Zona Pellucida like Domain Protein 1 (ZPLD1) Polymerization Is Regulated by Two Distinguished Hydrophobic Motifs
Abstract
:1. Introduction
2. Results
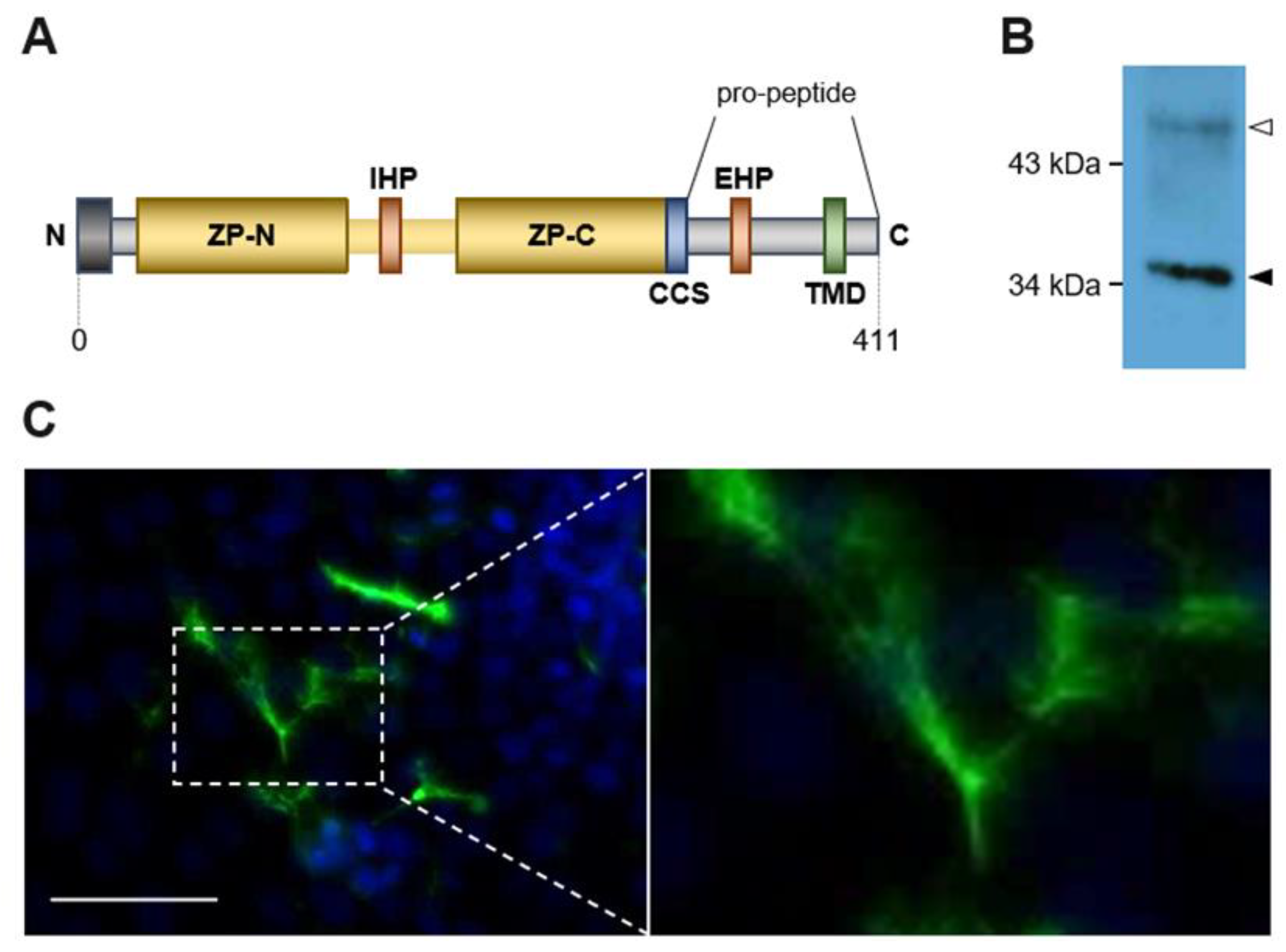
2.1. Transformed MDCK Cells Express Physiological Active ZPLD1
2.2. ZPLD1 Proteins Lacking the EHP or IHP Are Not Properly Processed
2.3. Depletion of EHP or IHP Leads to the Formation of Intracellular Polymeric Structures
2.4. Release of the C-Terminal Pro-Peptide and Hence Loss of the Putative EHP Is a Prerequisite for Proper Extracellular Polymerization
3. Discussion
4. Materials and Methods
4.1. Protein Sequence Analyses
4.2. ZPLD1 Expression Constructs
4.3. Cell Lines and Culture Conditions
4.4. Protein Purification and Western Blot
4.5. Immunofluorescence
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Dernedde, J.; Weise, C.; Müller, E.C.; Hagiwara, A.; Bachmann, S.; Suzuki, M.; Reutter, W.; Tauber, R.; Scherer, H. Cupulin Is a Zona Pellucida-like Domain Protein and Major Component of the Cupula from the Inner Ear. PLoS ONE 2014, 9, e111917. [Google Scholar] [CrossRef]
- Dohlman, G.F. The Shape and Function of the Cupula. J. Laryngol. Otol. 1969, 83, 43–53. [Google Scholar] [CrossRef]
- Takumida, M. Functional Morphology of the Crista Ampullaris: With Special Interests in Sensory Hairs and Cupula: A Review. Uchu Seibutsu Kagaku 2001, 15, 356–358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Helling, K.; Clarke, A.H.; Watanabe, N.; Scherer, H. Morphological Studies of the Form of the Cupula in the Semicircular Canal Ampulla. HNO 2000, 48, 822–827. [Google Scholar] [CrossRef] [PubMed]
- Scherer, H.; Watanabe, S. Introductory Remarks on This Issue. On the Role of the Ampulla in Disturbances of Vestibular Function. Uchu Seibutsu Kagaku 2001, 15, 350–352. [Google Scholar] [CrossRef] [Green Version]
- Iimura, Y.; Suzuki, M.; Otsuka, K.; Inagaki, T.; Konomi, U.; Shimizu, S. Effect of Cupula Shrinkage on the Semicircular Canal Activity. Acta Otolaryngol. 2010, 130, 1092–1096. [Google Scholar] [CrossRef]
- Vijayakumar, S.; Jones, S.M.; Jones, T.A.; Tian, C.; Johnson, K.R. Spontaneous Mutations of the ZPLD1 Gene in Mice Cause Semicircular Canal Dysfunction but Do Not Impair Gravity Receptor or Hearing Functions. Sci Rep. 2019, 9, 12430. [Google Scholar] [CrossRef] [Green Version]
