Thermostable Proteins from HaCaT Keratinocytes Identify a Wide Breadth of Intrinsically Disordered Proteins and Candidates for Liquid–Liquid Phase Separation
Abstract
1. Introduction
2. Results and Discussion
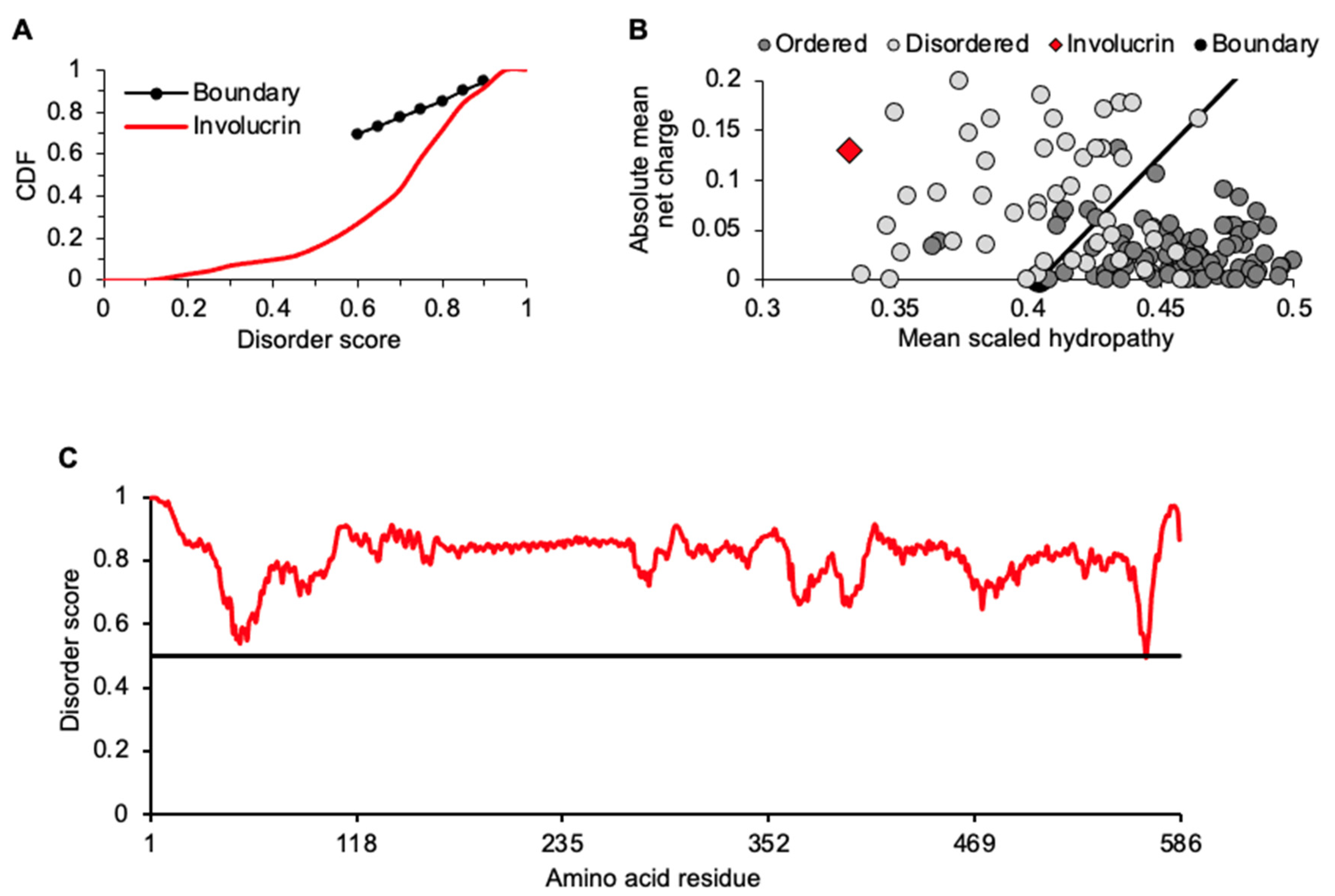
2.1. Retention of Involucrin in a Heat Lysate of Keratinocytes
2.2. Identification and Relationships among Keratinocyte-Expressed Proteins following Thermostability Fractionation
2.3. Intrinsic Disorder Traits of Thermostable Proteins Specific to or Highly Expressed in KCs
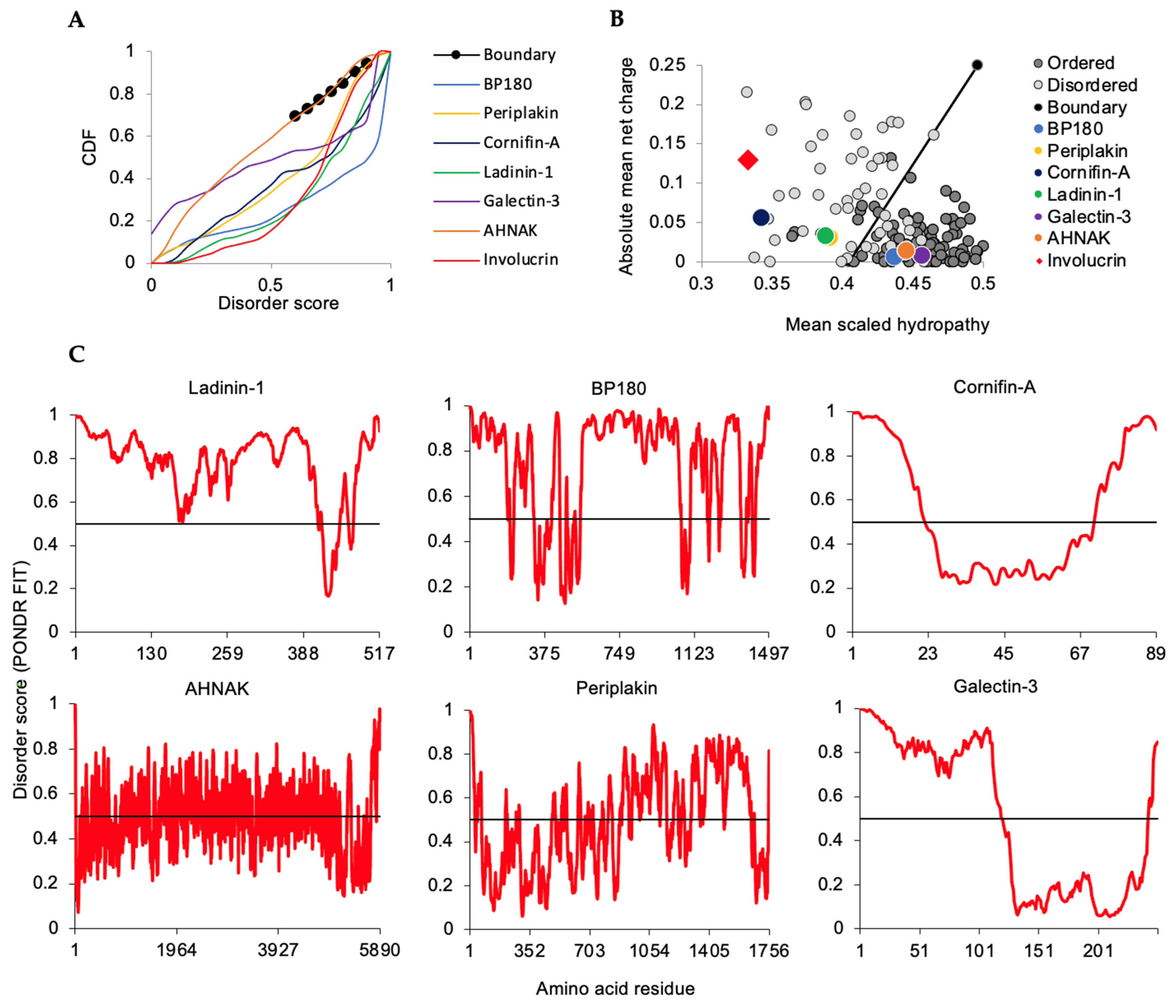
2.4. Intrinsic Disorder Traits of Broadly Expressed Proteins Retrieved in the KC Thermostable Lysate
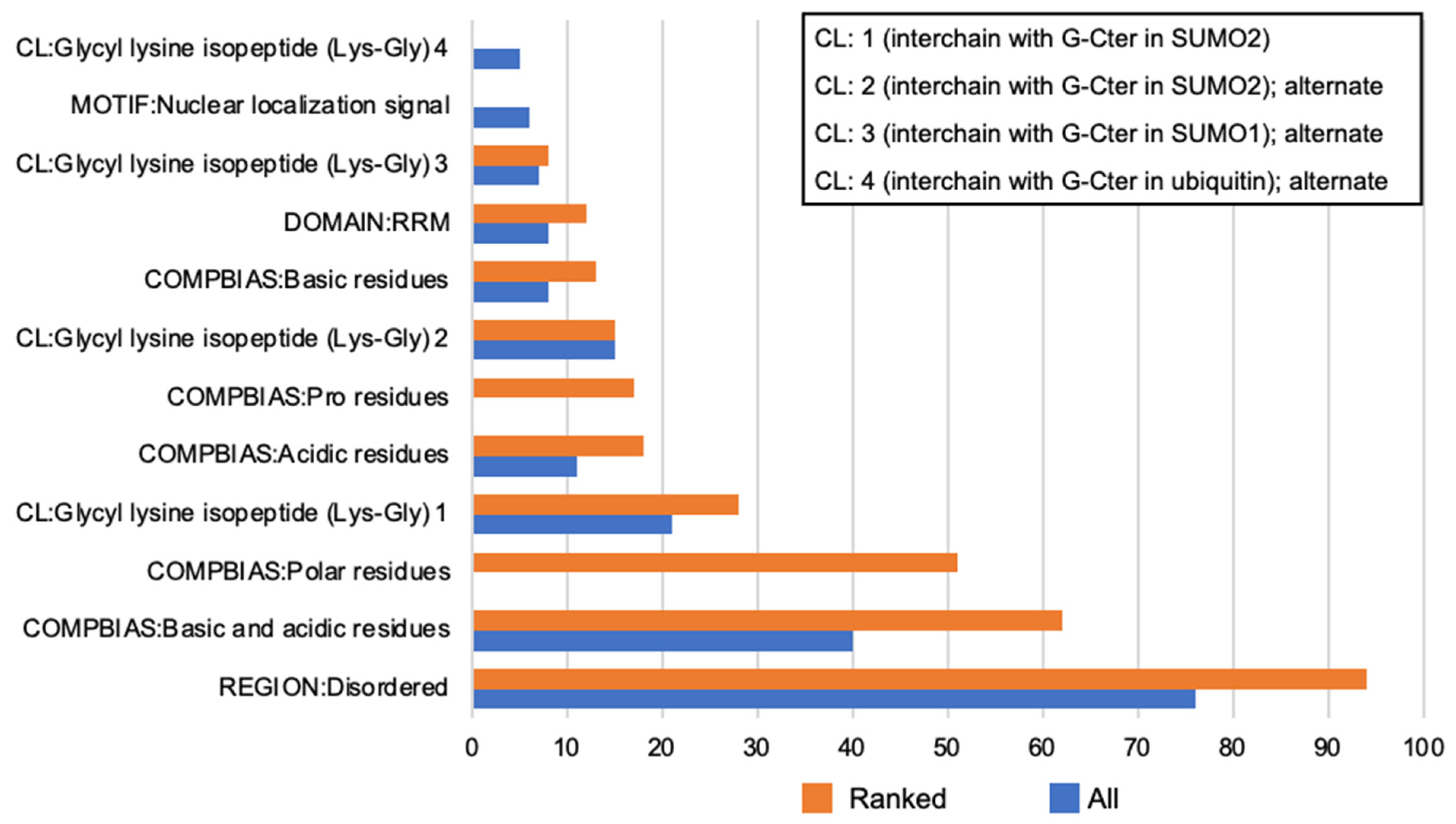
2.5. Keratinocyte Proteins: Database Intrinsic Disorder Assessment
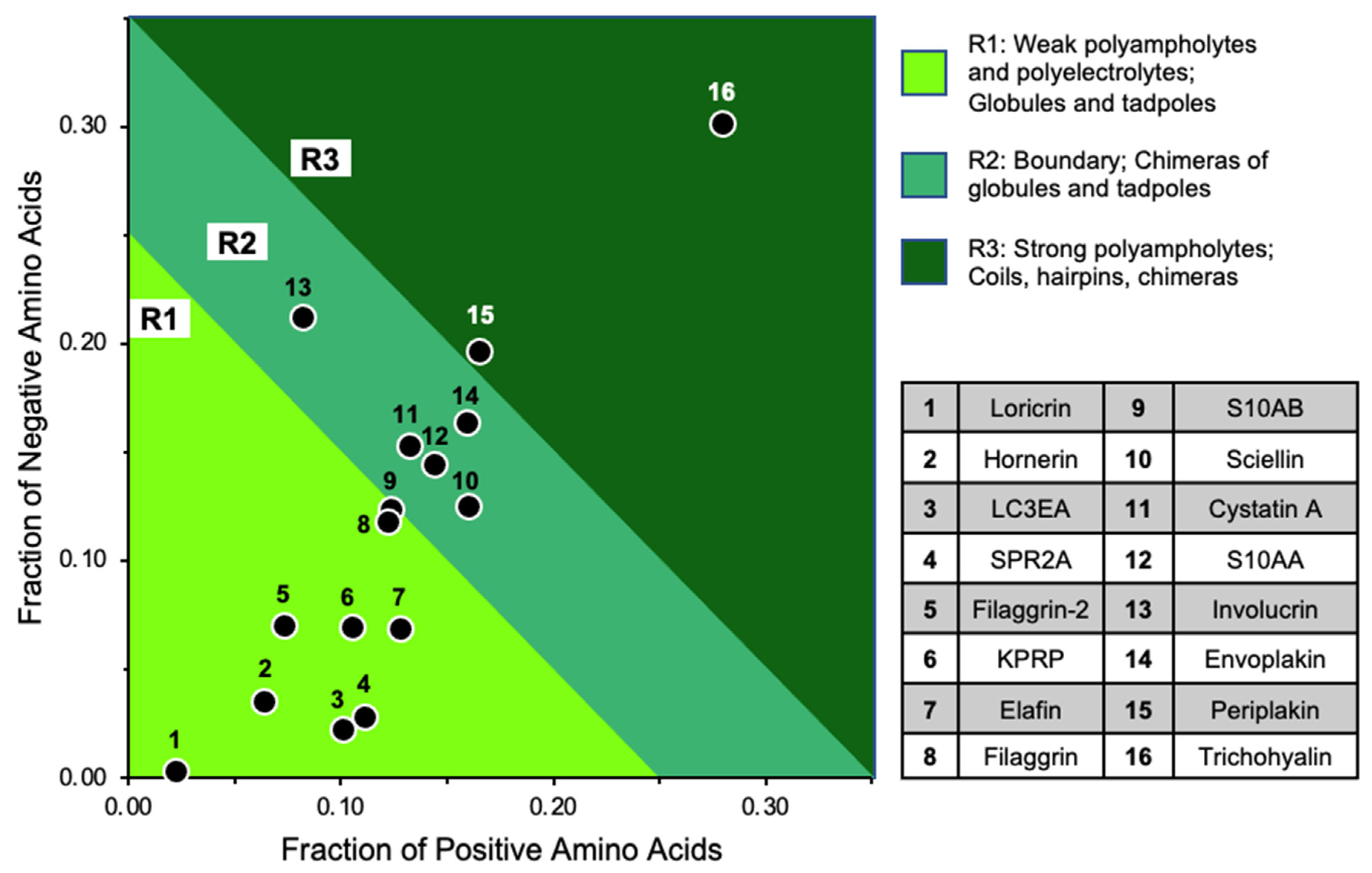
2.6. KC Cornified Envelope (CE) Components Are Candidates for LLPS
2.7. Conclusions
3. Materials and Methods
3.1. Cell Culture
3.2. Protein Preparation and Analysis
3.3. Liquid Chromatography (LC) Tandem Mass Spectrometry (MS, LC MS/MS)
3.4. Bioinformatics: Intrinsic Disorder, Sequence Retrieval, and Phase Separation Analyses
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Uversky, V.N. Protein intrinsic disorder and structure-function continuum. Prog. Mol. Biol. Transl. Sci. 2019, 166, 1–17. [Google Scholar] [PubMed]
- Cumberworth, A.; Lamour, G.; Babu, M.M.; Gsponer, J. Promiscuity as a functional trait: Intrinsically disordered regions as central players of interactomes. Biochem. J. 2013, 454, 361–369. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Huang, Y. Advantages of proteins being disordered. Protein. Sci. 2014, 23, 539–550. [Google Scholar] [CrossRef] [PubMed]
- Jespersen, N.; Barbar, E. Emerging Features of Linear Motif-Binding Hub Proteins. Trends Biochem. Sci. 2020, 45, 375–384. [Google Scholar] [CrossRef]
- Hu, G.; Wu, Z.; Uversky, V.N.; Kurgan, L. Functional Analysis of Human Hub Proteins and Their Interactors Involved in the Intrinsic Disorder-Enriched Interactions. Int. J. Mol. Sci. 2017, 18, 2761. [Google Scholar] [CrossRef]
- Phillips, A.H.; Kriwacki, R.W. Intrinsic protein disorder and protein modifications in the processing of biological signals. Curr. Opin. Struct. Biol. 2020, 60, 1–6. [Google Scholar] [CrossRef]
- Owen, I.; Shewmaker, F. The Role of Post-Translational Modifications in the Phase Transitions of Intrinsically Disordered Proteins. Int. J. Mol. Sci. 2019, 20, 5501. [Google Scholar] [CrossRef]
- Fung, H.Y.J.; Birol, M.; Rhoades, E. IDPs in macromolecular complexes: The roles of multivalent interactions in diverse assemblies. Curr. Opin. Struct. Biol. 2018, 49, 36–43. [Google Scholar] [CrossRef]
- Mulley, J.C.; Callen, D.F. New regional localisations for HAGH and PGP on human chromosome 16. Hum. Genet. 1986, 74, 423–424. [Google Scholar] [CrossRef]
- Bondos, S.E.; Dunker, A.K.; Uversky, V.N. Intrinsically disordered proteins play diverse roles in cell signaling. Cell Commun. Signal. 2022, 20, 20. [Google Scholar] [CrossRef]
- Wright, P.E.; Dyson, H.J. Intrinsically disordered proteins in cellular signalling and regulation. Nat. Rev. Mol. Cell Biol. 2015, 16, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Teilum, K.; Olsen, J.G.; Kragelund, B.B. On the specificity of protein-protein interactions in the context of disorder. Biochem. J. 2021, 478, 2035–2050. [Google Scholar] [CrossRef] [PubMed]
- Lefèvre-Utile, A.; Braun, C.; Haftek, M.; Aubin, F. Five Functional Aspects of the Epidermal Barrier. Int. J. Mol. Sci. 2021, 22, 11676. [Google Scholar] [CrossRef] [PubMed]
