QTL Mapping of Palmitic Acid Content Using Specific-Locus Amplified Fragment Sequencing (SLAF-Seq) Genotyping in Soybeans (Glycine max L.)
Abstract
:1. Introduction
2. Results
2.1. Phenotypic Evaluation
2.2. SLAF Library Construction and Genotyping of the RIL Population
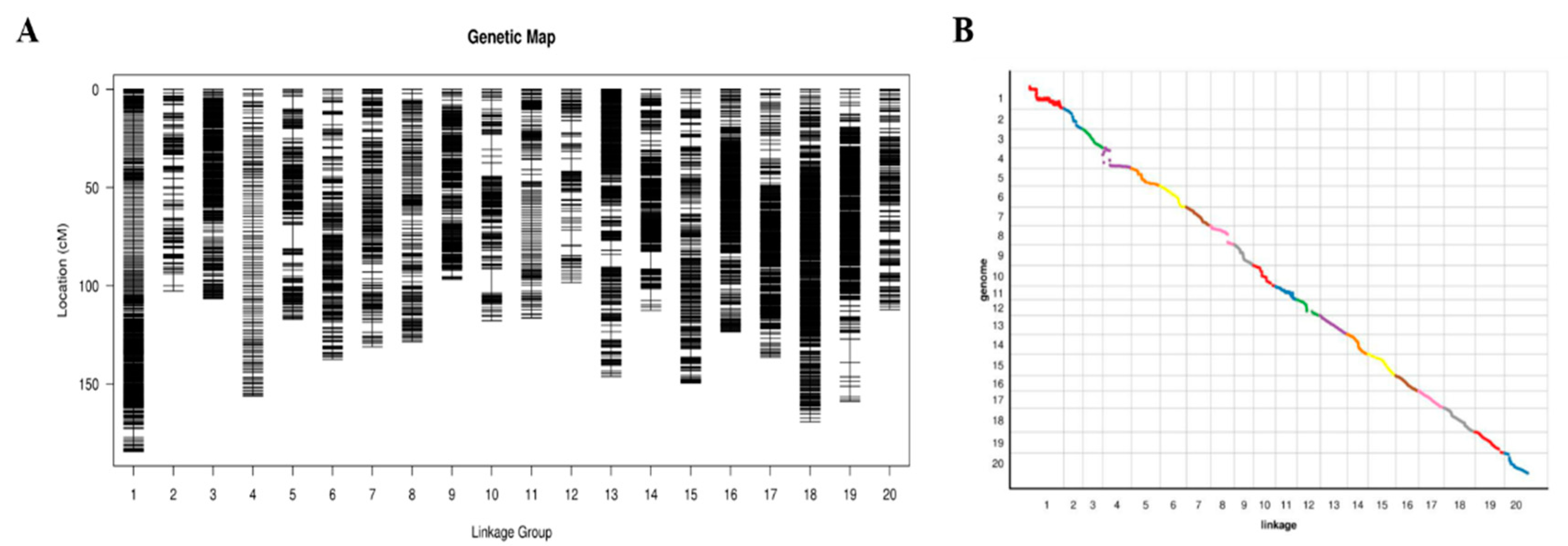
2.3. Construction of the Soybean Genetic Map
2.4. QTL Mapping of the RIL Population
2.5. Verification of Candidate Region and Screening of Linkage Markers
2.6. Analysis of Genome and Transcriptome
3. Discussion
3.1. Construction of the High-Density Genetic Map
3.2. QTL Mapping of Palmitic Acid Content
3.3. Analysis of Candidate Genes
4. Materials and Methods
4.1. Plant Material and Phenotyping
4.2. Genotype Analysis, Construction, and Genotyping of the SLAF Library, and Construction of the High-Density Genetic Map
4.3. QTL Analysis for Palmitic Acid Content
4.4. Validation of Candidate Region and Development of Linkage Marker
4.5. Re-Sequencing and RNA-Seq Analysis of Parents
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Crapiste, G.H.; Brevedan, M.I.; Carelli, A.A. Oxidation of sunflower oil during storage. J. Am. Oil Chem. Soc. 1999, 76, 1437. [Google Scholar] [CrossRef]
- Sala-Vila1, A.; Ros, E. Mounting evidence that increased consumption of a. linolenic acid, the vegetablen. 3 fatty acid, may benefit cardiovascular health. Clin. Lipidol. 2011, 6, 365–369. [Google Scholar] [CrossRef]
- Fan, S.; Li, B.; Yu, F.; Han, F.; Yan, S.; Wang, L.; Sun, J. Analysis of additive and epistatic quantitative trait loci underlying fatty acid concentrations in soybean seeds across multiple environments. Euphytica 2015, 206, 689–700. [Google Scholar] [CrossRef]
- Abdelghany, A.M.; Zhang, S.; Azam, M.; Shaibu, A.S.; Feng, Y.; Li, Y.; Tian, Y.; Hong, H.; Li, B.; Sun, J. Profiling of seed fatty acid composition in 1025 Chinese soybean accessions from diverse ecoregions. Crop. J. 2020, 8, 635–644. [Google Scholar] [CrossRef]
- Panthee, D.; Pantalone, V.; Saxton, A. Modifier QTL for fatty acid composition in soybean oil. Euphytica 2006, 152, 67–73. [Google Scholar] [CrossRef]
- Hu, F.B.; Stampfer, M.J.; Manson, J.E.; Rimm, E.; Colditz, G.A.; Rosner, B.A.; Hennekens, C.H.; Willett, W.C. Dietary fat intake and the risk of coronary heart disease in women. N. Engl. J. Med. 1997, 337, 1491–1499. [Google Scholar] [CrossRef] [PubMed]
- Thelen, J.J.; Ohlrogge, J.B. Metabolic engineering of fatty acid biosynthesis in plants. Metab. Eng. 2002, 4, 12–21. [Google Scholar] [CrossRef]
- Canakci, M.; Monyem, A.; Van Gerpen, J. Accelerated oxidation processes in biodiesel. Trans. ASAE 1999, 42, 1565. [Google Scholar] [CrossRef]
- Graef, G.; LaVallee, B.J.; Tenopir, P.; Tat, M.; Schweiger, B.; Kinney, A.J.; Van Gerpen, J.H.; Clemente, T.E. A high-oleic-acid and low-palmitic-acid soybean: Agronomic performance and evaluation as a feedstock for biodiesel. Plant Biotechnol. J. 2009, 7, 411–421. [Google Scholar] [CrossRef]
