Integrative Metabolomic and Transcriptomic Analysis Reveals the Mechanism of Specific Color Formation in Phoebe zhennan Heartwood
Abstract
1. Introduction
2. Results
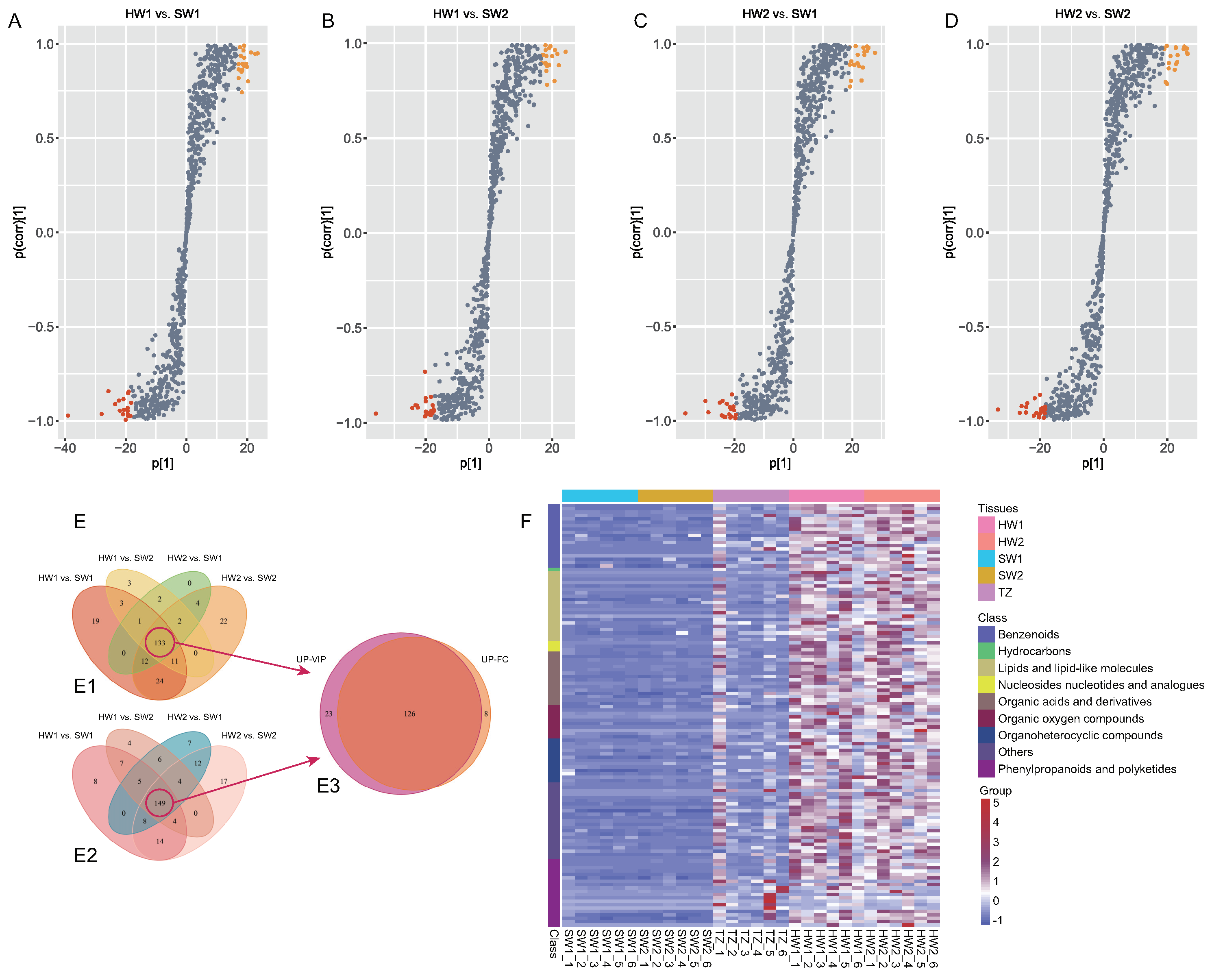
2.1. Comparison of DAMs among Sapwood, the Transition Zone, and Heartwood
2.2. Screening of Key Golden Thread-Related Metabolites in Nanmu Heartwood
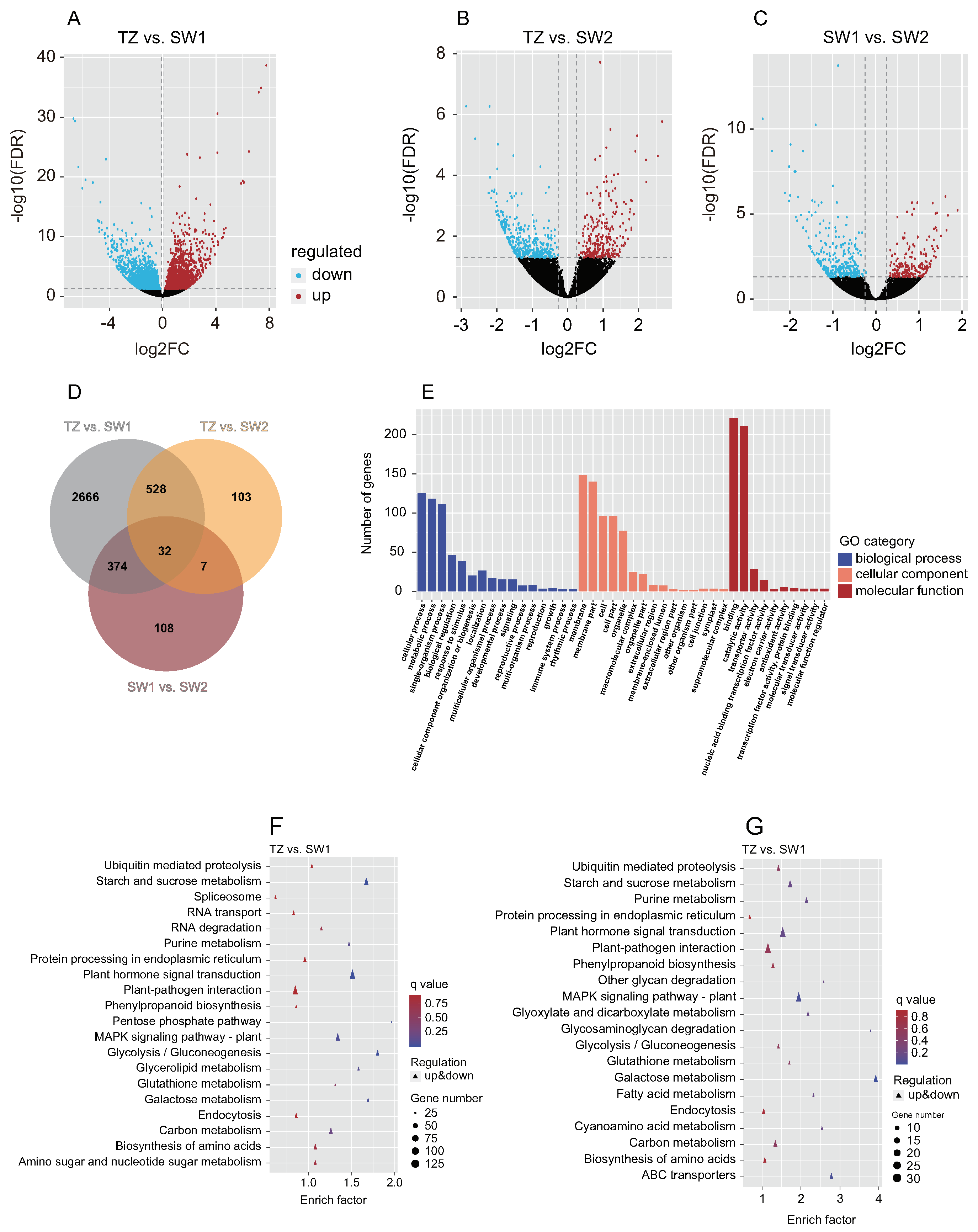
2.3. Transcriptome Analysis and DEG Identification in Different Tissues
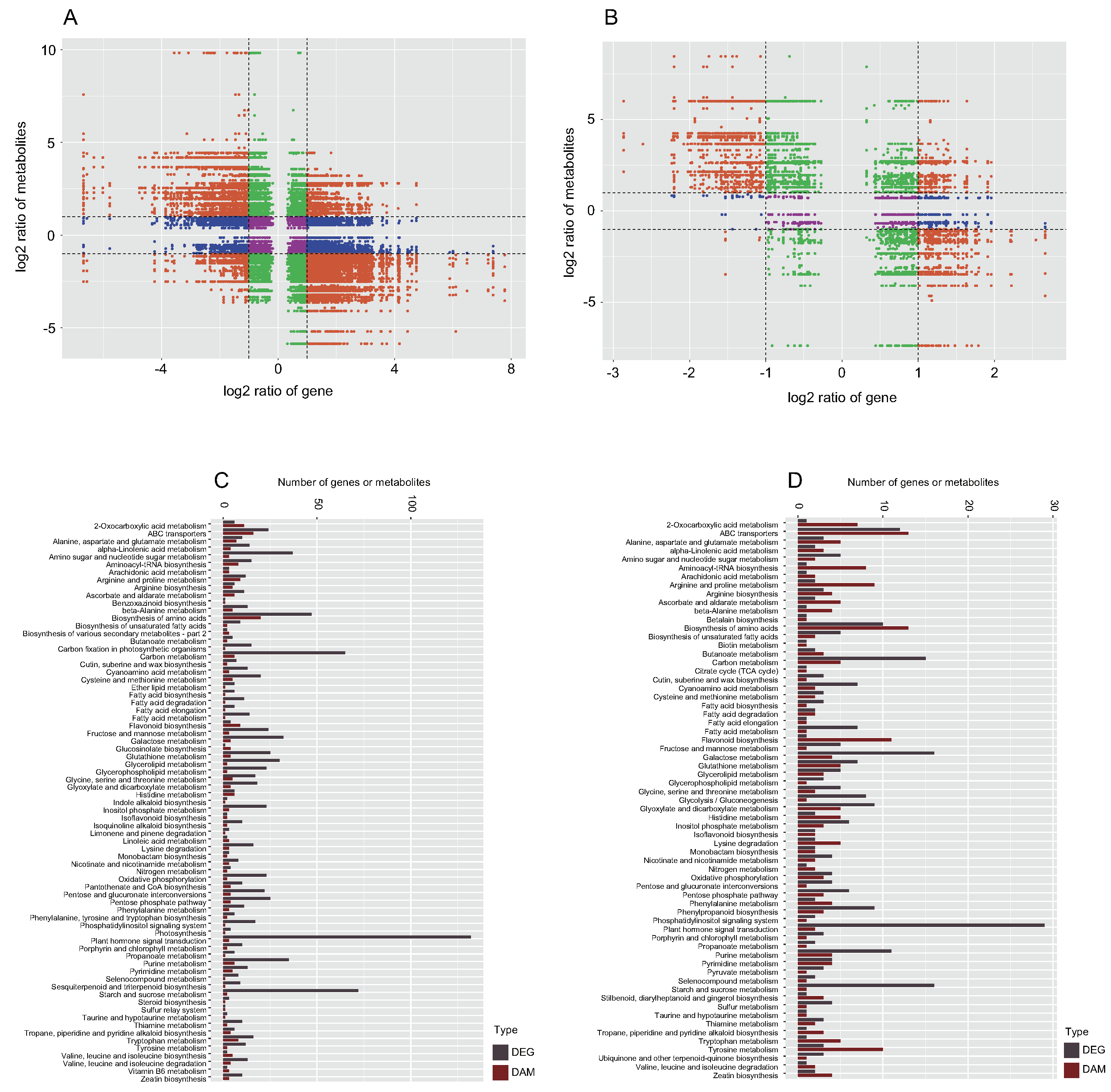
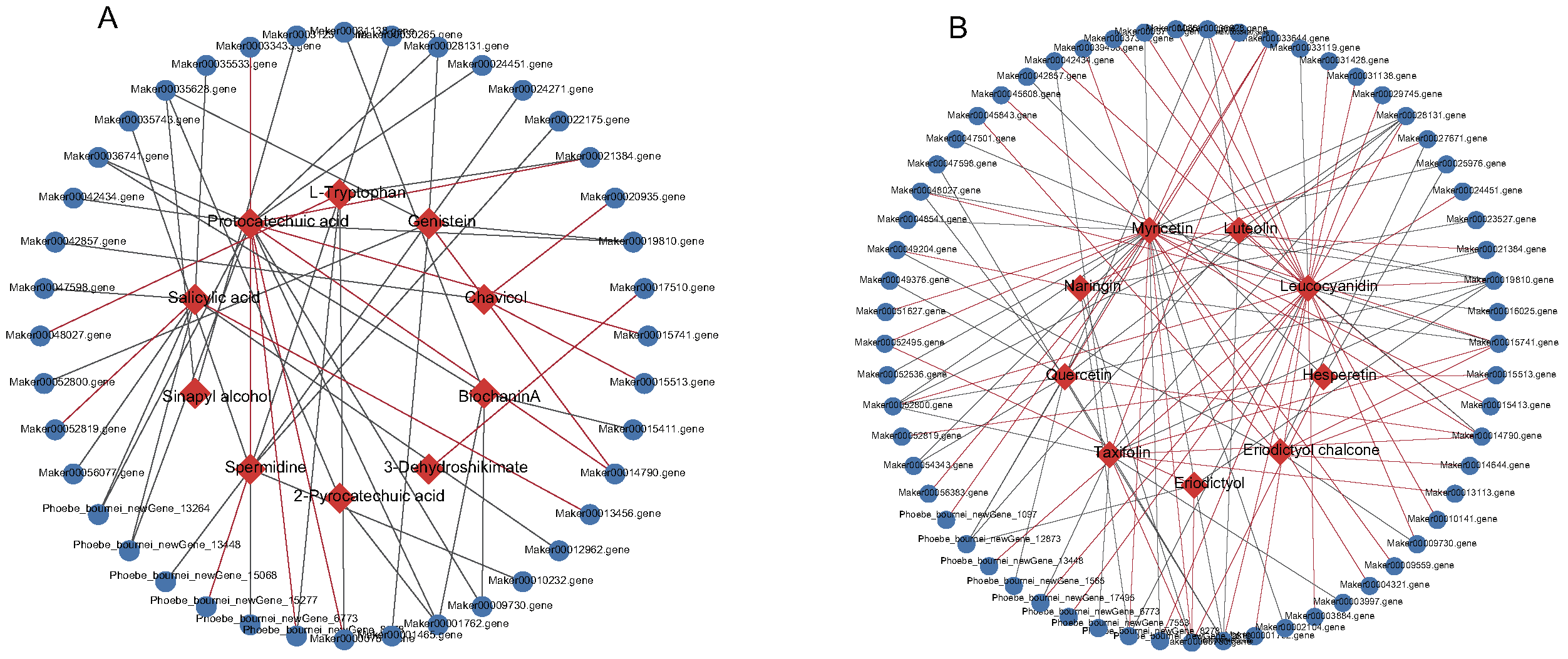
2.4. Correlation between Metabolites and Genes
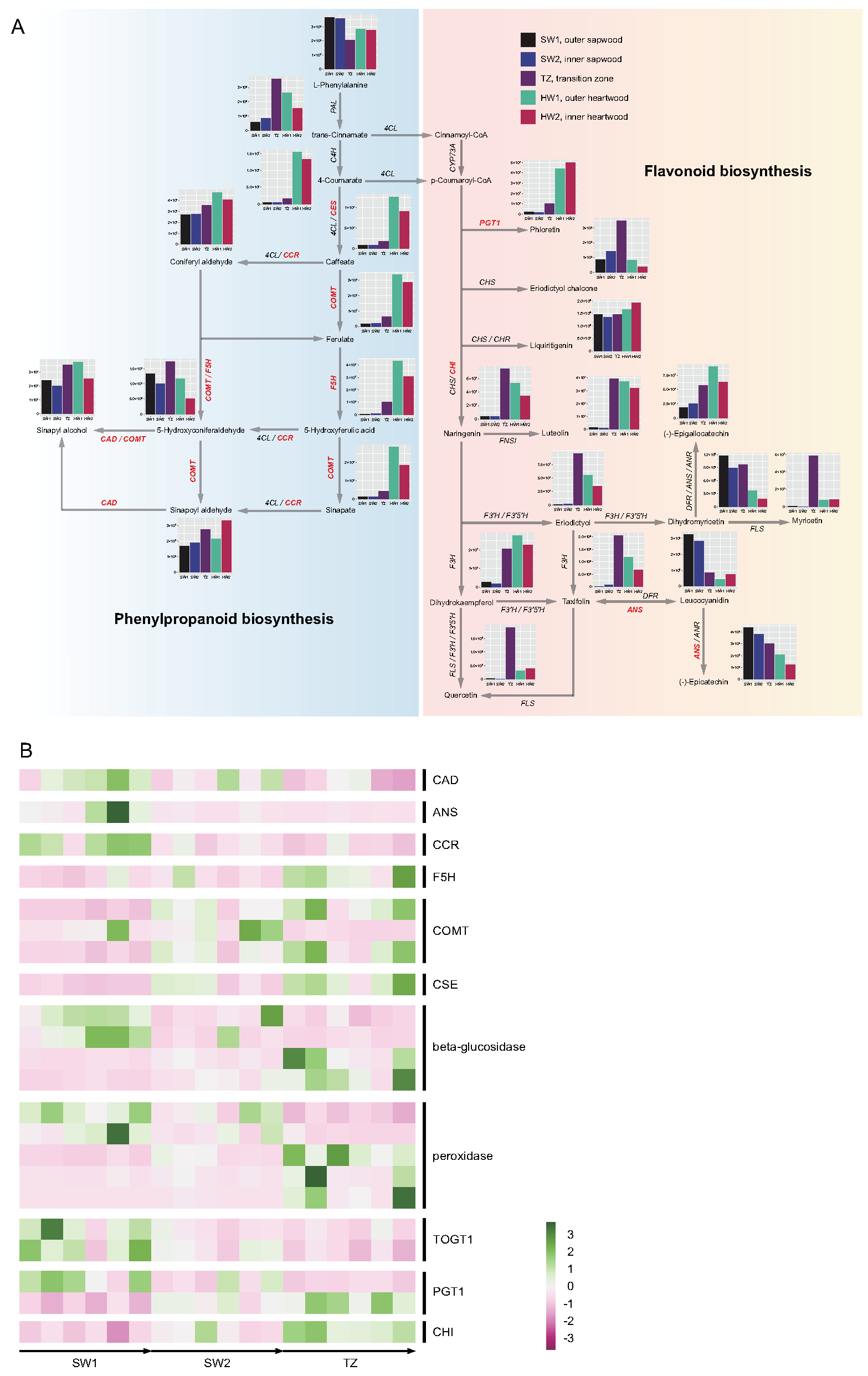
2.5. Phenylpropanoid and Flavonoid Biosynthesis Pathways Related to Nanmu Special Color Heartwood Formation
2.6. Transcription Factors Related to Phenylpropanoid and Flavonoid Biosynthesis
3. Discussion
4. Materials and Methods
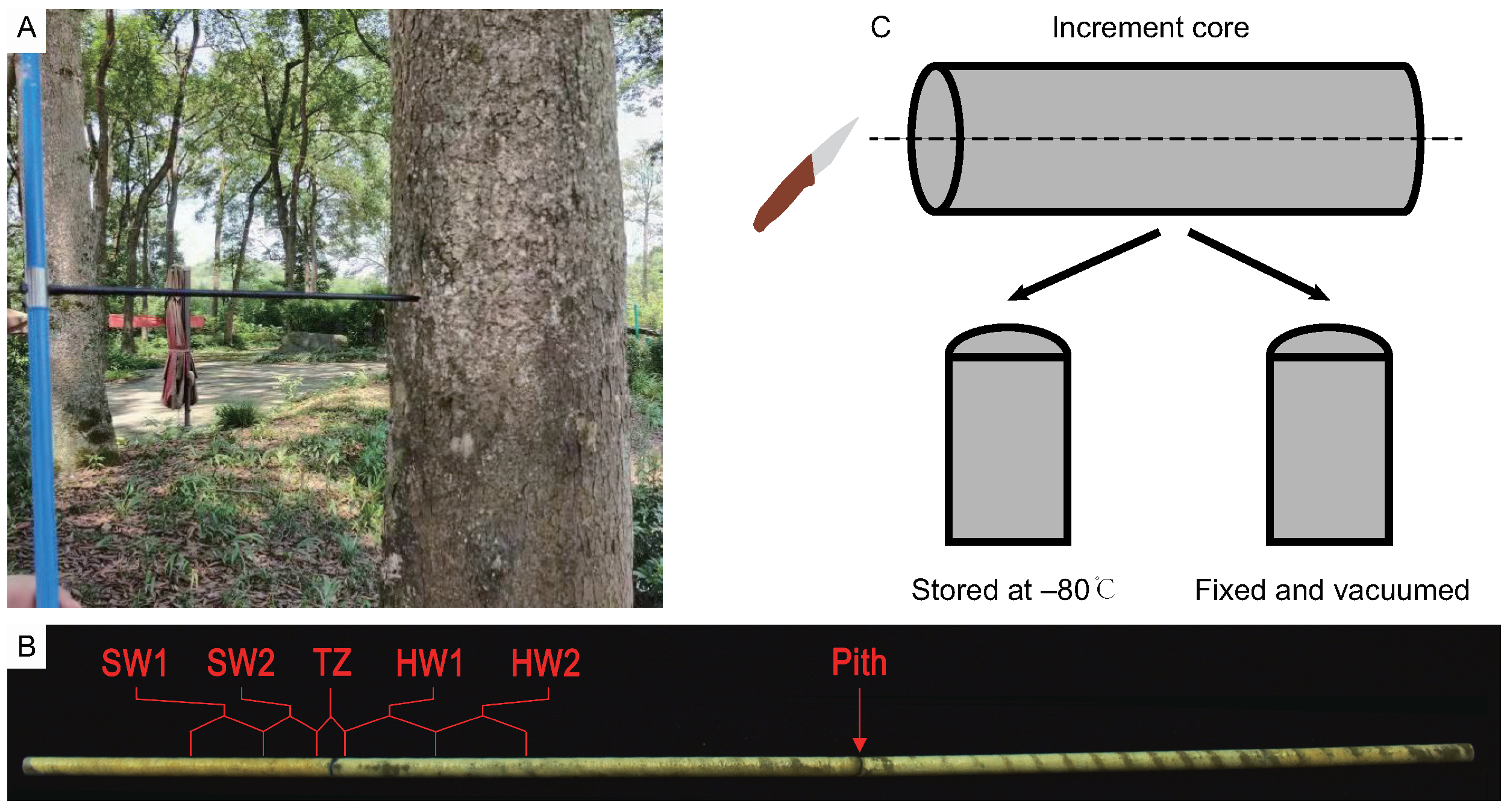
4.1. Plant Materials and Sampling
4.2. Microscopy Analyses and Flavonoid Content Measurement
4.3. Identification and Quantification of Metabolites
4.4. RNA-Seq Analysis
4.5. Real-Time Quantitative PCR
4.6. Integrative Transcriptomic and Metabolomic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, L.; Sun, X.; Wilson, I.W.; Shao, F.; Qiu, D. Identification of the genes involved in anthocyanin biosynthesis and accumulation in Taxus chinensis. Genes 2019, 10, 982. [Google Scholar] [CrossRef] [PubMed]
- Baar, J.; Wimmer, R.; D’Amico, S. Dependence of colour and discolouration on total extractive content of African Padauk and Jatoba. Wood Sci. Technol. 2014, 48, 1155–1165. [Google Scholar] [CrossRef]
- Ding, X.; Xiao, J.H.; Li, L.; Conran, J.G.; Li, J. Congruent species delimitation of two controversial gold-thread nanmu tree species based on morphological and restriction site-associated DNA sequencing data. J. Syst. Evol. 2019, 57, 234–246. [Google Scholar] [CrossRef]