- Lintzeri, D.A.; Karimian, N.; Blume-Peytavi, U.; Kottner, J. Epidermal thickness in healthy humans: A systematic review and meta-analysis. J. Eur. Acad. Derm. Venereol. 2022, 36, 1191–1200. [Google Scholar] [CrossRef]
- Rice, G.; Rompolas, P. Advances in resolving the heterogeneity and dynamics of keratinocyte differentiation. Curr. Opin. Cell Biol. 2020, 67, 92–98. [Google Scholar] [CrossRef]
- Eckhart, L.; Lippens, S.; Tschachler, E.; Declercq, W. Cell death by cornification. Biochim. Biophys. Acta 2013, 1833, 3471–3480. [Google Scholar] [CrossRef]
- Furue, M. Regulation of Filaggrin, Loricrin, and Involucrin by IL-4, IL-13, IL-17A, IL-22, AHR, and NRF2: Pathogenic Implications in Atopic Dermatitis. Int. J. Mol. Sci. 2020, 21, 5382. [Google Scholar] [CrossRef]
- McLean, W.H. Filaggrin failure-from ichthyosis vulgaris to atopic eczema and beyond. Br. J. Derm. 2016, 175 (Suppl. 2), 4–7. [Google Scholar] [CrossRef]
- Tuusa, J.; Kokkonen, N.; Tasanen, K. BP180/Collagen XVII: A Molecular View. Int. J. Mol. Sci. 2021, 22, 12233. [Google Scholar] [CrossRef] [PubMed]
- Shamilov, R.; Robinson, V.L.; Aneskievich, B.J. Seeing Keratinocyte Proteins through the Looking Glass of Intrinsic Disorder. Int. J. Mol. Sci. 2021, 22, 7912. [Google Scholar] [CrossRef]
- Quiroz, F.G.; Fiore, V.F.; Levorse, J.; Polak, L.; Wong, E.; Pasolli, H.A.; Fuchs, E. Liquid-liquid phase separation drives skin barrier formation. Science 2020, 367, eaax9554. [Google Scholar] [CrossRef] [PubMed]
- Tuusa, J.; Koski, M.K.; Ruskamo, S.; Tasanen, K. The intracellular domain of BP180/collagen XVII is intrinsically disordered and partially folds in an anionic membrane lipid-mimicking environment. Amino Acids 2020, 52, 619–627. [Google Scholar] [CrossRef] [PubMed]
- Latendorf, T.; Gerstel, U.; Wu, Z.; Bartels, J.; Becker, A.; Tholey, A.; Schröder, J.M. Cationic Intrinsically Disordered Antimicrobial Peptides (CIDAMPs) Represent a New Paradigm of Innate Defense with a Potential for Novel Anti-Infectives. Sci. Rep. 2019, 9, 3331. [Google Scholar] [CrossRef] [PubMed]
- Galea, C.A.; Pagala, V.R.; Obenauer, J.C.; Park, C.G.; Slaughter, C.A.; Kriwacki, R.W. Proteomic studies of the intrinsically unstructured mammalian proteome. J. Proteome. Res. 2006, 5, 2839–2848. [Google Scholar] [CrossRef]
- Galea, C.A.; High, A.A.; Obenauer, J.C.; Mishra, A.; Park, C.G.; Punta, M.; Schlessinger, A.; Ma, J.; Rost, B.; Slaughter, C.A.; et al. Large-scale analysis of thermostable, mammalian proteins provides insights into the intrinsically disordered proteome. J. Proteome. Res. 2009, 8, 211–226. [Google Scholar] [CrossRef] [PubMed]
- Skupien-Rabian, B.; Jankowska, U.; Swiderska, B.; Lukasiewicz, S.; Ryszawy, D.; Dziedzicka-Wasylewska, M.; Kedracka-Krok, S. Proteomic and bioinformatic analysis of a nuclear intrinsically disordered proteome. J. Proteom. 2016, 130, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Etoh, Y.; Simon, M.; Green, H. Involucrin acts as a transglutaminase substrate at multiple sites. Biochem.Biophys. Res. Commun. 1986, 136, 51–56. [Google Scholar] [CrossRef]
- Robinson, N.A.; LaCelle, P.T.; Eckert, R.L. Involucrin is a covalently crosslinked constituent of highly purified epidermal corneocytes: Evidence for a common pattern of involucrin crosslinking in vivo and in vitro. J. Investig. Dermatol. 1996, 107, 101–107. [Google Scholar] [CrossRef][Green Version]
- Rice, R.H.; Green, H. Presence in human epidermal cells of a soluble protein precursor of the cross-linked envelope: Activation of the cross-linking by calcium ions. Cell 1979, 18, 681–694. [Google Scholar] [CrossRef]
- Yaffe, M.B.; Beegen, H.; Eckert, R.L. Biophysical characterization of involucrin reveals a molecule ideally suited to function as an intermolecular cross-bridge of the keratinocyte cornified envelope. J. Biol. Chem. 1992, 267, 12233–12238. [Google Scholar] [CrossRef]
