Study of Degradation Kinetics and Structural Analysis of Related Substances of Ceftobiprole by HPLC with UV and MS/MS Detection
Abstract
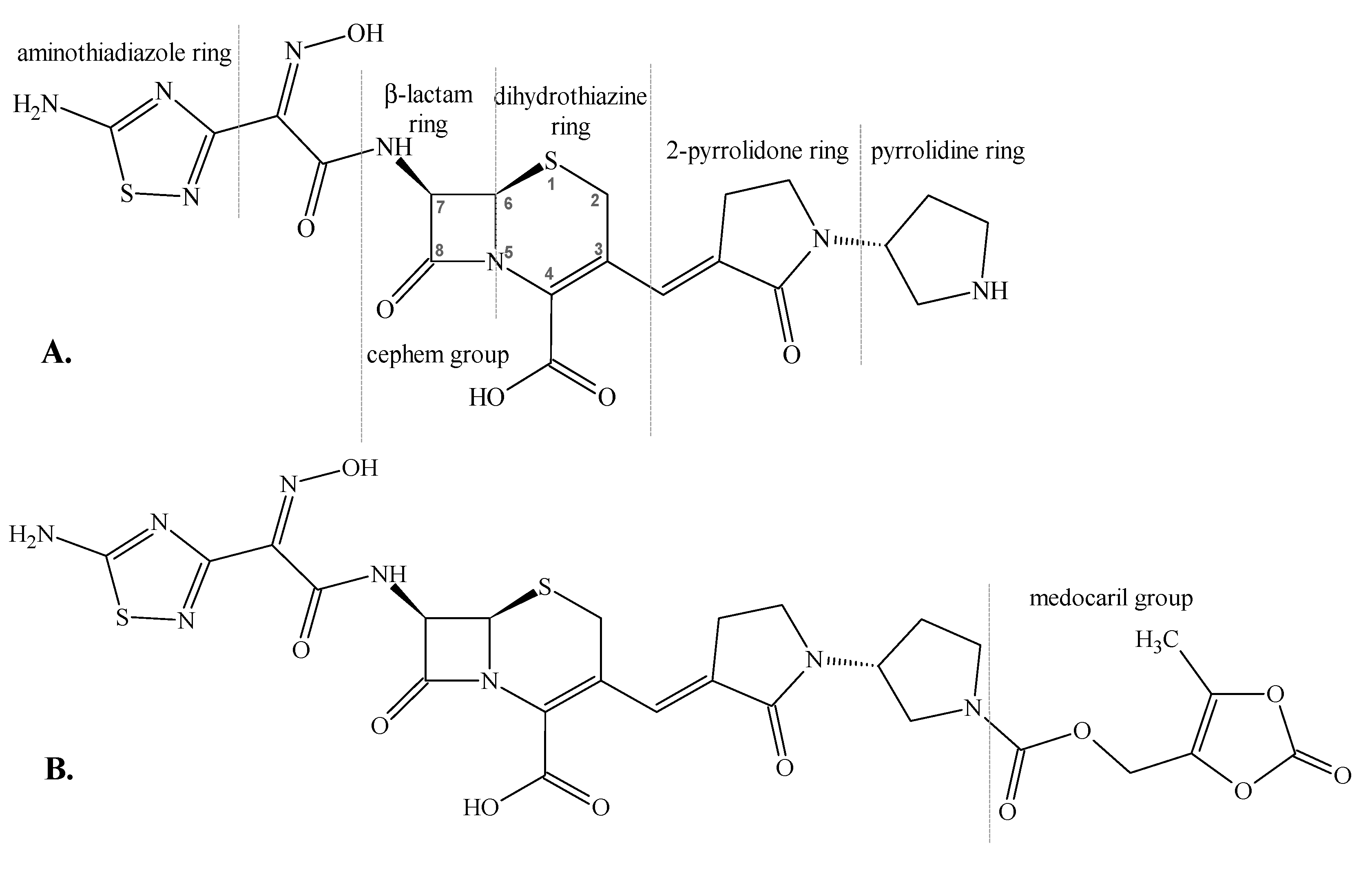
1. Introduction
2. Results and Discussion
2.1. Method Optimisation
2.2. Kinetics of Ceftobiprole Degradation in Solution-State Studies
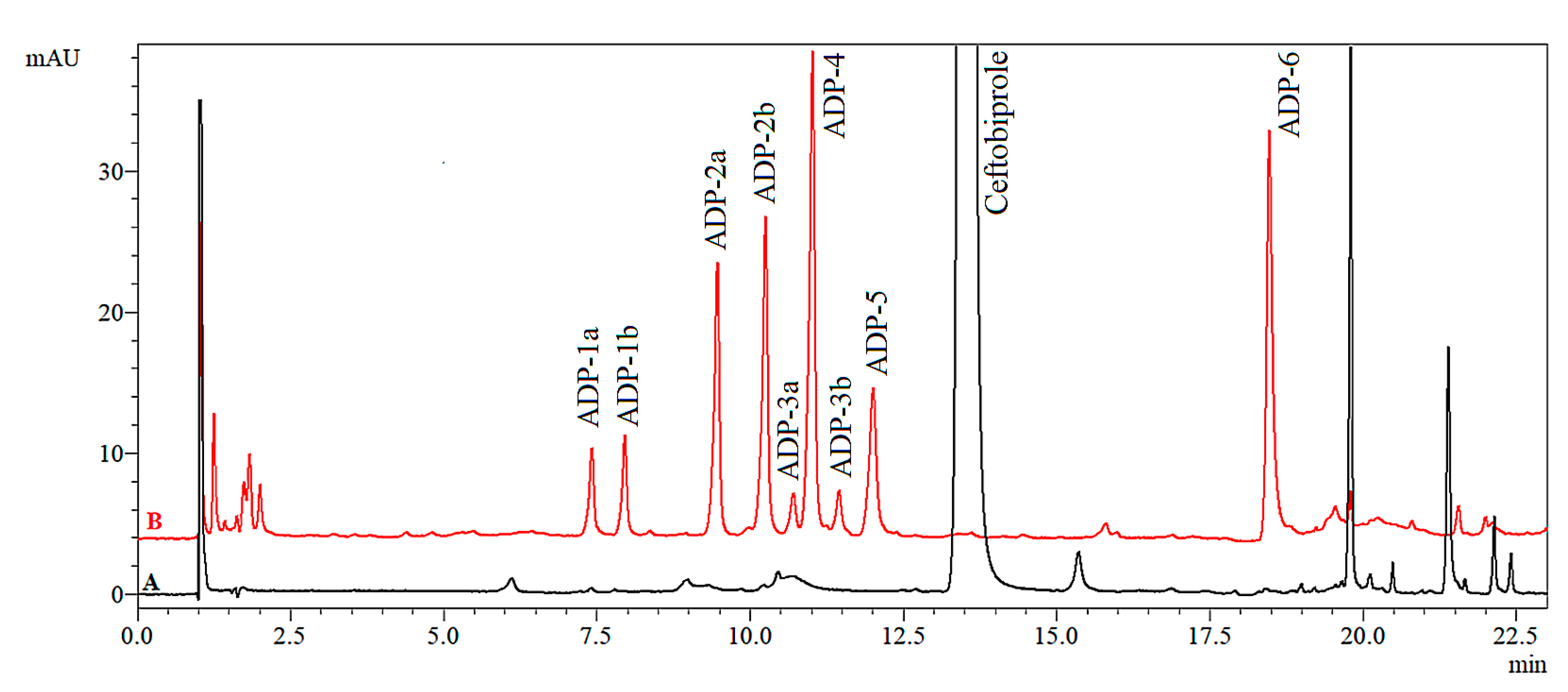
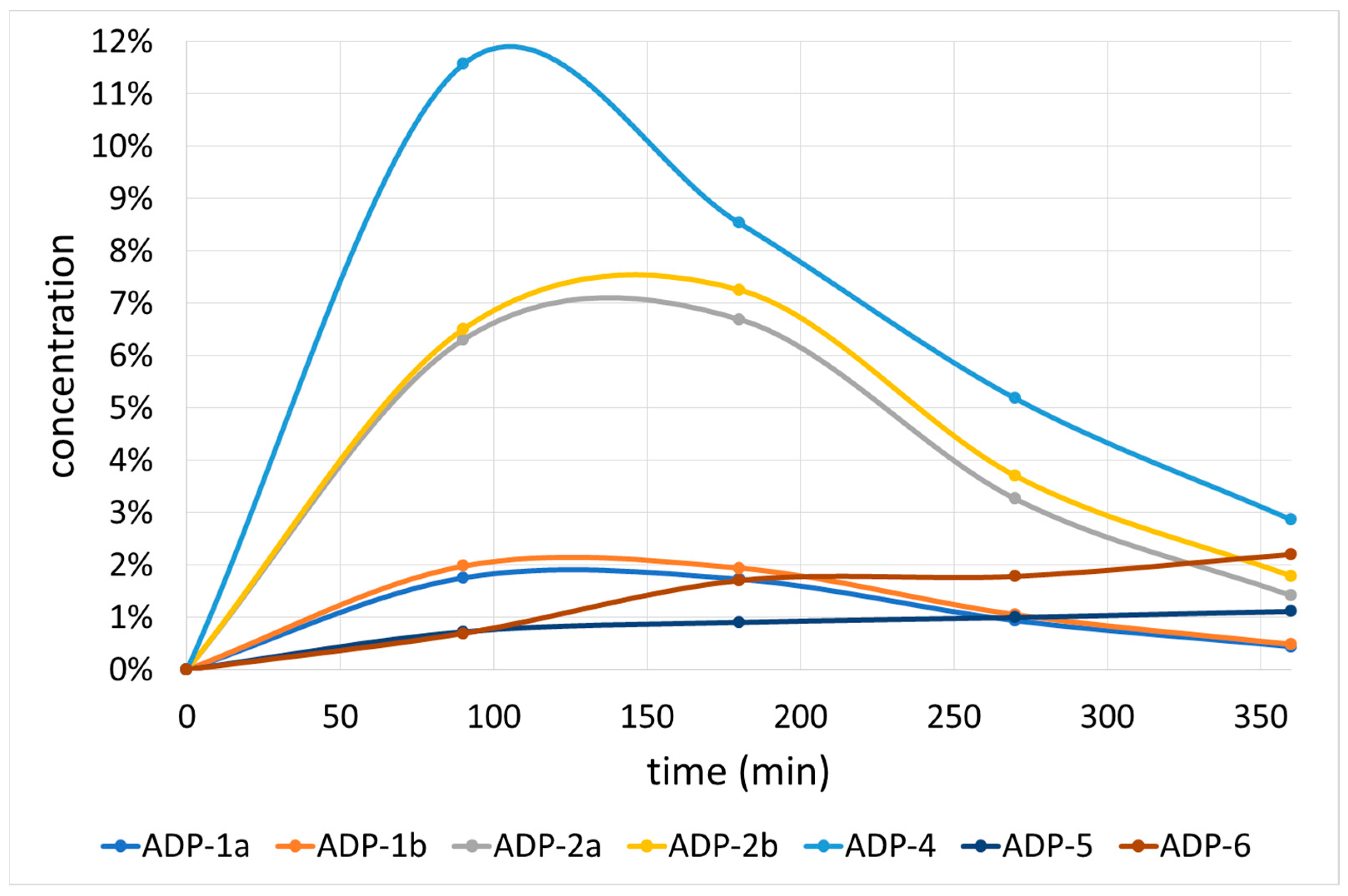
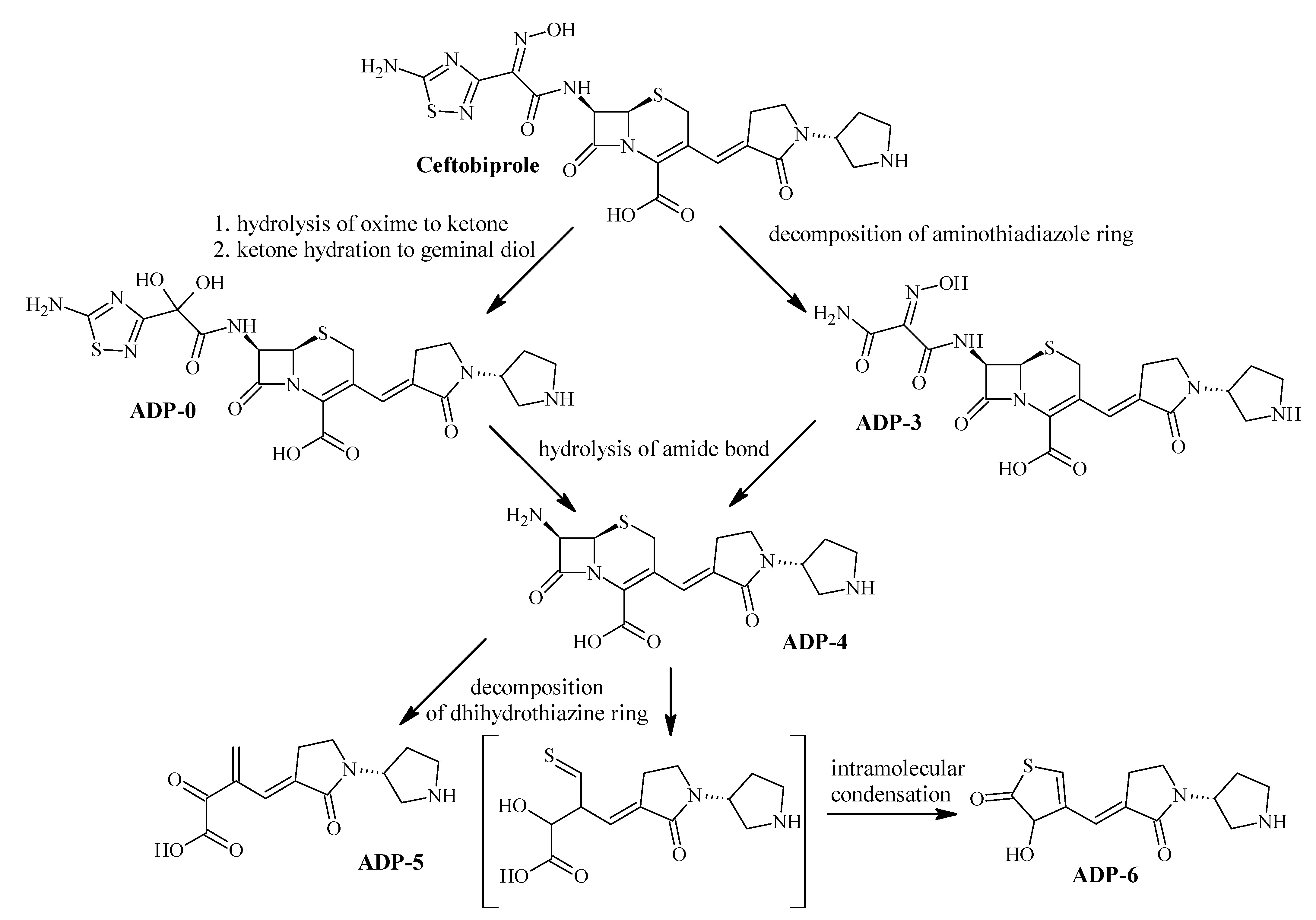
2.2.1. Acidic Degradation Products (ADPs)
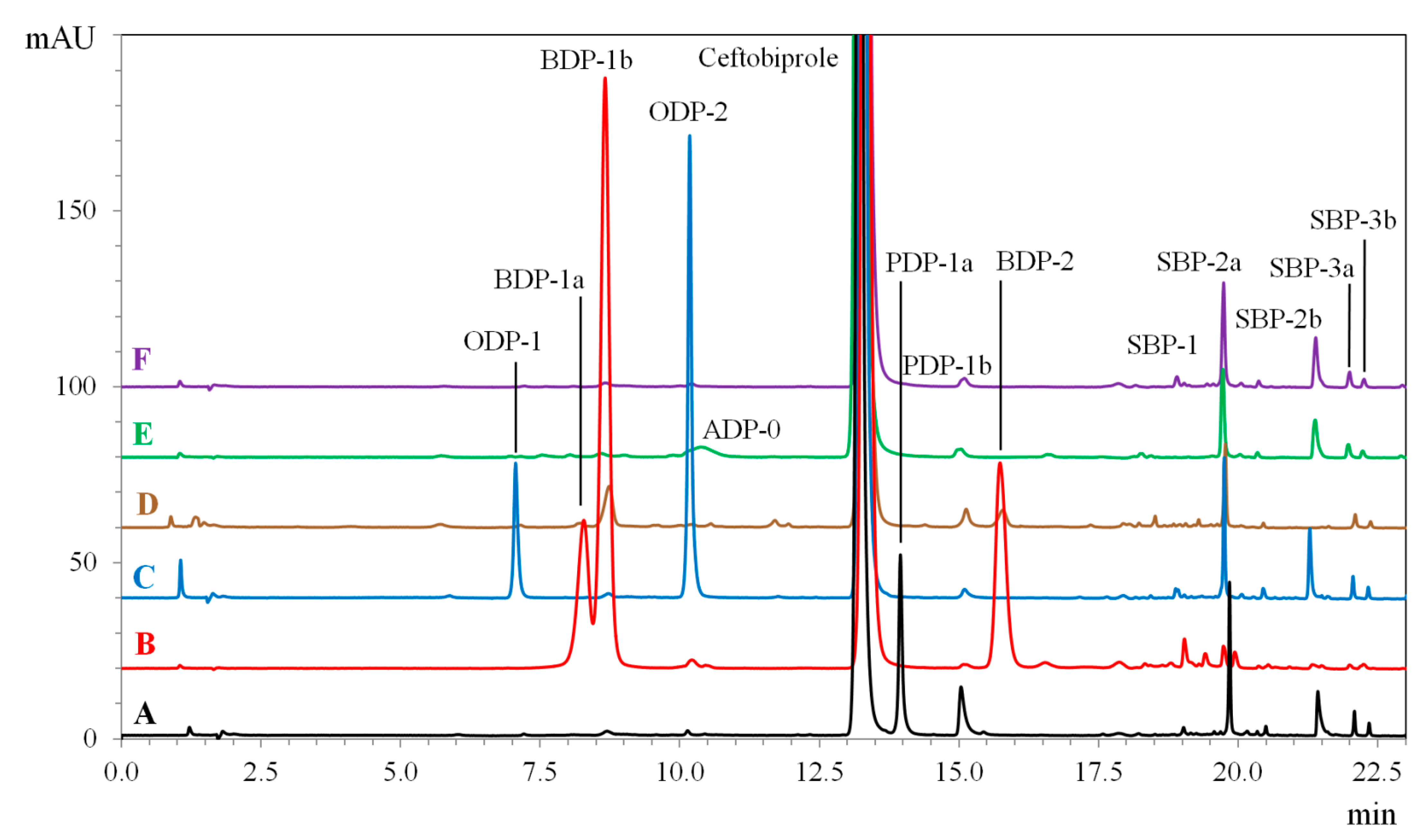
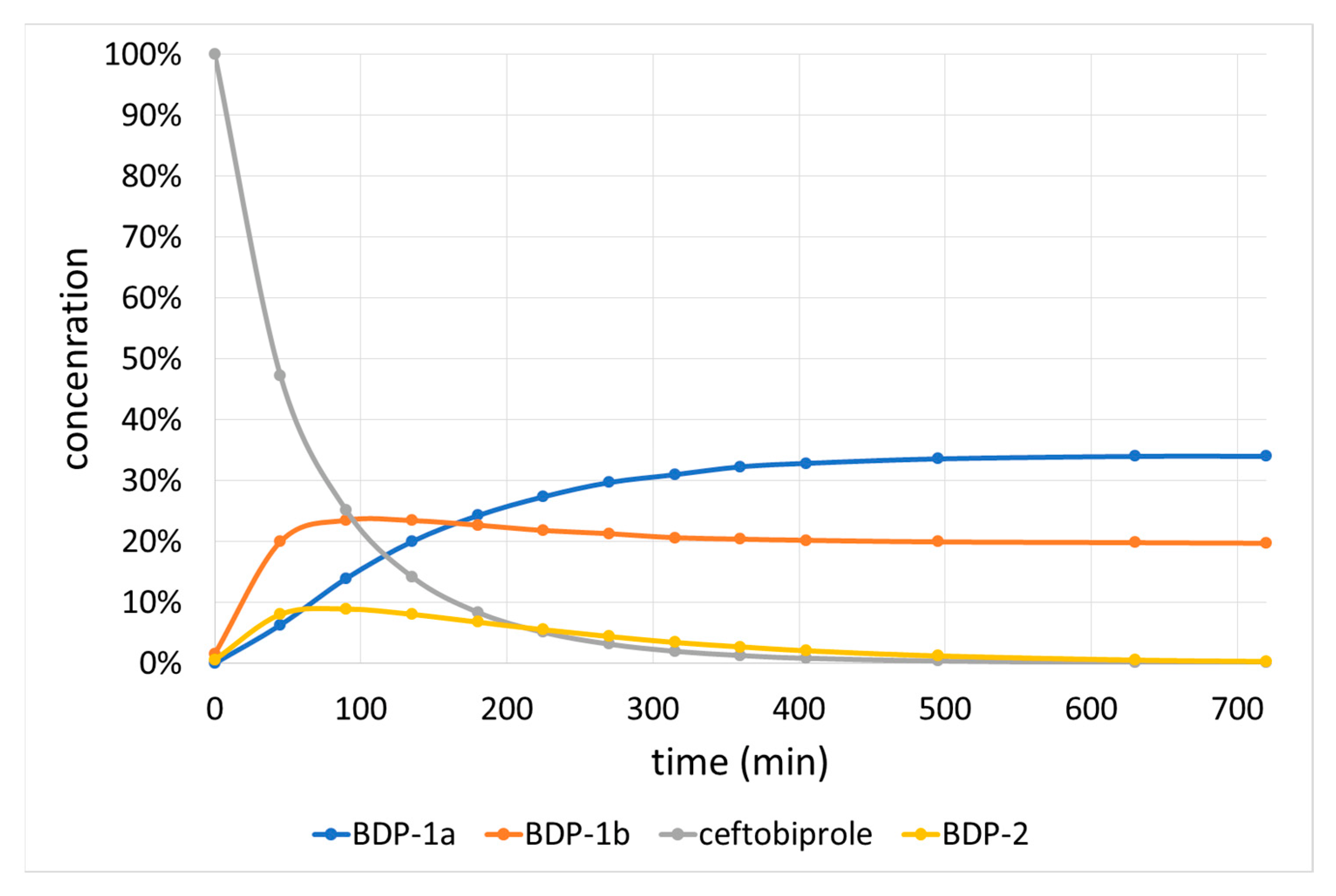
2.2.2. Basic Degradation Products (BDPs)
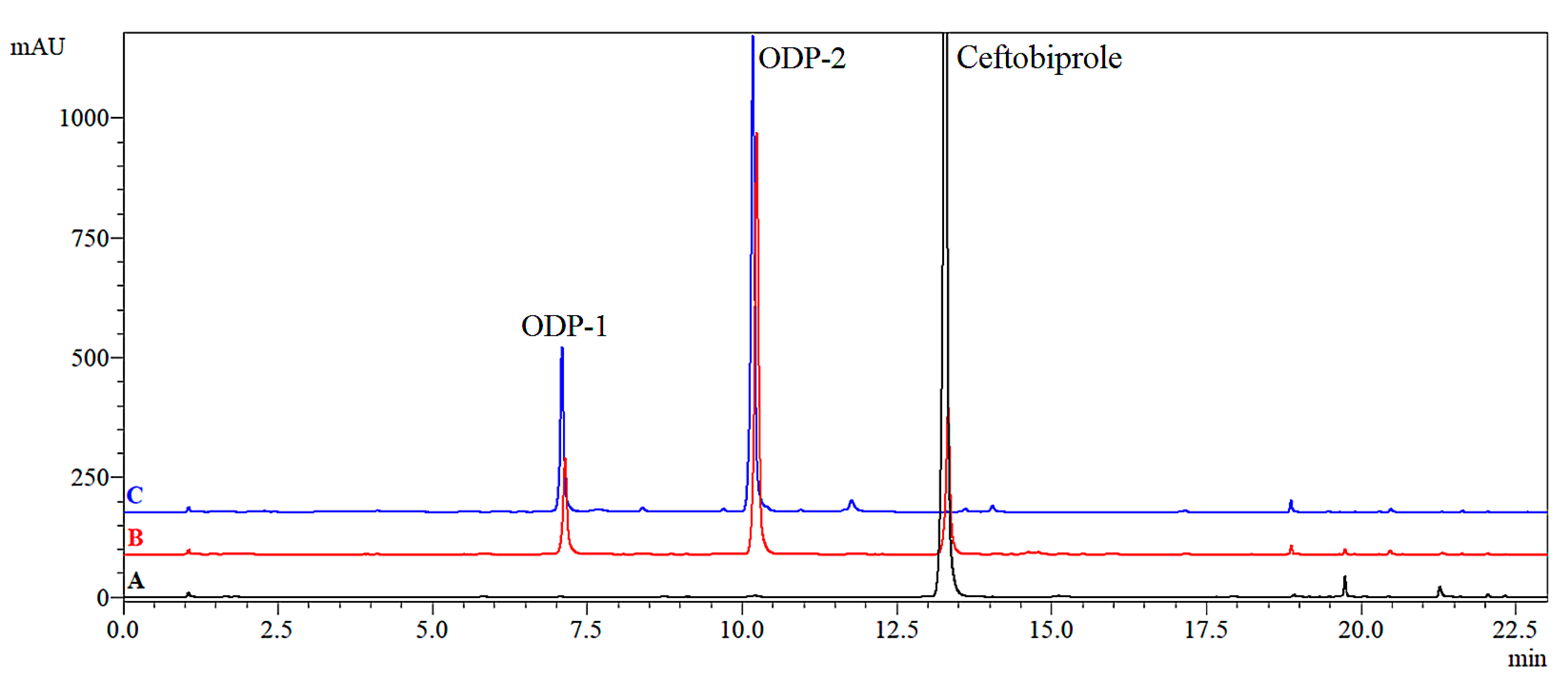
2.2.3. Oxidative Degradation Products (ODPs)
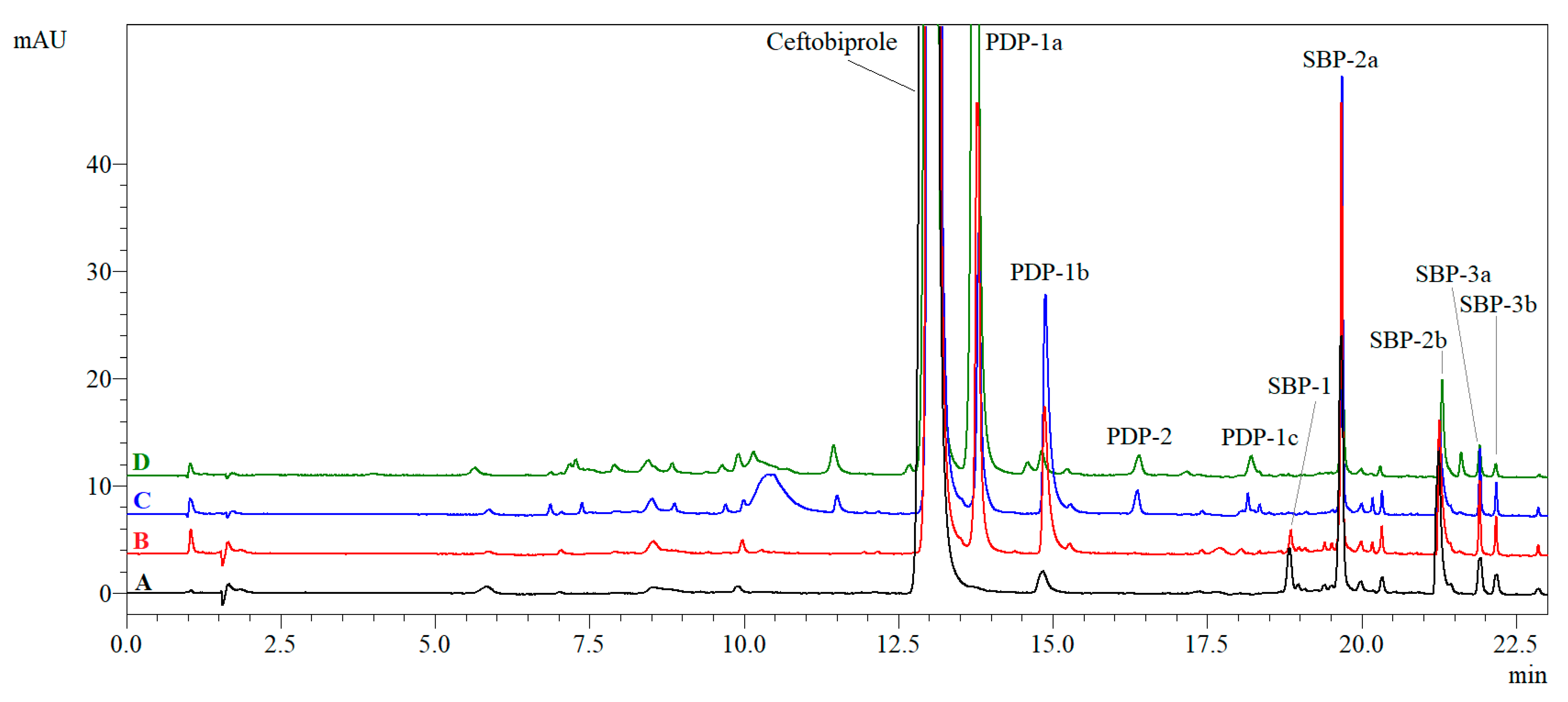
2.2.4. Photolytic Degradation Products (PDPs)
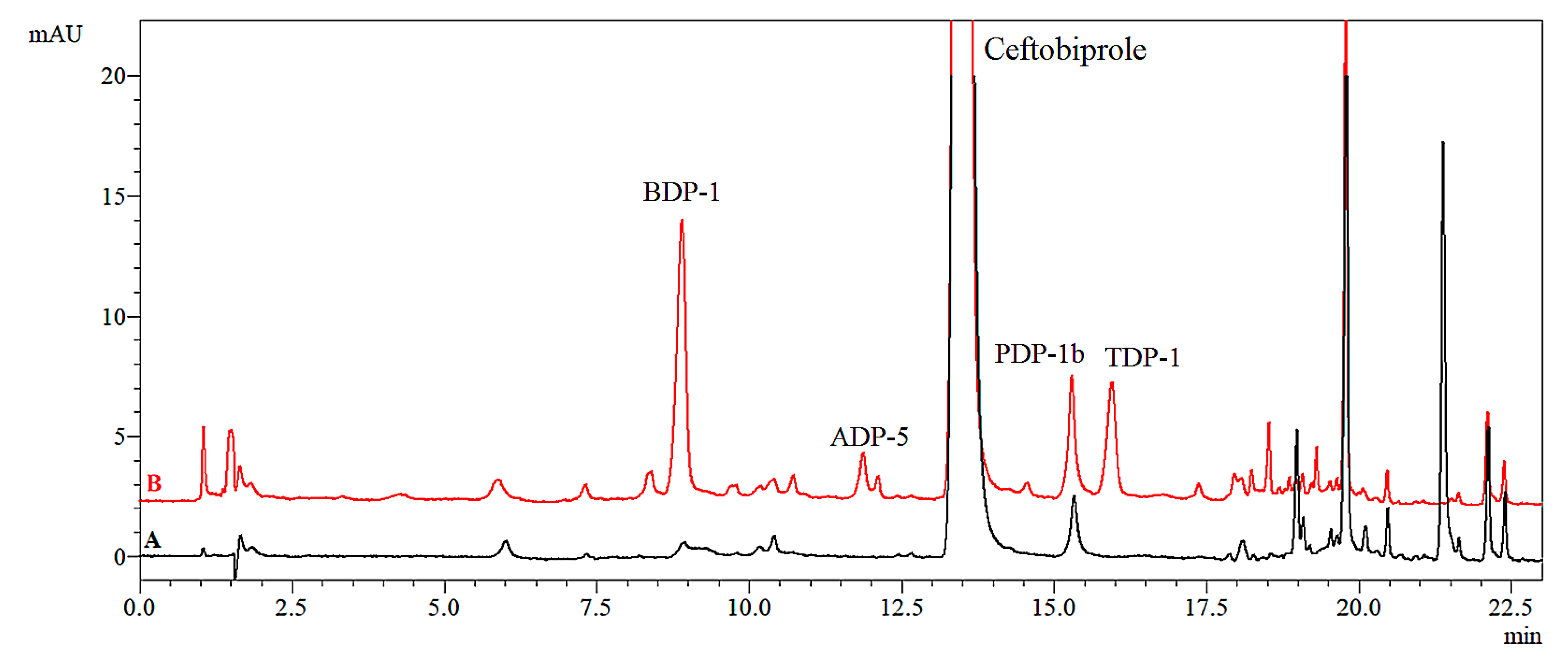
2.2.5. Thermolytic Degradation
2.3. Solid-State Degradation Kinetics
2.4. Mass Spectrometric Characterisation of the Impurities of Ceftobiprole
2.4.1. Identification of Synthesis By-Products (SBPs)
2.4.2. Identification of ADPs
2.4.3. Identification of BDPs
2.4.4. Identification of ODPs
2.4.5. Identification of PDPs and TDPs
