Protein Lipidation Types: Current Strategies for Enrichment and Characterization
Abstract
1. Introduction
2. Types of Protein Lipidation
2.1. Fatty Acylation
2.1.1. S-palmitoylation
2.1.2. N-palmitoylation
2.1.3. O-palmitoylation
2.1.4. N-myristoylation
2.1.5. Acylation of Other Saturated Fatty Acids
2.1.6. Acylation of Unsaturated Fatty Acids
2.2. N-lipoylation
2.3. S-prenylation
2.4. C-terminal Phosphatidylethanolaminylation
2.5. C-terminal Cholesterolyation
2.6. C-terminal GPI Anchoring
2.7. LDE Acylation
3. Detection of Protein Lipidation
3.1. Qualitative Methods
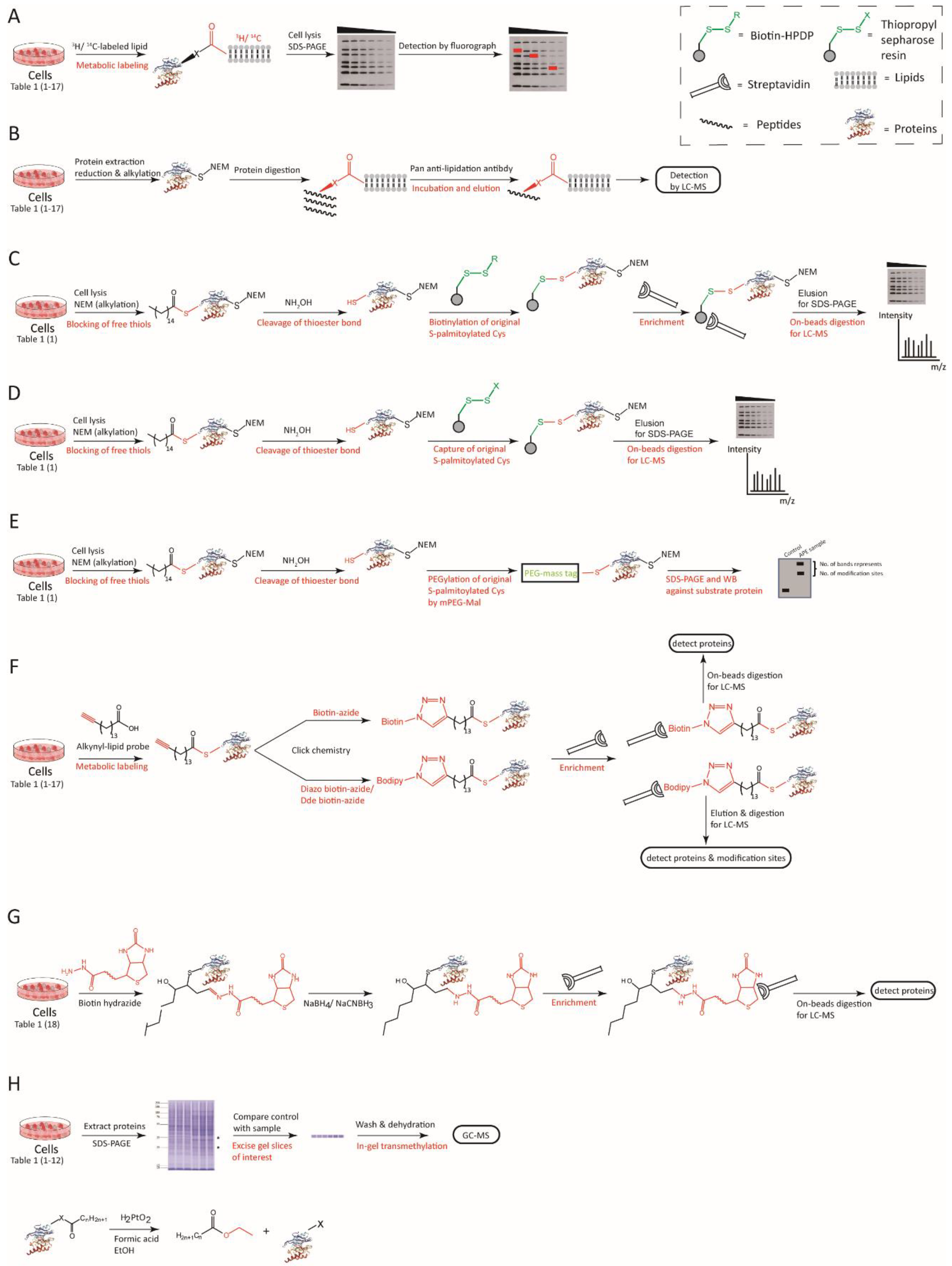
3.1.1. Radioactive Isotope-Labeling
3.1.2. Antibody Affinity Enrichment
3.1.3. Acyl-Biotin Exchange (ABE)
3.1.4. Click Chemistry
3.1.5. Biotin Hydrazide Affinity Capture
3.1.6. Lipid Esterification
3.1.7. Bioinformatics Tools
3.2. Quantitative Proteomics Methods
3.2.1. Stable Isotope Labeling with Amino Acids in Cell Culture (SILAC)
3.2.2. In Vitro Isotope Labeling
3.3. Dynamic Visualization Methods
4. Detection of PUFA-Modified Proteins
4.1. Difficulties in Detecting PUFA-Modified Proteins
4.2. Limitations in Current Methodology
4.3. Potential Solutions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| 14-PGJ2 | 15-Deoxy-D12, 14-prostaglandin J2 |
| 17-ODYA | 17-Octadecynoic acid |
| 2-HD | 2-trans-Hexadecenal |
| 4-HNE | 4-Hydroxy-2-nonenal |
| 4-ONE | 4-Oxo-2-nonenal |
| ABE | Acyl-biotin exchange |
| acyl-RAC | Acyl-resin–assisted capture |
| AKAP12 | A-kinase anchoring protein 12 |
| alk-LA | Alkynyl-linoleic acid |
| alk-LDEs | Alkynyl analogs of LDE |
| alk-PA | Alkynyl-palmitic acid |
| APE | Acyl-PEG exchange |
| APEGS | Acyl-pegyl exchange gel shift |
| APT | Acyl protein thioesterase |
| AQP0/MIP | Major intrinsic protein of lens fiber |
| BAX | BCL2-associated X, apoptosis regulator |
| BCKDHA | Branched-chain keto acid dehydrogenase E1 subunit alpha |
| Biotin-HPDP | Biotin-N-[6-(biotinamido)hexyl]-3′-(2′-pyridyldithio) propionamide |
| CD | Circular dichroism |
| Cysteine-SILAC | Cysteine-stable isotope labeling |
| Dde | 1-(4,4-Dimethyl-2,6-dioxocyclohex-1-ylidene)ethyl |
| EP300 | E1A-binding protein p300 |
| EThcD | Electron-transfer/higher-energy collision dissociation |
| FAS | Fas cell surface death receptor |
| FASN | Fatty acid synthase |
| FKBP4 | Peptidyl-prolyl cis-trans isomerase FKBP4 |
| FTase | Farnesyltransferase |
| FXR2 | FMR1 autosomal homolog 2 |
| GC-MS | Gas chromatography–mass spectrometry |
| GCV | Glycine cleavage system |
| GGTase-I | Geranylgeranyltransferase type I |
| GGTase-II | Geranylgeranyltransferase type II |
| GOAT | Ghrelin O-acyltransferase |
| GPI | Glycosylphosphatidylinositol |
| HH | Hedgehog family |
| HK1S | Hexokinase 1 variant in mammalian spermatozoa |
| HRAS | HRas proto-oncogene, GTPase |
| IA-alk | Iodoacetamide-alkyne |
| iFAT-MS | Isotope-coded fatty acid transmethylation–mass spectrometry |
| IMS | Ion-mobility spectrometry |
| iodoTMT | Iodoacetyl isobaric tandem mass tag |
| IP | Immunoprecipitation |
| isoTOP-ABPP | Isotopic tandem orthogonal proteolysis–activity-based protein profiling |
| ITPR1 | Inositol 1,4,5-triphosphate receptor type I |
| iTRAQ/TMT | Isobaric tag for relative and absolute quantitation/isobaric tandem mass tag |
| JAM3 | Junctional adhesion molecule 3 |
| KDH | α-Ketoglutarate |
| Khib | Lysine 2-hydroxyisobutyrylation |
| KRAS | KRas proto-oncogene, GTPase |
| LC3 | Microtubule-Associated Protein 1 Light Chain 3 Alpha |
| LC-MS | Liquid Chromatography-Mass Spectrometry |
| LDE | Lipid-Derived Electrophile |
| LPCAT1 | Lysophosphatidylcholine Acyltransferase 1 |
| MDA | Malonaldehyde |
| MS | Mass Spectrometry |
| NAT | N-terminal acetyltransferase |
| NEM | N-ethylmaleimide |
| NH2OH | Hydroxylamine |
| NMR | Nuclear Magnetic Resonance |
| NMT | N-myristoyl transferase |
| NRAS | NRas proto-oncogene, GTPase |
| PAT | Palmitoyl acyltransferase |
| PC | Photocleavable |
| PDH | Pyruvate dehydrogenase |
| PE | Phosphatidylethanolamine |
| PI-PLC | Phosphatidylinositol-specific phospholipase C |
| PLA | Proximity ligation assay |
| PRDX6 | Peroxiredoxin-6 |
| PRKAB1 | Protein kinase AMP-activated non-catalytic subunit beta 1 |
| PTM | Post-translational modification |
| PUFA | Polyunsaturated fatty acid |
| SFA | Saturated fatty acid |
| SILAC | Stable isotope labeling with amino acids in cell culture |
| SILAM | Stable isotope labeling of mammals |
| SIRT | Sirtuin |
| SNR | Signal-to-noise ratio |
| SRC | SRC proto-oncogene, non-receptor tyrosine kinase |
| VIM | Vimentin |
| WB | Western blotting |
References
- Mann, M.; Jensen, O.N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 2003, 21, 255–261. [Google Scholar] [CrossRef]
- Keenan, E.K.; Zachman, D.K.; Hirschey, M.D. Discovering the landscape of protein modifications. Mol. Cell 2021, 81, 1868–1878. [Google Scholar] [CrossRef]
- Zavialova, M.G.; Zgoda, V.G.; Nikolaev, E.N. Analysis of contribution of protein phosphorylation in the development of the diseases. Biomeditsinskaia Khimiia 2017, 63, 101–114. [Google Scholar] [CrossRef][Green Version]
- Cifani, P.; Kentsis, A. Towards comprehensive and quantitative proteomics for diagnosis and therapy of human disease. Proteomics 2017, 17, 1600079. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Zhang, Y.; Ren, J. Acetylation in cardiovascular diseases: Molecular mechanisms and clinical implications. Biochim. et Biophys. Acta (BBA)—Mol. Basis Dis. 2020, 1866, 165836. [Google Scholar] [CrossRef] [PubMed]
- Torres, M.P.; Dewhurst, H.; Sundararaman, N. Proteome-wide Structural Analysis of PTM Hotspots Reveals Regulatory Elements Predicted to Impact Biological Function and Disease. Mol. Cell. Proteom. 2016, 15, 3513–3528. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Sun, Y.; Niu, J.; Jarugumilli, G.K.; Wu, X. Protein Lipidation in Cell Signaling and Diseases: Function, Regulation, and Therapeutic Opportunities. Cell Chem. Biol. 2018, 25, 817–831. [Google Scholar] [CrossRef]
- Seo, J.; Lee, K.J. Post-translational modifications and their biological functions: Proteomic analysis and systematic approaches. J. Biochem. Mol. Biol. 2004, 37, 35–44. [Google Scholar] [CrossRef]
- Fukata, Y.; Fukata, M. Protein palmitoylation in neuronal development and synaptic plasticity. Nat. Rev. Neurosci. 2010, 11, 161–175. [Google Scholar] [CrossRef]