- Gianfrancesco, F.; Esposito, T.; Penco, S.; Maglione, V.; Liquori, C.L.; Patrosso, M.C.; Zuffardi, O.; Ciccodicola, A.; Marchuk, D.A.; Squitieri, F. ZPLD1 Gene Is Disrupted in a Patient with Balanced Translocation That Exhibits Cerebral Cavernous Mal-formations. Neuroscience 2008, 155, 345–349. [Google Scholar] [CrossRef]
- Liu, X.Q.; Fazio, J.; Hu, P.; Paterson, A.D. Identity-By-Descent Mapping for Diastolic Blood Pressure in Unrelated Mexican Americans. BMC Proc. 2016, 10 (Suppl. 7), 263–267. [Google Scholar] [CrossRef] [Green Version]
- Kobayashi, D.; Nishizawa, D.; Takasaki, Y.; Kasai, S.; Kakizawa, T.; Ikeda, K.; Fukuda, K.-I. Genome-Wide Association Study of Sensory Disturbances in the Inferior Alveolar Nerve after Bilateral Sagittal Split Ramus Osteotomy. Mol. Pain 2013, 9, 34. [Google Scholar] [CrossRef]
- Chen, Y.C.; Gonzalez, M.E.; Burman, B.; Zhao, X.; Anwar, T.; Tran, M.; Medhora, N.; Hiziroglu, A.B.; Lee, W.; Cheng, Y.H.; et al. Mesenchymal Stem/Stromal Cell Engulfment Reveals Metastatic Advantage in Breast Cancer. Cell Rep. 2019, 27, 3916–3926.e5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, Z.; Yuan, Q.; Zhao, J.; Wang, B.; Li, S.; Offringa, R.; Jin, X.; Wu, H. The m6A-Related mRNA Signature Predicts the Prognosis of Pancreatic Cancer Patients. Mol. Ther. Oncolytics 2020, 17, 460–470. [Google Scholar] [CrossRef] [PubMed]
- Glessner, J.T.; Bradfield, J.P.; Wang, K.; Takahashi, N.; Zhang, H.; Sleiman, P.M.; Mentch, F.D.; Klim, C.E.; Hou, C.; Thomas, K.A.; et al. A Genome-Wide Study Reveals Copy Number Variants Exclusive to Childhood Obesity Cases. Am. J. Hum. Genet. 2010, 87, 661–666. [Google Scholar] [CrossRef] [Green Version]
- Jovine, L.; Darie, C.C.; Litscher, E.S.; Wassarman, P.M. Zona Pellucida Domain Proteins. Annu. Rev. Biochem. 2005, 74, 83–114. [Google Scholar] [CrossRef] [Green Version]
- Pak, J.; Pu, Y.; Zhang, Z.T.; Hasty, D.L.; Wu, X.R. Tamm-Horsfall Protein Binds to Type 1 Fimbriated Escherichia coli and Prevents, E. coli from Binding to Uroplakin Ia and Ib Receptors. J. Biol. Chem. 2001, 276, 9924–9930. [Google Scholar] [CrossRef] [Green Version]
- Bork, P.; Sander, C. A Large Domain Common to Sperm Receptors (Zp2 and Zp3) and TGF-beta Type III Receptor. FEBS Lett. 1992, 300, 237–240. [Google Scholar] [CrossRef] [Green Version]
- Bokhove, M.; Jovine, L. Structure of Zona Pellucida Module Proteins. Curr. Top. Dev. Biol. 2018, 130, 413–442. [Google Scholar] [CrossRef]
