–16395. [Google Scholar] [CrossRef] [PubMed]
- El Yacoubi, B.; Lyons, B.; Cruz, Y.; Reddy, R.; Nordin, B.; Agnelli, F.; Williamson, J.R.; Schimmel, P.; Swairjo, M.A.; de Cré-cy-Lagard, V. The Universal YrdC/Sua5 Family Is Required for the Formation of Threonylcarbamoyladenosine in TRNA. Nucleic Acids Res. 2009, 37, 2894–2909. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.C.K.; Maisonneuve, P.; Szilard, R.K.; Lambert, J.P.; Ng, T.F.; Manczyk, N.; Huang, H.; Laister, R.; Caudy, A.A.; Gingras, A.C.; et al. Proteomic Analysis of the Human KEOPS Complex Identifies C14ORF142 as a Core Subunit Homologous to Yeast Gon7. Nucleic Acids Res. 2017, 45, 805–817. [Google Scholar] [CrossRef]
- Downey, M.; Houlsworth, R.; Maringele, L.; Rollie, A.; Brehme, M.; Galicia, S.; Guillard, S.; Partington, M.; Zubko, M.K.; Krogan, N.J.; et al. A Genome-Wide Screen Identifies the Evolutionarily Conserved KEOPS Complex as a Telomere Regulator. Cell 2006, 124, 1155–1168. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Tang, H.B.; Liu, N.N.; Tong, X.J.; Dang, W.; Duan, Y.M.; Fu, X.H.; Zhang, Y.; Peng, J.; Meng, F.L.; et al. Telomerase-Null Survivor Screening Identifies Novel Telomere Recombination Regulators. PLoS Genet. 2013, 9, e1003208. [Google Scholar] [CrossRef]
- Liu, Y.-Y.; He, M.-H.; Liu, J.-C.; Lu, Y.-S.; Peng, J.; Zhou, J.-Q. Yeast KEOPS Complex Regulates Telomere Length Independently of Its T6A Modification Function. J. Genet. Genom. 2018, 45, 247–257. [Google Scholar] [CrossRef]
- Peggion, C.; Lopreiato, R.; Casanova, E.; Ruzzene, M.; Facchin, S.; Pinna, L.A.; Carignani, G.; Sartori, G. Phosphorylation of the Saccharomyces Cerevisiae Grx4p Glutaredoxin by the Bud32p Kinase Unveils a Novel Signaling Pathway Involving Sch9p, a Yeast Member of the Akt/PKB Subfamily. FEBS J. 2008, 275, 5919–5933. [Google Scholar] [CrossRef]
- Kisseleva-Romanova, E.; Lopreiato, R.; Baudin-Baillieu, A.; Rousselle, J.-C.C.; Ilan, L.; Hofmann, K.; Namane, A.; Mann, C.; Libri, D. Yeast Homolog of a Cancer-Testis Antigen Defines a New Transcription Complex. EMBO J. 2006, 25, 3576–3585. [Google Scholar] [CrossRef]
- Costessi, A.; Mahrour, N.; Sharma, V.; Stunnenberg, R.; Stoel, M.A.; Tijchon, E.; Conaway, J.W.; Conaway, R.C.; Stunnenberg, H.G. The Human EKC/KEOPS Complex Is Recruited to Cullin2 Ubiquitin Ligases by the Human Tumour Antigen PRAME. PLoS ONE 2012, 7, e42822. [Google Scholar] [CrossRef]
- Weglenski, P. The Mechanism of Action of Proline Suppressors in Aspergillus Nidulans. J. Gen. Microbiol. 1967, 47, 77–85. [Google Scholar] [CrossRef] [Green Version]
- Bartnik, E.; Weglenski, P. Regulation of Arginine Catabolism in Aspergillus Nidulans. Nature 1974, 250, 590–592. [Google Scholar] [CrossRef]
- Dzikowska, A.; Grzelak, A.; Gawlik, J.; Szewczyk, E.; Mrozek, P.; Borsuk, P.; Koper, M.; Empel, J.; Szczesny, P.; Piłsyk, S.; et al. KAEA (SUDPRO), a Member of the Ubiquitous KEOPS/EKC Protein Complex, Regulates the Arginine Catabolic Pathway and the Expression of Several Other Genes in Aspergillus Nidulans. Gene 2015, 573, 310–320. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Hume, S.L.; Johnson, C.; Thompson, P.; Huang, J.; Gray, J.; Lamb, H.K.; Hawkins, A.R. The Transcription Repressor NmrA Is Subject to Proteolysis by Three Aspergillus Nidulans Proteases. Protein Sci. 2010, 19, 1405–1419. [Google Scholar] [CrossRef]