2.4.6. Identification of ODPs Formed in Solid-State Degradation
3. Materials and Methods
3.1. Materials
3.2. Solution-State Degradation Study
3.2.1. Acidic Hydrolysis with Temperature
3.2.2. Alkaline Hydrolysis
3.2.3. Oxidative Degradation
3.2.4. Photolytic Degradation
3.2.5. Thermolytic Degradation
3.3. Solid-State Degradation Study
- -
- At 50 and 60 °C for 7 and 21 days without control of humidity (Memmert oven, Memmert, GmbH + Co. KG, Schwabach FRG, Germany);
- -
- At 50 ± 2 °C/75 ± 5% RH (AtomControl Memmert, GmbH + Co. KG, Schwabach FRG, Germany), tested at seven-day intervals over a period of 21 days;
- -
- Under illumination with UV at 366 nm at room temperature, tested at seven-day intervals over a period of 21 days;
- -
- In a chamber simulating natural sunlight at an intensity of 5260 ± 316 lx for 10 and 21 days, at room temperature.
3.4. HPLC/UV Analysis
3.5. LC/MS/MS Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Reygaert, W.C. Ceftobiprole: An Emerging Therapeutic Option for Resistant and Complicated Infections. Clin. Med. Insights Ther. 2011, 3, 57–66. [Google Scholar] [CrossRef]
