TcdB of Clostridioides difficile Mediates RAS-Dependent Necrosis in Epithelial Cells
Abstract
1. Introduction
2. Results
2.1. Morphological Changes after Treatment with Toxins
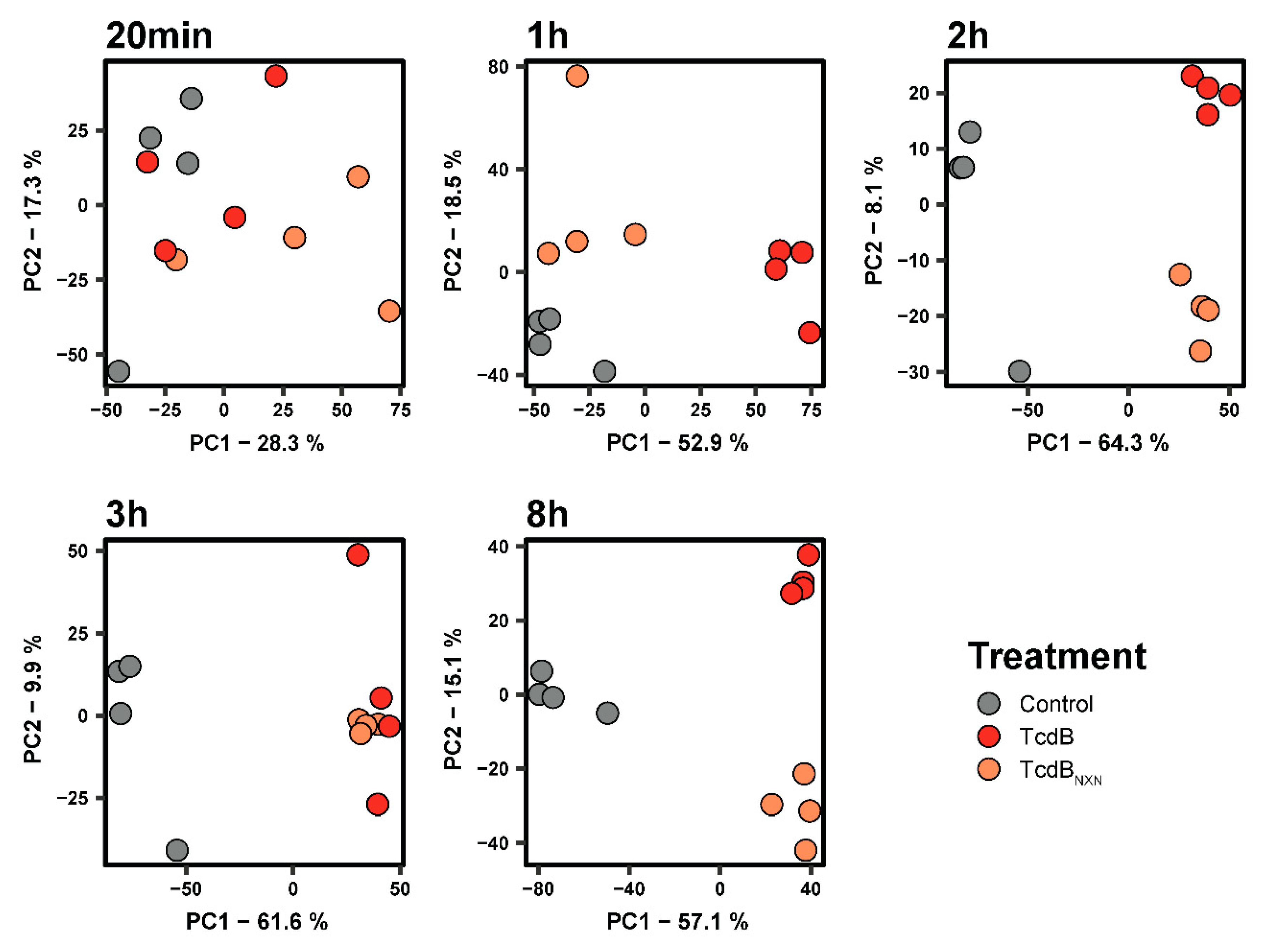
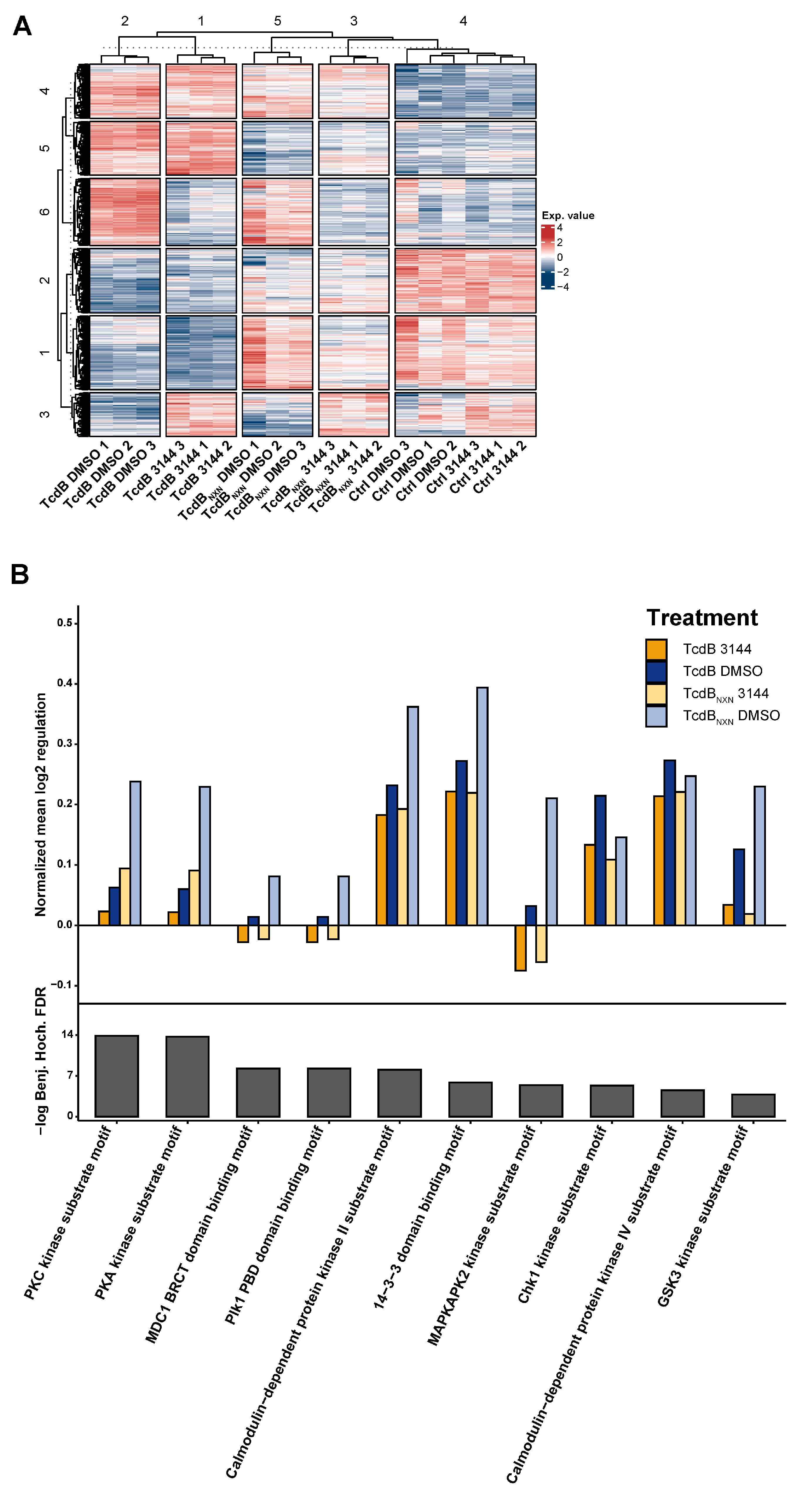
2.2. Comparative Phosphoproteomic Analysis
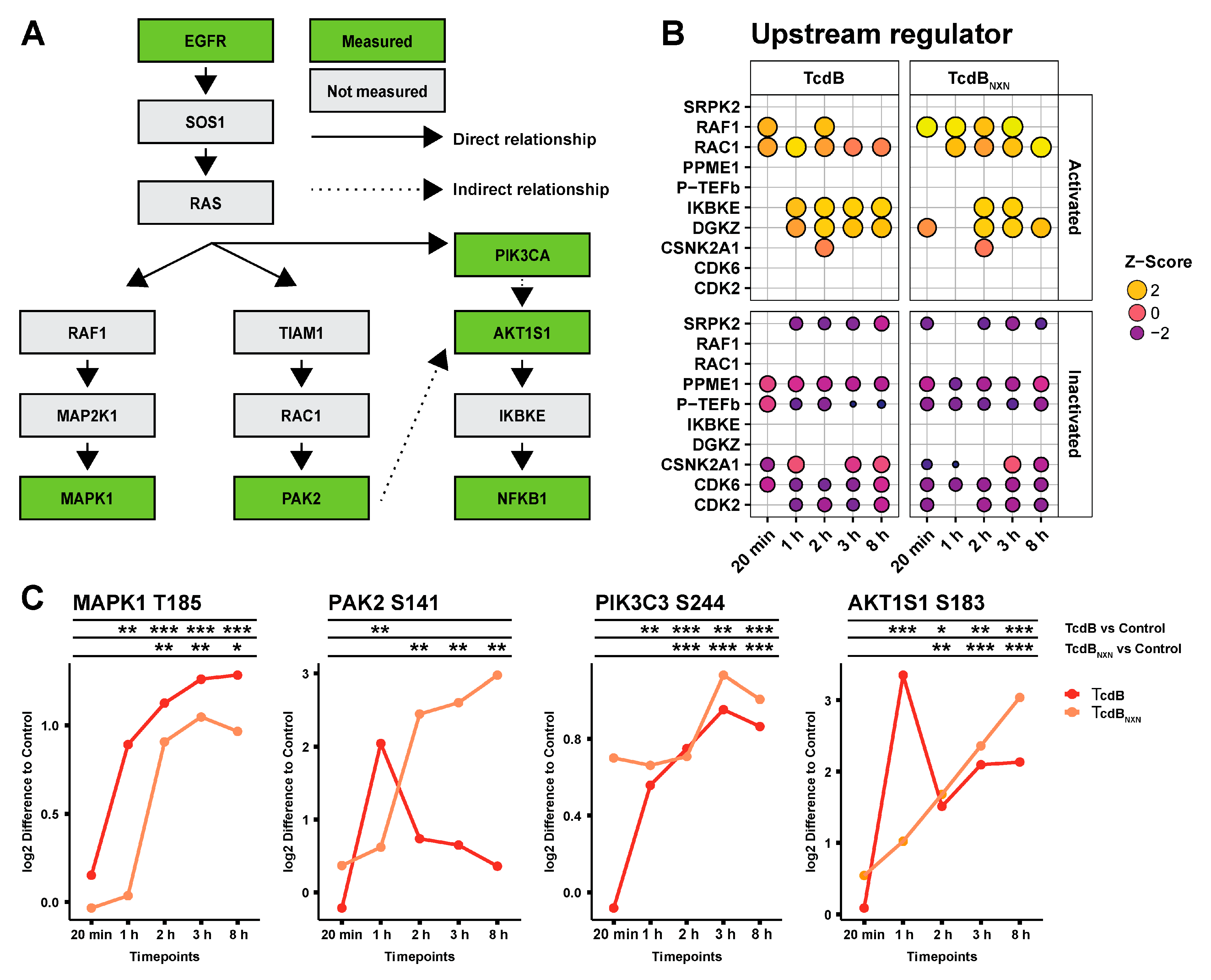
2.3. Kinetics of Phosphoproteome Alteration Point to RAS as Central Upstream Regulator
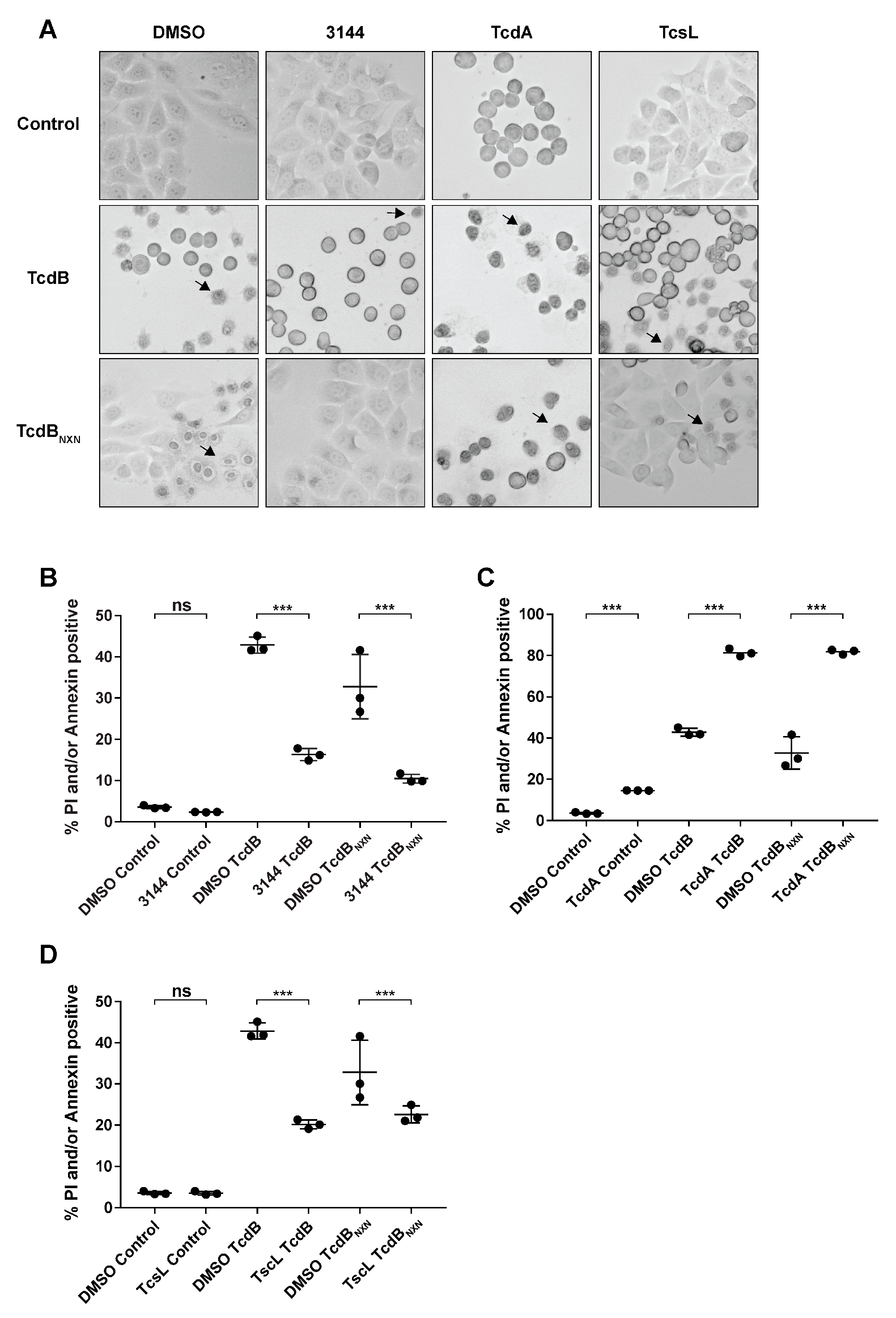
2.4. RAS Inhibition Led to a Reduction in Pyknosis
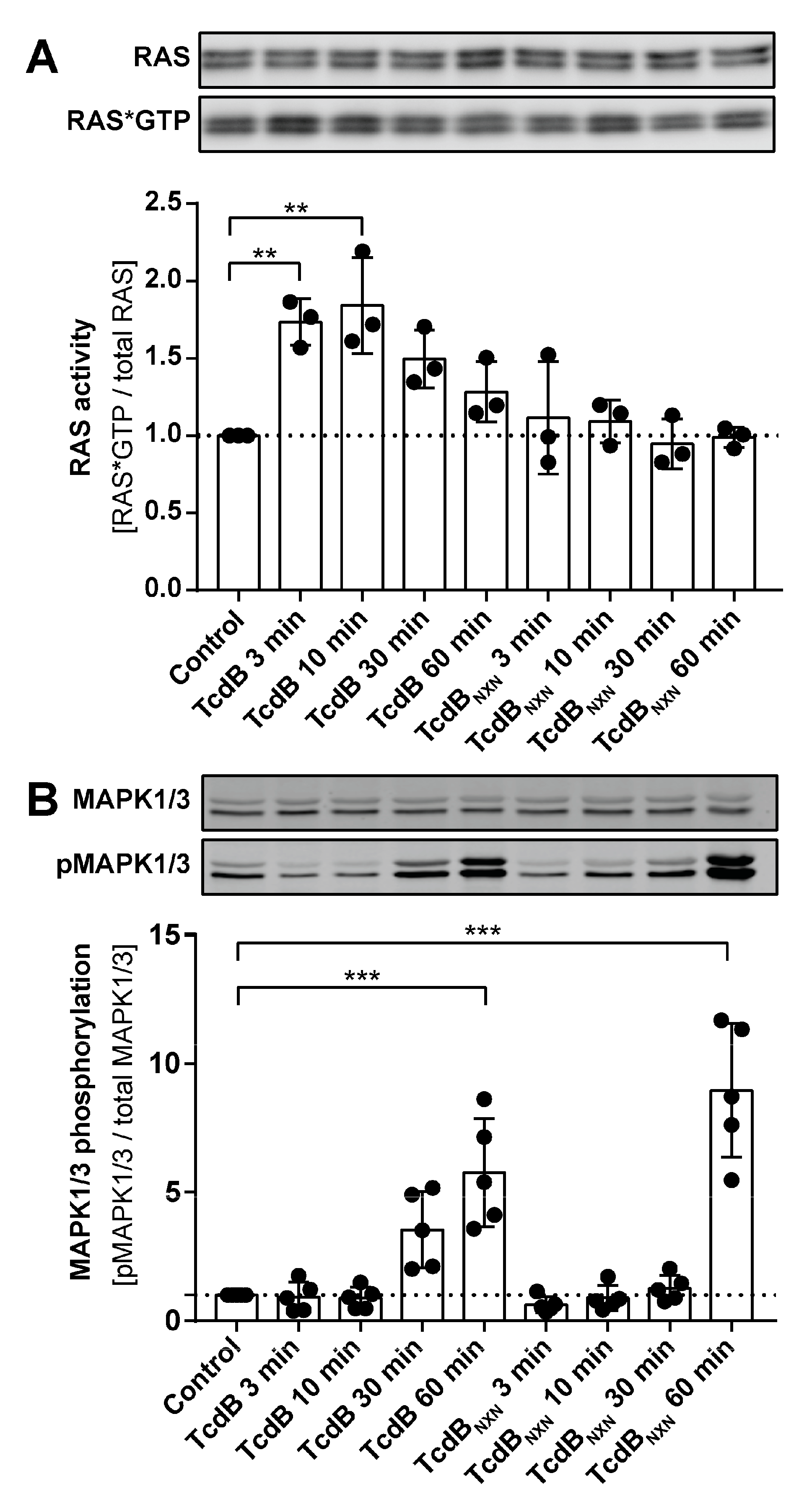
2.5. RAS Is Activated after TcdB Treatment
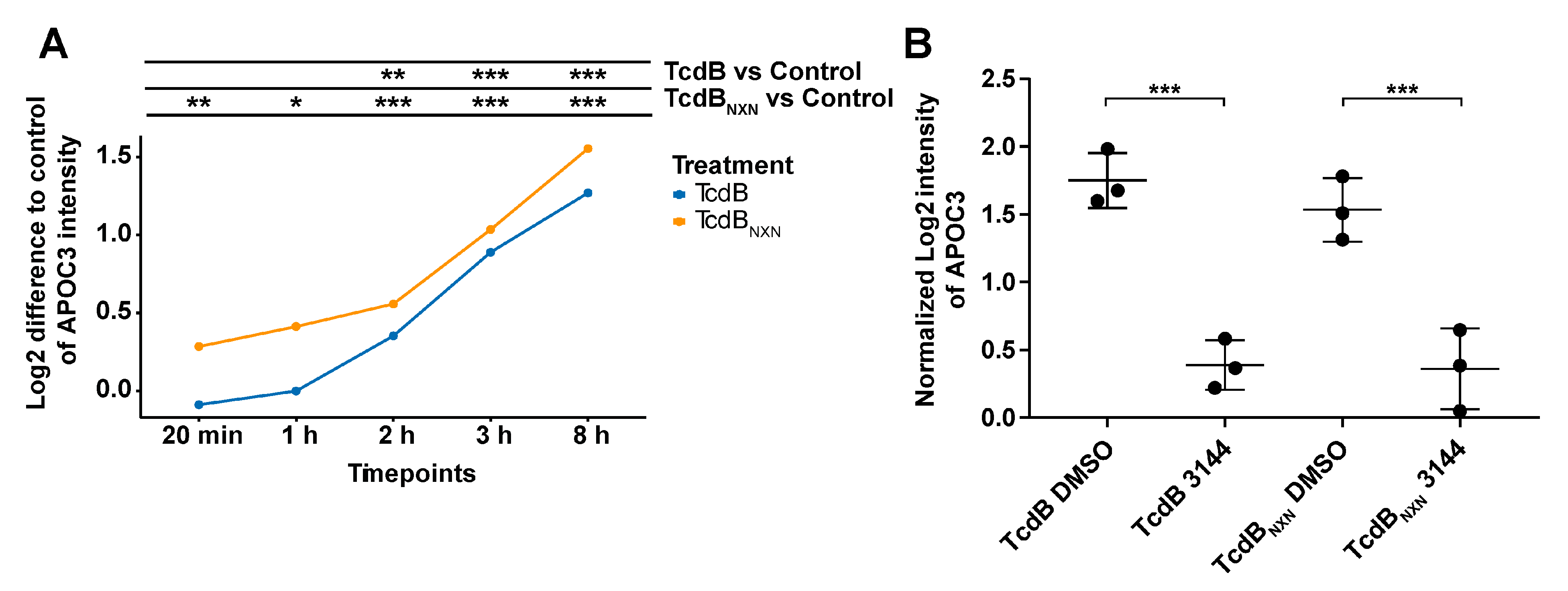
2.6. RAS Inhibition Influences Differences in Motif Phosphorylation and Reduced APOC3 Expression
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Protein Digestion
4.3. Phosphopeptide Enrichment and TMT Labeling
4.4. LC–MS Analysis
4.5. Flow Cytometry Analysis
4.6. DAPI Staining
4.7. RAS Activation Assay
4.8. Western Blotting
4.9. Data Processing
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Czepiel, J.; Dróżdż, M.; Pituch, H.; Kuijper, E.J.; Perucki, W.; Mielimonka, A.; Goldman, S.; Wultańska, D.; Garlicki, A.; Biesiada, G. Clostridium difficile infection: Review. Eur. J. Clin. Microbiol. Infect. Dis. 2019, 38, 1211–1221. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Antibiotic Resistance Threats in The United States. 2013. Available online: https://www.cdc.gov/drugresistance/threat-report-2013/pdf/ar-threats-2013-508.pdf (accessed on 14 June 2019).
- Aktories, K.; Schwan, C.; Jank, T. Clostridium difficile toxin biology. Ann. Rev. Microbiol. 2017, 71, 281–307. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Tian, S.; Zhang, J.; Liu, Z.; Robinson-McCarthy, L.; Miyashita, S.-I.; Breault, D.T.; Gerhard, R.; Oottamasathien, S.; Whelan, S.P.J.; et al. Sulfated glycosaminoglycans and low-density lipoprotein receptor contribute to Clostridium difficile toxin A entry into cells. Nat. Microbiol. 2019, 4, 1760–1769. [Google Scholar] [CrossRef] [PubMed]