- Todd, R.B. Regulation of Fungal Nitrogen Metabolism. In Biochemistry and Molecular Biology; Springer International Publishing: Cham, Switzerland, 2016; pp. 281–303. [Google Scholar]
- Berger, H.; Basheer, A.; Böck, S.; Reyes-Dominguez, Y.; Dalik, T.; Altmann, F.; Strauss, J. Dissecting Individual Steps of Nitrogen Transcription Factor Cooperation in the Aspergillus Nidulans Nitrate Cluster. Mol. Microbiol. 2008, 69, 1385–1398. [Google Scholar] [CrossRef]
- Lin, C.-J.; Smibert, P.; Zhao, X.; Hu, J.F.; Ramroop, J.; Kellner, S.M.; Benton, M.A.; Govind, S.; Dedon, P.C.; Sternglanz, R.; et al. An Extensive Allelic Series of Drosophila Kae1 Mutants Reveals Diverse and Tissue-Specific Requirements for T6A Biogenesis. RNA 2015, 21, 2103–2118. [Google Scholar] [CrossRef]
- Andrianopoulos, A.; Kourambas, S.; Sharp, J.A.; Davis, M.A.; Hynes, M.J. Characterization of the Aspergillus Nidulans NmrA Gene Involved in Nitrogen Metabolite Repression. J. Bacteriol. 1998, 180, 1973–1977. [Google Scholar] [CrossRef]
- Lamb, H.K.; Ren, J.; Park, A.; Johnson, C.; Leslie, K.; Cocklin, S.; Thompson, P.; Mee, C.; Cooper, A.; Stammers, D.K.; et al. Modulation of the Ligand Binding Properties of the Transcription Repressor NmrA by GATA-Containing DNA and Site-Directed Mutagenesis. Protein Sci. 2009, 13, 3127–3138. [Google Scholar] [CrossRef]
- Pfannmüller, A.; Leufken, J.; Studt, L.; Michielse, C.B.; Sieber, C.M.K.; Güldener, U.; Hawat, S.; Hippler, M.; Fufezan, C.; Tudzynski, B. Comparative Transcriptome and Proteome Analysis Reveals a Global Impact of the Nitrogen Regulators AreA and AreB on Secondary Metabolism in Fusarium Fujikuroi. PLoS ONE 2017, 12, e0176194. [Google Scholar] [CrossRef]
- Strittmatter, A.W.; Irniger, S.; Braus, G.H. Induction of JlbA MRNA Synthesis for a Putative BZIP Protein of Aspergillus Nidulans by Amino Acid Starvation. Curr. Genet. 2001, 39, 327–334. [Google Scholar] [CrossRef]
- Garcia-Barrio, M. Association of GCN1-GCN20 Regulatory Complex with the N-Terminus of EIF2alpha Kinase GCN2 Is Required for GCN2 Activation. EMBO J. 2000, 19, 1887–1899. [Google Scholar] [CrossRef] [Green Version]
- Bertram, P.G.; Choi, J.H.; Carvalho, J.; Chan, T.-F.; Ai, W.; Zheng, X.F.S. Convergence of TOR-Nitrogen and Snf1-Glucose Signaling Pathways onto Gln3. Mol. Cell. Biol. 2002, 22, 1246–1252. [Google Scholar] [CrossRef]
- Yang, X.J.; Seto, E. The Rpd3/Hda1 Family of Lysine Deacetylases: From Bacteria and Yeast to Mice and Men. Nat. Rev. Mol. Cell Biol. 2008, 9, 206–218. [Google Scholar] [CrossRef]
- Mizuguchi, G.; Shen, X.; Landry, J.; Wu, W.H.; Sen, S.; Wu, C. ATP-Driven Exchange of Histone H2AZ Variant Catalyzed by SWR1 Chromatin Remodeling Complex. Science 2004, 303, 343–348. [Google Scholar] [CrossRef]
- Crespo, J.L.; Powers, T.; Fowler, B.; Hall, M.N. The TOR-Controlled Transcription Activators GLN3, RTG1, and RTG3 Are Regulated in Response to Intracellular Levels of Glutamine. Proc. Natl. Acad. Sci. USA 2002, 99, 6784–6789. [Google Scholar] [CrossRef]
