Isolation and Optimization of a Broad-Spectrum Synthetic Antimicrobial Peptide, Ap920-WI, from Arthrobacter sp. H5 for the Biological Control of Plant Diseases
Abstract
:1. Introduction
2. Results
2.1. Screening of Antibacterial Genes from Arthrobacter sp. H5 Genome Library and Verification of Their Activity
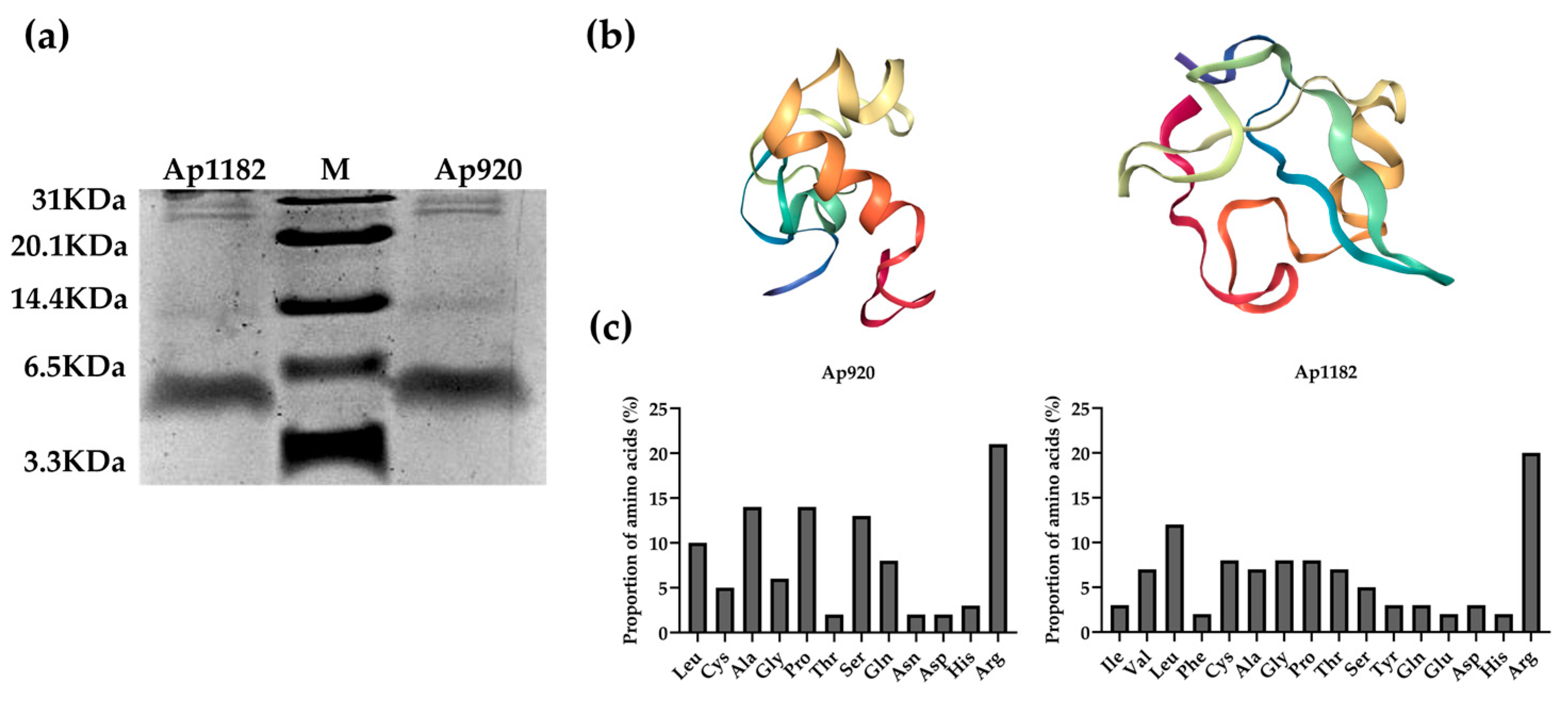
2.2. Purification of Antimicrobial Peptides Ap920 and Ap1182, Bioinformatics Analysis and Determination of Minimum Inhibitory Concentration (MIC) Value
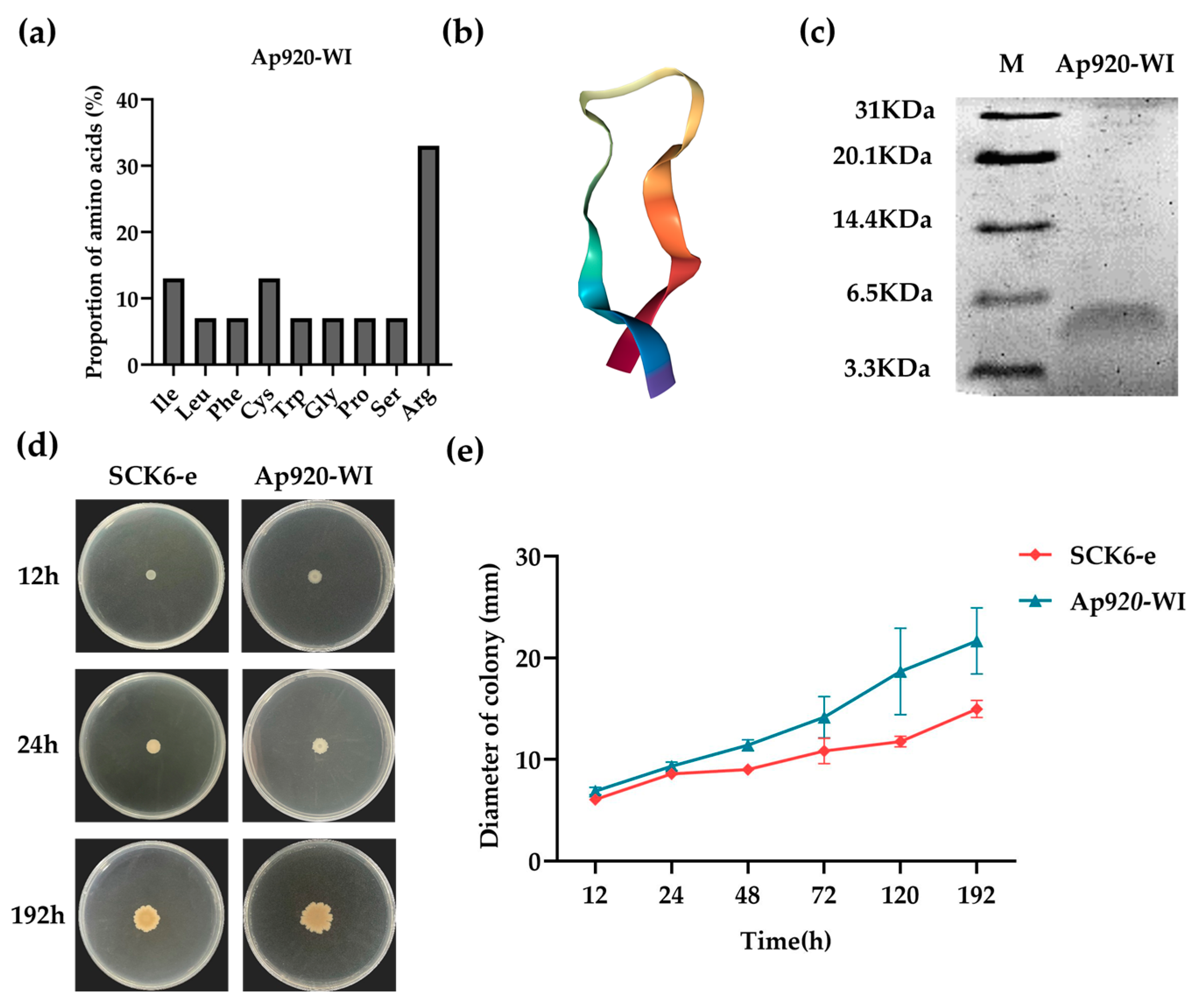
2.3. Optimization and Transformation of Ap920 Using Bioinformatics and Antimicrobial Peptide Properties
2.4. Ap920-WI Has Antibacterial Activities with Lower MIC Values
2.5. Ap920-WI Has Antifungal Activities with Low Concentrations for 50% of Maximal Effect (EC50) Value
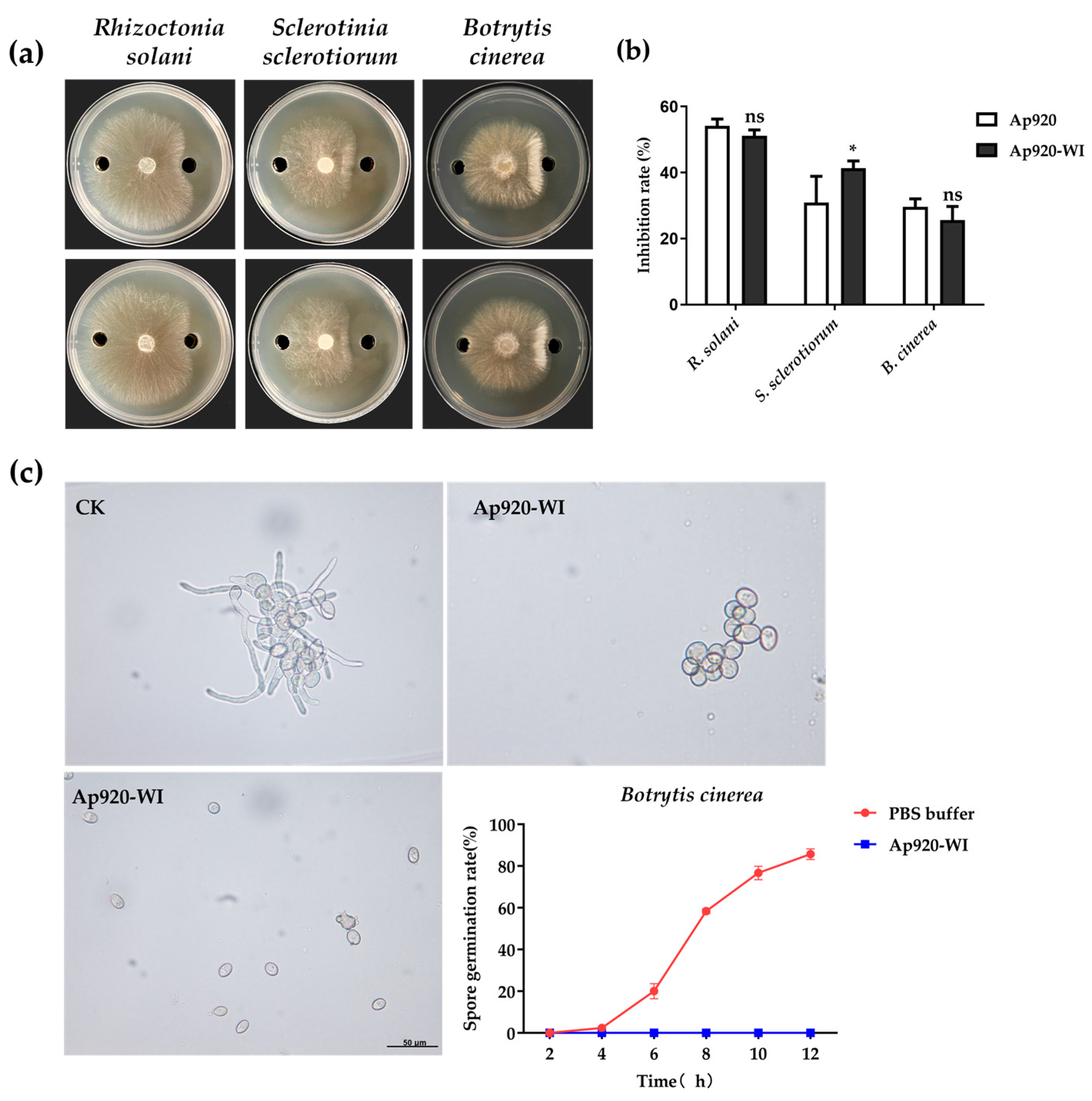
2.6. Inhibitory Effect of Ap920-WI on Mycelia Growth of Pathogenic Fungi
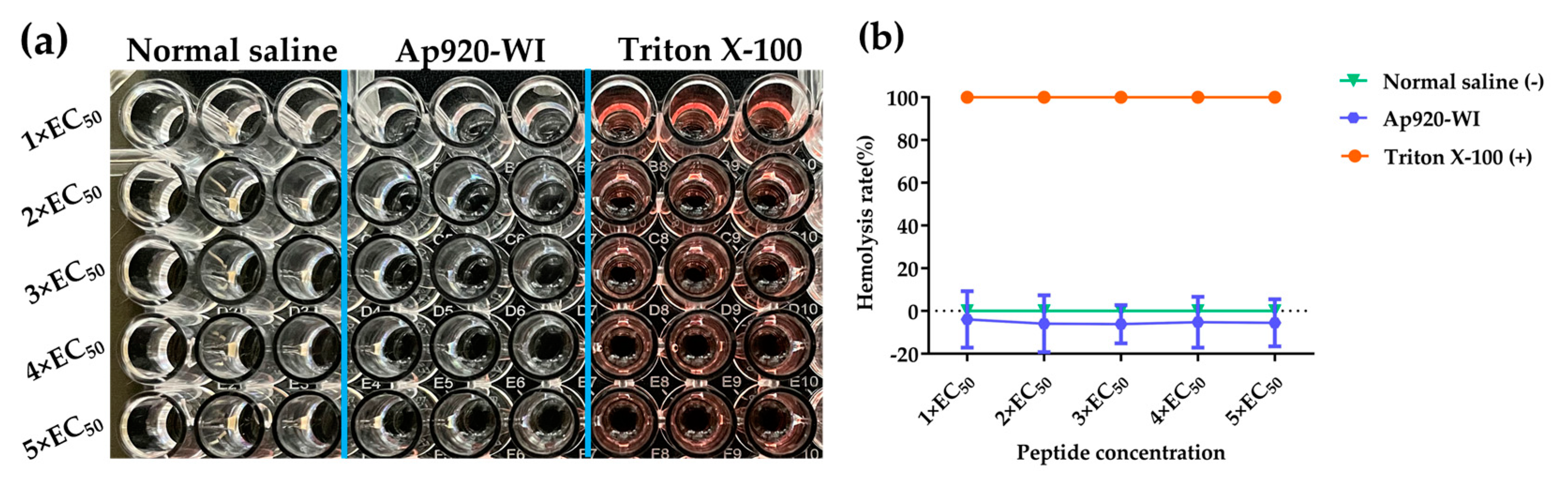
2.7. Ap920-WI Has No Cytotoxicity to Mammalian Cells
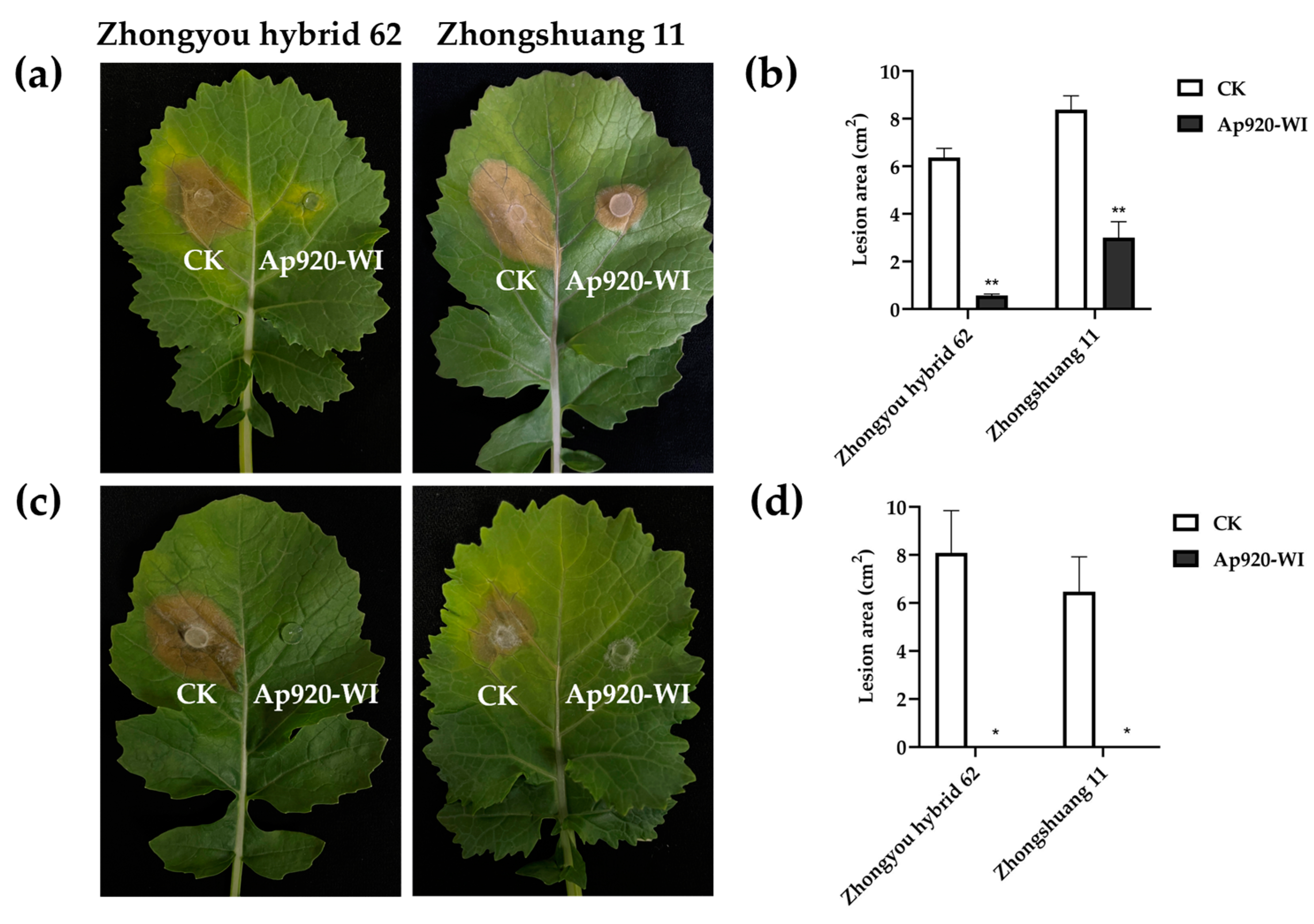
2.8. Ap920-WI Can Inhibit the Infection of S. sclerotiorum on Rape Leaves
3. Discussion
4. Materials and Methods
4.1. Plant Material and Pathogen Culture
4.2. Arthrobacter H5 Genome Library Construction
4.3. Candidate Gene Screening and Confirmation
4.4. Extracellular Protein Extraction
4.5. Antibacterial Activity Analysis
4.6. Construction of the Fusion Gene with His-tag+TEV Restriction Enzyme Site
4.7. Extracellular Protein Purification and Tricine-SDS-PAGE
4.8. Determination of MIC Value, Thermal Stability, and UV Stability
4.9. Bioinformatics Predictive Analysis and Optimization
4.10. Determination of the Inhibitory Effect of Antimicrobial Peptides on Pathogenic Fungi
4.11. Determination of Ap920-WI’s Inhibitory Effect on Botrytis Cinerea and Calculation of Its EC50 Value
4.12. Determination of Hemolytic Activity