- Karpiuk, I.; Michalska, K.; Bus, K.; Kiljan, M.; Tyski, S. Identification and determination of related substances of ceftaroline fosamil in medicinal product by high performance liquid chromatography with diode array detection and tandem mass spectrometry. J. Pharm. Biomed. Anal. 2017, 145, 651–660. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.; Sheng, H.; Saurí, J.; Xiang, R.; Martin, G. Structural elucidation of a dimeric impurity in the process development of ceftolozane using LC/HRMS and 2D-NMR. J. Pharm. Biomed. Anal. 2019, 174, 242–247. [Google Scholar] [CrossRef] [PubMed]
- Jamieson, C.; Drummond, F.; Hills, T.; Ozolina, L.; Gilchrist, M.; Seaton, R.A.; Santillo, M.; Wilkinson, A.-S.; Allwood, M.C. Assessment of ceftolozane/tazobactam stability in elastomeric devices and suitability for continuous infusion via outpatient parenteral antimicrobial therapy. JAC-Antimicrob. Resist. 2021, 3, dlab141. [Google Scholar] [CrossRef]
- Hamed, K.; Engelhardt, M.; Jones, M.E.; Saulay, M.; Holland, T.L.; Seifert, H.; Fowler, V.G., Jr. Ceftobiprole versus daptomycin in Staphylococcus aureus bacteremia: A novel protocol for a double-blind, Phase III trial. Future Microbiol. 2020, 15, 35–48. [Google Scholar] [CrossRef] [PubMed]
