Genetic Determinants of Antagonistic Interactions and the Response of New Endophytic Strain Serratia quinivorans KP32 to Fungal Phytopathogens
Abstract
:1. Introduction
2. Results
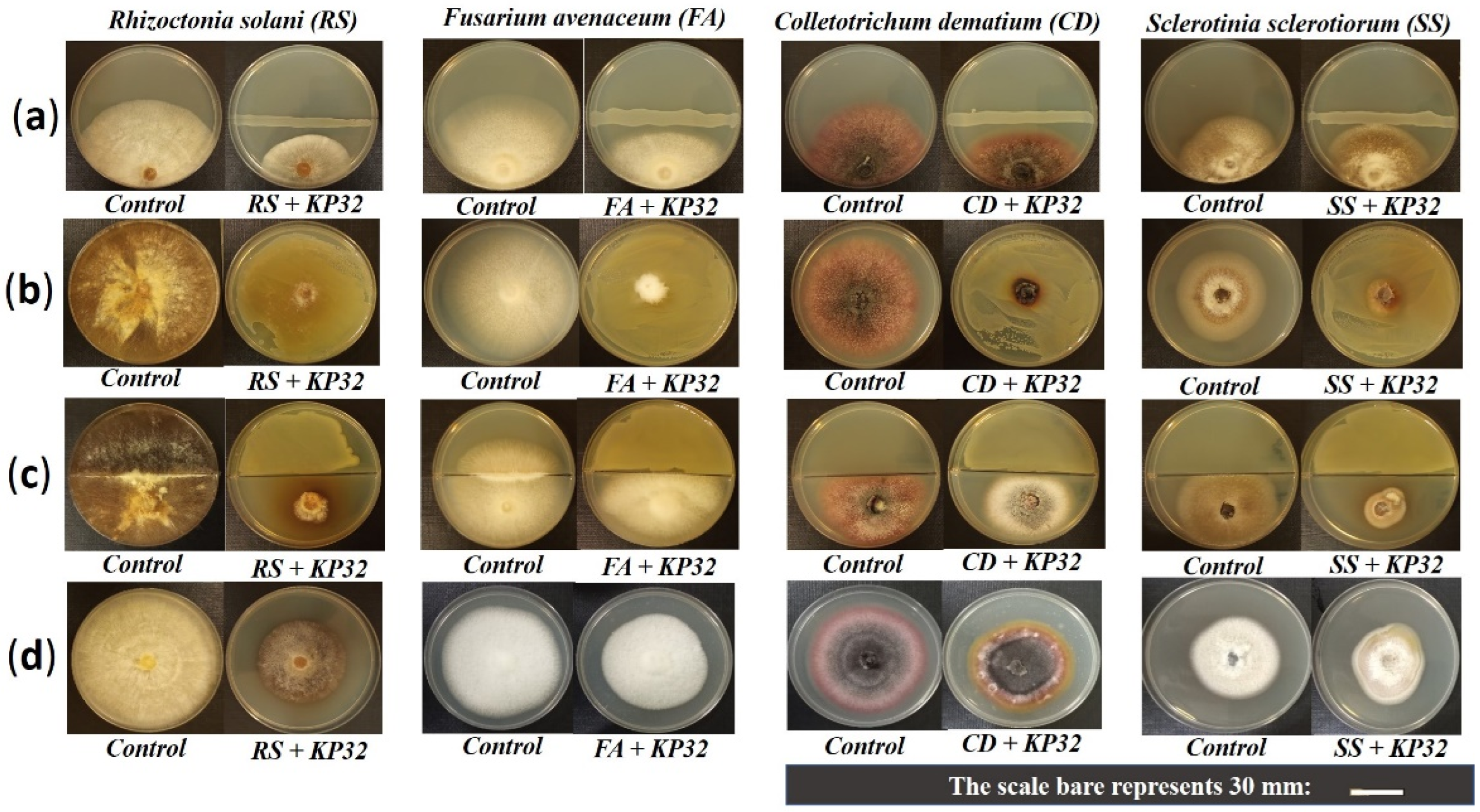
2.1. In Vitro Inhibition of Phytopathogens by the S. quinivornas KP32 Strain
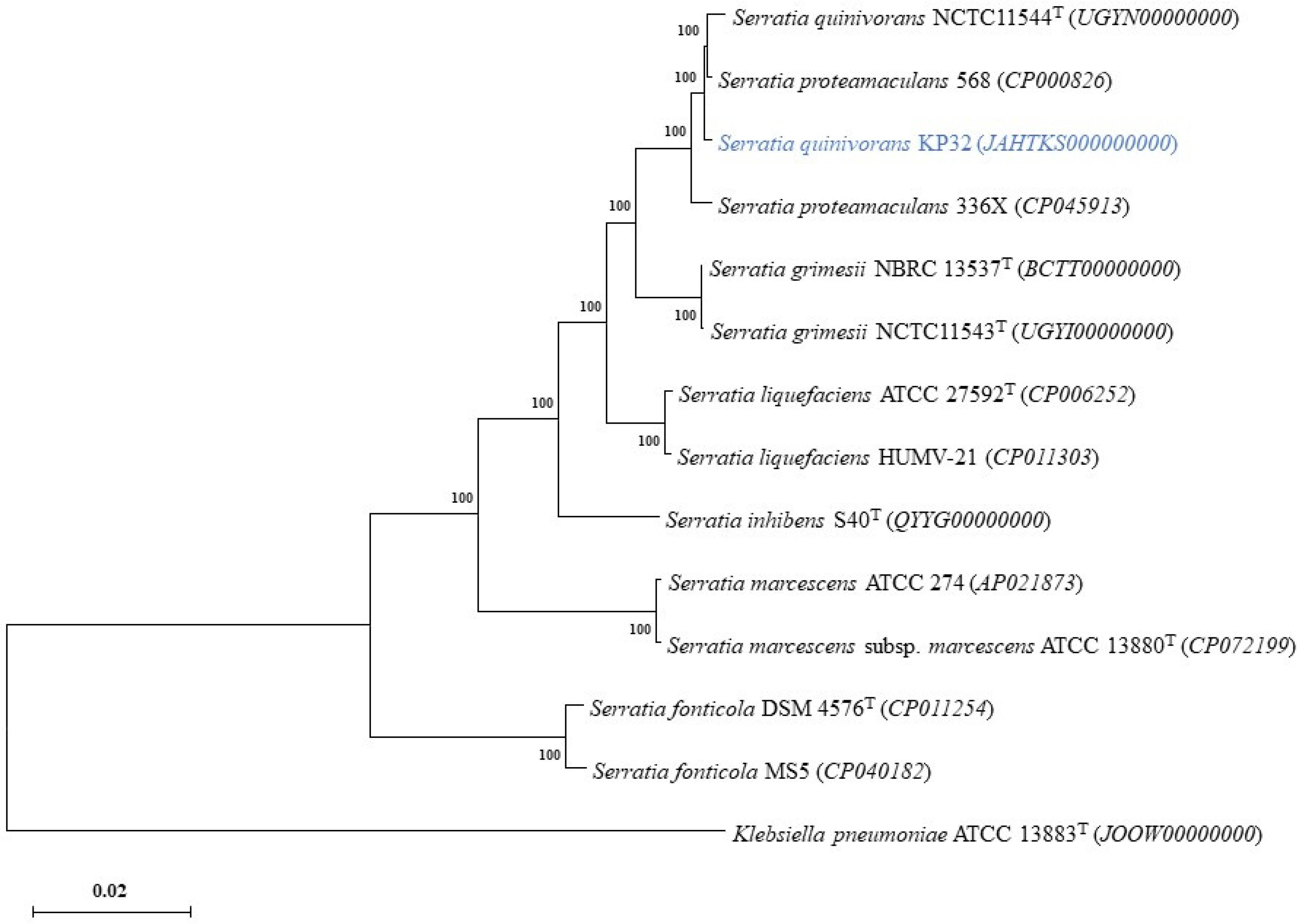
2.2. The Properties of Genome and Phylogenetic Analyses
2.3. Genes Essential for Biocontrol Activity in the KP32 Strain Genome
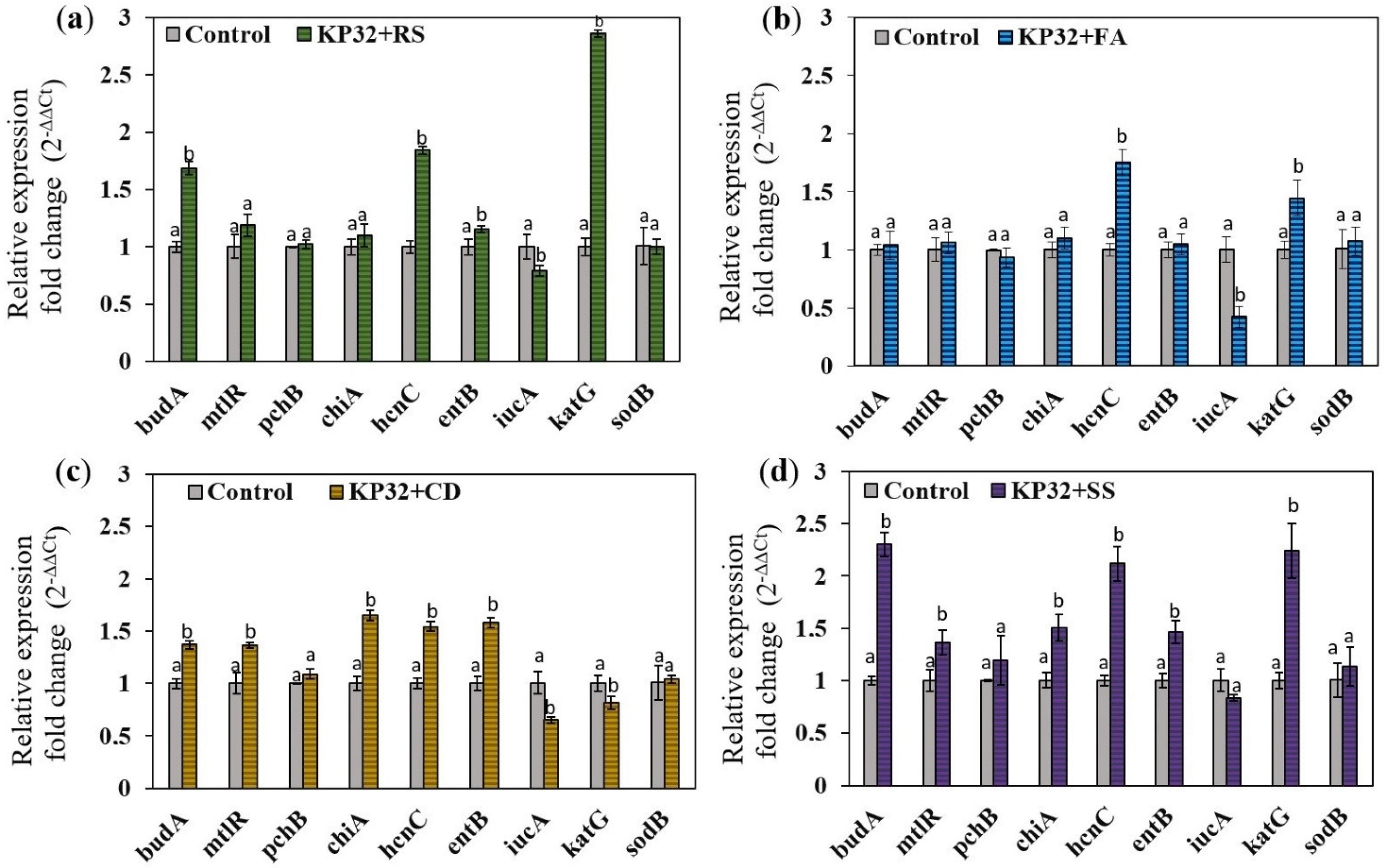
2.4. Determination of the Effect of Fungal Phytopathogens on the Expression of Genes Responsible for Antifungal Activity
2.5. Evaluation of Biocontrol Features of the KP32 Strain in a Biochemical Assay
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Growth Conditions
4.2. Pathogenic Fungi
4.3. In Vitro Screening of Antagonistic Behavior of the KP32 Strain against Fungal Phytopathogens
4.3.1. Dual-Culture Assay
4.3.2. Detection of Diffusible Metabolite Production
4.3.3. Detection of Volatile Metabolites Production
4.3.4. Evaluation of the Effect of KP32 Cell-Free Culture Filtrate
4.4. Genome Sequencing and Sequence Analysis
4.5. Phylogenetic Analysis
4.6. Evaluation of the Effect of Fungal Pathogens on the Expression Level of Genes Engaged in Antifungal Activity
4.6.1. Preparation of Fungal Filtrates
4.6.2. Influence of Fungal Filtrates on Bacterial Genes Expression
4.7. Evaluation of the Effect of Fungal Pathogens on Lytic and Antioxidant Enzymes Activity
4.7.1. Lytic Enzyme Activity
4.7.2. Antioxidant Enzyme Activity
4.8. Plant Growth Promotion Features of the KP32 Strain
4.9. Colonization Features of the KP32 Strain
Utilization of Selected Organic Compounds as the Sole Source of Carbon and Energy
4.10. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| BCA | Biological Control Agents |
| ISR | Induced Systemic Resistance |
| N-AHLs | N-acyl-homoserine lactones |
| PCR | Polymerase Chain Reaction |
| VOCs | Volatile Organic Compounds |
| CDS | Protein-encoding sequences |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| COG | Cluster of Orthologous Genes |
| NRPS | Nonribosomal Peptide-Synthetase |
| DHBA | Dihydroxybenzoic Acid |
| MFS | Major Facilitator Superfamily |
| IAA | Indole-3-acetic acid |
| TRP | Tryptophane |
| CBS | Cystathionine β synthase |
| CTH | Cystathionine-γ-lyase |
| GTS | Glutathione S-transferases |
| QS | Quorum sensing |
| CAZy | Carbohydrate-Active Enzymes |
| T2SS | Type II Secretion Systems |
| T6SS | Type VI Secretion Systems |
| T4PS | Type IV Pilus System |
| GI | Genomic Islands |
| RT-qPCR | quantitative reverse transcription PCR |
| SA | Salicylic Acid |
| ACC | 1-aminocyclopropane-1-carboxylate |
| DF | Dworkin and Foster |
| HCN | Hydrogen Cyanide |
| SOD | Superoxide Dismutase |
| CAT | Catalase |
| EPS | Exopolysaccharide |
| CRA | Congo Red Agar |
| SEM | Scanning Electron Microscopy |
| PGI | Percent Growth Inhibition |
| LB | Luria-Bertani broth |
| CV | Crystal Violet |
| PSI | Phosphate Solubilizing Index |
| CAS | Chrome azurol S |
| PGPB | Plant Growth Promoting Bacteria |
| PGP | Plant Growth Promotion |
| PDA | Potato Dextrose Agar |
| CWDEs | Cell Wall-Degrading Enzymes |
| DNS | 3,5-dinitro salicylic acid |
| LSD | Lowest Significant Difference |
References
- Marques-Pereira, C.; Proença, D.N.; Morais, P.V. Genome sequences of Serratia strains revealed common genes in both serratomolides gene clusters. Biology 2020, 9, 482. [Google Scholar] [CrossRef] [PubMed]
- Matilla, M.A.; Nogellova, V.; Morel, B.; Krell, T.; Salmond, G.P. Biosynthesis of the acetyl-CoA carboxylase-inhibiting antibiotic, andrimid in Serratia is regulated by Hfq and the LysR-type transcriptional regulator, AdmX. Environ. Microbiol. 2016, 18, 3635–3650. [Google Scholar] [CrossRef] [PubMed]
- AL-Ghanem, M.M. Serratia a novel source of secondary metabolites. Adv. Biotechnol. Microbiol. 2018, 11, 555814. [Google Scholar] [CrossRef]
- Niu, H.; Sun, Y.; Zhang, Z.; Zhao, D.; Wang, N.; Wang, L.; Guo, H. The endophytic bacterial entomopathogen Serratia marcescens promotes plant growth and improves resistance against Nilaparvata lugens in rice. Microbiol. Res. 2022, 256, 126956. [Google Scholar] [CrossRef] [PubMed]
- Vaughan, A.L.; Altermann, E.; Glare, T.R.; Hurst, M.R. Genome sequence of the entomopathogenic Serratia entomophila isolate 626 and characterisation of the species specific itaconate degradation pathway. BMC Genom. 2022, 23, 728. [Google Scholar] [CrossRef]
- Soenens, A.; Imperial, J. Biocontrol capabilities of the genus Serratia. Phytochem. Rev. 2020, 19, 577–587. [Google Scholar] [CrossRef]