4.13. Determination of Ap920-WI’s Effect on the Infection of S. sclerotiorum on Rapeseed
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wu, J.; Abbas, H.M.K.; Li, J.; Yuan, Y.; Liu, Y.; Wang, G.; Dong, A.W. Cell Membrane-Interrupting Antimicrobial Peptides from Isatis indigotica Fortune Isolated by a Bacillus subtilis Expression System. Biomolecules 2019, 10, 30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mookherjee, N.; Hancock, R.E. Cationic host defence peptides: Innate immune regulatory peptides as a novel approach for treating infections. Cell. Mol. Life Sci. 2007, 64, 922–933. [Google Scholar] [CrossRef]
- Campos, M.L.; de Souza, C.M.; de Oliveira, K.B.S.; Dias, S.C.; Franco, O.L. The role of antimicrobial peptides in plant immunity. J. Exp. Bot. 2018, 69, 4997–5011. [Google Scholar] [CrossRef] [Green Version]
- Mukhopadhyay, S.; Bharath Prasad, A.S.; Mehta, C.H.; Nayak, U.Y. Antimicrobial peptide polymers: No escape to ESKAPE pathogens-a review. World J. Microbiol. Biotechnol. 2020, 36, 131. [Google Scholar] [CrossRef]
- Lewies, A.; Du Plessis, L.H.; Wentzel, J.F. Antimicrobial Peptides: The Achilles' Heel of Antibiotic Resistance? Probiotics Antimicrob. Proteins 2019, 11, 370–381. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.Y.; Yan, Z.B.; Meng, Y.M.; Hong, X.Y.; Shao, G.; Ma, J.J.; Cheng, X.R.; Liu, J.; Kang, J.; Fu, C.Y. Antimicrobial peptides: Mechanism of action, activity and clinical potential. Mil. Med. Res. 2021, 8, 48. [Google Scholar] [CrossRef] [PubMed]
- Xuan, H.L.; Duc, T.D.; Thuy, A.M.; Chau, P.M.; Tung, T.T. Chemical approaches in the development of natural nontoxic peptide Polybia-MP1 as a potential dual antimicrobial and antitumor agent. Amino Acids 2021, 53, 843–852. [Google Scholar] [CrossRef]