- Taylor, P. Basilea Preps Key US Filing for MRSA Antibiotic Ceftobiprole. Available online: https://pharmaphorum.com/news/basilea-preps-key-us-filing-for-mrsa-antibiotic-ceftobiprole/ (accessed on 25 November 2022).
- Overcash, J.S.; Kim, C.; Keech, R.; Gumenchuk, I.; Ninov, B.; Gonzalez-Rojas, Y.; Waters, M.; Simeonov, S.; Engelhardt, M.; Saulay, M.; et al. Ceftobiprole Compared With Vancomycin Plus Aztreonam in the Treatment of Acute Bacterial Skin and Skin Structure Infections: Results of a Phase 3, Randomized, Double-blind Trial (TARGET). Clin. Infect. Dis. 2021, 73, e1507–e1517. [Google Scholar] [CrossRef] [PubMed]
- Lupia, T.; Pallotto, C.; Corcione, S.; Boglione, L.; De Rosa, F.G. Ceftobiprole Perspective: Current and Potential Future Indications. Antibiotics 2021, 10, 170. [Google Scholar] [CrossRef] [PubMed]
- European Medicines Agency Decision P/0311/2020. Available online: https://www.ema.europa.eu/en/medicines/human/paediatric-investigation-plans/emea-000205-pip02-11-m04 (accessed on 11 August 2022).
