Mathematical Modeling of ROS Production and Diode-like Behavior in the SDHA/SDHB Subcomplex of Succinate Dehydrogenases in Reverse Quinol-Fumarate Reductase Direction
Abstract
:1. Introduction
2. Results and Discussion
2.1. Eout Dependency of the Total Rate of Reverse Electron Transfer in SDH
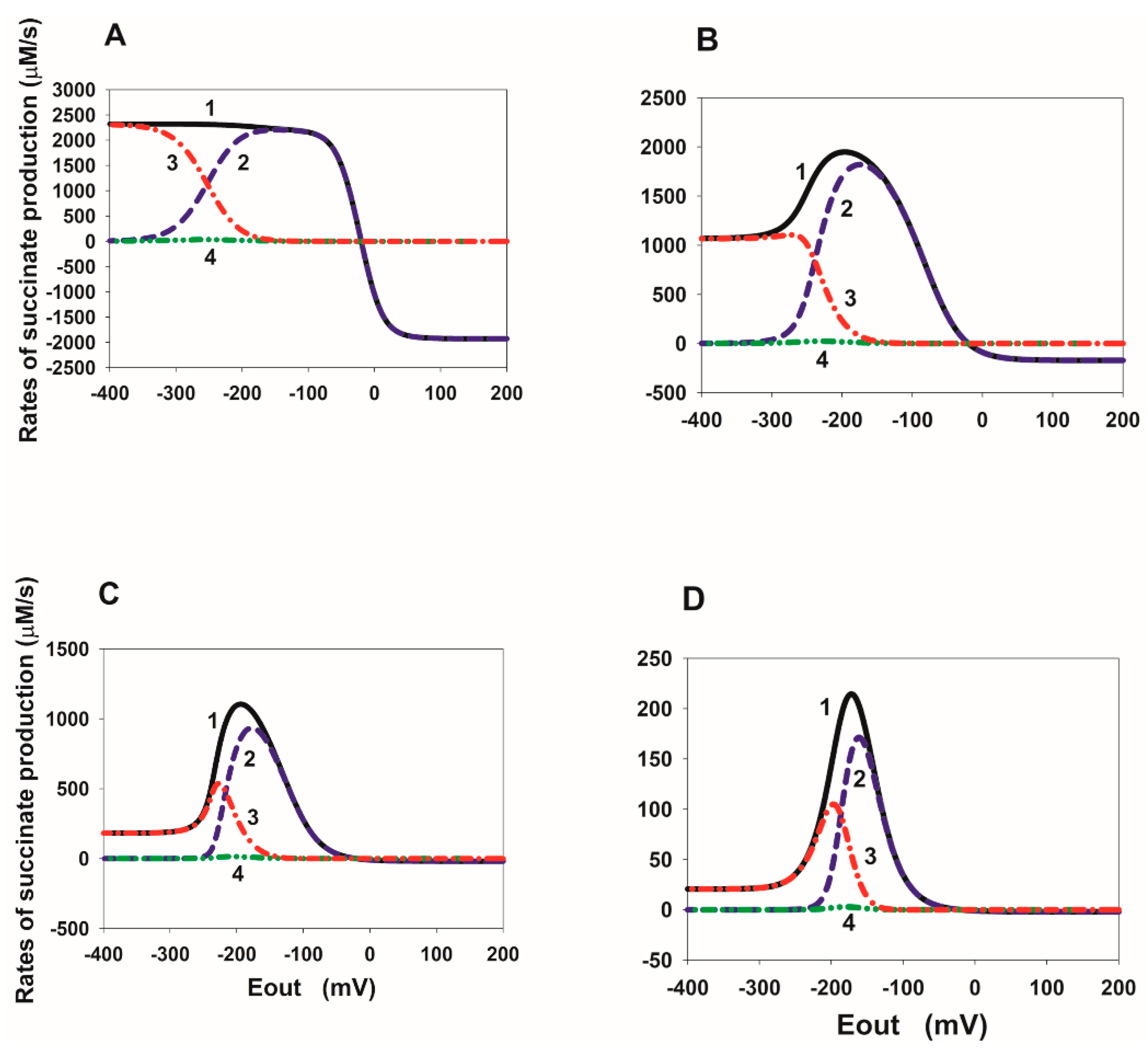
2.2. Eout Dependency of the Rate of Succinate Production in SDH
2.3. Computational Analysis of the ROS Production Rate in the SDHA/SDHB Subcomplex during Reverse Electron Transfer