- Mookherjee, N.; Anderson, M.A.; Haagsman, H.P.; Davidson, D.J. Antimicrobial host defence peptides: Functions and clinical potential. Nat. Rev. Drug Discov. 2020, 19, 311–332. [Google Scholar] [CrossRef]
- Castillo-Juarez, I.; Blancas-Luciano, B.E.; Garcia-Contreras, R.; Fernandez-Presas, A.M. Antimicrobial peptides properties beyond growth inhibition and bacterial killing. PeerJ 2022, 10, e12667. [Google Scholar] [CrossRef]
- Islam, M.S.; Mohamed, G.; Polash, S.A.; Hasan, M.A.; Sultana, R.; Saiara, N.; Dong, W. Antimicrobial Peptides from Plants: A cDNA-Library Based Isolation, Purification, Characterization Approach and Elucidating Their Modes of Action. Int. J. Mol. Sci. 2021, 22, 8712. [Google Scholar] [CrossRef]
- Fu, T.; Islam, M.S.; Ali, M.; Wu, J.; Dong, W. Two antimicrobial genes from Aegilops tauschii Cosson identified by the Bacillus subtilis expression system. Sci. Rep. 2020, 10, 13346. [Google Scholar] [CrossRef] [PubMed]
- Badosa, E.; Planas, M.; Feliu, L.; Montesinos, L.; Bonaterra, A.; Montesinos, E. Synthetic Peptides against Plant Pathogenic Bacteria. Microorganisms 2022, 10, 1784. [Google Scholar] [CrossRef] [PubMed]
- Tong, S.; Li, M.; Keyhani, N.O.; Liu, Y.; Yuan, M.; Lin, D.; Jin, D.; Li, X.; Pei, Y.; Fan, Y. Characterization of a fungal competition factor: Production of a conidial cell-wall associated antifungal peptide. PLoS Pathog. 2020, 16, e1008518. [Google Scholar] [CrossRef] [Green Version]
- Rajasekaran, K.; Cary, J.W.; Jaynes, J.M.; Cleveland, T.E. Disease resistance conferred by the expression of a gene encoding a synthetic peptide in transgenic cotton (Gossypium hirsutum L.) plants. Plant Biotechnol. J. 2005, 3, 545–554. [Google Scholar] [CrossRef] [PubMed]
- Qutb, A.M.; Wei, F.; Dong, W. Corrigendum: Prediction and Characterization of Cationic Arginine-Rich Plant Antimicrobial Peptide SM-985 from Teosinte (Zea mays ssp. mexicana). Front. Microbiol. 2021, 12, 667085. [Google Scholar] [CrossRef]