- Xiao, J.-H.; Ding, X.; Li, L.; Ma, H.; Ci, X.-Q.; van der Merwe, M.; Conran, J.G.; Li, J. Miocene diversification of a golden-thread nanmu tree species (Phoebe zhennan, Lauraceae) around the Sichuan Basin shaped by the East Asian monsoon. Ecol. Evol. 2020, 10, 10543–10557. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Qi, J.; Huang, X.; Zhou, N.; Hu, Y. Comparative analysis of modern and ancient buried Phoebe zhennan wood: Surface color, chemical components, infrared spectroscopy, and essential oil composition. J. For. Res. 2015, 26, 501–507. [Google Scholar] [CrossRef]
- Yeh, T.F.; Chu, J.H.; Liu, L.Y.; Chen, S.Y. Differential gene profiling of the heartwood formation process in Taiwania cryptomerioides Hayata xylem tissues. Int. J. Mol. Sci. 2020, 21, 960. [Google Scholar] [CrossRef] [PubMed]
- Rudman, P. Heartwood formation in trees. Nature 1966, 210, 608–610. [Google Scholar] [CrossRef]
- Plomion, C.; Leprovost, G.; Stokes, A. Wood formation in trees. Plant Physiol. 2001, 127, 1513–1523. [Google Scholar] [CrossRef]
- Shao, F.; Zhang, L.; Guo, J.; Liu, X.; Ma, W.; Wilson, I.W.; Qiu, D. A comparative metabolomics analysis of the components of heartwood and sapwood in Taxus chinensis (Pilger) Rehd. Sci. Rep. 2019, 9, 17647. [Google Scholar] [CrossRef]
- Beritognolo, I.; Magel, E.; Abdel-Latif, A.; Charpentier, J.-P.; Jay-Allemand, C.; Breton, C. Expression of genes encoding chalcone synthase, flavanone 3-hydroxylase and dihydroflavonol 4-reductase correlates with flavanol accumulation during heartwood formation in Juglans nigra. Tree Physiol. 2002, 22, 291–300. [Google Scholar] [CrossRef]