- Ruiz-Roig, C.; Noriega, N.; Duch, A.; Posas, F.; de Nadal, E. The Hog1 SAPK Controls the Rtg1/Rtg3 Transcriptional Complex Activity by Multiple Regulatory Mechanisms. Mol. Biol. Cell 2012, 23, 4286–4296. [Google Scholar] [CrossRef]
- Hoffmann, B.; Valerius, O.; Andermann, M.; Braus, G.H. Transcriptional Autoregulation and Inhibition of MRNA Translation of Amino Acid Regulator Gene CpcA of Filamentous Fungus Aspergillus Nidulans. Mol. Biol. Cell 2001, 12, 2846–2857. [Google Scholar] [CrossRef]
- Hortschansky, P.; Ando, E.; Tuppatsch, K.; Arikawa, H.; Kobayashi, T.; Kato, M.; Haas, H.; Brakhage, A.A. Deciphering the Combinatorial DNA-Binding Code of the CCAAT-Binding Complex and the Iron-Regulatory Basic Region Leucine Zipper (BZIP) Transcription Factor HapX. J. Biol. Chem. 2015, 290, 6058–6070. [Google Scholar] [CrossRef]
- Hortschansky, P.; Eisendle, M.; Al-Abdallah, Q.; Schmidt, A.D.; Bergmann, S.; Thön, M.; Kniemeyer, O.; Abt, B.; Seeber, B.; Werner, E.R.; et al. Interaction of HapX with the CCAAT-Binding Complex—A Novel Mechanism of Gene Regulation by Iron. EMBO J. 2007, 26, 3157–3168. [Google Scholar] [CrossRef] [PubMed]
- Todd, R.B.; Hynes, M.J.; Andrianopoulos, A. The Aspergillus Nidulans RcoA Gene Is Required for VeA-Dependent Sexual Development. Genetics 2006, 174, 1685–1688. [Google Scholar] [CrossRef]
- Hicks, J.; Lockington, R.A.; Strauss, J.; Dieringer, D.; Kubicek, C.P.; Kelly, J.; Keller, N. RcoA Has Pleiotropic Effects on Aspergillus Nidulans Cellular Development. Mol. Microbiol. 2001, 39, 1482–1493. [Google Scholar] [CrossRef]
- García, I.; Mathieu, M.; Nikolaev, I.; Felenbok, B.; Scazzocchio, C. Roles of the Aspergillus Nidulans Homologues of Tup1 and Ssn6 in Chromatin Structure and Cell Viability. FEMS Microbiol. Lett. 2008, 289, 146–154. [Google Scholar] [CrossRef]
- Davie, J.K.; Trumbly, R.J.; Dent, S.Y.R. Histone-Dependent Association of Tup1-Ssn6 with Repressed Genes In Vivo. Mol. Cell. Biol. 2002, 22, 693–703. [Google Scholar] [CrossRef] [PubMed]
- Davie, J.K.; Edmondson, D.G.; Coco, C.B.; Dent, S.Y.R. Tup1-Ssn6 Interacts with Multiple Class I Histone Deacetylases in Vivo. J. Biol. Chem. 2003, 278, 50158–50162. [Google Scholar] [CrossRef] [PubMed]
- Papamichos-Chronakis, M.; Conlan, R.S.; Gounalaki, N.; Copf, T.; Tzamarias, D. Hrs1/Med3 Is a Cyc8-Tup1 Corepressor Target in the RNA Polymerase II Holoenzyme. J. Biol. Chem. 2000, 275, 8397–8403. [Google Scholar] [CrossRef]
- Bayram, Ö.; Krappmann, S.; Ni, M.; Jin, W.B.; Helmstaedt, K.; Valerius, O.; Braus-Stromeyer, S.; Kwon, N.J.; Keller, N.P.; Yu, J.H.; et al. VelB/VeA/LaeA Complex Coordinates Light Signal with Fungal Development and Secondary Metabolism. Science 2008, 320, 1504–1506. [Google Scholar] [CrossRef] [PubMed]
- Kempf, C.; Bathe, F.; Fischer, R. Evidence That Two Pcl-like Cyclins Control Cdk9 Activity during Cell Differentiation in Aspergillus Nidulans Asexual Development. Eukaryot. Cell 2013, 12, 23–36. [Google Scholar] [CrossRef]
- Welkenhuysen, N.; Borgqvist, J.; Backman, M.; Bendrioua, L.; Goksör, M.; Adiels, C.B.; Cvijovic, M.; Hohmann, S. Single-Cell Study Links Metabolism with Nutrient Signaling and Reveals Sources of Variability. BMC Syst. Biol. 2017, 11, s12918–s017. [Google Scholar] [CrossRef]