- Zhou, Z.; Lakhssassi, N.; Knizia, D.; Cullen, M.A.; El Baz, A.; Embaby, M.G.; Liu, S.; Badad, O.; Vuong, T.D.; AbuGhazaleh, A.; et al. Genome-wide identification and analysis of soybean acyl-ACP thioesterase gene family reveals the role of GmFAT to improve fatty acid composition in soybean seed. Theor. Appl. Genet. 2021, 134, 3611–3623. [Google Scholar] [CrossRef]
- Fehr, W.R. Breeding for modified fatty acid composition in soybean. Crop Sci. 2007, 47, S72–S87. [Google Scholar] [CrossRef]
- Pham, A.-T.; Bilyeu, K.; Chen, P.; Boerma, H.R.; Li, Z. Characterization of the fan1 locus in soybean line A5 and development of molecular assays for high-throughput genotyping of FAD3 genes. Mol. Breed. 2014, 33, 895–907. [Google Scholar] [CrossRef]
- Pazdernik, D.L.; Killam, A.S.; Orf, J.H. Analysis of amino and fatty acid composition in soybean seed, using near infrared reflectance spectroscopy. Agron. J. 1997, 89, 679–685. [Google Scholar] [CrossRef]
- Naeve, S.L.; Proulx, R.A.; Hulke, B.S.; O’Neill, T.A. Sample size and heterogeneity effects on the analysis of whole soybean seed using near infrared spectroscopy. Agron. J. 2008, 100, 231–234. [Google Scholar] [CrossRef]
- Ambrose, A.; Lohumi, S.; Lee, W.-H.; Cho, B.K. Comparative nondestructive measurement of corn seed viability using Fourier transform near-infrared (FT-NIR) and Raman spectroscopy. Sens. Actuators B 2016, 224, 500–506. [Google Scholar] [CrossRef]
- Karn, A.; Heim, C.; Flint-Garcia, S.; Bilyeu, K.; Gillman, J. Development of rigorous fatty acid near-infrared spectroscopy quantitation methods in support of soybean oil improvement. J. Am. Oil Chem. Soc. 2017, 94, 69–76. [Google Scholar] [CrossRef]
- Rebetzke, G.; Pantalone, V.; Burton, J.; Carter, T.; Wilson, R. Genetic background and environment influence palmitate content of soybean seed oil. Crop Sci. 2001, 41, 1731–1736. [Google Scholar] [CrossRef]
- Song, W.; Yang, R.; Wu, T.; Wu, C.; Sun, S.; Zhang, S.; Jiang, B.; Tian, S.; Liu, X.; Han, T. Analyzing the effects of climate factors on soybean protein, oil contents, and composition by extensive and high-density sampling in China. J. Agric. Food Chem. 2016, 64, 4121–4130. [Google Scholar] [CrossRef] [PubMed]
- Cardinal, A.J.; Dewey, R.E.; Burton, J.W. Estimating the individual effects of the reduced palmitic acid fapnc and fap1 alleles on agronomic traits in two soybean populations. Crop Sci. 2008, 48, 633–639. [Google Scholar] [CrossRef]
- Cardinal, A.J.; Whetten, R.; Wang, S.; Auclair, J.; Hyten, D.; Cregan, P.; Bachlava, E.; Gillman, J.; Ramirez, M.; Dewey, R.; et al. Mapping the low palmitate fap1 mutation and validation of its effects in soybean oil and agronomic traits in three soybean populations. Theor. Appl. Genet. 2014, 127, 97–111. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Vries, B.D.; Fehr, W.R.; Welke, G.A.; Dewey, R.E. Molecular characterization of the mutant fap3 (A22) allele for reduced palmitate concentration in soybean. Crop Sci. 2011, 51, 1611–1616. [Google Scholar] [CrossRef]