- Na, X.; Kim, H.; Moyer, M.P.; Pothoulakis, C.; LaMont, J.T. gp96 is a human colonocyte plasma membrane binding protein for Clostridium difficile toxin A. Infect. Immun. 2008, 76, 2862–2871. [Google Scholar] [CrossRef]
- Schöttelndreier, D.; Langejürgen, A.; Lindner, R.; Genth, H. Low density lipoprotein receptor-related protein-1 (LRP1) is involved in the uptake of clostridioides difficile toxin A and serves as an internalizing receptor. Front. Cell. Infect. Microbiol. 2020, 10, 565465. [Google Scholar] [CrossRef]
- Yuan, P.; Zhang, H.; Cai, C.; Zhu, S.; Zhou, Y.; Yang, X.; He, R.; Li, C.; Guo, S.; Li, S.; et al. Chondroitin sulfate proteoglycan 4 functions as the cellular receptor for Clostridium difficile toxin B. Cell Res. 2015, 25, 157–168. [Google Scholar] [CrossRef]
- Tao, L.; Zhang, J.; Meraner, P.; Tovaglieri, A.; Wu, X.; Gerhard, R.; Zhang, X.; Stallcup, W.B.; Miao, J.; He, X.; et al. Frizzled are colonic epithelial receptors for Clostridium difficile toxin B. Nature 2016, 538, 350–355. [Google Scholar] [CrossRef]
- LaFrance, M.E.; Farrow, M.A.; Chandrasekaran, R.; Sheng, J.; Rubin, D.H.; Lacy, D.B. Identification of an epithelial cell receptor responsible for Clostridium difficile TcdB-induced cytotoxicity. Proc. Natl. Acad. Sci. USA 2015, 112, 7073–7078. [Google Scholar] [CrossRef]
- Orrell, K.E.; Zhang, Z.; Sugiman-Marangos, S.N.; Melnyk, R.A. Clostridium difficile toxins A and B: Receptors, pores, and translocation into cells. Crit. Rev. Biochem. Mol. Biol. 2017, 52, 461–473. [Google Scholar] [CrossRef]
- Zeiser, J.; Gerhard, R.; Just, I.; Pich, A. Substrate specificity of clostridial glucosylating toxins and their function on colonocytes analyzed by proteomics techniques. J. Proteome Res. 2013, 12, 1604–1618. [Google Scholar] [CrossRef]
- Jochim, N.; Gerhard, R.; Just, I.; Pich, A. Time-resolved cellular effects induced by TcdA from Clostridium difficile. Rapid Commun. Mass Spectrom. 2014, 28, 1089–1100. [Google Scholar] [CrossRef] [PubMed]
- Junemann, J.; Lämmerhirt, C.M.; Polten, F.; Just, I.; Gerhard, R.; Genth, H.; Pich, A. Quantification of small GTPase glucosylation by clostridial glucosylating toxins using multiplexed MRM analysis. Proteomics 2017, 17, 1700016. [Google Scholar] [CrossRef] [PubMed]