2.4. Dependence of the Rate of Succinate Production on the Fumarate Concentration in SDH
3. Methods and Models
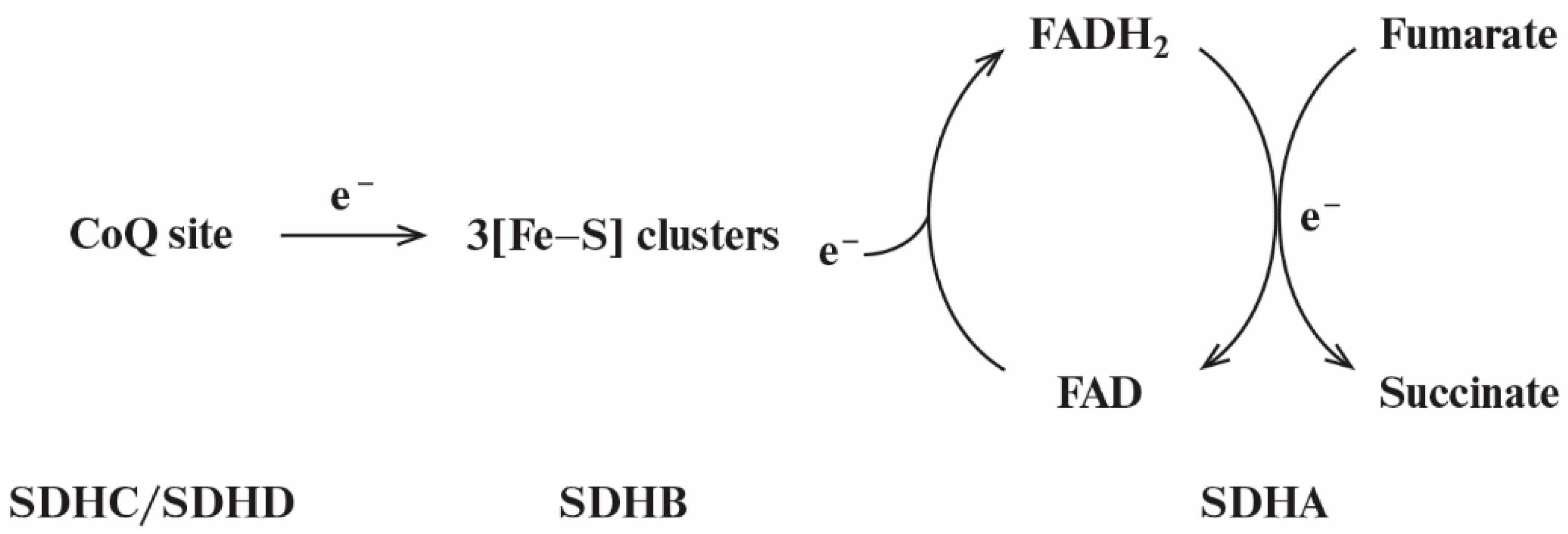
3.1. Kinetic Model of Reverse Electron Transfer in SDHA/SDHB Subunits of SDH
- (1)
- Keq4 ∙ Keq5 ∙ Keq6 ∙Keq7 ∙ Keq8 = 1;
- (2)
- Keq6 ∙ Keq10/(Keq11 ∙ Keq12) = 1;
- (3)
- Keq8 ∙ Keq9/(Keq13 ∙ Keq14) = 1;
- (4)
- Keq7 ∙ Keq12 ∙ Keq13 ∙Keq15 ∙ Keq16 = 1.
