Recent Advances in the Production of Genome-Edited Rats
Abstract
:1. Introduction
2. What Is GE Technology?
3. Delivery Method
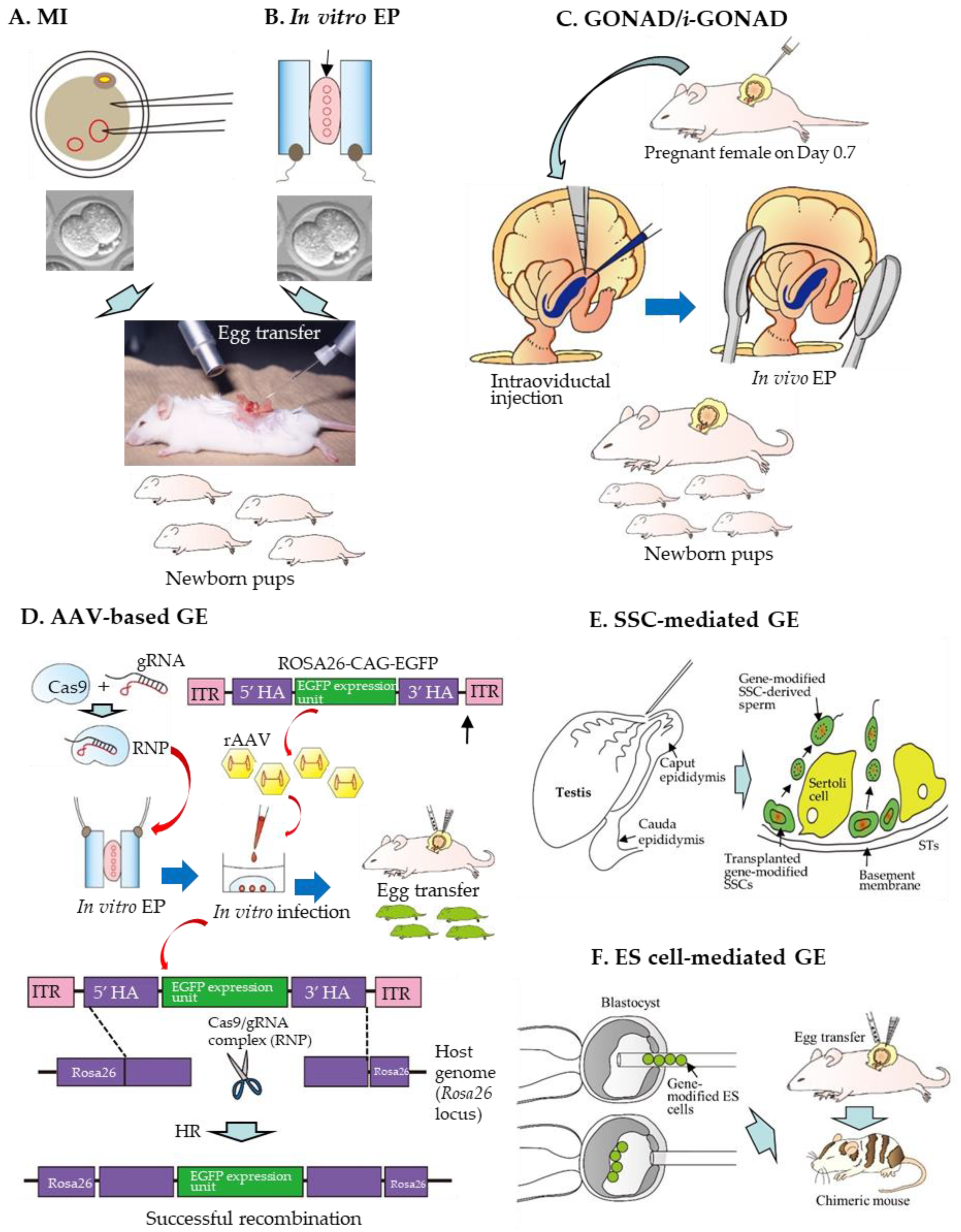
3.1. MI-Based Production of GE Rats
3.2. In Vitro EP-Based Production of GE Rats
3.3. GONAD-Based Production of GE Rats
3.4. rAAV-Mediated Production of GE Rats
3.5. SSC-Mediated Production of GE Rats
3.6. ES Cell-Mediated Production of GE Rats
4. Other Techniques and Factors to Modify the Rat Genome
4.1. Allele-Specific GE for the Correction of Mutated Phenotypes
4.2. Large Genomic Fragment Deletion
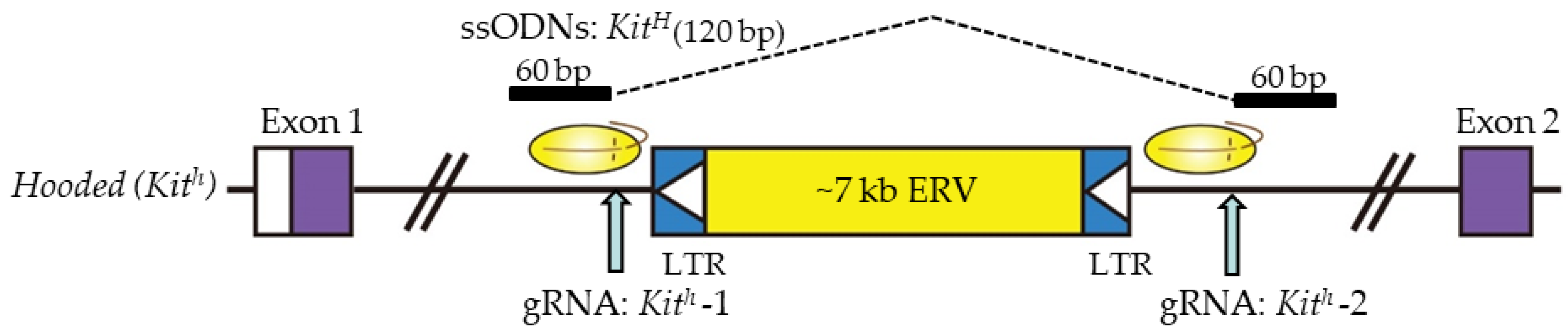
4.3. ssODN-Mediated KI with CRISPR/Cas9 for Large Genomic Regions in Rat Zygotes
4.4. Use of Inhibitor for Efficient KI
4.5. Choice of gRNA Is Very Important for Achieving High KI Efficiency in Rats
4.6. Base Editor Systems as An Efficient Tool Enabling Precise Genome Modifications
5. Disease Models in Rats
5.1. Models for Cardiovascular Diseases
5.2. Models for Neurological Diseases
5.3. Models for Muscular Diseases
5.4. Models for Pulmonary Diseases
5.5. Models for Metabolic Diseases
5.6. Models for Kidney Diseases
5.7. Models for Ophthalmology Diseases
5.8. Models for Hematological Systems
5.9. Others
6. Perspective
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Abcc6 | ATP binding cassette subtype C number 6 |
| ABE | Adenine base editor |
| Abhd6 | Alpha/beta-hydrolase domain 6 |
| Adora2a | Adenosine 2a receptor |
| α-Gal A | α-galactosidase A |
| Agxt | Alanine-glyoxylate aminotransferase |
| Ahr | Aryl hydrocarbon receptor |
| Aire | Autoimmune regulator |
| ALS | Amyotrophic lateral sclerosis |
| Angptl8 | Angiopoietin-like protein 8 |
| Anks3 | Ankyrin repeat and sterile alpha motif domain containing 3 |
| APECED | Autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy |
| ApoE | Apolipoprotein E |
| AS | Angelman syndrome |
| AsCpf1 | Acidaminococcus sp. Cpf1 |
| ASD | Autism spectrum disorder |
| Asip | Agouti signaling protein |
| Atg16l1 | Autophagy-related 16-like 1 |
| B2m | Beta-2-microglobulin |
| BE | Base-editing |
| BlatCas9 | Brevibacillus laterosporus Cas9 |
| Bmpr2 | Bone morphogenetic protein receptor 2 |
| BN | Brown Norway |
| C3 | Complement C3 |
| C9orf72 | Chromosome 9 open reading frame 72 |
| Cas12a | CRISPR-associated protein 12a |
| Calb2 | Calbindin 2 |
| Carns1 | Carnosine synthase 1 |
| Cas9n | Cas9 nickase |
| CBE | Cytosine base editor |
| Ccdc85c | Coiled-coil domain containing 85C |
| Cck | Cholecystokinin |
| CD | Crohn’s disease |
| CES2 | Carboxylesterase 2 |
| Ces2a | Carboxylesterase 2A |
| CF | Cystic fibrosis |
| Cftr | Cystic fibrosis transmembrane conductance regulator |
| CIPN | Chemotherapy-induced peripheral neuropathy |
| Cirp | Cold-inducible RNA-binding protein |
| CLICK | CRISPR with lssDNA inducing conditional knockout alleles |
| Cp | Ceruloplasmin |
| Cpf1 | CRISPR from Prevotella and Francisella1 |
| Cplx1 | Complexin I |
| Cre-ERT2 | Cre recombinase-estrogen receptor T2 |
| Crhr1 | Corticotropin releasing hormone receptor 1 |
| CRISPR/Cas9 | Clustered regularly interspaced palindrome repeats (CRISPR)/Caspase 9 (Cas9) |
| Cas12a | Type V CRISPR-effector protein (formerly Cpf1) |
| crRNA | CRISPR RNA |
| CS | Cockayne syndrome |
| Csb | Cockayne syndrome B |
| Cyp2c11 | Cytochrome P450, subfamily 2, polypeptide 11 |
| Cyp2d | Cytochrome P450 family 2 subfamily D member 6 |
| Cyp2e1 | Cytochrome P450 2E1 |
| Cyp2J2 | Cytochrome P450 family 2 subfamily J member 2 |
| Cyp2J3 | Cytochrome P450, family 2, subfamily j, polypeptide 3 |
| Cyp2J10 | Cytochrome P450, family 2, subfamily j, polypeptide 10 |
| Cyp3a1 | Cytochrome P450 3A1 |
| Cyp3a2 | Cytochrome P450 3A2 |
| Cyp24a1 | Cytochrome P450 family 24 subfamily A member 1 |
| Cyp27b1 | Cytochrome P450 family 27 subfamily B member 1 |
| DA | Dark agouti |
| Dat | Dopamine transporter |
| db | Diabetes |
| Dbndd1 | Dysbindin (dystrobrevin binding protein 1) domain containing 1 |
| Defb23 | Defensin beta 23 |
| Defb26 | Defensin beta 26 |
| Defb42 | Defensin beta 42 |
| dCas9 | Dead Cas9 |
| DdCBE | DddA-derived cytosine base editor |
| DddA | Bacterial toxin deaminase |
| Depdc5 | DEP domain-containing 5 |
| Des | Desmin |
| DdCBE | DddA-derived cytosine base editor |
| DMD | Duchenne muscular dystrophy |
| Dmd | Dystrophin-coding gene (dystrophin) |
| DKO | Double KO |
| DNA-PKcs | DNA-dependent protein kinase catalytic subunits |
| Dnah17 | Dynein axonemal heavy chain 17 |
| Drd1a | Dopamine receptor D1A |
| DSB | Double-stranded break |
| dsDNA | Double-stranded DNA |
| dsRed | Discosoma coral red fluorescent protein |
| EGFP | Enhanced green fluorescent protein |
| EndoMT | Endothelial-to-mesenchymal transition |
| EP | Electroporation |
| Epsti1 | Epithelial stromal interaction 1 |
| Erbb3 | Erb-B2 receptor tyrosine kinase 3 |
| ERV | Endogenous retroviral element |
| ES | Embryonic stem |
| Esr1 | Estrogen receptor 1 |
| ET | Egg transfer |
| F344 | Fisher 344 |
| F8 | Factor 8 |
| Fabp2 | Fatty acid binding protein 2 |
| Fah | Fumarylacetoacetate hydrolase |
| FD | Fabry disease |
| Fh | Fumarate hydratase |
| FH | Familial hypercholesterolemia |
| FHH | Fawn-hooded hypertensive |
| Fmr1 | Fragile X mental retardation 1 |
| FMRP | Fragile X mental retardation protein |
| FnCas9 | Francisella novicida Cas9 |
| Foxp3 | Forkhead box P3 |
| Foxn1 | Forkhead box N1 |
| FSG | F344-scid gamma |
| Fus | Fused in sarcoma |
| FXS | Fragile X syndrome |
| Gaa | Acid alpha-glucosidase |
| Gad1 | Glutamate decarboxylase 1 |
| GAD67 | Glutamate decarboxylase 67-kDa isoform |
| GE | Genome editing or genome-edited |
| GFP | Green fluorescent protein |
| G6pd | Glucose-6-phosphate dehydrogenase |
| GM | Genetically modified |
| GOI | Gene of interest |
| GONAD | Genome-editing via oviductal nucleic acids delivery |
| GR | Glucocorticoid receptors |
| GRdim | Glucocorticoid receptor mutant |
| Grin1 | Glutamate ionotropic receptor N-methyl-D-aspartate (NMDA) type subunit 1 |
| gRNA | Guide RNA |
| Gsd2 | Glycogen storage disease type 2 |
| GSDII | Glycogen storage disease type II (GSDII) |
| 2H2OP | Two-hit by gRNA and two-oligo with plasmid |
| HA | Hereditary aceruloplasminemia |
| HCD | High-cholesterol diet |
| HDR | Homology directed repair |
| Hemgn | Hemogen |
| hhy | Hemorrhagic hydrocephalus |
| Hprt1 | Hypoxanthine phosphoribosyltransferase 1 |
| HR | Homologous recombination |
| HT1 | Hereditary tyrosinemia type I |
| Htr7 | 5-hydroxytryptamine receptor 7 |
| IC | Intracytoplasmic injection |
| Ighm | Immunoglobulin heavy constant mu |
| IgM | Immunoglobulin M |
| i-GONAD | Improved GONAD |
| Il2rg | Interleukin 2 receptor gamma |
| indels | Insertion or deletion of nucleotides |
| iNOS | Inducible nitric oxide synthase |
| IRES | Internal ribosomal entry site |
| IVF | In vitro fertilized |
| Kat II | Kynurenine aminotransferase II |
| Kcnj16 | Potassium inwardly-rectifying channel, subfamily J, member 16 |
| Kcnk3 | Potassium channel subfamily K member 3 |
| KI | Knock-in |
| Kit | Kit proto-oncogene |
| KO | Knockout |
| L1cam | L1 cell adhesion molecule |
| LbCpf1 | Lachnospiraceae bacterium Cpf1 |
| Ldlr | Low-density lipoprotein receptor |
| Lrat | Lecithin retinol acyltransferase |
| Lepr | Leptin receptor |
| LE | Long–Evans |
| LEH | Long–Evans hooded |
| Lepr | Leptin receptor |
| LEW | Lewis |
| Lgr5 | Leucine-rich repeat-containing G-protein coupled receptor 5 |
| lncRNA | Long non-coding RNA |
| Lrat | Lecithin retinol acyltransferase |
| lssDNA | Long single-stranded DNA |
| Mc3r | Melanocortin 3 receptor |
| Mc4r | Melanocortin 4 receptor |
| Mdr1a | Multidrug resistance protein 1a |
| Mdr1b | Multidrug resistance protein 1b |
| Mdx | X-linked muscular dystrophy |
| Mecp2 | Methyl-CpG-binding protein 2 |
| MI | Microinjection |
| mitoTALE | TALE nickase enabling mitochondrial gene editing |
| Mkx | Mohawk homeobox |
| Mnx1 | Gene encoding homeobox protein HB9 |
| mtDNA | Mitochondrial DNA |
| mTORC1 | Rapamycin complex 1 signaling pathway |
| Myl4 | Myosin light-chain 4 |
| NAFLD | Nonalcoholic fatty liver disease |
| nCas9 | Cas9 nickase |
| Ndst1 | N-deacetylase |
| Ndst4 | N-sulfotransferase 4 |
| Nes | Nestin |
| Nfatc1 | Nuclear factor of activated T-cells |
| Ngln3 | Neuroligin 3 |
| NHEJ | Non-homologous end joining |
| Nkx3.1 | NK3 homeobox 1 |
| NmCas9 | Neisseria meningitidis Cas9 |
| Nr3c1 | Nuclear receptor subfamily 3, group C, member 1 |
| Nrf2 | Nuclear factor (erythroid-derived 2)-like-2 |
| Oatp1b2 | Organic anion transport polypeptide 1b2 |
| Ogdh | Oxoglutarate dehydrogenase |
| PAH | Pulmonary arterial hypertension |
| PAM | Protospacer adjacent motif |
| PB | PiggyBack transposon |
| PBS | Phosphate-buffered saline |
| PE | Prime editing |
| PGK | Phosphoglycerate kinase |
| PH | Pulmonary hypertension |
| PH1 | Primary hyperoxaluria type 1 |
| PI | Pronuclear injection |
| PMCA4 | Plasma membrane calmodulin–dependent calcium ATPase isoform 4 |
| PMS | Phelan–McDermid syndrome |
| Pnpla5 | Patatin-like phospholipase domain containing 5 |
| poly(A) | Polyadenine tails |
| Pomc | Proopiomelanocortin |
| Pp | Poring pulse |
| PPi | Pyrophosphate |
| Prf1 | Perforin 1 |
| Prkdc | DNA-dependent protein kinase catalytic subunit |
| Psen1 | Presenilin 1 |
| p52SHC/p46SHC | Adaptor protein 1 (shc1), encodes 3 main protein isoforms |
| Pvalb | Parvalbumin |
| PVG | Piebald Virol Glaxo |
| PXE | Pseudoxanthoma elasticum |
| Pxr | Pregnane X receptor |
| rAAV | Recombinant adeno-associated virus |
| Rab38 | Ras-related protein Rab-38 |
| Rag1 | Recombination activating gene 1 |
| Rag2 | Recombination activating gene 2 |
| RD | Retinal dystrophy |
| Ren | Renin |
| Rffl-lnc1 | A gene located within the 5′UTR intronic region of the rififylin (Rffl) gene |
| rGONAD | Rat-based i-GONAD |
| RNaseT2 | Ribonuclease T2 |
| RNP | Ribonucleoprotein |
| Rosa26 | Gt(ROSA)26Sor |
| RP | Retinitis pigmentosa |
| RS | Rett syndrome |
| SaCas9 | Staphylococcus aureus Cas9 |
| SCID | Severe combined immunodeficiency |
| SD | Sprague–Dawley |
| SERCA2a | Sarcoplasmic reticulum Ca2+-ATPase |
| sgRNA | Single-guide RNA |
| Shank3 | SH3 and multiple ankyrin repeat domains 3 |
| Sirpa | Signal regulatory protein alpha |
| Slc6 | Sodium- and chloride-dependent members of the solute carrier family 6 |
| Slc16a2 | Solute carrier family 16 member 2 coding for monocarboxylate transporter 8 (MCT8) |
| Slco1b2 | Solute carrier organic anion transporter family member 1B2 |
| SNP | Single-nucleotide polymorphism |
| Sox10 | Sry-box transcription factor 10 |
| Sox16 | SRY (sex determining region Y)-box 16 |
| SS | Dahl salt-sensitive |
| SSCs | Spermatogonial stem cells |
| ssDNA | Single-stranded DNA |
| ssODN | Single-stranded oligodeoxynucleotide |
| St1Cas9 | Streptococcus thermophilus Cas9 |
| STs | Seminiferous tubules |
| TAKE | Technique for Animal Knockout system by Electroporation |
| TALE | Transcriptional activator-like effector |
| TALENs | Transcription activator-like effector nucleases |
| Tet1 | Tet methylcytosine dioxygenase 1 |
| Tet2 | Tet methylcytosine dioxygenase 2 |
| Tet3 | Tet methylcytosine dioxygenase 3 |
| Tg | Transgenic |
| Th | Tyrosine hydroxylase |
| Thy1 | Thy-1 cell surface antigen |
| Tie2 | Tunica interna endothelial cell kinase 2 |
| Tlr4 | Toll-like receptor 4 |
| TM/Kyo | TRM or tremor rat |
| Tp | Transfer pulse |
| Tp53 | Tumor protein p53 |
| tracrRNA | Trans-activating CRISPR RNA |
| Trdmt1 | TRNA aspartic acid methyltransferase 1 |
| Trpa1 | Transient receptor potential family member ankyrin 1 |
| Trpv4 | Transient receptor potential cation channel subfamily V member 4 |
| Tyr | Tyrosinase |
| Ube3a | Ubiquitin protein ligase E3A |
| Uox | Urate oxidase (uricase) |
| Vapb | Vesicle-associated membrane protein-associated protein B/C |
| Vdr | Vitamin D receptor |
| Wfs1 | Wolfram syndrome 1 |
| WI | Wistar |
| WI-IM | Wistar–Imamichi |
| WKY | Wistar–Kyoto |
| Wnt1 | Wnt family member 1 |
| WS | Wolfram syndrome |
| WT | Wild-type |
| XLH | X-linked hydrocephalus |
| X-SCID | X-linked severe combined immunodeficiency |
| ZF | Zinc-finger |
| ZFNs | Zinc-finger nucleases |
| ZP | Zona pellucida |
References
- Kjell, J.; Olson, L. Rat models of spinal cord injury: From pathology to potential therapies. Dis. Models Mech. 2016, 9, 1125–1137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jacob, H.J. The Rat: A model used in biomedical research. In Rat Genomics: Methods and Protocols Methods in Molecular Biology; Anegon, I., Ed.; Humana Press: Totowa, NJ, USA, 2010; pp. 1–11. [Google Scholar] [CrossRef]
- Hammes, A.; Oberdorf-Maass, S.; Rother, T.; Nething, K.; Gollnick, F.; Linz, K.W.; Meyer, R.; Hu, K.; Han, H.; Gaudron, P.; et al. Overexpression of the sarcolemmal calcium pump in the myocardium of transgenic rats. Circ. Res. 1998, 83, 877–888. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vetter, R.; Rehfeld, U.; Reissfelder, C.; Weifl, W.; Wagner, K.D.; Günther, J.; Hammes, A.; Tschöpe, C.; Dillmann, W.; Paul, M. Transgenic overexpression of the sarcoplasmic reticulum Ca2+ ATPase improves reticular Ca2+ handling in normal and diabetic rat hearts. FASEB J. 2002, 16, 1657–1659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, K.R.; Capecchi, M.R. Site-directed mutagenesis by gene targeting in mouse embryo-derived stem cells. Cell 1987, 51, 503–512. [Google Scholar] [CrossRef]
- Doetschman, T.; Gregg, R.G.; Maeda, N.; Hooper, M.L.; Melton, D.W.; Thompson, S.; Smithies, O. Targetted correction of a mutant HPRT gene in mouse embryonic stem cells. Nature 1987, 330, 576–578. [Google Scholar] [CrossRef]
- Buehr, M.; Meek, S.; Blair, K.; Yang, J.; Ure, J.; Silva, J.; McLay, R.; Hall, J.; Ying, Q.L.; Smith, A. Capture of authentic embryonic stem cells from rat blastocysts. Cell 2008, 135, 1287–1298. [Google Scholar] [CrossRef]