- Hassan, M.F.; Qutb, A.M.; Dong, W. Prediction and Activity of a Cationic alpha-Helix Antimicrobial Peptide ZM-804 from Maize. Int. J. Mol. Sci. 2021, 22, 2643. [Google Scholar] [CrossRef]
- Kong, X.; Yang, M.; Abbas, H.M.K.; Wu, J.; Li, M.; Dong, W. Antimicrobial genes from Allium sativum and Pinellia ternata revealed by a Bacillus subtilis expression system. Sci. Rep. 2018, 8, 14514. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Islam, M.S.; Wang, J.; Zhao, Y.; Dong, W. Isolation of Potato Endophytes and Screening of Chaetomium globosum Antimicrobial Genes. Int. J. Mol. Sci. 2022, 23, 4611. [Google Scholar] [CrossRef]
- Park, S.A.; Bhatia, S.K.; Park, H.A.; Kim, S.Y.; Sudheer, P.; Yang, Y.H.; Choi, K.Y. Bacillus subtilis as a robust host for biochemical production utilizing biomass. Crit. Rev. Biotechnol. 2021, 41, 827–848. [Google Scholar] [CrossRef]
- Kaspar, F.; Neubauer, P.; Gimpel, M. Bioactive Secondary Metabolites from Bacillus subtilis: A Comprehensive Review. J. Nat. Prod. 2019, 82, 2038–2053. [Google Scholar] [CrossRef]
- Souza, C.C.; Guimaraes, J.M.; Pereira, S.D.S.; Mariuba, L.A.M. The multifunctionality of expression systems in Bacillus subtilis: Emerging devices for the production of recombinant proteins. Exp. Biol. Med. 2021, 246, 2443–2453. [Google Scholar] [CrossRef]
- Ye, J.; Li, Y.; Bai, Y.; Zhang, T.; Jiang, W.; Shi, T.; Wu, Z.; Zhang, Y.-H.P.J. A facile and robust T7-promoter-based high-expression of heterologous proteins in Bacillus subtilis. Bioresour. Bioprocess. 2022, 9, 56. [Google Scholar] [CrossRef]
- Chu, P.T.B.; Phan, T.T.P.; Nguyen, T.T.T.; Truong, T.T.T.; Schumann, W.; Nguyen, H.D. Potent IPTG-inducible integrative expression vectors for production of recombinant proteins in Bacillus subtilis. World J. Microbiol. Biotechnol. 2023, 39, 143. [Google Scholar] [CrossRef]
