Genome-Wide ChIPseq Analysis of AhR, COUP-TF, and HNF4 Enrichment in TCDD-Treated Mouse Liver
Abstract
:1. Introduction
2. Results
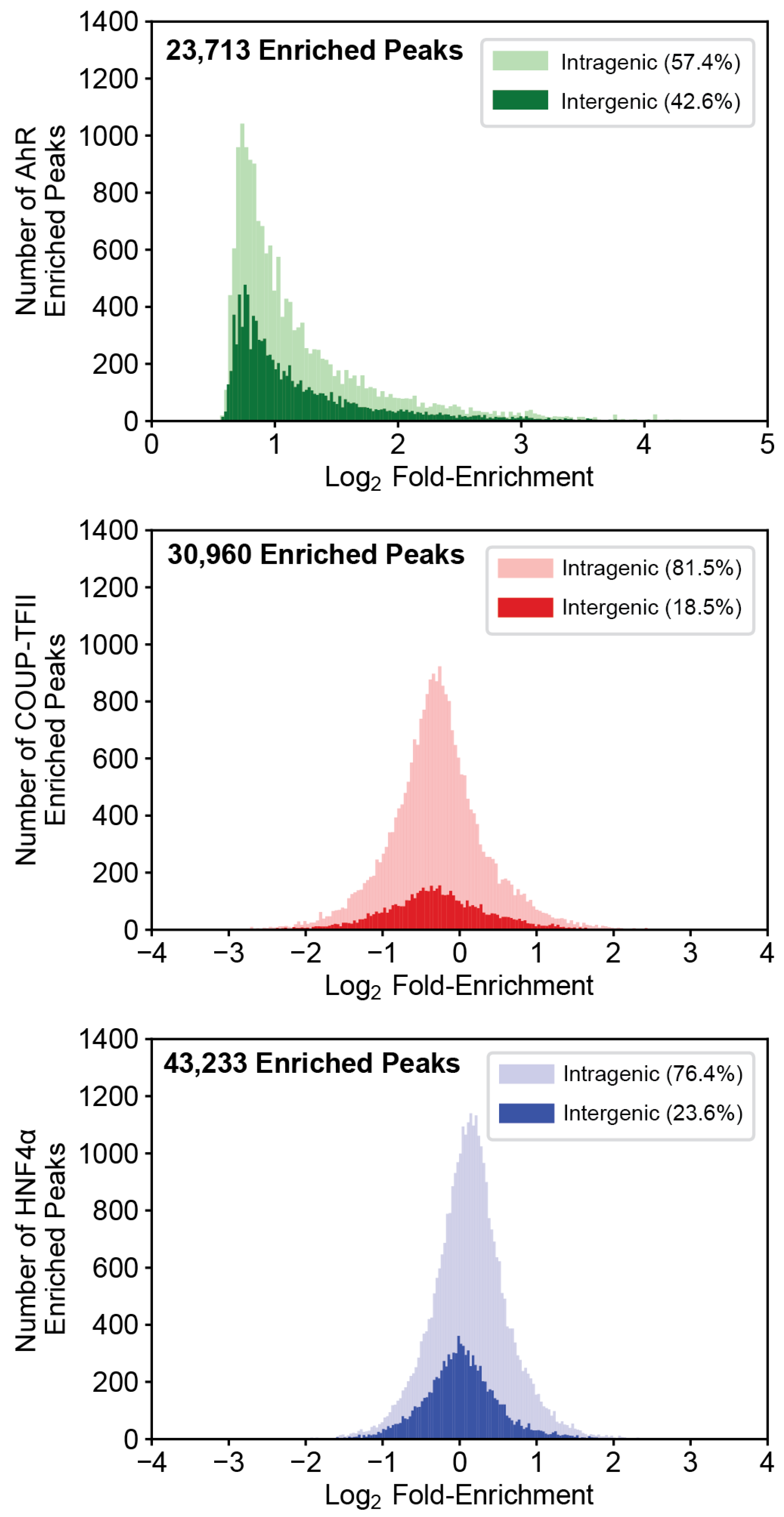
2.1. Evaluation of ChIPseq Datasets
2.2. Evaluation of Transcription Factor Gene Targets
2.3. Evaluation of Gene Transcriptional Changes
2.4. Functional Enrichment of Putatively AhR, COUP-TFII and HNF4α Co-Regulated Genes in Mice Treated with 30 µg/kg for 28 Days
2.5. Identification of AhR, COUP-TFII and HNF4α Binding Patterns in the Liver Secretome
2.6. Identification of AhR, COUP-TFII and HNF4α Binding Patterns in HNF4α-Dependent Genes
3. Discussion
4. Materials and Methods
4.1. Animal Treatment
4.2. ChIP-Sequencing
4.3. RNA Sequencing
4.4. Functional and Motif Enrichment Analyses
4.5. ChIPseq Transcription Factor Overrepresentation Analysis
4.6. Gene Set Enrichment Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Rothhammer, V.; Quintana, F.J. The aryl hydrocarbon receptor: An environmental sensor integrating immune responses in health and disease. Nat. Rev. Immunol. 2019, 19, 184–197. [Google Scholar] [CrossRef] [PubMed]
- Denison, M.S.; Soshilov, A.A.; He, G.; Degroot, D.E.; Zhao, B. Exactly the same but different: Promiscuity and diversity in the molecular mechanisms of action of the aryl hydrocarbon (dioxin) receptor. Toxicol. Sci. 2011, 124, 1–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dere, E.; Lo, R.; Celius, T.; Matthews, J.; Zacharewski, T.R. Integration of genome-wide computation DRE search, AhR ChIP-chip and gene expression analyses of TCDD-elicited responses in the mouse liver. BMC Genom. 2011, 12, 365. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ohtake, F.; Takeyama, K.; Matsumoto, T.; Kitagawa, H.; Yamamoto, Y.; Nohara, K.; Tohyama, C.; Krust, A.; Mimura, J.; Chambon, P.; et al. Modulation of oestrogen receptor signalling by association with the activated dioxin receptor. Nature 2003, 423, 545–550. [Google Scholar] [CrossRef] [PubMed]
- Murphy, K.A.; Villano, C.M.; Dorn, R.; White, L.A. Interaction between the aryl hydrocarbon receptor and retinoic acid pathways increases matrix metalloproteinase-1 expression in keratinocytes. J. Biol. Chem. 2004, 279, 25284–25293. [Google Scholar] [CrossRef] [Green Version]
- Wilson, S.R.; Joshi, A.D.; Elferink, C.J. The tumor suppressor Kruppel-like factor 6 is a novel aryl hydrocarbon receptor DNA binding partner. J. Pharmacol. Exp. Ther. 2013, 345, 419–429. [Google Scholar] [CrossRef] [Green Version]
- Larigot, L.; Juricek, L.; Dairou, J.; Coumoul, X. AhR signaling pathways and regulatory functions. Biochim. Open 2018, 7, 1–9. [Google Scholar] [CrossRef]