- Li, P.; Tong, C.; Mehrian-Shai, R.; Jia, L.; Wu, N.; Yan, Y.; Maxson, R.E.; Schulze, E.N.; Song, H.; Hsieh, C.L.; et al. Germline competent embryonic stem cells derived from rat blastocysts. Cell 2008, 135, 1299–1310. [Google Scholar] [CrossRef] [Green Version]
- Kawamata, M.; Ochiya, T. Generation of genetically modified rats from embryonic stem cells. Proc. Natl. Acad. Sci. USA 2010, 107, 14223–14228. [Google Scholar] [CrossRef] [Green Version]
- Meek, S.; Buehr, M.; Sutherland, L.; Thomson, A.; Mullins, J.J.; Smith, A.J.; Burdon, T. Efficient gene targeting by homologous recombination in rat embryonic stem cells. PLoS ONE 2010, 5, e14225. [Google Scholar] [CrossRef]
- Tong, C.; Li, P.; Wu, N.L.; Yan, Y.; Ying, Q.L. Production of p53 gene knockout rats by homologous recombination in embryonic stem cells. Nature 2010, 467, 211–213. [Google Scholar] [CrossRef] [Green Version]
- Lan, H.; Li, S.; Guo, Z.; Men, H.; Wu, Y.; Li, N.; Bryda, E.C.; Capecchi, M.R.; Wu, S. Efficient generation of selection-gene-free rat knockout models by homologous recombination in ES cells. FEBS Lett. 2016, 590, 3416–3424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kobayashi, T.; Yamaguchi, T.; Hamanaka, S.; Kato-Itoh, M.; Yamazaki, Y.; Ibata, M.; Sato, H.; Lee, Y.S.; Usui, J.; Knisely, A.S.; et al. Generation of rat pancreas in mouse by interspecific blastocyst injection of pluripotent stem cells. Cell 2010, 142, 787–799. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamamoto, S.; Ooshima, Y.; Nakata, M.; Yano, T.; Nishimura, N.; Nishigaki, R.; Satomi, Y.; Matsumoto, H.; Matsumoto, Y.; Takeyama, M. Efficient gene-targeting in rat embryonic stem cells by CRISPR/Cas and generation of human kynurenine aminotransferase II (KAT II) knock-in rat. Transgenic Res. 2015, 24, 991–1001. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Spitzer, S.; Agathou, S.; Karadottir, R.T.; Smith, A. Gene editing in rat embryonic stem cells to produce in vitro models and in vivo reporters. Stem Cell Rep. 2017, 9, 1262–1274. [Google Scholar] [CrossRef] [Green Version]
- Zan, Y.; Haag, J.D.; Chen, K.S.; Shepel, L.A.; Wigington, D.; Wang, Y.R.; Hu, R.; Lopez-Guajardo, C.C.; Brose, H.L.; Porter, K.I.; et al. Production of knockout rats using ENU mutagenesis and a yeast-based screening assay. Nat. Biotechnol. 2003, 21, 645–651. [Google Scholar] [CrossRef]
- Harrison, M.M.; Jenkins, B.V.; O’Connor-Giles, K.M.; Wildonger, J.A. CRISPR view of development. Genes Dev. 2014, 28, 1859–1872. [Google Scholar] [CrossRef] [Green Version]
- Hsu, P.D.; Lander, E.S.; Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 2014, 157, 1262–1278. [Google Scholar] [CrossRef] [Green Version]
- Sonoda, E.; Hochegger, H.; Saberi, A.; Taniguchi, Y.; Takeda, S. Differential usage of non-homologous end-joining and homologous recombination in double strand break repair. DNA Repair 2006, 5, 1021–1029. [Google Scholar] [CrossRef]
- Kim, Y.G.; Cha, J.; Chandrasegaran, S. Hybrid restriction enzymes: Zinc finger fusions to Fok I cleavage domain. Proc. Natl. Acad. Sci. USA 1996, 93, 1156–1160. [Google Scholar] [CrossRef] [Green Version]
- Wolfe, S.A.; Nekludova, L.; Pabo, C.O. DNA recognition by Cys2His2 zinc finger proteins. Annu. Rev. Biophys. Biomol. Struct. 2000, 29, 183–212. [Google Scholar] [CrossRef]
- Moscou, M.J.; Bogdanove, A.J. A simple cipher governs DNA recognition by TAL effectors. Science 2009, 326, 1501. [Google Scholar] [CrossRef] [PubMed]
- Christian, M.; Cermak, T.; Doyle, E.L.; Schmidt, C.; Zhang, F.; Hummel, A.; Bogdanove, A.J.; Voytas, D.F. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 2010, 186, 757–761. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, T.; Huang, S.; Jiang, W.Z.; Wright, D.; Spalding, M.H.; Weeks, D.P.; Yang, B. TAL nucleases (TALNs): Hybrid proteins composed of TAL effectors and FokI DNA-cleavage domain. Nucleic Acids Res. 2011, 39, 359–372. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Gaj, T.; Gersbach, C.A.; Barbas, C.F., III. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013, 31, 397–405. [Google Scholar] [CrossRef] [Green Version]
- Doudna, J.A.; Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 2014, 346, 1258096. [Google Scholar] [CrossRef]
- Geurts, A.M.; Cost, G.J.; Freyvert, Y.; Zeitler, B.; Miller, J.C.; Choi, V.M.; Jenkins, S.S.; Wood, A.; Cui, X.; Meng, X.; et al. Knockout rats via embryo microinjection of zinc-finger nucleases. Science 2009, 325, 433. [Google Scholar] [CrossRef] [Green Version]
- Mashimo, T.; Takizawa, A.; Voigt, B.; Yoshimi, K.; Hiai, H.; Kuramoto, T.; Serikawa, T. Generation of knockout rats with X-linked severe combined immunodeficiency (X-SCID) using zinc-finger nucleases. PLoS ONE 2010, 5, e8870. [Google Scholar] [CrossRef]
- Ménoret, S.; Iscache, A.L.; Tesson, L.; Rémy, S.; Usal, C.; Osborn, M.J.; Cost, G.J.; Brüggemann, M.; Buelow, R.; Anegon, I. Characterization of immunoglobulin heavy chain knockout rats. Eur. J. Immunol. 2010, 40, 2932–2941. [Google Scholar] [CrossRef]
- Moreno, C.; Hoffman, M.; Stodola, T.J.; Didier, D.N.; Lazar, J.; Geurts, A.M.; North, P.E.; Jacob, H.J.; Greene, A.S. Creation and characterization of a renin knockout rat. Hypertension 2011, 57, 614–619. [Google Scholar] [CrossRef] [Green Version]
- Tesson, L.; Usal, C.; Ménoret, S.; Leung, E.; Niles, B.J.; Remy, S.; Santiago, Y.; Vincent, A.I.; Meng, X.; Zhang, L.; et al. Knockout rats generated by embryo microinjection of TALENs. Nat. Biotechnol. 2011, 29, 695–696. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Ji, D.; Fisher, D.A.; Wu, Y.; Briner, D.M.; Weinstein, E.J. Targeted integration in rat and mouse embryos with zinc-finger nucleases. Nat. Biotechnol. 2011, 29, 64–67. [Google Scholar] [CrossRef]
- Mashimo, T.; Takizawa, A.; Kobayashi, J.; Kunihiro, Y.; Yoshimi, K.; Ishida, S.; Tanabe, K.; Yanagi, A.; Tachibana, A.; Hirose, J.; et al. Generation and characterization of severe combined immunodeficiency rats. Cell Rep. 2012, 2, 685–694. [Google Scholar] [CrossRef] [Green Version]
- Ménoret, S.; Fontanière, S.; Jantz, D.; Tesson, L.; Thinard, R.; Rémy, S.; Usal, C.; Ouisse, L.H.; Fraichard, A.; Anegon, I. Generation of Rag1-knockout immunodeficient rats and mice using engineered meganucleases. FASEB J. 2013, 27, 703–711. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.J.; Fisher, D.A.; Kouranova, E.; McCoy, A.; Forbes, K.; Wu, Y.; Henry, R.; Ji, D.; Chambers, A.; Warren, J.; et al. Whole-rat conditional gene knockout via genome editing. Nat. Methods 2013, 10, 638–640. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Qiu, Z.; Shao, Y.; Chen, Y.; Guan, Y.; Liu, M.; Li, Y.; Gao, N.; Wang, L.; Lu, X.; et al. Heritable gene targeting in the mouse and rat using a CRISPR-Cas system. Nat. Biotechnol. 2013, 31, 681–683. [Google Scholar] [CrossRef]
- Li, W.; Teng, F.; Li, T.; Zhou, Q. Simultaneous generation and germline transmission of multiple gene mutations in rat using CRISPR-Cas systems. Nat. Biotechnol. 2013, 31, 684–686. [Google Scholar] [CrossRef]
- Ferguson, C.; McKay, M.; Harris, R.A.; Homanics, G.E. Toll-like receptor 4 (Tlr4) knockout rats produced by transcriptional activator-like effector nuclease (TALEN)-mediated gene inactivation. Alcohol 2013, 47, 595–599. [Google Scholar] [CrossRef] [Green Version]
- Harrill, J.A.; Hukkanen, R.R.; Lawson, M.; Martin, G.; Gilger, B.; Soldatow, V.; Lecluyse, E.L.; Budinsky, R.A.; Rowlands, J.C.; Thomas, R.S. Knockout of the aryl hydrocarbon receptor results in distinct hepatic and renal phenotypes in rats and mice. Toxicol. Appl. Pharmacol. 2013, 272, 503–518. [Google Scholar] [CrossRef]
- Nielsen, L.N.; Wiinberg, B.; Häger, M.; Holmberg, H.L.; Hansen, J.J.; Roepstorff, K.; Tranholm, M. A novel F8−/− rat as a translational model of human hemophilia A. J. Thromb. Haemost. 2014, 12, 1274–1282. [Google Scholar] [CrossRef]
- Hamilton, S.M.; Green, J.R.; Veeraragavan, S.; Yuva, L.; McCoy, A.; Wu, Y.; Warren, J.; Little, L.; Ji, D.; Cui, X.; et al. Fmr1 and Nlgn3 knockout rats: Novel tools for investigating autism spectrum disorders. Behav. Neurosci. 2014, 128, 103–109. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaneko, T.; Sakuma, T.; Yamamoto, T.; Mashimo, T. Simple knockout by electroporation of engineered endonucleases into intact rat embryos. Sci. Rep. 2014, 4, 6382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Remy, S.; Tesson, L.; Menoret, S.; Usal, C.; De Cian, A.; Thepenier, V.; Thinard, R.; Baron, D.; Charpentier, M.; Renaud, J.B.; et al. Efficient gene targeting by homology-directed repair in rat zygotes using TALE nucleases. Genome Res. 2014, 24, 1371–1383. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, Y.; Ma, J.; Zhang, X.; Chen, W.; Yu, L.; Lu, Y.; Bai, L.; Shen, B.; Huang, X.; Zhang, L. Generation of eGFP and Cre knockin rats by CRISPR/Cas9. FEBS J. 2014, 281, 3779–3790. [Google Scholar] [CrossRef]
- Yoshimi, K.; Kaneko, T.; Voigt, B.; Mashimo, T. Allele-specific genome editing and correction of disease-associated phenotypes in rats using the CRISPR–Cas platform. Nat. Commun. 2014, 5, 4240. [Google Scholar] [CrossRef] [Green Version]
- Ma, Y.; Shen, B.; Zhang, X.; Lu, Y.; Chen, W.; Ma, J.; Huang, X.; Zhang, L. Heritable multiplex genetic engineering in rats using CRISPR/Cas9. PLoS ONE 2014, 9, e89413. [Google Scholar] [CrossRef]
- Rumi, M.A.K.; Dhakal, P.; Kubota, K.; Chakraborty, D.; Lei, T.; Larson, M.A.; Wolfe, M.W.; Roby, K.F.; Vivian, J.L.; Soares, M.J. Generation of Esr1-knockout rats using zinc finger nuclease-mediated genome editing. Endocrinology 2014, 155, 1991–1999. [Google Scholar] [CrossRef] [Green Version]
- De León, V.P.; Anne-Marie, M.; Laurent, T.; Ignacio, A.; Edith, H. Generation of TALEN-mediated GRdim knock-in rats by homologous recombination. PLoS ONE 2014, 9, e88146. [Google Scholar] [CrossRef]
- Nakamura, K.; Fujii, W.; Tsuboi, M.; Tanihata, J.; Teramoto, N.; Takeuchi, S.; Naito, K.; Yamanouchi, K.; Nishihara, M. Generation of muscular dystrophy model rats with a CRISPR/Cas system. Sci. Rep. 2014, 4, 5635. [Google Scholar] [CrossRef]
- Larcher, T.; Lafoux, A.; Tesson, L.; Remy, S.; Thepenier, V.; François, V.; Le Guiner, C.; Goubin, H.; Dutilleul, M.; Guigand, L.; et al. Characterization of dystrophin deficient rats: A new model for Duchenne muscular dystrophy. PLoS ONE 2014, 9, e110371. [Google Scholar] [CrossRef] [Green Version]
- Tuggle, K.L.; Birket, S.E.; Cui, X.; Hong, J.; Warren, J.; Reid, L.; Chambers, A.; Ji, D.; Gamber, K.; Chu, K.K.; et al. Characterization of defects in ion transport and tissue development in cystic fibrosis transmembrane conductance regulator (CFTR)-knockout rats. PLoS ONE 2014, 9, e91253. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ranchoux, B.; Antigny, F.; Rucker-Martin, C.; Hautefort, A.; Péchoux, C.; Bogaard, H.J.; Dorfmüller, P.; Remy, S.; Lecerf, F.; Planté, S.; et al. Endothelial-to-mesenchymal transition in pulmonary hypertension. Circulation 2015, 131, 1006–1018. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaneko, T.; Mashimo, T. Simple genome editing of rodent intact embryos by electroporation. PLoS ONE 2015, 10, e0142755. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.; Xie, D.; Huang, J.; Lv, F.; Shi, D.; Liu, Y.; Lin, L.; Geng, L.; Wu, Y.; Liang, D.; et al. Cold-inducible RNA-binding protein regulates cardiac repolarization by targeting transient outward potassium channels. Circ. Res. 2015, 116, 1655–1659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wei, S.; Zhang, Y.; Su, L.; He, K.; Wang, Q.; Zhang, Y.; Yang, D.; Yang, Y.; Ma, S. Apolipoprotein E-deficient rats develop atherosclerotic plaques in partially ligated carotid arteries. Atherosclerosis 2015, 243, 589–592. [Google Scholar] [CrossRef] [PubMed]
- Ménoret, S.; De Cian, A.; Tesson, L.; Remy, S.; Usal, C.; Boulé, J.B.; Boix, C.; Fontanière, S.; Crénéguy, A.; Nguyen, T.H.; et al. Homology-directed repair in rodent zygotes using Cas9 and TALEN engineered proteins. Sci. Rep. 2015, 5, 14410. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Shao, Y.; Guan, Y.; Li, L.; Wu, L.; Chen, F.; Liu, M.; Chen, H.; Ma, Y.; Ma, X.; et al. Large genomic fragment deletion and functional gene cassette knock-in via Cas9 protein mediated genome editing in one-cell rodent embryos. Sci. Rep. 2015, 5, 17517. [Google Scholar] [CrossRef] [Green Version]
- Bao, D.; Ma, Y.; Zhang, X.; Guan, F.; Chen, W.; Gao, K.; Qin, C.; Zhang, L. Preliminary characterization of a leptin receptor knockout rat created by CRISPR/Cas9 system. Sci. Rep. 2015, 5, 15942. [Google Scholar] [CrossRef] [Green Version]
- Chapman, K.M.; Medrano, G.A.; Jaichander, P.; Chaudhary, J.; Waits, A.E.; Nobrega, M.A.; Hotaling, J.M.; Ober, C.; Hamra, F.K. Targeted germline modifications in rats using CRISPR/Cas9 and spermatogonial stem cells. Cell Rep. 2015, 10, 1828–1835. [Google Scholar] [CrossRef] [Green Version]
- Zhai, W.; Hu, H.; Le, L.; Zhuang, F.; Wang, K.; Zhao, Y.; Wang, K.Z.; Liu, X.M.; Sun, D.; Wang, X.Y.; et al. Generation and analysis of the Rett syndrome-associated MeCP2-null rat model. Yi Chuan 2016, 38, 1004–1011. [Google Scholar] [CrossRef]
- Patterson, K.C.; Hawkins, V.E.; Arps, K.M.; Mulkey, D.K.; Olsen, M.L. MeCP2 deficiency results in robust Rett-like behavioural and motor deficits in male and female rats. Hum. Mol. Genet. 2016, 25, 3303–3320. [Google Scholar] [CrossRef]
- Zhang, L.; Shao, Y.; Li, L.; Tian, F.; Cen, J.; Chen, X.; Hu, D.; Zhou, Y.; Xie, W.; Zheng, Y.; et al. Efficient liver repopulation of transplanted hepatocyte prevents cirrhosis in a rat model of hereditary tyrosinemia type I. Sci. Rep. 2016, 6, 31460. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoshimi, K.; Kunihiro, Y.; Kaneko, T.; Nagahora, H.; Voigt, B.; Mashimo, T. ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes. Nat. Commun. 2016, 7, 10431. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Renaud, J.B.; Boix, C.; Charpentier, M.; De Cian, A.; Cochennec, J.; Duvernois-Berthet, E.; Perrouault, L.; Tesson, L.; Edouard, J.; Thinard, R.; et al. Improved genome editing efficiency and flexibility using modified oligonucleotides with TALEN and CRISPR-Cas9 nucleases. Cell Rep. 2016, 14, 2263–2272. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, D.; Zhong, Y.; Li, X.; Li, Y.; Li, X.; Cao, J.; Fan, Z.; Fan, H.; Yuan, L.; Xu, B.; et al. Generation of TALEN-mediated FH knockout rat model. Oncotarget 2016, 7, 61656–61669. [Google Scholar] [CrossRef] [Green Version]