- Cui, W.; Han, L.; Suo, F.; Liu, Z.; Zhou, L.; Zhou, Z. Exploitation of Bacillus subtilis as a robust workhorse for production of heterologous proteins and beyond. World J. Microbiol. Biotechnol. 2018, 34, 145. [Google Scholar] [CrossRef] [PubMed]
- Fu, G.; Yue, J.; Li, D.; Li, Y.; Lee, S.Y.; Zhang, D. An operator-based expression toolkit for Bacillus subtilis enables fine-tuning of gene expression and biosynthetic pathway regulation. Proc. Natl. Acad Sci. USA 2022, 119, e2119980119. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Fan, L.; Sun, J.; Lao, X.; Zheng, H. Computational resources and tools for antimicrobial peptides. J. Pept. Sci. 2017, 23, 4–12. [Google Scholar] [CrossRef] [PubMed]
- Gawde, U.; Chakraborty, S.; Waghu, F.H.; Barai, R.S.; Khanderkar, A.; Indraguru, R.; Shirsat, T.; Idicula-Thomas, S. CAMPR4: A database of natural and synthetic antimicrobial peptides. Nucleic. Acids Res. 2023, 51, D377–D383. [Google Scholar] [CrossRef]
- Usmani, S.S.; Kumar, R.; Bhalla, S.; Kumar, V.; Raghava, G.P.S. In Silico Tools and Databases for Designing Peptide-Based Vaccine and Drugs. Adv. Protein Chem. Struct. Biol. 2018, 112, 221–263. [Google Scholar] [CrossRef]
- Chen, X.; Wu, X.; Wang, S. An optimized antimicrobial peptide analog acts as an antibiotic adjuvant to reverse methicillin-resistant Staphylococcus aureus. NPJ Sci. Food 2022, 6, 57. [Google Scholar] [CrossRef]
- Torres, M.D.T.; Melo, M.C.R.; Flowers, L.; Crescenzi, O.; Notomista, E.; de la Fuente-Nunez, C. Mining for encrypted peptide antibiotics in the human proteome. Nat. Biomed. Eng. 2022, 6, 67–75. [Google Scholar] [CrossRef]
- Zhang, M.; Ouyang, J.; Fu, L.; Xu, C.; Ge, Y.; Sun, S.; Li, X.; Lai, S.; Ke, H.; Yuan, B.; et al. Hydrophobicity Determines the Bacterial Killing Rate of alpha-Helical Antimicrobial Peptides and Influences the Bacterial Resistance Development. J. Med. Chem. 2022, 65, 14701–14720. [Google Scholar] [CrossRef] [PubMed]