- Kajava, A.V. alpha-Helical solenoid model for the human involucrin. FEBS Lett. 2000, 473, 127–131. [Google Scholar] [CrossRef]
- van der Lee, R.; Buljan, M.; Lang, B.; Weatheritt, R.J.; Daughdrill, G.W.; Dunker, A.K.; Fuxreiter, M.; Gough, J.; Gsponer, J.; Jones, D.T.; et al. Classification of intrinsically disordered regions and proteins. Chem. Rev. 2014, 114, 6589–6631. [Google Scholar] [CrossRef] [PubMed]
- Xue, B.; Dunbrack, R.L.; Williams, R.W.; Dunker, A.K.; Uversky, V.N. PONDR-FIT: A meta-predictor of intrinsically disordered amino acids. Biochim. Biophys. Acta 2010, 1804, 996–1010. [Google Scholar] [CrossRef] [PubMed]
- Candi, E.; Schmidt, R.; Melino, G. The cornified envelope: A model of cell death in the skin. Nat. Rev. Mol. Cell Biol. 2005, 6, 328–340. [Google Scholar] [CrossRef]
- Boczonadi, V.; Määttä, A. Functional Analysis of Periplakin and Envoplakin, Cytoskeletal Linkers, and Cornified Envelope Precursor Proteins. Methods Enzym. 2016, 569, 309–329. [Google Scholar]
- Kalinin, A.; Marekov, L.N.; Steinert, P.M. Assembly of the epidermal cornified cell envelope. J. Cell Sci. 2001, 114, 3069–3070. [Google Scholar] [CrossRef]
- Park, H.; Yamanaka, T.; Nukina, N. Proteomic analysis of heat-stable proteins revealed an increased proportion of proteins with compositionally biased regions. Sci. Rep. 2022, 12, 4347. [Google Scholar] [CrossRef]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 2022, 50, W216–W221. [Google Scholar] [CrossRef]
- Vertegaal, A.C.O. Signalling mechanisms and cellular functions of SUMO. Nat. Rev. Mol. Cell Biol. 2022, 23, 715–731. [Google Scholar] [CrossRef]
- Banani, S.F.; Rice, A.M.; Peeples, W.B.; Lin, Y.; Jain, S.; Parker, R.; Rosen, M.K. Compositional Control of Phase-Separated Cellular Bodies. Cell 2016, 166, 651–663. [Google Scholar] [CrossRef]
- Calabretta, S.; Richard, S. Emerging Roles of Disordered Sequences in RNA-Binding Proteins. Trends Biochem. Sci. 2015, 40, 662–672. [Google Scholar] [CrossRef] [PubMed]
- Martin, E.W.; Holehouse, A.S. Intrinsically disordered protein regions and phase separation: Sequence determinants of assembly or lack thereof. Emerg. Top. Life Sci. 2020, 4, 307–329. [Google Scholar] [PubMed]
- Moll, R.; Moll, I. Epidermal adhesion molecules and basement membrane components as target structures of autoimmunity. Virchows Arch 1998, 432, 487–504. [Google Scholar] [CrossRef] [PubMed]
- Masunaga, T.; Shimizu, H.; Ishiko, A.; Fujiwara, T.; Hashimoto, T.; Nishikawa, T. Desmoyokin/AHNAK protein localizes to the non-desmosomal keratinocyte cell surface of human epidermis. J. Investig. Dermatol. 1995, 104, 941–945. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Hsu, D.K.; Chen, H.Y.; Yang, R.Y.; Carraway, K.L., 3rd; Isseroff, R.R.; Liu, F.T. Galectin-3 regulates intracellular trafficking of EGFR through Alix and promotes keratinocyte migration. J. Investig. Dermatol. 2012, 132, 2828–2837. [Google Scholar] [CrossRef] [PubMed]
- Bouameur, J.E.; Favre, B.; Borradori, L. Plakins, a versatile family of cytolinkers: Roles in skin integrity and in human diseases. J. Investig. Dermatol. 2014, 134, 885–894. [Google Scholar] [CrossRef]
- Fujimoto, W.; Nakanishi, G.; Arata, J.; Jetten, A.M. Differential expression of human cornifin alpha and beta in squamous differentiating epithelial tissues and several skin lesions. J. Investig. Dermatol. 1997, 108, 200–204. [Google Scholar] [CrossRef]
- Trzeciak, M.; Sakowicz-Burkiewicz, M.; Wesserling, M.; Dobaczewska, D.; Glen, J.; Nowicki, R.; Pawelczyk, T. Expression of Cornified Envelope Proteins in Skin and Its Relationship with Atopic Dermatitis Phenotype. Acta Derm.-Venereol. 2017, 97, 36–41. [Google Scholar] [CrossRef]