- Greaves, J.; Chamberlain, L.H. New links between S-acylation and cancer. J. Pathol. 2014, 233, 4–6. [Google Scholar] [CrossRef]
- Yeste-Velasco, M.; Linder, M.E.; Lu, Y.J. Protein S-palmitoylation and cancer. Biochim. Biophys. Acta Rev. Cancer 2015, 1856, 107–120. [Google Scholar] [CrossRef] [PubMed]
- Chavda, B.; Arnott, J.A.; Planey, S.L. Targeting protein palmitoylation: Selective inhibitors and implications in disease. Expert Opin. Drug Discov. 2014, 9, 1005–1019. [Google Scholar] [CrossRef] [PubMed]
- Roth, A.F.; Wan, J.; Bailey, A.O.; Sun, B.; Kuchar, J.A.; Green, W.N.; Phinney, B.S.; Yates, J.R., 3rd; Davis, N.G. Global analysis of protein palmitoylation in yeast. Cell 2006, 125, 1003–1013. [Google Scholar] [CrossRef] [PubMed]
- Peng, T.; Thinon, E.; Hang, H.C. Proteomic analysis of fatty-acylated proteins. Curr. Opin. Chem. Biol. 2016, 30, 77–86. [Google Scholar] [CrossRef]
- Hannoush, R.N.; Sun, J. The chemical toolbox for monitoring protein fatty acylation and prenylation. Nat. Chem. Biol. 2010, 6, 498–506. [Google Scholar] [CrossRef]
- Mueller, T.M.; Meador-Woodruff, J.H. Post-translational protein modifications in schizophrenia. NPJ Schizophr. 2020, 6, 5. [Google Scholar] [CrossRef]
- Hong, M.; Zhang, Y.; Hu, F. Membrane protein structure and dynamics from NMR spectroscopy. Annu. Rev. Phys. Chem. 2012, 63, 1–24. [Google Scholar] [CrossRef]
- Miles, A.J.; Wallace, B.A. Circular dichroism spectroscopy of membrane proteins. Chem. Soc. Rev. 2016, 45, 4859–4872. [Google Scholar] [CrossRef]
- Hanashima, S.; Nakane, T.; Mizohata, E. Heavy Atom Detergent/Lipid Combined X-ray Crystallography for Elucidating the Structure-Function Relationships of Membrane Proteins. Membranes 2021, 11, 823. [Google Scholar] [CrossRef]
- Clabbers, M.T.B.; Xu, H. Macromolecular crystallography using microcrystal electron diffraction. Acta Crystallogr. D Struct. Biol. 2021, 77, 313–324. [Google Scholar] [CrossRef]
- Aldini, G.; Domingues, M.R.; Spickett, C.M.; Domingues, P.; Altomare, A.; Sanchez-Gomez, F.J.; Oeste, C.L.; Perez-Sala, D. Protein lipoxidation: Detection strategies and challenges. Redox Biol. 2015, 5, 253–266. [Google Scholar] [CrossRef]
- Linder, M.E.; Deschenes, R.J. Palmitoylation: Policing protein stability and traffic. Nat. Rev. Mol. Cell Biol. 2007, 8, 74–84. [Google Scholar] [CrossRef] [PubMed]
- Resh, M.D. Trafficking and signaling by fatty-acylated and prenylated proteins. Nat. Chem. Biol. 2006, 2, 584–590. [Google Scholar] [CrossRef]
- Braun, P.E.; Radin, N.S. Interactions of lipids with a membrane structural protein from myelin. Biochemistry 1969, 8, 4310–4318. [Google Scholar] [CrossRef]
- Schmidt, M.F.; Schlesinger, M.J. Fatty acid binding to vesicular stomatitis virus glycoprotein: A new type of post-translational modification of the viral glycoprotein. Cell 1979, 17, 813–819. [Google Scholar] [CrossRef]
- Schlesinger, M.J.; Magee, A.I.; Schmidt, M.F. Fatty acid acylation of proteins in cultured cells. J. Biol. Chem. 1980, 255, 10021–10024. [Google Scholar] [CrossRef]
- Stoffyn, P.; Folch-Pi, J. On the type of linkage binding fatty acids present in brain white matter proteolipid apoprotein. Biochem. Biophys. Res. Commun. 1971, 44, 157–161. [Google Scholar] [CrossRef]
- Chen, B.; Zheng, B.; DeRan, M.; Jarugumilli, G.K.; Fu, J.; Brooks, Y.S.; Wu, X. ZDHHC7-mediated S-palmitoylation of Scribble regulates cell polarity. Nat. Chem. Biol. 2016, 12, 686–693. [Google Scholar] [CrossRef]
- Hernandez, J.L.; Davda, D.; Cheung See Kit, M.; Majmudar, J.D.; Won, S.J.; Gang, M.; Pasupuleti, S.C.; Choi, A.I.; Bartkowiak, C.M.; Martin, B.R. APT2 Inhibition Restores Scribble Localization and S-Palmitoylation in Snail-Transformed Cells. Cell Chem. Biol. 2017, 24, 87–97. [Google Scholar] [CrossRef]
- Chan, P.; Han, X.; Zheng, B.; DeRan, M.; Yu, J.; Jarugumilli, G.K.; Deng, H.; Pan, D.; Luo, X.; Wu, X. Autopalmitoylation of TEAD proteins regulates transcriptional output of the Hippo pathway. Nat. Chem. Biol. 2016, 12, 282–289. [Google Scholar] [CrossRef]
- Lanyon-Hogg, T.; Faronato, M.; Serwa, R.A.; Tate, E.W. Dynamic Protein Acylation: New Substrates, Mechanisms, and Drug Targets. Trends Biochem. Sci. 2017, 42, 566–581. [Google Scholar] [CrossRef]
- Blanc, M.; David, F.; Abrami, L.; Migliozzi, D.; Armand, F.; Burgi, J.; van der Goot, F.G. SwissPalm: Protein Palmitoylation database. F1000Research 2015, 4, 261. [Google Scholar] [CrossRef] [PubMed]
- Blanc, M.; David, F.P.A.; van der Goot, F.G. SwissPalm 2: Protein S-Palmitoylation Database. Methods Mol. Biol. 2019, 2009, 203–214. [Google Scholar] [PubMed]
- Dowal, L.; Yang, W.; Freeman, M.R.; Steen, H.; Flaumenhaft, R. Proteomic analysis of palmitoylated platelet proteins. Blood 2011, 118, e62–e73. [Google Scholar] [CrossRef]
- Kang, R.; Wan, J.; Arstikaitis, P.; Takahashi, H.; Huang, K.; Bailey, A.O.; Thompson, J.X.; Roth, A.F.; Drisdel, R.C.; Mastro, R.; et al. Neural palmitoyl-proteomics reveals dynamic synaptic palmitoylation. Nature 2008, 456, 904–909. [Google Scholar] [CrossRef] [PubMed]
- Martin, B.R.; Wang, C.; Adibekian, A.; Tully, S.E.; Cravatt, B.F. Global profiling of dynamic protein palmitoylation. Nat Methods 2011, 9, 84–89. [Google Scholar] [CrossRef]
- Rocks, O.; Gerauer, M.; Vartak, N.; Koch, S.; Huang, Z.P.; Pechlivanis, M.; Kuhlmann, J.; Brunsveld, L.; Chandra, A.; Ellinger, B.; et al. The Palmitoylation Machinery Is a Spatially Organizing System for Peripheral Membrane Proteins. Cell 2010, 141, 458–471. [Google Scholar] [CrossRef] [PubMed]
- Adachi, N.; Hess, D.T.; McLaughlin, P.; Stamler, J.S. S-Palmitoylation of a Novel Site in the beta2-Adrenergic Receptor Associated with a Novel Intracellular Itinerary. J. Biol. Chem. 2016, 291, 20232–20246. [Google Scholar] [CrossRef]
- Rossin, A.; Durivault, J.; Chakhtoura-Feghali, T.; Lounnas, N.; Gagnoux-Palacios, L.; Hueber, A.O. Fas palmitoylation by the palmitoyl acyltransferase DHHC7 regulates Fas stability. Cell Death Differ. 2015, 22, 643–653. [Google Scholar] [CrossRef]
- Frohlich, M.; Dejanovic, B.; Kashkar, H.; Schwarz, G.; Nussberger, S. S-palmitoylation represents a novel mechanism regulating the mitochondrial targeting of BAX and initiation of apoptosis. Cell Death Dis. 2014, 5, e1057. [Google Scholar] [CrossRef]