- Erdmann, J.; Junemann, J.; Schröder, A.; Just, I.; Gerhard, R.; Pich, A. Glucosyltransferase-dependent and -independent effects of TcdB on the proteome of HEp-2 cells. Proteomics 2017, 17, 1600435. [Google Scholar] [CrossRef]
- Junemann, J.; Birgin, G.; Erdmann, J.; Schröder, A.; Just, I.; Gerhard, R.; Pich, A. Toxin A of the nosocomial pathogen Clostridium difficile induces primary effects in the proteome of HEp-2 cells. Proteomics Clin. Appl. 2017, 11, 1600031. [Google Scholar] [CrossRef] [PubMed]
- Wohlan, K.; Goy, S.; Olling, A.; Srivaratharajan, S.; Tatge, H.; Genth, H.; Gerhard, R. Pyknotic cell death induced by Clostridium difficile TcdB: Chromatin condensation and nuclear blister are induced independently of the glucosyltransferase activity. Cell. Microbiol. 2014, 16, 1678–1692. [Google Scholar] [CrossRef] [PubMed]
- Chumbler, N.M.; Farrow, M.A.; Lapierre, L.A.; Franklin, J.L.; Haslam, D.B.; Haslam, D.; Goldenring, J.R.; Lacy, D.B. Clostridium difficile Toxin B causes epithelial cell necrosis through an autoprocessing-independent mechanism. PLoS Pathog. 2012, 8, e1003072. [Google Scholar] [CrossRef]
- Bilverstone, T.W.; Garland, M.; Cave, R.J.; Kelly, M.L.; Tholen, M.; Bouley, D.M.; Kaye, P.; Minton, N.P.; Bogyo, M.; Kuehne, S.A.; et al. The glucosyltransferase activity of C. difficile Toxin B is required for disease pathogenesis. PLoS Pathog. 2020, 16, e1008852. [Google Scholar] [CrossRef]
- Wadhwa, A.; Al Nahhas, M.F.; Dierkhising, R.A.; Patel, R.; Kashyap, P.; Pardi, D.S.; Khanna, S.; Grover, M. High risk of post-infectious irritable bowel syndrome in patients with Clostridium difficile infection. Aliment. Pharmacol. Ther. 2016, 44, 576–582. [Google Scholar] [CrossRef]
- Kordus, S.L.; Thomas, A.K.; Lacy, D.B. Clostridioides difficile toxins: Mechanisms of action and antitoxin therapeutics. Nat. Rev. Microbiol. 2022, 20, 285–298. [Google Scholar] [CrossRef]