- Todd, R.B.; Fraser, J.A.; Wong, K.H.; Davis, M.A.; Hynes, M.J. Nuclear Accumulation of the GATA Factor AreA in Response to Complete Nitrogen Starvation by Regulation of Nuclear Export. Eukaryot. Cell 2005, 4, 1646–1653. [Google Scholar] [CrossRef]
- Nayak, T.; Szewczyk, E.; Oakley, C.E.; Osmani, A.; Ukil, L.; Murray, S.L.; Hynes, M.J.; Osmani, S.A.; Oakley, B.R. A Versatile and Efficient Gene-Targeting System for Aspergillus Nidulans. Genetics 2006, 172, 1557–1566. [Google Scholar] [CrossRef]
- Schmitt, M.E.; Brown, T.A.; Trumpower, B.L. A Rapid and Simple Method for Preparation of RNA from Saccharomyces Cerevisiae. Nucleic Acids Res. 1990, 18, 3091–3092. [Google Scholar] [CrossRef] [Green Version]
- Trapnell, C.; Pachter, L.; Salzberg, S.L. TopHat: Discovering Splice Junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast Gapped-Read Alignment with Bowtie 2. Nat. Methods 2012, 9, 1–24. [Google Scholar] [CrossRef]
- Morgan, M.; Anders, S.; Lawrence, M.; Aboyoun, P.; Pagès, H.; Gentleman, R. ShortRead: A Bioconductor Package for Input, Quality Assessment and Exploration of High-Throughput Sequence Data. Bioinformatics 2009, 25, 2607–2608. [Google Scholar] [CrossRef] [PubMed]
- Seabold, S.; Perktold, J. Statsmodels: Econometric and Statistical Modeling with Python. In Proceedings of the 9th Python in Science Conference, Austin, TX, USA, 28 June–3 July 2010. [Google Scholar]
- Robinson Mark, D.; McCarthy Davis, J.; Smyth Gordon, K. EdgeR: A Bioconductor Package for Differential Expression Analysis of Digital Gene Expression Data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Robinson, M.D.; Oshlack, A. A Scaling Normalization Method for Differential Expression Analysis of RNA-Seq Data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef]
- Cerqueira, G.C.; Arnaud, M.B.; Inglis, D.O.; Skrzypek, M.S.; Binkley, G.; Simison, M.; Miyasato, S.R.; Binkley, J.; Orvis, J.; Shah, P.; et al. The Aspergillus Genome Database: Multispecies Curation and Incorporation of RNA-Seq Data to Improve Structural Gene Annotations. Nucleic Acids Res. 2014, 42, D705–D710. [Google Scholar] [CrossRef]
- Stajich, J.E.; Harris, T.; Brunk, B.P.; Brestelli, J.; Fischer, S.; Harb, O.S.; Kissinger, J.C.; Li, W.; Nayak, V.; Pinney, D.F.; et al. FungiDB: An Integrated Functional Genomics Database for Fungi. Nucleic Acids Res. 2012, 40, D675–D681. [Google Scholar] [CrossRef]
- Basenko, E.Y.; Pulman, J.A.; Shanmugasundram, A.; Harb, O.S.; Crouch, K.; Starns, D.; Warrenfeltz, S.; Aurrecoechea, C.; Stoeckert, C.J.; Kissinger, J.C.; et al. FungiDB: An Integrated Bioinformatic Resource for Fungi and Oomycetes. J. Fungi 2018, 4, 39. [Google Scholar] [CrossRef]
- Cherry, J.M.; Hong, E.L.; Amundsen, C.; Balakrishnan, R.; Binkley, G.; Chan, E.T.; Christie, K.R.; Costanzo, M.C.; Dwight, S.S.; Engel, S.R.; et al. Saccharomyces Genome Database: The Genomics Resource of Budding Yeast. Nucleic Acids Res. 2012, 40, D700–D705. [Google Scholar] [CrossRef]
- Edgar, R.; Domrachev, M.; Lash, A.E. The Gene Expression Omnibus: NCBI Gene Expression and Hybridization Array Data Repository. Nucleic Acids Res. 2002, 30, 207–210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Engler, C.; Kandzia, R.; Marillonnet, S. A One Pot, One Step, Precision Cloning Method with High Throughput Capability. PLoS ONE 2008, 3, e3647. [Google Scholar] [CrossRef]