- Electronic Medicines Compendium (emc). Zevtera 500 mg Powder for Concentrate for Solution for Infusion—Summary of Product Characteristics. Available online: https://www.medicines.org.uk/emc/product/9164 (accessed on 25 November 2022).
- Zhanel, G.G.; Lam, A.; Schweizer, F.; Thomson, K.; Walkty, A.; Rubinstein, E.; Gin, A.S.; Hoban, D.J.; Noreddin, A.M.; Karlowsky, J.A. Ceftobiprole. A Review of a Broad-Spectrum and Anti-MRSA Cephalosporin. Am. J. Clin. Dermatol. 2008, 9, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Giacobbe, D.R.; De Rosa, F.G.; Del Bono, V.; Grossi, P.A.; Pea, F.; Petrosillo, N.; Rossolini, G.M.; Tascini, C.; Tumbarello, M.; Viale, P.; et al. Ceftobiprole: Drug evaluation and place in therapy. Expert Rev. Anti-Infect. Ther. 2019, 17, 689–698. [Google Scholar] [CrossRef] [PubMed]
- International Council on Harmonisation of Technical Requirements for Registration of Pharmaceuticals for Human Use. ICH Topic Q1A (R2). Stability Testing of New Drug Substances and Products. Available online: https://database.ich.org/sites/default/files/Q1A%28R2%29%20Guideline.pdf (accessed on 25 November 2022).