- Farrow, M.A.; Chumbler, N.M.; Lapierre, L.A.; Franklin, J.L.; Rutherford, S.A.; Goldenring, J.R.; Lacy, D.B. Clostridium difficile toxin B-induced necrosis is mediated by the host epithelial cell NADPH oxidase complex. Proc. Natl. Acad. Sci. USA 2013, 110, 18674–18679. [Google Scholar] [CrossRef]
- Beer, L.-A.; Tatge, H.; Reich, N.; Tenspolde, M.; Olling, A.; Goy, S.; Rottner, K.; Alekov, A.K.; Gerhard, R. Early cell death induced by Clostridium difficile TcdB: Uptake and Rac1-glucosylation kinetics are decisive for cell fate. Cell. Microbiol. 2018, 20, e12865. [Google Scholar] [CrossRef] [PubMed]
- Zewinger, S.; Reiser, J.; Jankowski, V.; Alansary, D.; Hahm, E.; Triem, S.; Klug, M.; Schunk, S.J.; Schmit, D.; Kramann, R.; et al. Apolipoprotein C3 induces inflammation and organ damage by alternative inflammasome activation. Nat. Immunol. 2020, 21, 30–41. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, Y.; Sakimura, A.; Park, C.M.; Tomaru, R.; Tanaka, T.; Ozawa, T.; Zhou, Y.; Narita, K.; Kishi, H.; Muraguchi, A.; et al. Feedback control of ErbB2 via ERK-mediated phosphorylation of a conserved threonine in the juxtamembrane domain. Sci. Rep. 2016, 6, 31502. [Google Scholar] [CrossRef] [PubMed]
- Cargnello, M.; Roux, P.P. Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol. Mol. Biol. Rev. 2011, 75, 50–83. [Google Scholar] [CrossRef] [PubMed]
- Junemann, J.; Just, I.; Gerhard, R.; Pich, A. Quantitative phosphoproteome analysis of clostridioides difficile toxin B treated human epithelial cells. Front. Microbiol. 2018, 9, 3083. [Google Scholar] [CrossRef] [PubMed]
- Suryadinata, R.; Sadowski, M.; Sarcevic, B. Control of cell cycle progression by phosphorylation of cyclin-dependent kinase (CDK) substrates. Biosci. Rep. 2010, 30, 243–255. [Google Scholar] [CrossRef]
- Yang, Z.; He, N.; Zhou, Q. Brd4 recruits P-TEFb to chromosomes at late mitosis to promote G1 gene expression and cell cycle progression. Mol. Cell. Biol. 2008, 28, 967–976. [Google Scholar] [CrossRef]
- Li, X.; Yang, S.; Zhang, M.; Xie, S.; Xie, Z. Downregulation of SRPK2 promotes cell cycle arrest though E2F1 in non-small cell lung cancer. Eur. J. Histochem. 2019, 63, 3067. [Google Scholar] [CrossRef]
