Double-Labeling Method for Visualization and Quantification of Membrane-Associated Proteins in Lactococcus lactis
Abstract
:1. Introduction
2. Results
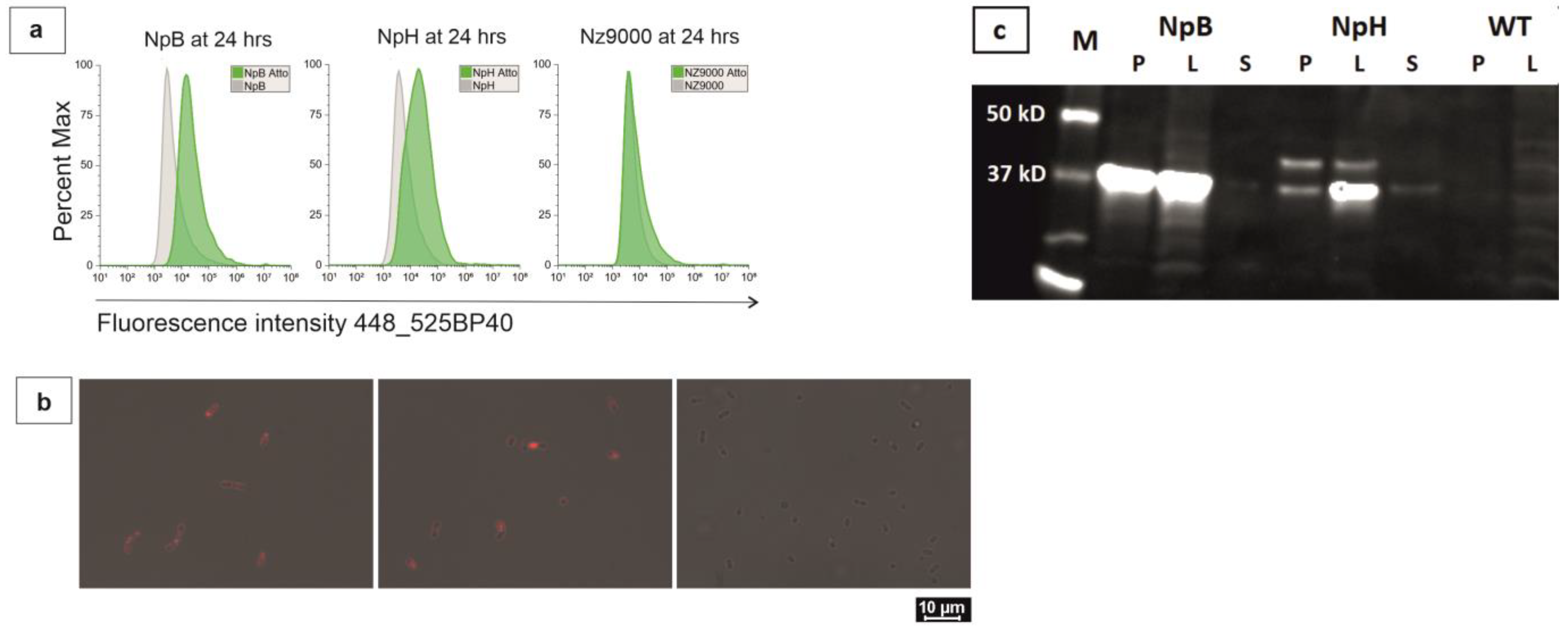
2.1. Overproduction and Overexpression of Recombinant Homologous Proteins
2.2. Trafficking of Membrane-Associated Proteins in Live Bacterial Cells
2.3. Optimization of Biarsenical Technology for Membrane-Associated Proteins
2.4. Quantification of Overexpressed Proteins
3. Discussion
3.1. Real-Time Detection and Quantification of Protein Expression in L. lactis Cytoplasm and on Cell Surface
3.2. Ellman’s Reagent in Biarsenical Labeling Technology
3.3. Determination of Translational Pathways for Membrane Associated Proteins in L. lactis
4. Materials and Methods
4.1. Genetic Designation and Engineering
4.2. Cell Culture and Plasmid Transformation
4.3. Plasmid Selection, Isolation, and Sequencing
4.4. Cell Induction for Protein Overexpression
4.5. Cell Disruption and Protein Measurement
4.6. Protein Analysis and Purification
4.7. Surface Protein Staining
4.8. Blocking of Surface-Anchored Tetracysteine Tags
4.9. Intracellular Protein Staining
4.10. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Robinson, K.; Chamberlain, L.M.; Schofield, K.M.; Wells, J.M.; Le Page, R.W. Oral vaccination of mice against tetanus with recombinant Lactococcus lactis. Nat. Biotechnol. 1997, 15, 653–657. [Google Scholar] [CrossRef]
- Gilbert, C.; Robinson, K.; Le Page, R.W.F.; Wells, J.M. Heterologous Expression of an Immunogenic Pneumococcal Type 3 Capsular Polysaccharide in Lactococcus lactis. Infect. Immun. 2000, 68, 3251–3260. [Google Scholar] [CrossRef] [Green Version]
- Dillard, J.P.; Yother, J. Analysis of Streptococcus pneumoniae sequences cloned into Escherichia coli: Effect of promoter strength and transcription terminators. J. Bacteriol. 1991, 173, 5105–5109. [Google Scholar] [CrossRef] [Green Version]
- Perez, C.A.; Eichwald, C.; Burrone, O.; Mendoza, D. Rotavirus vp7 antigen produced by Lactococcus lactis induces neutralizing antibodies in mice. J. Appl. Microbiol. 2005, 99, 1158–1164. [Google Scholar] [CrossRef] [PubMed]
- Xin, K.-Q.; Hoshino, Y.; Toda, Y.; Igimi, S.; Kojima, Y.; Jounai, N.; Ohba, K.; Kushiro, A.; Kiwaki, M.; Hamajima, K.; et al. Immunogenicity and protective efficacy of orally administered recombinant Lactococcus lactis expressing surface-bound HIV Env. Blood 2003, 102, 223–228. [Google Scholar] [CrossRef] [Green Version]
- Korepanova, A.; Gao, F.P.; Hua, Y.; Qin, H.; Nakamoto, R.K.; Cross, T.A. Cloning and expression of multiple integral membrane proteins from Mycobacterium tuberculosis in Escherichia coli. Protein Sci. 2009, 14, 148–158. [Google Scholar] [CrossRef] [PubMed]