- Marsan, E.; Ishida, S.; Schramm, A.; Weckhuysen, S.; Muraca, G.; Lecas, S.; Liang, N.; Treins, C.; Pende, M.; Roussel, D.; et al. Depdc5 knockout rat: A novel model of mTORopathy. Neurobiol. Dis. 2016, 89, 180–189. [Google Scholar] [CrossRef]
- Ma, Y.; Chen, W.; Zhang, X.; Yu, L.; Dong, W.; Pan, S.; Gao, S.; Huang, X.; Zhang, L. Increasing the efficiency of CRISPR/Cas9-mediated precise genome editing in rats by inhibiting NHEJ and using Cas9 protein. RNA Biol. 2016, 13, 605–612. [Google Scholar] [CrossRef] [Green Version]
- Goto, T.; Hara, H.; Nakauchi, H.; Hochi, S.; Hirabayashi, M. Hypomorphic phenotype of Foxn1 gene-modified rats by CRISPR/Cas9 system. Transgenic Res. 2016, 25, 533–544. [Google Scholar] [CrossRef]
- You, P.; Hu, H.; Chen, Y.; Zhao, Y.; Yang, Y.; Wang, T.; Xing, R.; Shao, Y.; Zhang, W.; Li, D.; et al. Effects of melanocortin 3 and 4 receptor deficiency on energy homeostasis in rats. Sci. Rep. 2016, 6, 34938. [Google Scholar] [CrossRef] [Green Version]
- Priestley, J.R.C.; Kautenburg, K.E.; Casati, M.C.; Endres, B.T.; Geurts, A.M.; Lombard, J.H. The NRF2 knockout rat: A new animal model to study endothelial dysfunction, oxidant stress, and microvascular rarefaction. Am. J. Physiol. Heart Circ. Physiol. 2016, 310, H478–H487. [Google Scholar] [CrossRef] [Green Version]
- Taguchi, K.; Takaku, M.; Egner, P.A.; Morita, M.; Kaneko, T.; Mashimo, T.; Kensler, T.W.; Yamamoto, M. Generation of a new model rat: Nrf2 knockout rats are sensitive to aflatoxin B1 toxicity. Toxicol. Sci. 2016, 152, 40–52. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Tang, Y.; Lu, J.; Shao, Y.; Qin, X.; Li, Y.; Wang, L.; Li, D.; Liu, M. Characterization of novel cytochrome P450 2E1 knockout rat model generated by CRISPR/Cas9. Biochem. Pharmacol. 2016, 105, 80–90. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.; Ito, Y.; Shinohara, M.; Yamashita, S.; Ichinose, S.; Kishida, A.; Oyaizu, T.; Kayama, T.; Nakamichi, R.; Koda, N.; et al. Gene targeting of the transcription factor Mohawk in rats causes heterotopic ossification of Achilles tendon via failed tenogenesi. Proc. Natl. Acad. Sci. USA 2016, 113, 7840–7845. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Q.; Kingman, J.; van de Wetering, K.; Tannouri, S.; Sundberg, J.P.; Uitto, J. Abcc6 knockout rat model highlights the role of liver in PPi homeostasis in Pseudoxanthoma elasticum. J. Investig. Dermatol. 2017, 137, 1025–1032. [Google Scholar] [CrossRef] [PubMed]
- Plaas, M.; Seppa, K.; Reimets, R.; Jagomäe, T.; Toots, M.; Koppel, T.; Vallisoo, T.; Nigul, M.; Heinla, I.; Meier, R.; et al. Wfs1-deficient rats develop primary symptoms of Wolfram syndrome: Insulin-dependent diabetes, optic nerve atrophy and medullary degeneration. Sci. Rep. 2017, 7, 10220. [Google Scholar] [CrossRef] [Green Version]
- Tian, Y.; Yang, C.; Shang, S.; Cai, Y.; Deng, X.; Zhang, J.; Shao, F.; Zhu, D.; Liu, Y.; Chen, G.; et al. Loss of FMRP impaired hippocampal long-term plasticity and spatial learning in rats. Front. Mol. Neurosci. 2017, 10, 269. [Google Scholar] [CrossRef] [Green Version]
- Peng, W.; Li, M.; Li, H.; Tang, K.; Zhuang, J.; Zhang, J.; Xiao, J.; Jiang, H.; Li, D.; Yu, Y.; et al. Dysfunction of myosin light-chain 4 (MYL4) leads to heritable atrial cardiomyopathy with electrical, contractile, and structural components: Evidence from genetically-engineered rats. J. Am. Heart Assoc. 2017, 6, e007030. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Lu, W.; Gao, N.; Long, Y.; Shao, Y.; Liu, M.; Chen, H.; Ye, S.; Ma, X.; Liu, M.; et al. Generation of obese rat model by transcription activator-like effector nucleases targeting the leptin receptor gene. Sci. China Life Sci. 2017, 60, 152–157. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Gao, Q.; Zhang, X.; Huang, L.; Xu, K.; Hu, Y.; Liu, L.; Mu, Y.; Li, K. PNPLA5-knockout rats induced by CRISPR/Cas9 exhibit abnormal bleeding and lipid level. J. Integr. Agric. 2017, 16, 169–180. [Google Scholar] [CrossRef] [Green Version]
- Harony-Nicolas, H.; Kay, M.; du Hoffmann, J.; Klein, M.E.; Bozdagi-Gunal, O.; Riad, M.; Daskalakis, N.P.; Sonar, S.; Castillo, P.E.; Hof, P.R.; et al. Oxytocin improves behavioral and electrophysiological deficits in a novel Shank3-deficient rat. eLife 2017, 6, e18904. [Google Scholar] [CrossRef]
- Cheng, X.; Waghulde, H.; Mell, B.; Morgan, E.E.; Pruett-Miller, S.M.; Joe, B. Positional cloning of quantitative trait nucleotides for blood pressure and cardiac QT-interval by targeted CRISPR/Cas9 editing of a novel long non-coding RNA. PLoS Genet. 2017, 13, e1006961. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Lai, D.H.; Wilson, R.A.; Chen, Y.F.; Wang, L.F.; Yu, Z.L.; Li, M.Y.; He, P.; Hide, G.; Sun, X.; et al. Nitric oxide blocks the development of the human parasite Schistosoma japonicum. Proc. Natl. Acad. Sci. USA 2017, 114, 10214–10219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Palygin, O.; Levchenko, V.; Ilatovskaya, D.V.; Pavlov, T.S.; Pochynyuk, O.M.; Jacob, H.J.; Geurts, A.M.; Hodges, M.R.; Staruschenko, A. Essential role of Kir5.1 channels in renal salt handling and blood pressure control. JCI Insight 2017, 2, e92331. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Shao, Y.; Qin, X.; Liu, D.; Chen, A.; Li, D.; Liu, M.; Wang, X. CRISPR knockout rat cytochrome P450 3A1/2 model for advancing drug metabolism and pharmacokinetics research. Sci. Rep. 2017, 7, 42922. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ossart, J.; Moreau, A.; Autrusseau, E.; Ménoret, S.; Martin, J.C.; Besnard, M.; Ouisse, L.H.; Tesson, L.; Flippe, L.; Kisand, K.; et al. Breakdown of immune tolerance in AIRE-deficient rats induces a severe autoimmune polyendocrinopathy–candiasis–ectodermal dystrophy–like autoimmune disease. J. Immunol. 2018, 201, 874–887. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Jiang, X.; Xu, M.; Wang, H.; Sang, X.; Qin, M.; Bao, P.; Wang, R.; Zhang, C.; Lu, H.; et al. Sleep and circadian abnormalities precede cognitive deficits in R521C FUS knockin rats. Neurobiol. Aging 2018, 72, 159–170. [Google Scholar] [CrossRef]
- Ménoret, S.; Ouisse, L.-H.; Tesson, L.; Delbos, F.; Garnier, D.; Remy, S.; Usal, C.; Concordet, J.P.; Giovannangeli, C.; Chenouard, V.; et al. Generation of immunodeficient rats with Rag1 and Il2rg gene deletions and human tissue grafting models. Transplantation 2018, 102, 1271–1278. [Google Scholar] [CrossRef]
- Izumi, R.; Kusakabe, T.; Noguchi, M.; Iwakura, H.; Tanaka, T.; Miyazawa, T.; Aotani, D.; Hosoda, K.; Kangawa, K.; Nakao, K. CRISPR/Cas9-mediated Angptl8 knockout suppresses plasma triglyceride concentrations and adiposity in rats. J. Lipid Res. 2018, 59, 1575–1585. [Google Scholar] [CrossRef] [Green Version]
- Takabayashi, S.; Aoshima, T.; Kabashima, K.; Aoto, K.; Ohtsuka, M.; Sato, M. i-GONAD (improved genome-editing via oviductal nucleic acids delivery), a convenient in vivo tool to produce genome-edited rats. Sci. Rep. 2018, 8, 12059. [Google Scholar] [CrossRef]
- Kobayashi, T.; Namba, M.; Koyano, T.; Fukushima, M.; Sato, M.; Ohtsuka, M.; Matsuyama, M. Successful production of genome-edited rats by the rGONAD method. BMC Biotechnol. 2018, 18, 19. [Google Scholar] [CrossRef] [Green Version]
- Zhao, Y.; Yang, Y.; Xing, R.; Cui, X.; Xiao, Y.; Xie, L.; You, P.; Wang, T.; Zeng, L.; Peng, W.; et al. Hyperlipidemia induces typical atherosclerosis development in Ldlr and Apoe deficient rats. Atherosclerosis 2018, 271, 26–35. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Wang, L.; Guo, N.; Wang, S.; Yang, L.; Li, Y.; Wang, M.; Yin, S.; Han, H.; Zeng, L.; et al. Cas9-nickase-mediated genome editing corrects hereditary tyrosinemia in rats. J. Biol. Chem. 2018, 293, 6883–6892. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, R.; Fang, X.; He, L.; Shao, Y.; Guo, N.; Wang, L.; Liu, M.; Li, D.; Geng, H. Generation of a primary hyperoxaluria type 1 disease model via CRISPR/Cas9 system in rats. Curr. Mol. Med. 2018, 18, 436–447. [Google Scholar] [CrossRef]
- Miyasaka, Y.; Uno, Y.; Yoshimi, K.; Kunihiro, Y.; Yoshimura, T.; Tanaka, T.; Ishikubo, H.; Hiraoka, Y.; Takemoto, N.; Tanaka, T.; et al. CLICK: One-step generation of conditional knockout mice. BMC Genom. 2018, 19, 318. [Google Scholar] [CrossRef] [Green Version]
- Sinkevicius, K.W.; Morrison, T.R.; Kulkarni, P.; Cagliostro, M.K.C.; Iriah, S.; Malmberg, S.; Sabrick, J.; Honeycutt, J.A.; Askew, K.L.; Trivedi, M.; et al. RNaseT2 knockout rats exhibit hippocampal neuropathology and deficits in memory. Dis. Models Mech. 2018, 11, dmm032631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, L.; Zhang, X.; Wang, L.; Yin, S.; Zhu, B.; Xie, L.; Duan, Q.; Hu, H.; Zheng, R.; Wei, Y.; et al. Increasing targeting scope of adenosine base editors in mouse and rat embryos through fusion of TadA deaminase with Cas9 variants. Protein Cell 2018, 9, 814–819. [Google Scholar] [CrossRef] [Green Version]
- Miller, J.J.; Aoki, K.; Moehring, F.; Murphy, C.A.; O’Hara, C.L.; Tiemeyer, M.; Stucky, C.L.; Dahms, N.M. Neuropathic pain in a Fabry disease rat model. JCI Insight 2018, 3, e99171. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fan, Z.; Li, L.; Li, X.; Zhang, M.; Zhong, Y.; Li, Y.; Yu, D.; Cao, J.; Zhao, J.; Xiaoming, D.; et al. Generation of an oxoglutarate dehydrogenase knockout rat model and the effect of a high-fat diet. RSC Adv. 2018, 8, 16636–16644. [Google Scholar] [CrossRef] [Green Version]
- Leo, D.; Sukhanov, I.; Zoratto, F.; Illiano, P.; Caffino, L.; Sanna, F.; Messa, G.; Emanuele, M.; Esposito, A.; Dorofeikova, M.; et al. Pronounced hyperactivity, cognitive dysfunctions, and BDNF dysregulation in dopamine transporter knock-out rats. J. Neurosci. 2018, 38, 1959–1972. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.; Zhang, L.; Xie, M.; Li, Y.; Huang, P.; Saunders, T.L.; Fox, D.A.; Rosenquist, R.; Lin, F. Role of complement in a rat model of paclitaxel-induced peripheral neuropathy. J. Immunol. 2018, 200, 4094–4101. [Google Scholar] [CrossRef]
- Noto, F.K.; Adjan-Steffey, V.; Tong, M.; Ravichandran, K.; Zhang, W.; Arey, A.; McClain, C.B.; Ostertag, E.; Mazhar, S.; Sangodkar, J.; et al. Sprague dawley Rag2-null rats created from engineered spermatogonial stem cells are immunodeficient and permissive to human xenografts. Mol. Cancer Ther. 2018, 17, 2481–2489. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, C.; Zhou, Y.; Xie, S.; Yin, Q.; Tang, C.; Ni, Z.; Fei, J.; Zhang, Y. CRISPR/Cas9-mediated genome editing reveals the synergistic effects of β-defensin family members on sperm maturation in rat epididymis. FASEB J. 2018, 32, 1354–1363. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, Y.; Yu, L.; Zhang, X.; Xin, C.; Huang, S.; Bai, L.; Chen, W.; Gao, R.; Li, J.; Pan, S.; et al. Highly efficient and precise base editing by engineered dCas9-guide tRNA adenosine deaminase in rats. Cell Discov. 2018, 4, 39. [Google Scholar] [CrossRef]
- Wei, Y.; Yang, L.; Zhang, X.; Sui, D.; Wang, C.; Wang, K.; Shan, M.; Guo, D.; Wang, H. Generation and characterization of a CYP2C11-null rat model by using the CRISPR/Cas9 method. Drug Metab. Dispos. 2018, 46, 525–531. [Google Scholar] [CrossRef] [PubMed]
- Dreano, E.; Bacchetta, M.; Simonin, J.; Galmiche, L.; Usal, C.; Slimani, L.; Sadoine, J.; Tesson, L.; Anegon, I.; Concordet, J.P.; et al. Characterization of two rat models of cystic fibrosis—KO and F508del CFTR—Generated by Crispr-Cas9. Anim. Models Exp. Med. 2019, 2, 297–311. [Google Scholar] [CrossRef]
- Miller, J.J.; Aoki, K.; Mascari, C.A.; Beltrame, A.K.; Sokumbi, O.; North, P.E.; Tiemeyer, M.; Kriegel, A.J.; Dahms, N.M. α-Galactosidase A-deficient rats accumulate glycosphingolipids and develop cardiorenal phenotypes of Fabry disease. FASEB J. 2019, 33, 418–429. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, T.J.; Lan, X.Y.; Wei, M.P.; Zhai, F.J.; Boeckers, T.M.; Wang, J.N.; Yuan, S.; Jin, M.Y.; Xie, Y.F.; Dang, W.W.; et al. Altered behaviors and impaired synaptic function in a novel rat model with a complete Shank3 deletion. Front. Cell. Neurosci. 2019, 13, 111. [Google Scholar] [CrossRef]
- Lambert, M.; Capuano, V.; Boet, A.; Tesson, L.; Bertero, T.; Nakhleh, M.K.; Remy, S.; Anegon, I.; Pechoux, C.; Hautefort, A.; et al. Characterization of Kcnk3-mutated rat, a novel model of pulmonary hypertension. Circ. Res. 2019, 125, 678–695. [Google Scholar] [CrossRef]
- Lee, J.G.; Ha, C.H.; Yoon, B.; Cheong, S.A.; Kim, G.; Lee, D.J.; Woo, D.C.; Kim, Y.H.; Nam, S.Y.; Lee, S.W.; et al. Knockout rat models mimicking human atherosclerosis created by Cpf1-mediated gene targeting. Sci. Rep. 2019, 9, 2628. [Google Scholar] [CrossRef] [Green Version]
- Honda, A.; Tachibana, R.; Hamada, K.; Morita, K.; Mizuno, N.; Morita, K.; Asano, M. Efficient derivation of knock-out and knock-in rats using embryos obtained by in vitro fertilization. Sci. Rep. 2019, 9, 11571. [Google Scholar] [CrossRef]
- Liang, C.; Zhao, J.; Lu, J.; Zhang, Y.; Ma, X.; Shang, X.; Li, Y.; Ma, X.; Liu, M.; Wang, X. Development and characterization of MDR1 (Mdr1a/b) CRISPR/Cas9 knockout rat model. Drug Metab. Dispos. 2019, 47, 71–79. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Emmert, A.S.; Vuong, S.M.; Shula, C.; Lindquist, D.; Yuan, W.; Hu, Y.C.; Mangano, F.T.; Goto, J. Characterization of a novel rat model of X-linked hydrocephalus by CRISPR-mediated mutation in L1cam. J. Neurosurg. 2019, 132, 945–958. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Wu, Z.; Liu, L.; Liu, J.; Wang, Y. Rat model of Cockayne syndrome neurological disease. Cell Rep. 2019, 29, 800–809.e5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kenawi, M.; Rouger, E.; Island, M.L.; Leroyer, P.; Robin, F.; Rémy, S.; Tesson, L.; Anegon, I.; Nay, K.; Derbré, F.; et al. Ceruloplasmin deficiency does not induce macrophagic iron overload: Lessons from a new rat model of hereditary aceruloplasminemia. FASEB J. 2019, 33, 13492–13502. [Google Scholar] [CrossRef] [Green Version]