- Fettucciari, K.; Macchioni, L.; Davidescu, M.; Scarpelli, P.; Palumbo, C.; Corazzi, L.; Marchegiani, A.; Cerquetella, M.; Spaterna, A.; Marconi, P.; et al. Clostridium difficile toxin B induces senescence in enteric glial cells: A potential new mechanism of Clostridium difficile pathogenesis. Biochim. Biophys. Acta Mol. Cell Res. 2018, 1865, 1945–1958. [Google Scholar] [CrossRef]
- Yin, M.; Wang, X.; Lu, J. Advances in IKBKE as a potential target for cancer therapy. Cancer Med. 2020, 9, 247–258. [Google Scholar] [CrossRef]
- Li, Y.; Xu, S.; Xu, Q.; Chen, Y. Clostridium difficile toxin B induces colonic inflammation through the TRIM46/DUSP1/MAPKs and NF-κB signalling pathway. Artif. Cells Nanomed. Biotechnol. 2020, 48, 452–462. [Google Scholar] [CrossRef] [PubMed]
- Kyriakis, J.M.; App, H.; Zhang, X.F.; Banerjee, P.; Brautigan, D.L.; Rapp, U.R.; Avruch, J. Raf-1 activates MAP kinase-kinase. Nature 1992, 358, 417–421. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Pease, D.R.; Chen, S.; Zhang, B.; Phee, H. P21-activated kinase 2 is essential in maintenance of peripheral Foxp3+ regulatory T cells. Immunology 2018, 154, 309–321. [Google Scholar] [CrossRef] [PubMed]
- Nascimento, E.B.M.; Snel, M.; Guigas, B.; van der Zon, G.C.M.; Kriek, J.; Maassen, J.A.; Jazet, I.M.; Diamant, M.; Ouwens, D.M. Phosphorylation of PRAS40 on Thr246 by PKB/AKT facilitates efficient phosphorylation of Ser183 by mTORC1. Cell. Signal. 2010, 22, 961–967. [Google Scholar] [CrossRef]
- Porta, C.; Paglino, C.; Mosca, A. Targeting PI3K/Akt/mTOR Signaling in Cancer. Front. Oncol. 2014, 4, 64. [Google Scholar] [CrossRef]
- Genth, H.; Junemann, J.; Lämmerhirt, C.M.; Lücke, A.-C.; Schelle, I.; Just, I.; Gerhard, R.; Pich, A. Difference in mono-o-glucosylation of ras subtype GTPases between toxin A and toxin B from clostridioides difficile strain 10463 and lethal toxin from clostridium sordellii strain 6018. Front. Microbiol. 2018, 9, 3078. [Google Scholar] [CrossRef]
- Chumbler, N.M.; Farrow, M.A.; Lapierre, L.A.; Franklin, J.L.; Lacy, D.B. Clostridium difficile toxins TcdA and TcdB cause colonic tissue damage by distinct mechanisms. Infect. Immun. 2016, 84, 2871–2877. [Google Scholar] [CrossRef]