- Schlegel, S.; Hjelm, A.; Baumgarten, T.; Vikström, D.; de Gier, J.-W. Bacterial-based membrane protein production. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2014, 1843, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaur, J.; Bachhawat, A.K. A modified Western blot protocol for enhanced sensitivity in the detection of a membrane protein. Anal. Biochem. 2009, 384, 348–349. [Google Scholar] [CrossRef]
- Chekli, Y.; Peron-Cane, C.; Dell’arciprete, D.; Allemand, J.-F.; Li, C.; Ghigo, J.-M.; Gautier, A.; Lebreton, A.; Desprat, N.; Beloin, C. Visualizing the dynamics of exported bacterial proteins with the chemogenetic fluorescent reporter FAST. Sci. Rep. 2020, 10, 15791. [Google Scholar] [CrossRef]
- Waldo, G.S.; Standish, B.M.; Berendzen, J.; Terwilliger, T.C. Rapid protein-folding assay using green fluorescent protein. Nat. Biotechnol. 1999, 17, 691–696. [Google Scholar] [CrossRef]
- Zhang, A.; Gonzalez, S.M.; Cantor, E.J.; Chong, S. Construction of a mini-intein fusion system to allow both direct monitoring of soluble protein expression and rapid purification of target proteins. Gene 2001, 275, 241–252. [Google Scholar] [CrossRef]
- Berlec, A.; Zadravec, P.; Jevnikar, Z.; Štrukelj, B. Identification of candidate carrier proteins for surface display on Lactococcus lactis by theoretical and experimental analyses of the surface proteome. Appl. Environ. Microbiol. 2011, 77, 1292–1300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clausen, T.; Southan, C.; Ehrmann, M. The HtrA Family of Proteases: Implications for Protein Composition and Cell Fate. Mol. Cell 2002, 10, 443–455. [Google Scholar] [CrossRef]
- Poquet, I.; Saint, V.; Seznec, E.; Simoes, N.; Bolotin, A.; Gruss, A. HtrA is the unique surface housekeeping protease in Lactococcus lactis and is required for natural protein processing. Mol. Microbiol. 2000, 35, 1042–1051. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Bruchez, M.P. Advances in chemical labeling of proteins in living cells. Cell Tissue Res. 2015, 360, 179–194. [Google Scholar] [CrossRef] [Green Version]
- Hogg, K.; Thomas, J.; Ashford, D.; Cartwright, J.; Coldwell, R.; Weston, D.J.; Pillmoor, J.; Surry, D.; O’toole, P. Quantification of proteins by flow cytometry: Quantification of human hepatic transporter P-gp and OATP1B1 using flow cytometry and mass spectrometry. Methods 2015, 82, 38–46. [Google Scholar] [CrossRef] [PubMed]
- BD Biosciences. Introduction to Flow Cytometry: A Learning Guide. December 2002. Available online: https://www.bu.edu/flow-cytometry/files/2010/10/BD-Flow-Cytom-Learning-Guide.pdf (accessed on 20 January 2019).