- Boverhof, D.R.; Burgoon, L.D.; Tashiro, C.; Chittim, B.; Harkema, J.R.; Jump, D.B.; Zacharewski, T.R. Temporal and dose-dependent hepatic gene expression patterns in mice provide new insights into TCDD-Mediated hepatotoxicity. Toxicol. Sci. 2005, 85, 1048–1063. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pierre, S.; Chevallier, A.; Teixeira-Clerc, F.; Ambolet-Camoit, A.; Bui, L.C.; Bats, A.S.; Fournet, J.C.; Fernandez-Salguero, P.; Aggerbeck, M.; Lotersztajn, S.; et al. Aryl hydrocarbon receptor-dependent induction of liver fibrosis by dioxin. Toxicol. Sci. 2014, 137, 114–124. [Google Scholar] [CrossRef] [Green Version]
- Fernandez-Salguero, P.M.; Hilbert, D.M.; Rudikoff, S.; Ward, J.M.; Gonzalez, F.J. Aryl-hydrocarbon receptor-deficient mice are resistant to 2,3,7,8-tetrachlorodibenzo-p-dioxin-induced toxicity. Toxicol. Appl. Pharmacol. 1996, 140, 173–179. [Google Scholar] [CrossRef]
- Nault, R.; Doskey, C.M.; Fader, K.A.; Rockwell, C.E.; Zacharewski, T. Comparison of Hepatic NRF2 and Aryl Hydrocarbon Receptor Binding in 2,3,7,8-Tetrachlorodibenzo-p-dioxin-Treated Mice Demonstrates NRF2-Independent PKM2 Induction. Mol. Pharmacol. 2018, 94, 876–884. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Doskey, C.M.; Fader, K.A.; Nault, R.; Lydic, T.; Matthews, J.; Potter, D.; Sharratt, B.; Williams, K.; Zacharewski, T. 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) alters hepatic polyunsaturated fatty acid metabolism and eicosanoid biosynthesis in female Sprague-Dawley rats. Toxicol. Appl. Pharmacol. 2020, 398, 115034. [Google Scholar] [CrossRef] [PubMed]
- Cholico, G.N.; Fling, R.R.; Zacharewski, N.A.; Fader, K.A.; Nault, R.; Zacharewski, T.R. Thioesterase induction by 2,3,7,8-tetrachlorodibenzo-p-dioxin results in a futile cycle that inhibits hepatic β-oxidation. Sci. Rep. 2021, 11, 15689. [Google Scholar] [CrossRef] [PubMed]
- Fader, K.A.; Nault, R.; Zhang, C.; Kumagai, K.; Harkema, J.R.; Zacharewski, T.R. 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD)-elicited effects on bile acid homeostasis: Alterations in biosynthesis, enterohepatic circulation, and microbial metabolism. Sci. Rep. 2017, 7, 5921. [Google Scholar] [CrossRef] [PubMed]