- Cardinal, A.J.; Burton, J.W.; Camacho-Roger, A.M.; Yang, J.H.; Wilson, R.F.; Dewey, R.E. Molecular analysis of soybean lines with low palmitic acid content in the seed oil. Crop Sci. 2007, 47, 304–310. [Google Scholar] [CrossRef]
- Buhr, T.; Sato, S.; Ebrahim, F.; Xing, A.; Zhou, Y.; Mathiesen, M.; Schweiger, B.; Kinney, A.; Staswick, P.; Clemente, T. Ribozyme termination of RNA transcripts down-regulate seed fatty acid genes in transgenic soybean. Plant J. 2002, 30, 155–163. [Google Scholar] [CrossRef]
- Li, B.; Fan, S.; Yu, F.; Chen, Y.; Zhang, S.; Han, F.; Yan, S.; Wang, L.; Sun, J. High-resolution mapping of QTL for fatty acid composition in soybean using specific-locus amplified fragment sequencing. Theor. Appl. Genet. 2017, 130, 1467–1479. [Google Scholar] [CrossRef]
- Schmutz, J.; Cannon, S.B.; Schlueter, J.; Ma, J.; Mitros, T.; Nelson, W.; Hyten, D.L.; Song, Q.; Thelen, J.J.; Cheng, J.; et al. Genome sequence of the palaeopolyploid soybean. Nature 2010, 463, 178–183. [Google Scholar] [CrossRef] [PubMed]
- Soranzo, N.; Spector, T.D.; Mangino, M.; Kühnel, B.; Rendon, A.; Teumer, A.; Willenborg, C.; Wright, B.; Chen, L.; Li, M.; et al. A genome-wide meta-analysis identifies 22 loci associated with eight hematological parameters in the HaemGen consortium. Nat. Genet. 2009, 41, 1182–1190. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Liu, M.; Wang, Y.; Li, X.; Cheng, S.; Shu, L.; Yu, Z.; Kong, J.; Zhao, T.; Gai, J. Characterizing two inter-specific bin maps for the exploration of the QTLs/genes that confer three soybean evolutionary traits. Front. Plant Sci. 2016, 7, 1248. [Google Scholar] [CrossRef]
- Cao, Y.; Li, S.; Wang, Z.; Chang, F.; Kong, J.; Gai, J.; Zhao, T. Identification of major quantitative trait loci for seed oil content in soybeans by combining linkage and genome-wide association mapping. Front. Plant Sci. 2017, 8, 1222. [Google Scholar] [CrossRef]
- Liu, N.; Li, M.; Hu, X.; Ma, Q.; Mu, Y.; Tan, Z.; Xia, Q.; Zhang, G.; Nian, H. Construction of high-density genetic map and QTL mapping of yield-related and two quality traits in soybean RILs population by RAD-sequencing. BMC Genom. 2017, 18, 1–13. [Google Scholar] [CrossRef]
- Kong, L.; Lu, S.; Wang, Y.; Fang, C.; Wang, F.; Nan, H.; Su, T.; Li, S.; Zhang, F.; Li, X.; et al. Quantitative trait locus mapping of flowering time and maturity in soybean using next-generation sequencing-based analysis. Front. Plant Sci. 2018, 9, 995. [Google Scholar] [CrossRef] [Green Version]
- Watanabe, S.; Shimizu, T.; Machita, K.; Tsubokura, Y.; Xia, Z.; Yamada, T.; Hajika, M.; Ishimoto, M.; Katayose, Y.; Harada, K.; et al. Development of a high-density linkage map and chromosome segment substitution lines for Japanese soybean cultivar Enrei. DNA Res. 2018, 25, 123–136. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Han, D.; Guo, Y.; Yan, H.; Wei, Z.; Tian, Y.; Qiu, L. QTL mapping pod dehiscence resistance in soybean (Glycine max L. Merr.) using specific-locus amplified fragment sequencing. Theor. Appl. Genet. 2019, 132, 2253–2272. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Dong, Y.; Wu, H.; Hu, B.; Zhai, H.; Yang, J.; Xia, Z. Positional cloning of the flowering time QTL qFT12-1 reveals the link between the clock related PRR homolog with photoperiodic response in soybeans. Front. Plant Sci. 2019, 10, 1303. [Google Scholar] [CrossRef] [PubMed]