- Bryksin, A.V.; Matsumura, I. Overlap Extension PCR Cloning: A Simple and Reliable Way to Create Recombinant Plasmids. Biotechniques 2010, 48, 463–465. [Google Scholar] [CrossRef]
- Perkins, D.N.; Pappin, D.J.C.; Creasy, D.M.; Cottrell, J.S. Probability-based Protein Identification by Searching Sequence Databases Using Mass Spectrometry Data-Perkins-1999-ELECTROPHORESIS-Wiley Online Library. Electrophoresis 1999, 20, 3551–3567. [Google Scholar] [CrossRef]
- Deutsch, E.W.; Bandeira, N.; Sharma, V.; Perez-Riverol, Y.; Carver, J.J.; Kundu, D.J.; García-Seisdedos, D.; Jarnuczak, A.F.; Hewapathirana, S.; Pullman, B.S.; et al. The ProteomeXchange Consortium in 2020: Enabling “big Data” Approaches in Proteomics. Nucleic Acids Res. 2020, 48, D1145–D1152. [Google Scholar] [CrossRef]
- Perez-Riverol, Y.; Bai, J.; Bandla, C.; García-Seisdedos, D.; Hewapathirana, S.; Kamatchinathan, S.; Kundu, D.J.; Prakash, A.; Frericks-Zipper, A.; Eisenacher, M.; et al. The PRIDE Database Resources in 2022: A Hub for Mass Spectrometry-Based Proteomics evidences. Nucleic Acids Res. 2022, 50, D543–D552. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2001; ISBN 9780879695774. [Google Scholar]
- Boedi, S.; Reyes-Dominguez, Y.; Strauss, J. Chromatin Immunoprecipitation Analysis in Filamentous Fungi. Methods Mol. Biol. 2012, 944, 221–236. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and Accurate Long-Read Alignment with Burrows-Wheeler Transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef]
- Yates, A.D.; Allen, J.; Amode, R.M.; Azov, A.G.; Barba, M.; Becerra, A.; Bhai, J.; Campbell, L.I.; Carbajo Martinez, M.; Chakiachvili, M.; et al. Ensembl Genomes 2022: An Expanding Genome Resource for Non-Vertebrates. Nucleic Acids Res. 2022, 50, D996–D1003. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017; p. 1. [Google Scholar]
- Zhang, Y.; Liu, T.; Meyer, C.A.; Eeckhoute, J.; Johnson, D.S.; Bernstein, B.E.; Nussbaum, C.; Myers, R.M.; Brown, M.; Li, W.; et al. Model-Based Analysis of ChIP-Seq (MACS). Genome Biol. 2008, 9, R137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quevillon, E.; Silventoinen, V.; Pillai, S.; Harte, N.; Mulder, N.; Apweiler, R.; Lopez, R. InterProScan: Protein Domains Identifier. Nucleic Acids Res. 2005, 33, W116–W120. [Google Scholar] [CrossRef]
- Thumuluri, V.; Almagro Armenteros, J.J.; Johansen, A.R.; Nielsen, H.; Winther, O. DeepLoc 2.0: Multi-Label Subcellular Localization Prediction Using Protein Language Models. Nucleic Acids Res. 2022, 50, W228–W234. [Google Scholar] [CrossRef]
- Blom, N.; Gammeltoft, S.; Brunak, S. Sequence and Structure-Based Prediction of Eukaryotic Protein Phosphorylation Sites. J. Mol. Biol. 1999, 294, 1351–1362. [Google Scholar] [CrossRef]
- Blom, N.; Sicheritz-Pontén, T.; Gupta, R.; Gammeltoft, S.; Brunak, S. Prediction of Post-Translational Glycosylation and Phosphorylation of Proteins from the Amino Acid Sequence. Proteomics 2004, 4, 1633–1649. [Google Scholar] [CrossRef]


| Gene AN Number | Gene Name | Yeast Homologue Gene Name 1 | Protein Function 2 |
|---|---|---|---|