- Fader, K.A.; Nault, R.; Ammendolia, D.A.; Harkema, J.R.; Williams, K.J.; Crawford, R.B.; Kaminski, N.E.; Potter, D.; Sharratt, B.; Zacharewski, T.R. 2,3,7,8-Tetrachlorodibenzo-p-dioxin alters lipid metabolism and depletes immune cell populations in the jejunum of C57BL/6 mice. Toxicol. Sci. 2015, 148, 567–580. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Angrish, M.M.; Mets, B.D.; Jones, A.D.; Zacharewski, T.R. Dietary fat is a lipid source in 2,3,7,8-tetrachlorodibenzo-rho-dioxin (TCDD)-elicited hepatic steatosis in C57BL/6 mice. Toxicol. Sci. 2012, 128, 377–386. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nault, R.; Fader, K.A.; Lydic, T.A.; Zacharewski, T.R. Lipidomic Evaluation of Aryl Hydrocarbon Receptor-Mediated Hepatic Steatosis in Male and Female Mice Elicited by 2,3,7,8-Tetrachlorodibenzo-p-dioxin. Chem. Res. Toxicol. 2017, 30, 1060–1075. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nault, R.; Fader, K.A.; Harkema, J.R.; Zacharewski, T. Loss of liver-specific and sexually dimorphic gene expression by aryl hydrocarbon receptor activation in C57BL/6 mice. PLoS ONE 2017, 12, e0184842. [Google Scholar] [CrossRef]
- Angrish, M.M.; Dominici, C.Y.; Zacharewski, T.R. TCDD-elicited effects on liver, serum, and adipose lipid composition in C57BL/6 mice. Toxicol. Sci. 2013, 131, 108–115. [Google Scholar] [CrossRef] [Green Version]
- Lu, H.; Cui, W.; Klaassen, C.D. Nrf2 protects against 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD)-induced oxidative injury and steatohepatitis. Toxicol. Appl. Pharmacol. 2011, 256, 122–135. [Google Scholar] [CrossRef] [Green Version]
- Yeager, R.L.; Reisman, S.A.; Aleksunes, L.M.; Klaassen, C.D. Introducing the “TCDD-inducible AhR-Nrf2 gene battery”. Toxicol. Sci. 2009, 111, 238–246. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Parviz, F.; Matullo, C.; Garrison, W.D.; Savatski, L.; Adamson, J.W.; Ning, G.; Kaestner, K.H.; Rossi, J.M.; Zaret, K.S.; Duncan, S.A. Hepatocyte nuclear factor 4α controls the development of a hepatic epithelium and liver morphogenesis. Nat. Genet. 2003, 34, 292–296. [Google Scholar] [CrossRef] [PubMed]
- Bonzo, J.A.; Ferry, C.H.; Matsubara, T.; Kim, J.H.; Gonzalez, F.J. Suppression of hepatocyte proliferation by hepatocyte nuclear factor 4α in adult mice. J. Biol. Chem. 2012, 287, 7345–7356. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ashraf, U.M.; Sanchez, E.R.; Kumarasamy, S. COUP-TFII revisited: Its role in metabolic gene regulation. Steroids 2019, 141, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Fader, K.A.; Nault, R.; Doskey, C.M.; Fling, R.R.; Zacharewski, T.R. 2,3,7,8-Tetrachlorodibenzo-p-dioxin abolishes circadian regulation of hepatic metabolic activity in mice. Sci. Rep. 2019, 9, 6514. [Google Scholar] [CrossRef]