- Lahlali, R.; Ezrari, S.; Radouane, N.; Kenfaoui, J.; Esmaeel, Q.; El Hamss, H.; Belabess, Z.; Barka, E.A. Biological control of plant pathogens: A global perspective. Microorganisms 2022, 10, 596. [Google Scholar] [CrossRef]
- Toffolatti, S.L.; Maffi, D.; Serrati, L.; Vercesi, A. Histological and ultrastructural studies on the curative effects of mandipropamid on Plasmopara viticola. J. Phytopathol. 2010, 159, 201–207. [Google Scholar] [CrossRef]
- Hazarika, D.J.; Goswami, G.; Gautom, T.; Parveen, A.; Das, P.; Barooah, M.; Chandra Boro, R. Lipopeptide mediated biocontrol activity of endophytic Bacillus subtilis against fungal phytopathogens. BMC Microbiol. 2019, 19, 71. [Google Scholar] [CrossRef]
- Wang, M.; Xing, Y.; Wang, J.; Xu, Y.; Wang, G. The role of the chi1 gene from the endophytic bacteria Serratia proteamaculans 336x in the biological control of wheat take-all. Can. J. Microbiol. 2014, 60, 533–540. [Google Scholar] [CrossRef]
- Dhar Purkayastha, G.; Mangar, P.; Saha, A.; Saha, D. Evaluation of the biocontrol efficacy of a Serratia marcescens strain indigenous to tea rhizosphere for the management of root rot disease in tea. PLoS ONE 2018, 13, e0191761. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suryadi, Y.; Susilowati, D.N.; Fauziah, F. Management of Plant Diseases by PGPR-Mediated Induced Resistance with Special Reference to Tea and Rice Crops. In Plant Growth Promoting Rhizobacteria for Sustainable Stress Management. Microorganisms for Sustainability; Sayyed, R., Ed.; Springer: Singapore, 2019; Volume 13, pp. 65–110. [Google Scholar]
- Kshetri, L.; Naseem, F.; Pandey, P. Role of Serratia sp. as Biocontrol Agent and Plant Growth Stimulator, with Prospects of Biotic Stress Management in Plant. In Plant Growth Promoting Rhizobacteria for Sustainable Stress Management. Microorganisms for Sustainability; Sayyed, R., Ed.; Springer: Singapore, 2019; Volume 13, pp. 169–200. [Google Scholar]
- Su, C.; Xiang, Z.; Liu, Y.; Zhao, X.; Sun, Y.; Li, Z.; Li, L.; Chang, F.; Chen, T.; Wen, X.; et al. Analysis of the genomic sequences and metabolites of Serratia surfactantfaciens sp. nov. YD25T that simultaneously produces prodigiosin and serrawettin W2. BMC Genom. 2016, 17, 865. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, X.; Jia, J.; Atkinson, S.; Cámara, M.; Gao, K.; Li, H.; Cao, J. Biocontrol potential of an endophytic Serratia sp. G3 and its mode of action. World J. Microbiol. Biotechnol. 2010, 26, 1465–1471. [Google Scholar] [CrossRef]
- Kamensky, M.; Ovadis, M.; Chet, I.; Chernin, L. Soil-borne strain IC14 of Serratia plymuthica with multiple mechanisms of antifungal activity provides biocontrol of Botrytis cinerea and Sclerotinia sclerotiorum diseases. Soil Biol. Biochem. 2003, 35, 323–331. [Google Scholar] [CrossRef]
- De Vleeschauwer, D.; Hö, M. Using Serratia plymuthica to control fungal pathogens of plant. CAB Rev. 2007, 2, 46. [Google Scholar] [CrossRef] [Green Version]
- Neupane, S.; Finlay, R.D.; Alström, S.; Elfstrand, M.; Högberg, N. Transcriptional responses of the bacterial antagonist Serratia plymuthica to the fungal phytopathogen Rhizoctonia solani. Environ. Microbiol. Rep. 2015, 7, 123–127. [Google Scholar] [CrossRef]
- Guitiérrez-Román, M.I.; Holguín-Meléndez, F.; Bello-Mendoza, R.; Guillén-Navarro, K.; Dunn, M.F.; Huerta-Palacios, G. Production of prodigiosin and chitinases by tropical Serratia marcescens strains with potential to control plant pathogens. World J. Microbiol. Biotechnol. 2012, 28, 145–153. [Google Scholar] [CrossRef]
- Ferraz, H.G.M.; Resende, R.S.; Moreira, P.C.; Silveira, P.R.; Milagres, E.A.; Oliveira, J.R.; Rodrigues, F.A. Antagonistic rhizobacteria and jasmonic acid induce resistance against tomato bacterial spot. Plant Prot. Sci. 2015, 74, 417–427. [Google Scholar] [CrossRef] [Green Version]
- Kumar, A.; Radhakrishnan, E.; Droby, S.; Singh, V.; Singh, S.; White, J. Entry, colonization, and distribution of endophytic microorganisms in plants. In Microbial Endophytes: Functional Biology and Applications, 1st ed.; Kumar, A., Radhakrishnan, E.K., Eds.; Woodhead Publishing: Cambridge, UK, 2019; Volume 1, pp. 1–33. [Google Scholar]
- Nelkner, J.; Tejerizo, G.T.; Hassa, J.; Lin, T.W.; Witte, J.; Verwaaijen, B.; Winkler, A.; Bunk, B.; Spröer, C.; Overmann, J.; et al. Genetic potential of the biocontrol agent Pseudomonas brassicacearum (formerly P. trivialis) 3Re2-7 unraveled by genome sequencing and mining, comparative genomics and transcriptomics. Genes 2019, 10, 601. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Yang, Y.; Dubern, J.F.; Li, H.; Halliday, N.; Chernin, L.; Gao, K.; Cámara, M.; Liu, X. Regulation of GacA in Pseudomonas chlororaphis Strains Shows a Niche Specificity. PLoS ONE 2015, 10, e0137553. [Google Scholar] [CrossRef]
