Genome Integrity and Neurological Disease
Abstract
:1. Nervous Systems and the Brain
2. Neurodegenerative Diseases
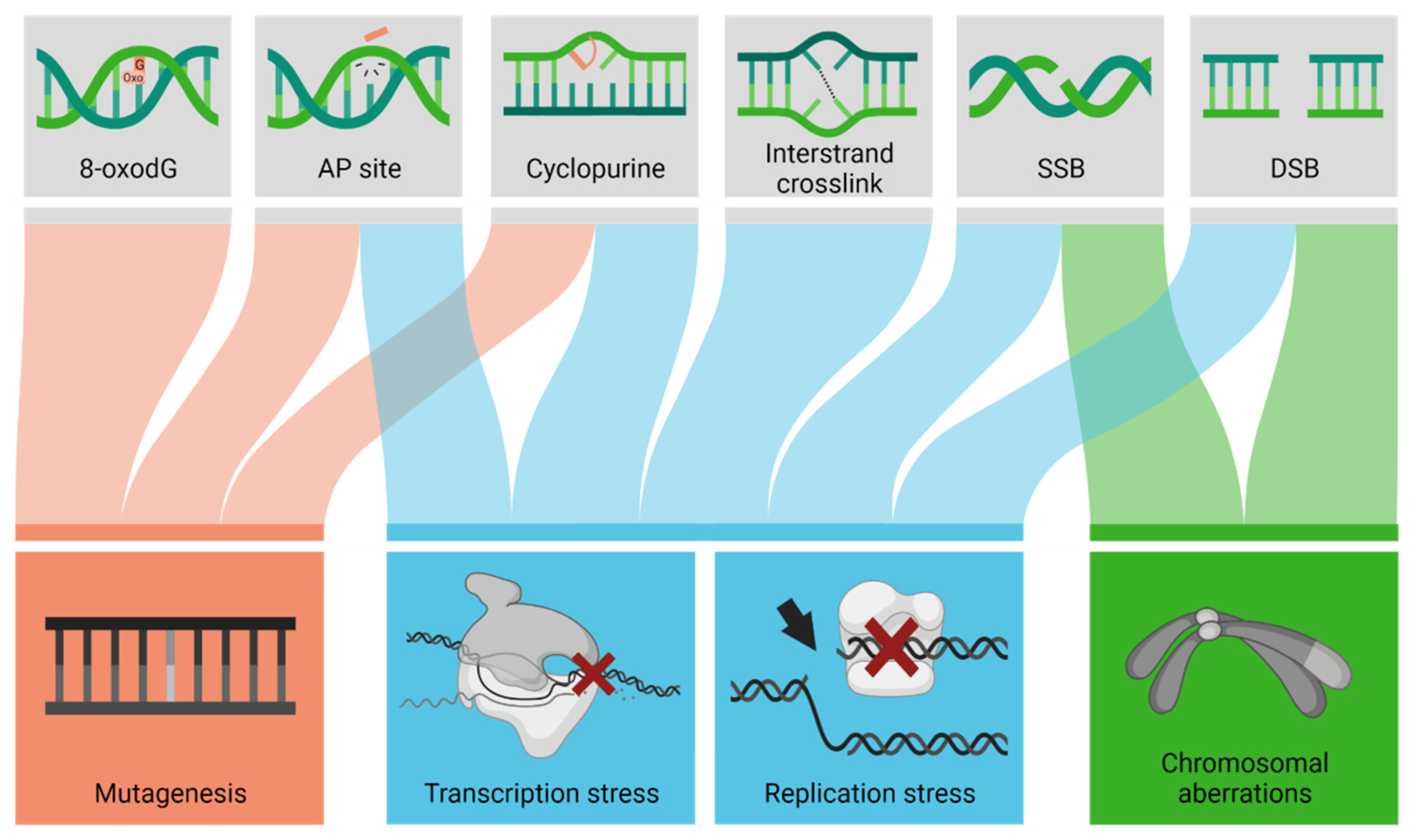
3. Exogenous and Endogenous Threats to the Brain: Oxidative DNA Damage
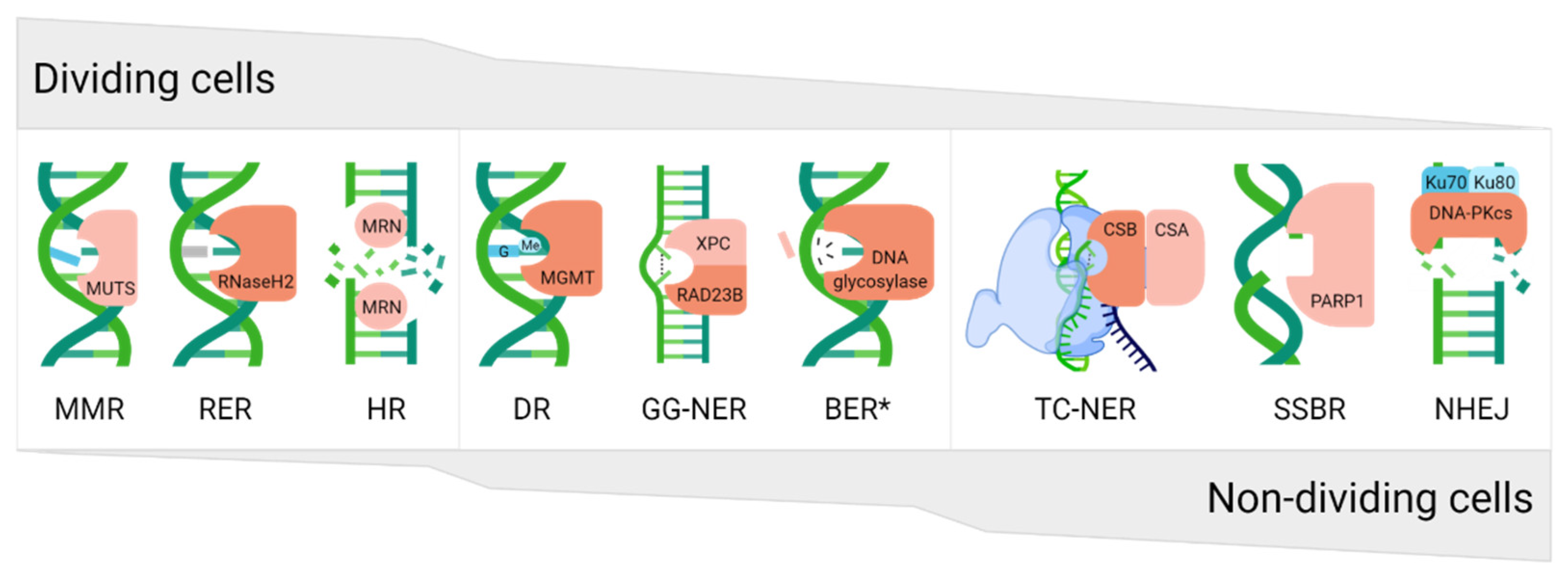
4. DNA Repair Mechanisms
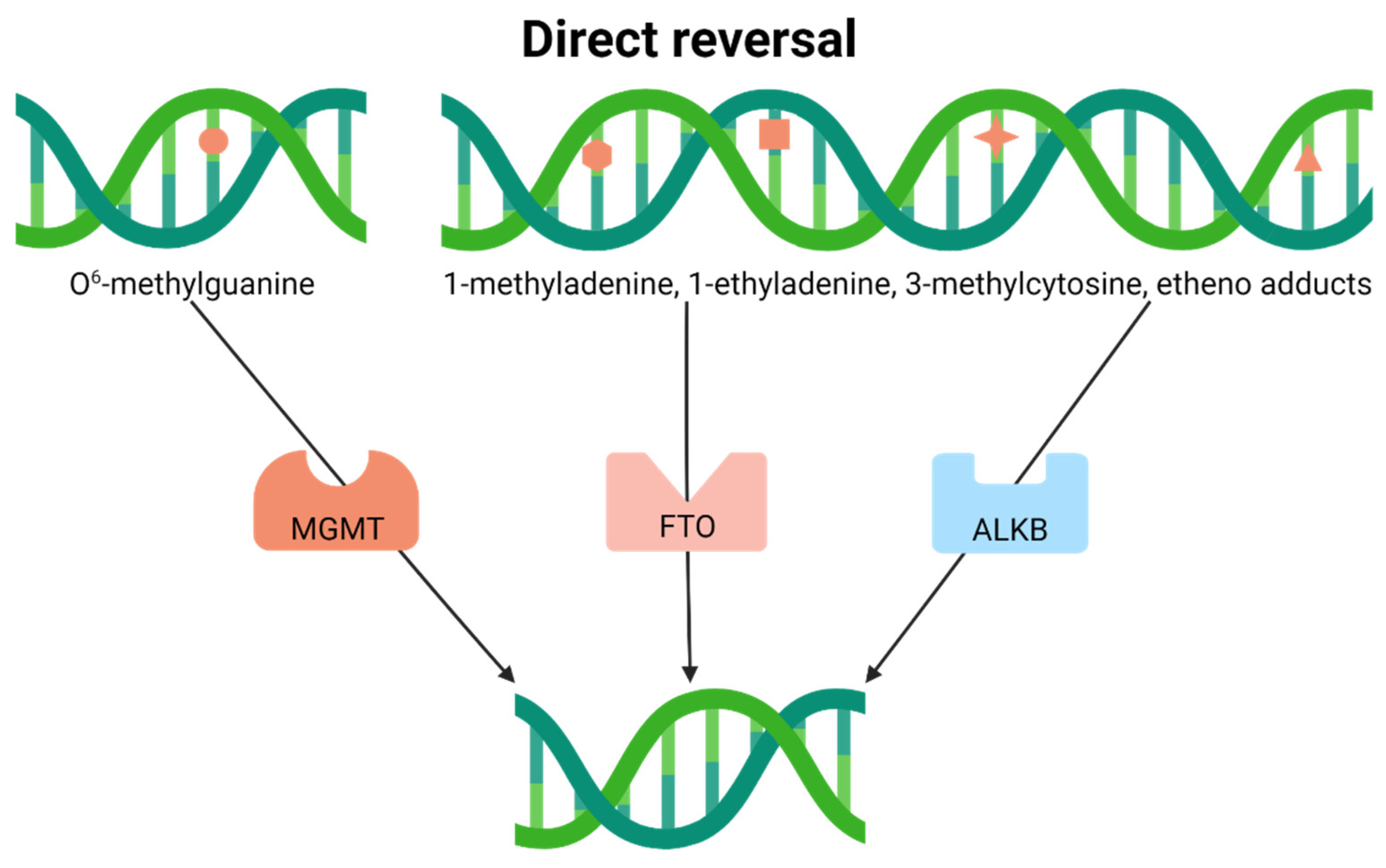
4.1. Direct Reversal
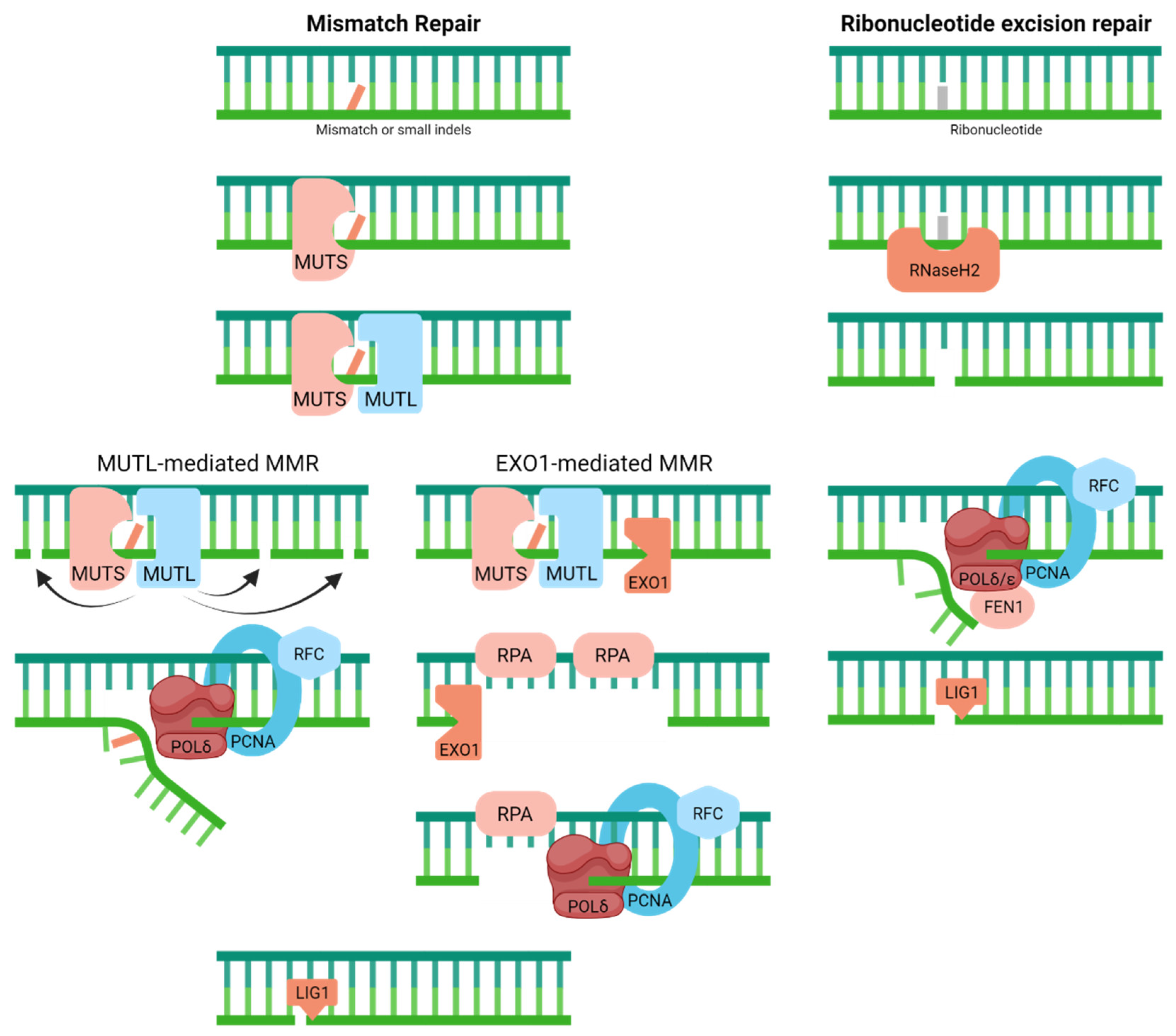
4.2. Mismatch Repair
4.3. Ribonucleotide Excision Repair
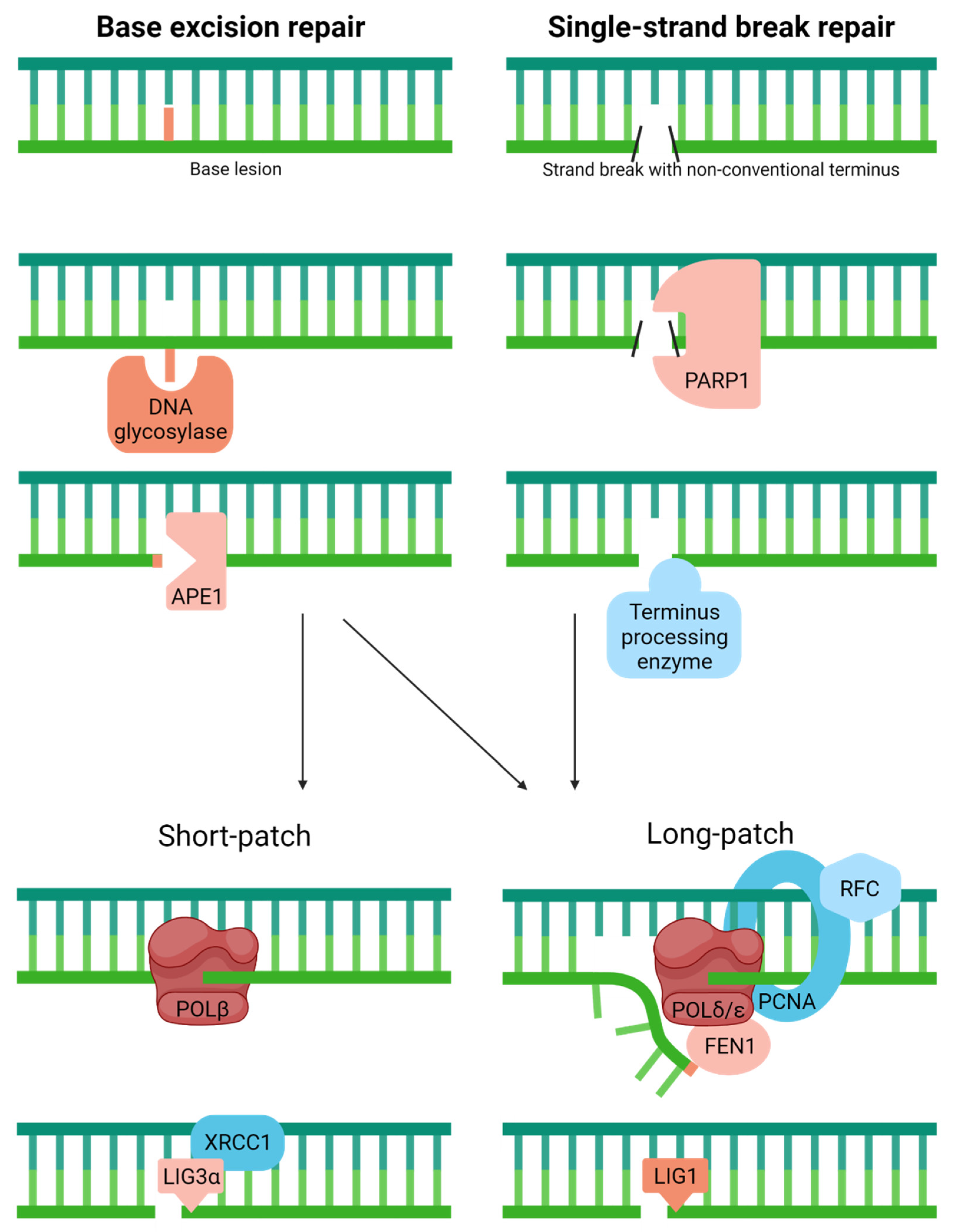
4.4. Base Excision Repair
4.5. Single-Strand Break Repair
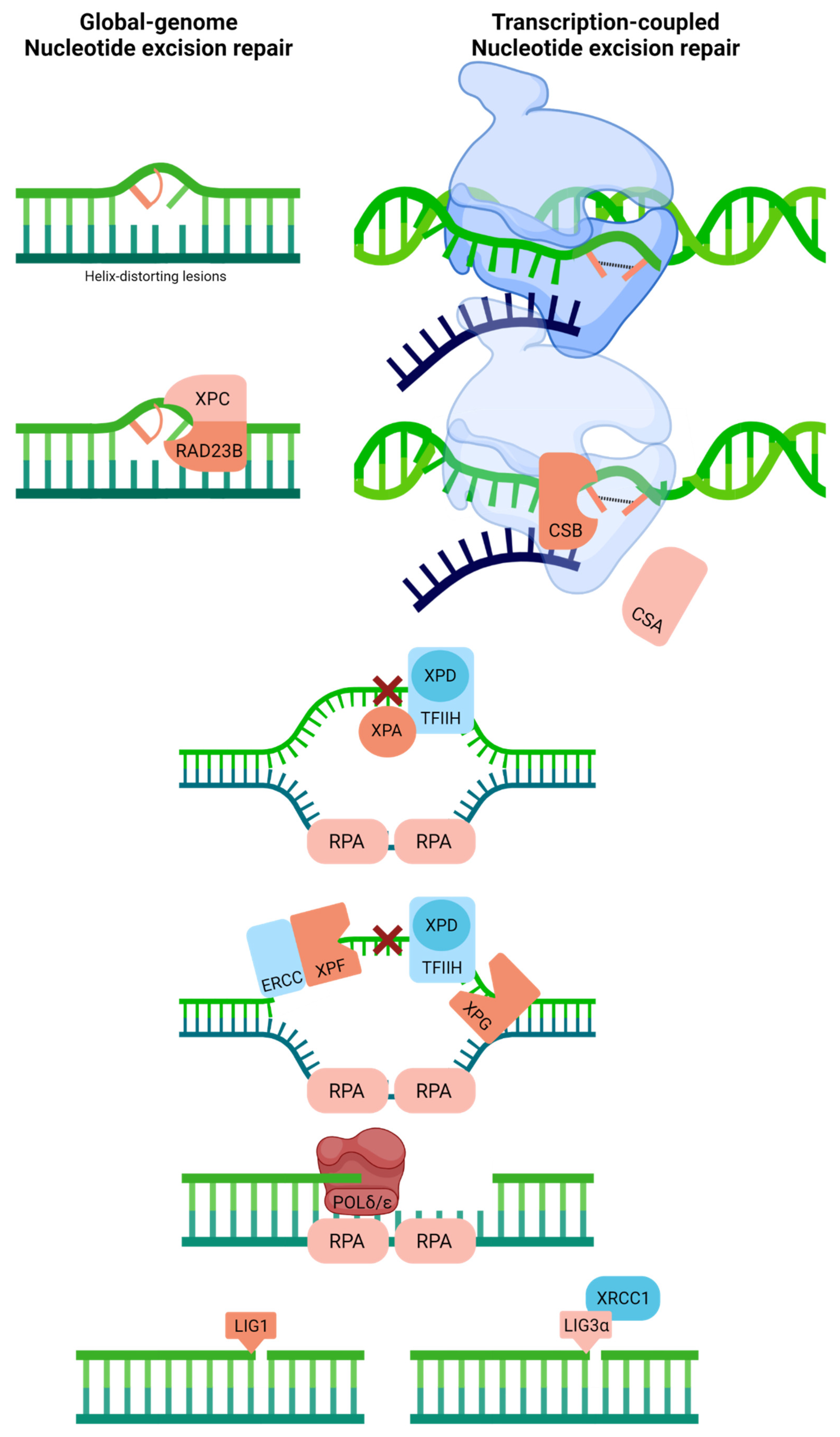
4.6. Nucleotide Excision Repair
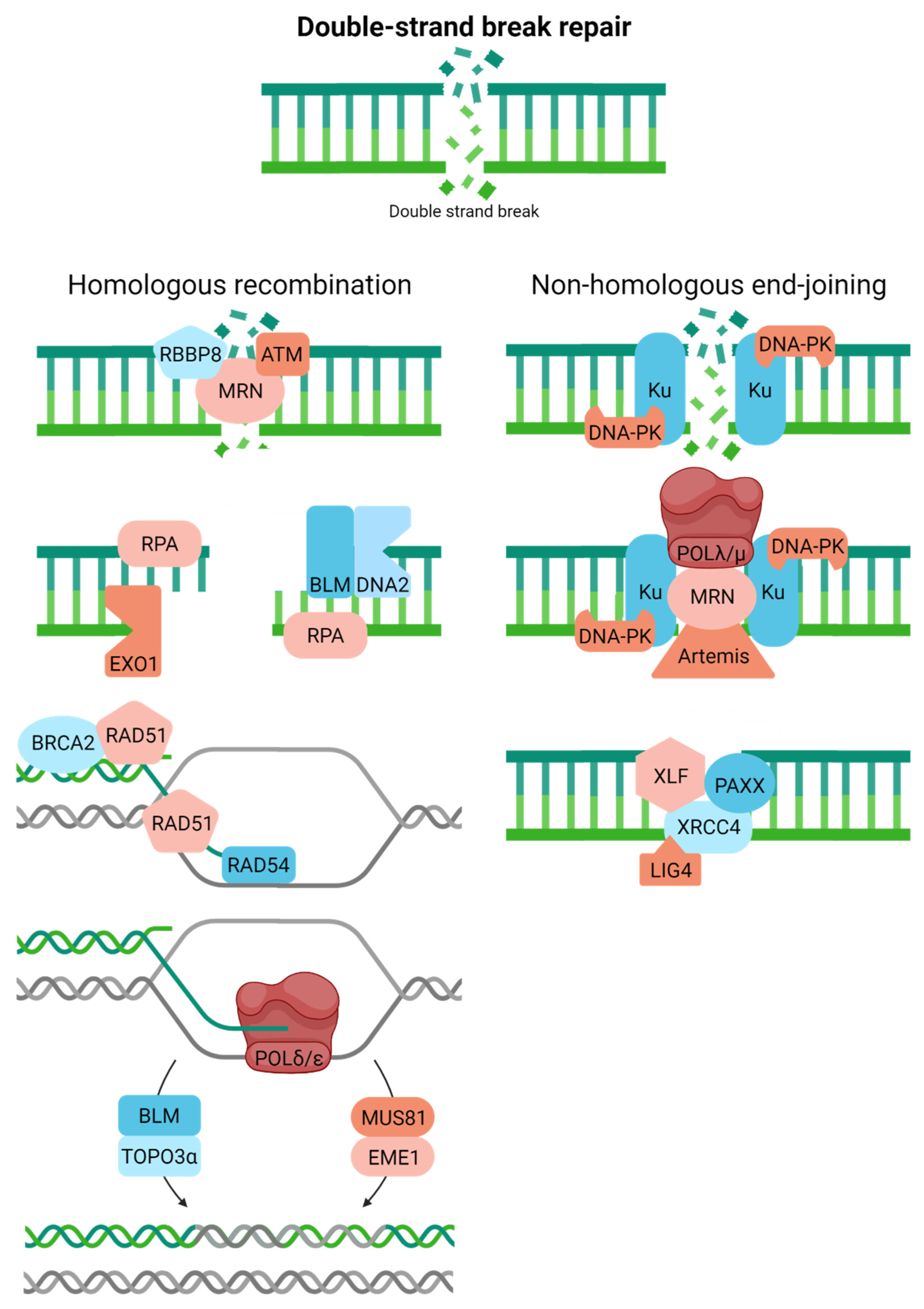
4.7. Double-Strand Break Repair
5. DNA Repair Defects in Neurodegenerative Disease
5.1. Inherited Disorders Involving Defects in DNA Repair and Neurological Abnormalities
5.2. Associations of DNA Repair Defects with Classic Neurological Diseases
5.3. DNA Damage, DNA Repair, and Neuronal Cell Function
6. Summary and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Liebeskind, B.J.; Hillis, D.M.; Zakon, H.H.; Hofmann, H.A. Complex Homology and the Evolution of Nervous Systems. Trends Ecol. Evol. 2016, 31, 127–135. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laurent, G. On the value of model diversity in neuroscience. Nat. Rev. Neurosci. 2020, 21, 395–396. [Google Scholar] [CrossRef] [PubMed]
- Sousa, A.M.M.; Meyer, K.A.; Santpere, G.; Gulden, F.O.; Sestan, N. Evolution of the Human Nervous System Function, Structure, and Development. Cell 2017, 170, 226–247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lisman, J. The challenge of understanding the brain: Where we stand in 2015. Neuron 2015, 86, 864–882. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jäkel, S.; Dimou, L. Glial Cells and Their Function in the Adult Brain: A Journey through the History of Their Ablation. Front. Cell. Neurosci. 2017, 11, 24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- López-Otín, C.; Blasco, M.A.; Partridge, L.; Serrano, M.; Kroemer, G. The hallmarks of aging. Cell 2013, 153, 1194–1217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jellinger, K.A. Basic mechanisms of neurodegeneration: A critical update. J. Cell. Mol. Med. 2010, 14, 457–487. [Google Scholar] [CrossRef]
- Lindahl, T. Instability and decay of the primary structure of DNA. Nature 1993, 362, 709–715. [Google Scholar] [CrossRef]
- Deavall, D.G.; Martin, E.A.; Horner, J.M.; Roberts, R. Drug-Induced Oxidative Stress and Toxicity. J. Toxicol. 2012, 2012, 645460. [Google Scholar] [CrossRef] [Green Version]
- Gidron, Y.; Russ, K.; Tissarchondou, H.; Warner, J. The relation between psychological factors and DNA-damage: A critical review. Biol. Psychol. 2006, 72, 291–304. [Google Scholar] [CrossRef]
- De Bont, R.; van Larebeke, N. Endogenous DNA damage in humans: A review of quantitative data. Mutagenesis 2004, 19, 169–185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Voulgaridou, G.P.; Anestopoulos, I.; Franco, R.; Panayiotidis, M.I.; Pappa, A. DNA damage induced by endogenous aldehydes: Current state of knowledge. Mutat. Res. 2011, 711, 13–27. [Google Scholar] [CrossRef] [PubMed]
- Rydberg, B.; Lindahl, T. Nonenzymatic methylation of DNA by the intracellular methyl group donor S-adenosyl-L-methionine is a potentially mutagenic reaction. EMBO J. 1982, 1, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Reisz, J.A.; Bansal, N.; Qian, J.; Zhao, W.; Furdui, C.M. Effects of ionizing radiation on biological molecules--mechanisms of damage and emerging methods of detection. Antioxid. Redox Signal. 2014, 21, 260–292. [Google Scholar] [CrossRef] [PubMed]
- Harman, D. Aging: A Theory Based on Free Radical and Radiation Chemistry. J. Gerontol. 1956, 11, 298–300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kregel, K.C.; Zhang, H.J. An integrated view of oxidative stress in aging: Basic mechanisms, functional effects, and pathological considerations. Am. J. Physiol.-Regul. Integr. Comp. Physiol. 2007, 292, R18–R36. [Google Scholar] [CrossRef] [PubMed]
- Thau, L.; Reddy, V.; Singh, P. Anatomy, Central Nervous System. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Kadry, H.; Noorani, B.; Cucullo, L. A blood–brain barrier overview on structure, function, impairment, and biomarkers of integrity. Fluids Barriers CNS 2020, 17, 69. [Google Scholar] [CrossRef]
- Sokoloff, L. The brain as a chemical machine. Prog. Brain Res. 1992, 94, 19–33. [Google Scholar] [CrossRef]
- Kausar, S.; Wang, F.; Cui, H. The Role of Mitochondria in Reactive Oxygen Species Generation and Its Implications for Neurodegenerative Diseases. Cells 2018, 7, 274. [Google Scholar] [CrossRef] [Green Version]
- Lee, K.H.; Cha, M.; Lee, B.H. Neuroprotective Effect of Antioxidants in the Brain. Int. J. Mol. Sci. 2020, 21, 7152. [Google Scholar] [CrossRef]
- Huang, R.; Zhou, P.-K. DNA damage repair: Historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Signal Transduct. Target. Ther. 2021, 6, 254. [Google Scholar] [CrossRef] [PubMed]
- Dizdaroglu, M.; Jaruga, P. Mechanisms of free radical-induced damage to DNA. Free. Radic. Res. 2012, 46, 382–419. [Google Scholar] [CrossRef] [PubMed]
- Cadet, J.; Davies, K.J.A.; Medeiros, M.H.; Di Mascio, P.; Wagner, J.R. Formation and repair of oxidatively generated damage in cellular DNA. Free. Radic. Biol. Med. 2017, 107, 13–34. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, V.; Wilson, D.M. 3rd DNA Damage and Associated DNA Repair Defects in Disease and Premature Aging. Am. J. Hum. Genet. 2019, 105, 237–257. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Folch, J.; Junyent, F.; Verdaguer, E.; Auladell, C.; Pizarro, J.G.; Beas-Zarate, C.; Pallàs, M.; Camins, A. Role of cell cycle re-entry in neurons: A common apoptotic mechanism of neuronal cell death. Neurotox. Res. 2012, 22, 195–207. [Google Scholar] [CrossRef]
- Stein, A.; Sia, E.A. Mitochondrial DNA repair and damage tolerance. Front. Biosci. 2017, 22, 920–943. [Google Scholar] [CrossRef]
- Tano, K.; Shiota, S.; Collier, J.; Foote, R.S.; Mitra, S. Isolation and structural characterization of a cDNA clone encoding the human DNA repair protein for O6-alkylguanine. Proc. Natl. Acad. Sci. USA 1990, 87, 686–690. [Google Scholar] [CrossRef] [Green Version]
