Dimerisation of the Yeast K+ Translocation Protein Trk1 Depends on the K+ Concentration
Abstract
1. Introduction
2. Results
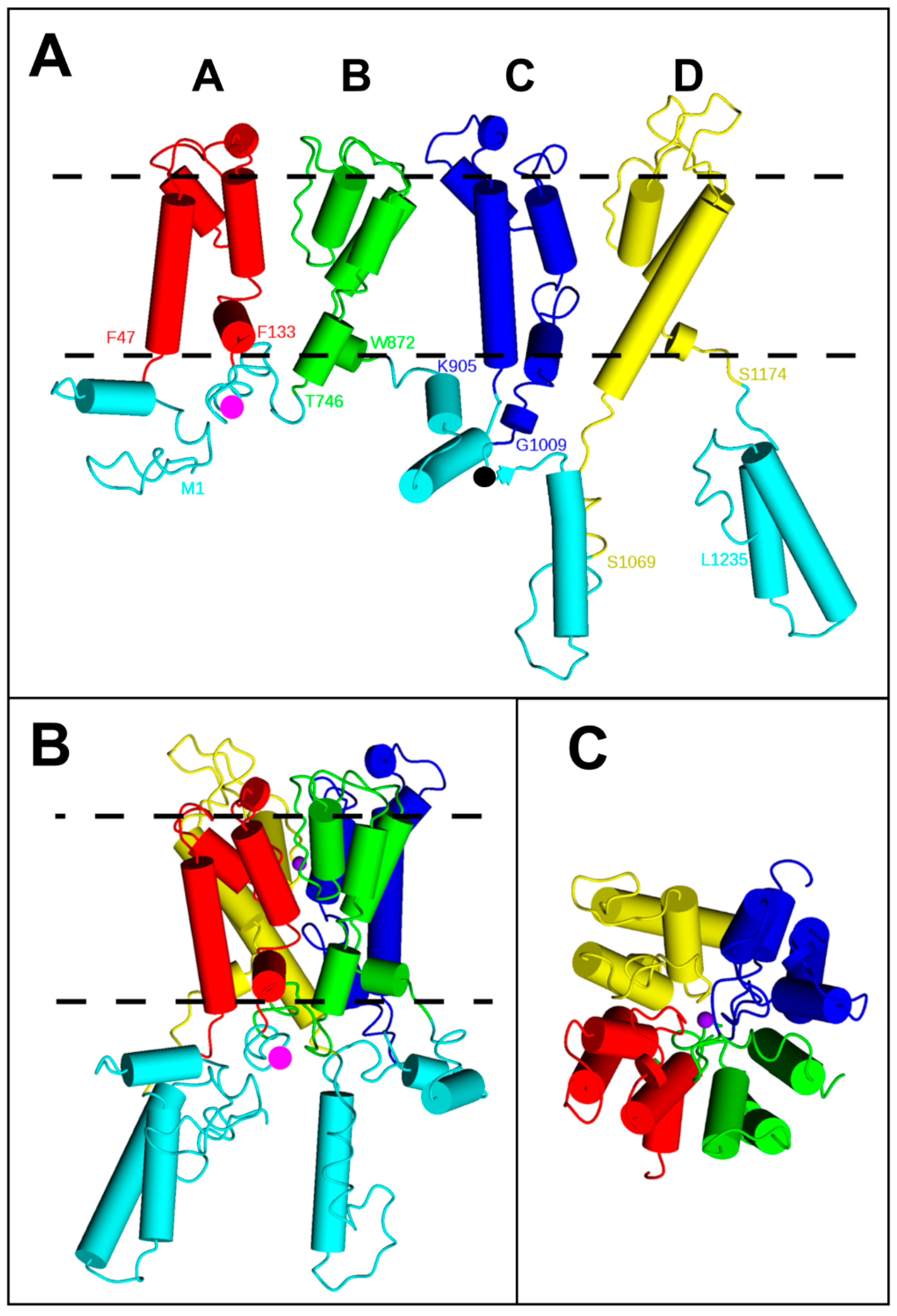
2.1. BiFC Indicates That Trk1[ΔLHL] Can Exist as a Dimer with an Interface Formed by MPMs C and D
2.2. Single Molecule Fluorescence Microscopy and Stepwise Photobleaching
2.3. Quantification of BiFC Fluorescence
3. Discussion
4. Material and Methods
4.1. Strains and Growth Conditions
4.2. Plasmid Construction and Yeast Transformation
4.3. Fluorescence Microscopy
4.4. Single Molecule Fluorescence Microscopy (SMFM)
4.5. Quantification of BiFC
4.6. Molecular Modelling and MD Simulations
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bertl, A.; Ramos, J.; Ludwig, J.; Lichtenberg-Fraté, H.; Reid, J.; Bihler, H.; Calero, F.; Martinez, P.; Ljungdahl, P.O. Characterization of potassium transport in wild-type and isogenic yeast strains carrying all combinations of trk1, trk2 and tok1 null mutations. Mol. Microbiol. 2003, 47, 767–780. [Google Scholar] [CrossRef] [PubMed]
- Gaber, R.F.; Styles, C.A.; Fink, G.R. TRK1 encodes a plasma membrane protein required for high-affinity potassium transport in Saccharomyces cerevisiae. Mol. Cell. Biol. 1988, 8, 2848–2859. [Google Scholar] [CrossRef] [PubMed]
- Ramos, J.; Alijo, R.; Haro, R.; Rodriguez-Navarro, A. TRK2 is not a low-affinity potassium transporter in Saccharomyces cerevisiae. J. Bacteriol. 1994, 176, 249–252. [Google Scholar] [CrossRef] [PubMed]
- Petrezsélyová, S.; Ramos, J.; Sychrová, H. Trk2 transporter is a relevant player in K+ supply and plasma-membrane potential control in Saccharomyces cerevisiae. Folia Microbiol. 2011, 56, 23–28. [Google Scholar] [CrossRef]
- Zimmermannová, O.; Felcmanová, K.; Rosas-Santiago, P.; Papoušková, K.; Pantoja, O.; Sychrová, H. Erv14 cargo receptor participates in regulation of plasma-membrane potential, intracellular pH and potassium homeostasis via its interaction with K+-specific transporters Trk1 and Tok1. Biochim. Et Biophys. Acta Mol. Cell Res. 2019, 1866, 1376–1388. [Google Scholar] [CrossRef]
- Zimmermannova, O.; Felcmanova, K.; Sacka, L.; Colinet, A.-S.; Morsomme, P.; Sychrova, H. K+-specific importers Trk1 and Trk2 play different roles in Ca2+ homeostasis and signalling in Saccharomyces cerevisiae cells. FEMS Yeast Res. 2021, 21, 1–12. [Google Scholar] [CrossRef]
- Diskowski, M.; Mikusevic, V.; Stock, C.; Hänelt, I. Functional diversity of the superfamily of K+ transporters to meet various requirements. Biol. Chem. 2015, 396, 1003–1014. [Google Scholar] [CrossRef]
- Huang, C.; Pedersen, B.P.; Stokes, D.L. Crystal Structure of the Potassium Importing KdpFABC Membrane Complex. Nature 2017, 546, 681–685. [Google Scholar] [CrossRef]