- Hautefort, A.; Mendes-Ferreira, P.; Sabourin, J.; Manaud, G.; Bertero, T.; Rucker-Martin, C.; Riou, M.; Adão, R.; Manoury, B.; Lambert, M.; et al. Bmpr2 mutant rats develop pulmonary and cardiac characteristics of pulmonary arterial hypertension. Circulation 2019, 139, 932–948. [Google Scholar] [CrossRef]
- Demireva, E.Y.; Xie, H.; Flood, E.D.; Thompson, J.M.; Seitz, B.M.; Watts, S.W. Creation of the 5-hydroxytryptamine receptor 7 knockout rat as a tool for cardiovascular research. Physiol. Genom. 2019, 51, 290–301. [Google Scholar] [CrossRef]
- Scheimann, J.R.; Moloney, R.D.; Mahbod, P.; Morano, R.L.; Fitzgerald, M.; Hoskins, O.; Packard, B.A.; Cotella, E.M.; Hu, Y.C.; Herman, J.P. Conditional deletion of glucocorticoid receptors in rat brain results in sex-specific deficits in fear and coping behaviors. eLife 2019, 8, e44672. [Google Scholar] [CrossRef]
- Pettibone, J.R.; Yu, J.Y.; Derman, R.C.; Faust, T.W.; Hughes, E.D.; Filipiak, W.E.; Saunders, T.L.; Ferrario, C.R.; Berke, J.D. Knock-in rat lines with Cre recombinase at the dopamine D1 and adenosine 2a receptor loci. eNeuro 2019, 6, ENEURO.0163-19.2019. [Google Scholar] [CrossRef] [Green Version]
- Shi, Q.; Mattson, J.G.; Fahs, S.A.; Geurts, A.M.; Weiler, H.; Montgomery, R.R. The severe spontaneous bleeding phenotype in a novel hemophilia A rat model is rescued by platelet FVIII expression. Blood Adv. 2020, 4, 55–65. [Google Scholar] [CrossRef]
- Takabayashi, S.; Aoshima, T.; Kobayashi, Y.; Takagi, H.; Akasaka, E.; Sato, M. Successful i-GONAD in Brown Norway rats by modification of in vivo electroporation conditions. OBM Genet. 2020, 4, 9. [Google Scholar] [CrossRef]
- Dodge, A.; Peters, M.M.; Greene, H.E.; Dietrick, C.; Botelho, R.; Chung, D.; Willman, J.; Nenninger, A.W.; Ciarlone, S.; Kamath, S.G.; et al. Generation of a novel rat model of Angelman syndrome with a complete Ube3a gene deletion. Autism Res. 2020, 13, 397–409. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Zhao, X.M.; Liu, J.; Wang, Y.Y.; Xiong, L.L.; He, X.Y.; Wang, T.H. Complexin I knockout rats exhibit a complex neurobehavioral phenotype including profound ataxia and marked deficits in lifespan. Pflugers Arch. 2020, 472, 117–133. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Schiel, J.A.; Maksimova, E.; Strezoska, Z.; Zhao, G.; Anderson, E.M.; Wu, Y.; Warren, J.; Bartels, A.; van Brabant Smith, A.; et al. ErCas12a CRISPR-MAD7 for model generation in human cells, mice, and rats. CRISPR J. 2020, 3, 97–108. [Google Scholar] [CrossRef] [PubMed]
- Fujihara, K.; Yamada, K.; Ichitani, Y.; Kakizaki, T.; Jiang, W.; Miyata, S.; Suto, T.; Kato, D.; Saito, S.; Watanabe, M.; et al. CRISPR/Cas9-engineered Gad1 elimination in rats leads to complex behavioral changes: Implications for schizophrenia. Transl. Psychiatry 2020, 10, 426. [Google Scholar] [CrossRef]
- Langer, H.T.; Mossakowski, A.A.; Willis, B.J.; Grimsrud, K.N.; Wood, J.A.; Lloyd, K.; Zbinden-Foncea, H.; Baar, K. Generation of desminopathy in rats using CRISPR-Cas9. J. Cachexia Sarcopenia Muscle 2020, 11, 1364–1376. [Google Scholar] [CrossRef]
- Zheng, R.; Li, Y.; Wang, L.; Fang, X.; Zhang, J.; He, L.; Yang, L.; Li, D.; Geng, H. CRISPR/Cas9–mediated metabolic pathway reprogramming in a novel humanized rat model ameliorates primary hyperoxaluria type 1. Kidney Int. 2020, 98, 947–957. [Google Scholar] [CrossRef]
- Yu, Y.; Zhang, N.; Dong, X.; Fan, N.; Wang, L.; Xu, Y.; Chen, H.; Duan, W. Uricase-deficient rat is generated with CRISPR/Cas9 technique. PeerJ 2020, 8, e8971. [Google Scholar] [CrossRef]
- Nishikawa, M.; Yasuda, K.; Takamatsu, M.; Abe, K.; Okamoto, K.; Horibe, K.; Mano, H.; Nakagawa, K.; Tsugawa, N.; Hirota, Y.; et al. Generation of novel genetically modified rats to reveal the molecular mechanisms of vitamin D actions. Sci. Rep. 2020, 10, 5677. [Google Scholar] [CrossRef]
- Birket, S.E.; Davis, J.M.; Fernandez-Petty, C.M.; Henderson, A.G.; Oden, A.M.; Tang, L.; Wen, H.; Hong, J.; Fu, L.; Chambers, A.; et al. Ivacaftor reverses airway mucus abnormalities in a rat model harboring a humanized G551D-CFTR. Am. J. Respir. Crit. Care Med. 2020, 202, 1271–1282. [Google Scholar] [CrossRef]
- Bae, H.S.; Jin, Y.K.; Ham, S.; Kim, H.K.; Shin, H.; Cho, G.B.; Lee, K.J.; Lee, H.; Kim, K.M.; Koo, O.J.; et al. CRISRP/Cas9-mediated knockout of Mct8 reveals a functional involvement of Mct8 in testis and sperm development in a rat. Sci. Rep. 2020, 10, 11148. [Google Scholar] [CrossRef]
- Sanna, F.; Bratzu, J.; Serra, M.P.; Leo, D.; Quartu, M.; Boi, M.; Espinoza, S.; Gainetdinov, R.R.; Melis, M.R.; Argiolas, A. Altered sexual behavior in dopamine transporter (DAT) knockout male rats: A behavioral, neurochemical and intracerebral microdialysis study. Front. Behav. Neurosci. 2020, 14, 58. [Google Scholar] [CrossRef] [PubMed]
- Reese, R.M.; Dourado, M.; Anderson, K.; Warming, S.; Stark, K.L.; Balestrini, A.; Suto, E.; Lee, W.; Riol-Blanco, L.; Shields, S.D.; et al. Behavioral characterization of a CRISPR-generated TRPA1 knockout rat in models of pain, itch, and asthma. Sci. Rep. 2020, 10, 979. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Chen, A.; Ma, X.; Shang, X.; Zhang, Y.; Guo, Y.; Liu, M.; Wang, X. Generation and characterization of cytochrome P450 2J3/10 CRISPR/Cas9 knockout rat model. Drug Metab. Dispos. 2020, 48, 1129–1136. [Google Scholar] [CrossRef] [PubMed]
- Konishi, S.; Tanaka, N.; Mashimo, T.; Yamamoto, T.; Sakuma, T.; Kaneko, T.; Tanaka, M.; Izawa, T.; Yamate, J.; Kuwamura, M. Pathological characteristics of Ccdc85c knockout rats: A rat model of genetic hydrocephalus. Exp. Anim. 2020, 69, 26–33. [Google Scholar] [CrossRef]
- Tambini, M.D.; D’Adamio, L. Knock-in rats with homozygous PSEN1L435F Alzheimer mutation are viable and show selective γ-secretase activity loss causing low Aβ40/42 and high Aβ43. J. Biol. Chem. 2020, 295, 7442–7451. [Google Scholar] [CrossRef] [Green Version]
- Dong, W.; Zhang, L.; Sun, C.; Gao, X.; Guan, F.; Li, J.; Chen, W.; Ma, Y.; Zhang, L. Knock in of a hexanucleotide repeat expansion in the C9orf72 gene induces ALS in rats. Anim. Models Exp. Med. 2020, 3, 237–244. [Google Scholar] [CrossRef]
- Kitagawa, A.; Kizub, I.; Jacob, C.; Michael, K.; D’Alessandro, A.; Reisz, J.A.; Grzybowski, M.; Geurts, A.M.; Rocic, P.; Gupte, R.; et al. CRISPR-mediated single nucleotide polymorphism modeling in rats reveals insight into reduced cardiovascular risk associated with Mediterranean G6PD variant. Hypertension 2020, 76, 523–532. [Google Scholar] [CrossRef]
- Ma, X.; Shang, X.; Qin, X.; Lu, J.; Liu, M.; Wang, X. Characterization of organic anion transporting polypeptide 1b2 knockout rats generated by CRISPR/Cas9: A novel model for drug transport and hyperbilirubinemia disease. Acta Pharm. Sin. B 2020, 10, 850–860. [Google Scholar] [CrossRef]
- Yoshimi, K.; Oka, Y.; Miyasaka, Y.; Kotani, Y.; Yasumura, M.; Uno, Y.; Hattori, K.; Tanigawa, A.; Sato, M.; Oya, M.; et al. Combi-CRISPR: Combination of NHEJ and HDR provides efficient and precise plasmid-based knock-ins in mice and rats. Hum. Genet. 2021, 140, 277–287. [Google Scholar] [CrossRef]
- Aoshima, T.; Kobayashi, Y.; Takagi, H.; Iijima, K.; Sato, M.; Takabayashi, S. Modification of improved-genome editing via oviductal nucleic acids delivery (i-GONAD)-mediated knock-in in rats. BMC Biotechnol. 2021, 21, 63. [Google Scholar] [CrossRef]
- Koster, C.; van den Hurk, K.T.; Lewallen, C.F.; Talib, M.; Ten Brink, J.B.; Boon, C.; Bergen, A.A. The Lrat−/− rat: CRISPR/Cas9 construction and phenotyping of a new animal model for retinitis pigmentosa. Int. J. Mol. Sci. 2021, 22, 7234. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, K.; Nishikawa, M.; Okamoto, K.; Horibe, K.; Mano, H.; Yamaguchi, M.; Okon, R.; Nakagawa, K.; Tsugawa, N.; Okano, T.; et al. Elucidation of metabolic pathways of 25-hydroxyvitamin D3 mediated by CYP24A1 and CYP3A using Cyp24a1 knockout rats generated by CRISPR/Cas9 system. J. Biol. Chem. 2021, 296, 100668. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.; Kim, U.; Yang, H.; Ryu, B.; Kim, J.; Sakuma, T.; Yamamoto, T.; Park, J.H. TALEN-mediated generation of Nkx3.1 knockout rat model. Prostate 2021, 81, 182–193. [Google Scholar] [CrossRef] [PubMed]
- Noguchi, K.; Kadekawa, K.; Nishijima, S.; Sakanashi, M.; Okitsu-Sakurayama, S.; Higa-Nakamine, S.; Yamamoto, H.; Sugaya, K. Phenotypic characterization of the endocannabinoid-degrading enzyme alpha/beta-hydrolase domain 6 knockout rat. Cannabis Cannabinoid Res. 2021. [Google Scholar] [CrossRef]
- Miller, B.; Kostrominova, T.Y.; Geurts, A.M.; Sorokin, A. Double p52Shc/p46Shc rat knockout demonstrates severe gait abnormalities accompanied by dilated cardiomyopathy. Int. J. Mol. Sci. 2021, 22, 5237. [Google Scholar] [CrossRef]
- De Souza Gonçalves, L.; Sales, L.P.; Saito, T.R.; Campos, J.C.; Fernandes, A.L.; Natali, J.; Jensen, L.; Arnold, A.; Ramalho, L.; Bechara, L.; et al. Histidine dipeptides are key regulators of excitation-contraction coupling in cardiac muscle: Evidence from a novel CARNS1 knockout rat model. Redox Biol. 2021, 44, 102016. [Google Scholar] [CrossRef]
- Soga, M.; Izumi, T.; Nanchi, I.; Horita, N.; Yamamoto, M.; Kawasaki, S.; Ogawa, K.; Fujita, M.; Morioka, Y. Suppression of joint pain in transient receptor potential vanilloid 4 knockout rats with monoiodoacetate-induced osteoarthritis. Pain Rep. 2021, 6, e951. [Google Scholar] [CrossRef]
- Liu, J.; Shang, X.; Huang, S.; Xu, Y.; Lu, J.; Zhang, Y.; Liu, Z.; Wang, X. Construction and characterization of CRISPR/Cas9 knockout rat model of carboxylesterase 2a gene. Mol. Pharmacol. 2021, 100, 480–490. [Google Scholar] [CrossRef]
- Zhang, H.; Zheng, Q.; Chen-Tsai, R.Y. Establishment of a Cre-rat resource for creating conditional and physiological relevant models of human diseases. Transgenic Res. 2021, 30, 91–104. [Google Scholar] [CrossRef]
- Chesney, K.L.; Men, H.; Hankins, M.A.; Bryda, E.C. The Atg16l1 gene: Characterization of wild type, knock-in, and knock-out phenotypes in rats. Physiol. Genom. 2021, 53, 269–281. [Google Scholar] [CrossRef]
- Chen, L.; Ouyang, J.; Li, X.; Xiao, X.; Sun, W.; Li, S.; Zhou, L.; Liao, Y.; Zhang, Q. DNAH17 is essential for rat spermatogenesis and fertility. J. Genet. 2021, 100, 14. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.; Chen, X.; Guo, J.; Zhang, X.; Sun, H.; Wang, J.; Qian, X.; Li, B.; Tan, L.; Yu, L.; et al. Precision modeling of mitochondrial disease in rats via DdCBE-mediated mtDNA editing. Cell Discov. 2021, 7, 95. [Google Scholar] [CrossRef] [PubMed]
- Gordon, J.W.; Scangos, G.A.; Plotkin, D.J.; Barbosa, J.A.; Ruddle, F.H. Genetic transformation of mouse embryos by microinjection of purified DNA. Proc. Natl. Acad. Sci. USA 1980, 77, 7380–7384. [Google Scholar] [CrossRef] [Green Version]
- Clark, J.F.; Dinsmore, C.J.; Soriano, P. A most formidable arsenal: Genetic technologies for building a better mouse. Genes Dev. 2020, 34, 1256–1286. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Yang, H.; Shivalila, C.S.; Dawlaty, M.M.; Cheng, A.W.; Zhang, F.; Jaenisch, R. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 2013, 153, 910–918. [Google Scholar] [CrossRef] [Green Version]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex genome engineering using CRISPR/Cas systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef] [Green Version]
- Sato, M.; Takabayashi, S.; Akasaka, E.; Nakamura, S. Recent advances and future perspectives of in vivo targeted delivery of genome-editing reagents to germ cells, embryos, and fetuses in mice. Cells 2020, 9, 799. [Google Scholar] [CrossRef] [Green Version]
- Shao, Y.; Guan, Y.; Wang, L.; Qiu, Z.; Liu, M.; Chen, Y.; Wu, L.; Li, Y.; Ma, X.; Liu, M.; et al. CRISPR/Cas-mediated genome editing in the rat via direct injection of one-cell embryos. Nat. Protoc. 2014, 9, 2493–2512. [Google Scholar] [CrossRef]
- Ménoret, S.; Tesson, L.; Rémy, S.; Usal, C.; Thépenier, V.; Thinard, R.; Ouisse, L.H.; De Cian, A.; Giovannangeli, C.; Concordet, J.P.; et al. Gene targeting in rats using transcription activator-like effector nucleases. Methods 2014, 69, 102–107. [Google Scholar] [CrossRef]
- Tesson, L.; Remy, S.; Ménoret, S.; Usal, C.; Thinard, R.; Savignard, C.; De Cian, A.; Giovannangeli, C.; Concordet, J.P.; Anegon, I. Genome editing in rats using TALE nucleases. Methods Mol. Biol. 2016, 1338, 245–259. [Google Scholar] [CrossRef]
- Young, J.L.; Dean, D.A. Electroporation-mediated gene delivery. Adv. Genet. 2015, 89, 49–88. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, M.; Takemoto, T. Electroporation enables the efficient mRNA delivery into the mouse zygotes and facilitates CRISPR/Cas9-based genome editing. Sci. Rep. 2015, 5, 11315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qin, W.; Dion, S.L.; Kutny, P.M.; Zhang, Y.; Cheng, A.W.; Jillette, N.L.; Malhotra, A.; Geurts, A.M.; Chen, Y.G.; Wang, H. Efficient CRISPR/Cas9-mediated genome editing in mice by zygote electroporation of nuclease. Genetics 2015, 200, 423–430. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, S.; Lee, B.; Lee, A.Y.; Modzelewski, A.J.; He, L. Highly efficient mouse genome editing by CRISPR ribonucleoprotein electroporation of zygotes. J. Biol. Chem. 2016, 291, 14457–14467. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tröder, S.E.; Ebert, L.K.; Butt, L.; Assenmacher, S.; Schermer, B.; Zevnik, B. An optimized electroporation approach for efficient CRISPR/Cas9 genome editing in murine zygotes. PLoS ONE 2018, 13, e0196891. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Teixeira, M.; Py, B.F.; Bosc, C.; Laubreton, D.; Moutin, M.J.; Marvel, J.; Flamant, F.; Markossian, S. Electroporation of mice zygotes with dual guide RNA/Cas9 complexes for simple and efficient cloning-free genome editing. Sci. Rep. 2018, 8, 474. [Google Scholar] [CrossRef] [Green Version]