- Brocco, V.F.; Paes, J.B.; Costa, L.G.d.; Kirker, G.T.; Brazolin, S. Wood color changes and termiticidal properties of teak heartwood extract used as a wood preservative. Holzforschung 2020, 74, 233–245. [Google Scholar] [CrossRef]
- Cao, S.; Zhang, Z.; Sun, Y.; Li, Y.; Zheng, H. Profiling of widely targeted metabolomics for the identification of secondary metabolites in heartwood and sapwood of the red-heart chinese fir (Cunninghamia Lanceolata). Forests 2020, 11, 897. [Google Scholar] [CrossRef]
- Yang, X.; Yu, X.; Liu, Y.; Shi, Z.; Li, L.; Xie, S.; Zhu, G.; Zhao, P. Comparative metabolomics analysis reveals the color variation between heartwood and sapwood of Chinese fir (Cunninghamia lanceolata (Lamb.) Hook. Ind. Crop. Prod. 2021, 169, 113656. [Google Scholar] [CrossRef]
- Déjardin, A.; Laurans, F.; Arnaud, D.; Breton, C.; Pilate, G.; Leplé, J.-C. Wood formation in Angiosperms. Comptes Rendus Biol. 2010, 333, 325–334. [Google Scholar] [CrossRef]
- Gierlinger, N.; Jacques, D.; Grabner, M.; Wimmer, R.; Schwanninger, M.; Rozenberg, P.; Pâques, L.E. Colour of larch heartwood and relationships to extractives and brown-rot decay resistance. Trees 2004, 18, 102–108. [Google Scholar] [CrossRef]
- Takahashi, K.; Mori, K. Relationships between blacking phenomenon and norlignans of sugi (Cryptomeria japonica) heartwood III: Coloration of norlignans with alkaline treatment. J. Wood Sci. 2006, 52, 134–139. [Google Scholar] [CrossRef]
- Celedon, J.M.; Bohlmann, J. An extended model of heartwood secondary metabolism informed by functional genomics. Tree Physiol. 2018, 38, 311–319. [Google Scholar] [CrossRef]
- Amusant, N.; Beauchene, J.; Fournier, M.; Janin, G.; Thevenon, M.-F. Decay resistance in Dicorynia guianensis Amsh.: Analysis of inter-tree and intra-tree variability and relations with wood colour. Ann. For. Sci. 2004, 61, 373–380. [Google Scholar] [CrossRef]