- Rydberg, B.; Spurr, N.; Karran, P. cDNA cloning and chromosomal assignment of the human O6-methylguanine-DNA methyltransferase. cDNA expression in Escherichia coli and gene expression in human cells. J. Biol. Chem. 1990, 265, 9563–9569. [Google Scholar] [CrossRef]
- Koike, G.; Maki, H.; Takeya, H.; Hayakawa, H.; Sekiguchi, M. Purification, structure, and biochemical properties of human O6-methylguanine-DNA methyltransferase. J. Biol. Chem. 1990, 265, 14754–14762. [Google Scholar] [CrossRef]
- Pegg, A.E. Multifaceted roles of alkyltransferase and related proteins in DNA repair, DNA damage, resistance to chemotherapy, and research tools. Chem. Res. Toxicol. 2011, 24, 618–639. [Google Scholar] [CrossRef] [Green Version]
- Fedeles, B.I.; Singh, V.; Delaney, J.C.; Li, D.; Essigmann, J.M. The AlkB Family of Fe(II)/α-Ketoglutarate-dependent Dioxygenases: Repairing Nucleic Acid Alkylation Damage and Beyond. J. Biol. Chem. 2015, 290, 20734–20742. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alemu, E.A.; He, C.; Klungland, A. ALKBHs-facilitated RNA modifications and de-modifications. DNA Repair 2016, 44, 87–91. [Google Scholar] [CrossRef] [Green Version]
- Duncan, T.; Trewick, S.C.; Koivisto, P.; Bates, P.A.; Lindahl, T.; Sedgwick, B. Reversal of DNA alkylation damage by two human dioxygenases. Proc. Natl. Acad. Sci. USA 2002, 99, 16660–16665. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aas, P.A.; Otterlei, M.; Falnes, P.O.; Vågbø, C.B.; Skorpen, F.; Akbari, M.; Sundheim, O.; Bjørås, M.; Slupphaug, G.; Seeberg, E.; et al. Human and bacterial oxidative demethylases repair alkylation damage in both RNA and DNA. Nature 2003, 421, 859–863. [Google Scholar] [CrossRef] [PubMed]
- Zdżalik, D.; Domańska, A.; Prorok, P.; Kosicki, K.; van den Born, E.; Falnes, P.; Rizzo, C.J.; Guengerich, F.P.; Tudek, B. Differential repair of etheno-DNA adducts by bacterial and human AlkB proteins. DNA Repair 2015, 30, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Westbye, M.P.; Feyzi, E.; Aas, P.A.; Vågbø, C.B.; Talstad, V.A.; Kavli, B.; Hagen, L.; Sundheim, O.; Akbari, M.; Liabakk, N.B.; et al. Human AlkB homolog 1 is a mitochondrial protein that demethylates 3-methylcytosine in DNA and RNA. J. Biol. Chem. 2008, 283, 25046–25056. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Modrich, P. Mechanisms in E. coli and Human Mismatch Repair (Nobel Lecture). Angew. Chem. Int. Ed. 2016, 55, 8490–8501. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drummond, J.T.; Li, G.M.; Longley, M.J.; Modrich, P. Isolation of an hMSH2-p160 heterodimer that restores DNA mismatch repair to tumor cells. Science 1995, 268, 1909–1912. [Google Scholar] [CrossRef]
- Acharya, S.; Wilson, T.; Gradia, S.; Kane, M.F.; Guerrette, S.; Marsischky, G.T.; Kolodner, R.; Fishel, R. hMSH2 forms specific mispair-binding complexes with hMSH3 and hMSH6. Proc. Natl. Acad. Sci. USA 1996, 93, 13629–13634. [Google Scholar] [CrossRef] [Green Version]
- Genschel, J.; Modrich, P. Mechanism of 5′-directed excision in human mismatch repair. Mol. Cell 2003, 12, 1077–1086. [Google Scholar] [CrossRef]
- Zhang, Y.; Yuan, F.; Presnell, S.R.; Tian, K.; Gao, Y.; Tomkinson, A.E.; Gu, L.; Li, G.M. Reconstitution of 5′-directed human mismatch repair in a purified system. Cell 2005, 122, 693–705. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kadyrov, F.A.; Dzantiev, L.; Constantin, N.; Modrich, P. Endonucleolytic function of MutLalpha in human mismatch repair. Cell 2006, 126, 297–308. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kadyrov, F.A.; Genschel, J.; Fang, Y.; Penland, E.; Edelmann, W.; Modrich, P. A possible mechanism for exonuclease 1-independent eukaryotic mismatch repair. Proc. Natl. Acad. Sci. USA 2009, 106, 8495–8500. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kellner, V.; Luke, B. Molecular and physiological consequences of faulty eukaryotic ribonucleotide excision repair. EMBO J. 2020, 39, e102309. [Google Scholar] [CrossRef]
- Zhou, Z.X.; Williams, J.S.; Lujan, S.A.; Kunkel, T.A. Ribonucleotide incorporation into DNA during DNA replication and its consequences. Crit. Rev. Biochem. Mol. Biol. 2021, 56, 109–124. [Google Scholar] [CrossRef] [PubMed]
- Cerritelli, S.M.; Crouch, R.J. RNase H2-RED carpets the path to eukaryotic RNase H2 functions. DNA Repair 2019, 84, 102736. [Google Scholar] [CrossRef]
- Sparks, J.L.; Chon, H.; Cerritelli, S.M.; Kunkel, T.A.; Johansson, E.; Crouch, R.J.; Burgers, P.M. RNase H2-Initiated Ribonucleotide Excision Repair. Mol. Cell 2012, 47, 980–986. [Google Scholar] [CrossRef] [Green Version]
- Krokan, H.E.; Bjørås, M. Base excision repair. Cold Spring Harb. Perspect. Biol. 2013, 5, a012583. [Google Scholar] [CrossRef]
- Hindi, N.N.; Elsakrmy, N.; Ramotar, D. The base excision repair process: Comparison between higher and lower eukaryotes. Cell. Mol. Life Sci. 2021, 78, 7943–7965. [Google Scholar] [CrossRef]
- Li, M.; Wilson, D.M. 3rd Human apurinic/apyrimidinic endonuclease 1. Antioxid. Redox Signal. 2014, 20, 678–707. [Google Scholar] [CrossRef] [Green Version]
- Beard, W.A. DNA polymerase β: Closing the gap between structure and function. DNA Repair 2020, 93, 102910. [Google Scholar] [CrossRef] [PubMed]
- London, R.E. XRCC1-Strategies for coordinating and assembling a versatile DNA damage response. DNA Repair 2020, 93, 102917. [Google Scholar] [CrossRef] [PubMed]
- Caldecott, K.W. XRCC1 protein; Form and function. DNA Repair 2019, 81, 102664. [Google Scholar] [CrossRef] [PubMed]
- Robertson, A.B.; Klungland, A.; Rognes, T.; Leiros, I. DNA repair in mammalian cells: Base excision repair: The long and short of it. Cell. Mol. Life Sci. 2009, 66, 981–993. [Google Scholar] [CrossRef] [PubMed]
- Yoon, G.; Caldecott, K.W. Nonsyndromic cerebellar ataxias associated with disorders of DNA single-strand break repair. Handb. Clin. Neurol. 2018, 155, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Andres, S.N.; Schellenberg, M.J.; Wallace, B.D.; Tumbale, P.; Williams, R.S. Recognition and repair of chemically heterogeneous structures at DNA ends. Environ. Mol. Mutagen. 2015, 56, 1–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abbotts, R.; Wilson, D.M. 3rd Coordination of DNA single strand break repair. Free. Radic. Biol. Med. 2017, 107, 228–244. [Google Scholar] [CrossRef]