- Oldfield, C.J.; Cheng, Y.; Cortese, M.S.; Brown, C.J.; Uversky, V.N.; Dunker, A.K. Comparing and combining predictors of mostly disordered proteins. Biochemistry 2005, 44, 1989–2000. [Google Scholar] [CrossRef]
- Shtivelman, E.; Cohen, F.E.; Bishop, J.M. A human gene (AHNAK) encoding an unusually large protein with a 1.2-microns polyionic rod structure. Proc. Natl. Acad. Sci. USA 1992, 89, 5472–5476. [Google Scholar] [CrossRef]
- Aneskievich, B.J.; Shamilov, R.; Vinogradova, O. Intrinsic disorder in integral membrane proteins. Prog. Mol. Biol. Transl. Sci. 2021, 183, 101–134. [Google Scholar] [PubMed]
- Lin, Y.H.; Qiu, D.C.; Chang, W.H.; Yeh, Y.Q.; Jeng, U.S.; Liu, F.T.; Huang, J.R. The intrinsically disordered N-terminal domain of galectin-3 dynamically mediates multisite self-association of the protein through fuzzy interactions. J. Biol. Chem. 2017, 292, 17845–17856. [Google Scholar] [CrossRef] [PubMed]
- Chiu, Y.P.; Sun, Y.C.; Qiu, D.C.; Lin, Y.H.; Chen, Y.Q.; Kuo, J.C.; Huang, J.R. Liquid-liquid phase separation and extracellular multivalent interactions in the tale of galectin-3. Nat. Commun. 2020, 11, 1229. [Google Scholar] [CrossRef] [PubMed]
- Bhopatkar, A.A.; Uversky, V.N.; Rangachari, V. Granulins modulate liquid-liquid phase separation and aggregation of the prion-like C-terminal domain of the neurodegeneration-associated protein TDP-43. J. Biol. Chem. 2020, 295, 2506–2519. [Google Scholar] [CrossRef]
- Roth, L.; Srivastava, S.; Lindzen, M.; Sas-Chen, A.; Sheffer, M.; Lauriola, M.; Enuka, Y.; Noronha, A.; Mancini, M.; Lavi, S.; et al. SILAC identifies LAD1 as a filamin-binding regulator of actin dynamics in response to EGF and a marker of aggressive breast tumors. Sci. Signal. 2018, 11, eaan0949. [Google Scholar] [CrossRef]
- Edqvist, P.H.; Fagerberg, L.; Hallström, B.M.; Danielsson, A.; Edlund, K.; Uhlén, M.; Pontén, F. Expression of human skin-specific genes defined by transcriptomics and antibody-based profiling. J. Histochem. Cytochem. 2015, 63, 129–141. [Google Scholar] [CrossRef]
- Motoki, K.; Megahed, M.; LaForgia, S.; Uitto, J. Cloning and chromosomal mapping of mouse ladinin, a novel basement membrane zone component. Genomics 1997, 39, 323–330. [Google Scholar] [CrossRef]
- Vernon, R.M.; Forman-Kay, J.D. First-generation predictors of biological protein phase separation. Curr. Opin. Struct. Biol. 2019, 58, 88–96. [Google Scholar] [CrossRef]
- Shen, B.; Chen, Z.; Yu, C.; Chen, T.; Shi, M.; Li, T. Computational Screening of Phase-separating Proteins. Genom. Proteom. Bioinform. 2021, 19, 13–24. [Google Scholar] [CrossRef]
- Pancsa, R.; Vranken, W.; Mészáros, B. Computational resources for identifying and describing proteins driving liquid-liquid phase separation. Brief Bioinform. 2021, 22, bbaa408. [Google Scholar] [CrossRef]
- Lin, Z.; Zhao, J.; Nitoiu, D.; Scott, C.A.; Plagnol, V.; Smith, F.J.; Wilson, N.J.; Cole, C.; Schwartz, M.E.; McLean, W.I.; et al. Loss-of-function mutations in CAST cause peeling skin, leukonychia, acral punctate keratoses, cheilitis, and knuckle pads. Am. J. Hum. Genet. 2015, 96, 440–447. [Google Scholar] [CrossRef] [PubMed]
- Alkhalifah, A.; Chiaverini, C.; Del Giudice, P.; Supsrisunjai, C.; Hsu, C.K.; Liu, L.; Charlesworth, A.; McGrath, J.A.; Lacour, J.P. PLACK syndrome resulting from a new homozygous insertion mutation in CAST. J. Dermatol. Sci. 2017, 88, 256–258. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.H.; Varadi, M.; Tompa, P.; Pauwels, K. Affinity purification of human m-calpain through an intrinsically disordered inhibitor, calpastatin. PLoS ONE 2017, 12, e0174125. [Google Scholar] [CrossRef] [PubMed]