- Nault, R.; Fader, K.A.; Kirby, M.P.; Ahmed, S.; Matthews, J.; Jones, A.D.; Lunt, S.Y.; Zacharewski, T.R. Pyruvate Kinase Isoform Switching and Hepatic Metabolic Reprogramming by the Environmental Contaminant 2,3,7,8-Tetrachlorodibenzo-p-Dioxin. Toxicol. Sci. 2016, 149, 358–371. [Google Scholar] [CrossRef] [Green Version]
- Franko, A.; Hartwig, S.; Kotzka, J.; Ruoß, M.; Nüssler, A.K.; Königsrainer, A.; Häring, H.U.; Lehr, S.; Peter, A. Identification of the secreted proteins originated from primary human hepatocytes and HepG2 cells. Nutrients 2019, 11, 1795. [Google Scholar] [CrossRef] [Green Version]
- Durinck, S.; Spellman, P.T.; Birney, E.; Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 2009, 4, 1184–1191. [Google Scholar] [CrossRef] [Green Version]
- Walesky, C.; Edwards, G.; Borude, P.; Gunewardena, S.; O’Neil, M.; Yoo, B.; Apte, U. Hepatocyte nuclear factor 4 alpha deletion promotes diethylnitrosamine-induced hepatocellular carcinoma in rodents. Hepatology 2013, 57, 2480–2490. [Google Scholar] [CrossRef] [Green Version]
- Forgacs, A.L.; Kent, M.N.; Makley, M.K.; Mets, B.; DelRaso, N.; Jahns, G.L.; Burgoon, L.D.; Zacharewski, T.R.; Reo, N.V. Comparative metabolomic and genomic analyses of TCDD-elicited metabolic disruption in mouse and rat liver. Toxicol. Sci. 2012, 125, 41–55. [Google Scholar] [CrossRef] [Green Version]
- Nault, R.; Fader, K.A.; Ammendolia, D.A.; Dornbos, P.; Potter, D.; Sharratt, B.; Kumagai, K.; Harkema, J.R.; Lunt, S.Y.; Matthews, J.; et al. Dose-Dependent Metabolic Reprogramming and Differential Gene Expression in TCDD-Elicited Hepatic Fibrosis. Toxicol. Sci. 2016, 154, 253–266. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fader, K.A.; Nault, R.; Kirby, M.P.; Markous, G.; Matthews, J.; Zacharewski, T.R. Convergence of hepcidin deficiency, systemic iron overloading, heme accumulation, and REV-ERBalpha/beta activation in aryl hydrocarbon receptor-elicited hepatotoxicity. Toxicol. Appl. Pharmacol. 2017, 321, 1–17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fling, R.R.; Doskey, C.M.; Fader, K.A.; Nault, R.; Zacharewski, T.R. 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) dysregulates hepatic one carbon metabolism during the progression of steatosis to steatohepatitis with fibrosis in mice. Sci. Rep. 2020, 10, 14831. [Google Scholar] [CrossRef] [PubMed]
- Orlowska, K.; Fling, R.R.; Nault, R.; Zacharewski, T. 2,3,7,8-Tetrachlorodibenzo-p-dioxin elicited decreases in cobalamin inhibits methylmalonyl-CoA mutase activity redirecting propionyl-CoA metabolism to the β–oxidation-like pathway resulting in hepatic accumulation of the toxic intermediate acrylyl-CoA. bioRxiv 2021. [Google Scholar] [CrossRef]