- Parsons, J.L.; Dianova, I.I.; Dianov, G.L. APE1 is the major 3′-phosphoglycolate activity in human cell extracts. Nucleic Acids Res. 2004, 32, 3531–3536. [Google Scholar] [CrossRef] [Green Version]
- Wiederhold, L.; Leppard, J.B.; Kedar, P.; Karimi-Busheri, F.; Rasouli-Nia, A.; Weinfeld, M.; Tomkinson, A.E.; Izumi, T.; Prasad, R.; Wilson, S.H.; et al. AP endonuclease-independent DNA base excision repair in human cells. Mol. Cell 2004, 15, 209–220. [Google Scholar] [CrossRef]
- Ahel, I.; Rass, U.; El-Khamisy, S.F.; Katyal, S.; Clements, P.M.; McKinnon, P.J.; Caldecott, K.W.; West, S.C. The neurodegenerative disease protein aprataxin resolves abortive DNA ligation intermediates. Nature 2006, 443, 713–716. [Google Scholar] [CrossRef]
- Karimi-Busheri, F.; Daly, G.; Robins, P.; Canas, B.; Pappin, D.J.; Sgouros, J.; Miller, G.G.; Fakhrai, H.; Davis, E.M.; Le Beau, M.M.; et al. Molecular characterization of a human DNA kinase. J. Biol. Chem. 1999, 274, 24187–24194. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jilani, A.; Ramotar, D.; Slack, C.; Ong, C.; Yang, X.M.; Scherer, S.W.; Lasko, D.D. Molecular cloning of the human gene, PNKP, encoding a polynucleotide kinase 3′-phosphatase and evidence for its role in repair of DNA strand breaks caused by oxidative damage. J. Biol. Chem. 1999, 274, 24176–24186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Khamisy, S.F.; Saifi, G.M.; Weinfeld, M.; Johansson, F.; Helleday, T.; Lupski, J.R.; Caldecott, K.W. Defective DNA single-strand break repair in spinocerebellar ataxia with axonal neuropathy-1. Nature 2005, 434, 108–113. [Google Scholar] [CrossRef] [PubMed]
- Balakrishnan, L.; Bambara, R.A. Flap endonuclease 1. Annu. Rev. Biochem. 2013, 82, 119–138. [Google Scholar] [CrossRef] [Green Version]
- Dehé, P.M.; Gaillard, P.H.L. Control of structure-specific endonucleases to maintain genome stability. Nat. Rev. Mol. Cell Biol. 2017, 18, 315–330. [Google Scholar] [CrossRef]
- Faridounnia, M.; Folkers, G.E.; Boelens, R. Function and Interactions of ERCC1-XPF in DNA Damage Response. Molecules 2018, 23, 3205. [Google Scholar] [CrossRef] [Green Version]
- Jung, K.Y.; Kodama, T.; Greenberg, M.M. Repair of the major lesion resulting from C5′-oxidation of DNA. Biochemistry 2011, 50, 6273–6279. [Google Scholar] [CrossRef] [Green Version]
- Çaglayan, M.; Prasad, R.; Krasich, R.; Longley, M.J.; Kadoda, K.; Tsuda, M.; Sasanuma, H.; Takeda, S.; Tano, K.; Copeland, W.C.; et al. Complementation of aprataxin deficiency by base excision repair enzymes in mitochondrial extracts. Nucleic Acids Res. 2017, 45, 10079–10088. [Google Scholar] [CrossRef]
- Ellenberger, T.; Tomkinson, A.E. Eukaryotic DNA ligases: Structural and functional insights. Annu. Rev. Biochem. 2008, 77, 313–338. [Google Scholar] [CrossRef] [Green Version]
- Mullenders, L.H.F. Solar UV damage to cellular DNA: From mechanisms to biological effects. Photochem. Photobiol. Sci. 2018, 17, 1842–1852. [Google Scholar] [CrossRef]
- Cleaver, J.E.; Lam, E.T.; Revet, I. Disorders of nucleotide excision repair: The genetic and molecular basis of heterogeneity. Nat. Rev. Genet. 2009, 10, 756–768. [Google Scholar] [CrossRef] [PubMed]
- Sugasawa, K.; Ng, J.M.; Masutani, C.; Iwai, S.; van der Spek, P.J.; Eker, A.P.; Hanaoka, F.; Bootsma, D.; Hoeijmakers, J.H. Xeroderma pigmentosum group C protein complex is the initiator of global genome nucleotide excision repair. Mol. Cell 1998, 2, 223–232. [Google Scholar] [CrossRef]
- Sugasawa, K. Molecular mechanisms of DNA damage recognition for mammalian nucleotide excision repair. DNA Repair 2016, 44, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Mellon, I.; Bohr, V.A.; Smith, C.A.; Hanawalt, P.C. Preferential DNA repair of an active gene in human cells. Proc. Natl. Acad. Sci. USA 1986, 83, 8878–8882. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Henning, K.A.; Li, L.; Iyer, N.; McDaniel, L.D.; Reagan, M.S.; Legerski, R.; Schultz, R.A.; Stefanini, M.; Lehmann, A.R.; Mayne, L.V.; et al. The Cockayne syndrome group A gene encodes a WD repeat protein that interacts with CSB protein and a subunit of RNA polymerase II TFIIH. Cell 1995, 82, 555–564. [Google Scholar] [CrossRef] [Green Version]
- Troelstra, C.; van Gool, A.; de Wit, J.; Vermeulen, W.; Bootsma, D.; Hoeijmakers, J.H. ERCC6, a member of a subfamily of putative helicases, is involved in Cockayne’s syndrome and preferential repair of active genes. Cell 1992, 71, 939–953. [Google Scholar] [CrossRef] [Green Version]
- D’Souza, A.; Blee, A.M.; Chazin, W.J. Mechanism of action of nucleotide excision repair machinery. Biochem. Soc. Trans. 2022, 50, 375–386. [Google Scholar] [CrossRef]
- Scully, R.; Panday, A.; Elango, R.; Willis, N.A. DNA double-strand break repair-pathway choice in somatic mammalian cells. Nat. Rev. Mol. Cell Biol. 2019, 20, 698–714. [Google Scholar] [CrossRef]
- Jachimowicz, R.D.; Goergens, J.; Reinhardt, H.C. DNA double-strand break repair pathway choice—From basic biology to clinical exploitation. Cell Cycle 2019, 18, 1423–1434. [Google Scholar] [CrossRef]
- Jasin, M.; Rothstein, R. Repair of strand breaks by homologous recombination. Cold Spring Harb. Perspect. Biol. 2013, 5, a012740. [Google Scholar] [CrossRef]
- Sun, Y.; McCorvie, T.J.; Yates, L.A.; Zhang, X. Structural basis of homologous recombination. Cell. Mol. Life Sci. 2020, 77, 3–18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elbakry, A.; Löbrich, M. Homologous Recombination Subpathways: A Tangle to Resolve. Front. Genet. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Pannunzio, N.R.; Watanabe, G.; Lieber, M.R. Nonhomologous DNA end-joining for repair of DNA double-strand breaks. J. Biol. Chem. 2018, 293, 10512–10523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaplin, A.K.; Blundell, T.L. Structural biology of multicomponent assemblies in DNA double-strand-break repair through non-homologous end joining. Curr. Opin. Struct. Biol. 2020, 61, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Sallmyr, A.; Tomkinson, A.E. Repair of DNA double-strand breaks by mammalian alternative end-joining pathways. J. Biol. Chem. 2018, 293, 10536–10546. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rolig, R.L.; McKinnon, P.J. Linking DNA damage and neurodegeneration. Trends Neurosci. 2000, 23, 417–424. [Google Scholar] [CrossRef]