- Fredericks, G.J.; Hoffmann, F.W.; Rose, A.H.; Osterheld, H.J.; Hess, F.M.; Mercier, F.; Hoffmann, P.R. Stable expression and function of the inositol 1,4,5-triphosphate receptor requires palmitoylation by a DHHC6/selenoprotein K complex. Proc. Natl. Acad. Sci. USA 2014, 111, 16478–16483. [Google Scholar] [CrossRef] [PubMed]
- Aramsangtienchai, P.; Spiegelman, N.A.; Cao, J.; Lin, H.N. S-Palmitoylation of Junctional Adhesion Molecule C Regulates Its Tight Junction Localization and Cell Migration. J. Biol. Chem. 2017, 292, 5325–5334. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Smith, D.A.; Memarzadeh, S.; Lowell, C.A.; Cooper, J.A.; Witte, O.N. Differential transformation capacity of Src family kinases during the initiation of prostate cancer. Proc. Natl. Acad. Sci. USA 2011, 108, 6579–6584. [Google Scholar] [CrossRef] [PubMed]
- Varland, S.; Osberg, C.; Arnesen, T. N-terminal modifications of cellular proteins: The enzymes involved, their substrate specificities and biological effects. Proteomics 2015, 15, 2385–2401. [Google Scholar] [CrossRef] [PubMed]
- Madsen, A.S.; Andersen, C.; Daoud, M.; Anderson, K.A.; Laursen, J.S.; Chakladar, S.; Huynh, F.K.; Colaco, A.R.; Backos, D.S.; Fristrup, P.; et al. Investigating the Sensitivity of NAD+-dependent Sirtuin Deacylation Activities to NADH. J. Biol. Chem. 2016, 291, 7128–7141. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Spiegelman, N.A.; Nelson, O.D.; Jing, H.; Lin, H. SIRT6 regulates Ras-related protein R-Ras2 by lysine defatty-acylation. eLife 2017, 6, 39. [Google Scholar] [CrossRef]
- Zou, C.; Ellis, B.M.; Smith, R.M.; Chen, B.B.; Zhao, Y.; Mallampalli, R.K. Acyl-CoA:lysophosphatidylcholine acyltransferase I (Lpcat1) catalyzes histone protein O-palmitoylation to regulate mRNA synthesis. J. Biol. Chem. 2011, 286, 28019–28025. [Google Scholar] [CrossRef]
- Branton, W.D.; Rudnick, M.S.; Zhou, Y.; Eccleston, E.D.; Fields, G.B.; Bowers, L.D. Fatty acylated toxin structure. Nature 1993, 365, 496–497. [Google Scholar] [CrossRef]
- Maurer-Stroh, S.; Eisenhaber, B.; Eisenhaber, F. N-terminal N-myristoylation of proteins: Prediction of substrate proteins from amino acid sequence. J. Mol. Biol. 2002, 317, 541–557. [Google Scholar] [CrossRef]
- Liu, Z.; Yang, T.; Li, X.; Peng, T.; Hang, H.C.; Li, X.D. Integrative chemical biology approaches for identification and characterization of “erasers” for fatty-acid-acylated lysine residues within proteins. Angew. Chem. Int. Ed. Engl. 2015, 54, 1149–1152. [Google Scholar] [CrossRef]
- Teng, Y.B.; Jing, H.; Aramsangtienchai, P.; He, B.; Khan, S.; Hu, J.; Lin, H.; Hao, Q. Efficient demyristoylase activity of SIRT2 revealed by kinetic and structural studies. Sci. Rep. 2015, 5, 8529. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, C.; Weinert, B.T.; Nishida, Y.; Verdin, E.; Mann, M. The growing landscape of lysine acetylation links metabolism and cell signalling. Nat. Rev. Mol. Cell Biol. 2014, 15, 536–550. [Google Scholar] [CrossRef] [PubMed]
- Brett, K.; Kordyukova, L.V.; Serebryakova, M.V.; Mintaev, R.R.; Alexeevski, A.V.; Veit, M. Site-specific S-acylation of influenza virus hemagglutinin: The location of the acylation site relative to the membrane border is the decisive factor for attachment of stearate. J. Biol. Chem. 2014, 289, 34978–34989. [Google Scholar] [CrossRef] [PubMed]
- Kordyukova, L.V.; Serebryakova, M.V.; Baratova, L.A.; Veit, M. S acylation of the hemagglutinin of influenza viruses: Mass spectrometry reveals site-specific attachment of stearic acid to a transmembrane cysteine. J. Virol. 2008, 82, 9288–9292. [Google Scholar] [CrossRef]
- Gutierrez, J.A.; Solenberg, P.J.; Perkins, D.R.; Willency, J.A.; Knierman, M.D.; Jin, Z.; Witcher, D.R.; Luo, S.; Onyia, J.E.; Hale, J.E. Ghrelin octanoylation mediated by an orphan lipid transferase. Proc. Natl. Acad. Sci. USA 2008, 105, 6320–6325. [Google Scholar] [CrossRef]
- Yang, J.; Brown, M.S.; Liang, G.; Grishin, N.V.; Goldstein, J.L. Identification of the acyltransferase that octanoylates ghrelin, an appetite-stimulating peptide hormone. Cell 2008, 132, 387–396. [Google Scholar] [CrossRef]
- Takada, R.; Satomi, Y.; Kurata, T.; Ueno, N.; Norioka, S.; Kondoh, H.; Takao, T.; Takada, S. Monounsaturated fatty acid modification of Wnt protein: Its role in Wnt secretion. Dev. Cell 2006, 11, 791–801. [Google Scholar] [CrossRef]
- Zhai, L.; Chaturvedi, D.; Cumberledge, S. Drosophila wnt-1 undergoes a hydrophobic modification and is targeted to lipid rafts, a process that requires porcupine. J. Biol. Chem. 2004, 279, 33220–33227. [Google Scholar] [CrossRef]
- Kakugawa, S.; Langton, P.F.; Zebisch, M.; Howell, S.; Chang, T.H.; Liu, Y.; Feizi, T.; Bineva, G.; O—Reilly, N.; Snijders, A.P.; et al. Notum deacylates Wnt proteins to suppress signalling activity. Nature 2015, 519, 187–192. [Google Scholar] [CrossRef]
- Schey, K.L.; Gutierrez, D.B.; Wang, Z.; Wei, J.; Grey, A.C. Novel fatty acid acylation of lens integral membrane protein aquaporin-0. Biochemistry 2010, 49, 9858–9865. [Google Scholar] [CrossRef]
- Muszbek, L.; Laposata, M. Covalent modification of proteins by arachidonate and eicosapentaenoate in platelets. J. Biol. Chem. 1993, 268, 18243–18248. [Google Scholar] [CrossRef]
- Mathias, R.A.; Greco, T.M.; Oberstein, A.; Budayeva, H.G.; Chakrabarti, R.; Rowland, E.A.; Kang, Y.B.; Shenk, T.; Cristea, I.M. Sirtuin 4 Is a Lipoamidase Regulating Pyruvate Dehydrogenase Complex Activity. Cell 2014, 159, 1615–1625. [Google Scholar] [CrossRef] [PubMed]
- Rowland, E.A.; Greco, T.M.; Snowden, C.K.; McCabe, A.L.; Silhavy, T.J.; Cristea, I.M. Sirtuin Lipoamidase Activity Is Conserved in Bacteria as a Regulator of Metabolic Enzyme Complexes. Mbio 2017, 8, e01096-17. [Google Scholar] [CrossRef] [PubMed]
- Moores, S.L.; Schaber, M.D.; Mosser, S.D.; Rands, E.; Ohara, M.B.; Garsky, V.M.; Marshall, M.S.; Pompliano, D.L.; Gibbs, J.B. Sequence Dependence of Protein Isoprenylation. J. Biol. Chem. 1991, 266, 14603–14610. [Google Scholar] [CrossRef]
- Zhang, F.L.; Casey, P.J. Protein prenylation: Molecular mechanisms and functional consequences. Annu. Rev. Biochem. 1996, 65, 241–269. [Google Scholar] [CrossRef]
- Nakatogawa, H.; Ichimura, Y.; Ohsumi, Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 2007, 130, 165–178. [Google Scholar] [CrossRef]
- Ray, A.; Jatana, N.; Thukral, L. Lipidated proteins: Spotlight on protein-membrane binding interfaces. Prog. Biophys. Mol. Biol. 2017, 128, 74–84. [Google Scholar] [CrossRef]