- Kiss, R.; Kovács, D.; Tompa, P.; Perczel, A. Local structural preferences of calpastatin, the intrinsically unstructured protein inhibitor of calpain. Biochemistry 2008, 47, 6936–6945. [Google Scholar] [CrossRef]
- Peng, Z.; Mizianty, M.J.; Kurgan, L. Genome-scale prediction of proteins with long intrinsically disordered regions. Proteins 2014, 82, 145–158. [Google Scholar] [CrossRef]
- Brodsky, S.; Jana, T.; Barkai, N. Order through disorder: The role of intrinsically disordered regions in transcription factor binding specificity. Curr. Opin. Struct. Biol. 2021, 71, 110–115. [Google Scholar] [CrossRef]
- Lu, F.; Lionnet, T. Transcription Factor Dynamics. Cold Spring Harb Perspect. Biol. 2021, 13, a040949. [Google Scholar] [CrossRef]
- Avecilla, A.R.C.; Quiroz, F.G. Cracking the Skin Barrier: Liquid-Liquid Phase Separation Shines under the Skin. JID Innov. 2021, 1, 100036. [Google Scholar] [CrossRef]
- Steinert, P.M.; Marekov, L.N. Direct evidence that involucrin is a major early isopeptide cross-linked component of the keratinocyte cornified cell envelope. J. Biol. Chem. 1997, 272, 2021–2030. [Google Scholar] [CrossRef]
- Takahashi, M.; Tezuka, T.; Katunuma, N. Phosphorylated cystatin α is a natural substrate of epidermal transglutaminase for formation of skin cornified envelope. FEBS Lett. 1992, 308, 79–82. [Google Scholar] [CrossRef]
- DiRusso, C.J.; Dashtiahangar, M.; Gilmore, T.D. Scaffold Proteins as Dynamic Integrators of Biological Processes. J. Biol. Chem. 2022, 298, 102628. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Jiang, L. Liquid-Liquid Phase Separation Bridges Physics, Chemistry, and Biology. Langmuir 2022, 38, 9043–9049. [Google Scholar] [CrossRef] [PubMed]
- Mohanty, P.; Kapoor, U.; Devarajan, D.S.; Phan, T.M.; Rizuan, A.; Mittal, J. Principles Governing the Phase Separation of Multidomain Proteins. Biochemistry 2022, 61, 2443–2455. [Google Scholar] [CrossRef]
- Kalinin, A.E.; Kajava, A.V.; Steinert, P.M. Epithelial barrier function: Assembly and structural features of the cornified cell envelope. Bioessays 2002, 24, 789–800. [Google Scholar] [CrossRef] [PubMed]
- Hohl, D.; Mehrel, T.; Lichti, U.; Turner, M.L.; Roop, D.R.; Steinert, P.M. Characterization of human loricrin. Structure and function of a new class of epidermal cell envelope proteins. J. Biol. Chem. 1991, 266, 6626–6636. [Google Scholar] [CrossRef]
- Holehouse, A.S.; Das, R.K.; Ahad, J.N.; Richardson, M.O.; Pappu, R.V. CIDER: Resources to Analyze Sequence-Ensemble Relationships of Intrinsically Disordered Proteins. Biophys. J. 2017, 112, 16–21. [Google Scholar] [CrossRef]
- Das, R.K.; Pappu, R.V. Conformations of intrinsically disordered proteins are influenced by linear sequence distributions of oppositely charged residues. Proc. Natl. Acad. Sci. USA 2013, 110, 13392–13397. [Google Scholar] [CrossRef]
- Martin, E.W.; Holehouse, A.S.; Grace, C.R.; Hughes, A.; Pappu, R.V.; Mittag, T. Sequence Determinants of the Conformational Properties of an Intrinsically Disordered Protein Prior to and upon Multisite Phosphorylation. J. Am. Chem. Soc. 2016, 138, 15323–15335. [Google Scholar] [CrossRef]
- Yoneda, K.; McBride, O.W.; Korge, B.P.; Kim, I.G.; Steinert, P.M. The cornified cell envelope: Loricrin and transglutaminases. J. Dermatol. 1992, 19, 761–764. [Google Scholar] [CrossRef]
- Vanhoutteghem, A.; Djian, P.; Green, H. Ancient origin of the gene encoding involucrin, a precursor of the cross-linked envelope of epidermis and related epithelia. Proc. Natl. Acad. Sci. USA 2008, 105, 15481–15486. [Google Scholar] [CrossRef]