- Cleaver, J.E. Defective repair replication of DNA in xeroderma pigmentosum. Nature 1968, 218, 652–656. [Google Scholar] [CrossRef]
- Setlow, R.B.; Regan, J.D.; German, J.; Carrier, W.L. Evidence that xeroderma pigmentosum cells do not perform the first step in the repair of ultraviolet damage to their DNA. Proc. Natl. Acad. Sci. USA 1969, 64, 1035–1041. [Google Scholar] [CrossRef] [Green Version]
- Rizza, E.R.H.; DiGiovanna, J.J.; Khan, S.G.; Tamura, D.; Jeskey, J.D.; Kraemer, K.H. Xeroderma Pigmentosum: A Model for Human Premature Aging. J. Investig. Dermatol. 2021, 141, 976–984. [Google Scholar] [CrossRef]
- Kumar, N.; Raja, S.; Van Houten, B. The involvement of nucleotide excision repair proteins in the removal of oxidative DNA damage. Nucleic Acids Res. 2020, 48, 11227–11243. [Google Scholar] [CrossRef]
- Kühbacher, U.; Duxin, J.P. How to fix DNA-protein crosslinks. DNA Repair 2020, 94, 102924. [Google Scholar] [CrossRef] [PubMed]
- McKinnon, P.J. DNA repair deficiency and neurological disease. Nat. Rev. Neurosci. 2009, 10, 100–112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Madabhushi, R.; Pan, L.; Tsai, L.H. DNA damage and its links to neurodegeneration. Neuron 2014, 83, 266–282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rothblum-Oviatt, C.; Wright, J.; Lefton-Greif, M.A.; McGrath-Morrow, S.A.; Crawford, T.O.; Lederman, H.M. Ataxia telangiectasia: A review. Orphanet J. Rare Dis. 2016, 11, 159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kastan, M.B.; Zhan, Q.; el-Deiry, W.S.; Carrier, F.; Jacks, T.; Walsh, W.V.; Plunkett, B.S.; Vogelstein, B.; Fornace, A.J., Jr. A mammalian cell cycle checkpoint pathway utilizing p53 and GADD45 is defective in ataxia-telangiectasia. Cell 1992, 71, 587–597. [Google Scholar] [CrossRef]
- Rotman, G.; Shiloh, Y. ATM: A mediator of multiple responses to genotoxic stress. Oncogene 1999, 18, 6135–6144. [Google Scholar] [CrossRef] [Green Version]
- Shiloh, Y. The cerebellar degeneration in ataxia-telangiectasia: A case for genome instability. DNA Repair 2020, 95, 102950. [Google Scholar] [CrossRef]
- Lavin, M.F.; Kozlov, S.; Gatei, M.; Kijas, A.W. ATM-Dependent Phosphorylation of All Three Members of the MRN Complex: From Sensor to Adaptor. Biomolecules 2015, 5, 2877–2902. [Google Scholar] [CrossRef] [Green Version]
- Taylor, A.M.R.; Rothblum-Oviatt, C.; Ellis, N.A.; Hickson, I.D.; Meyer, S.; Crawford, T.O.; Smogorzewska, A.; Pietrucha, B.; Weemaes, C.; Stewart, G.S. Chromosome instability syndromes. Nat. Rev. Dis. Primers 2019, 5, 64. [Google Scholar] [CrossRef]
- Alt, F.W.; Schwer, B. DNA double-strand breaks as drivers of neural genomic change, function, and disease. DNA Repair 2018, 71, 158–163. [Google Scholar] [CrossRef]
- Amirifar, P.; Ranjouri, M.R.; Yazdani, R.; Abolhassani, H.; Aghamohammadi, A. Ataxia-telangiectasia: A review of clinical features and molecular pathology. Pediatr. Allergy Immunol. 2019, 30, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Bennett, C.L.; La Spada, A.R. Novel Helicase at the Interface of RNA Transcriptome Regulation and Neurobiology: From Normal Function to Pathological Roles in Motor Neuron Disease and Cerebellar Degeneration. Adv. Neurobiol. 2018, 20, 265–281. [Google Scholar] [CrossRef] [PubMed]
- Brambati, A.; Zardoni, L.; Nardini, E.; Pellicioli, A.; Liberi, G. The dark side of RNA:DNA hybrids. Mutat. Res. Rev. Mutat. Res. 2020, 784, 108300. [Google Scholar] [CrossRef] [PubMed]
- Perego, M.G.L.; Taiana, M.; Bresolin, N.; Comi, G.P.; Corti, S. R-Loops in Motor Neuron Diseases. Mol. Neurobiol. 2019, 56, 2579–2589. [Google Scholar] [CrossRef] [PubMed]
- Fang, E.F.; Scheibye-Knudsen, M.; Chua, K.F.; Mattson, M.P.; Croteau, D.L.; Bohr, V.A. Nuclear DNA damage signalling to mitochondria in ageing. Nat. Rev. Mol. Cell Biol. 2016, 17, 308–321. [Google Scholar] [CrossRef] [Green Version]
- Coppedè, F.; Migliore, L. DNA damage in neurodegenerative diseases. Mutat. Res. 2015, 776, 84–97. [Google Scholar] [CrossRef]
- Czarny, P.; Bialek, K.; Ziolkowska, S.; Strycharz, J.; Sliwinski, T. DNA damage and repair in neuropsychiatric disorders. What do we know and what are the future perspectives? Mutagenesis 2020, 35, 79–106. [Google Scholar] [CrossRef]
- Gupta, S.; You, P.; SenGupta, T.; Nilsen, H.; Sharma, K. Crosstalk between Different DNA Repair Pathways Contributes to Neurodegenerative Diseases. Biology 2021, 10, 163. [Google Scholar] [CrossRef]
- Sykora, P.; Misiak, M.; Wang, Y.; Ghosh, S.; Leandro, G.S.; Liu, D.; Tian, J.; Baptiste, B.A.; Cong, W.N.; Brenerman, B.M.; et al. DNA polymerase β deficiency leads to neurodegeneration and exacerbates Alzheimer disease phenotypes. Nucleic Acids Res. 2015, 43, 943–959. [Google Scholar] [CrossRef]
- Shanbhag, N.M.; Evans, M.D.; Mao, W.; Nana, A.L.; Seeley, W.W.; Adame, A.; Rissman, R.A.; Masliah, E.; Mucke, L. Early neuronal accumulation of DNA double strand breaks in Alzheimer’s disease. Acta Neuropathol. Commun. 2019, 7, 77. [Google Scholar] [CrossRef] [Green Version]
- Thadathil, N.; Delotterie, D.F.; Xiao, J.; Hori, R.; McDonald, M.P.; Khan, M.M. DNA Double-Strand Break Accumulation in Alzheimer’s Disease: Evidence from Experimental Models and Postmortem Human Brains. Mol. Neurobiol. 2021, 58, 118–131. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Curle, A.J.; Haider, A.M.; Balmus, G. The role of DNA damage response in amyotrophic lateral sclerosis. Essays Biochem. 2020, 64, 847–861. [Google Scholar] [CrossRef] [PubMed]
- Mitra, J.; Hegde, M.L. A Commentary on TDP-43 and DNA Damage Response in Amyotrophic Lateral Sclerosis. J. Exp. Neurosci. 2019, 13, 1179069519880166. [Google Scholar] [CrossRef]
- Wang, H.; Guo, W.; Mitra, J.; Hegde, P.M.; Vandoorne, T.; Eckelmann, B.J.; Mitra, S.; Tomkinson, A.E.; Van Den Bosch, L.; Hegde, M.L. Mutant FUS causes DNA ligation defects to inhibit oxidative damage repair in Amyotrophic Lateral Sclerosis. Nat. Commun. 2018, 9, 3683. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, W.Y.; Pan, L.; Su, S.C.; Quinn, E.J.; Sasaki, M.; Jimenez, J.C.; Mackenzie, I.R.; Huang, E.J.; Tsai, L.H. Interaction of FUS and HDAC1 regulates DNA damage response and repair in neurons. Nat. Neurosci. 2013, 16, 1383–1391. [Google Scholar] [CrossRef] [PubMed]