3.2. Computational Model of Reverse Electron Transfer in SDHA/SDHB Subunits of SDH. Mathematical Model
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Markevich, N.I.; Galimova, M.H.; Markevich, L.N. Hysteresis and bistability in the succinate-CoQ reductase activity and reactive oxygen species production in the mitochondrial respiratory complex II. Redox Biol. 2020, 37, 101630. [Google Scholar] [CrossRef] [PubMed]
- Sucheta, A.; Ackrell, B.A.; Cochran, B.; Armstrong, F.A. Diode-like behaviour of a mitochondrial electron-transport enzyme. Nature 1992, 356, 361–362. [Google Scholar] [CrossRef] [PubMed]
- Spinelli, J.B.; Rosen, P.C.; Sprenger, H.G.; Puszynska, A.M.; Mann, J.L.; Roessler, J.M.; Cangelosi, A.L.; Henne, A.; Condon, K.J.; Zhang, T.; et al. Fumarate is a terminal electron acceptor in the mammalian electron transport chain. Science 2021, 374, 1227–1237. [Google Scholar] [CrossRef] [PubMed]
- Chouchani, E.T.; Pell, V.R.; Gaude, E.; Aksentijević, D.; Sundier, S.Y.; Robb, E.L.; Logan, A.; Nadtochiy, S.M.; Ord, E.N.J.; Smith, A.C.; et al. Ischaemic accumulation of succinate controls reperfusion injury through mitochondrial ROS. Nature 2014, 515, 431–435. [Google Scholar] [CrossRef] [Green Version]
- Chinopoulos, C. Succinate in ischemia: Where does it come from? Int. J. Biochem. Cell Biol. 2019, 115, 105580. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, Y.T.; Miller, J.H.; Day, M.M.; Munger, J.C.; Brookes, P.S. Accumulation of Succinate in Cardiac Ischemia Primarily Occurs via Canonical Krebs Cycle Activity. Cell Rep. 2018, 23, 2617–2628. [Google Scholar] [CrossRef] [Green Version]
- Pershad, H.R.; Hirst, J.; Cochran, B.; Ackrell, B.A.; Armstrong, F.A. Voltammetric studies of bidirectional catalytic electron transport in Escherichia coli succinate dehydrogenase: Comparison with the enzyme from beef heart mitochondria. Biochim. Biophys. Acta 1999, 1412, 262–272. [Google Scholar] [CrossRef] [Green Version]
- Hearon, J.Z. The kinetics of linear systems with special reference to periodic Reactions. Bull. Math. Biophys. 1953, 15, 121–141. [Google Scholar] [CrossRef]
- Bonomi, F.; Pagani, S.; Cerletti, P.; Giori, C. Modification of the Thermodynamic Properties of the Electron-Transferring Groups in Mitochondrial Succinate Dehydrogenase upon Binding of Succinate. Eur. J. Biochem. 1983, 134, 439–445. [Google Scholar] [CrossRef]
- Quinlan, C.L.; Orr, A.L.; Perevoshchikova, I.V.; Treberg, J.R.; Ackrell, B.A.; Brand, M.D. Mitochondrial complex II can generate reactive oxygen species at high rates in both the forward and reverse reaction. J. Biol. Chem. 2012, 287, 27255–27264. [Google Scholar] [CrossRef]
- Grivennikova, V.G.; Kozlovsky, V.S.; Vinogradov, A.D. Respiratory complex II: ROS production and the kinetics of ubiquinone reduction. Biochim. Biophys. Acta 2017, 1858, 109–117. [Google Scholar] [CrossRef] [PubMed]
- Manhas, N.; Duong, Q.V.; Lee, P.; Richardson, J.; Robertson, J.D.; Moxley, M.A.; Bazil, J.N. Computationally modeling mammalian succinate dehydrogenase kinetics identifies the origins and primary determinants of ROS production. J. Biol. Chem. 2020, 295, 15262–15279. [Google Scholar] [CrossRef] [PubMed]