- Chen, S.; Sun, S.; Moonen, D.; Lee, C.; Lee, A.Y.; Schaffer, D.V.; He, L. CRISPR-READI: Efficient generation of knockin mice by CRISPR RNP electroporation and AAV donor infection. Cell Rep. 2019, 27, 3780–3789.e4. [Google Scholar] [CrossRef] [Green Version]
- Wei, J.; Gaynor, P.; Cole, S.; Brophy, B.; Oback, B.; Laible, G. Developing the laboratory conditions for bovine zygote-mediated genome editing by electroporation. In Proceedings of the World Congress on Genetics Applied to Livestock Production; 2018. Available online: http://www.wcgalp.org/system/files/proceedings/2018/developing-condition-bovine-zygote-mediated-genome-editing-electroporation.pdf (accessed on 30 January 2022).
- Tanihara, F.; Takemoto, T.; Kitagawa, E.; Rao, S.; Do, L.T.; Onishi, A.; Yamashita, Y.; Kosugi, C.; Suzuki, H.; Sembon, S.; et al. Somatic cell reprogramming-free generation of genetically modified pigs. Sci. Adv. 2016, 2, e1600803. [Google Scholar] [CrossRef] [Green Version]
- Tanihara, F.; Hirata, M.; Nguyen, N.T.; Le, Q.A.; Wittayarat, M.; Fahrudin, M.; Hirano, T.; Otoi, T. Generation of CD163-edited pig via electroporation of the CRISPR/Cas9 system into porcine in vitro-fertilized zygotes. Anim. Biotechnol. 2021, 32, 147–154. [Google Scholar] [CrossRef]
- Sato, M.; Jin, H.; Akasaka, E.; Miyoshi, M. In vitro electroporation in the presence of CRISPR/Cas9 reagents as a safe and useful method for producing biallelic knock out porcine embryos. OBM Genet. 2021, 5, 15. [Google Scholar] [CrossRef]
- Nakagawa, Y.; Sakuma, T.; Takeo, T.; Nakagata, N.; Yamamoto, T. Electroporation-mediated genome editing in vitrified/warmed mouse zygotes created by IVF via ultra-superovulation. Exp. Anim. 2018, 67, 535–543. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaneko, T.; Nakagawa, Y. Genome editing of rodents by electroporation of CRISPR/Cas9 into frozen-warmed pronuclear-stage embryos. Cryobiology 2020, 92, 231–234. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Kim, D.; Cho, S.W.; Kim, J.; Kim, J.S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 2014, 24, 1012–1019. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hakim, B.A.; Tyagi, V.; Agnihotri, S.K.; Amar Nath, A.; Agrawal, A.K.; Jain, A.; Singh, D.; Konwar, R.; Sachdev, M. Electroporation of mouse follicles, oocytes and embryos without manipulating zona pellucida. J. Dev. Biol. 2021, 9, 13. [Google Scholar] [CrossRef] [PubMed]
- Bagheri, H.; Friedman, H.; Shao, H.; Chong, Y.; Lo, C.A.; Emran, F.; Kays, I.; Yang, X.J.; Cooper, E.; Chen, B.E.; et al. TIE: A method to electroporate long DNA templates into preimplantation embryos for CRISPR-Cas9 gene editing. CRISPR J. 2018, 1, 223–229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaneko, T. Genome editing of rat. Methods Mol. Biol. 2017, 1630, 101–108. [Google Scholar] [CrossRef]
- Remy, S.; Chenouard, V.; Tesson, L.; Usal, C.; Ménoret, S.; Brusselle, L.; Heslan, J.M.; Nguyen, T.H.; Bellien, J.; Merot, J.; et al. Generation of gene-edited rats by delivery of CRISPR/Cas9 protein and donor DNA into intact zygotes using electroporation. Sci. Rep. 2017, 7, 16554. [Google Scholar] [CrossRef]
- Takahashi, G.; Gurumurthy, C.B.; Wada, K.; Miura, H.; Sato, M.; Ohtsuka, M. GONAD: Genome-editing via Oviductal Nucleic Acids Delivery system: A novel microinjection independent genome engineering method in mice. Sci. Rep. 2015, 5, 11406. [Google Scholar] [CrossRef] [Green Version]
- Ohtsuka, M.; Sato, M.; Miura, H.; Takabayashi, S.; Matsuyama, M.; Koyano, T.; Arifin, N.; Nakamura, S.; Wada, K.; Gurumurthy, C.B. i-GONAD: A robust method for in situ germline genome engineering using CRISPR nucleases. Genome Biol. 2018, 19, 25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hirose, M.; Honda, A.; Fulka, H.; Tamura-Nakano, M.; Matoba, S.; Tomishima, T.; Mochida, K.; Hasegawa, A.; Nagashima, K.; Inoue, K.; et al. Acrosin is essential for sperm penetration through the zona pellucida in hamsters. Proc. Natl. Acad. Sci. USA 2020, 117, 2513–2518. [Google Scholar] [CrossRef]
- Gurumurthy, C.B.; Sato, M.; Nakamura, A.; Inui, M.; Kawano, N.; Islam, M.A.; Ogiwara, S.; Takabayashi, S.; Matsuyama, M.; Nakagawa, S.; et al. Creation of CRISPR-based germline-genome-engineered mice without ex vivo handling of zygotes by i-GONAD. Nat. Protoc. 2019, 14, 2452–2482. [Google Scholar] [CrossRef] [PubMed]
- Ohtsuka, M.; Sato, M. i-GONAD: A method for generating genome-edited animals without ex vivo handling of embryos. Dev. Growth Differ. 2019, 61, 306–315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Namba, M.; Kobayashi, T.; Koyano, T.; Kohno, M.; Ohtsuka, M.; Matsuyama, M. GONAD: A new method for germline genome-editing in mice and rats. Dev. Growth Differ. 2021, 63, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Daya, S.; Berns, K.I. Gene therapy using adeno-associated virus vectors. Clin. Microbiol. Rev. 2008, 21, 583–593. [Google Scholar] [CrossRef] [Green Version]
- Mizuno, N.; Mizutani, E.; Sato, H.; Kasai, M.; Ogawa, A.; Suchy, F.; Yamaguchi, T.; Nakauchi, H. Intra-embryo gene cassette knockin by CRISPR/Cas9-mediated genome editing with adeno-associated viral vector. iScience 2018, 9, 286–297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoon, Y.; Wang, D.; Tai, P.W.L.; Riley, J.; Gao, G.; Rivera-Pérez, J.A. Streamlined ex vivo and in vivo genome editing in mouse embryos using recombinant adeno-associated viruses. Nat. Commun. 2018, 9, 412. [Google Scholar] [CrossRef]
- Edraki, A.; Mir, A.; Ibraheim, R.; Gainetdinov, I.; Yoon, Y.; Song, C.Q.; Cao, Y.; Gallant, J.; Xue, W.; Rivera-Pérez, J.A.; et al. A compact, high-accuracy Cas9 with a dinucleotide PAM for in vivo genome editing. Mol. Cell 2019, 73, 714–726.e4. [Google Scholar] [CrossRef] [Green Version]
- Romeo, C.; Chen, S.H.; Goulding, E.; Van Gorder, L.; Schwartz, M.; Walker, M.; Scott, G.; Scappini, E.; Ray, M.; Martin, N.P. AAV diffuses across zona pellucida for effortless gene delivery to fertilized eggs. Biochem. Biophys. Res. Commun. 2020, 526, 85–90. [Google Scholar] [CrossRef]
- Wang, D.; Niu, Y.; Ren, L.; Kang, Y.; Tai, P.W.L.; Si, C.; Mendonca, C.A.; Ma, H.; Gao, G.; Ji, W. Gene delivery to nonhuman primate preimplantation embryos using recombinant adeno-associated virus. Adv. Sci. 2019, 6, 1900440. [Google Scholar] [CrossRef]
- Cronkhite, J.T.; Norlander, C.; Furth, J.K.; Levan, G.; Garbers, D.L.; Hammer, R.E. Male and female germline specific expression of anEGFP reporter gene in a unique strain of transgenic rats. Dev. Biol. 2005, 284, 171–183. [Google Scholar] [CrossRef] [Green Version]
- Wu, Y.; Liang, D.; Wang, Y.; Bai, M.; Tang, W.; Bao, S.; Yan, Z.; Li, D.; Li, J. Correction of a genetic disease in mouse via use of CRISPR-Cas9. Cell Stem Cell 2013, 13, 659–662. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujii, W.; Kawasaki, K.; Sugiura, K.; Naito, K. Efficient generation of large-scale genome-modified mice using gRNA and CAS9 endonuclease. Nucleic Acids Res. 2013, 41, e187. [Google Scholar] [CrossRef] [PubMed]
- Brandl, C.; Ortiz, O.; Röttig, B.; Wefers, B.; Wurst, W.; Kühn, R. Creation of targeted genomic deletions using TALEN or CRISPR/Cas nuclease pairs in one-cell mouse embryos. FEBS Open Bio 2015, 5, 26–35. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Birling, M.C.; Schaeffer, L.; André, P.; Lindner, L.; Maréchal, D.; Ayadi, A.; Sorg, T.; Pavlovic, G.; Hérault, Y. Efficient and rapid generation of large genomic variants in rats and mice using CRISMERE. Sci. Rep. 2017, 7, 43331. [Google Scholar] [CrossRef] [Green Version]
- Mao, Z.; Bozzella, M.; Seluanov, A.; Gorbunova, V. Comparison of nonhomologous end joining and homologous recombination in human cells. DNA Repair 2008, 7, 1765–1771. [Google Scholar] [CrossRef] [Green Version]
- Maruyama, T.; Dougan, S.K.; Truttmann, M.C.; Bilate, A.M.; Ingram, J.R.; Ploegh, H.L. Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat. Biotechnol. 2015, 33, 538–542. [Google Scholar] [CrossRef]
- Singh, P.; Schimenti, J.C.; Bolcun-Filas, E. A mouse geneticist’s practical guide to CRISPR applications. Genetics 2015, 199, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Chu, V.T.; Weber, T.; Wefers, B.; Wurst, W.; Sander, S.; Rajewsky, K.; Kühn, R. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat. Biotechnol. 2015, 33, 543–548. [Google Scholar] [CrossRef] [Green Version]
- Song, J.; Yang, D.; Xu, J.; Zhu, T.; Chen, Y.E.; Zhang, J. RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency. Nat. Commun. 2016, 7, 10548. [Google Scholar] [CrossRef] [Green Version]
- Gerlach, M.; Kraft, T.; Brenner, B.; Petersen, B.; Niemann, H.; Montag, J. Efficient knock-in of a point mutation in porcine fibroblasts using the CRISPR/Cas9-GMNN fusion gene. Genes 2018, 9, 296. [Google Scholar] [CrossRef] [Green Version]
- Raveux, A.; Vandormael-Pournin, S.; Cohen-Tannoudji, M. Optimization of the production of knock-in alleles by CRISPR/Cas9 microinjection into the mouse zygote. Sci. Rep. 2017, 7, 42661. [Google Scholar] [CrossRef] [PubMed]
- Graf, R.; Li, X.; Chu, V.T.; Rajewsky, K. sgRNA sequence motifs blocking efficient CRISPR/Cas9-mediated gene editing. Cell Rep. 2019, 26, 1098–1103. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Labun, K.; Montague, T.G.; Gagnon, J.A.; Thyme, S.B.; Valen, E. CHOPCHOP v2: A web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res. 2016, 44, W272–W276. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.W.; Kim, S.; Kim, J.M.; Kim, J.S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 2013, 31, 230–232. [Google Scholar] [CrossRef] [PubMed]
- Mali, P.; Yang, L.; Esvelt, K.M.; Aach, J.; Guell, M.; DiCarlo, J.E.; Norville, J.E.; Church, G.M. RNA-guided human genome engineering via Cas9. Science 2013, 339, 823–826. [Google Scholar] [CrossRef] [Green Version]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420–424. [Google Scholar] [CrossRef] [Green Version]
- Nishida, K.; Arazoe, T.; Yachie, N.; Banno, S.; Kakimoto, M.; Tabata, M.; Mochizuki, M.; Miyabe, A.; Araki, M.; Hara, K.Y.; et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 2016, 353, aaf8729. [Google Scholar] [CrossRef]
- Gaudelli, N.M.; Komor, A.C.; Rees, H.A.; Packer, M.S.; Badran, A.H.; Bryson, D.I.; Liu, D.R. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 2017, 551, 464–471. [Google Scholar] [CrossRef]
- Ryu, S.M.; Koo, T.; Kim, K.; Lim, K.; Baek, G.; Kim, S.T.; Kim, H.S.; Kim, D.E.; Lee, H.; Chung, E.; et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 2018, 36, 536–539. [Google Scholar] [CrossRef]
- Liu, Z.; Lu, Z.; Yang, G.; Huang, S.; Li, G.; Feng, S.; Liu, Y.; Li, J.; Yu, W.; Zhang, Y.; et al. Efficient generation of mouse models of human diseases via ABE- and BE-mediated base editing. Nat. Commun. 2018, 9, 2338. [Google Scholar] [CrossRef] [Green Version]
- Mok, B.Y.; de Moraes, M.H.; Zeng, J.; Bosch, D.E.; Kotrys, V.; Raguram, A.; Hsu, F.; Radey, M.C.; Peterson, S.B.; Mootha, V.K.; et al. A bacterial cytidine deaminase toxin enables CRISPR-free mitochondrial base editing. Nature 2020, 583, 631–637. [Google Scholar] [CrossRef]
- Asada, H.; Kawamura, Y.; Maruyama, K.; Kume, H.; Ding, R.G.; Kanbara, N.; Kuzume, H.; Sanbo, M.; Yagi, T.; Obata, K. Cleft palate and decreased brain gamma-aminobutyric acid in mice lacking the 67-kDa isoform of glutamic acid decarboxylase. Proc. Natl. Acad. Sci. USA 1997, 94, 6496–6499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chua, S.C., Jr.; Chung, W.K.; Wu-Peng, X.S.; Zhang, Y.; Liu, S.M.; Tartaglia, L.; Leibel, R.L. Phenotypes of mouse diabetes and rat fatty due to mutations in the OB (leptin) receptor. Science 1996, 271, 994–996. [Google Scholar] [CrossRef]
- Argilés, J.M. The obese Zucker rat: A choice for fat metabolism 1968-1988: Twenty years of research on the insights of the Zucker mutation. Prog. Lipid Res. 1989, 28, 53–66. [Google Scholar] [CrossRef]
- Wu, X.; Wakamiya, M.; Vaishnav, S.; Geske, R.; Montgomery, C., Jr.; Jones, P.; Bradley, A.; Caskey, C.T. Hyperuricemia and urate nephropathy in urate oxidase-deficient mice. Proc. Natl. Acad. Sci. USA 1994, 91, 742–746. [Google Scholar] [CrossRef] [Green Version]
- Nakade, S.; Yamamoto, T.; Sakuma, T. Cas9, Cpf1 and C2c1/2/3-What’s next? Bioengineered 2017, 8, 265–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anzalone, A.V.; Randolph, P.B.; Davis, J.R.; Sousa, A.A.; Koblan, L.W.; Levy, J.M.; Chen, P.J.; Wilson, C.; Newby, G.A.; Raguram, A.; et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 2019, 576, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Li, X.; He, S.; Huang, S.; Li, C.; Chen, Y.; Liu, Z.; Huang, X.; Wang, X. Efficient generation of mouse models with the prime editing system. Cell Discov. 2020, 6, 27. [Google Scholar] [CrossRef] [PubMed]
- Barbaric, I.; Gaynor Miller, G.; Dear, T.N. Appearances can be deceiving: Phenotypes of knockout mice. Brief. Funct. Genom. 2007, 6, 91–103. [Google Scholar] [CrossRef] [Green Version]


| Method for Gene Modification | GE Tool (Mode for Gene Modification) | Rat Strain | Outcome | Target Gene | References |
|---|---|---|---|---|---|
| MI DNA or mRNA | ZFNs (indels) | SS FHH SD | Knockout (KO) rats first produced using ZFN technology with high frequency (25 to 100% disruption); showing successful germ-line transmission. | IgM Rab38 | Geurts et al. (2009) [28] |