- Sanna, S.; Esposito, S.; Masala, A.; Sini, P.; Nieddu, G.; Galioto, M.; Fais, M.; Iaccarino, C.; Cestra, G.; Crosio, C. HDAC1 inhibition ameliorates TDP-43-induced cell death in vitro and in vivo. Cell Death Dis. 2020, 11, 369. [Google Scholar] [CrossRef] [PubMed]
- Milanese, C.; Cerri, S.; Ulusoy, A.; Gornati, S.V.; Plat, A.; Gabriels, S.; Blandini, F.; Di Monte, D.A.; Hoeijmakers, J.H.; Mastroberardino, P.G. Activation of the DNA damage response in vivo in synucleinopathy models of Parkinson’s disease. Cell Death Dis. 2018, 9, 818. [Google Scholar] [CrossRef] [Green Version]
- Gonzalez-Hunt, C.P.; Sanders, L.H. DNA damage and repair in Parkinson’s disease: Recent advances and new opportunities. J. Neurosci. Res. 2021, 99, 180–189. [Google Scholar] [CrossRef]
- Sepe, S.; Milanese, C.; Gabriels, S.; Derks, K.W.; Payan-Gomez, C.; van Ijcken, W.; Rijksen, Y.M.; Nigg, A.L.; Moreno, S.; Cerri, S.; et al. Inefficient DNA Repair Is an Aging-Related Modifier of Parkinson’s Disease. Cell Rep. 2016, 15, 1866–1875. [Google Scholar] [CrossRef] [Green Version]
- SenGupta, T.; Palikaras, K.; Esbensen, Y.Q.; Konstantinidis, G.; Galindo, F.J.N.; Achanta, K.; Kassahun, H.; Stavgiannoudaki, I.; Bohr, V.A.; Akbari, M.; et al. Base excision repair causes age-dependent accumulation of single-stranded DNA breaks that contribute to Parkinson disease pathology. Cell Rep. 2021, 36, 109668. [Google Scholar] [CrossRef]
- Singh, R.N.; Howell, M.D.; Ottesen, E.W.; Singh, N.N. Diverse role of survival motor neuron protein. Biochim. Biophys. Acta-Gene Regul. Mech. 2017, 1860, 299–315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abugable, A.A.; Morris, J.L.M.; Palminha, N.M.; Zaksauskaite, R.; Ray, S.; El-Khamisy, S.F. DNA repair and neurological disease: From molecular understanding to the development of diagnostics and model organisms. DNA Repair 2019, 81, 102669. [Google Scholar] [CrossRef] [PubMed]
- Maiuri, T.; Suart, C.E.; Hung, C.L.K.; Graham, K.J.; Barba Bazan, C.A.; Truant, R. DNA Damage Repair in Huntington’s Disease and Other Neurodegenerative Diseases. Neurotherapeutics 2019, 16, 948–956. [Google Scholar] [CrossRef]
- McLoughlin, H.S.; Moore, L.R.; Paulson, H.L. Pathogenesis of SCA3 and implications for other polyglutamine diseases. Neurobiol. Dis. 2020, 134, 104635. [Google Scholar] [CrossRef] [PubMed]
- Iyer, R.R.; Pluciennik, A. DNA Mismatch Repair and its Role in Huntington’s Disease. J. Huntingt. Dis. 2021, 10, 75–94. [Google Scholar] [CrossRef] [PubMed]
- Suberbielle, E.; Sanchez, P.E.; Kravitz, A.V.; Wang, X.; Ho, K.; Eilertson, K.; Devidze, N.; Kreitzer, A.C.; Mucke, L. Physiologic brain activity causes DNA double-strand breaks in neurons, with exacerbation by amyloid-β. Nat. Neurosci. 2013, 16, 613–621. [Google Scholar] [CrossRef] [Green Version]
- Madabhushi, R.; Gao, F.; Pfenning, A.R.; Pan, L.; Yamakawa, S.; Seo, J.; Rueda, R.; Phan, T.X.; Yamakawa, H.; Pao, P.C.; et al. Activity-Induced DNA Breaks Govern the Expression of Neuronal Early-Response Genes. Cell 2015, 161, 1592–1605. [Google Scholar] [CrossRef] [Green Version]
- Li, X.; Marshall, P.R.; Leighton, L.J.; Zajaczkowski, E.L.; Wang, Z.; Madugalle, S.U.; Yin, J.; Bredy, T.W.; Wei, W. The DNA Repair-Associated Protein Gadd45γ Regulates the Temporal Coding of Immediate Early Gene Expression within the Prelimbic Prefrontal Cortex and Is Required for the Consolidation of Associative Fear Memory. J. Neurosci. Off. J. Soc. Neurosci. 2019, 39, 970–983. [Google Scholar] [CrossRef]
- Saavedra-Rodríguez, L.; Vázquez, A.; Ortiz-Zuazaga, H.G.; Chorna, N.E.; González, F.A.; Andrés, L.; Rodríguez, K.; Ramírez, F.; Rodríguez, A.; Peña de Ortiz, S. Identification of flap structure-specific endonuclease 1 as a factor involved in long-term memory formation of aversive learning. J. Neurosci. Off. J. Soc. Neurosci. 2009, 29, 5726–5737. [Google Scholar] [CrossRef]
- Goldberg, S.; Visochek, L.; Giladi, E.; Gozes, I.; Cohen-Armon, M. PolyADP-ribosylation is required for long-term memory formation in mammals. J. Neurochem. 2009, 111, 72–79. [Google Scholar] [CrossRef]
- Wu, W.; Hill, S.E.; Nathan, W.J.; Paiano, J.; Callen, E.; Wang, D.; Shinoda, K.; van Wietmarschen, N.; Colón-Mercado, J.M.; Zong, D.; et al. Neuronal enhancers are hotspots for DNA single-strand break repair. Nature 2021, 593, 440–444. [Google Scholar] [CrossRef] [PubMed]
- Reid, D.A.; Reed, P.J.; Schlachetzki, J.C.M.; Nitulescu, I.I.; Chou, G.; Tsui, E.C.; Jones, J.R.; Chandran, S.; Lu, A.T.; McClain, C.A.; et al. Incorporation of a nucleoside analog maps genome repair sites in postmitotic human neurons. Science 2021, 372, 91–94. [Google Scholar] [CrossRef]
- Weren, R.D.; Ligtenberg, M.J.; Geurts van Kessel, A.; De Voer, R.M.; Hoogerbrugge, N.; Kuiper, R.P. NTHL1 and MUTYH polyposis syndromes: Two sides of the same coin? J. Pathol. 2018, 244, 135–142. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Livingston, J.H.; Crow, Y.J. Neurologic Phenotypes Associated with Mutations in TREX1, RNASEH2A, RNASEH2B, RNASEH2C, SAMHD1, ADAR1, and IFIH1: Aicardi-Goutières Syndrome and Beyond. Neuropediatrics 2016, 47, 355–360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aditi; Downing, S.M.; Schreiner, P.A.; Kwak, Y.D.; Li, Y.; Shaw, T.I.; Russell, H.R.; McKinnon, P.J. Genome instability independent of type I interferon signaling drives neuropathology caused by impaired ribonucleotide excision repair. Neuron 2021, 109, 3962–3979.e3966. [Google Scholar] [CrossRef] [PubMed]
- Suberbielle, E.; Djukic, B.; Evans, M.; Kim, D.H.; Taneja, P.; Wang, X.; Finucane, M.; Knox, J.; Ho, K.; Devidze, N.; et al. DNA repair factor BRCA1 depletion occurs in Alzheimer brains and impairs cognitive function in mice. Nat. Commun. 2015, 6, 8897. [Google Scholar] [CrossRef] [PubMed]
- Mazina, O.M.; Keskin, H.; Hanamshet, K.; Storici, F.; Mazin, A.V. Rad52 Inverse Strand Exchange Drives RNA-Templated DNA Double-Strand Break Repair. Mol. Cell 2017, 67, 19–29.e13. [Google Scholar] [CrossRef] [Green Version]
- Welty, S.; Teng, Y.; Liang, Z.; Zhao, W.; Sanders, L.H.; Greenamyre, J.T.; Rubio, M.E.; Thathiah, A.; Kodali, R.; Wetzel, R.; et al. RAD52 is required for RNA-templated recombination repair in post-mitotic neurons. J. Biol. Chem. 2018, 293, 1353–1362. [Google Scholar] [CrossRef] [Green Version]