| AN5170 | rosA | UME6 | Putative Zn(II)2Cys6 transcription factor; negative regulator of sexual development. Homologue of yeast Ume6p, encoding histone Rpd3p deacetylase subunit, general regulator of chromatin structure and transcriptional repressor. |
| AN1848 | nosA | UME6 | Putative Zn(II)2Cys6 transcription factor; negative regulator of sexual development. Homologue of yeast Ume6p, encoding histone Rpd3p deacetylase subunit, general regulator of chromatin structure and transcriptional repressor. |
| AN1812 | jlbA | – | bZIP transcription factor involved in amino acid biosynthesis. |
| AN8168 | nmrA | – | Co-repressor of general nitrogen regulator AreA |
| AN0948 | – | GCN20 | Protein with predicted ATPase activity. Homologue of yeast GCN20p, positive regulator of Gcn2p kinase; forms a complex with Gcn1p; proposed to stimulate Gcn2p activation by an uncharged tRNA. |
| AN1298 | – | RTG3 | Homologue of yeast Rtg3p, bHLH/Zip transcription factor for retrograde (RTG) and TOR pathways. |
| AN6076 | – | SWR1 | Protein with helicase domain of SNF2 type. Homologue of yeast Swr1p, Swi2/Snf2-related ATPase; catalytic subunit of SWR1 complex, which exchanges histone variant H2AZ (Htz1p) for chromatin-bound histone H2A. |
| AN10854 | – | SNF4 | Putative serine-threonine protein kinase. Homologue of yeast Snf4p, activating subunit of AMP-activated Snf1p kinase complex; activates glucose-repressed genes, represses glucose-induced genes. |
| AN2366 | – | – | Putative trypsin-like protease involved in proteolytic cleavage of NmrA, a regulatory protein involved in repression of nitrogen metabolism. |
| Gene AN Number | Gene Name | Protein Function (from FungiDB) | Yeast Homologue Gene Name | Protein Function of Yeast Homologue (from SGD) | DNAse * +/− |
|---|---|---|---|---|---|
| AN2845 | pccA | Putative component of EKC/KEOPS complex | PCC1 | Component of KEOPS complex | + |
| AN2513 | pipA | Putative component of EKC/KEOPS complex | BUD32 | Protein kinase, component of KEOPS complex | + |
| AN11910 | cgi121 | Putative component of EKC/KEOPS complex | CGI121 | Component of KEOPS complex | − |
| AN11901 | gonG | Putative component of EKC/KEOPS complex | GON7 | Component of KEOPS complex | + |
| AN6505 | rcoA | WD40 repeat protein | TUP1 | Chromatin-silencing transcriptional regulator; forms complex with Cyc8p, involved in establishment of repressive chromatin structure | − |
| AN12054 | ssnF | Orthologue of S. cerevisiae Ssn6p | CYC8 | Chromatin-silencing transcriptional regulator; acts together with Tup1p | − |
| AN2181 | – | Orthologue(s) have RNA polymerase II transcription factor activity, TBP-class protein binding | TOA2 | TFIIA small subunit; involved in transcriptional activation, acts as anti-repressor or co-activator | + |
| AN2412 | cmkA | Calcium/calmodulin-dependent protein kinase A | CMK2 | Calmodulin-dependent protein kinase | − |
| AN6305 | pkaA | cAMP-dependent protein kinase catalytic subunit | TPK2 | Catalytic subunit of cAMP-dependent protein kinase | + |
| AN4987 | pkaR | Putative protein kinase A (PKA) regulatory subunit | BCY1 | Regulatory subunit of cyclic AMP-dependent protein kinase PKA | + |