| MI mRNA | ZFNs (indels) | F344/Stm TM/Kyo | KO rats generated as a model for X-linked severe combined immunodeficiency (X-SCID) with rates greater than 20%; showing successful germ-line transmission. | Il2rg | Mashimo et al. (2010) [29] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated to show the absence of antibody-mediated hyperacute allograft rejection; can be used for biomedical research. | IgM | Ménoret et al. (2010) [30] |
| MI mRNA | ZFNs (indels) | SS | Renin-angiotensin system-related KO rats first produced. | Ren | Moreno et al. (2011) [31] |
| MI mRNA or plasmid DNA | TALENs (indels) | Unknown | KO rats first produced using TALEN technology. | IgM | Tesson et al. (2011) [32] |
| MI mRNA/plasmid DNA | ZFN (KI) | SD LEH | Knock-in (KI) rats first produced using ZFN technology; showing successful germ-line transmission of KI phenotype. | Mdr1a Pxr | Cui et al. (2011) [33] |
| MI mRNA | ZFNs (indels) | F344/Stm WI | KO rats lacking either DNA-dependent protein kinase catalytic subunit (Prkdc) or Prkdc and interleukin 2 receptor gamma (Il2rg) generated; double knockout (DKO) F344-scid gamma (FSG) rats showed more immunocompromised phenotypes, such as the abolishment of natural killer cells. | Prkdc Il2rg | Mashimo et al. (2012) [34] |
| MI plasmid DNA | Engineered meganuclease | SD | KO immunodeficient rats first produced using meganucleases; showing decreased numbers of immature and mature T and B lymphocytes and normal natural killer cells. | Rag1 | Ménoret et al. (2013) [35] |
| MI mRNA/plasmid DNA | ZFNs (KI) | SD | LoxP-flanked (floxed) rats first produced using ZFN technology; can be useful for Cre-dependent gene disruption in vivo. | Grin1 Crhr1 Tp53 | Brown et al. (2013) [36] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | SD | KO rats first produced using CRISPR/Cas9 technology with multiple gene mutations (melanocortin 3 receptor (Mc3r) and melanocortin 4 receptor (Mc4r)) in a germ-line competent manner. | Mc3r Mc4r | Li D et al. (2013) [37] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | SD | KO rats first produced using CRISPR/Cas9 technology with multiple gene mutations (Tet methylcytosine dioxygenase 1 (Tet1), Tet methylcytosine dioxygenase 2 (Tet2) and Tet methylcytosine dioxygenase 3 (Tet3)) in a germ-line competent manner. | Tet1 Tet2 Tet3 | Li W et al. (2013) [38] |
| MI mRNAs | TALENs (indels) | WI | KO rats generated with a markedly attenuated response to a lipopolysaccharide challenge; can be used as a model for studying ethanol action and general inflammatory conditions including septic shock. | Tlr4 | Ferguson et al. (2013) [39] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated to examine the roles of aryl hydrocarbon receptor (Ahr); showing alterations to the urinary tract, including hydronephrosis and hydroureter; suggesting a role of Ahr in tissue development, homeostasis, and toxicity. | Ahr | Harrill et al. (2013) [40] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated as a model for hemophilia A; showing a prolonged activated partial thromboplastin time and clot formation time; can be useful for the study to treat hemophilia A. | F8 | Nielsen et al. (2013) [41] |
| MI mRNA | ZFNs (indels) | Unknown | KO rat lines generated as a model for autism spectrum disorder (ASD); showing abnormalities in ASD-relevant phenotypes including juvenile play, perseverative behaviors, and sensorimotor gating. | Fmrp Nlgn3 | Hamilton et al. (2013) [42] |
| In vitro EP mRNA/gRNA | ZFN (indels) TALEN (indels) CRISPR/Cas9 (indels) | F344/Stm | KO rats first produced using in vitro electroporation (EP) (called “Technique for Animal Knockout system by Electroporation (TAKE)”); in vitro EP resulted in an embryonic survival rate (91%) and a GE rate (73%). | Il2rg | Kaneko et al. (2014) [43] |
| MI mRNA/DNA donor | TALENs (KI) | SD | Homology-directed repair (HDR)-modified KI rats generated with multiple gene mutations with high efficiency (0.62%–5.13%). | Hprt1 Rosa26 Ighm | Remy et al. (2014) [44] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (KI) | SD | KI rats first generated using CRISPR/Cas9-based two-cut strategy with an efficiency up to 54%. | Trdmt1 Nes Cck | Ma et al. (2014) [45] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels, KI) | F344/Stm DA PVG | Using single-stranded oligodeoxynucleotide (ssODN) donor as templates, three recessive phenotypes (including the albino phenotype by single-nucleotide polymorphism (SNP) exchange, the non-agouti phenotype by integration of a 19-bp DNA fragment, and the hooded phenotype by removal of a retrovirus-derived insertional DNA fragment) were successfully corrected. | Tyr Asip Kit | Yoshimi et al. (2014) [46] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | SD | KO rats generated with multiple gene mutations (four genes) in a germ-line competent manner. | ApoE B2m Prf1 Prkdc | Ma et al. (2014) [47] |
| MI mRNAs | ZFNs (indels) | SD | KO rats generated to examine the function of estrogen receptor 1 (Esr1); showing infertility, due to small testes in males, and large polycystic ovaries, thread-like uteri, and poorly developed mammary glands in females; can be a new tool for investigating the pathophysiology of estrogen action. | Esr1 | Rumi et al. (2014) [48] |
| MI mRNAs/DNA donor | TALENs (KI) | SD | KI rats (carrying a pA476T mutation) first produced using TALEN technology with an efficiency of 17%. | Nr3c1 | Ponce de Leon et al. (2014) [49] |
| MI mRNA/gRNAs | CRISPR/Cas9 (indels) | WI-IM | KO rats generated as a model for Duchenne muscular dystrophy (DMD); showing a decline in muscle strength, and the emergence of degenerative/regenerative phenotypes in the skeletal muscle, heart, and diaphragm; can be useful for developing therapeutic methods to treat DMD. | Dmd | Nakamura et al. (2014) [50] |
| MI mRNA | TALENs (indels) | SD | KO rats generated as a model for DMD; showing reduction in muscle strength and a decrease in spontaneous motor activity with dilated cardiomyopathy; can be useful as an animal model of DMD. | Dmd | Larcher et al. (2014) [51] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated as a model for cystic fibrosis (CF); showing abnormalities in the ileum and increased intracellular mucus in the proximal nasal septa as well as reduced airway surface liquid and periciliary liquid depth; recapitulating many aspects of CF disease. | Cftr | Tuggle et al. (2014) [52] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated to know that the bone morphogenetic protein receptor 2 (Bmpr2) mutations are linked to pulmonary arterial hypertension (PAH); displaying an intense pulmonary vascular remodeling at 3 months; suggesting that endothelial-to-mesenchymal transition (EndoMT) is linked to alterations in BPMR2 signaling. | Bmpr2 | Ranchoux et al. (2015) [53] |
| In vitro EP mRNA/gRNA/ssODN | CRISPR/Cas9 (indels, KI) | WI | KI and KO rats produced through in vitro EP in the presence of CRISPR/Cas9 components with efficiencies of 33% and 88%, respectively. | Il2rg | Kaneko and Mashimo (2015) [54] |
| MI mRNAs | TALENs (indels) | Unknown | KO rats generated to examine the function of cold-inducible RNA-binding protein (Cirp) in heart; CIRP modulates cardiac repolarization by negatively adjusting the expression and function of Ito channels. | Cirp | Li et al. (2015) [55] |
| MI mRNAs | TALENs (indels) | SD | KO rats displayed typical dyslipidemia, although no obvious atherosclerotic lesion was noted in the enface aortas and aortic root; can be a novel model for dyslipidemia and is used in the research of atherosclerosis. | ApoE | Wei et al. (2015) [56] |
| MI protein/gRNA | CRISPR/Cas9 (KI) TALENs (KI) | SD | Cas9 protein was more efficient at HDR than Cas9 mRNA, while TALEN protein was less efficient than TALEN mRNA for inducing HDR. | Rosa26 Foxp3 Anks3 | Ménoret et al. (2015) [57] |
| MI protein/sgRNA/DNA donor | CRISPR/Cas9 (KI) | SD | A DNA cassette (composed of a green fluorescent protein (GFP) reporter sequence flanked by the two pairs of lox sites) was successfully inserted in the reverse orientation into the target (leucine-rich repeat-containing G-protein coupled receptor 5 (Lgr5)) locus, which was later inverted by Cre-mediated recombination. | Lgr5 | Wang et al. (2015) [58] |
| ES cell-mediated GE plasmid DNA | CRISPR/Cas9 (KI) | ES cells | The first production of KI rats using CRISPR/Cas9-engineered embryonic stem (ES) cells; in vitro EP of Cas9 plasmid, sgRNA plasmid, and a KI vector led to enhanced homologous recombination (HR) efficiency up to 36%. | Kat II | Yamamoto et al. (2015) [14] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated as a model for obesity; showing obesity, hyperphagia, hyperglycemia, glucose intolerance, hyperinsulinemia, and dyslipidemia, as well as a decrease in bone volume and bone mineral density of the femur; can be useful for the research on obesity and diabetes. | Lepr | Bao et al. (2015) [59] |
| SSC-mediated GE plasmid DNA | CRISPR/Cas9 (indels) | SD-derived SSCs | The first creation of spermatogonial stem cell (SSC)-derived GE KO rats; when recipient male rats were transplanted with engineered SSCs into seminiferous tubules (STs), and then mated with wild-type (WT) females, the resulting progeny harbored indels with ~10% efficiency. | Epsti1 Erbb3 | Chapman et al. (2015) [60] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated as a model for Rett syndrome (RS); showing body weight loss, anxiety tendency, and cognitive deficits; recapitulating the major symptoms of RS patients. | Mecp2 | Zhai et al. (2016) [61] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated as a model for RS; showing behavioral and motor deficits in male and female rats; can be used in RS research. | Mecp2 | Patterson et al. (2016) [62] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated as a model for hereditary tyrosinemia type I (HT1); showing major phenotypic manifestations of human HT1, including hypertyrosinemia, liver failure, and renal tubular damage; also showing remarkable liver fibrosis and cirrhosis; can be used as a model of HT1 with liver cirrhosis. | Fah | Zhang et al. (2016) [63] |
| MI mRNA/gRNA/ lssODN | CRISPR/Cas9 (KI) | F344/Stm | The ssODN-mediated KI rats were successfully generated through two-hit by gRNA and two-oligo with plasmid (2H2OP) system; can be applied to any target site with any donor vector without the need to construct homology arms, thus simplifying genome engineering in living organisms. | Tyr Thy1 Rosa26 Cyp2d Sirpa | Yoshimi et al. (2016) [64] |
| MI mRNA/gRNA/ ssODN | CRISPR/Cas9 (KI) | SD | Usage of chemically modified (phosphorothioate-modified) ssODN donors is shown to be effective in improving KI efficiency. | Cftr | Renaud et al. (2016) [65] |
| MI mRNA | TALENs (indels) | SD | Homozygous KO rats are embryonically lethal; both male and female fumarate hydratase (Fh)+/- KO rats exhibited reduced litter size; also showing hematopoietic and kidney dysfunction; Fh+/- KO rats can be useful for further functional fumarate hydratase studies. | Fh | Yu et al. (2016) [66] |
| MI mRNA | TALENs (indels) | F344 | Homozygous KO rats are embryonically lethal; DEP domain-containing 5 (Depdc5)(+/-) rats developed normally and exhibited no spontaneous electroclinical seizures, but had altered cortical neuron excitability and firing patterns; can be a model for mTORopathy. | Depdc5 | Marsan et al. (2016) [67] |
| MI protein (or mRNA)/sgRNA/donor plasmid | CRISPR/Cas9 (KI) | SD | Both Scr7 (non-homologous end joining (NHEJ) inhibitor) and Cas9 protein were used to increase KI efficiency; consequently, this combination successfully increased the HR-mediated precise modification. | Fabp2 Dbndd1 | Ma et al. (2016) [68] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | WI | KO rats generated to examine the function of forkhead box N1 (Foxn1) that is known to a critical factor for differentiation of thymic and skin epithelial cells; showing thymus deficiency and incomplete hairless. | Foxn1 | Goto et al. (2016) [69] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | SD | Mc3r and Mc4r single-KO or DKO rats generated to examine Mc3r and Mc4r more deeply; Mc3r KO rats displayed hypophagia and decreased body weight, while Mc4r KO and DKO exhibited hyperphagia and increased body weight; both Mc4r KO and DKO are good models for obesity and diabetes research. | Mc3r Mc4r | You et al. (2016) [70] |
| MI mRNA | TALENs (indels) | SD | KO rats generated to evaluate the role of reduced nuclear factor (erythroid-derived 2)-like-2 (NRF2)-regulated antioxidant defenses; suppression of NRF2 antioxidant defenses plays a role in the development of salt-induced oxidant stress, endothelial dysfunction, and microvessel rarefaction; can be used as therapeutics to ameliorate vascular oxidant stress in humans. | Nrf2 | Priestley et al. (2016) [71] |
| MI mRNA | ZFNs (indels) | F344/Stm | KO rats generated to evaluate the role of reduced NRF2-regulated antioxidant defenses were sensitive to aflatoxin B1 toxicity; can be a new model animal in toxicology. | Nrf2 | Taguchi et al. (2016) [72] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | Unknown | KO rats generated to investigate the functions of cytochrome P450 2E1 (CYP2E1) were viable and fertile and did not display any obvious physiological abnormities; can be a powerful tool for the study of CYP2E1 in the chemical metabolism, toxicity, carcinogenicity. | Cyp2e1 | Wang et al. (2016) [73] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | WI | KO rats generated to examine the function of Mohawk homeobox (Mkx) in tenogenesis; showing systemic hypoplasia of tendons and earlier heterotopic ossification of the Achilles tendon; can be used for studying tendon physiology and tissue engineering. | Mkx | Suzuki et al. (2016) [74] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated as a model for pseudoxanthoma elasticum, a heritable ectopic mineralization disorder; showing reduced levels of plasma inorganic pyrophosphate (PPi), suggesting a critical role of hepatic ATP binding cassette subtype C number 6 (ABCC6) in contributing to plasma PPi levels. | Abcc6 | Li et al. (2017) [75] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated as model of Wolfram syndrome (WS), a rare autosomal-recessive disorder characterized by juvenile-onset diabetes, optic atrophy, and hearing loss; exhibiting the core symptoms of WS. | Wfs1 | Plaas et al. (2017) [76] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats specifically impairs long-term synaptic plasticity and hippocampus-dependent learning in a manner resembling the key symptoms of Fragile X syndrome (FXS). | Fmr1 | Tian et al. (2017) [77] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | Unknown | KO rats generated to investigate the role of myosin light-chain 4 (MYL4) in atrial cardiomyopathy; showing progressive atrial cardiomyopathy; suggesting that Myl4 is as a key gene required for atrial contractile, electrical, and structural integrity. | Myl4 | Peng et al. (2017) [78] |