- Klinge, C.M.; Kaur, K.; Swanson, H.I. The aryl hydrocarbon receptor interacts with estrogen receptor alpha and orphan receptors COUP-TFI and ERRα1. Arch. Biochem. Biophys. 2000, 373, 163–174. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.H.; Ing, N.H.; Tsai, S.Y.; O’Malley, B.W.; Tsai, M.J. The COUP-TFs compose a family of functionally related transcription factors. Gene Expr. 1991, 1, 207–216. [Google Scholar]
- Polvani, S.; Pepe, S.; Milani, S.; Galli, A. COUP-TFII in Health and Disease. Cells 2019, 9, 101. [Google Scholar] [CrossRef] [Green Version]
- Kliewer, S.A.; Umesono, K.; Heyman, R.A.; Mangelsdorf, D.J.; Dyck, J.A.; Evans, R.M. Retinoid X receptor-COUP-TF interactions modulate retinoic acid signaling. Proc. Natl. Acad. Sci. USA 1992, 89, 1448–1452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kruse, S.W.; Suino-Powell, K.; Zhou, X.E.; Kretschman, J.E.; Reynolds, R.; Vonrhein, C.; Xu, Y.; Wang, L.; Tsai, S.Y.; Tsai, M.J.; et al. Identification of COUP-TFII orphan nuclear receptor as a retinoic acid-activated receptor. PLoS Biol. 2008, 6, 2002–2015. [Google Scholar] [CrossRef] [PubMed]
- Schaeffer, E.; Guillou, F.; Part, D.; Zakin, M.M. A different combination of transcription factors modulates the expression of the human transferrin promoter in liver and Sertoli cells. J. Biol. Chem. 1993, 268, 23399–23408. [Google Scholar] [CrossRef]
- Stroup, D.; Chiang, J.Y.L. HNF4 and COUP-TFII interact to modulate transcription of the cholesterol 7α-hydroxylase gene (CYP7A1). J. Lipid Res. 2000, 41, 1–11. [Google Scholar] [CrossRef]
- Pineda Torra, I.; Jamshidi, Y.; Flavell, D.M.; Fruchart, J.-C.; Staels, B. Characterization of the Human PPARα Promoter: Identification of a Functional Nuclear Receptor Response Element. Mol. Endocrinol. 2002, 16, 1013–1028. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nault, R.; Fader, K.A.; Kopec, A.K.; Harkema, J.R.; Zacharewski, T.R.; Luyendyk, J.P. Coagulation-driven hepatic fibrosis requires protease activated receptor-1 (PAR-1) in a mouse model of TCDD-elicited steatohepatitis. Toxicol. Sci. 2016, 154, 381–391. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duncan, S.A.; Manova, K.; Chen, W.S.; Hoodless, P.; Weinstein, D.C.; Bachvarova, R.F.; Darnell, J.E. Expression of transcription factor HNF-4 in the extraembryonic endoderm, gut, and nephrogenic tissue of the developing mouse embryo: HNF-4 is a marker for primary endoderm in the implanting blastocyst. Proc. Natl. Acad. Sci. USA 1994, 91, 7598–7602. [Google Scholar] [CrossRef] [Green Version]