- Chen, M.H.; Li, Y.J.; Kawakami, T.; Xu, S.M.; Chuang, P.T. Palmitoylation is required for the production of a soluble multimeric Hedgehog protein complex and long-range signaling in vertebrates. Genes Dev. 2004, 18, 641–659. [Google Scholar] [CrossRef]
- Goetz, J.A.; Singh, S.; Suber, L.M.; Kull, F.J.; Robbins, D.J. A highly conserved amino-terminal region of sonic hedgehog is required for the formation of its freely diffusible multimeric form. J. Biol. Chem. 2006, 281, 4087–4093. [Google Scholar] [CrossRef]
- Yu, S.C.; Guo, Z.W.; Johnson, C.; Gu, G.F.; Wu, Q.Y. Recent progress in synthetic and biological studies of GPI anchors and GPI-anchored proteins. Curr. Opin. Chem. Biol. 2013, 17, 1006–1013. [Google Scholar] [CrossRef]
- Masuishi, Y.; Nomura, A.; Okayama, A.; Kimura, Y.; Arakawa, N.; Hirano, H. Mass spectrometric identification of glycosylphosphatidylinositol-anchored peptides. J. Proteome Res. 2013, 12, 4617–4626. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Qin, W.; Wang, C. Chemoproteomic profiling of protein modifications by lipid-derived electrophiles. Curr. Opin. Chem. Biol. 2016, 30, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Nagahara, N.; Matsumura, T.; Okamoto, R.; Kajihara, Y. Protein cysteine modifications: (1) medical chemistry for proteomics. Curr. Med. Chem. 2009, 16, 4419–4444. [Google Scholar] [CrossRef] [PubMed]
- George, J.; Soares, C.; Montersino, A.; Beique, J.C.; Thomas, G.M. Palmitoylation of LIM Kinase-1 ensures spine-specific actin polymerization and morphological plasticity. eLife 2015, 4, e06327. [Google Scholar] [CrossRef] [PubMed]
- Pepinsky, R.B.; Zeng, C.; Wen, D.; Rayhorn, P.; Baker, D.P.; Williams, K.P.; Bixler, S.A.; Ambrose, C.M.; Garber, E.A.; Miatkowski, K.; et al. Identification of a palmitic acid-modified form of human Sonic hedgehog. J. Biol. Chem. 1998, 273, 14037–14045. [Google Scholar] [CrossRef]
- Buglino, J.A.; Resh, M.D. Hhat is a palmitoylacyltransferase with specificity for N-palmitoylation of Sonic Hedgehog. J. Biol. Chem. 2008, 283, 22076–22088. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Moriya, K.; Nagatoshi, K.; Ota, Y.; Ezure, T.; Ando, E.; Tsunasawa, S.; Utsumi, T. Strategy for comprehensive identification of human N-myristoylated proteins using an insect cell-free protein synthesis system. Proteomics 2010, 10, 1780–1793. [Google Scholar] [CrossRef]
- Patwardhan, P.; Resh, M.D. Myristoylation and membrane binding regulate c-Src stability and kinase activity. Mol. Cell Biol. 2010, 30, 4094–4107. [Google Scholar] [CrossRef]
- Thinon, E.; Serwa, R.A.; Broncel, M.; Brannigan, J.A.; Brassat, U.; Wright, M.H.; Heal, W.P.; Wilkinson, A.J.; Mann, D.J.; Tate, E.W. Global profiling of co- and post-translationally N-myristoylated proteomes in human cells. Nat. Commun. 2014, 5, 4919. [Google Scholar] [CrossRef]
- Liang, J.; Xu, Z.X.; Ding, Z.; Lu, Y.; Yu, Q.; Werle, K.D.; Zhou, G.; Park, Y.Y.; Peng, G.; Gambello, M.J.; et al. Myristoylation confers noncanonical AMPK functions in autophagy selectivity and mitochondrial surveillance. Nat. Commun. 2015, 6, 7926. [Google Scholar] [CrossRef]
- Stackpole, E.E.; Akins, M.R.; Fallon, J.R. N-myristoylation regulates the axonal distribution of the Fragile X-related protein FXR2P. Mol. Cell. Neurosci. 2014, 62, 42–50. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kumar, S.; Parameswaran, S.; Sharma, R.K. Novel myristoylation of the sperm-specific hexokinase 1 isoform regulates its atypical localization. Biol. Open 2015, 4, 1679–1687. [Google Scholar] [CrossRef][Green Version]
- Wright, M.H.; Heal, W.P.; Mann, D.J.; Tate, E.W. Protein myristoylation in health and disease. J. Chem. Biol. 2010, 3, 19–35. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Chen, W.; Cobb, M.H.; Zhao, Y.M. PTMap-A sequence alignment software for unrestricted, accurate, and full-spectrum identification of post-translational modification sites. Proc. Natl. Acad. Sci. USA 2009, 106, 761–766. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Sabari, B.R.; Garcia, B.A.; Allis, C.D.; Zhao, Y. SnapShot: Histone modifications. Cell 2014, 159, 458–458.e1. [Google Scholar] [CrossRef] [PubMed]
- DeMar, J.C., Jr.; Anderson, R.E. Identification and quantitation of the fatty acids composing the CoA ester pool of bovine retina, heart, and liver. J. Biol. Chem. 1997, 272, 31362–31368. [Google Scholar] [CrossRef]
- Liang, X.; Nazarian, A.; Erdjument-Bromage, H.; Bornmann, W.; Tempst, P.; Resh, M.D. Heterogeneous fatty acylation of Src family kinases with polyunsaturated fatty acids regulates raft localization and signal transduction. J. Biol. Chem. 2001, 276, 30987–30994. [Google Scholar] [CrossRef]
- Dizhoor, A.M.; Ericsson, L.H.; Johnson, R.S.; Kumar, S.; Olshevskaya, E.; Zozulya, S.; Neubert, T.A.; Stryer, L.; Hurley, J.B.; Walsh, K.A. The Nh2 Terminus of Retinal Recoverin Is Acylated by a Small Family of Fatty-Acids. J. Biol. Chem. 1992, 267, 16033–16036. [Google Scholar] [CrossRef]
- Kokame, K.; Fukada, Y.; Yoshizawa, T.; Takao, T.; Shimonishi, Y. Lipid Modification at the N-Terminus of Photoreceptor G-Protein Alpha-Subunit. Nature 1992, 359, 749–752. [Google Scholar] [CrossRef]
- Pereira-Leal, J.B.; Hume, A.N.; Seabra, M.C. Prenylation of Rab GTPases: Molecular mechanisms and involvement in genetic disease. FEBS Lett. 2001, 498, 197–200. [Google Scholar] [CrossRef]
- Rowland, E.A.; Snowden, C.K.; Cristea, I.M. Protein lipoylation: An evolutionarily conserved metabolic regulator of health and disease. Curr. Opin. Chem. Biol. 2018, 42, 76–85. [Google Scholar] [CrossRef] [PubMed]
- Reed, L.J. A trail of research from lipoic acid to alpha-keto acid dehydrogenase complexes. J. Biol. Chem. 2001, 276, 38329–38336. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Zhang, X.; Chen, X.; Aramsangtienchai, P.; Tong, Z.; Lin, H. Protein Lipidation: Occurrence, Mechanisms, Biological Functions, and Enabling Technologies. Chem. Rev. 2018, 118, 919–988. [Google Scholar] [CrossRef]
- Liu, M.; Sjogren, A.K.; Karlsson, C.; Ibrahim, M.X.; Andersson, K.M.; Olofsson, F.J.; Wahlstrom, A.M.; Dalin, M.; Yu, H.; Chen, Z.; et al. Targeting the protein prenyltransferases efficiently reduces tumor development in mice with K-RAS-induced lung cancer. Proc. Natl. Acad. Sci. USA 2010, 107, 6471–6476. [Google Scholar] [CrossRef]