- Caneschi, W.L.; Sanchez, A.B.; Felestrino, É.B.; de Carvalho Lemes, C.G.; Cordeiro, I.F.; Fonseca, N.P.; Villa, M.M.; Vieira, I.T.; Moraes, L.Â.G.; de Almeida Barbosa Assis, R.; et al. Serratia liquefaciens FG3 isolated from a metallophyte plant sheds light on the evolution and mechanisms of adaptive traits in extreme environments. Sci. Rep. 2019, 9, 18006. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matteoli, F.P.; Passarelli-Araujo, H.; Reis, R.J.A.; da Rocha, L.O.; de Souza, E.M.; Aravind, L.; Olivares, F.L.; Venancio, T.M. Genome sequencing and assessment of plant growth-promoting properties of a Serratia marcescens strain isolated from vermicompost. BMC Genom. 2018, 16, 750. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barnhart, M.M.; Chapman, M.R. Curli biogenesis and function. Annu. Rev. Microbiol. 2006, 60, 131–147. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rezzonico, F.; Smits, T.H.; Duffy, B. Detection of AI-2 receptors in genomes of Enterobacteriaceae suggests a role of type-2 quorum sensing in closed ecosystems. Sensors 2012, 12, 6645–6665. [Google Scholar] [CrossRef] [Green Version]
- Eida, A.A.; Bougouffa, S.; L’Haridon, F.; Alam, I.; Weisskopf, L.; Bajic, V.B.; Saad, M.M.; Hirt, H. Genome Insights of the Plant-Growth Promoting Bacterium Cronobacter muytjensii JZ38 With Volatile-Mediated Antagonistic Activity Against Phytophthora infestans. Front. Microbiol. 2020, 11, 369. [Google Scholar] [CrossRef] [Green Version]
- Andrés-Barrao, C.; Lafi, F.F.; Alam, I.; de Zélicourt, A.; Eida, A.A.; Bokhari, A.; Alzubaidy, H.; Bajic, V.B.; Hirt, H.; Saad, M.M. Complete genome sequence analysis of Enterobacter sp. SA187, a plant multi-stress tolerance promoting endophytic bacterium. Front. Microbiol. 2017, 8, 2023. [Google Scholar] [CrossRef] [Green Version]
- Green, E.R.; Mecsas, J. Bacterial Secretion Systems: An Overview. Microbiol. Spectr. 2016, 4, 1–19. [Google Scholar] [CrossRef] [Green Version]
- Trunk, K.; Peltier, J.; Liu, Y.C.; Dill, B.D.; Walker, L.; Gow, N.A.R.; Stark, M.J.R.; Quinn, J.; Strahl, H.; Trost, M.; et al. The type VI secretion system deploys antifungal effectors against microbial competitors. Nat. Microbiol. 2018, 3, 920–931. [Google Scholar] [CrossRef]
- Wolska, K.; Jakubczak, A. Wykrywanie biofilmu Pseudomonas aeruginosa na biomateriałach medycznych. Med. Doświadczalna Mikrobiol. 2003, 55, 371–378. [Google Scholar]
- Stepanovic, S.; Vukovic, D.; Hola, V.; Di Bonaventura, G.; Djukic, S.; Cirkovic, I.; Ruzicka, F. Quantification of biofilm in microtiter plates: Overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS 2007, 115, 891–899. [Google Scholar] [CrossRef]
- Sharma, M.; Saleh, D.; Charron, J.-B.; Jabaji, S. A crosstalk between Brachypodium root exudates, organic acids, and Bacillus velezensis B26, a growth promoting bacterium. Front. Microbiol. 2020, 11, 575578. [Google Scholar] [CrossRef] [PubMed]
- He, D.-C.; He, M.-H.; Amalin, D.M.; Liu, W.; Alvindia, D.G.; Zhan, J. Biological Control of Plant Diseases: An Evolutionary and Eco-Economic Consideration. Pathogens 2021, 10, 1311. [Google Scholar] [CrossRef] [PubMed]
- Chandra, H.; Kumari, P.; Bisht, R.; Prasad, R.; Yadav, S. Plant growth promoting Pseudomonas aeruginosa from Valeriana wallichii displays antagonistic potential against three phytopathogenic fungi. Mol. Biol. Rep. 2020, 47, 6015–6026. [Google Scholar] [CrossRef] [PubMed]
- Afzal, I.; Shinwari, Z.K.; Sikandar, S.; Shahzad, S. Plant beneficial endophytic bacteria: Mechanisms, diversity, host range and genetic determinants. Microbiol. Res. 2019, 221, 36–49. [Google Scholar] [CrossRef] [PubMed]
- Shen, S.S.; Choi, O.H.; Lee, S.M.; Park, C.S. In vitro and in vivo activities of biocontrol agent, Serratia plymuthica A21-4, against Phytophthora capsici. Plant Pathol. J. 2002, 18, 221–224. [Google Scholar] [CrossRef] [Green Version]
- Frankowski, J.; Lorito, M.; Scala, F.; Schmid, R.; Berg, G.; Bahl, H. Purification and properties of two chitinolytic enzymes of Serratia plymuthica HRO-C48. Arch. Microbiol. 2001, 176, 421–426. [Google Scholar] [CrossRef]
- Roberts, D.; Lohrke, S.; Meyer, S.; Buyer, J.; Bowers, J.; Baker, C.; Li, W.; de Souza, J.; Lewis, J.; Chung, S. Biocontrol agents applied individually and in combination for suppression of soilborne diseases of cucumber. Crop Prot. 2005, 24, 141–155. [Google Scholar] [CrossRef] [Green Version]
- Parani, K.; Shetty, G.P.; Saha, B.K. Isolation of Serratia marcescens SR1 as a source of chitinase having potentiality of using as a biocontrol agent. Indian J. Microbiol. 2011, 51, 247–250. [Google Scholar] [CrossRef] [Green Version]