| AN5973 | pkcB | Protein with similarity to protein kinase C | YPK1 | Serine/threonine protein kinase | + |
| AN1545 | – | Putative regulatory subunit of protein phosphatase 2A (PP2A) | CDC55 | Non-essential regulatory subunit B of protein phosphatase 2A (PP2A) | + |
| Gene AN Number | Gene Name | Protein Function (from FungiDB) | Yeast Homologue Gene Name * | Protein Function of Yeast Homologue (from SGD) | Chromatin Region Interacting with KaeA |
|---|---|---|---|---|---|
| AN0733 | hhtA | Histone H3 | HHT1 | Histone H3; core histone protein required for chromatin assembly, part of heterochromatin-mediated telomeric and HM silencing | 5′ UTR |
| AN0734 | H4.1 | Histone H4.1 | HHF2 | Histone H4; core histone protein required for chromatin assembly and chromosome function; contributes to telomeric silencing | 5′ UTR |
| AN3468 | H2A.X | Histone H2A | HTA1 | Histone H2A; core histone protein required for chromatin assembly and chromosome function | 5′ UTR |
| AN1007 | niiA | Putative nitrite reductase with predicted role in nitrogen metabolism | – | – | Promoter region |
| AN2436 | aclB | Putative ATP citrate synthase with predicted role in TCA cycle | – | – | 5′ UTR |
| AN2435 | aclA | Putative ATP citrate synthase with predicted role in TCA cycle | – | – | 5′ UTR |
| AN3675 | cpcA | Transcription factor of c-Jun family of Gcn4p-like transcriptional activators | GCN4 | bZIP transcriptional activator of amino acid biosynthetic genes; responds to amino acid starvation | 5′ UTR |
| AN4159 | glnA | Putative ammonium glutamate ligase with predicted role in glutamate and glutamine metabolism | GLN1 | Glutamine synthetase (GS); synthesizes glutamine from glutamate and ammonia; with Glt1p, forms secondary pathway for glutamate biosynthesis from ammonia | 5′ UTR |
| AN4376 | gdhA | Putative glutamate dehydrogenase associated with NADP | GDH1 | NADP(+)-dependent glutamate dehydrogenase | 5′ UTR |
| AN5823 | sidA | L-ornithine N5-monooxygenase | – | – | 5′ UTR |
| AN7463 | meaA | Main ammonium transporter in A. nidulans | MEP3 | Ammonium permease with high capacity and low affinity | 5′ UTR |
| AN8041 | gpdA | Glyceraldehyde-3-phosphate dehydrogenase with predicted role in gluconeogenesis and glycolysis | TDH3 | Glyceraldehyde-3-phosphate dehydrogenase (GAPDH), isozyme 3; involved in glycolysis and gluconeogenesis | 5′UTR |
| AN8251 | hapX | bZIP transcription factor | – | – | exon |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gawlik, J.; Koper, M.; Bogdanowicz, A.; Weglenski, P.; Dzikowska, A. Nuclear Functions of KaeA, a Subunit of the KEOPS Complex in Aspergillus nidulans. Int. J. Mol. Sci. 2022, 23, 11138. https://doi.org/10.3390/ijms231911138
Gawlik J, Koper M, Bogdanowicz A, Weglenski P, Dzikowska A. Nuclear Functions of KaeA, a Subunit of the KEOPS Complex in Aspergillus nidulans. International Journal of Molecular Sciences. 2022; 23(19):11138. https://doi.org/10.3390/ijms231911138
Chicago/Turabian StyleGawlik, Joanna, Michal Koper, Albert Bogdanowicz, Piotr Weglenski, and Agnieszka Dzikowska. 2022. "Nuclear Functions of KaeA, a Subunit of the KEOPS Complex in Aspergillus nidulans" International Journal of Molecular Sciences 23, no. 19: 11138. https://doi.org/10.3390/ijms231911138