- Yoshida, K.; Futamura, N.; Nishiguchi, M. Collection of expressed genes from the transition zone of Cryptomeria japonica in the dormant season. J. Wood Sci. 2012, 58, 89–103. [Google Scholar] [CrossRef]
- Huang, Z.L.; Tsai, C.J.; Harding, S.A.; Meilan, R.; Woeste, K. A cross-species transcriptional profile analysis of heartwood formation in black walnut. Plant Mol. Biol. Report. 2010, 28, 222–230. [Google Scholar] [CrossRef]
- Yang, J.; Kamdem, D.P.; Keathley, D.E.; Han, K.H. Seasonal changes in gene expression at the sapwood—Heartwood transition zone of black locust (Robinia pseudoacacia) revealed by cDNA microarray analysis. Tree Physiol. 2004, 24, 461–474. [Google Scholar] [CrossRef] [PubMed]
- Cao, S.; Deng, H.; Zhao, Y.; Zhang, Z.; Tian, Y.; Sun, Y.; Li, Y.; Zheng, H. Metabolite profiling and transcriptome analysis unveil the mechanisms of red-heart Chinese fir [Cunninghamia lanceolata (Lamb.) Hook] heartwood coloration. Front. Plant Sci. 2022, 13, 854716. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Rao, S.; Li, Q.; Xu, M.; Wang, A.; Zhang, H.; Chen, J. The coloring mechanism of a novel golden variety in Populus deltoides based on the RGB color mode. For. Res. 2021, 1, 1–13. [Google Scholar] [CrossRef]
- Su, Y.; Zhang, J.; Xu, Z.; Li, J.; Wang, P.; Song, Z.; Tian, G.; Li, L.; Song, J.; Wang, J. Integrative analysis of metabolome and transcriptome reveals the mechanism of color formation in white root (Salvia miltiorrhiza). Ind. Crop. Prod. 2021, 170, 113784. [Google Scholar] [CrossRef]
- Hofmann, T.; Guran, R.; Zitka, O.; Visi-Rajczi, E.; Albert, L. Liquid chromatographic/mass spectrometric study on the role of beech (Fagus sylvatica L.) wood polyphenols in red heartwood formation. Forests 2022, 13, 10. [Google Scholar] [CrossRef]