- Stock, C.; Hielkema, L.; Tascón, I.; Wunnicke, D.; Oostergetel, G.T.; Azkargorta, M.; Paulino, C.; Hänelt, I. Cryo-EM structures of KdpFABC suggest a K+ transport mechanism via two inter-subunit half-channels. Nat. Commun. 2018, 9, 1–10. [Google Scholar] [CrossRef]
- Durell, S.R.; Guy, H.R. Structural models of the KtrB, TrkH, and Trk1,2 symporters based on the structure of the KcsA K+ channel. Biophys. J. 1999, 77, 789–807. [Google Scholar] [CrossRef]
- Durell, S.R.; Hao, Y.; Nakamura, T.; Bakker, E.P.; Guy, H.R. Evolutionary relationship between K+ channels and symporters. Biophys. J. 1999, 77, 775–788. [Google Scholar] [CrossRef] [PubMed]
- Doyle, D.A.; Cabral, J.M.; Pfuetzner, R.A.; Kuo, A.; Gulbis, J.M.; Cohen, S.L.; Chait, B.T.; MacKinnon, R. The Structure of the PotassiumChannel: Molecular Basis of K+ Conduction and Selectivity. Science 1998, 280, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kale, D.; Spurny, P.; Shamayeva, K.; Spurna, K.; Kahoun, D.; Ganser, D.; Zayats, V.; Ludwig, J. The S. cerevisiae cation translocation protein Trk1 is functional without its “long hydrophilic loop” but LHL regulates cation translocation activity and selectivity. Biochim. Et Biophys. Acta Biomembr. 2019, 1861, 1476–1488. [Google Scholar] [CrossRef] [PubMed]
- Zayats, V.; Stockner, T.; Pandey, S.K.; Wörz, K.; Ettrich, R.; Ludwig, J. A refined atomic scale model of the Saccharomyces cerevisiae K+-translocation protein Trk1p combined with experimental evidence confirms the role of selectivity filter glycines and other key residues. Biochim. Et Biophys. Acta Biomembr. 2015, 1848, 1183–1195. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cao, Y.; Jin, X.; Huang, H.; Derebe, M.G.; Levin, E.J.; Kabaleeswaran, V.; Pan, Y.; Punta, M.; Love, J.; Weng, J.; et al. Crystal structure of a potassium ion transporter, TrkH. Nature 2011, 471, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Vieira-Pires, R.S.; Szollosi, A.; Morais-Cabral, J.H. The structure of the KtrAB potassium transporter. Nature 2013, 496, 323–328. [Google Scholar] [CrossRef]
- Kuroda, T.; Bihler, H.; Bashi, E.; Slayman, C.L.; Rivetta, A. Chloride channel function in the yeast TRK-potassium transporters. J. Membr. Biol. 2004, 198, 177–192. [Google Scholar] [CrossRef]
- Pardo, J.P.; González-Andrade, M.; Allen, K.; Kuroda, T.; Slayman, C.L.; Rivetta, A. A structural model for facultative anion channels in an oligomeric membrane protein: The yeast TRK (K+) system. Pflug. Arch. Eur. J. Physiol. 2015, 467, 2447–2460. [Google Scholar] [CrossRef]
- Ariño, J.; Aydar, E.; Drulhe, S.; Ganser, D.; Jorrín, J.; Kahm, M.; Krause, F.; Petrezsélyová, S.; Yenush, L.; Zimmermannová, O. Systems Biology of Monovalent Cation Homeostasis in Yeast. The Translucent Contribution. Adv. Microb. Physiol. 2014, 64, 1–63. [Google Scholar] [CrossRef]