- Chakraborty, U.; Chakraborty, B.N.; Chakraborty, A.P. Influence of Serratia marcescens TRS-1 on growth promotion and induction of resistance in Camellia sinensis against Fomes lamaoensis. J. Plant Interact. 2010, 5, 261–272. [Google Scholar] [CrossRef]
- Varma, A.; Bakshi, M.; Lou, B.; Hartmann, A.; Oelmueller, R. Piriformospora indica: A Novel Plant Growth Promoting Mycorrhizal Fungus. Agric. Res. 2012, 1, 117–131. [Google Scholar] [CrossRef]
- Dashti, N.; Prithiviraj, B.; Hynes, R.K.; Smith, D.L. Root and Rhizosphere Colonization of Soybean (Glycine max (L.) Merr.) by Plant-Growth-Promoting Rhizobacteria at Low Root Zone Temperatures and under Short-Season Conditions. J. Agron. Crop Sci. 2000, 185, 15–22. [Google Scholar] [CrossRef]
- Berg, G.; Roskot, N.; Steidle, A.; Eberl, L.; Zock, A.; Smalla, K. Plant-dependent genotypic and phenotypic diversity of antagonistic rhizobacteria isolated from different Verticillium host plants. Appl. Environ. Microbiol. 2002, 68, 3328–3338. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, A.; Lin, R.; Zhang, D.; Qin, P.; Xu, L.; Ai, P.; Ding, L.; Wang, Y.; Chen, Y.; Liu, Y.; et al. The evolution and pathogenic mechanisms of the rice sheath blight pathogen. Nat. Commun. 2013, 4, 1424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marzouk, T.; Chaouachi, M.; Sharma, A.; Jallouli, S.; Mhamdi, R.; Kaushik, N.; Djébali, N. Biocontrol of Rhizoctonia solani using volatile organic compounds of solanaceae seed-borne endophytic bacteria. Postharvest Biol. Technol. 2021, 181, 111655. [Google Scholar] [CrossRef]
- Gkarmiri, K.; Finlay, R.D.; Alström, S.; Thomas, E.; Cubeta, M.A.; Högberg, N. Transcriptomic changes in the plant pathogenic fungus Rhizoctonia solani AG-3 in response to the antagonistic bacteria Serratia proteamaculans and Serratia plymuthica. BMC Genom. 2015, 16, 630. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, P.; Kwok, A.H.Y.; Jiang, J.; Ran, T.; Xu, D.; Wang, W.; Leung, F.C. Comparative Genome Analyses of Serratia marcescens FS14 Reveals Its High Antagonistic Potential. PLoS ONE 2015, 10, e0123061. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Delgado, N.; Olivera, M.; Cádiz, F.; Bravo, G.; Montenegro, I.; Madrid, A.; Fuentealba, C.; Pedreschi, R.; Salgado, E.; Besoain, X. Volatile Organic Compounds (VOCs) Produced by Gluconobacter cerinus and Hanseniaspora osmophila Displaying Control Effect against Table Grape-Rot Pathogens. Antibiotics 2021, 10, 663. [Google Scholar] [CrossRef]
- Weise, T.; Thurmer, A.; Brady, S.; Kai, M.; Daniel, R.; Gottschalk, G.; Piechulla, B. VOC emission of various Serratia species and isolates and genome analysis of Serratia plymuthica 4Rx13. FEMS Microbiol. Lett. 2014, 352, 45–53. [Google Scholar] [CrossRef] [Green Version]
- Nascimento, F.; Vicente, C.; Cock, P.; Tavares, M.; Rossi, M.; Hasegawa, K.; Mota, M. From plants to nematodes: Serratia grimesii BXF1 genome reveals an adaptation to the modulation of multi-species interactions. Microb. Genom. 2018, 4, e000178. [Google Scholar] [CrossRef]
- Yi, H.S.; Ahn, Y.R.; Song, G.C.; Ghim, S.Y.; Lee, S.; Lee, G.; Ryu, C.M. Impact of a Bacterial Volatile 2,3-Butanediol on Bacillus subtilis Rhizosphere Robustness. Front. Microbiol. 2016, 7, 993. [Google Scholar] [CrossRef] [Green Version]
- van der Lelie, D.; Taghavi, S.; Monchy, S.; Schwender, J.; Miller, L.; Ferrieri, R.; Rogers, A.; Wu, X.; Zhu, W.; Weyens, N.; et al. Poplar and its Bacterial Endophytes: Coexistence and Harmony. Crit. Rev. Plant Sci. 2009, 28, 346–358. [Google Scholar] [CrossRef]
- Taghavi, S.; Garafola, C.; Monchy, S.; Newman, L.; Hoffman, A.; Weyens, N.; Barac, T.; Vangronsveld, J.; van der Lelie, D. Genome survey and characterization of endophytic bacteria exhibiting a beneficial effect on growth and development of poplar trees. Appl. Environ. Microbiol. 2009, 75, 748–757. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Umesha, S.; Singh, K.P.P.; Singh, R. Microbial biotechnology and sustainable agriculture. In Biotechnology for Sustainable Agriculture; Lakhan Singh, R., Mondal, S., Eds.; Woodhead Publishing: Cambridge, UK, 2018; pp. 185–205. [Google Scholar]
- Peng, G.; Zhao, X.; Li, Y.; Wang, R.; Huang, Y.; Qi, G. Engineering Bacillus velezensis with high production of acetoin primes strong induced systemic resistance in Arabidopsis thaliana. Microbiol. Res. 2019, 227, 126297. [Google Scholar] [CrossRef]