- Sjogren, A.K.M.; Andersson, K.M.E.; Khan, O.; Olofsson, F.J.; Karlsson, C.; Bergo, M.O. Inactivating GGTase-I reduces disease phenotypes in a mouse model of K-RAS-induced myeloproliferative disease. Leukemia 2011, 25, 186–189. [Google Scholar] [CrossRef]
- Porter, J.A.; Ekker, S.C.; Park, W.J.; von Kessler, D.P.; Young, K.E.; Chen, C.H.; Ma, Y.; Woods, A.S.; Cotter, R.J.; Koonin, E.V.; et al. Hedgehog patterning activity: Role of a lipophilic modification mediated by the carboxy-terminal autoprocessing domain. Cell 1996, 86, 21–34. [Google Scholar] [CrossRef]
- Zeng, X.; Goetz, J.A.; Suber, L.M.; Scott, W.J.; Schreiner, C.M.; Robbins, D.J. A freely diffusible form of Sonic hedgehog mediates long-range signalling. Nature 2001, 411, 716–720. [Google Scholar] [CrossRef]
- Orlean, P.; Menon, A.K. Thematic review series: Lipid posttranslational modifications. GPI anchoring of protein in yeast and mammalian cells, or: How we learned to stop worrying and love glycophospholipids. J. Lipid Res. 2007, 48, 993–1011. [Google Scholar] [PubMed]
- Tsai, Y.H.; Liu, X.; Seeberger, P.H. Chemical biology of glycosylphosphatidylinositol anchors. Angew. Chem. Int. Ed. Engl. 2012, 51, 11438–11456. [Google Scholar] [CrossRef]
- Gamage, D.G.; Hendrickson, T.L. GPI transamidase and GPI anchored proteins: Oncogenes and biomarkers for cancer. Crit. Rev. Biochem. Mol. Biol. 2013, 48, 446–464. [Google Scholar] [CrossRef]
- Bautista, J.M.; Marin-Garcia, P.; Diez, A.; Azcarate, I.G.; Puyet, A. Malaria proteomics: Insights into the parasite-host interactions in the pathogenic space. J. Proteom. 2014, 97, 107–125. [Google Scholar] [CrossRef]
- Puig, B.; Altmeppen, H.; Glatzel, M. The GPI-anchoring of PrP: Implications in sorting and pathogenesis. Prion 2014, 8, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Weerapana, E.; Blewett, M.M.; Cravatt, B.F. A chemoproteomic platform to quantitatively map targets of lipid-derived electrophiles. Nat. Methods 2014, 11, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Boveris, A.; Navarro, A. Brain mitochondrial dysfunction in aging. IUBMB Life 2008, 60, 308–314. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Liu, Y.; Hou, X.; Ye, Z.; Wang, C. Quantitative and Site-Specific Chemoproteomic Profiling of Targets of Acrolein. Chem. Res. Toxicol. 2019, 32, 467–473. [Google Scholar] [CrossRef]
- Galligan, J.J.; Rose, K.L.; Beavers, W.N.; Hill, S.; Tallman, K.A.; Tansey, W.P.; Marnett, L.J. Stable histone adduction by 4-oxo-2-nonenal: A potential link between oxidative stress and epigenetics. J. Am. Chem. Soc. 2014, 136, 11864–11866. [Google Scholar] [CrossRef]
- Cui, Y.W.; Li, X.; Lin, J.W.; Hao, Q.; Li, X.D. Histone Ketoamide Adduction by 4-Oxo-2-nonenal Is a Reversible Posttranslational Modification Regulated by Sirt2. ACS Chem. Biol. 2017, 12, 47–51. [Google Scholar] [CrossRef]
- Bantscheff, M.; Lemeer, S.; Savitski, M.M.; Kuster, B. Quantitative mass spectrometry in proteomics: Critical review update from 2007 to the present. Anal. Bioanal. Chem. 2012, 404, 939–965. [Google Scholar] [CrossRef]
- Bantscheff, M.; Schirle, M.; Sweetman, G.; Rick, J.; Kuster, B. Quantitative mass spectrometry in proteomics: A critical review. Anal. Bioanal. Chem. 2007, 389, 1017–1031. [Google Scholar] [CrossRef]
- Held, J.M.; Gibson, B.W. Regulatory control or oxidative damage? Proteomic approaches to interrogate the role of cysteine oxidation status in biological processes. Mol. Cell Proteom. 2012, 11, R111.013037. [Google Scholar] [CrossRef]
- Hao, G.; Derakhshan, B.; Shi, L.; Campagne, F.; Gross, S.S. SNOSID, a proteomic method for identification of cysteine S-nitrosylation sites in complex protein mixtures. Proc. Natl. Acad. Sci. USA 2006, 103, 1012–1017. [Google Scholar] [CrossRef] [PubMed]
- Bolla, J.R.; Agasid, M.T.; Mehmood, S.; Robinson, C.V. Membrane Protein-Lipid Interactions Probed Using Mass Spectrometry. Annu. Rev. Biochem. 2019, 88, 85–111. [Google Scholar] [CrossRef] [PubMed]
- Urner, L.H.; Schulze, M.; Maier, Y.B.; Hoffmann, W.; Warnke, S.; Liko, I.; Folmert, K.; Manz, C.; Robinson, C.V.; Haag, R.; et al. A new azobenzene-based design strategy for detergents in membrane protein research. Chem. Sci. 2020, 11, 3538–3546. [Google Scholar] [CrossRef]
- Morgner, N.; Montenegro, F.; Barrera, N.P.; Robinson, C.V. Mass spectrometry—From peripheral proteins to membrane motors. J. Mol. Biol. 2012, 423, 1–13. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Giannakouros, T.; Armstrong, J.; Magee, A.I. Protein prenylation in Schizosaccharomyces pombe. FEBS Lett. 1992, 297, 103–106. [Google Scholar] [CrossRef]
- Goldstein, J.L.; Brown, M.S. Regulation of the mevalonate pathway. Nature 1990, 343, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Swarthout, J.T.; Lobo, S.; Farh, L.; Croke, M.R.; Greentree, W.K.; Deschenes, R.J.; Linder, M.E. DHHC9 and GCP16 constitute a human protein fatty acyltransferase with specificity for H- and N-Ras. J. Biol. Chem. 2005, 280, 31141–31148. [Google Scholar] [CrossRef]
- Schmidt, M.F.; Bracha, M.; Schlesinger, M.J. Evidence for covalent attachment of fatty acids to Sindbis virus glycoproteins. Proc. Natl. Acad. Sci. USA 1979, 76, 1687–1691. [Google Scholar] [CrossRef]
- Martin, D.D.O.; Vilas, G.L.; Prescher, J.A.; Rajaiah, G.; Falck, J.R.; Bertozzi, C.R.; Berthiaume, L.G. Rapid detection, discovery, and identification of post-translationally myristoylated proteins during apoptosis using a bio-orthogonal azidomyristate analog. FASEB J. 2008, 22, 797–806. [Google Scholar] [CrossRef]
- Fang, C.; Zhang, X.; Zhang, L.; Gao, X.; Yang, P.; Lu, H. Identification of Palmitoylated Transitional Endoplasmic Reticulum ATPase by Proteomic Technique and Pan Antipalmitoylation Antibody. J. Proteome Res. 2016, 15, 956–962. [Google Scholar] [CrossRef]
- Huang, H.; Tang, S.; Ji, M.; Tang, Z.Y.; Shimada, M.; Liu, X.J.; Qi, S.K.; Locasale, J.W.; Roeder, R.G.; Zhao, Y.M.; et al. EP300-Mediated Lysine 2-Hydroxyisobutyrylation Regulates Glycolysis. Mol. Cell 2018, 70, 663. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.M.; Peng, Z.; Chen, H.Y.; Pan, T.T.; Hu, X.N.; Wang, F.; Luo, T. Posttranslational lysine 2-hydroxyisobutyrylation of human sperm tail proteins affects motility. Hum. Reprod. 2020, 35, 494–503. [Google Scholar] [CrossRef] [PubMed]