- Mishra, G.; Collings, D.; Altaner, C. Physiological changes during heartwood formation in young Eucalyptus bosistoana trees. IAWA J. 2018, 39, 382–394. [Google Scholar] [CrossRef]
- Yazaki, Y. Wood colors and their coloring matters: A review. Nat. Prod. Commun. 2015, 10, 1934578X1501000332. [Google Scholar] [CrossRef]
- Mei, X.; Wan, S.; Lin, C.; Zhou, C.; Hu, L.; Deng, C.; Zhang, L. Integration of metabolome and transcriptome reveals the relationship of Benzenoid–Phenylpropanoid pigment and aroma in purplet tea flowers. Front. Plant Sci. 2021, 12, 762330. [Google Scholar] [CrossRef]
- Dehon, L.; Macheix, J.J.; Durand, M. Involvement of peroxidases in the formation of the brown coloration of heartwood in Juglans nigra. J. Exp. Bot. 2002, 53, 303–311. [Google Scholar] [CrossRef]
- Yang, J.; Park, S.; Kamdem, D.P.; Keathley, D.E.; Retzel, E.; Paule, C.; Kapur, V.; Han, K.-H. Novel gene expression profiles define the metabolic and physiological processes characteristic of wood and its extractive formation in a hardwood tree species, Robinia pseudoacacia. Plant Mol. Biol. 2003, 52, 935–956. [Google Scholar] [CrossRef]
- Huang, G.; Liao, X.; Han, Q.; Zhou, Z.; Liang, K.; Li, G.; Yang, G.; Tembrock, L.R.; Wang, X.; Wu, Z. Integrated metabolome and transcriptome analyses reveal dissimilarities in the anthocyanin synthesis pathway between different developmental leaf color transitions in Hopea hainanensis (Dipterocarpaceae). Front. Plant Sci. 2022, 13, 830413. [Google Scholar] [CrossRef] [PubMed]
- Nakatsuka, T.; Nishihara, M.; Mishiba, K.; Yamamura, S. Temporal expression of flavonoid biosynthesis-related genes regulates flower pigmentation in gentian plants. Plant Sci. 2005, 168, 1309–1318. [Google Scholar] [CrossRef]