- Levin, P.A.; Angert, E.R. Small but Mighty: Cell Size and Bacteria. Cold Spring Harb. Perspect. Biol. 2015, 7, a019216. [Google Scholar] [CrossRef] [Green Version]
- Schär-Zammaretti, P.; Ubbink, J. The cell wall of lactic acid bacteria: Surface constituents and macromolecular conformations. Biophys. J. 2003, 85, 4076–4092. [Google Scholar] [CrossRef] [Green Version]
- Zadravec, P.; Mavrič, A.; Matijašić, B.B.; Štrukelj, B.; Berlec, A. Engineering BmpA as a carrier for surface display of IgG-binding domain on Lactococcus lactis. Protein Eng. Des. Select. 2014, 27, 21–27. [Google Scholar] [CrossRef] [Green Version]
- Brajtenbach, D.; Puls, J.-S.; de Opitz, C.L.M.; Sass, P.; Kubitscheck, U.; Grein, F. Quantitative Analysis of Microscopy Data to Evaluate Bacterial Responses to Antibiotic Treatment; Springer: Berlin/Heidelberg, Germany, 2022; pp. 231–257. [Google Scholar] [CrossRef]
- Lepore, A.; Taylor, H.; Landgraf, D.; Okumus, B.; Jaramillo-Riveri, S.; McLaren, L.; Bakshi, S.; Paulsson, J.; El Karoui, M. Quantification of very low-abundant proteins in bacteria using the HaloTag and epi-fluorescence microscopy. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Van Teeffelen, S.; Shaevitz, J.W.; Gitai, Z. Image analysis in fluorescence microscopy: Bacterial dynamics as a case study. Bioessays 2012, 34, 427–436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Sharkawey, A. Calculate the Corrected Total Cell Fluorescence (CTCF). ResearchGate Working paper. May 2016. [Google Scholar] [CrossRef]
- Hoffmann, C.; Gaietta, G.; Bünemann, M.; Adams, S.R.; Oberdorff-Maass, S.; Behr, B.; Vilardaga, J.-P.; Tsien, R.Y.; Ellisman, M.H.; Lohse, M.J. A FlAsH-based FRET approach to determine G protein–coupled receptor activation in living cells. Nature Methods 2005, 2, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Ju, W.; Morishita, W.; Tsui, J.; Gaietta, G.; Deerinck, T.J.; Adams, S.R.; Garner, C.; Tsien, R.Y.; Ellisman, M.H.; Malenka, R.C. Activity-dependent regulation of dendritic synthesis and trafficking of AMPA receptors. Nat. Neurosci. 2004, 7, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Crivat, G.; Tokumasu, F.; Sa, J.M.; Hwang, J.; Wellems, T.E. Tetracysteine-based fluorescent tags to study protein localization and trafficking in Plasmodium falciparum-infected erythrocytes. PLoS ONE 2011, 6, e22975. [Google Scholar] [CrossRef] [Green Version]
- Kottegoda, S.; Aoto, P.C.; Sims, C.E.; Allbritton, N.L. Biarsenical−Tetracysteine Motif as a Fluorescent Tag for Detection in Capillary Electrophoresis. Anal. Chem. 2008, 80, 5358–5366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luedtke, N.W.; Dexter, R.J.; Fried, D.B.; Schepartz, A. Surveying polypeptide and protein domain conformation and association with FlAsH and ReAsH. Nat. Chem. Biol. 2007, 3, 779–784. [Google Scholar] [CrossRef] [PubMed]
- Winther, J.R.; Thorpe, C. Quantification of thiols and disulfides. Biochim. Biophys. Acta (BBA) Gen. Subj. 2014, 1840, 838–846. [Google Scholar] [CrossRef] [Green Version]
- Ellman, G.L. Tissue sulfhydryl groups. Arch. Biochem. 1959, 82, 70–77. [Google Scholar] [CrossRef]