- Fu, L.H.; Hu, K.D.; Hu, L.Y.; Li, Y.H.; Hu, L.B.; Yan, H.; Liu, Y.S.; Zhang, H. An antifungal role of hydrogen sulfide on the postharvest pathogens Aspergillus niger and Penicillium italicum. PLoS ONE 2014, 9, e104206. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Popova, A.A.; Koksharova, O.A.; Lipasova, V.A.; Zaitseva, J.V.; Katkova-Zhukotskaya, O.A.; Eremina, S.I.; Mironov, A.S.; Chernin, L.S.; Khmel, I.A. Inhibitory and toxic effects of volatiles emitted by strains of Pseudomonas and Serratia on growth and survival of selected microorganisms, Caenorhabditis elegans, and Drosophila melanogaster. Biomed. Res. Int. 2014, 2014, 125704. [Google Scholar] [CrossRef] [Green Version]
- Etminani, F.; Harighi, B. Isolation and Identification of Endophytic Bacteria with Plant Growth Promoting Activity and Biocontrol Potential from Wild Pistachio Trees. Plant Pathol. J. 2018, 34, 208–217. [Google Scholar] [CrossRef]
- Dorjey, S.; Dolkar, D.; Sharma, R. Plant growth promoting rhizobacteria Pseudomonas: A review. Int. J. Curr. Microbiol. 2017, 6, 1335–1344. [Google Scholar] [CrossRef]
- Purushotham, P.; Arun, P.V.; Prakash, J.S.; Podile, A.R. Chitin binding proteins act synergistically with chitinases in Serratia proteamaculans 568. PLoS ONE 2012, 7, e36714. [Google Scholar] [CrossRef]
- Compant, S.; Duffy, B.; Nowak, J.; Clément, C.; Barka, E.A. Use of plant growth promoting bacteria for biocontrol of plant diseases: Principles, mechanisms of action, and future prospects. Appl. Environ. Microbiol. 2005, 71, 4951–4959. [Google Scholar] [CrossRef] [Green Version]
- Someya, N.; Nakajima, M.; Watanabe, K.; Hibi, T.; Akutsu, K. Potential of Serratia marcescens strain B2 for biological control of rice sheath blight. Biocontrol Sci. Technol. 2005, 15, 105–109. [Google Scholar] [CrossRef]
- Ben Slama, H.; Triki, M.A.; Bouket, A.C.; Mefteh, B.F.; Alenezi, F.N.; Luptakova, L.; Cherif-Silini, H.; Vallat, A.; Oszako, T.; Gharsallah, N.; et al. Screening of the High Rhizosphere Competent Limoniastrum monopetalum’ Culturable Endophyte Microbiota Allows the Recovery of Multifaceted and Versatile Biocontrol Agents. Microorganisms 2019, 7, 249. [Google Scholar] [CrossRef]
- Kumar, A.; Vandana, R.S.; Singh, M.; Pandey, K.D. Plant growth promoting rhizobacteria (PGPR). A promising approach to disease management. In Microbes and Environmental Management; Studium Press: New Delhi, India, 2019; pp. 195–209. [Google Scholar]
- Faltin, F.; Lottmann, J.; Grosch, R.; Berg, G. Strategy to select and assess antagonistic bacteria for biological control of Rhizoctonia solani Kühn. Can. J. Microbiol. 2004, 50, 811–820. [Google Scholar] [CrossRef] [PubMed]
- Shanmugaiah, V.; Nithya, K.; Harikrishnan, H.; Jayaprakashvel, M.; Balasubramanian, N. Biocontrol mechanisms of siderophores against bacterial plant pathogens. In Sustainable Approaches to Controlling Plant Pathogenic Bacteria, 1st ed.; Kannan, V.R., Bastas, K.K., Eds.; CRC Press: Boca Raton, FL, USA, 2015; pp. 167–190. [Google Scholar]
- Ahmed, E.; Holmström, S.J. Siderophores in environmental research: Roles and applications. Microbial. Biotechnol. 2014, 7, 196–208. [Google Scholar] [CrossRef] [PubMed]
- Schiessl, K.T.; Janssen, E.M.; Kraemer, S.M.; McNeill, K.; Ackermann, M. Magnitude and Mechanism of Siderophore-Mediated Competition at Low Iron Solubility in the Pseudomonas aeruginosa Pyochelin System. Front. Microbiol. 2017, 8, 1964. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bigirimana, J.; Höfte, M. Induction of systemic resistance to Colletotrichum lindemuthianum in bean by a benzothiadiazole derivative and rhizobacteria. Phytoparasitica 2002, 30, 159–168. [Google Scholar] [CrossRef]
- Sharma, A.; Johri, B.N. Growth promoting influence of siderophore producing Pseudomonas strains GRP3A and PRS9 in maize (Zea mays L.) under iron limiting conditions. Microbiol. Res. 2003, 158, 243–248. [Google Scholar] [CrossRef]
- Press, C.M.; Loper, J.E.; Kloepper, J.W. Role of iron in rhizobacteria mediated induced systemic resistance of cucumber. Phytopathology 2001, 91, 593–598. [Google Scholar] [CrossRef] [Green Version]
- Singh, D.P.; Gupta, V.K.; Prabha, R. Microbial Interventions in Agriculture and Environment: Rhizosphere, Microbiome and Agro-Ecology; Springer: Singapore, 2019. [Google Scholar]
- Saha, R.; Saha, N.; Donofrio, R.S.; Bestervelt, L.L. Microbial siderophores: A mini review. J. Basic Microbiol. 2013, 53, 303–317. [Google Scholar] [CrossRef]
- Bargaz, A.; Lyamlouli, K.; Chtouki, M.; Zeroual, Y.; Dhiba, D. Soil Microbial Resources for Improving Fertilizers Efficiency in an Integrated Plant Nutrient Management System. Front. Microbiol. 2018, 9, 1606. [Google Scholar] [CrossRef] [Green Version]