- Muir, S.R.; Collins, G.J.; Robinson, S.; Hughes, S.; Bovy, A.; Ric De Vos, C.H.v.; van Tunen, A.J.; Verhoeyen, M.E. Overexpression of petunia chalcone isomerase in tomato results in fruit containing increased levels of flavonols. Nat. Biotechnol. 2001, 19, 470–474. [Google Scholar] [CrossRef]
- Hamberger, B.; Ellis, M.; Friedmann, M.; de Azevedo Souza, C.; Barbazuk, B.; Douglas, C.J. Genome-wide analyses of phenylpropanoid-related genes in Populus trichocarpa, Arabidopsis thaliana, and Oryza sativa: The Populus lignin toolbox and conservation and diversification of angiosperm gene families. Botany 2007, 85, 1182–1201. [Google Scholar]
- Vogt, T. Phenylpropanoid Biosynthesis. Mol. Plant 2010, 3, 2–20. [Google Scholar] [CrossRef] [PubMed]
- Latchman, D.S. Transcription factors: An overview. Int. J. Biochem. Cell Biol. 1997, 29, 1305–1312. [Google Scholar] [CrossRef]
- Schwechheimer, C.; Bevan, M. The regulation of transcription factor activity in plants. Trends Plant Sci. 1998, 3, 378–383. [Google Scholar] [CrossRef]
- Zhuang, H.; Lou, Q.; Liu, H.; Han, H.; Wang, Q.; Tang, Z.; Ma, Y.; Wang, H. Differential regulation of anthocyanins in green and purple turnips revealed by combined de novo transcriptome and metabolome analysis. Int. J. Mol. Sci. 2019, 20, 4387. [Google Scholar] [CrossRef]
- Feller, A.; Machemer, K.; Braun, E.L.; Grotewold, E. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J. 2011, 66, 94–116. [Google Scholar] [CrossRef]
- Kampe, A.; Magel, E. New insights into heartwood and heartwood formation. In Cellular Aspects of Wood Formation; Fromm, J., Ed.; Plant Cell Monographs; Springer: Berlin/Heidelberg, Germany, 2013; Volume 20, p. 71. [Google Scholar]
- Nakada, R.; Fukatsu, E. Seasonal variation of heartwood formation in Larix kaempferi. Tree Physiol. 2012, 32, 1497–1508. [Google Scholar] [CrossRef]