- Baertschi, S.W.; Jansen, P.J.; Alsante, K.M. Stress testing: A predictive tool. In Pharmaceutical Stress Testing: Predicting Drug Degradation, 2nd ed.; Baertschi, S.W., Alsante, K.M., Reed, R.A., Eds.; CRC Press: Boca Raton, FL, USA, 2011; pp. 10–48. [Google Scholar]
- Binert-Kusztal, Ż.; Starek, M.; Żandarek, J.; Dąbrowska, M. Development of TLC Chromatographic-Densitometric Procedure for Qualitative and Quantitative Analysis of Ceftobiprole. Processes 2021, 9, 708. [Google Scholar] [CrossRef]
- Binert-Kusztal, Ż.; Żandarek, J.; Starek, M.; Dąbrowska, M. Optimizations of the Conditions for Ceftobiprole Determination in a Complex Matrix. Processes 2022, 10, 2196. [Google Scholar] [CrossRef]
- Lima, B.; Bodeau, S.; Quinton, M.-C.; Leven, C.; Lemaire-Hurtel, A.-S.; Bennis, Y. Validation and Application of an HPLC-DAD Method for Routine Therapeutic Drug Monitoring of Ceftobiprole. Antimicrob. Agents Chemother. 2019, 63, e00515-19. [Google Scholar] [CrossRef] [PubMed]
- Llopis, B.; Bleibtreu, A.; Schlemmer, D.; Robidou, P.; Paccoud, O.; Tissot, N.; Noé, G.; Junot, H.; Luyt, C.-É.; Funck-Brentano, C.; et al. Simple and accurate quantitative analysis of cefiderocol and ceftobiprole in human plasma using liquid chromatography-isotope dilution tandem mass spectrometry: Interest for their therapeutic drug monitoring and pharmacokinetic studies. Clin. Chem. Lab. Med. (CCLM) 2021, 59, 1800–1810. [Google Scholar] [CrossRef]
- Sahoo, B.M.; Banik, B.K. Therapeutic Potentials of β-Lactam. In Synthetic Approaches to Nonaromatic Nitrogen Heterocycles; Phillips, A.M.M.M.F., Ed.; John Wiley & Sons Ltd.: New York, NY, USA, 2020; pp. 59–88. [Google Scholar]
- Chemicalize. Available online: https://chemicalize.com/welcome (accessed on 25 November 2022).
- Michalska, K.; Widyńska, W.; Bus, K.; Bocian, W.; Tyski, S. The solution and solid-state degradation study followed by identification of tedizolid related compounds in medicinal product by high performance liquid chromatography with diode array and tandem mass spectrometry detection. J. Pharm. Biomed. Anal. 2021, 194, 113783. [Google Scholar] [CrossRef] [PubMed]
- Baptista, L.; Clemente da Silva, E.; Arbilla, G. Oxidation mechanism of dimethyl sulfoxide (DMSO) by OH radical in liquid phase. Phys. Chem. Chem. Phys. 2008, 10, 6867–6879. [Google Scholar] [CrossRef] [PubMed]
- Fasani, E.; Albini, A. Photostability stress testing. In Pharmaceutical Stress Testing: Predicting Drug Degradation, 2nd ed.; Baertschi, S.W., Alsante, K.M., Reed, R.A., Eds.; CRC Press: Boca Raton, FL, USA, 2011; pp. 192–217. [Google Scholar]
- International Council on Harmonisation of Technical Requirements for Registration of Pharmaceuticals for Human Use. ICH Topic Q1B. Photostability Testing of New Active Substances and Medicinal Products. Available online: https://database.ich.org/sites/default/files/Q1B%20Guideline.pdf (accessed on 25 November 2022).
- Hughes, D.L. Patent Review of Manufacturing Routes to Fifth-Generation Cephalosporin Drugs. Part 2, Ceftaroline Fosamil and Ceftobiprole Medocaril. Org. Process Res. Dev. 2017, 21, 800–815. [Google Scholar] [CrossRef]
- Yang, Y. Side Reactions in Peptide Synthesis; Academic Press: Oxford, UK, 2016. [Google Scholar]
- Young, P.R.; Howell, L.G.; Owen, T.C. Pyruvamide semicarbazone formation. Kinetics, mechanism, and pertinence to pyruvamide-dependent histidine decarboxylase. J. Am. Chem. Soc. 1975, 97, 6544–6551. [Google Scholar] [CrossRef] [PubMed]
- Baertschi, S.W.; Alsante, K.M.; Santafianos, D. Stress testing: The chemistry of drug degradation. In Pharmaceutical Stress Testing: Predicting Drug Degradation, 2nd ed.; Baertschi, S.W., Alsante, K.M., Reed, R.A., Eds.; CRC Press: Boca Raton, FL, USA, 2011; pp. 49–141. [Google Scholar]
- Council of Europe. Cefixime. Ph. Eur. 10.5, 1188 (04/2018). In European Pharmacopoeia; Council of Europe: Strasbourg, France, 2022. [Google Scholar]
- Council of Europe. Cefprozil. Ph Eur. 10.5, 2342 (01/2017). In European Pharmacopoeia; Council of Europe: Strasbourg, France, 2022. [Google Scholar]
- Paul Harmon, G.B. Oxidative Susceptibility Testing. In Pharmaceutical Stress Testing: Predicting Drug Degradation, 2nd ed.; Steven, W., Baertschi, K.M.A., Robert, A.R., Eds.; CRC Press: Boca Raton, FL, USA, 2011; pp. 168–191. [Google Scholar]
- Council of Europe. Cefradine. Ph. Eur. 10.5, 0814 (01/2014). In European Pharmacopoeia; Council of Europe: Strasbourg, France, 2022. [Google Scholar]




| Column | Kinetex Biphenyl 150 × 2.1 mm; 1.7 μm | |
| Column temperature | 40 °C | |
| Autosampler temperature | 4 °C—for method optimization 25 °C—for kinetic studies | |
| Mobile phase | A: 20 mM ammonium acetate buffer, pH = 5.8 B: ACN | |
| Flow rate | 0.3 mL/min | |
| Gradient | Time (min) | %B |
| 0 | 0 | |
| 15 | 10 | |
| 28 | 90 | |
| 33 | 90 | |
| 33.1 | 0 | |
| 45 | 0 | |
| Injection volume | 1 μL—for kinetic studies 20 µL—for LC/MS studies | |