- Ramos, J.; Haro, R.; Rodriguez-Navarro, A. Regulation of potassium fluxes in Saccharomyces cerevisiae. BBA Biomembr. 1990, 1029, 211–217. [Google Scholar] [CrossRef]
- Ramos, J.; Rodriguez-Navarro, A. Regulation and interconversion of the potassium transport systems of Saccharomyces cerevisiae as revealed by rubidium transport. Eur. J. Biochem. 1986, 154, 307–311. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Navarro, A.; Ramos, J. Dual System for potassium transport in Saccharomyces cerevisiae. J. Bacteriol. 1984, 159, 940–945. [Google Scholar] [CrossRef] [PubMed]
- Masaryk, J.; Sychrová, H. Yeast Trk1 Potassium Transporter Gradually Changes Its Affinity in Response to Both External and Internal Signals. J. Fungi 2022, 8, 432. [Google Scholar] [CrossRef] [PubMed]
- Boles, E.; Hollenberg, C.P. The molecular genetics of hexose transport in yeasts. FEMS Microbiol. Rev. 1997, 21, 85–111. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-H.; Roy, A.; Jouandot, D.; Cho, K.H. The glucose signaling network in yeast. Biochim. Et Biophys. Acta Gen. Subj. 2013, 1830, 5204–5210. [Google Scholar] [CrossRef]
- Pacheco, A.; Donzella, L.; Hernandez-Lopez, M.J.; Almeida, M.J.; Prieto, J.A.; Randez-Gil, F.; Morrissey, J.P.; Sousa, M.J. Hexose transport in Torulaspora delbrueckii: Identification of Igt1, a new dual-affinity transporter. FEMS Yeast Res. 2020, 20, 1–10. [Google Scholar] [CrossRef]
- Fu, H.H.; Luan, S. AtKUP1: A dual-affinity K+ transporter from Arabidopsis. Plant Cell 1998, 10, 63–73. [Google Scholar] [CrossRef]
- Kim, E.J.; Kwak, J.M.; Uozumi, N.; Schroeder, J.I. AtKUP1: An arabidopsis gene encoding high-affinity potassium transport activity. Plant Cell 1998, 10, 51–62. [Google Scholar] [CrossRef]
- Tsay, Y.F.; Schroeder, J.I.; Feldmann, K.A.; Crawford, N.M. The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell 1993, 72, 705–713. [Google Scholar] [CrossRef]
- Sun, J.; Bankston, J.R.; Payandeh, J.; Hinds, T.R.; Zagotta, W.N.; Zheng, N. Crystal structure of the plant dual-affinity nitrate transporter NRT1.1. Nature 2014, 507, 73–77. [Google Scholar] [CrossRef]
- Brachmann, C.; Davies, A.; Cost, G.J.; Caputo, E.; Li, J.; Hieter, P.; Boeke, J.D. Designer deletion strains derived from Saccharomyces cerevisiae S288C: A useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 1998, 14, 115–132. [Google Scholar] [CrossRef]
- Zahrádka, J.; Sychrová, H. Plasma-membrane hyperpolarization diminishes the cation efflux via Nha1 antiporter and Ena ATPase under potassium-limiting conditions. FEMS Yeast Res. 2012, 12, 439–446. [Google Scholar] [CrossRef] [PubMed]