- Jovine, L.; Qi, H.; Williams, Z.; Litscher, E.S.; Wassarman, P.M. A Duplicated Motif Controls Assembly of Zona Pellucida Domain Proteins. Proc. Natl. Acad. Sci. USA 2004, 101, 5922–5927. [Google Scholar] [CrossRef] [Green Version]
- Stanisich, J.J.; Zyla, D.S.; Afanasyev, P.; Xu, J.; Kipp, A.; Olinger, E.; Devuyst, O.; Pilhofer, M.; Boehringer, D.; Glockshuber, R. The Cryo-EM Structure of the Human Uromodulin Filament Core Reveals a Unique Assembly Mechanism. eLife 2020, 9, e60265. [Google Scholar] [CrossRef]
- Stsiapanava, A.; Xu, C.; Brunati, M.; Zamora-Caballero, S.; Schaeffer, C.; Bokhove, M.; Han, L.; Hebert, H.; Carroni, M.; Ya-sumasu, S.; et al. Cryo-EM Structure of Native Human Uromodulin, a Zona Pellucida Module Polymer. EMBO J. 2020, 39, e106807. [Google Scholar] [CrossRef]
- Bokhove, M.; Nishimura, K.; Brunati, M.; Han, L.; de Sanctis, D.; Rampoldi, L.; Jonive, L. A Structured Interdomain Linker Directs Self-Polymerization of Human Uromodulin. Proc. Natl. Acad. Sci. USA 2016, 113, 1552–1557. [Google Scholar] [CrossRef] [Green Version]
- Han, L.; Monné, M.; Okumura, H.; Schwend, T.; Cherry, A.L.; Flot, D.; Matsuda, T.; Jovine, L. Insights into Egg Coat Assembly and Egg-Sperm Interaction from the X-ray Structure of Full-Length ZP3. Cell 2010, 143, 404–415. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schaeffer, C.; Santambrogio, S.; Perucca, S.; Casari, G.; Rampoldi, L. Analysis of Uromodulin Polymerization Provides New Insights into the Mechanisms Regulating ZP Domain-Mediated Protein Assembly. Mol. Biol. Cell 2009, 20, 589–599. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, M.; Gold, L.; Dorward, H.; Liang, L.F.; Hoodbhoy, T.; Boja, E.; Fales, H.M.; Dean, J. Mutation of a Conserved Hydro-phobic Patch Prevents Incorporation of ZP3 into the Zona Pellucida Surrounding Mouse Eggs. Mol. Cell Biol. 2003, 23, 8982–8991. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Darie, C.C.; Biniossek, M.L.; Jovine, L.; Litscher, E.S.; Wassarman, P.M. Structural Characterization of Fish Egg Vitelline En-velope Proteins by Mass Spectrometry. Biochemistry 2004, 43, 7459–7478. [Google Scholar] [CrossRef]
- Darie, C.C.; Biniossek, M.L.; Gawinowicz, M.A.; Milgrom, Y.; Thumfart, J.O.; Jovine, L.; Litscher, E.; Wassarman, P.M. Mass Spectrometric Evidence That Proteolytic Processing of Rainbow Trout Egg Vitelline Envelope Proteins Takes Place on the Egg. J. Biol. Chem. 2005, 280, 37585–37598. [Google Scholar] [CrossRef] [Green Version]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the Sensitivity of Progressive Multiple Sequence Alignment through Sequence Weighting, Position-Specific Gap Penalties and Weight Matrix Choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [Green Version]