| MI mRNA | TALENs (indels) | SD | KO rats generated as obese rat model; showing early onset of obesity and infertility. | Lepr | Chen et al. (2017) [79] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to determine the function of patatin-like phospholipase domain containing 5 (Pnpla5); were viable, but showed a variety of abnormalities related to lipid metabolism; can be a possibly model for the treatment of cardiovascular diseases. | Pnpla5 | Liu et al. (2017) [80] |
| MI mRNA | ZFNs (indels) | SD | KO rats generated as a model for Phelan–McDermid syndrome (PMS); exhibit impaired long-term social recognition memory and attention, and reduced synaptic plasticity in the hippocampal-medial prefrontal cortex pathway. | Shank3 | Harony-Nicolas et al. (2017) [81] |
| MI mRNA/gRNA (or mRNA/gRNA/ssODN) | CRISPR/Cas9 (KI, indels) | SS | GM rats generated to assess the candidacy of a novel predicted long non-coding RNA (lncRNA) showing strong association of cardiac QT-interval; KO rats showed aberrant, short QT-intervals, and elevated blood pressure; KI rats showed the rescued phenotype. | Rffl-lnc1 | Cheng et al. (2017) [82] |
| MI mRNA | TALENs (indels) | SD | KO rats generated to examine why rats are naturally resistant to Schistosoma japonicum infection; high expression levels of inducible nitric oxide synthase (iNOS) in rats play an important role in blocking the egg-induced granuloma formation of S. japonicum; providing insights for understanding the pathogenesis of human schistosomiasis. | iNOS | Shen et al. (2017) [83] |
| MI mRNA | ZFNs (indels) | SS | KO rats generated to define the importance of Kir5.1 (encoded by potassium inwardly-rectifying channel, subfamily J, member 16 (Kcnj16)) in blood pressure control under salt-induced hypertension; exhibiting hypokalemia and reduced blood pressure, and 100% mortality within a few days triggered by salt wasting; demonstrating that Kir5.1 channels are key regulators of renal salt handling. | Kcnj16 | Palygin et al. (2017) [84] |
| ES cell-mediated GE plasmid | CRISPR/Cas9 (KI) | ES cells | KI rats derived from the CRISPR/Cas9-engineered ES cells; two clones showing non-disruptive KI of Discosoma coral red fluorescent protein (dsRed) at the Sry-box transcription factor 10 (Sox10) locus produced germ-line chimeras; showing expression of dsRed in the Sox10-expressing cells (oligodendrocyte lineage cells) in vivo. | Sox10 | Chen et al. (2017) [15] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SD | DKO rats generated to investigate cytochrome P450 3A (CYP3A) functions; were viable and fertile, and had no obvious physiological abnormities; can be a powerful tool for the study of the physiological and pharmacological roles of CYP3A, especially in drug and chemical metabolism in vivo. | Cyp3a1 Cyp3a2 | Lu et al. (2017) [85] |
| MI mRNA | ZFNs (indels) | BN | KO rats generated as a model for autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy (APECED); displaying several of the key symptoms of APECED, including alopecia, skin depigmentation, and nail dystrophy | Aire | Ossart et al. (2018) [86] |
| MI mRNA/gRNA/ssODN | CRISPR/Cas9 (KI) | SD | KI rats (carrying a Fused in sarcoma (Fus)-point mutation (R521C)) generated as a model for amyotrophic lateral sclerosis (ALS); developing adult-onset learning and memory behavioral deficits cognitive impairment; suggesting a new role of FUS in sleep and circadian regulation. | Fus | Zhang et al. (2018) [87] |
| MI mRNA | TALENs (indels) | SD | Immunodeficient rats with DKO rats generated for obtaining more profound immunosuppressed phenotype; can be useful as recipients for long-term studies concerning tissue humanization of different tissues. | Il2rg Rag1 | Ménoret et al. (2018) [88] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | F344/Stm | KO rats generated to clarify the roles of angiopoietin-like protein 8 (ANGPTL8) in glucose and lipid metabolism; showing lower body weight and fat content, associated with impaired lipogenesis in adipocytes; can be an important therapeutic target for obesity and dyslipidemia. | Angptl8 | Izumi et al. (2018) [89] |
| i-GONAD protein/gRNA (or protein/gRNA/ssODN) | CRISPR/Cas9 (indels, KI) | SD LEW BN | Improved GONAD (i-GONAD) was found useful for producing KO and KI rats with efficiencies of 62 to 97% and 9%, respectively. | Tyr Sox16 | Takabayashi et al. (2018) [90] |
| i-GONAD (rGONAD) protein/gRNA (or protein/gRNA/ssODN) | CRISPR/Cas9 (indels, KI) | WKY DA | Most suitable condition for in vivo gene delivery towards rat preimplantation embryos was assessed; i-GONAD was found useful for producing KO and KI rats with relative high efficiency. | Tyr | Kobayashi et al. (2018) [91] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | Unknown | DKO rats (lacking both apolipoprotein E (ApoE) and low-density lipoprotein receptor (Ldlr)) generated as a model for familial hypercholesterolemia; showing hypercholesterolemia, hepatosteatosis, and atherosclerosis. | ApoE Ldlr | Zhao et al. (2018) [92] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to investigate whether Cas9 nickase (Cas9n)-mediated GE can efficiently correct the HT1; tail-vein injection of adenoviral vectors carrying Cas9n, repair donor template, and sgRNA successfully corrected the phenotype of fumarylacetoacetate hydrolase (Fah) KO rats. | Fah | Shao et al. (2018) [93] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | Unknown | KO rats generated as a model for primary hyperoxaluria type 1 (PH1); showing crystalluria with abnormal phenotypes, such as a slight dilatation of renal tubules with mild fibrosis in the kidney; suggesting impairment of glyoxylic acid metabolism; can be a useful tool for the development and evaluation of drugs and therapeutics. | Agxt | Zheng et al. (2018) [94] |
| In vitro EP (protein/gRNAs/lssDNA) | CRISPR/Cas9 (KI) | F344 | Efficient approaches enabling the quick generation of floxed alleles in mice and rats were taken using in vitro EP in the presence of Cas9 protein, gRNAs, and long single-stranded DNA (lssDNA); homozygous KI oocytes carrying tissue-specific Cre gene were successfully obtained with high efficiency. | Vapb | Miyasaka et al. (2018) [95] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated as a model for RNase T2 deficiency; showing no evidence of cystic lesions, but hippocampal neuroinflammation, altered lysosomal function, and cognitive defects; can be a useful model to study the RNase T2 function. | RNase T2 | Sinkevicius et al. (2018) [96] |
| MI mRNA/sgRNA | ABEs (indels) | Unknown | KO rats generated as a model for Glycogen storage disease type II (GSDII) with an efficiency of 85%; the adenine base editor (ABE) system is a powerful and convenient tool to introduce precise base conversions in rodents. | Gaa | Yang et al. (2018) [97] |
| MI plasmid DNA | CRISPR/Cas9 (indels) | DA | KO rats generated as a model for Fabry disease (FD); showing substantial serum and tissue accumulation of α-galactosyl glycosphingolipids and had pronounced mechanical pain behavior. | α-Gal A | Miller et al. (2018) [98] |
| MI mRNA | TALENs (indels) | SD | Oxoglutarate dehydrogenase (Ogdh)−/− rats are lethal; Ogdh+/− rats had higher body weight; treatment of Ogdh+/− rats with a high-fat diet resulted in liver dysfunction; Ogdh+/− rats can be used as model for investigations of metabolic syndrome and obesity-related hepatic carcinogenesis. | Ogdh | Fan et al. (2018) [99] |
| MI mRNA | ZFNs (indels) | WI | KO rats developed normally, but exhibited spontaneous locomotor hyperactivity and cognitive dysfunctions; can be a model for human diseases involving aberrant dopamine functions. | Dat | Leo et al. (2018) [100] |
| MI plasmid | CRISPR/Cas9 (indels) | F344 SD | KO rats generated to know the role of complement in chemotherapy-induced peripheral neuropathy (CIPN); suggesting that complement may be a new target for the development of novel therapeutics to manage CIPN. | C3 | Xu et al. (2018) [101] |
| SSC-mediated GE plasmid | TALENs (indels) | SD-derived SSCs | SSCs-derived GE KO rats showed immunodeficiency with lacked mature B and T cells; allowing growth of human tumors; can be used for xenograft studies. | Rag2 | Noto et al. (2018) [102] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | Unknown | Multiple-gene KO rats generated to examine the function of defensin beta (Defb) expressed abundantly in the epididymis; showing subfertility and decreased motility of sperm; suggesting that Defb family members affect sperm maturation by a synergistic pattern in the epididymis. | Defb23 Defb26 Defb42 | Zhang et al. (2018) [103] |
| MI mRNA/sgRNA | ABE (indels) | SD | For hemogen (Hemgn) targeting, out of 15 rats obtained, 14 contained an A to G conversion at the 14th base distal from the protospacer adjacent motif (PAM), indicating the high base-editing efficiency. | Hemgn Ndst1 Ndst4 | Ma et al. (2018) [104] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to assess the roles of cytochrome P450, subfamily 2, polypeptide 11 (CYP2C11); were viable and had no obvious abnormalities, with the exception of reduced fertility; can be a valuable tool to study the in vivo function of CYP2C11. | Cyp2c11 | Wei et al. (2018) [105] |
| MI mRNA/gRNA (or mRNA/gRNA ssODN) | CRISPR/Cas9 (indels, KI) | Unknown | KI (carrying F508del mutation) and KO rats generated as a model for CF; both rats exhibit CF phenotypic anomalies such as vas deferens agenesis and tooth enamel defects; can be used as a novel resource to advance the development of CF therapeutics. | Cftr | Dreano et al. (2019) [106] |
| MI plasmid DNA | CRISPR/Cas9 (indels) | DA | KO rats generated as a model for FD; showing pronounced renal tubule dysfunction and mitral valve thickening; can be further used to study disease mechanisms and test therapies. | α-Gal A | Miller et al. (2019) [107] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated a model for ASDs; showing impaired social memory (but not impaired social interaction behaviors), impaired learning and memory, increased anxiety-like behavior, increased mechanical pain threshold, and decreased thermal sensation. | Shank3 | Song et al. (2019) [108] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated as a model for pulmonary hypertension (PH); showing PH pathogenesis associated with low serum-albumin concentration; can be used for understanding the mechanisms of PH and testing therapeutic molecules. | Kcnk3 | Lambert et al. (2019) [109] |
| MI mRNA/sgRNA | CRISPR-Cpf1 (indels) | SD | ApoE KO rats displayed hyperlipidemia and aortic lesions; showing that the Cpf1 system can target single or multiple genes efficiently; these rats can be helpful for understanding initial-stage atherosclerosis. | ApoE Ldlr | Lee et al. (2019) [110] |
| In vitro EP (protein/sgRNA/plasmid DNA) rAAV | CRISPR/Cas9 (KI, indels) | BN | Superovulation is successfully conducted to Brown Norway (BN), which is a very difficult rat strain to superovulate; in vitro EP or infection with recombinant adeno-associated virus (rAAV) enables KO or KI in the in vitro fertilized (IVF) rat embryos efficiently. | Tyr | Honda et al. (2019) [111] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats exhibited complete loss of multidrug resistance protein 1 (MDR1) in the liver, small intestine, brain, and kidney; can be useful for studying the function of MDR1 in drug absorption, tumor multidrug, and drug target validation. | Mdr1a/b | Liang et al. (2019) [112] |
| MI protein/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated as a model for X-linked hydrocephalus (XLH); hemizygous male mutants developed hydrocephalus and delayed development. | L1cam | Emmert et al. (2019) [113] |
| MI mRNA/sgRNA/ssODN | CRISPR/Cas9 (KI) | SD | KI rats (carrying a R571X mutation) generated to mimic Cockayne syndrome (CS); showing atrophy and dysmyelination in the cerebellar cortex; RNA-seq suggests that transcription dysregulation could contribute to the CS pathogenesis. | Csb | Xu et al. (2019) [114] |
| MI protein/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to understand the mechanisms leading to iron excess in HA; showing decreased iron concentration and transferrin saturation; suggesting that the reported role of ceruloplasmin (Cp) cannot fully explain the iron hepatosplenic phenotype in hereditary aceruloplasminemia (HA). | Cp | Kenawi et al. (2019) [115] |
| MI mRNA | ZFNs | SD | KO rats generated as a model for PAH; showing age-dependent spontaneous PAH with a low penetrance (16%-27%), similar to that in humans; can be a promising tool to study the pathogenesis of PAH. | Bmpr2 | Hautefort et al. (2019) [116] |
| In vitro EP protein/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to examine the role of 5-hydroxytryptamine receptor 7 (Htr7) in blood pressure regulation; neither the male nor female KO rats were different in body size, fat weights, or mass of organs (kidney, heart, and brain) important to blood pressure; can be used to investigate the importance of Htr7 in blood pressure regulation. | Htr7 | Demireva et al. (2019) [117] |