- Qi, Z.; Huang, L.; Zhu, R.; Xin, D.; Liu, C.; Han, X.; Jiang, H.; Hong, W.; Hu, G.; Zheng, H.; et al. A high-density genetic map for soybean based on specific length amplified fragment sequencing. PLoS ONE 2014, 9, e104871. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Cheng, Y.; Ma, Z.; Liu, X.; Ma, Q.; Xia, Q.; Zhang, G.; Mu, Y.; Nian, H. Fine-mapping of QTLs for individual and total isoflavone content in soybean (Glycine max L.) using a high-density genetic map. Theor. Appl. Genet. 2018, 131, 555–568. [Google Scholar] [CrossRef] [PubMed]
- Patil, G.; Vuong, T.D.; Kale, S.; Valliyodan, B.; Deshmukh, R.; Zhu, C.; Wu, X.; Bai, Y.; Yungbluth, D.; Lu, F. Dissecting genomic hotspots underlying seed protein, oil, and sucrose content in an interspecific mapping population of soybean using high-density linkage mapping. Plant Biotechnol. J. 2018, 16, 1939–1953. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Li, H.; Wang, J.; Zhang, H.; Hu, Z.; Chu, S.; Lv, H.; Yu, D. High-density genetic mapping identifies new major loci for tolerance to low-phosphorus stress in soybean. Front. Plant Sci. 2016, 7, 372. [Google Scholar] [CrossRef]
- Lee, J.S.; Kim, S.; Ha, B.-K.; Kang, S. Positional mapping and identification of novel quantitative trait locus responsible for UV-B radiation tolerance in soybean [Glycine max (L.) Merr.]. Mol. Breed. 2016, 36, 50. [Google Scholar] [CrossRef]
- Niu, J.; Guo, N.; Sun, J.; Li, L.; Cao, Y.; Li, S.; Huang, J.; Zhao, J.; Zhao, T.; Xing, H. Fine mapping of a resistance gene RpsHN that controls Phytophthorasojae using recombinant inbred lines and secondary populations. Front. Plant Sci. 2017, 8, 538. [Google Scholar] [CrossRef]
- Ren, H.; Han, J.; Wang, X.; Zhang, B.; Yu, L.; Gao, H.; Hong, H.; Sun, R.; Tian, Y.; Qi, X.; et al. QTL mapping of drought tolerance traits in soybean with SLAF sequencing. Crop J. 2020, 8, 977–989. [Google Scholar] [CrossRef]
- Wilkes, J.; Saski, C.; Klepadlo, M.; Fallen, B.; Agudelo, P. Quantitative trait loci associated with Rotylenchulusreniformis host suitability in soybean. Phytopathology 2020, 110, 1511–1521. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Liu, D.; Zhang, X.; Li, W.; Liu, H.; Hong, W.; Jiang, C.; Guan, N.; Ma, C.; Zeng, H.; et al. SLAF-seq: An efficient method of large-scale de novo SNP discovery and genotyping using high-throughput sequencing. PLoS ONE 2013, 8, e58700. [Google Scholar] [CrossRef] [PubMed]