- Redza-Dutordoir, M.; Averill-Bates, D.A. Activation of apoptosis signalling pathways by reactive oxygen species. Biochim. Biophys. Acta 2016, 1863, 2977–2992. [Google Scholar] [CrossRef]
- Rusanescu, G.; Gotoh, T.; Tian, X.; Feig, L.A. Regulation of Ras signaling specificity by protein kinase C. Mol. Cell. Biol. 2001, 21, 2650–2658. [Google Scholar] [CrossRef][Green Version]
- Liu, Z.; Zhang, S.; Chen, P.; Tian, S.; Zeng, J.; Perry, K.; Dong, M.; Jin, R. Structural basis for selective modification of Rho and Ras GTPases by Clostridioides difficile toxin B. Sci. Adv. 2021, 7, eabi4582. [Google Scholar] [CrossRef]
- Huelsenbeck, J.; Dreger, S.; Gerhard, R.; Barth, H.; Just, I.; Genth, H. Difference in the cytotoxic effects of toxin B from Clostridium difficile strain VPI 10463 and toxin B from variant Clostridium difficile strain 1470. Infect. Immun. 2007, 75, 801–809. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Yang, C.; Guo, G.; Li, N.; Yu, W. Motif-All: Discovering all phosphorylation motifs. BMC Bioinform. 2011, 12, S22. [Google Scholar] [CrossRef] [PubMed]
- Molina, J.R.; Adjei, A.A. The Ras/Raf/MAPK pathway. J. Thorac. Oncol. 2006, 1, 7–9. [Google Scholar] [CrossRef]
- McCubrey, J.A.; Steelman, L.S.; Bertrand, F.E.; Davis, N.M.; Sokolosky, M.; Abrams, S.L.; Montalto, G.; D’Assoro, A.B.; Libra, M.; Nicoletti, F.; et al. GSK-3 as potential target for therapeutic intervention in cancer. Oncotarget 2014, 5, 2881–2911. [Google Scholar] [CrossRef] [PubMed]
- Farrow, M.A.; Chumber, N.M.; Bloch, S.C.; King, M.; Moton-Melancon, K.; Shupe, J.; Washington, M.K.; Spiller, B.W.; Lacy, D.B. Small molecule inhibitor screen reveals calcium channel signaling as a mechanistic mediator of Clostridium difficile TcdB-induced necrosis. ACS Chem. Biol. 2020, 15, 1212–1221. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, A.; Contreras, C.; Sánchez, A.; Prieto, D. Role of Phosphatidylinositol 3-Kinase (PI3K), Mitogen-Activated Protein Kinase (MAPK), and Protein Kinase C (PKC) in calcium signaling pathways linked to the α1-adrenoceptor in resistance arteries. Front. Physiol. 2019, 10, 55. [Google Scholar] [CrossRef]