- Xuan, J.; Feng, W.; Wang, J.; Wang, R.; Zhang, B.; Bo, L.; Chen, Z.S.; Yang, H.; Sun, L. Antimicrobial peptides for combating drug-resistant bacterial infections. Drug Resist. Updates 2023, 68, 100954. [Google Scholar] [CrossRef] [PubMed]
- Duarte-Mata, D.I.; Salinas-Carmona, M.C. Antimicrobial peptides immune modulation role in intracellular bacterial infection. Front. Immunol. 2023, 14, 1119574. [Google Scholar] [CrossRef] [PubMed]
- Moretta, A.; Salvia, R.; Scieuzo, C.; Di Somma, A.; Vogel, H.; Pucci, P.; Sgambato, A.; Wolff, M.; Falabella, P. A bioinformatic study of antimicrobial peptides identified in the Black Soldier Fly (BSF) Hermetia illucens (Diptera: Stratiomyidae). Sci. Rep. 2020, 10, 16875. [Google Scholar] [CrossRef]
- Liscano, Y.; Onate-Garzon, J.; Delgado, J.P. Peptides with Dual Antimicrobial-Anticancer Activity: Strategies to Overcome Peptide Limitations and Rational Design of Anticancer Peptides. Molecules 2020, 25, 4245. [Google Scholar] [CrossRef]
- Wang, T.; Zou, C.; Wen, N.; Liu, X.; Meng, Z.; Feng, S.; Zheng, Z.; Meng, Q.; Wang, C. The effect of structural modification of antimicrobial peptides on their antimicrobial activity, hemolytic activity, and plasma stability. J. Pept. Sci. 2021, 27, e3306. [Google Scholar] [CrossRef]
- Bhattacharjya, S.; Straus, S.K. Design, Engineering and Discovery of Novel alpha-Helical and beta-Boomerang Antimicrobial Peptides against Drug Resistant Bacteria. Int. J. Mol. Sci. 2020, 21, 5773. [Google Scholar] [CrossRef]
- Bobde, S.S.; Alsaab, F.M.; Wang, G.; Van Hoek, M.L. Ab initio Designed Antimicrobial Peptides Against Gram-Negative Bacteria. Front. Microbiol. 2021, 12, 715246. [Google Scholar] [CrossRef]
- Waghu, F.H.; Idicula-Thomas, S. Collection of antimicrobial peptides database and its derivatives: Applications and beyond. Protein Sci. 2019, 29, 36–42. [Google Scholar] [CrossRef] [Green Version]