- Davey, J.W.; Hohenlohe, P.A.; Etter, P.D.; Boone, J.Q.; Catchen, J.M.; Blaxter, M.L. Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat. Rev. Genet. 2011, 12, 499–510. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Reif, J.C.; Ma, Y.; Hong, H.; Liu, Z.; Chang, R.; Qiu, L. Targeted association mapping demonstrating the complex molecular genetics of fatty acid formation in soybean. BMC Genom. 2015, 16, 841. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Du, W.; Yu, W.; Zhang, W.; Huang, F.; Cheng, H.; Yu, D. Genome-wide association analysis discovered new loci and candidate genes associated with low-phosphorus tolerance based on shoot mineral elements concentrations in soybean. Mol. Genet. Genom. 2022, 297, 843–858. [Google Scholar] [CrossRef]
- Song, Q.; Jenkins, J.; Jia, G.; Hyten, D.L.; Pantalone, V.; Jackson, S.A.; Schmutz, J.; Cregan, P.B. Construction of high resolution genetic linkage maps to improve the soybean genome sequence assembly Glyma1. 01. BMC Genom. 2016, 17, 33. [Google Scholar] [CrossRef]
- Rebetzke, G.J.; Burton, J.W.; Carter Jr, T.E.; Wilson, R.F. Changes in agronomic and seed characteristics with selection for reduced palmitic acid content in soybean. Crop Sci. 1998, 38, 297–302. [Google Scholar] [CrossRef]
- Reinprecht, Y.; Poysa, V.W.; Yu, K.; Rajcan, I.; Ablett, G.R.; Pauls, K.P. Seed and agronomic QTL in low linolenic acid, lipoxygenase-free soybean (Glycine max (L.) Merrill) germplasm. Genome 2006, 49, 1510–1527. [Google Scholar] [CrossRef]
- Hyten, D.L.; Pantalone, V.R.; Saxton, A.M.; Schmidt, M.E.; Sams, C.E. Molecular mapping and identification of soybean fatty acid modifier quantitative trait loci. J. Am. Oil Chem. Soc. 2004, 81, 1115–1118. [Google Scholar] [CrossRef]
- Li, Z.; Wilson, R.F.; Rayford, W.E.; Boerma, H.R. Molecular mapping genes conditioning reduced palmitic acid content in N87-2122-4 soybean. Crop Sci. 2002, 42, 373–378. [Google Scholar]
- Leamy, L.J.; Zhang, H.; Li, C.; Chen, C.Y.; Song, B.-H. A genome-wide association study of seed composition traits in wild soybean (Glycine soja). BMC Genom. 2017, 18, 18. [Google Scholar] [CrossRef] [Green Version]
- Priolli, R.; Carvalho, C.; Bajay, M.; Pinheiro, J.B.; Vello, N.A. Genome analysis to identify SNPs associated with oil content and fatty acid components in soybean. Euphytica 2019, 215, 54. [Google Scholar] [CrossRef]
- Priolli, R.H.G.; Campos, J.; Stabellini, N.; Pinheiro, J.B.; Vello, N.A. Association mapping of oil content and fatty acid components in soybean. Euphytica 2015, 203, 83–96. [Google Scholar] [CrossRef]
- Huai, Y.Y.; Zhang, S.R.; Wu, T.T.; Muhammad, A.; Li, J.; Sun, S.; Han, T.F.; Li Bin Sun, J.M. Potential evaluation of molecular markers related to major nutritional quality traits in soybean breeding. Acta Agr. Sin. 2022, 48, 1957–1976. [Google Scholar]
- Bachleda, N.; Pham, A.; Li, Z. Identifying FATB1a deletion that causes reduced palmitic acid content in soybean N87-2122-4 to develop a functional marker for marker-assisted selection. Mol. Breed. 2016, 36, 45. [Google Scholar] [CrossRef]