| MI mRNA/two sgRNAs/donor plasmid containing two flanking loxP sites | CRISPR/Cas9 (KI) | SD | KI rats generated to create a conditional glucocorticoid receptors (GR) knockdown using a dual sgRNA strategy; affording high-precision knockdown of GR across a variety of contexts, ranging from neuronal depletion to circuit-wide manipulations. | Nr3c1 | Scheimann et al. (2019) [118] |
| MI mRNA/ sgRNA/donor plasmid | CRISPR/Cas9 (KI) | LE | Two KI rat lines (D1-Cre and A2a-Cre) generated for Cre-mediated gene manipulation; will be used to study both normal brain functions and neurological and psychiatric pathophysiology. | Drd1a Adora2a | Pettibone et al. (2019) [119] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SS | KO rats generated as a model for hemophilia; showing spontaneous bleeding in the soft tissue, muscles, or joints occurred in 100% of factor 8 (F8)-/- rats; bone marrow reconstitution rescued the spontaneous bleeding in rat HA hemophilia. | F8 | Shi et al. (2020) [120] |
| i-GONAD protein/gRNAs | CRISPR/Cas9 (indels) | BN | i-GONAD under EP conditions with current values ranging from 150 to 200 mA enabled production of KO rats with efficiencies ranging from 75% to 24%. | Tyr | Takabayashi et al. (2020) [121] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated as a model for Angelman syndrome (AS); showing deficits in motor coordination as well as learning and memory, phenotypically mirroring human AS; can be a model for the study of AS. | Ube3a | Dodge et al. (2020) [122] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to explore the function of complexin I (Cplx1); showing a complex neurobehavioral phenotype including profound ataxia, dystonia, movement and exploratory deficits, and increased anxiety and sensory deficits, but had normal cognitive function. | Cplx1 | Xu et al. (2020) [123] |
| MI mRNA (or protein)/gRNAs/plasmid donor | Cas12a/Cpf1 (MAD7) (indels, KI) | SD | MAD7 is capable of generating indels, small DNA insertion, and fluorescent gene tagging of endogenous genes, such as Cas9; causing indels with 20% efficiency in rats; MAD7 can expand the CRISPR toolbox for genome engineering. | Calb2 | Liu et al. (2020) [124] |
| In vitro EP mRNA/gRNA (or mRNA/gRNA/ lssDNA) | CRISPR/Cas9 (indels, KI) | LE | KO rats generated to know the role of glutamate decarboxylase 1 (GAD1) deficiency in pathogenesis of schizophrenia; showing complex behavioral changes, such as hypoactivity in a novel environment and decreased preference for social novelty; can be a model for studying the mechanism of schizophrenia. | Gad1 | Fujihara et al. (2020) [125] |
| MI protein/sgRNA/ssODN | CRISPR/Cas9 (KI) | SD | KI rats (carrying a R350P mutation) generated as a model for desminopathy; showing the phenotype of desminopathy; can be a useful tool for understanding of the disease and testing therapeutic approaches to delay disease progression. | Des | Langer et al. (2020) [126] |
| MI mRNA/sgRNA/ssODN | CRISPR/Cas9 (KI) | SD | KI rats (carrying a D205N mutation) generated as a model for primary hyperoxaluria type 1 (PH1); showing hyperoxaluria at 1 month of age and exhibited severe renal calcium oxalate deposition after ethylene glycol challenge; can be a useful model for understanding the disease and developing therapeutics. | Agxt | Zheng et al. (2020) [127] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated a model for hyperuricemia; were apparently healthy with more than a 95% survival up to one year and showed renal abnormality such as mild glomerular/tubular lesions; can be useful for studying hyperuricemia. | Uox | Yu et al. (2020) [128] |
| MI mRNA/sgRNA (or mRNA/sgRNA/ssODN) | CRISPR/Cas9 (KI, indels) | WI | Cytochrome P450 family 27 subfamily B member 1 (Cyp27b1) KO, vitamin D receptor (Vdr) KO, and Vdr KI (carrying a R270L mutation) rats generated as a model for human type II rickets; showing growth retardation and abnormal bone formation; can be useful for elucidating the molecular mechanism of vitamin D action. | Cyp27b1 Vdr | Nishikawa et al. (2020) [129] |
| MI mRNA/donor DNA | ZFNs (KI) | SD | KI rats (carrying a G551D variant) generated as a model for CF; showing that the epithelia recapitulates the expected absence of cystic fibrosis transmembrane conductance regulator (CFTR) activity, which was restored with ivacaftor, a medication used to treat CF. | Cftr | Birket et al. (2020) [130] |
| In vitro EP protein/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to investigate the role of monocarboxylate transporter 8 (MCT8) (encoded by solute carrier family 16 member 2 coding for monocarboxylate transporter 8 (Slc16a2)) during spermatogenesis; showing growth delay during postnatal development with reduced sperm motility and viability; suggesting the role of MCT8 in spermatogenesis. | Slc16a2 | Bae et al. (2020) [131] |
| MI mRNA | ZFNs (indels) | WI | KO rats generated to examine whether dopamine plays a key role in sexual behavior; showing more rapid acquisition of stable sexual activity levels and to higher levels of sexual motivation and activity; can be confirmed that dopamine has a key role in sexual behavior. | Dat | Sanna et al. (2020) [132] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to enable more sophisticated modeling of pain, itch, and asthma; showing apparently normal behavioral responses to pain and itch, although transient receptor potential family member ankyrin 1 (TRPA1)-dependent immune cell infiltration into the lung was seen under the asthma-inducing condition; useful for studying that TRPA1 can be as a drug target. | Trpa1 | Reese et al. (2020) [133] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to study the function of cytochrome P450 family 2 subfamily J (Cyp2J) isoform genes in vivo; were viable and fertile; can be a useful tool to study the function of Cyp2J in drug metabolism and cardiovascular disease. | Cyp2J2 Cyp2J3 Cyp2J10 | Lu et al. (2020) [134] |
| MI mRNA | TALENs (indels) | F344 | KO rats generated to study the role of coiled-coil domain containing 85C (Ccdc85c) in the pathogenesis of spontaneous hydrocephalus shown in hemorrhagic hydrocephalus (hhy) mice; showing non-obstructive hydrocephalus, subcortical heterotopia, and intracranial hemorrhage; suggesting a role of Ccdc85c in cerebral development. | Ccdc85c | Konishi et al. (2020) [135] |
| MI mRNA/gRNA/ssODN | CRISPR/Cas9 (KI) | LE | KI rats (carrying a Psen1LF mutation) generated to determine potential early pathogenic changes caused by this mutation; survived into adulthood; showing increased levels of Aβ43, a longer and potentially more amyloidogenic Aβ form. | Psen1 | Tambini and D’Adamio (2020) [136] |
| MI protein/sgRNAs/donor DNA | CRISPR/Cas9 (KI) | SD | KI rats (carrying GGGGCC (G4C2) repeats) generated as a model for ALS; showing motor deficits from 4 months of age; can be used for investigating the neurotoxicity in chromosome 9 open reading frame 72 (C9orf72)-related ALS. | C9orf72 | Dong et al. (2020) [137] |
| MI protein/sgRNAs/donor DNA | CRISPR/Cas9 (KI) | SD | KI rats generated as a model for the Mediterranean SNP G6PDS188F; showing glucose-6-phosphate dehydrogenase (G6PD) activity, but not expression, was reduced to 20% of WT littermates; replacement of a single amino acid (S188F) in G6PD, a rate-limiting enzyme in the pentose phosphate pathway, may be related to pathogenesis of vascular diseases. | G6pd | Kitagawa et al. (2020) [138] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to examine the role of solute carrier organic anion transporter family member 1B2 (Slco1b2); in those rats, the serum levels of bilirubin and bile acids, the substrates of organic anion transporting polypeptide organic anion transport polypeptide 1b2 (OATP1B2), increased; can be a disease model to study hyperbilirubinemia-related diseases. | Slco1b2 | Ma et al. (2020) [139] |
| MI, in vitro EP mRNA/sgRNAs/dsDNA donor vectors (or mRNA/sgRNAs/lssDNA) | CRISPR/Cas9 (KI) | F344 | A new powerful method, called Combi-CRISPR, was developed for achieving plasmid-based KI in rat embryos using the CRISPR/Cas9 system; in other words, this method is the combination of highly efficient editing via NHEJ and the low-efficiency but precise editing via HDR. | Pvalb Th | Yoshimi et al. (2021) [140] |
| i-GONAD protein/gRNA/ssODN | CRISPR/Cas9 (KI) | SD | Effects of 3 gRNAs (which recognize different portions of the target locus, but also overlap each other in the target locus) and of commercially available KI-enhancing drugs on the KI efficiency; KI efficiency largely depends on the type of gRNA used; none of drugs are significantly effective for KI efficiency. | Tyr | Aoshima et al. (2021) [141] |
| MI protein/gRNA (or protein/gRNA/ssODN) | CRISPR/Cas9 (indels, KI) | BN | KO rats generated as a model for retinal dystrophy (RD); showing reduced vision and structural abnormalities, such as overall retinal thinning; can be new animal model for retinal dystrophy. | Lrat | Koster et al. (2021) [142] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | WI | KO rats generated to investigate cytochrome P450 family 24 subfamily A member 1 (CYP24A1)-dependent or -independent metabolism of 25(OH)D3, the prohormone of calcitriol; found to be valuable for metabolic studies of vitamin D and its analogs. | Cyp24a1 | Yasuda et al. (2021) [143] |
| In vitro EP mRNA | TALENs (indels) | SD | KO rats generated to elucidate the functions of NK3 homeobox 1 (Nkx3.1); showing reduced fertility, decreased prostate weights, and increased epithelial cell layers; can be a model for studying the role of Nkx3.1 in decreased prostate weights, fertility, and breast cancer, as well as in prostate cancer. | Nkx3.1 | Lee et al. (2021) [144] |
| MI mRNA/sgRNA | CRISPR/Cas9 (indels) | Unknown | KO rats generated to elucidate the roles of alpha/beta-hydrolase domain 6 (ABHD6) in vivo; showing normal appearance, but caused more frequent urination in the stimulated bladder. | Abhd6 | Noguchi et al. (2021) [145] |
| MI plasmid | CRISPR/Cas9 (indels) | SS | KO rats lacking adaptor protein 1 (shc1), encodes 3 main protein isoforms (p52Shc and p46Shc isoforms) generated to further investigate the function of Shc1; showing severe gait abnormalities accompanied by dilated cardiomyopathy. | p52Shc p46Shc | Miller et al. (2021) [146] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | WI | KO rats generated to investigate the in vivo roles of carnosine synthase 1 (Carns1); showing a significant impairment of contractile function in the cardiac muscle; suggesting a role of high-cholesterol diet (HCD) in the regulation of Ca2+ handling and excitation–contraction coupling of cardiac muscle. | Carns1 | de Souza Gonçalves et al. (2021) [147] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | SD | KO rats generated to examine the function of transient receptor potential cation channel subfamily V member 4 (Trpv4) in regulating osteoarthritic pain; showing suppression of joint pain under the monoiodoacetate-induced osteoarthritic pain model; suggesting that inhibition of TRPV4 might be a novel potent analgesic strategy for treating osteoarthritic pain. | Trpv4 | Soga et al. (2021) [148] |
| MI mRNA/sgRNAs | CRISPR/Cas9 (indels) | SD | KO rats generated to examine the role of carboxylesterase 2A (Ces2a) in drug metabolism mediated by carboxylesterase 2 (CES2); showing obesity, insulin resistance, and liver fat accumulation, which are consistent with the symptoms of nonalcoholic fatty liver disease (NAFLD); can be used as a model for NAFLD. | Ces2a | Liu et al. (2021) [149] |
| MI protein/two plasmids (one containing gRNA and the other containing Cre-ERT2-mCherry cassette as donor plasmid) | CRISPR/Cas9 (KI) | SD | Generation of six Cre driver rats allowing for the controlled gene expression or conditional KO in a temporal and spatial manner through the Cre-ERT2/loxP system. | Wnt1 Pomc Mnx1 Drd1a Gad67 Tie2 | Zhang et al. (2021) [150] |
| MI mRNA/gRNA/ssODN | CRISPR/Cas9 (KI) | SD F344 | KI rats (carrying a T300A mutation) generated as a model for Crohn’s disease (CD); showing morphological abnormalities in both Paneth and goblet cells, but do not develop spontaneous intestinal permeability or inflammatory bowel disease; can be used for the study of both autophagy and CD susceptibility. | Atg16l1 | Chesney et al. (2021) [151] |
| MI mRNA/gRNA | CRISPR/Cas9 (indels) | SD | KO rats generated to examine the function of dynein axonemal heavy chain 17 (Dnah17) in spermatogenesis; showing infertility with significantly decreased number of sperm; DNAH17 is critical for spermatogenesis. | Dnah17 | Chen et al. (2021) [152] |
| MI PB vector carrying DdCBE pair/PB transposase mRNA | DdCBE (gene correction) | SD | DddA-derived cytosine base editor (DdCBE) was applied to explore the possible production of GE rats as a mitochondrial disease model with pathogenic mitochondrial DNA (mtDNA) mutations; MI of piggyBack transposon (PB) vector carrying DdCBE pair with PB transposase mRNA resulted in production of pups with mtDNA mutations with 36% efficiency. | G7755 | Qi et al. (2021) [153] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sato, M.; Nakamura, S.; Inada, E.; Takabayashi, S. Recent Advances in the Production of Genome-Edited Rats. Int. J. Mol. Sci. 2022, 23, 2548. https://doi.org/10.3390/ijms23052548
Sato M, Nakamura S, Inada E, Takabayashi S. Recent Advances in the Production of Genome-Edited Rats. International Journal of Molecular Sciences. 2022; 23(5):2548. https://doi.org/10.3390/ijms23052548
Chicago/Turabian StyleSato, Masahiro, Shingo Nakamura, Emi Inada, and Shuji Takabayashi. 2022. "Recent Advances in the Production of Genome-Edited Rats" International Journal of Molecular Sciences 23, no. 5: 2548. https://doi.org/10.3390/ijms23052548