- McGuffin, L.J.; Bryson, K.; Jones, D.T. The PSIPRED Protein Structure Prediction Server. Bioinformation 2000, 16, 404–405. [Google Scholar] [CrossRef] [Green Version]
- Drozdetskiy, A.; Cole, C.; Procter, J.; Barton, G.J. JPred4: A Protein Secondary Structure Prediction Server. Nucleic Acids Res. 2015, 43, W389–W394. [Google Scholar] [CrossRef]
- Pollastri, G.; McLysaght, A. Porter: A New, Accurate Server for Protein Secondary Structure Prediction. Bioinformation 2005, 21, 1719–1720. [Google Scholar] [CrossRef] [Green Version]
- Cheng, J.; Randall, A.Z.; Sweredoski, M.J.; Baldi, P. SCRATCH: A Protein Structure and Structural Feature Prediction Server. Nucleic Acids Res. 2005, 33, W72–W76. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rost, B.; Liu, J. The PredictProtein Server. Nucleic Acids Res. 2003, 31, 3300–3304. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Varadi, M.; Anyango, S.; Deshpande, M.; Nair, S.; Natassia, C.; Yordanova, G.; Yuan, D.; Stroe, O.; Wood, G.; Laydon, A.; et al. AlphaFold Protein Structure Database: Massively Expanding the Structural Coverage of Protein-Sequence Space with High-Accuracy models. Nucleic Acids Res. 2021, 50, D439–D444. [Google Scholar] [CrossRef] [PubMed]
- PyMOL|Pymol.org. Available online: https://pymol.org/2/#page-top (accessed on 20 October 2022).




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Knepper, M.I.; Dernedde, J. Zona Pellucida like Domain Protein 1 (ZPLD1) Polymerization Is Regulated by Two Distinguished Hydrophobic Motifs. Int. J. Mol. Sci. 2022, 23, 13894. https://doi.org/10.3390/ijms232213894
Knepper MI, Dernedde J. Zona Pellucida like Domain Protein 1 (ZPLD1) Polymerization Is Regulated by Two Distinguished Hydrophobic Motifs. International Journal of Molecular Sciences. 2022; 23(22):13894. https://doi.org/10.3390/ijms232213894
Chicago/Turabian StyleKnepper, Marie Isabell, and Jens Dernedde. 2022. "Zona Pellucida like Domain Protein 1 (ZPLD1) Polymerization Is Regulated by Two Distinguished Hydrophobic Motifs" International Journal of Molecular Sciences 23, no. 22: 13894. https://doi.org/10.3390/ijms232213894