- Stewart, G.S.; Wang, B.; Bignell, C.R.; Taylor, A.M.R.; Elledge, S.J. MDC1 is a mediator of the mammalian DNA damage checkpoint. Nature 2003, 421, 961–966. [Google Scholar] [CrossRef]
- Stieglitz, F.; Gerhard, R.; Pich, A. The binary toxin of clostridioides difficile alters the proteome and phosphoproteome of HEp-2 cells. Front. Microbiol. 2021, 12, 725612. [Google Scholar] [CrossRef]
- Mangan, M.S.; Gorki, F.; Krause, K.; Heinz, A.; Ebert, T.; Jahn, D.; Hiller, K.; Hornung, V.; Mauer, M.; Schmidt, F.I.; et al. Clostridium difficile toxin B activates the NLRP3 inflammasome in human macrophages, demonstrating a novel regulatory mechanism for the Pyrin inflammasome. bioRxiv 2021. [Google Scholar] [CrossRef]
- Beer, L.-A.; Tatge, H.; Schneider, C.; Ruschig, M.; Hust, M.; Barton, J.; Thiemann, S.; Fühner, V.; Russo, G.; Gerhard, R. The binary toxin CDT of Clostridium difficile as a tool for intracellular delivery of bacterial glucosyltransferase domains. Toxins 2018, 10, 225. [Google Scholar] [CrossRef]
- Zecha, J.; Satpathy, S.; Kanashova, T.; Avanessian, S.C.; Kane, M.H.; Clauser, K.R.; Mertins, P.; Carr, S.A.; Kuster, B. TMT Labeling for the masses: A robust and cost-efficient, in-solution labeling approach. Mol. Cell. Proteom. 2019, 18, 1468–1478. [Google Scholar] [CrossRef] [PubMed]
- Dreissigacker, U.; Mueller, M.S.; Unger, M.; Siegert, P.; Genze, F.; Gierschik, P.; Giehl, K. Oncogenic K-Ras down-regulates Rac1 and RhoA activity and enhances migration and invasion of pancreatic carcinoma cells through activation of p38. Cell. Signal. 2006, 18, 1156–1168. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized P.P.B.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium; Bateman, A.; Martin, M.-J.; Orchard, S.; Magrane, M.; Agivetova, R.; Ahmad, S.; Alpi, E.; Bowler-Barnett, E.H.; Britto, R.; et al. UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res. 2021, 49, D480–D489. [Google Scholar] [CrossRef]