- Yasuhara, T.; Kato, R.; Hagiwara, Y.; Shiotani, B.; Yamauchi, M.; Nakada, S.; Shibata, A.; Miyagawa, K. Human Rad52 Promotes XPG-Mediated R-loop Processing to Initiate Transcription-Associated Homologous Recombination Repair. Cell 2018, 175, 558–570.e511. [Google Scholar] [CrossRef] [Green Version]
- McDevitt, S.; Rusanov, T.; Kent, T.; Chandramouly, G.; Pomerantz, R.T. How RNA transcripts coordinate DNA recombination and repair. Nat. Commun. 2018, 9, 1091. [Google Scholar] [CrossRef]
| DNA Repair Gene | Full Name | Protein Biochemical Function | Genetic Disease/Disorder * | |
|---|---|---|---|---|
| Modulation of Nucleotide Pools | ||||
| DNPH1 | 2′-Deoxynucleoside 5′-Phosphate N-Hydrolase 1 | Hydrolase for 5-hydroxymethyl deoxyuridine | NLE | |
| DUT | dUTP Pyrophosphatase | dUTPase | NLE | |
| NUDT1 (MTH1) | Nudix Hydrolase 1 (MutT Homolog 1) | 8-oxoGTPase | NLE | |
| PARK7 (DJ1) | Park7 Gene (Oncogene DJ1) | Guanine glycation repair | Parkinson disease 7 | ● |
| Direct Reversal | ||||
| ALKB3 (DEPC1) | AlkB Homolog 3 | 1-meA dioxygenase | NLE | |
| ALKBH2 (ABH2) | AlkB Homolog 2 | 1-meA dioxygenase | NLE | |
| MGMT | Methylguanine-DNA Methyltransferase | O6-meG alkyltransferase | NLE | |
| Mismatch Repair | ||||
| EXO1 | Exonuclease 1 | 5′ exonuclease | NLE | |
| MLH1 | MutL Homolog 1 | MutL homologs, forming heterodimer | Colorectal cancer, hereditary nonpolyposis, type 2 Mismatch repair cancer syndrome 1 Muir–Torre syndrome | |
| MSH2 | MutS Homolog 2 | Mismatch and loop recognition | Colorectal cancer, hereditary nonpolyposis, type 1 Mismatch repair cancer syndrome 2 Muir–Torre syndrome | |
| MSH3 | MutS Homolog 3 | Loop recognition | Endometrial carcinoma, somatic Familial adenomatous polyposis 4 | |
| MSH6 | MutS Homolog 6 | Mismatch recognition | Endometrial carcinoma, familial Colorectal cancer, hereditary nonpolyposis, type 5 Mismatch repair cancer syndrome 3 | |
| PMS2 | PMS1 Homolog 2 | MutL homologs, forming heterodimer | Colorectal cancer, hereditary nonpolyposis, type 4 Mismatch repair cancer syndrome 4 | |
| Base Excision Repair | ||||
| APEX1 (APE1) | Apurinic Endonuclease 1 | AP endonuclease | NLE | |
| MBD4 | Methyl-CpG-Binding Domain Protein 4 | DNA glycosylases: major altered base released: U or T opposite G at CpG sequences | NLE | |
| MPG (AAG) | N-Methylpurine DNA Glycosylase (3-Alkyladenine DNA Glycosylase) | DNA glycosylases: major altered base released: 3-meA, ethenoA, hypoxanthine | NLE | |
| MUTYH (MUTY) | MutY DNA Glycosylase | DNA glycosylases: major altered base released: A opposite 8-oxoG | Adenomas, multiple colorectal Gastric cancer, somatic | |
| NEIL1 | Endonuclease VIII-Like 1 | DNA glycosylases: major altered base released: Removes thymine glycol | NLE | |
| NEIL2 | Endonuclease VIII-Like 2 | DNA glycosylases: major altered base released: Removes oxidative products of pyrimidines | NLE | |
| NEIL3 | Endonuclease VIII-Like 3 | DNA glycosylases: major altered base released: Removes oxidative products of pyrimidines | NLE | |
| NTHL1 (NTH1) | Endonuclease III-Like 1 | DNA glycosylases: major altered base released: Ring-saturated or fragmented pyrimidines | Familial adenomatous polyposis 3 | |
| OGG1 | 8-Oxoguanine DNA Glycosylase | DNA glycosylases: major altered base released: 8-oxoG opposite C | Renal cell carcinoma, clear cell, somatic | |
| POLB | DNA Polymerase β | BER in nuclear DNA | NLE | |
| SMUG1 | Single-Strand-Selective Monofunctional Uracil-DNA Glycosylase 1 | DNA glycosylases: major altered base released: U | NLE | |
| TDG | Thymine-DNA Glycosylase | DNA glycosylases: major altered base released: U, T or ethenoC opposite G | NLE | |
| UNG | Uracil-DNA Glycosylase | DNA glycosylases: major altered base released: U | Immunodeficiency with hyper IgM, type 5 | |
| Ribonucleotide Excision Repair | ||||
| RNASEH2 | Ribonuclease H2 | RNA-DNA ribonuclease | Aicardi–Goutieres syndrome 2 | ● |
| Aicardi–Goutieres syndrome 3 | ● | |||
| Aicardi–Goutieres syndrome 4 | ● | |||
| TOP1 | DNA Topoisomerase I | Alter DNA topology by breaking single DNA strand | DNA topoisomerase U, camptothecin-resistant | |
| Nucleotide Excision Repair | ||||
| CETN2 | Centrin 2 | Binds DNA distortions | NLE | |
| DDB1 | DNA Damage-Binding Protein 1 | Complex defective in XP group E | NLE | |
| DDB2 (XPE) | DNA Damage-Binding Protein 2 | Complex defective in XP group E | Xeroderma pigmentosum, group E, DDB-negative subtype | ● |
| ERCC1 | Excision Repair Complementing Defective in Chinese Hamster 1 | 5′ incision DNA binding subunit | Cerebrooculofacioskeletal syndrome 4 | ● |
| ERCC2 (XPD) | Excision Repair Complementing Defective in Chinese Hamster 2 (XPD Gene) | 5′ to 3′ DNA helicase | Cerebrooculofacioskeletal syndrome 2 # | ● |
| Trichothiodystrophy 1, photosensitive | ||||
| Xeroderma pigmentosum, group D | ● | |||
| ERCC3 (XPB) | Excision Repair, Complementing Defective in Chinese Hamster 3 (XPB Gene) | 3′ to 5′ DNA helicase | Trichothiodystrophy 2, photosensitive | |
| Xeroderma pigmentosum, group B | ● | |||
| ERCC4 (XPF) | Excision Repair Complementing Defective in Chinese Hamster 4 (XPF Gene) | 5′ incision catalytic subunit | Fanconi anemia, complementation group Q | |
| Xeroderma pigmentosum, group F | ● | |||
| Xeroderma pigmentosum, type F/Cockayne syndrome | ● | |||
| XFE progeroid syndrome | ● | |||
| ERCC5 (XPG) | Excision Repair Complementing Defective in Chinese Hamster 5 (XPG Gene) | 3′ incision | Cerebrooculofacioskeletal syndrome 3 | ● |
| Xeroderma pigmentosum, group G | ● | |||
| Xeroderma pigmentosum, group G/Cockayne syndrome | ● | |||
| ERCC6 (CSB) | Excision Repair Cross-Complementing Group 6 | Cockayne syndrome and UV-Sensitive Syndrome; Needed for transcription-coupled NER | Susceptibility to lung cancer | |
| Susceptibility to macular degeneration, age related, 5 | ||||
| Cerebrooculofacioskeletal syndrome 1 | ● | |||
| Cockayne syndrome, type B | ● | |||
| De Sanctis–Cacchione syndrome | ● | |||
| Premature ovarian failure 11 | ||||
| UV-sensitive syndrome 1 | ||||
| ERCC8 (CSA) | Excision Repair Cross-Complementing Group 8 | Cockayne syndrome and UV-Sensitive Syndrome; Needed for transcription-coupled NER | Cockayne syndrome, type A | ● |
| UV-sensitive syndrome 2 | ||||
| RAD23A | RAD23 Homolog A | Substitutes for RAD23B | NLE | |
| RAD23B | RAD 23 Homolog B | Binds DNA distortions | NLE | |