- Uversky, V.N. Intrinsically disordered proteins in overcrowded milieu: Membrane-less organelles, phase separation, and intrinsic disorder. Curr. Opin. Struct. Biol. 2017, 44, 18–30. [Google Scholar] [CrossRef] [PubMed]
- Chu, X.; Sun, T.; Li, Q.; Xu, Y.; Zhang, Z.; Lai, L.; Pei, J. Prediction of liquid-liquid phase separating proteins using machine learning. BMC Bioinform. 2022, 23, 72. [Google Scholar] [CrossRef] [PubMed]
- Ditlev, J.A.; Case, L.B.; Rosen, M.K. Who’s In and Who’s Out-Compositional Control of Biomolecular Condensates. J. Mol. Biol. 2018, 430, 4666–4684. [Google Scholar] [CrossRef] [PubMed]
- Sevilla, L.M.; Nachat, R.; Groot, K.R.; Klement, J.F.; Uitto, J.; Djian, P.; Määttä, A.; Watt, F.M. Mice deficient in involucrin, envoplakin, and periplakin have a defective epidermal barrier. J. Cell Biol. 2007, 179, 1599–1612. [Google Scholar] [CrossRef]
- Saito, Y.; Kimura, W. Roles of Phase Separation for Cellular Redox Maintenance. Front. Genet. 2021, 12, 691946. [Google Scholar] [CrossRef]
- Huang, S.; Xu, B.; Liu, Y. Calcium promotes α-synuclein liquid-liquid phase separation to accelerate amyloid aggregation. Biochem. Biophys. Res. Commun. 2022, 603, 13–20. [Google Scholar] [CrossRef]
- André, A.A.M.; Spruijt, E. Liquid-Liquid Phase Separation in Crowded Environments. Int. J. Mol. Sci. 2020, 21, 5908. [Google Scholar] [CrossRef]
- Rudraiah, S.; Shamilov, R.; Aneskievich, B.J. TNIP1 reduction sensitizes keratinocytes to post-receptor signalling following exposure to TLR agonists. Cell Signal. 2018, 45, 81–92. [Google Scholar] [CrossRef]
- Mohibi, S.; Zhang, J.; Chen, X. PABPN1, a Target of p63, Modulates Keratinocyte Differentiation through Regulation of p63α mRNA Translation. J. Investig. Dermatol. 2020, 140, 2166–2177.e6. [Google Scholar] [CrossRef]
- Shamilov, R.; Ackley, T.W.; Aneskievich, B.J. Enhanced Wound Healing-and Inflammasome-Associated Gene Expression in TNFAIP3-Interacting Protein 1- (TNIP1-) Deficient HaCaT Keratinocytes Parallels Reduced Reepithelialization. Mediat. Inflamm. 2020, 2020, 5919150. [Google Scholar] [CrossRef]
- Shamilov, R.; Vinogradova, O.; Aneskievich, B.J. The Anti-Inflammatory Protein TNIP1 Is Intrinsically Disordered with Structural Flexibility Contributed by Its AHD1-UBAN Domain. Biomolecules 2020, 10, 1531. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- Klein, J.B.; Knepper, M.A. Protein Mass Spectrometry Made Simple. J. Am. Soc. Nephrol. 2018, 29, 158–51587. [Google Scholar] [CrossRef] [PubMed]
- Leuenberger, P.; Ganscha, S.; Kahraman, A.; Cappelletti, V.; Boersema, P.J.; von Mering, C.; Claassen, M.; Picotti, P. Cell-wide analysis of protein thermal unfolding reveals determinants of thermostability. Science 2017, 355, eaai7825. [Google Scholar] [CrossRef]
- Xue, B.; Oldfield, C.J.; Dunker, A.K.; Uversky, V.N. CDF it all: Consensus prediction of intrinsically disordered proteins based on various cumulative distribution functions. FEBS Lett. 2009, 583, 1469–1474. [Google Scholar] [CrossRef]
- Huang, F.; Oldfield, C.J.; Xue, B.; Hsu, W.L.; Meng, J.; Liu, X.; Shen, L.; Romero, P.; Uversky, V.N.; Dunker, A. Improving protein order-disorder classification using charge-hydropathy plots. BMC Bioinform. 2014, 15 (Suppl. 17), S4. [Google Scholar] [CrossRef]
- Oates, M.E.; Romero, P.; Ishida, T.; Ghalwash, M.; Mizianty, M.J.; Xue, B.; Dosztányi, Z.; Uversky, V.N.; Obradovic, Z.; Kurgan, L.; et al. D²P²: Database of disordered protein predictions. Nucleic. Acids Res. 2013, 41, D508–D516. [Google Scholar] [CrossRef]
- Heger, A.; Holm, L. Rapid automatic detection and alignment of repeats in protein sequences. Proteins 2000, 41, 224–237. [Google Scholar] [CrossRef]