- Nagai, T.; Ibata, K.; Park, E.S.; Kubota, M.; Mikoshiba, K.; Miyawaki, A. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 2002, 20, 87–90. [Google Scholar] [CrossRef] [PubMed]
- Güldener, U.; Heck, S.; Fiedler, T.; Beinhauer, J.; Hegemann, J.H. A New Efficient Gene Disruption Cassette for Repeated Use in Budding Yeast. Nucleic Acids Res. 1996, 24, 2519–2524. [Google Scholar] [CrossRef]
- Ho, S.N.; Hunt, H.D.; Horton, R.M.; Pullen, J.K.; Pease, L.R. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 1989, 77, 51–59. [Google Scholar] [CrossRef]
- Gietz, R.D.; Schiestl, R.H. Frozen competent yeast cells that can be transformed with high efficiency using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2007, 2, 1–4. [Google Scholar] [CrossRef]
- Shamayeva, K.; Spurna, K.; Kulik, N.; Kale, D.; Munko, O.; Spurny, P.; Zayats, V.; Ludwig, J. MPM motifs of the yeast SKT protein Trk1 can assemble to form a functional K+-translocation system. Biochim. Et Biophys. Acta Biomembr. 2021, 1863, 183513. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Buchroithner, B.; Mayr, S.; Hauser, F.; Priglinger, E.; Stangl, H.; Santa-Maria, A.R.; Deli, M.A.; Der, A.; Klar, T.A.; Axmann, M.; et al. Dual channel microfluidics for mimicking the blood-brain barrier. ACS Nano 2021, 15, 2984–2993. [Google Scholar] [CrossRef]
- Hauser, F.; Jacak, J. Real-time 3D single-molecule localization microscopy analysis using lookup tables. Biomed. Opt. Express 2021, 12, 4955. [Google Scholar] [CrossRef]
- Falk, T.; Mai, D.; Bensch, R.; Çiçek, Ö.; Abdulkadir, A.; Marrakchi, Y.; Böhm, A.; Deubner, J.; Jäckel, Z.; Seiwald, K.; et al. U-Net: Deep learning for cell counting, detection, and morphometry. Nat. Methods 2019, 16, 67–70. [Google Scholar] [CrossRef] [PubMed]
- Krieger, E.; Koraimann, G.; Vriend, G. Increasing the Precision of Comparative Models with YASARA NOVA—A Self-Parameterizing Force Field. PROTEINS: Struct. Funct. Genet. 2002, 47, 393–402. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Rauscher, S.; Nawrocki, G.; Ran, T.; Feig, M.; de Groot, B.L.; Grubmüller, H.; MacKerell, A.D., Jr. CHARMM36m: An improved force field for folded and intrinsically disordered proteins. Nat. Methods 2016, 14, 71–73. [Google Scholar] [CrossRef] [PubMed]
- Krieger, E.; Nielsen, J.E.; Spronk, C.A.; Vriend, G. Fast empirical pKa prediction by Ewald summation. J. Mol. Graph. Model. 2006, 25, 481–486. [Google Scholar] [CrossRef] [PubMed]
- Schneidman-Duhovny, D.; Inbar, Y.; Nussinov, R.; Wolfson, H.J. PatchDock and SymmDock: Servers for rigid and symmetric docking. Nucleic Acids Research 2005, 33 (Suppl. 2), 363–367. [Google Scholar] [CrossRef] [PubMed]