- Wang, X.; Jiang, G.-L.; Green, M.; Scott, R.A.; Hyten, D.L.; Cregan, P.B. Quantitative trait locus analysis of saturated fatty acids in a population of recombinant inbred lines of soybean. Mol. Breed. 2012, 30, 1163–1179. [Google Scholar] [CrossRef]
- Wilson, R.F. The role of genomics and biotechnology in achieving global food security for high-oleic vegetable oil. J. Oleo Sci. 2012, 61, 357–367. [Google Scholar] [CrossRef]
- Cahoon, E.B.; Clemente, T.E.; Damude, H.G.; Kinney, A.J. Modifying vegetable oils for food and non-food purposes. In Oil Crops; Vollmann, J., Rajcan, I., Eds.; Springer: New York, NY, USA, 2009; pp. 31–56. [Google Scholar]
- Thapa, R.; Carrero-Colón, M.; Hudson, K.A. New alleles of FATB1A to reduce palmitic acid levels in soybean. Crop Sci. 2016, 56, 1076–1080. [Google Scholar] [CrossRef]
- Gillman, J.D.; Stacey, M.G.; Cui, Y.; Berg, H.R.; Stacey, G. Deletions of the SACPD-C locus elevate seed stearic acid levels but also result in fatty acid and morphological alterations in nitrogen fixing nodules. BMC Plant Biol. 2014, 14, 143. [Google Scholar] [CrossRef] [PubMed]
- Bueno, R.D.; God, P.I.V.G.; Prata, I.O.; Pereira, P.H.S.; Teixeira, A.I.; Piovesan, N.D.; De Barros, E.G. Association of candidate genes for fatty acid content in soybean by temperature-switch PCR (TSP) genotyping. Crop Breed. Appl. Biotechnol. 2018, 18, 244–251. [Google Scholar] [CrossRef]
- Saghaimaroof, M.A.; Soliman, K.M.; Jorgensen, R.A.; Allard, R.W. Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc. Natl. Acad. Sci. USA 1984, 81, 8014–8018. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Ye, G.; Wang, J. A modified algorithm for the improvement of composite interval mapping. Genetics 2007, 175, 361–374. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Ribaut, J.-M.; Li, Z.; Wang, J. Inclusive composite interval mapping (ICIM) for digenic epistasis of quantitative traits in biparental populations. Theor. Appl. Genet. 2008, 116, 243–260. [Google Scholar] [CrossRef] [PubMed]
- Broman, K.W.; Wu, H.; Sen, Ś.; Churchill, G.A. R/qtl: QTL mapping in experimental crosses. Bioinformatics 2003, 19, 889–890. [Google Scholar] [CrossRef] [Green Version]
- Fehr, W.; Caviness, C. Stages of soybean development. In Iowa Agricultural and Home Economics Experiment Station Special Report; Iowa State University: Ames, IA, USA, 1977; pp. 3–11. [Google Scholar]




| Year | Parents | RILs | ||||||
|---|---|---|---|---|---|---|---|---|
| DN42 | Hobbit | Mean | SD | Range | Skewness | Kurtosis | CV (%) | |
| 2017 | 11.99 | 9.32 | 9.97 | 0.8 | 1.69–11.2 | −6.17 | 61.25 | 0.08 |
| 2018 | 11.99 | 8.12 | 9.52 | 0.76 | 1.57–11.03 | −6.17 | 63.94 | 0.08 |
| 2019 | 10.86 | 8.92 | 9.92 | 0.81 | 1.9–11.69 | −5.25 | 51.7 | 0.08 |
| Chromosome | Number of Markers | Map Distance (cM) | Average Map Distance (cM) | Gap < 5 cM (%) | Max Gap (cM) |
|---|---|---|---|---|---|
| 1 | 539 | 184.4 | 0.34 | 100 | 4.53 |
| 2 | 170 | 102.71 | 0.6 | 99.41 | 6.15 |
| 3 | 754 | 106.6 | 0.14 | 100 | 2.93 |
| 4 | 136 | 156.16 | 1.15 | 100 | 3.24 |
| 5 | 319 | 117.18 | 0.37 | 99.06 | 12.36 |