- Wishart, D.S.; Tzur, D.; Knox, C.; Eisner, R.; Guo, A.C.; Young, N.; Cheng, D.; Jewell, K.; Arndt, D.; Sawhney, S. HMDB: The human metabolome database. Nucleic Acids Res. 2007, 35, D521–D526. [Google Scholar] [CrossRef]
- Horai, H.; Arita, M.; Kanaya, S.; Nihei, Y.; Ikeda, T.; Suwa, K.; Ojima, Y.; Tanaka, K.; Tanaka, S.; Aoshima, K. MassBank: A public repository for sharing mass spectral data for life sciences. J. Mass Spectrom. 2010, 45, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Sud, M.; Fahy, E.; Cotter, D.; Brown, A.; Dennis, E.A.; Glass, C.K.; Merrill, A.H., Jr.; Murphy, R.C.; Raetz, C.R.; Russell, D.W. Lmsd: Lipid maps structure database. Nucleic Acids Res. 2007, 35, D527–D532. [Google Scholar] [CrossRef] [PubMed]
- Abdelrazig, S.; Safo, L.; Rance, G.A.; Fay, M.W.; Theodosiou, E.; Topham, P.D.; Kim, D.-H.; Fernández-Castané, A. Metabolic characterisation of Magnetospirillum gryphiswaldense MSR-1 using LC-MS-based metabolite profiling. RSC Adv. 2020, 10, 32548–32560. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Wishart, D.S. Web-based inference of biological patterns, functions and pathways from metabolomic data using MetaboAnalyst. Nat. Protoc. 2011, 6, 743–760. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.-P.; Sun, W.-H.; Xiong, Y.-F.; Jiang, Y.-T.; Liu, X.-D.; Liao, X.-Y.; Zhang, D.-Y.; Jiang, S.-Z.; Li, Y.; Liu, B.; et al. The Phoebe genome sheds light on the evolution of magnoliids. Hortic. Res. 2020, 7, 146. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Varet, H.; Brillet-Guéguen, L.; Coppée, J.-Y.; Dillies, M.-A. SARTools: A DESeq2-and EdgeR-based R pipeline for comprehensive differential analysis of RNA-Seq data. PLoS ONE 2016, 11, e0157022. [Google Scholar] [CrossRef]