- Hayhurst, G.P.; Lee, Y.-H.; Lambert, G.; Ward, J.M.; Gonzalez, F.J. Hepatocyte Nuclear Factor 4α (Nuclear Receptor 2A1) Is Essential for Maintenance of Hepatic Gene Expression and Lipid Homeostasis. Mol. Cell. Biol. 2001, 21, 1393–1403. [Google Scholar] [CrossRef] [Green Version]
- Morimoto, A.; Kannari, M.; Tsuchida, Y.; Sasaki, S.; Saito, C.; Matsuta, T.; Maeda, T.; Akiyama, M.; Nakamura, T.; Sakaguchi, M.; et al. An HNF4α–microRNA-194/192 signaling axis maintains hepatic cell function. J. Biol. Chem. 2017, 292, 10574–10585. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.; Chen, J.; Ko, C.I.; Fan, Y.; Carreira, V.; Chen, Y.; Xia, Y.; Medvedovic, M.; Puga, A. Disruption of aryl hydrocarbon receptor homeostatic levels during embryonic stem cell differentiation alters expression of homeobox transcription factors that control cardiomyogenesis. Environ. Health Perspect. 2013, 121, 1334–1343. [Google Scholar] [CrossRef] [Green Version]
- Boitano, A.E.; Wang, J.; Romeo, R.; Bouchez, L.C.; Parker, A.E.; Sutton, S.E.; Walker, J.R.; Flaveny, C.A.; Perdew, G.H.; Denison, M.S.; et al. Aryl Hydrocarbon Receptor Antagonists Promote the Expansion of Human Hematopoietic Stem Cells. Science 2010, 329, 1345–1348. [Google Scholar] [CrossRef] [Green Version]
- Singh, K.P.; Garrett, R.W.; Casado, F.L.; Gasiewicz, T.A. Aryl hydrocarbon receptor-null allele mice have hematopoietic stem/progenitor cells with abnormal characteristics and functions. Stem Cells Dev. 2011, 20, 769–784. [Google Scholar] [CrossRef] [Green Version]
- Gazit, R.; Garrison, B.S.; Rao, T.N.; Shay, T.; Costello, J.; Ericson, J.; Kim, F.; Collins, J.J.; Regev, A.; Wagers, A.J.; et al. Transcriptome analysis identifies regulators of hematopoietic stem and progenitor cells. Stem Cell Rep. 2013, 1, 266–280. [Google Scholar] [CrossRef] [Green Version]
- Metidji, A.; Omenetti, S.; Crotta, S.; Li, Y.; Nye, E.; Ross, E.; Li, V.; Maradana, M.R.; Schiering, C.; Stockinger, B. The Environmental Sensor AHR Protects from Inflammatory Damage by Maintaining Intestinal Stem Cell Homeostasis and Barrier Integrity. Immunity 2018, 49, 353–362.e5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sato, T.; Clevers, H. Growing self-organizing mini-guts from a single intestinal stem cell: Mechanism and applications. Science 2013, 340, 1190–1194. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schneider, A.J.; Branam, A.M.; Peterson, R.E. Intersection of AHR and Wnt signaling in development, health, and disease. Int. J. Mol. Sci. 2014, 15, 17852–17885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Percie Du Sert, N.; Hurst, V.; Ahluwalia, A.; Alam, S.; Avey, M.T.; Baker, M.; Browne, W.J.; Clark, A.; Cuthill, I.C.; Dirnagl, U.; et al. The ARRIVE guidelines 2.0: Updated guidelines for reporting animal research. BMC Vet. Res. 2020, 16, 242. [Google Scholar] [CrossRef]