- Freire, J.M.; Almeida Dias, S.; Flores, L.; Veiga, A.S.; Castanho, M.A. Mining viral proteins for antimicrobial and cell-penetrating drug delivery peptides. Bioinformatics 2015, 31, 2252–2256. [Google Scholar] [CrossRef] [Green Version]
- Yaseen, A.G.S.; Akhtar, N.; Amin, I.; Minhas, F. HemoNet: Predicting hemolytic activity of peptides with integrated feature learning. J. Bioinform. Comput. Biol. 2021, 19, 5. [Google Scholar] [CrossRef] [PubMed]
- Pineda-Castaneda, H.M.; Huertas-Ortiz, K.A.; Leal-Castro, A.L.; Vargas-Casanova, Y.; Parra-Giraldo, C.M.; Garcia-Castaneda, J.E.; Rivera-Monroy, Z.J. Designing Chimeric Peptides: A Powerful Tool for Enhancing Antibacterial Activity. Chem. Biodivers. 2021, 18, e2000885. [Google Scholar] [CrossRef]
- Deng, T.; Ge, H.; He, H.; Liu, Y.; Zhai, C.; Feng, L.; Yi, L. The heterologous expression strategies of antimicrobial peptides in microbial systems. Protein Expr. Purif. 2017, 140, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Koch, P.; Schmitt, S.; Heynisch, A.; Gumpinger, A.; Wuthrich, I.; Gysin, M.; Shcherbakov, D.; Hobbie, S.N.; Panke, S.; Held, M. Optimization of the antimicrobial peptide Bac7 by deep mutational scanning. BMC Biol. 2022, 20, 114. [Google Scholar] [CrossRef]
- Ma, Y.; Guo, Z.; Xia, B.; Zhang, Y.; Liu, X.; Yu, Y.; Tang, N.; Tong, X.; Wang, M.; Ye, X.; et al. Identification of antimicrobial peptides from the human gut microbiome using deep learning. Nat. Biotechnol. 2022, 40, 921–931. [Google Scholar] [CrossRef]
- Clarke, J.D. Cetyltrimethyl ammonium bromide (CTAB) DNA miniprep for plant DNA isolation. Cold Spring Harb. Protoc. 2009, 2009, pdb-prot5177. [Google Scholar] [CrossRef]