- Sharma, S.B.; Sayyed, R.Z.; Trivedi, M.H.; Gobi, T.A. Phosphate solubilizing microbes: Sustainable approach for managing phosphorus deficiency in agricultural soils. Springerplus 2013, 2, 587–600. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anzuay, M.S.; Ludueña, L.M.; Angelini, J.G.; Fabra, A.; Taurian, T. Beneficial effects of native phosphate solubilizing bacteria on peanut (Arachis hypogaea L.) growth and phosphorus acquisition. Symbiosis 2015, 66, 89–97. [Google Scholar] [CrossRef]
- Dipak, P.; Sankar, S. Isolation and characterization of phosphate solubilizing bacterium Pseudomonas aeruginosa KUPSB12 with antibacterial potential from river Ganga, India. Ann. Agrar. Sci. 2017, 15, 130–136. [Google Scholar] [CrossRef] [Green Version]
- Zeng, Q.; Wu, X.; Wang, J.; Ding, X. Phosphate solubilization and gene expression of phosphate-solubilizing bacterium Burkholderia multivorans WS-FJ9 under different levels of soluble phosphate. J. Microbiol. Biotechnol. 2017, 27, 844–855. [Google Scholar] [CrossRef]
- Pande, A.; Pandey, P.; Mehra, S.; Singh, M.; Kaushik, S. Phenotypic and genotypic characterization of phosphate solubilizing bacteria and their efficiency on the growth of maize. J. Genet. Eng. Biotechnol. 2017, 15, 379–391. [Google Scholar] [CrossRef]
- Zhang, S.; Moyne, A.L.; Reddy, M.S.; Kloepper, J.W. The role of salicylic acid in induced systemic resistance elicited by plant growth-promoting rhizobacteria against blue mold of tobacco. Biol. Control 2002, 25, 288–296. [Google Scholar] [CrossRef]
- Dubuis, C.; Keel, C.; Haas, D. Dialogues of root-colonizing biocontrol pseudomonads. Eur. J. Plant Pathol. 2007, 119, 311–328. [Google Scholar] [CrossRef] [Green Version]
- Raaijmakers, J.M.; Paulitz, T.C.; Steinberg, C.; Alabouvette, C.; Moënne-Loccoz, Y. The rhizosphere: A playground and battlefeld for soilborne pathogens and benefcial microorganisms. Plant Soil 2009, 321, 341–361. [Google Scholar] [CrossRef] [Green Version]
- Danese, P.N.; Pratt, L.A.; Kolter, R. Exopolysaccharide production is required for development of Escherichia coli K-12 biofilm architecture. J. Bacteriol. 2000, 182, 3593–3596. [Google Scholar] [CrossRef] [Green Version]
- Zhu, M.-L.; Wu, X.-Q.; Wang, Y.-H.; Dai, Y. Role of Biofilm Formation by Bacillus pumilus HR10 in Biocontrol against Pine Seedling Damping-Off Disease Caused by Rhizoctonia solani. Forests 2020, 11, 652. [Google Scholar] [CrossRef]
- Shehata, H.R.; Ettinger, C.L.; Eisen, J.A.; Raizada, M.N. Genes Required for the Anti-fungal Activity of a Bacterial Endophyte Isolated from a Corn Landrace Grown Continuously by Subsistence Farmers Since 1000 BC. Front. Microbiol. 2016, 7, 1548. [Google Scholar] [CrossRef] [Green Version]
- Kandel, S.L.; Joubert, P.M.; Doty, S.L. Bacterial Endophyte Colonization and Distribution within Plants. Microorganisms 2017, 5, 77. [Google Scholar] [CrossRef] [PubMed]
- Hover, T.; Maya, T.; Ron, S.; Sandovsky, H.; Shadkchan, Y.; Kijner, N.; Mitiagin, Y.; Fichtman, B.; Harel, A.; Shanks, R.M.; et al. Mechanisms of Bacterial (Serratia marcescens) Attachment to, Migration along, and Killing of Fungal Hyphae. Appl. Environ. Microbiol. 2016, 82, 2585–2594. [Google Scholar] [CrossRef] [Green Version]
- Chlebek, D.; Pinski, A.; Żur, J.; Michalska, J.; Hupert-Kocurek, K. Genome Mining and Evaluation of the Biocontrol Potential of Pseudomonas fluorescens BRZ63, a New Endophyte of Oilseed Rape (Brassica napus L.) against Fungal Pathogens. Int. J. Mol. Sci. 2020, 21, 8740. [Google Scholar] [CrossRef] [PubMed]
- Huerta-Cepas, J.; Szklarczyk, D.; Heller, D.; Hernández-Plaza, A.; Forslund, S.K.; Cook, H.; Mende, D.R.; Letunic, I.; Rattei, T.; Jensen, L.J.; et al. EggNOG 5.0: A hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 2019, 47, D309–D314. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kanehisa, M.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016, 44, D457–D462. [Google Scholar] [CrossRef] [Green Version]
- Blin, K.; Shaw, S.; Steinke, K.; Villebro, R.; Ziemert, N.; Lee, S.Y.; Medema, M.H.; Weber, T. AntiSMASH 5.0: Updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019, 47, W81–W87. [Google Scholar] [CrossRef] [Green Version]
- Yin, Y.; Mao, X.; Yang, J.; Chen, X.; Mao, F.; Xu, Y. dbCAN: A web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 2012, 40, W445–W451. [Google Scholar] [CrossRef]
- Zhang, H.; Yohe, T.; Huang, L.; Entwistle, S.; Wu, P.; Yang, Z.; Busk, P.K.; Xu, Y.; Yin, Y. dbCAN2: A meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 2018, 46, W95–W101. [Google Scholar] [CrossRef] [Green Version]
- Bertelli, C.; Laird, M.R.; Williams, K.P.; Simon Fraser University Research Computing Group; Lau, B.Y.; Hoad, G.; Winsor, G.L.; Brinkman, F.S.L. IslandViewer 4: Expanded prediction of genomic islands for larger-scale datasets. Nucleic Acids Res. 2017, 45, W30–W35. [Google Scholar] [CrossRef]