| UVSSA (KIAA1530) | UV-Stimulated Scaffold Protein A | Cockayne syndrome and UV-Sensitive Syndrome; Needed for transcription-coupled NER | UV-sensitive syndrome 3 | |
| XPA | Xeroderma Pigmentosum Complementation Group A | Binds damaged DNA in preincision complex | Xeroderma pigmentosum, group A | ● |
| XPC | Xeroderma Pigmentosum Complementation Group C | Binds DNA distortions | Xeroderma pigmentosum, group C | |
| Strand Break Processing Factors and Helicases | ||||
| APEX2 | Apurinic/Apyrimidinic Endonuclease 2 | AP endonuclease | NLE | |
| APLF | Aprataxin- and PNKP-Like Factor | Accessory factor for DNA end-joining | NLE | |
| APTX | Aprataxin | Processing of DNA single-strand interruptions | Ataxia, early-onset, with oculomotor apraxia and hypoalbuminemia | ● |
| BLM | Bloom | Bloom syndrome helicase | Bloom syndrome | |
| LIG1 | Ligase 1 | DNA ligase | NLE | |
| LIG3 | Ligase III | DNA Ligase III | NLE | |
| PNKP | Polynucleotide Kinase 3′ Phosphatase | Converts some DNA breaks to ligatable ends | Charcot-Marie-Tooth disease, type 2B2 | ● |
| Ataxia-oculomotor apraxia 4 | ● | |||
| Microcephaly, seizures, and developmental delay | ● | |||
| RECQL (RECQ1) | RECQ Protein-Like | DNA helicase | ||
| RECQL4 | RECQ Protein-Like 4 | DNA helicase | Baller-Gerold syndrome RAPADILINO syndrome Rothmund–Thomson syndrome, type 2 | |
| RECQL5 | RECQ Protein-Like 5 | DNA helicase | NLE | |
| SPRTN (Spartan) | AprT-Like N-Terminal Domain Protein | Reads ubiquitylation | Ruijs–Aalfs syndrome | |
| TDP1 | Tyrosyl-DNA Phosphodiesterase 1 | Removes 3′-tyrosylphosphate and 3′-phosphoglycolate from DNA; human disorder SCAN1 | Spinocerebellar ataxia, autosomal recessive, with axonal neuropathy 1 | ● |
| TDP2 (TTRAP) | Tyrosyl-DNA Phosphodiesterase 2 (TRAF- and TNF Receptor-Associated Protein) | 5′- and 3′-tyrosyl DNA phosphodiesterase | Spinocerebellar ataxia, autosomal recessive 23 | ● |
| WRN | Werner | Werner syndrome helicase/3′–exonuclease | Werner syndrome | ● |
| XRCC1 | X-ray Repair Cross Complementing 1 | Scaffold LIG3 accessory factor | Spinocerebellar ataxia, autosomal recessive 26 # | ● |
| Non-Homologous End-Joining | ||||
| DCLRE1C (Artemis) | DNA Cross-Link Repair Protein 1C | Nuclease | Omenn syndrome Severe combined immunodeficiency, Athabascan type | |
| LIG4 | Ligase IV | Ligase | Resistance to Multiple myeloma | |
| LIG4 syndrome | ● | |||
| NHEJ1 (XLF, Cernunnos) | Nonhomologous End-Joining Factor 1 (XRCC4-Like Factor) | End joining factor | Severe combined immunodeficiency with microcephaly, growth retardation, and sensitivity to ionizing radiation | ● |
| PRKDC (DNA-PKcs) | Protein Kinase DNA-Activated Catalytic Subunit (DNA-Dependent Protein Kinase) | DNA-dependent protein kinase catalytic subunit | Immunodeficiency 26, with or without neurologic abnormalities | ● |
| XRCC4 | X-ray Repair Cross Complementing 4 | Ligase accessory factor | Short stature, microcephaly, and endocrine dysfunction | ● |
| XRCC5 (Ku80) | X-ray Repair Cross Complementing 5 (Ku Antigen, 80-KD Subunit) | DNA end binding subunit | NLE | |
| XRCC6 (Ku70) | X-ray Repair Cross Complementing 6 (Ku Antigen, 70-KD Subunit) | DNA end binding subunit | NLE | |
| Homologous Recombination | ||||
| BARD1 | BRCA1-Associated Ring Domain 1 | BRCA1-associated | Susceptibility to breast cancer | |
| BRCA1 | Breast Cancer 1 Gene | Accessory factor for transcription and recombination, E3 Ubiquitin ligase | Breast-ovarian cancer, familial, 1 Susceptibility to pancreatic cancer, 4 Fanconi anemia, complementation group S | |
| EME1 (MMS4L) | Essential Meiotic Structure-Specific Endonuclease 1 | Subunits of structure-specific DNA nuclease | NLE | |
| EME2 | Essential Meiotic Structure-Specific Endonuclease 2 | Subunits of structure-specific DNA nuclease | NLE | |
| GEN1 | GEN1 Homolog of Drosophila | Nuclease cleaving Holliday junctions | NLE | |
| HELQ (HEL308) | Helicase PolQ-Like | DNA helicase in RAD51 paralog complex | NLE | |
| MRE11A | MRE11 Homolog | 3′ exonuclease, defective in ATLD (ataxia-telangiectasia-like disorder) | Ataxia-telangiectasia-like disorder 1 | ● |
| MUS81 | MUS81 Structure-Specific Endonuclease Subunit | Subunits of structure-specific DNA nuclease | NLE | |
| NBN (NBS1) | Nibrin | Mutated in Nijmegen breakage syndrome | Aplastic anemia | |
| Leukemia, acute lymphoblastic | ||||
| Nijmegen breakage syndrome | ● | |||
| RAD50 | RAD50 Double-Strand Break Repair Protein | ATPase in complex with MRE11A, NBS1 | Nijmegen breakage syndrome-like disorder | ● |
| RAD51 | RAD51 Recombinase | Homologous pairing | Susceptibility to breast cancer | |
| Fanconi anemia, complementation group R | ||||
| Mirror movements 2 | ● | |||
| RAD51B | RAD51 Paralog B | RAD51 homolog | NLE | |
| RAD51D | RAD51 Paralog D | RAD51 homolog | Susceptibility to breast-ovarian cancer, familial, 4 | |
| RAD52 | RAD52 Homolog | Accessory factors for recombination | NLE | |
| RAD54B | RAD54 Homolog B | Accessory factors for recombination | Colon cancer, somatic Lymphoma, non-Hodgkin, somatic | |
| RAD54L | RAD54-Like | Accessory factors for recombination | Breast cancer, invasive ductal Adenocarcinoma, colonic, somatic Lymphoma, non-Hodgkin, somatic | |
| RBBP8 (CtIP) | Retinoblastoma-Binding Protein 8 | Promotes DNA end resection | Jawad syndrome | ● |
| Pancreatic carcinoma, somatic | ||||
| Seckel syndrome 2 | ● | |||
| SLX1A (GIYD1) | SLX1 Homolog A (GIY-YIG Domain Containing Protein 1) | Subunit of SLX1-SLX4 structure-specific nuclease, two identical tandem genes in the human genome | NLE | |
| SLX1B (GIYD2) | SLX1 Homolog B (GIY-YIG Domain Containing Protein 2) | Subunit of SLX1-SLX4 structure-specific nuclease, two identical tandem genes in the human genome | NLE | |
| SWI5 | SWI5 Homologous Recombination Repair Protein | Accessory factor for loading RAD51 | NLE | |
| XRCC2 | X-ray Repair Cross Complementing 2 | DNA break and crosslink repair | Fanconi anemia, complementation group U # Premature ovarian failure 17 # Spermatogenic failure | |
| XRCC3 | X-ray Repair Cross Complementing 3 | DNA break and crosslink repair | Susceptibility to breast cancer Melanoma, cutaneous malignant, 6 | |
| Fanconi Anemia Pathway | ||||
| BRCA2 (FANCD1) | BRCA2 Gene (Fanconi Anemia, Complementation Group D1) | Cooperation with RAD51, essential function | Susceptibility to breast cancer, male | |
| Breast-ovarian cancer, familial, 2 | ||||