- Ge, H.; Li, B.; Chen, W.; Xu, Q.; Chen, S.; Zhang, H.; Wu, J.; Zhen, Q.; Li, Y.; Yong, L.; et al. Differential occurrence of lysine 2-hydroxyisobutyrylation in psoriasis skin lesions. J. Proteom. 2019, 205, 103420. [Google Scholar] [CrossRef] [PubMed]
- Yin, D.; Jiang, N.; Zhang, Y.; Wang, D.; Sang, X.; Feng, Y.; Chen, R.; Wang, X.; Yang, N.; Chen, Q. Global Lysine Crotonylation and 2-Hydroxyisobutyrylation in Phenotypically Different Toxoplasma gondii Parasites. Mol. Cell Proteom. 2019, 18, 2207–2224. [Google Scholar] [CrossRef]
- Drisdel, R.C.; Green, W.N. Labeling and quantifying sites of protein palmitoylation. Biotechniques 2004, 36, 276–285. [Google Scholar] [CrossRef]
- Marin, E.P.; Derakhshan, B.; Lam, T.T.; Davalos, A.; Sessa, W.C. Endothelial cell palmitoylproteomic identifies novel lipid-modified targets and potential substrates for protein acyl transferases. Circ. Res. 2012, 110, 1336–1344. [Google Scholar] [CrossRef]
- Forrester, M.T.; Hess, D.T.; Thompson, J.W.; Hultman, R.; Moseley, M.A.; Stamler, J.S.; Casey, P.J. Site-specific analysis of protein S-acylation by resin-assisted capture. J. Lipid Res. 2011, 52, 393–398. [Google Scholar] [CrossRef]
- Percher, A.; Ramakrishnan, S.; Thinon, E.; Yuan, X.Q.; Yount, J.S.; Hang, H.C. Mass-tag labeling reveals site-specific and endogenous levels of protein S-fatty acylation. Proc. Natl. Acad. Sci. USA 2016, 113, 4302–4307. [Google Scholar] [CrossRef]
- Drisdel, R.C.; Alexander, J.K.; Sayeed, A.; Green, W.N. Assays of protein palmitoylation. Methods 2006, 40, 127–134. [Google Scholar] [CrossRef]
- Roth, A.F.; Wan, J.; Green, W.N.; Yates, J.R.; Davis, N.G. Proteomic identification of palmitoylated proteins. Methods 2006, 40, 135–142. [Google Scholar] [CrossRef]
- Wang, Q.; Chan, T.R.; Hilgraf, R.; Fokin, V.V.; Sharpless, K.B.; Finn, M.G. Bioconjugation by copper(I)-catalyzed azide-alkyne [3 + 2] cycloaddition. J. Am. Chem. Soc. 2003, 125, 3192–3193. [Google Scholar] [CrossRef] [PubMed]
- Hannoush, R.N.; Arenas-Ramirez, N. Imaging the Lipidome: Omega-Alkynyl Fatty Acids for Detection and Cellular Visualization of Lipid-Modified Proteins. ACS Chem. Biol. 2009, 4, 581–587. [Google Scholar] [CrossRef] [PubMed]
- Broncel, M.; Serwa, R.A.; Ciepla, P.; Krause, E.; Dallman, M.J.; Magee, A.I.; Tate, E.W. Multifunctional reagents for quantitative proteome-wide analysis of protein modification in human cells and dynamic profiling of protein lipidation during vertebrate development. Angew. Chem. Int. Ed. Engl. 2015, 54, 5948–5951. [Google Scholar] [CrossRef] [PubMed]
- Wright, M.H.; Clough, B.; Rackham, M.D.; Rangachari, K.; Brannigan, J.A.; Grainger, M.; Moss, D.K.; Bottrill, A.R.; Heal, W.P.; Broncel, M.; et al. Validation of N-myristoyltransferase as an antimalarial drug target using an integrated chemical biology approach. Nat. Chem. 2014, 6, 112–121. [Google Scholar] [CrossRef] [PubMed]
- Chesarino, N.M.; Hach, J.C.; Chen, J.L.; Zaro, B.W.; Rajaram, M.V.; Turner, J.; Schlesinger, L.S.; Pratt, M.R.; Hang, H.C.; Yount, J.S. Chemoproteomics reveals Toll-like receptor fatty acylation. BMC Biol. 2014, 12, 91. [Google Scholar] [CrossRef] [PubMed]
- Kostiuk, M.A.; Corvi, M.M.; Keller, B.O.; Plummer, G.; Prescher, J.A.; Hangauer, M.J.; Bertozzi, C.R.; Rajaiah, G.; Falck, J.R.; Berthiaume, L.G. Identification of palmitoylated mitochondrial proteins using a bio-orthogonal azido-palmitate analogue. FASEB J. 2008, 22, 721–732. [Google Scholar] [CrossRef]
- Vila, A.; Tallman, K.A.; Jacobs, A.T.; Liebler, D.C.; Porter, N.A.; Marnett, L.J. Identification of protein targets of 4-hydroxynonenal using click chemistry for ex vivo biotinylation of azido and alkynyl derivatives. Chem. Res. Toxicol. 2008, 21, 432–444. [Google Scholar] [CrossRef]
- Jarugumilli, G.K.; Choi, J.R.; Chan, P.; Yu, M.; Sun, Y.; Chen, B.; Niu, J.; DeRan, M.; Zheng, B.; Zoeller, R.; et al. Chemical Probe to Identify the Cellular Targets of the Reactive Lipid Metabolite 2- trans-Hexadecenal. ACS Chem. Biol. 2018, 13, 1130–1136. [Google Scholar] [CrossRef]
- Ciepla, P.; Konitsiotis, A.D.; Serwa, R.A.; Masumoto, N.; Leong, W.P.; Dallman, M.J.; Magee, A.I.; Tate, E.W. New chemical probes targeting cholesterylation of Sonic Hedgehog in human cells and zebrafish. Chem. Sci. 2014, 5, 4249–4259. [Google Scholar] [CrossRef]
- Zheng, B.; Jarugumilli, G.K.; Chen, B.; Wu, X. Chemical Probes to Directly Profile Palmitoleoylation of Proteins. Chembiochem 2016, 17, 2022–2027. [Google Scholar] [CrossRef]
- Codreanu, S.G.; Kim, H.Y.; Porter, N.A.; Liebler, D.C. Biotinylated probes for the analysis of protein modification by electrophiles. Methods Mol. Biol. 2012, 803, 77–95. [Google Scholar] [PubMed]
- Kim, H.Y.; Tallman, K.A.; Liebler, D.C.; Porter, N.A. An azido-biotin reagent for use in the isolation of protein adducts of lipid-derived electrophiles by streptavidin catch and photorelease. Mol. Cell Proteom. 2009, 8, 2080–2089. [Google Scholar] [CrossRef] [PubMed]
- Szychowski, J.; Mahdavi, A.; Hodas, J.J.; Bagert, J.D.; Ngo, J.T.; Landgraf, P.; Dieterich, D.C.; Schuman, E.M.; Tirrell, D.A. Cleavable biotin probes for labeling of biomolecules via azide-alkyne cycloaddition. J. Am. Chem. Soc. 2010, 132, 18351–18360. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.L.; Verhelst, S.H.L. Cleavable trifunctional biotin reagents for protein labelling, capture and release. Chem. Commun. 2013, 49, 5366–5368. [Google Scholar] [CrossRef] [PubMed]
- Hang, H.C.; Linder, M.E. Exploring protein lipidation with chemical biology. Chem. Rev. 2011, 111, 6341–6358. [Google Scholar] [CrossRef]
- Soreghan, B.A.; Yang, F.; Thomas, S.N.; Hsu, J.; Yang, A.J. High-throughput proteomic-based identification of oxidatively induced protein carbonylation in mouse brain. Pharm. Res. 2003, 20, 1713–1720. [Google Scholar] [CrossRef] [PubMed]
- Codreanu, S.G.; Zhang, B.; Sobecki, S.M.; Billheimer, D.D.; Liebler, D.C. Global analysis of protein damage by the lipid electrophile 4-hydroxy-2-nonenal. Mol. Cell Proteom. 2009, 8, 670–680. [Google Scholar] [CrossRef]
- Dong, L.; Li, J.; Li, L.; Li, T.; Zhong, H. Comparative analysis of S-fatty acylation of gel-separated proteins by stable isotope-coded fatty acid transmethylation and mass spectrometry. Nat. Protoc. 2011, 6, 1377–1390. [Google Scholar] [CrossRef]
- Sorek, N.; Yalovsky, S. Analysis of protein S-acylation by gas chromatography-coupled mass spectrometry using purified proteins. Nat. Protoc. 2010, 5, 834–840. [Google Scholar] [CrossRef]