- Farag, R.S.; Daw, Z.Y.; Hewedi, F.M.; El-Baroty, G.S.A. Antimicrobial Activity of Some Egyptian Spice Essential Oils. J. Food Prot. 1989, 52, 665–667. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Cui, X.; Zhang, M.; Bai, B.; Yang, Y.; Fan, S. The antibacterial mechanism of perilla rosmarinic acid. Biotechnol. Appl. Biochem. 2022, 69, 1757–1764. [Google Scholar] [CrossRef]


| Strains of Bacterium | Minimum Inhibitory Concentration (MIC: μg/mL) | ||
|---|---|---|---|
| Ap920 | Ap1182 | Ap920-WI | |
| Clavibacter fangii | 52 | 57 | 83 |
| Clavibacter michiganesis | 84 | 88 | 78 |
| Xanthomonas oryzae pv. oryzicola | 137 | 132 | 66 |
| Rastonia solanacearum | 67 | 64 | 37 |
| Xanthomonas oryzae pv. oryzae | 128 | 115 | 100 |
| Strains of Fungus | Regression Equation | EC50 (μg/mL) | Correlation Coefficient (R-Value) |
|---|---|---|---|
| Rhizoctonia solani | Y = 0.3996 + 3.1572X | 29 | 0.9958 |
| Sclerotinia sclerotiorum | Y = 1.3984 + 2.0141X | 44 | 0.9805 |
| Botrytis cinerea | Y = 1.9214 + 1.8783X | 61 | 0.9834 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, L.; Islam, M.S.; Song, P.; Zhu, L.; Dong, W. Isolation and Optimization of a Broad-Spectrum Synthetic Antimicrobial Peptide, Ap920-WI, from Arthrobacter sp. H5 for the Biological Control of Plant Diseases. Int. J. Mol. Sci. 2023, 24, 10598. https://doi.org/10.3390/ijms241310598
Zhao L, Islam MS, Song P, Zhu L, Dong W. Isolation and Optimization of a Broad-Spectrum Synthetic Antimicrobial Peptide, Ap920-WI, from Arthrobacter sp. H5 for the Biological Control of Plant Diseases. International Journal of Molecular Sciences. 2023; 24(13):10598. https://doi.org/10.3390/ijms241310598
Chicago/Turabian StyleZhao, Li, Md. Samiul Islam, Pei Song, Li Zhu, and Wubei Dong. 2023. "Isolation and Optimization of a Broad-Spectrum Synthetic Antimicrobial Peptide, Ap920-WI, from Arthrobacter sp. H5 for the Biological Control of Plant Diseases" International Journal of Molecular Sciences 24, no. 13: 10598. https://doi.org/10.3390/ijms241310598