| Glioblastoma 3 | ● | |||
| Medulloblastoma | ● | |||
| Pancreatic cancer 2 | ||||
| Prostate cancer | ||||
| Fanconi anemia, complementation group D1 | ||||
| Wilms tumor | ||||
| BRIP1 (FANCJ) | BRCA1-Interacting Protein 1 (Fanconi Anemia, Complementation Group J) | DNA helicase, BRCA1-interacting | Susceptibility to breast cancer, early-onset Fanconi anemia, complementation group J | |
| FAAP20 | Fanconi Anemia-Associated Protein, 20-KD Subunit (Chromosome 1 Open Reading Frame 86) | Tolerance and repair of DNA crosslinks and other adducts in DNA: FANCA-associated | NLE | |
| FAAP24 | FA Core Complex-Associated Protein 24 | Tolerance and repair of DNA crosslinks and other adducts in DNA: FAAP24 | NLE | |
| FAAP100 | Fanconi Anemia-Associated Protein, 100-KD Subunit | Part of FA core complex | NLE | |
| FANCA | Fanconi Anemia, Complementation Group A | Tolerance and repair of DNA crosslinks and other adducts in DNA: FANCA | Fanconi anemia, complementation group A | |
| FANCB | Fanconi Anemia, Complementation Group B | Tolerance and repair of DNA crosslinks and other adducts in DNA: FANCB | Fanconi anemia, complementation group B | |
| FANCC | Fanconi Anemia, Complementation Group C | Tolerance and repair of DNA crosslinks and other adducts in DNA: FANCC | Fanconi anemia, complementation group C | |
| FANCD2 | Fanconi Anemia, Complementation Group D2 | Target for monoubiquitination | Fanconi anemia, complementation group D2 | |
| FANCE | Fanconi Anemia, Complementation Group E | Tolerance and repair of DNA crosslinks and other adducts in DNA: FANCE | Fanconi anemia, complementation group E | |
| FANCG (XRCC9) | Fanconi Anemia, Complementation Group G (X-ray Repair Cross Complementing 9) | Tolerance and repair of DNA crosslinks and other adducts in DNA: FANCG | Fanconi anemia, complementation group G | |
| FANCI | Fanconi Anemia, Complementation Group I | Target for monoubiquitination | Fanconi anemia, complementation group I | |
| FANCL | Fanconi Anemia, Complementation Group L | Tolerance and repair of DNA crosslinks and other adducts in DNA: FANCL | Fanconi anemia, complementation group L | |
| FANCM | Fanconi Anemia, Complementation Group M | Helicase/translocase | Premature ovarian failure 15 # Spermatogenic failure 28 | |
| PALB2 (FANCN) | Partner and Localizer of BRCA2 (Fanconi Anemia, Complementation Group N) | Co-localizes with BRCA2 (FANCD1) | Susceptibility to breast cancer Susceptibility to pancreatic cancer, 3 Fanconi anemia, complementation group N | |
| RAD51C (FANCO) | RAD51 Paralog C (Fanconi Anemia, Complementation Group O) | Rad51 homolog, FANCO | Susceptibility to breast-ovarian cancer, familial, 3 Fanconi anemia, complementation group O | |
| SLX4 (FANCP) | SLX4 Structure-Specific Endonuclease Subunit (Fanconi Anemia, Complementation Group P) | Nuclease subunit/scaffold SLX4, FANCP | Fanconi anemia, complementation group P | |
| UBE2T (FANCT) | Ubiquitin-Conjugating Enzyme E2T (Fanconi Anemia, Complementation Group T) | E2 ligase for FANCL | Fanconi anemia, complementation group T | |
| DNA Damage Response Proteins | ||||
| ATM | Ataxia-Telangiectasia Mutated Gene | Ataxia telangiectasia | Susceptibility to breast cancer | |
| Ataxia-telangiectasia | ● | |||
| Lymphoma, B-cell non-Hodgkin, somatic | ||||
| Lymphoma, mantle cell, somatic | ||||
| T-cell prolymphocytic leukemia, somatic | ||||
| ATR | ATR Serine/Threonine Kinase | ATM- and PI-3K-like essential kinase | Cutaneous telangiectasia and cancer syndrome, familial # | |
| Seckel syndrome 1 | ● | |||
| ATRIP | ATR-Interacting Protein | ATR-interacting protein | NLE | |
| CHEK1 | Checkpoint Kinase 1 | Effector kinases | NLE | |
| CHEK2 | Checkpoint Kinase 2 | Effector kinases | Susceptibility to breast and colorectal cancer Susceptibility to breast cancer Susceptibility to prostate cancer, familial Li-Fraumeni syndrome Osteosarcoma, somatic | |
| MDC1 | Mediator of DNA Damage Checkpoint Protein 1 | Mediator of DNA damage checkpoint | NLE | |
| PARP1 (ADPRT) | Poly(ADP-Ribose) Polymerase 1 (ADP-Ribosyltransferase 1) | Protects strand interruptions | NLE | |
| PARP2 (ADPRT2) | Poly(ADP-Ribose) Polymerase 2 (ADP-Ribosyltransferase 2) | PARP-like enzyme | NLE | |
| PARP3 (ADPRT3) | Poly(ADP-Ribose) Polymerase 3 (ADP-Ribosyltransferase 3) | PARP-like enzyme | NLE | |
| TP53 | Tumor Protein 53 | Regulation of the cell cycle | Adrenocortical carcinoma, pediatric | |
| Basal cell carcinoma 7 | ||||
| Choroid plexus papilloma | ||||
| Colorectal cancer | ||||
| Glioma susceptibility 1 | ● | |||
| Osteosarcoma | ||||
| Bone marrow failure syndrome 5 | ||||
| Breast cancer, somatic | ||||
| Hepatocellular carcinoma, somatic | ||||
| Li-Fraumeni syndrome | ||||
| Nasopharyngeal carcinoma, somatic | ||||
| Pancreatic cancer, somatic | ||||
| TP53BP1 (53BP1) | Tumor Protein p53-Binding Protein 1 | Chromatin-binding checkpoint protein | NLE | |
| Accessory Proteins | ||||
| HUS1 | Hydroxyurea-Sensitive 1 | Subunits of PCNA-like sensor of damaged DNA | NLE | |
| PCNA | Proliferating Cell Nuclear Antigen | Sliding clamp for pol delta and pol epsilon | Ataxia-telangiectasia-like disorder 2 # | ● |
| RAD1 | RAD1 Checkpoint DNA Exonuclease | Subunits of PCNA-like sensor of damaged DNA | NLE | |
| RAD9A | RAD9A Checkpoint Clamp Component A | Subunits of PCNA-like sensor of damaged DNA | NLE | |
| RAD17 (RAD24) | RAD17 Checkpoint Clamp Loader Component (Homolog of RAD24) | RFC-like DNA damage sensor | NLE | |
| RPA1 | Replication Protein A1 | Binds DNA in preincision complex | NLE | |
| RPA2 | Replication Protein A2 | Binds DNA in preincision complex | NLE | |
| RPA3 | Replication Protein A3 | Binds DNA in preincision complex | NLE | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Scheijen, E.E.M.; Wilson, D.M., III. Genome Integrity and Neurological Disease. Int. J. Mol. Sci. 2022, 23, 4142. https://doi.org/10.3390/ijms23084142
Scheijen EEM, Wilson DM III. Genome Integrity and Neurological Disease. International Journal of Molecular Sciences. 2022; 23(8):4142. https://doi.org/10.3390/ijms23084142
Chicago/Turabian StyleScheijen, Elle E. M., and David M. Wilson, III. 2022. "Genome Integrity and Neurological Disease" International Journal of Molecular Sciences 23, no. 8: 4142. https://doi.org/10.3390/ijms23084142