- Riddles, P.W.; Blakeley, R.L.; Zerner, B. [8] Reassessment of Ellman’s reagent. Methods Enzymol. 1983, 91, 49–60. [Google Scholar] [CrossRef]
- Rudner, L.; Nydegger, S.; Coren, L.V.; Nagashima, K.; Thali, M.; Ott, D.E. Dynamic fluorescent imaging of human immunodeficiency virus type 1 gag in live cells by biarsenical labeling. J. Virol. 2005, 79, 4055–4065. [Google Scholar] [CrossRef] [Green Version]
- Hoffmann, C.; Gaietta, G.; Zürn, A.; Adams, S.R.; Terrillon, S.; Ellisman, M.H.; Tsien, R.Y.; Lohse, M.J. Fluorescent labeling of tetracysteine-tagged proteins in intact cells. Nat. Protoc. 2010, 5, 1666–1677. [Google Scholar] [CrossRef] [PubMed]
- Cline, D.J.; Redding, S.E.; Brohawn, S.G.; Psathas, J.N.; Schneider, J.P.; Thorpe, C. New Water-Soluble Phosphines as Reductants of Peptide and Protein Disulfide Bonds: Reactivity and Membrane Permeability. Biochemistry 2004, 43, 15195–15203. [Google Scholar] [CrossRef]
- Chatzi, K.E.; Sardis, M.F.; Economou, A.; Karamanou, S. SecA-mediated targeting and translocation of secretory proteins. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2014, 1843, 1466–1474. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huber, D.; Boyd, D.; Xia, Y.; Olma, M.H.; Gerstein, M.; Beckwith, J. Use of Thioredoxin as a Reporter to Identify a Subset of Escherichia coli Signal Sequences That Promote Signal Recognition Particle-Dependent Translocation. J. Bacteriol. 2005, 187, 2983–2991. [Google Scholar] [CrossRef] [Green Version]
- Zhang, D.; Sweredoski, M.J.; Graham, R.L.J.; Hess, S.; Shan, S. Novel Proteomic Tools Reveal Essential Roles of SRP and Importance of Proper Membrane Protein Biogenesis. Mol. Cell. Proteom. 2012, 11, 011585. [Google Scholar] [CrossRef] [Green Version]
- Lee, C.; Beckwith, J. Cotranslational and Posttranslational Protein Translocation in Prokaryotic Systems. Annu. Rev. Cell Biol. 1986, 2, 315–332. [Google Scholar] [CrossRef]
- Steinberg, R.; Knüpffer, L.; Origi, A.; Asti, R.; Koch, H.-G. Co-translational protein targeting in bacteria. FEMS Microbiol. Lett. 2018, 365, fny095. [Google Scholar] [CrossRef]
- Holo, H.; Nes, I.F. High-frequency transformation by electroporation of Lactococcus lactis subsp. cremoris grown with glycine in osmotically stabilized media. Appl. Environ. Microbiol. 1989, 55, 3119–3123. [Google Scholar] [CrossRef] [PubMed] [Green Version]







Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hoang, M.N.; Peterbauer, C. Double-Labeling Method for Visualization and Quantification of Membrane-Associated Proteins in Lactococcus lactis. Int. J. Mol. Sci. 2023, 24, 10586. https://doi.org/10.3390/ijms241310586
Hoang MN, Peterbauer C. Double-Labeling Method for Visualization and Quantification of Membrane-Associated Proteins in Lactococcus lactis. International Journal of Molecular Sciences. 2023; 24(13):10586. https://doi.org/10.3390/ijms241310586
Chicago/Turabian StyleHoang, Mai Ngoc, and Clemens Peterbauer. 2023. "Double-Labeling Method for Visualization and Quantification of Membrane-Associated Proteins in Lactococcus lactis" International Journal of Molecular Sciences 24, no. 13: 10586. https://doi.org/10.3390/ijms241310586