- Bustin, S.A.; Beaulieu, J.F.; Huggett, J.; Jaggi, R.; Kibenge, F.S.B.; Olsvik, P.A.; Penning, L.C.; Toegel, S. MIQE précis: Practical implementation of minimum standard guidelines for fluorescence-based quantitative real-time PCR experiments. BMC Mol. Biol. 2010, 11, 74. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.L.; Chen, J.; Tian, Q.; Wang, S.; Xia, X.; Yin, W. Identification and validation of reference genes for Populus euphratica gene expression analysis during abiotic stresses by quantitative real-time PCR. Physiol. Plant. 2014, 152, 529–545. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, H.; An, W.; Gu, Y.; Peng, J.; Jiang, Y.; Li, J.; Chen, L.; Zhu, P.; He, F.; Zhang, F.; et al. Integrative Metabolomic and Transcriptomic Analysis Reveals the Mechanism of Specific Color Formation in Phoebe zhennan Heartwood. Int. J. Mol. Sci. 2022, 23, 13569. https://doi.org/10.3390/ijms232113569
Yang H, An W, Gu Y, Peng J, Jiang Y, Li J, Chen L, Zhu P, He F, Zhang F, et al. Integrative Metabolomic and Transcriptomic Analysis Reveals the Mechanism of Specific Color Formation in Phoebe zhennan Heartwood. International Journal of Molecular Sciences. 2022; 23(21):13569. https://doi.org/10.3390/ijms232113569
Chicago/Turabian StyleYang, Hanbo, Wenna An, Yunjie Gu, Jian Peng, Yongze Jiang, Jinwu Li, Lianghua Chen, Peng Zhu, Fang He, Fan Zhang, and et al. 2022. "Integrative Metabolomic and Transcriptomic Analysis Reveals the Mechanism of Specific Color Formation in Phoebe zhennan Heartwood" International Journal of Molecular Sciences 23, no. 21: 13569. https://doi.org/10.3390/ijms232113569
APA StyleYang, H., An, W., Gu, Y., Peng, J., Jiang, Y., Li, J., Chen, L., Zhu, P., He, F., Zhang, F., Xiao, J., Liu, M., & Wan, X. (2022). Integrative Metabolomic and Transcriptomic Analysis Reveals the Mechanism of Specific Color Formation in Phoebe zhennan Heartwood. International Journal of Molecular Sciences, 23(21), 13569. https://doi.org/10.3390/ijms232113569