- De Matos Simoes, R.; Emmert-Streib, F. Bagging statistical network inference from large-scale gene expression data. PLoS ONE 2012, 7, e33624. [Google Scholar] [CrossRef] [Green Version]
- Heinz, S.; Benner, C.; Spann, N.; Bertolino, E.; Lin, Y.C.; Laslo, P.; Cheng, J.X.; Murre, C.; Singh, H.; Glass, C.K. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 2010, 38, 576–589. [Google Scholar] [CrossRef] [Green Version]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [Green Version]







| TermID | DEGs | All Genes in Term | Adj. p-Value |
|---|---|---|---|
| TZ_DIURNAL_GENES | 1222 | 5613 | 0 |
| TZ_SCS_PORTALHEP | 326 | 581 | 6.91 × 10−269 |
| TZ_SCS_MIDCENTRALHEP | 292 | 563 | 1.76 × 10−226 |
| TZ_SCS_CENTRALHEP | 203 | 466 | 3.59 × 10−136 |
| GO_CC_MM_MITOCHONDRION | 308 | 1328 | 6.80 × 10−120 |
| KEGG_MM_METABOLIC_PATHWAYS | 290 | 1180 | 1.59 × 10−119 |
| HEPG2_SECRETOME | 216 | 757 | 1.53 × 10−101 |
| PHH_SECRETOME | 211 | 726 | 7.27 × 10−101 |
| TZ_LIVER | 103 | 181 | 3.77 × 10−83 |
| GO_CC_MM_CYTOSOL | 210 | 927 | 3.02 × 10−78 |
| TZ_SCS_MIDPORTALHEP | 81 | 149 | 8.88 × 10−63 |
| GO_MF_MM_ATP_BINDING | 240 | 1454 | 3.22 × 10−61 |
| TZ_MALE_ENRICHED | 113 | 367 | 1.08 × 10−55 |
| GO_BP_MM_METABOLIC_PROCESS | 131 | 521 | 2.05 × 10−53 |
| GO_MF_MM_METAL_ION_BINDING | 168 | 871 | 1.64 × 10−51 |
| TZ_SCS_MACROPHAGES | 171 | 904 | 2.96 × 10−51 |
| GO_MF_MM_PROTEIN_HOMODIMERIZATION_ACTIVITY | 125 | 509 | 9.79 × 10−50 |
| TZ_SCS_ENDOTHELIALCELLS | 119 | 461 | 9.79 × 10−50 |
| GO_BP_MM_OXIDATION-REDUCTION_PROCESS | 128 | 535 | 1.25 × 10−49 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cholico, G.N.; Nault, R.; Zacharewski, T.R. Genome-Wide ChIPseq Analysis of AhR, COUP-TF, and HNF4 Enrichment in TCDD-Treated Mouse Liver. Int. J. Mol. Sci. 2022, 23, 1558. https://doi.org/10.3390/ijms23031558
Cholico GN, Nault R, Zacharewski TR. Genome-Wide ChIPseq Analysis of AhR, COUP-TF, and HNF4 Enrichment in TCDD-Treated Mouse Liver. International Journal of Molecular Sciences. 2022; 23(3):1558. https://doi.org/10.3390/ijms23031558
Chicago/Turabian StyleCholico, Giovan N., Rance Nault, and Tim R. Zacharewski. 2022. "Genome-Wide ChIPseq Analysis of AhR, COUP-TF, and HNF4 Enrichment in TCDD-Treated Mouse Liver" International Journal of Molecular Sciences 23, no. 3: 1558. https://doi.org/10.3390/ijms23031558