- Yan, J.; Mizianty, M.J.; Filipow, P.L.; Uversky, V.N.; Kurgan, L. RAPID: Fast and accurate sequence-based prediction of intrinsic disorder content on proteomic scale. Biochim. Biophys. Acta 2013, 1834, 1671–1680. [Google Scholar] [CrossRef]
- Bolognesi, B.; Lorenzo Gotor, N.; Dhar, R.; Cirillo, D.; Baldrighi, M.; Tartaglia, G.G.; Lehner, B. A Concentration-Dependent Liquid Phase Separation Can Cause Toxicity upon Increased Protein Expression. Cell Rep. 2016, 16, 222–231. [Google Scholar] [CrossRef]
- Hardenberg, M.; Horvath, A.; Ambrus, V.; Fuxreiter, M.; Vendruscolo, M. Widespread occurrence of the droplet state of proteins in the human proteome. Proc. Natl. Acad. Sci USA 2020, 117, 33254–33262. [Google Scholar] [CrossRef] [PubMed]
- Vernon, R.M.; Chong, P.A.; Tsang, B.; Kim, T.H.; Bah, A.; Farber, P.; Lin, H.; Forman-Kay, J.D. Pi-Pi contacts are an overlooked protein feature relevant to phase separation. eLife 2018, 7, e31486. [Google Scholar] [CrossRef] [PubMed]
- Geiger, F.; Acker, J.; Papa, G.; Wang, X.; E Arter, W.; Saar, K.L.; A Erkamp, N.; Qi, R.; Bravo, J.P.; Strauss, S.; et al. Liquid-liquid phase separation underpins the formation of replication factories in rotaviruses. EMBO J. 2021, 40, e107711. [Google Scholar] [CrossRef] [PubMed]
- Saar, K.L.; Morgunov, A.S.; Qi, R.; Arter, W.E.; Krainer, G.; Lee, A.A.; Knowles, T.P.J. Learning the molecular grammar of protein condensates from sequence determinants and embeddings. Proc. Natl. Acad. Sci. USA 2021, 118, e2019053118. [Google Scholar] [CrossRef] [PubMed]



| Protein | UniProt # | AA | Protein Role | PONDR-FIT (0–1) | RAPID Disorder % | SLIDER IDR (0–1) | catGRAN a |
|---|---|---|---|---|---|---|---|
| Involucrin b | P07476 | 585 | CE component, cornification c | 0.812 | 65.81 | 0.917 | 0.821 |
| Ladinin-1 | O00515 | 517 | ECM and cadherin binding | 0.774 | 68.28 | 0.929 | 0.333 |
| BP180 | Q9UMD9 | 1497 | Transmembrane attachment to ECM | 0.736 | 40.75 | 0.913 | 3.184 |
| Cornifin A d | P35321 | 89 | CE component, cornification | 0.551 | 79.78 | 0.848 | −2.527 |
| AHNAK/Desmoyokin d | Q09666 | 5890 | Scaffolding, nuclear-cytoplasmic shuttling | 0.494 | 48.37 | 0.851 | 2.942 |
| Periplakin | O60437 | 1756 | CE component, scaffolding | 0.491 | 30.30 | 0.903 | 0.857 |
| Galectin-3 d | P17931 | 250 | Cell–cell, cell–ECM interaction, splicing | 0.501 | 18.00 | 0.487 | 1.571 |
| Protein | UniProt # | AA | Protein Role | PONDR-FIT (0–1) | RAPID Disorder % | SLIDER IDR (0–1) | catGRAN a |
|---|---|---|---|---|---|---|---|
| Prelamin A/C | P02545 | 664 | Nuclear membrane support, scaffolding | 0.620 | 34.79 | 0.9015 | 0.904 |
| TAB182 | Q9C0C2 | 1729 | Nuclear and cyto-plasmic RNA-binding | 0.734 | 51.13 | 0.9281 | 1.975 |
| Calpastatin | P20810 | 708 | Calpain protease inhibitor | 0.804 | 78.39 | 0.9565 | 1.010 |
| SERBP1 | Q8NC51 | 408 | mRNA stability, PML formation | 0.751 | 60.54 | 0.9049 | 2.829 |
| Nucleolin 1 | P19338 | 710 | RNA-binding, pre-rRNA transcription | 0.633 | 62.39 | 0.9357 | 2.495 |
| ADIRF | Q15847 | 76 | Transcription factor | 0.810 | 50.00 | 0.6675 | −0.330 |
| Protein | UniProt # | AA | Kappa | Omega | FCR | NCPR | Hydropathy | Dis. Prom. | PR |
|---|---|---|---|---|---|---|---|---|---|
| Loricrin | P23490 | 312 | 0.252 | 1.787 | 0.026 | 0.019 | 4.244 | 0.827 | 1 |
| Hornerin | Q86YZ3 | 2850 | 0.162 | 0.823 | 0.099 | 0.029 | 3.136 | 0.911 | 1 |
| LC3EA | Q5TA76 | 89 | 0.309 | 0.420 | 0.124 | 0.079 | 3.602 | 0.775 | 1 |
| SPR2A a | P35326 | 72 | 0.151 | 0.115 | 0.139 | 0.083 | 3.193 | 0.764 | 1 |
| Filaggrin 2 | Q5D862 | 2391 | 0.227 | 0.621 | 0.144 | 0.003 | 3.134 | 0.883 | 1 |
| KPRP a | Q5T749 | 579 | 0.185 | 0.172 | 0.174 | 0.036 | 3.711 | 0.710 | 1 |