- Markevich, N.I.; Markevich, L.N.; Hoek, J.B. Computational Modeling Analysis of Generation of Reactive Oxygen Species by Mitochondrial Assembled and Disintegrated Complex II. Front. Physiol. 2020, 11, 557721. [Google Scholar] [CrossRef] [PubMed]
- Siebels, I.; Dröse, S. Q-site inhibitor induced ROS production of mitochondrial complex II is attenuated by TCA cycle dicarboxylates. Biochim. Biophys. Acta 2013, 1827, 1156–1164. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kotlyar, A.B.; Vinogradov, A.D. Interaction of the membrane-bound succinate dehydrogenase with substrate and competitive inhibitors. Biochim. Biophys. Acta 1984, 784, 24–34. [Google Scholar] [CrossRef] [PubMed]
- Maklashina, E.; Cecchini, G.; Dikanov, S.A. Defining a Direction: Electron Transfer and Catalysis in Escherichia Coli Complex II Enzymes. Biochim. Biophys. Acta 2013, 1827, 668–678. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ohnishi, T.; Lim, J.; Winter, D.B.; King, T.E. Thermodynamic and EPR characteristics of a HiPIP-type iron–sulfur center in the succinate dehydrogenase of the respiratory chain. J. Biol. Chem. 1976, 251, 2105–2109. [Google Scholar] [CrossRef]
- Ohnishi, T.; Salerno, J.C. Thermodynamic and EPR characteristics of two ferredoxin-type iron-sulfur centers in the succinate-ubiquinone reductase segment of the respiratory chain. J. Biol. Chem. 1976, 251, 2094–2104. [Google Scholar] [CrossRef]
- Ohnishi, T.; King, T.E.; Salerno, J.C.; Blum, H.; Bowyer, J.R.; Maida, T. Thermodynamic and electron paramagnetic resonance characterization of flavin in succinate dehydrogenase. J. Biol. Chem. 1981, 256, 5577–5582. [Google Scholar] [CrossRef]
- Murphy, M.P. How mitochondria produce reactive oxygen species. Biochem. J. 2009, 417, 1–13. [Google Scholar] [CrossRef]
- Markevich, N.I.; Hoek, J.B. Computational modeling analysis of mitochondrial superoxide production under varying substrate conditions and upon inhibition of different segments of the electron transport chain. Biochim. Biophys. Acta 2015, 1847, 656–679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Markevich, N.I.; Hoek, J.B. Computational modeling analysis of acute and chronic ethanol- induced oxidative stress. Math. Biol. Bioinformat. 2014, 9, 63–88. [Google Scholar]
- Jin, Z.Q.; Zhou, H.Z.; Cecchini, G.; Gray, M.O.; Karliner, J.S. MnSOD in mouse heart:Acute responses ischemic preconditioning and ischemia–reperfusion injury. Am. J. Physiol. Heart Circ. Physiol. 2005, 288, H2986–H2994. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wojtovich, A.P.; Nehrke, K.W.; Brookes, P.S. The mitochondrial complex II and ATP- sensitive potassium channel interaction: Quantitation of the channel in heart mitochondria. Acta Biochim. 2010, 57, 431–434. [Google Scholar] [CrossRef]
- Deichmann, U.; Schuster, S.; Mazat, J.P.; Cornish-Bowden, A. Commemorating the 1913 Michaelis-Menten paper Die Kinetik der Invertinwirkung: Three perspectives. FEBS J. 2014, 281, 435–463. [Google Scholar] [CrossRef]
- Vinnakota, K.C.; Bassingthwaighte, J.B. Myocardial density and composition: A basis for calculating intracellular metabolite concentrations. Am. J. Physiol. Heart Circ. Physiol. 2004, 286, H1742–H1749. [Google Scholar] [CrossRef] [Green Version]