- Avram, O.; Rapoport, D.; Portugez, S.; Pupko, T. M1CR0B1AL1Z3R—A user-friendly web server for the analysis of large-scale microbial genomics data. Nucleic Acids Res. 2019, 47, W88–W92. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Talavera, G.; Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 2007, 56, 564–577. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Vicente, C.S.L.; Nascimento, F.X.; Ikuyo, Y.; Cock, P.J.; Mota, M.; Hasegawa, K. The genome and genetics of a high oxidative stress tolerant Serratia sp. LCN16 isolated from the plant parasitic nematode Bursaphelenchus xylophilus. BMC Genom. 2016, 17, 301. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zur, J.; Pinski, A.; Wojcieszynska, D.; Smułek, W.; Guzik, U. Diclofenac degradation—Enzymes, genetic background and cellular alterations triggered in diclofenac-metabolizing strain Pseudomonas moorei KB4. Int. J. Mol. Sci. 2020, 21, 6786. [Google Scholar] [CrossRef] [PubMed]
- Dąbrowska, G.B.; Tylman-Mojżeszek, W.; Mierek-Adamska, A.; Richert, A.; Hrynkiewicz, K. Potential of Serratia plymuthica IV-11-34 strain for biodegradation of polylactide and poly(ethylene terephthalate). Int. J. Biol. Macromol. 2021, 193, 145–153. [Google Scholar] [CrossRef] [PubMed]
- Taylor, S.; Wakem, M.; Dijkman, G.; Alsarraj, M.; Nguyen, M. A practical approach to RT-qPCR-Publishing data that conform to the MIQE guidelines. Methods 2010, 50, S1–S5. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Ghose, T.K. Measurement of cellulase activities. Pure Appl. Chem. 1987, 59, 257–268. [Google Scholar] [CrossRef]
- Rais, A.; Jabeen, Z.; Shair, F.; Hafeez, F.Y.; Hassan, M.N. Bacillus spp., a bio-control agent enhances the activity of antioxidant defense enzymes in rice against Pyricularia oryzae. PLoS ONE 2017, 12, e0187412. [Google Scholar] [CrossRef] [Green Version]
- Zarei, M.; Aminzadeh, S.; Zolgharnein, H.; Safahieh, A.; Daliri, M.; Noghabi, K.A.; Ghoroghi, A.; Motallebi, A. Characterization of a chitinase with antifungal activity from a native Serratia marcescens B4A. Braz. J. Microbiol. 2011, 42, 1017–1029. [Google Scholar] [CrossRef] [Green Version]
- Zhang, C.; Bruins, M.E.; Yang, Z.Q.; Liu, S.T.; Rao, P.F. A new formula to calculate activity of superoxide dismutase in indirect assays. Anal. Biochem. 2016, 503, 65–67. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, G.; Pandey, S.; Ray, A.K.; Kumar, R. Bioremediation of heavy metals by a novel bacterial strain Enterobacter cloaca and its antioxidant enzyme activity, flocculant production and protein expression in presence of lead, cadmium and nickel. Water Air Soil Pollut. 2015, 226, 91. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of proteins utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef] [PubMed]
- Johnston-Monje, D.; Raizada, M.N. Conservation and diversity of seed associated endophytes in Zea across boundaries of evolution, ethnography and ecology. PLoS ONE 2013, 6, e20396. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Syamala, M.; Sivaji, M. Functional characterization of various plant growth promoting activity of Pseudomonas fluorescens and Bacillus subtilis from Aloe vera rhizosphere. J. Pharmacogn. Phytochem. 2017, 6, 120–122. [Google Scholar]
- Sandhya, V.; Shrivastava, M.; Ali, S.Z.; Prasad, V.S.S.K. Endophytes from maize with plant growth promotion and biocontrol activity under drought stress. Russ. Agric. Sci. 2017, 43, 22–34. [Google Scholar] [CrossRef]
- Cappuccino, J.G.; Sherman, N. Biochemical Activities of Microorganisms. Microbiology, A Laboratory Manual; The Benjamin/Cummings Publishing Co.: San Francisco, CA, USA, 1992; pp. 188–247. [Google Scholar]
- Schwyn, B.; Neilands, J.B. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 1987, 160, 47–56. [Google Scholar] [CrossRef]
- Pikovskaya, R.I. Mobilization of phosphorus in soil in connection with vital activity of some microbial species. Microbiology 1948, 17, 362–370. [Google Scholar]
- Naveed, M.; Mitter, B.; Yousaf, S.; Pastar, M.; Afzal, M.; Sessitsch, A. The endophyte Enterobacter sp. FD17: A maize growth enhancer selected based on rigorous testing of plant beneficial traits and colonization characteristics. Biol. Fertil. Soils 2014, 50, 249–262. [Google Scholar] [CrossRef]
- Freeman, D.J.; Falkiner, F.R.; Keane, C.T. New method for detecting slime production by coagulase negative staphylococci. J. Clin. Pathol. 1989, 42, 872–874. [Google Scholar] [CrossRef] [Green Version]
- Taghadosi, R.; Shakibaie, M.R.; Masoumi, S. Biochemical detection of N-Acyl homoserine lactone from biofilm-forming uropathogenic Escherichia coli isolated from urinary tract infection samples. Rep. Biochem. Mol. Biol. 2015, 3, 56–61. [Google Scholar] [PubMed]