| Elafin | P19957 | 117 | 0.124 | 0.227 | 0.197 | 0.060 | 4.664 | 0.598 | 1 |
| Filaggrin | P20930 | 4061 | 0.164 | 0.235 | 0.240 | 0.005 | 2.827 | 0.919 | 1 |
| S10AB | P31949 | 105 | 0.149 | 0.213 | 0.248 | 0 | 4.160 | 0.619 | 1 |
| Sciellin | O95171 | 688 | 0.143 | 0.134 | 0.285 | 0.035 | 3.455 | 0.657 | 2 |
| Cystatin A | P01040 | 98 | 0.146 | 0.146 | 0.286 | −0.020 | 3.822 | 0.633 | 2 |
| S10AA | P60903 | 97 | 0.098 | 0.231 | 0.289 | 0 | 4.139 | 0.598 | 2 |
| Involucrin | P07476 | 585 | 0.144 | 0.060 | 0.294 | −0.130 | 2.996 | 0.776 | 2 |
| Envoplakin | Q92817 | 2033 | 0.129 | 0.116 | 0.323 | −0.004 | 3.700 | 0.716 | 2 |
| Periplakin | O60437 | 1756 | 0.138 | 0.086 | 0.362 | −0.031 | 3.518 | 0.695 | 3 |
| Trichohyalin | Q07283 | 1943 | 0.116 | 0.110 | 0.581 | −0.021 | 1.995 | 0.838 | 3 |
| Protein | UniProt # | AA | PONDR-FIT (0–1) | RAPID Disorder % | catGRAN. a | FuzDrop pLLPS (0–1) b | P Score c | DeePhase d | PSPred. d |
|---|---|---|---|---|---|---|---|---|---|
| Hornerin | Q86YZ3 | 2850 | 0.923 | 57.72 | 5.572 | 1.000 | 11.91 | 0.88 | 0.9527 |
| Filaggrin 2 | Q5D862 | 2391 | 0.896 | 49.14 | 4.418 | 0.9999 | 12.03 | 0.87 | 0.9904 |
| Filaggrin | P20930 | 4061 | 0.895 | 55.58 | 3.553 | 1.000 | 4.73 | 0.83 | 0.9908 |
| Loricrin | P23490 | 312 | 0.840 | 62.82 | 7.808 | 0.9985 | 12.31 | 0.81 | 0.9958 |
| Involucrin | P07476 | 585 | 0.812 | 65.81 | 0.821 | 0.9921 | 2.18 | 0.78 | 0.4945 |
| Trichohyalin | Q07283 | 1943 | 0.753 | 73.60 | 1.312 | 0.9947 | 0.66 | 0.72 | 0.2190 |
| LCE3A | Q5TA76 | 89 | 0.625 | 18.18 | −0.779 | 0.9978 | NA | 0.59 | 0.9596 |
| Sciellin | O95171 | 688 | 0.620 | 41.57 | 1.306 | 0.9527 | 2.10 | 0.78 | 0.9546 |
| KPRP | Q5T749 | 579 | 0.571 | 29.02 | −0.591 | 0.9867 | 5.72 | 0.82 | 0.9096 |
| SPR2A | P35326 | 72 | 0.530 | 65.28 | −3.342 | 0.9868 | NA | 0.58 | 0.7971 |
| Periplakin | O60437 | 1756 | 0.491 | 30.30 | 0.857 | 0.6168 | 0.38 | 0.59 | 0.2852 |
| Envoplakin | Q92817 | 2033 | 0.478 | 26.41 | 0.780 | 0.8022 | 2.00 | 0.81 | 0.6487 |
| S100-A11 | P31949 | 105 | 0.374 | 17.14 | −0.481 | 0.1275 | NA | 0.11 | 0.0125 |
| Cystatin A | P01040 | 98 | 0.363 | 27.55 | 0.410 | 0.1673 | NA | 0.10 | 0.0050 |
| Elafin | P19957 | 117 | 0.350 | 30.77 | −0.180 | 0.2376 | NA | 0.17 | 0.0574 |
| S100-A10 | P60903 | 97 | 0.336 | 16.49 | −0.373 | 0.2124 | NA | 0.08 | 0.0033 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Samulevich, M.L.; Shamilov, R.; Aneskievich, B.J. Thermostable Proteins from HaCaT Keratinocytes Identify a Wide Breadth of Intrinsically Disordered Proteins and Candidates for Liquid–Liquid Phase Separation. Int. J. Mol. Sci. 2022, 23, 14323. https://doi.org/10.3390/ijms232214323
Samulevich ML, Shamilov R, Aneskievich BJ. Thermostable Proteins from HaCaT Keratinocytes Identify a Wide Breadth of Intrinsically Disordered Proteins and Candidates for Liquid–Liquid Phase Separation. International Journal of Molecular Sciences. 2022; 23(22):14323. https://doi.org/10.3390/ijms232214323
Chicago/Turabian StyleSamulevich, Michael L., Rambon Shamilov, and Brian J. Aneskievich. 2022. "Thermostable Proteins from HaCaT Keratinocytes Identify a Wide Breadth of Intrinsically Disordered Proteins and Candidates for Liquid–Liquid Phase Separation" International Journal of Molecular Sciences 23, no. 22: 14323. https://doi.org/10.3390/ijms232214323
APA StyleSamulevich, M. L., Shamilov, R., & Aneskievich, B. J. (2022). Thermostable Proteins from HaCaT Keratinocytes Identify a Wide Breadth of Intrinsically Disordered Proteins and Candidates for Liquid–Liquid Phase Separation. International Journal of Molecular Sciences, 23(22), 14323. https://doi.org/10.3390/ijms232214323