- Hackenbrock, C.R.; Chazotte, B.; Gupte, S.S. The random collision model and a critical assessment of diffusion and collision in mitochondrial electron transport. J. Bioenerg. Biomembr. 1986, 18, 331–368. [Google Scholar] [CrossRef]
- Wu, F.; Yang, F.; Vinnakota, K.C.; Beard, D.A. Computer modeling of mitochondrial tricarboxylic acid cycle, oxidative phosphorylation, metabolite transport, and electrophysiology. J. Biol. Chem. 2007, 282, 24525–24537. [Google Scholar] [CrossRef] [Green Version]
- Schwerzmann, K.; Cruz-Orive, L.M.; Eggman, R.; Sanger, A.; Weibel, E.R. Molecular architecture of the inner membrane of mitochondria from rat liver: A combined biochemical and stereological study. J. Cell. Biol. 1986, 102, 97–103. [Google Scholar] [CrossRef]









| No | Reaction | Rate Equation |
|---|---|---|
| 1 | e− + [3Fe-4S] = [3Fe-4S]− | V1 = k1 ∙ ([3Fe-4S] ∙ exp(-α ∙ F ∙ Eout/RT) − [3Fe-4S]− ∙ exp((1-α) ∙ F ∙ Eout/RT)/Keq1) |
| 2 | [4Fe-4S] + [3Fe-4S]− = [4Fe-4S]− + [3Fe-4S] | V2 = k2 ∙ ([4Fe-4S] ∙ [3Fe-4S]− − [4Fe-4S]− ∙ [3Fe-4S]/Keq2) |
| 3 | [2Fe-2S] + [4Fe-4S]− = [2Fe-2S]− + [4Fe-4S] | V3 = k3 ∙ ([2Fe-2S] ∙ [4Fe-4S]− − [2Fe-2S]− ∙ [4Fe-4S]/Keq3) |
| 4 | FAD + fum = FAD.fum | V4 = k4 ∙ (FAD ∙ fum − FAD.fum/Keq4) |
| 5 | FAD.fum + [2Fe-2S]− + H + = FADH•.fum + [2Fe-2S] | V5 = k5 ∙ (FAD.fum ∙ [2Fe-2S]− ∙ H + − FADH•.fum ∙ [2Fe-2S]/Keq5) |
| 6 | FADH•.fum + [2Fe-2S]− + H + = FADH2.fum + [2Fe-2S] | V6 = k6 ∙ (FADH•.fum ∙ [2Fe-2S]− ∙ H + − FADH2.fum ∙ [2Fe-2S]/Keq6) |
| 7 | FADH2.fum = FAD.suc | V7 = k7 ∙ (FADH2.fum − FAD.suc/Keq7) |
| 8 | FAD.suc = FAD + suc | V8 = k9 ∙ (FAD.suc − FAD ∙ suc/Keq8) |
| 9 | FAD + [2Fe-2S]− + H + = FADH• + [2Fe-2S] | V9 = k9 ∙ (FAD ∙ [2Fe-2S]− ∙ H + − FADH• ∙ [2Fe-2S]/Keq9) |
| 10 | FADH• + fum = FADH•.fum | V10 = k10 ∙ (FADH• ∙ fum − FADH•.fum/Keq10) |
| 11 | FADH• + [2Fe-2S]− + H + = FADH2 + [2Fe-2S] | V11 = k11 ∙ (FADH• ∙ [2Fe-2S]− ∙ H + − FADH2 ∙ [2Fe-2S]/Keq11) |
| 12 | FADH2 + fum = FADH2.fum | V12 = k12 ∙ (FADH2 ∙ fum − FADH2.fum/Keq12) |
| 13 | FAD.suc + [2Fe-2S]− + H + = FADH•.suc + [2Fe-2S] | V13 = k13 ∙ (FAD.suc ∙ [2Fe-2S]− ∙ H + − FADH•.suc ∙ [2Fe-2S]/Keq13) |
| 14 | FADH•.suc = suc + FADH• | V14 = k14 ∙ (FADH•.suc − FADH• ∙ suc/Keq14) |
| 15 | FADH•.suc + [2Fe-2S]− + H + = FADH2.suc + [2Fe-2S] | V15 = k15 ∙ (FADH•.suc ∙ [2Fe-2S]− ∙ H + − FADH2.suc ∙ [2Fe-2S]/Keq15) |
| 16 | FADH2.suc = suc + FADH2 | V16 = k16 ∙ (FADH2.suc − FADH2 ∙ suc/Keq16) |
| Hydrogen peroxide (H2O2) production by SDH | ||
| 17 | FADH2 + O2 = FAD + H2O2 | V17 = k17 ∙ (FADH2 ∙ O2 − FAD ∙ H2O2/Keq17) |
| Superoxide anion (O2∙−) production by the subcomplex SDHA/SDHB of SDH | ||
| 18 | FADH2 + O2 = FADH• + O2∙− + H + | V18 = k18 ∙ (FADH2 ∙ O2 − FADH• ∙ O2∙− ∙ H + /Keq18) |
| 19 | FADH• + O2 = FAD + O2∙− + H + | V19 = k19 ∙ (FADH• ∙ O2 − FAD ∙ O2∙− ∙ H + /Keq19) |
| 20 | [3Fe-4S]− + O2 = [3Fe-4S] + O2∙− | V20 = k20 ∙ ([3Fe-4S]− ∙ O2 − [3Fe-4S] ∙ O2∙−/Keq20) |
| Superoxide anion dismutation in the mitochondrial matrix | ||
| 21 | 2 O2∙− + 2H + → O2 + H2O2 | V21 = Vmax21 ∙ O2∙−/(Km21 + O2∙−) |
| Release of hydrogen peroxide (H2O2) from the mitochondrial matrix to cytosol | ||
| 22 | H2O2 → | V22 = k22 ∙ H2O2 |