- Sorek, N.; Akerman, A.; Yalovsky, S. Analysis of protein prenylation and S-acylation using gas chromatography-coupled mass spectrometry. Methods Mol. Biol. 2013, 1043, 121–134. [Google Scholar]
- Xue, Y.; Chen, H.; Jin, C.; Sun, Z.; Yao, X. NBA-Palm: Prediction of palmitoylation site implemented in Naïve Bayes algorithm. BMC Bioinform. 2006, 7, 458. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Wen, L.; Gao, X.; Jin, C.; Xue, Y.; Yao, X. CSS-Palm 2.0: An updated software for palmitoylation sites prediction. Protein Eng. Des. Sel. 2008, 21, 639–644. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Xue, Y.; Yao, X.; Xu, Y. CSS-Palm: Palmitoylation site prediction with a clustering and scoring strategy (CSS). Bioinformatics 2006, 22, 894–896. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.-B.; Wu, L.-Y.; Wang, Y.-C.; Deng, N.-Y. Prediction of palmitoylation sites using the composition of k-spaced amino acid pairs. Protein Eng. Des. Sel. 2009, 22, 707–712. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Adams, R.M.; Chourey, K.; Hurst, G.B.; Hettich, R.L.; Pan, C. Systematic comparison of label-free, metabolic labeling, and isobaric chemical labeling for quantitative proteomics on LTQ Orbitrap Velos. J. Proteome Res. 2012, 11, 1582–1590. [Google Scholar] [CrossRef] [PubMed]
- Tom, C.T.; Martin, B.R. Fat chance! Getting a grip on a slippery modification. ACS Chem. Biol. 2013, 8, 46–57. [Google Scholar] [CrossRef]
- Ong, S.-E.; Blagoev, B.; Kratchmarova, I.; Kristensen, D.B.; Steen, H.; Pandey, A.; Mann, M. Stable Isotope Labeling by Amino Acids in Cell Culture, SILAC, as a Simple and Accurate Approach to Expression Proteomics. Mol. Cell. Proteom. 2002, 1, 376. [Google Scholar] [CrossRef]
- Wan, J.; Savas, J.N.; Roth, A.F.; Sanders, S.S.; Singaraja, R.R.; Hayden, M.R.; Yates, J.R., 3rd; Davis, N.G. Tracking brain palmitoylation change: Predominance of glial change in a mouse model of Huntington’s disease. Chem. Biol. 2013, 20, 1421–1434. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, L.; Ji, G.; Lei, Q.; Fang, C.; Lu, H. Site-Specific Quantification of Protein Palmitoylation by Cysteine-Stable Isotope Metabolic Labeling. Anal Chem 2018, 90, 10543–10550. [Google Scholar] [CrossRef]
- Weerapana, E.; Wang, C.; Simon, G.M.; Richter, F.; Khare, S.; Dillon, M.B.; Bachovchin, D.A.; Mowen, K.; Baker, D.; Cravatt, B.F. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 2010, 468, 790–795. [Google Scholar] [CrossRef]
- Hemsley, P.A.; Weimar, T.; Lilley, K.S.; Dupree, P.; Grierson, C.S. A proteomic approach identifies many novel palmitoylated proteins in Arabidopsis. New Phytol. 2013, 197, 805–814. [Google Scholar] [CrossRef] [PubMed]
- Shakir, S.; Vinh, J.; Chiappetta, G. Quantitative analysis of the cysteine redoxome by iodoacetyl tandem mass tags. Anal. Bioanal. Chem. 2017, 409, 3821–3830. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.X.; Hannoush, R.N. Single-cell imaging of Wnt palmitoylation by the acyltransferase porcupine. Nat. Chem. Biol. 2014, 10, 61–68. [Google Scholar] [CrossRef]
- Dursina, B.; Reents, R.; Delon, C.; Wu, Y.W.; Kulharia, M.; Thutewohl, M.; Veligodsky, A.; Kalinin, A.; Evstifeev, V.; Ciobanu, D.; et al. Identification and specificity profiling of protein prenyltransferase inhibitors using new fluorescent phosphoisoprenoids. J. Am. Chem. Soc. 2006, 128, 2822–2835. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Hannoush, R.N. Single-cell in situ imaging of palmitoylation in fatty-acylated proteins. Nat. Protoc. 2014, 9, 2607–2623. [Google Scholar] [CrossRef]
- Ismail, V.S.; Mosely, J.A.; Tapodi, A.; Quinlan, R.A.; Sanderson, J.M. The lipidation profile of aquaporin-0 correlates with the acyl composition of phosphoethanolamine lipids in lens membranes. Biochim. Biophys. Acta 2016, 1858, 2763–2768. [Google Scholar] [CrossRef]
- Kim, M.S.; Pinto, S.M.; Getnet, D.; Nirujogi, R.S.; Manda, S.S.; Chaerkady, R.; Madugundu, A.K.; Kelkar, D.S.; Isserlin, R.; Jain, S.; et al. A draft map of the human proteome. Nature 2014, 509, 575–581. [Google Scholar] [CrossRef]
- Wilhelm, M.; Schlegl, J.; Hahne, H.; Gholami, A.M.; Lieberenz, M.; Savitski, M.M.; Ziegler, E.; Butzmann, L.; Gessulat, S.; Marx, H.; et al. Mass-spectrometry-based draft of the human proteome. Nature 2014, 509, 582–587. [Google Scholar] [CrossRef]
- Fuentes, N.R.; Kim, E.; Fan, Y.Y.; Chapkin, R.S. Omega-3 fatty acids, membrane remodeling and cancer prevention. Mol. Asp. Med. 2018, 64, 79–91. [Google Scholar] [CrossRef]
- Yang, W.S.; Kim, K.J.; Gaschler, M.M.; Patel, M.; Shchepinov, M.S.; Stockwell, B.R. Peroxidation of polyunsaturated fatty acids by lipoxygenases drives ferroptosis. Proc. Natl. Acad. Sci. USA 2016, 113, E4966–E4975. [Google Scholar] [CrossRef]
- Shaikh, S.R.; Edidin, M. Polyunsaturated fatty acids, membrane organization, T cells, and antigen presentation. Am. J. Clin. Nutr. 2006, 84, 1277–1289. [Google Scholar] [CrossRef] [PubMed]
- Shaikh, S.R.; Kinnun, J.J.; Leng, X.; Williams, J.A.; Wassall, S.R. How polyunsaturated fatty acids modify molecular organization in membranes: Insight from NMR studies of model systems. Biochim. Biophys. Acta 2015, 1848, 211–219. [Google Scholar] [CrossRef] [PubMed]
- Cravatt, B.F.; Simon, G.M.; Yates, J.R. 3rd, The biological impact of mass-spectrometry-based proteomics. Nature 2007, 450, 991–1000. [Google Scholar] [CrossRef] [PubMed]
- Aebersold, R.; Mann, M. Mass spectrometry-based proteomics. Nature 2003, 422, 198–207. [Google Scholar] [CrossRef]
- Domon, B.; Aebersold, R. Mass spectrometry and protein analysis. Science 2006, 312, 212–217. [Google Scholar] [CrossRef]
- Zhou, B.; Wang, Y.; Yan, Y.; Mariscal, J.; Di Vizio, D.; Freeman, M.R.; Yang, W. Low-Background Acyl-Biotinyl Exchange Largely Eliminates the Coisolation of Non-S-Acylated Proteins and Enables Deep S-Acylproteomic Analysis. Anal. Chem. 2019, 91, 9858–9866. [Google Scholar] [CrossRef]
- Schulte-Zweckel, J.; Dwivedi, M.; Brockmeyer, A.; Janning, P.; Winter, R.; Triola, G. A hydroxylamine probe for profiling S-acylated fatty acids on proteins. Chem. Commun. 2019, 55, 11183–11186. [Google Scholar] [CrossRef]
- Windsor, K.; Genaro-Mattos, T.C.; Kim, H.Y.; Liu, W.; Tallman, K.A.; Miyamoto, S.; Korade, Z.; Porter, N.A. Probing lipid-protein adduction with alkynyl surrogates: Application to Smith-Lemli-Opitz syndrome. J. Lipid Res. 2013, 54, 2842–2850. [Google Scholar] [CrossRef]