| Detection wavelength | 320 nm | |
| Time Point | 10% DMSO | 0.1 M HCl |
|---|---|---|
| % Drug Remaining | % Drug Remaining | |
| 0 | 100.0% | 100.0% |
| 3 h | 99.5% | 98.0% |
| 6 h | 99.4% | 96.9% |
| 12 h | 99.2% | 94.3% |
| 24 h | 98.9% | 89.6% |
| Degradation Conditions | Solvent | DPs Formed after 6 h | k (s−1) | t90 (min) | R2 | Ea (kJ/mol) | |
|---|---|---|---|---|---|---|---|
| Acidic hydrolysis | 25 °C | 0.1 M HCl | ADP-0 (0.94%) | 1.31 × 10−6 | 1340.2 | 0.9899 | na |
| 25 °C | 1 M HCl | The rate constant is determined from the extrapolation of Arrhenius plot | 1.77 × 10−6 | 993.8 | na | 102.3 | |
| 50 °C | ADP-1a (0.95%), ADP-1b (1.03%) ADP-2a (2.48%), ADP-2b (2.59%), ADP-4 (11.10%) | 4.34 × 10−5 | 40.5 | 0.9999 | |||
| 60 °C | ADP-1a (1.30%), ADP-1b (1.42%) ADP-2a (5.18%), ADP-2b (5.52%), ADP-4 (9.13%), ADP-5 (0.81%), ADP-6 (1.69%) | 1.33 × 10−4 | 13.2 | 0.9972 | |||
| 70 °C | ADP-1a (0.44%), ADP-1b (0.48%) ADP-2a (1.42%), ADP-2b (1.79%), ADP-3a (0.25%), ADP-4 (2.86%), ADP-3b (0.23%), ADP-5 (1.11%), ADP-6 (2.20%) | 3.99 × 10−4 | 4.4 | 0.9796 | |||
| Alkaline hydrolysis | 25 °C | 0.01 M NaOH | BDP-1a (32.28%), BDP-1b (20.39%), BDP-2 (2.70%) | 1.78 × 10−4 | 9.8 | 0.9909 | na |
| Oxidation | 3% H2O2 | 10% DMSO | ODP-1 (7.83%), ODP-2 (27.33%) | 2.56 × 10−5 | 68.7 | 0.9991 | na |
| 0.1 M HCl | ODP-1 (20.91%), ODP-2 (58.12%) | 1.33 × 10−4 | 13.2 | 0.9932 | na | ||
| Photolysis | 254 nm | 10% DMSO | PDP-1a (3.03%), PDP-1b * (1.26%) | 4.28 × 10−6 | 410.0 | 0.9991 | na |
| 0.1 M HCl | PDP-1a (1.68%), PDP-1b * (1.90%), PDP-2 (0.16%), PDP-1c (0.09%) | 4.72 × 10−6 | 372.3 | 0.9843 | na | ||
| 366 nm | 10% DMSO | PDP-1a (15.48%), PDP-1b * (0.26%), PDP-1c (0.05%) | 3.03 × 10−5 | 58.0 | 0.9896 | na | |
| 0.1 M HCl | PDP-1a (10.61%), PDP-1b * (0.19%), PDP-2 (0.15%), PDP-1c (0.15%) | 3.43 × 10−5 | 51.2 | 0.9851 | na | ||
| Thermolysis | 25 °C | 10% DMSO | The rate constant is determined from the extrapolation of Arrhenius plot | 9.74 × 10−8 | 18,023.1 | na | 106.3 |
| 50 °C | BDP-1b (0.45%) | 2.62 × 10−6 | 669.4 | 0.9837 | |||
| 60 °C | BDP-1b (1.32%), PDP-1b * (0.31%), TDP-1 (0.15%) | 9.25 × 10−6 | 189.9 | 0.9983 | |||
| 70 °C | BDP-1b (1.32%), ADP-5 (0.35%), PDP-1b * (0.60%), TDP-1 (0.83%) | 2.63 × 10−5 | 66.9 | 0.9962 | |||
| Condition | Exposure | DPs Detected | k (s−1) | t90 (d) | R2 |
|---|---|---|---|---|---|
| thermal | 21 d at 50 °C/uncontrolled RH | BDP-1b (7 d: 0.20%, 21 d: 0.15%), ODP-2 (7 d: 0.10%, 21 d: 0.12%), PDP-1b (7 d: 0.33%, 21 d: 0.32%) | 6.92 × 10−9 | 176.2 | 0.9949 |
| 21 d at 60 °C/uncontrolled RH | BDP-1b (7 d: 0.25%, 21 d: 0.16%), ODP-2 (7 d: 0.10%, 21 d: 0.12%), PDP-1b (7 d: 0.39%, 21 d: 0.34%) | 9.34 × 10−9 | 130.5 | 0.9842 | |
| thermal, high humidity | 21 d at 50 °C/75% RH | BDP-1b (7 d: 0.21%, 14 d: 0.40%, 21 d: 0.30%), ODP-2 (7 d: 0.06%, 14 d: 0.07%, 21 d: 0.07%), PDP-1b (7 d: 0.31%, 14 d: 0.27%, 21 d: 0.30%) | 3.61 × 10−9 | 337.7 | 0.9894 |
| photolysis, VIS | 1.26–2.65 million lux-hours | ODP-1 (10 d: 0.33%, 21 d: 0.64%), ODP-S1 (10 d: 0.06%, 21 d: 0.14%), ODP-S2 (10 d: 0.33%, 21 d: 0.65%), ODP-2 (10 d: 0.30%, 21 d: 0.51%), ODP-S3 (10 d: 0.23%, 21 d: 0.43%), PDP-1b (10 d: 0.39%, 21 d: 0.49%), ODP-S4 (10 d: 0.43%, 21 d: 0.69%) | 1.68 × 10−7 | 7.2 | 0.8415 |
| photolysis, UV | 21 d, 366 nm | ODP-1 (7 d: 0.26%, 14 d: 0.33%, 21 d: 0.45%), ODP-S1 (7 d: 0.06%, 14 d: 0.05%, 21 d: 0.07%), ODP-S2 (7 d: 0.28%, 14 d: 0.23%, 0.32%), ODP-2 (7 d: 0.25%, 14 d: 0.29%, 21 d: 0.37%), ODP-S3 (7 d: 0.19%, 14 d: 0.15%, 21 d: 0.22%), PDP-1b (7 d: 0.40%, 14 d: 0.34%, 21 d: 0.36%), ODP-S4 (7 d: 0.02%, 14 d: 0.32%, 21 d: 0.33%) | 2.60 × 10−8 | 46.9 | 0.9893 |
| Name | RRT | Experimental m/z | Proposed Molecular Formula of [M + H]+ | Error (ppm) | mSigma | Experimental m/z of Major Fragment Ions | Proposed [M + H]+ Formula of Fragment Ions | Proposed Structure |
|---|---|---|---|---|---|---|---|---|
| Ceftobiprole | 1.00 | 535.1167 | C20H23N8O6S2 | 1.8 | 26.3 | 473.1182 365.1264 321.1372 308.1063 291.1207 264.1168 220.1435 203.1179 | C19H21N8O3S2 C16H21N4O4S C15H21N4O2S C14H18N3O3S C14H17N3O4 C13H18N3OS C12H18N3O C12H15N2O |  |
| SBP-1 | 1.42 | 567.1431 | C21H27N8O7S2 | 1.4 | 6.3 | 523.1537 489.1660 370.1519 319.1763 264.1171 235.0907 220.1445 202.1335 | C20H27N8O5S2 C20H25N8O5S C18H20N5O4 C16H23N4O3 C13H18N3OS C12H15N2OS C12H18N3O C12H16N3 |  |
| SBP-2a SBP-2b | 1.48 1.59 | 591.1791 591.1799 | C24H31N8O6S2 | 1.9 0.5 | 17.6 36.8 | 535.1176 491.1248 473.1152 365.1288 321.1348 308.1060 291.1206 264.1167 228.0182 220.1430 203.1173 | C20H23N8O6S2 C19H23N8O4S2 C19H21N8O3S2 C16H21N4O4S C15H21N4O2S C14H18N3O3S C14H17N3O4 C13H18N3OS C6H6N5O3S C12H18N3O C12H15N2O | 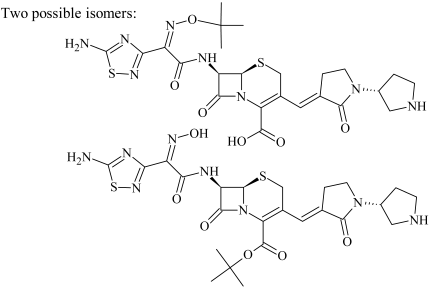 |
| SBP-3a SBP-3b | 1.65 1.67 | 701.1926 701.1933 | C33H33N8O6S2 | 4.7 3.7 | 18.6 13.1 | 394.0962 308.1044 291.1213 264.1164 247.1234 220.1444 203.1180 167.0857 | C21H18N2O4S C14H18N3O3S C14H17N3O4 C13H18N3OS C13H17N3O2 C12H18N3O C12H15N2O C13H12 | 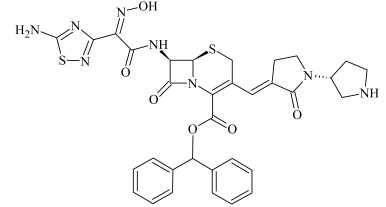 |