| Reaction No | Midpoint Potential Em = E, (mV) | Equilibrium Constant Keq | kforward | Other Parameters | Reference |
|---|---|---|---|---|---|
| 1 | E([3Fe-4S]) = 60 | 11.023 | 1∙103 s−1 | α = 0.5 α is a certain coefficient that varies from 0 to 1. Eout—electrode potential. F, R and T have a usual meaning. | [17] a |
| 2 | E([4Fe-4S]) = −260 E([3Fe-4S]) = 60 | 2.78∙10−6 | 1∙104 µM−1·s−1 | pH = 7.4 | [17,18] a |
| 3 | E([2Fe-2S]) = 0 E([4Fe-4S]) = −260 | 3.29∙104 | 1∙104 µM−1·s−1 | pH = 7.4 | [18] a |
| 4 | 4.17∙10−3 µM−1 | 1 s−1 | [15] b | ||
| 5 | E(FAD/FADH•) = −127 E([2Fe-2S]) = 0 | 0.03 e µM−1 | 1∙103 µM−2·s−1 | pH = 7 pH = 7.4 | [18,19] a |
| 6 | E(FADH•/FADH2) = −31 E([2Fe-2S]) = 0 | 0.289 e µM−1 | 1∙103 µM−2·s−1 | pH = 7 pH = 7.4 | [18,19] a |
| 7 | 2778 d | 2.78∙106 s−1 | |||
| 8 | 10 µM | 0.5 s−1 | [19] b | ||
| 9 | E(FAD/FADH•) = −127 E([2Fe-2S]) = 0 | 0.006 | 1∙103 µM−2·s−1 | pH = 7 pH = 7.4 | [18,19] a |
| 10 | 0.02 µM−1 | 103 s−1 | [15] b | ||
| 11 | E(FADH•/FADH2) = −31 E([2Fe-2S]) = 0 | 0.289 µM−1 | 1∙103 µM−2·s−1 | pH = 7 pH = 7.4 | [18,19] a |
| 12 | 0.02 µM−1 | 103 s−1 | [15] b | ||
| 13 | E(FAD/FADH•) = −127 E([2Fe-2S]) = 0 | 2.4∙10−4 e µM−1 | 1∙103 µM−2·s−1 | pH = 7 pH = 7.4 | [18,19] a |
| 14 | 250 µM | 10 s−1 | [15] b | ||
| 15 | E(FADH•/FADH2) = −31 E([2Fe-2S]) = 0 | 0.289 e µM−1 | 1∙103 µM−2·s−1 | pH = 7 pH = 7.4 | [18,19] a |
| 16 | 250 µM | 10 s−1 | [15] b | ||
| Hydrogen peroxide (H2O2) production by Complex II | |||||
| 17 | E(O2/H2O2) = 690 E(FAD/FADH2) = −79 | 5.2∙1026 | 0.01 µM−1·s−1 | pH = 7 | [19] a |
| Superoxide anion (O2∙−) production by the subcomplex SDHA/SDHB of SDH | |||||
| 18 | E(O2/O2∙−) = −160 E(FADH•/FADH2) = −31 | 6∙10−3 | 0.01 µM−1·s−1 | pH = 7 | [20] a [19] a |
| 19 | E(O2/O2∙−) = −160 E(FAD/FADH•) = −127 | 0.267 | 0.1 µM−1·s−1 | pH = 7 pH = 7.4 | [20] a [19] a |
| 20 | E(O2/O2∙−) = −160 E([3Fe-4S]) = 60 | 1.5∙10−4 | 1·10−3 µM−1·s−1 | pH = 7 pH = 7.4 | [20] a [17] a |
| Accompanying reactions in the matrix and inner membrane | |||||
| Superoxide anion dismutation in the mitochondrial matrix | |||||
| 21 | Vmax21 = 5.6∙104 µM·s−1 f Km21 = 50 µM | [21] d | |||
| Efflux of hydrogen peroxide (H2O2) from the mitochondrial matrix to cytoplasm | |||||
| 22 | 30 s−1 | [22] c | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Markevich, N.I.; Markevich, L.N. Mathematical Modeling of ROS Production and Diode-like Behavior in the SDHA/SDHB Subcomplex of Succinate Dehydrogenases in Reverse Quinol-Fumarate Reductase Direction. Int. J. Mol. Sci. 2022, 23, 15596. https://doi.org/10.3390/ijms232415596
Markevich NI, Markevich LN. Mathematical Modeling of ROS Production and Diode-like Behavior in the SDHA/SDHB Subcomplex of Succinate Dehydrogenases in Reverse Quinol-Fumarate Reductase Direction. International Journal of Molecular Sciences. 2022; 23(24):15596. https://doi.org/10.3390/ijms232415596
Chicago/Turabian StyleMarkevich, Nikolay I., and Lubov N. Markevich. 2022. "Mathematical Modeling of ROS Production and Diode-like Behavior in the SDHA/SDHB Subcomplex of Succinate Dehydrogenases in Reverse Quinol-Fumarate Reductase Direction" International Journal of Molecular Sciences 23, no. 24: 15596. https://doi.org/10.3390/ijms232415596