- Madden, T.L.; Tatusov, R.L.; Zhang, J. Applications of network BLAST server. Computer methods for macromolecular sequence analysis. Methods Enzymol. 1996, 266, 131–141. [Google Scholar] [PubMed]
- Cuff, J.A.; Clamp, M.E.; Siddiqui, A.S.; Finlay, M.; Barton, G. JPred: A consensus secondary structure prediction server. Bioinformatics 1998, 14, 892–893. [Google Scholar] [CrossRef]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER suite: Protein structure and function prediction. Nat. Methods 2014, 12, 7–8. [Google Scholar] [CrossRef]
- Canutescu, A.A.; Dunbrack, R.L. Cyclic coordinate descent: A robotics algorithm for protein loop closure. Protein Sci. 2003, 12, 963–972. [Google Scholar] [CrossRef]
- Konagurthu, A.; Whisstock, J.; Stuckey, P.; Lesk, A. MUSTANG: A Multiple Structural Alignment Algorithm. Proteins Struct. Funct. Bioinform. 2006, 14, 659–664. [Google Scholar] [CrossRef]








| VC/Trk1[ΔLHL] | VC/Trk1[ΔLHL] | Trk1[ΔLHL]/VC | Trk1[ΔLHL]/VN | ||
|---|---|---|---|---|---|
| VN/Trk1[ΔLHL] | Trk1[ΔLHL]/VN | VN/Trk1[ΔLHL] | Trk1[ΔLHL]/VC | ||
| Result | NO BiFC | BiFC | BiFC | BiFC | Consistent |
| Possible arrangement | |||||
| Dimer AB/BA | no | no | no | no | no |
| Dimer BC/CB | yes | no | no | no | no |
| Dimer CD/DC | yes | yes | yes | yes | yes |
| Dimer DA/AD | yes | yes | yes | yes | yes |
| Tetramer A | yes | no | yes | no | no |
| Tetramer B | no | yes | yes | no | no |
| Tetramer C | no | yes | yes | yes | no |
| Tetramer D | yes | yes | yes | yes | yes |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kulik, N.; Kale, D.; Spurna, K.; Shamayeva, K.; Hauser, F.; Milic, S.; Janout, H.; Zayats, V.; Jacak, J.; Ludwig, J. Dimerisation of the Yeast K+ Translocation Protein Trk1 Depends on the K+ Concentration. Int. J. Mol. Sci. 2023, 24, 398. https://doi.org/10.3390/ijms24010398
Kulik N, Kale D, Spurna K, Shamayeva K, Hauser F, Milic S, Janout H, Zayats V, Jacak J, Ludwig J. Dimerisation of the Yeast K+ Translocation Protein Trk1 Depends on the K+ Concentration. International Journal of Molecular Sciences. 2023; 24(1):398. https://doi.org/10.3390/ijms24010398
Chicago/Turabian StyleKulik, Natalia, Deepika Kale, Karin Spurna, Katsiaryna Shamayeva, Fabian Hauser, Sandra Milic, Hannah Janout, Vasilina Zayats, Jaroslaw Jacak, and Jost Ludwig. 2023. "Dimerisation of the Yeast K+ Translocation Protein Trk1 Depends on the K+ Concentration" International Journal of Molecular Sciences 24, no. 1: 398. https://doi.org/10.3390/ijms24010398
APA StyleKulik, N., Kale, D., Spurna, K., Shamayeva, K., Hauser, F., Milic, S., Janout, H., Zayats, V., Jacak, J., & Ludwig, J. (2023). Dimerisation of the Yeast K+ Translocation Protein Trk1 Depends on the K+ Concentration. International Journal of Molecular Sciences, 24(1), 398. https://doi.org/10.3390/ijms24010398