- Tyanova, S.; Temu, T.; Sinitcyn, P.; Carlson, A.; Hein, M.Y.; Geiger, T.; Mann, M.; Cox, J. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 2016, 13, 731–740. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing. 2013. Available online: https://www.r-project.org (accessed on 28 January 2022).
- Gu, Z.; Eils, R.; Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 2016, 32, 2847–2849. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: Berlin/Heidelberg, Germany, 2016; ISBN 9783319242774. [Google Scholar]
- Jensen, L.J.; Kuhn, M.; Stark, M.; Chaffron, S.; Creevey, C.; Muller, J.; Doerks, T.; Julien, P.; Roth, A.; Simonovic, M.; et al. STRING 8—A global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res. 2009, 37, D412–D416. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Vizcaíno, J.A.; Côté, R.G.; Csordas, A.; Dianes, J.A.; Fabregat, A.; Foster, J.M.; Griss, J.; Alpi, E.; Birim, M.; Contell, J.; et al. The PRoteomics IDEntifications (PRIDE) database and associated tools: Status in 2013. Nucleic Acids Res. 2013, 41, D1063–D1069. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stieglitz, F.; Gerhard, R.; Hönig, R.; Giehl, K.; Pich, A. TcdB of Clostridioides difficile Mediates RAS-Dependent Necrosis in Epithelial Cells. Int. J. Mol. Sci. 2022, 23, 4258. https://doi.org/10.3390/ijms23084258
Stieglitz F, Gerhard R, Hönig R, Giehl K, Pich A. TcdB of Clostridioides difficile Mediates RAS-Dependent Necrosis in Epithelial Cells. International Journal of Molecular Sciences. 2022; 23(8):4258. https://doi.org/10.3390/ijms23084258
Chicago/Turabian StyleStieglitz, Florian, Ralf Gerhard, Rabea Hönig, Klaudia Giehl, and Andreas Pich. 2022. "TcdB of Clostridioides difficile Mediates RAS-Dependent Necrosis in Epithelial Cells" International Journal of Molecular Sciences 23, no. 8: 4258. https://doi.org/10.3390/ijms23084258
APA StyleStieglitz, F., Gerhard, R., Hönig, R., Giehl, K., & Pich, A. (2022). TcdB of Clostridioides difficile Mediates RAS-Dependent Necrosis in Epithelial Cells. International Journal of Molecular Sciences, 23(8), 4258. https://doi.org/10.3390/ijms23084258