- Zhu, J.; Warner, E.; Parikh, N.D.; Lubman, D.M. Glycoproteomic markers of hepatocellular carcinoma-mass spectrometry based approaches. Mass Spectrom. Rev. 2019, 38, 265–290. [Google Scholar] [CrossRef]
- Zhang, Y.; Xie, X.; Zhao, X.; Tian, F.; Lv, J.; Ying, W.; Qian, X. Systems analysis of singly and multiply O-glycosylated peptides in the human serum glycoproteome via EThcD and HCD mass spectrometry. J. Proteom. 2018, 170, 14–27. [Google Scholar] [CrossRef]
- Yu, Q.; Shi, X.; Feng, Y.; Kent, K.C.; Li, L. Improving data quality and preserving HCD-generated reporter ions with EThcD for isobaric tag-based quantitative proteomics and proteome-wide PTM studies. Anal. Chim. Acta 2017, 968, 40–49. [Google Scholar] [CrossRef] [PubMed]
- Meier, F.; Park, M.A.; Mann, M. Trapped Ion Mobility Spectrometry and Parallel Accumulation-Serial Fragmentation in Proteomics. Mol. Cell Proteom. 2021, 20, 100138. [Google Scholar] [CrossRef] [PubMed]
- Winter, D.L.; Wilkins, M.R.; Donald, W.A. Differential Ion Mobility-Mass Spectrometry for Detailed Analysis of the Proteome. Trends Biotechnol. 2019, 37, 198–213. [Google Scholar] [CrossRef] [PubMed]
- Kanu, A.B.; Dwivedi, P.; Tam, M.; Matz, L.; Hill, H.H., Jr. Ion mobility-mass spectrometry. J. Mass Spectrom. 2008, 43, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Creese, A.J.; Smart, J.; Cooper, H.J. Large-scale analysis of peptide sequence variants: The case for high-field asymmetric waveform ion mobility spectrometry. Anal. Chem. 2013, 85, 4836–4843. [Google Scholar] [CrossRef]
- Zhang, Z.; Wu, S.; Stenoien, D.L.; Pasa-Tolic, L. High-throughput proteomics. Annu. Rev. Anal. Chem. 2014, 7, 427–454. [Google Scholar] [CrossRef]
- Pan, J.; Borchers, C.H. Top-down mass spectrometry and hydrogen/deuterium exchange for comprehensive structural characterization of interferons: Implications for biosimilars. Proteomics 2014, 14, 1249–1258. [Google Scholar] [CrossRef]
- Zinnel, N.F.; Pai, P.J.; Russell, D.H. Ion mobility-mass spectrometry (IM-MS) for top-down proteomics: Increased dynamic range affords increased sequence coverage. Anal. Chem. 2012, 84, 3390–3397. [Google Scholar] [CrossRef]

| Modification | Lipid | Structure | Linkage | Modified Residue | References | |
|---|---|---|---|---|---|---|
| 1 | S-palmitoylation | Palmitic acid (C16:0) |  | Thioester | Cysteine | [22,23,28,29] |
| 2 | N-terminal palmitoylation | Palmitic acid (C16:0) | Amide | N-terminal Cysteine | [31,44] | |
| 3 | Nε-palmitoylation | Palmitic acid (C16:0) | Amide | Lysine | [45,46] | |
| 4 | O-palmitoylation | Palmitic acid (C16:0) | Oxyester | Serine | [47] | |
| Threonine | [48] | |||||
| 5 | N-terminal myristoylation | Myristic acid (C14:0) |  | Amide | N-terminal Glycine | [49] |
| 6 | Nε-myristoylation | Myristic acid (C14:0) | Amide | Lysine | [50,51,52] | |
| 7 | S-stearoylation | Stearic acid (C18:0) |  | Thioester | Cysteine | [53,54] |
| 8 | O-octanoylation | Octanoic acid (C8:0) |  | Oxyester | Serine | [55,56] |
| 9 | O-palmitoleoylation | Palmitoleic acid (C16:1n7) |  | Oxyester | Serine | [57,58,59] |
| 10 | N-oleoylation | Oleic acid (C18:1n9) |  | Amide | Lysine | [60] |
| 11 | Unnamed | Arachidonic acid (C20:4n6) |  | Yet unknown | Yet unknown | [61] |
| 12 | Unnamed | Eicosapentaenoic acid (C20:5n3) |  | Yet unknown | Yet unknown | [61] |
| 13 | N-lipoylation | Lipoic acid |  | Amide | Lysine | [62,63] |
| 14 | S-prenylation | Isoprenoid |  | Untitled | C-terminal Cysteine | [64,65] |
 | ||||||
| 15 | C-terminal phosphatidyl-ethanolaminylation | PE |  | Amide | C-terminal Glycine | [66,67] |
| 16 | C-terminal cholesterolyation | Cholesterol |  | Oxyester | C-terminus | [68,69] |
| 17 | C-terminal GPI anchor | GPI |  | Amide | C-terminus | [70,71] |
| 18 | LDE acylation | LDE |  | Carbonyls | Nucleophilic residues | [72,73] |
| Aldehydes | ||||||
| Radioactive Isotope-Labeling | Antibody Affinity Enrichment | ABE | Click Chemistry | Biotin Hydrazide Affinity Capture | Lipid Esterification | |
|---|---|---|---|---|---|---|
| Procedures | 3H/14C metabolic labeling, radiography | Pan-antibody detection of modified moieties | Block-free thiols, cleavage thioester bonds, capture-exposed thiols, IP with streptavidin, WB or elution for MS | alkynyl/azide-lipid probe incorporation, click reaction, IP with streptavidin, elution for MS | Carbonyl group and biotin-hydrazide linkage, capture and analyze LDEs | Dissociative lipids with esterification, GC-MS analysis |
| Applications | Detection of lipidated proteins | Detection of lipidated proteins | Detection of Cysteine S-acylation | Detection of lipidated proteins | Detection of protein lipidation with LDEs | Detection of lipidation |
| Advantages | Direct detection of lipidated proteins without altering the lipid structure | Amenable for protein enrichment | Efficiently distinguishes S-palmitoylation | Availability of alkynyl/azide-lipid probes | Simple method for LDE detection | Quantification of lipid species |
| Disadvantages | Radioactive exposure, limited by the availability of radio-labeled fatty acid | Limited by the availability of pan-antibodies | High background | Interference with endogenous lipidation | Unable to identify the modified sites, high background | Unable to identify the modified sites, high background |
| Throughput | Low | High | High | High | High | High |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, R.; Chen, Y.Q. Protein Lipidation Types: Current Strategies for Enrichment and Characterization. Int. J. Mol. Sci. 2022, 23, 2365. https://doi.org/10.3390/ijms23042365
Wang R, Chen YQ. Protein Lipidation Types: Current Strategies for Enrichment and Characterization. International Journal of Molecular Sciences. 2022; 23(4):2365. https://doi.org/10.3390/ijms23042365
Chicago/Turabian StyleWang, Rong, and Yong Q. Chen. 2022. "Protein Lipidation Types: Current Strategies for Enrichment and Characterization" International Journal of Molecular Sciences 23, no. 4: 2365. https://doi.org/10.3390/ijms23042365
APA StyleWang, R., & Chen, Y. Q. (2022). Protein Lipidation Types: Current Strategies for Enrichment and Characterization. International Journal of Molecular Sciences, 23(4), 2365. https://doi.org/10.3390/ijms23042365