| ADP-0 | 538.1165 | C20H24N7O7S2 | 1.6 | 14.8 | 476.1171 458.1067 448.1201 308.1063 291.1209 264.1164 220.1441 203.1178 | C19H22N7O4S2 C19H20N7O3S2 C18H22N7O3S2 C14H18N3O3S C14H17N3O4 C13H18N3OS C12H18N3O C12H15N2O |  | |
| ADP-1a ADP-1b | 0.54 0.58 | 506.1439 506.1459 | C20H24N7O7S | 2.7 −1.2 | 7.2 80.0 | --------- | --------------- | Due to low intensity of fragmentation mass spectrum, the structural formula cannot be determined. Three tentative structural proposals are shown in Figure S2. |
| ADP-2a ADP-2b | 0.69 0.75 | 507.1297 507.1303 | C20H23N6O8S | −1.0 −2.0 | 12.3 13.5 | 464.1230 446.1141 291.1201 264.1177 247.1288 220.1424 203.1171 179.0814 | C19H22N5O7S C19H20N5O6S C14H17N3O4 C13H18N3OS C13H17N3O2 C12H18N3O C12H15N2O C9H11N2O2 | The structure cannot be unambiguously determined. Many structural formulas can be proposed based on the fragmentation mass spectrum (three examples are shown in Figure 11). |
| ADP-3a ADP-3b | 0.78 0.84 | 479.1348 479.1342 | C19H23N6O7S | −1.0 0.4 | 16.5 125.6 | 418.1190 365.1334 321.1399 291.1194 264.1172 223.0672 203.1189 179.0820 | C18H20N5O5S C16H21N4O4S C15H21N4O2S C14H17N3O4 C13H18N3OS C10H12N2O4 C12H15N2O C9H11N2O2 | 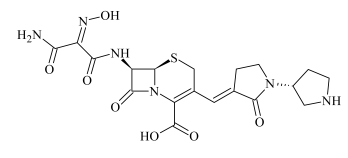 |
| ADP-4 | 0.81 | 365.1274 | C16H21N4O4S | 1.0 | 18.9 | 293.1406 291.1203 259.1537 232.1437 220.1424 207.0592 203.1168 165.1029 | C14H21N4OS C14H17N3O4 C14H19N4O C13H18N3O C12H18N3O C10H11N2OS C12H15N2O C9H13N2O | 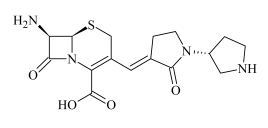 |
| ADP-5 | 0.88 | 265.1178 | C13H17N2O4 | 1.9 | 2.8 | 221.1281 203.1173 193.1312 154.0873 152.0745 137.0610 | C12H17N2O2 C12H15N2O C11H17N2O C8H12NO2 C8H10NO2 C8H8O2 | 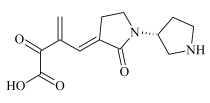 |
| ADP-6 | 1.16 | 281.0950 | C13H17N2O3S | 1.7 | 22.3 | 263.0846 253.0885 235.0739 219.1124 194.0272 152.0932 | C13H15N2O2S C12H17N2O2S C12H15N2OS C12H15N2O2 C9H8NO2S C8H12N2O |  |
| BDP-1a BDP-1b | 0.62 0.65 | 553.1279 553.1284 | C20H25N8O7S2 | 0.6 −0.4 | 41.2 35.3 | 509.1376 465.1485 431.1616 352.1189 308.1058 287.1505 264.1688 220.1428 202.1337 | C19H25N8O5S2 C18H25N8O3S2 C18H23N8O3S C15H20N4O4S C14H18N3O3S C15H19N4O2 C13H18N3OS C12H18N3O C12H16N3 |  |
| BDP-2 | 1.18 | 535.1174 | C20H23N8O6S2 | 0.5 | 14.1 | 308.1066 291.1202 264.1163 246.1062 228.0172 203.1171 | C14H18N3O3S C14H17N3O4 C13H18N3OS C13H16N3S C6H5N5O3S C12H15N2O |  |
| ODP-1 | 0.53 | 551.1118 | C20H23N8O7S2 | 1.3 | 28.5 | 489.1130 441.1436 357.0770 319.1211 291.1216 247.1311 220.1429 218.1279 | C19H21N8O4S2 C19H21N8O3S C15H13N6O3S C15H19N4O2S C14H17N3O4 C13H17N3O2 C12H18N3O C12H16N3O | 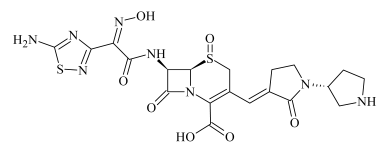 |
| ODP-2 | 0.77 | 551.1118 | C20H23N8O7S2 | 1.4 | 31.7 | 507.1217 489.1132 463.1332 441.1458 417.1445 319.1212 291.1220 247.1311 218.1279 | C19H23N8O5S2 C19H21N8O4S2 C18H23N8O3S2 C19H21N8O3S C17H21N8O3S C15H19N4O2S C14H17N3O4 C13H17N3O2 C12H16N3O |  |
| ODP-S1 ODP-S2 ODP-S3 | 0.59 0.74 0.90 | 551.1112 551.1123 551.1122 | C20H23N8O7S2 | 2.4 0.4 0.7 | 33.0 10.2 15.6 | --------- | --------------- | Due to low intensity of fragmentation mass spectrum, the structural formula cannot be determined. |
| ODP-S4 | 1.44 | [M + 2H]2+ 550.1035 | [M + 2H]2+ C40H44N16O14S4 | 2.2 | 28.3 | --------- | --------------- | Due to low intensity of fragmentation mass spectrum, the structural formula cannot be determined. |
| PDP-1a PDP-1b PDP-1c TDP-1 | 1.06 1.14 1.40 1.19 | 535.1174 535.1175 535.1178 535.1172 | C20H23N8O6S2 | 0.4 0.3 −0.2 0.9 | 34.2 34.3 18.4 16.9 | The same as for ceftobiprole | Isomers of ceftobiprole | |
| PDP-2 | 1.26 | 503.1447 | C20H23N8O6S | 1.8 | 20.7 | --------- | --------------- | Due to low intensity of fragmentation mass spectrum, the structural formula cannot be determined. Two tentative structural proposals are shown in Figure S3. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boczar, D.; Bus, K.; Michalska, K. Study of Degradation Kinetics and Structural Analysis of Related Substances of Ceftobiprole by HPLC with UV and MS/MS Detection. Int. J. Mol. Sci. 2022, 23, 15252. https://doi.org/10.3390/ijms232315252
Boczar D, Bus K, Michalska K. Study of Degradation Kinetics and Structural Analysis of Related Substances of Ceftobiprole by HPLC with UV and MS/MS Detection. International Journal of Molecular Sciences. 2022; 23(23):15252. https://doi.org/10.3390/ijms232315252
Chicago/Turabian StyleBoczar, Dariusz, Katarzyna Bus, and Katarzyna Michalska. 2022. "Study of Degradation Kinetics and Structural Analysis of Related Substances of Ceftobiprole by HPLC with UV and MS/MS Detection" International Journal of Molecular Sciences 23, no. 23: 15252. https://doi.org/10.3390/ijms232315252
APA StyleBoczar, D., Bus, K., & Michalska, K. (2022). Study of Degradation Kinetics and Structural Analysis of Related Substances of Ceftobiprole by HPLC with UV and MS/MS Detection. International Journal of Molecular Sciences, 23(23), 15252. https://doi.org/10.3390/ijms232315252