| 6 | 356 | 137.53 | 0.39 | 99.72 | 5.46 |
| 7 | 401 | 131.11 | 0.33 | 99.75 | 5.23 |
| 8 | 190 | 128.56 | 0.68 | 100 | 3.26 |
| 9 | 420 | 96.8 | 0.23 | 100 | 3.24 |
| 10 | 269 | 117.89 | 0.44 | 98.88 | 12.47 |
| 11 | 145 | 116.45 | 0.8 | 99.31 | 5.2 |
| 12 | 134 | 98.46 | 0.73 | 96.99 | 7.57 |
| 13 | 684 | 146.24 | 0.21 | 99.85 | 6.49 |
| 14 | 699 | 112.55 | 0.16 | 99.71 | 8.86 |
| 15 | 414 | 149.5 | 0.36 | 100 | 4.95 |
| 16 | 845 | 123.57 | 0.15 | 100 | 2.93 |
| 17 | 969 | 136.54 | 0.14 | 100 | 4.53 |
| 18 | 1119 | 169.34 | 0.15 | 100 | 2.43 |
| 19 | 1093 | 158.84 | 0.15 | 99.73 | 11.84 |
| 20 | 324 | 112.15 | 0.35 | 100 | 4.53 |
| Total | 9980 | 2602.58 | 0.39 | 19.62 | 5.91 |
| QTL | Year | Chr. | Left Marker | Right Marker | Genetic Position (cM) | Physical Position (bp) | LOD | ADD | PVE (%) | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Start | End | Start | End | ||||||||
| qPA10-1 | 2017 | 10 | Marker7632755 | Marker7723217 | 76.341 | 76.341 | 39,427,048 | 39,517,256 | 2.298 | 0.174 | 4.151 |
| qPA15-1 | 2018 | 15 | Marker5381863 | Marker5394023 | 43.494 | 43.78 | 9,218,139 | 9,387,049 | 6.528 | 0.259 | 10.066 |
| qPA15-2 | 2019 | 15 | Marker5381863 | Marker5394023 | 43.494 | 43.78 | 9,218,139 | 9,387,049 | 3.999 | 0.261 | 9.526 |
| Gene | SNP | Indel | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Intergenic | Upstream | Downstream | Nonsynonymous SNV | Synonymous SNV | Intronic | 3′UTR | Intergenic | Upstream | Downstream | Non-Frameshift Deletion | Non-Frameshift Insertion | 5′ UTR | |
| Glyma.15g119200 | 5 | 10 | 2 | 3 | |||||||||
| Glyma.15g119300 | 3 | 8 | 1 | 2 | 1 | ||||||||
| Glyma.15g119400 | 3 | 1 | 1 | 4 | 3 | 1 | |||||||
| Glyma.15g119500 | 7 | 6 | 1 | 3 | 4 | 7 | 1 | 3 | |||||
| Glyma.15g119600 | 14 | 23 | 2 | 5 | |||||||||
| Glyma.15g119700 | 15 | 13 | 2 | 1 | 4 | 3 | 1 | ||||||
| Glyma.15g119800 | 21 | 20 | 5 | 8 | 3 | ||||||||
| Total | 319 | 68 | 80 | 12 | 1 | 1 | 11 | 77 | 17 | 23 | 1 | 1 | 5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xue, Y.; Gao, H.; Liu, X.; Tang, X.; Cao, D.; Luan, X.; Zhao, L.; Qiu, L. QTL Mapping of Palmitic Acid Content Using Specific-Locus Amplified Fragment Sequencing (SLAF-Seq) Genotyping in Soybeans (Glycine max L.). Int. J. Mol. Sci. 2022, 23, 11273. https://doi.org/10.3390/ijms231911273
Xue Y, Gao H, Liu X, Tang X, Cao D, Luan X, Zhao L, Qiu L. QTL Mapping of Palmitic Acid Content Using Specific-Locus Amplified Fragment Sequencing (SLAF-Seq) Genotyping in Soybeans (Glycine max L.). International Journal of Molecular Sciences. 2022; 23(19):11273. https://doi.org/10.3390/ijms231911273
Chicago/Turabian StyleXue, Yongguo, Huawei Gao, Xinlei Liu, Xiaofei Tang, Dan Cao, Xiaoyan Luan, Lin Zhao, and Lijuan Qiu. 2022. "QTL Mapping of Palmitic Acid Content Using Specific-Locus Amplified Fragment Sequencing (SLAF-Seq) Genotyping in Soybeans (Glycine max L.)" International Journal of Molecular Sciences 23, no. 19: 11273. https://doi.org/10.3390/ijms231911273