- Michalska, J.; Piński, A.; Żur, J.; Mrozik, A. Selecting Bacteria Candidates for the Bioaugmentation of Activated Sludge to Improve the Aerobic Treatment of Landfill Leachate. Water 2020, 12, 140. [Google Scholar] [CrossRef]

| Attribute | Value |
|---|---|
| Genome size (bp) | 5,456,872 |
| Contigs | 107 |
| G + C content (%) | 64 |
| Genes (total) | 5194 |
| CDSs (total) | 5098 |
| Genes (coding) | 5024 |
| Protein genes | 5024 |
| RNA genes | 126 |
| rRNAs | 35 |
| tRNAs | 72 |
| ncRNAs | 12 |
| Pseudogenes | 44 |
| Genes assigned to COGs | 5018 |
| Genes assigned to KEGG pathways | 3342 |
| BioProject ID | PRJNA743191 |
| BioSample ID | SAMN20003760 |
| GenBank accession number | JAHTKS000000000.1 |
| Enzyme’s Activity | KP32 | KP32 + RS | KP32 + FA | KP32 + CD | KP32 + SS |
|---|---|---|---|---|---|
| Protease (U mL−1) | 10.32 ± 0.45 a | 24.03 ± 0.21 b | 9.05 ± 0.09 c | 10.89 ± 0.10 d | 4.98 ± 0.24 e |
| Amylase (U mL−1) | 0.61 ± 0.25 a | 0.64 ± 0.03 a | 0.64 ± 0.01 a | 0.21 ± 0.11 b | 0.11 ± 0.14 c |
| Cellulase (U mL−1) | 0.00 ± 0.00 a | 0.00 ± 0.00 a | 0.00 ± 0.00 a | 0.00 ± 0.00 a | 0.00 ± 0.00 a |
| Chitinase (U mL−1) | 0.84 ± 0.12 a | 0.85 ± 0.04 b | 0.89 ± 0.21 a | 2.12 ± 0.10 c | 1.32 ± 05 d |
| Catalase (U mg−1 of protein) | 2.84 ± 0.25 a | 8.49 ± 0.07 b | 4.05 ± 0.51 c | 4.89 ± 0.10 c | 2.47 ± 0.14 d |
| Superoxide dismutase (U mg−1 of protein) | 0.43 ± 0.06 a | 0.32 ± 0.09 a | 0.60 ± 0.08 b | 0.32 ± 0.11 c | 0.24 ± 0.02 d |
| Features | Strain KP32 |
|---|---|
| Plant growth promotion | |
| Acetoin and 2,3-butanediol production | + |
| IAA production (μg/mL) | 14.32 ± 0.12 |
| SA production (μg/mL) | 5.43 ± 0.89 |
| ACC deaminase production | + |
| Ammonia production | + |
| HCN production | + |
| Siderophore production | + |
| Phosphate solubilization (PSI) | 2.75 ± 11 |
| Colonization properties | |
| Autoaggregation (%) | |
| 2 h | 4.08 ± 0.21 |
| 24 h | 34.2 ± 0.09 |
| Biofilm formation (OD590 of crystal violet) | |
| 24 h | 0.297 ± 0.02 |
| 48 h | 0.40 ± 0.12 |
| 72 h | 0.52 ± 0.05 |
| Motility (mm) | |
| Swimming (0.3%) | 3.20 ± 0.3 |
| Swarming (0.5%) | 2.10 ± 0.07 |
| Twitching (1%) | 0.78 ± 0.15 |
| Exopolysacharydes production | + |
| N-AHLs production | + |
| Gene | Protein | Forward (5′-3’) | Reverse (5′-3’) | Tm (°C) | PCR Efficiency (%) | Slope | Product Size (bp) |
|---|---|---|---|---|---|---|---|
| gyrA | DNA gyrase subunit A | TGCGCTATATGCTGGTGGAC | GCAATTTTGGACATGCGCAC | 53.8 51.8 | 97.24 | −3.396 | 100 |
| gyrB | DNA gyrase subunit B | CGGCGGCAAATTTGATGACA | AACCAGTTCCAGCTTCTCGG | 51.8 53.8 | 99.00 | −3.351 | 100 |
| chiA | Chitinase | TGGAATGGCGATACCGGTAC | CCTTAAAGTTTGCCGTGCCC | 53.8 53.8 | 101.35 | −3.297 | 100 |
| budA | Alpha-acetolactate decarboxylase | CGGTGTTTACGAAGGGGAGG | GAAGGCGATCAGTTCACCGT | 55.9 53.8 | 90.94 | −3.567 | 100 |
| hcnC | Hydrogen cyanide synthase | ACAGCACTATCGACATGCCG | CCAGTCCAGCAGCGGATAAT | 53.8 53.8 | 106.76 | −3.178 | 100 |
| iucA | Aerobactic synthase | GTATGCCCCGGAATACCAGG | CTGGGTCAGCGGATATGCTT | 55.9 53.8 | 109.00 | −3.116 | 100 |
| entB | Enterobactin synthase | GATCAAGCAGGTGGTGGAGA | ATCGCTCTGCTGATTTGGCT | 53.8 51.8 | 108.20 | −3.147 | 100 |
| pchB | Isochorismate pyruvate lyase | TCATTAAGCTGATCGCCCGG | ATGGCCTCAAAGCGCTCTTT | 53.8 51.8 | 105.15 | −3.202 | 100 |
| katG | Catalase | GTTCACATTCCCAACTGCGC | ATCACCTTATTCCAGGCGGC | 53.8 53.8 | 105.35 | −3.201 | 100 |
| sodB | Superoxide dismutase | CGGCGGCATCTTCAACAATG | GGCCAGTTTACCTTCAGGCT | 53.8 53.8 | 104.53 | −3.219 | 100 |
| mtlR | Mannitol dehydrogenase | TCCCTTAAGTGAACGCCTCG | ATCGTGGCCAAACACCGTAT | 51.8 53.8 | 110.28 | −3.098 | 100 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chlebek, D.; Grebtsova, V.; Piński, A.; Żur-Pińska, J.; Hupert-Kocurek, K. Genetic Determinants of Antagonistic Interactions and the Response of New Endophytic Strain Serratia quinivorans KP32 to Fungal Phytopathogens. Int. J. Mol. Sci. 2022, 23, 15561. https://doi.org/10.3390/ijms232415561
Chlebek D, Grebtsova V, Piński A, Żur-Pińska J, Hupert-Kocurek K. Genetic Determinants of Antagonistic Interactions and the Response of New Endophytic Strain Serratia quinivorans KP32 to Fungal Phytopathogens. International Journal of Molecular Sciences. 2022; 23(24):15561. https://doi.org/10.3390/ijms232415561
Chicago/Turabian StyleChlebek, Daria, Valeriia Grebtsova, Artur Piński, Joanna Żur-Pińska, and Katarzyna Hupert-Kocurek. 2022. "Genetic Determinants of Antagonistic Interactions and the Response of New Endophytic Strain Serratia quinivorans KP32 to Fungal Phytopathogens" International Journal of Molecular Sciences 23, no. 24: 15561. https://doi.org/10.3390/ijms232415561







