Establishment of a Rapid Detection Method for Yeast-like Symbionts in Brown Planthopper Based on Droplet Digital PCR Technology
Abstract
:1. Introduction
2. Results
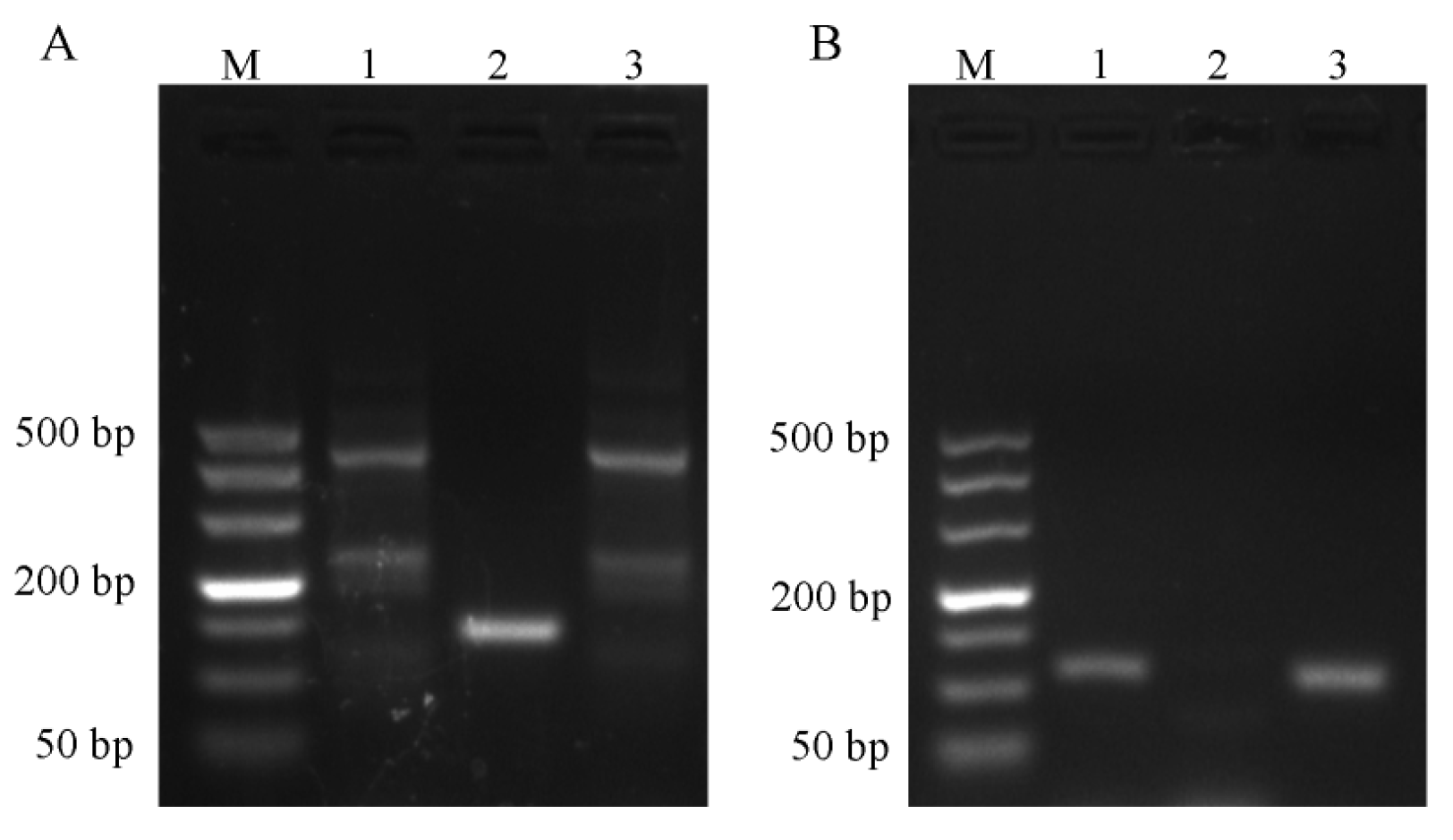
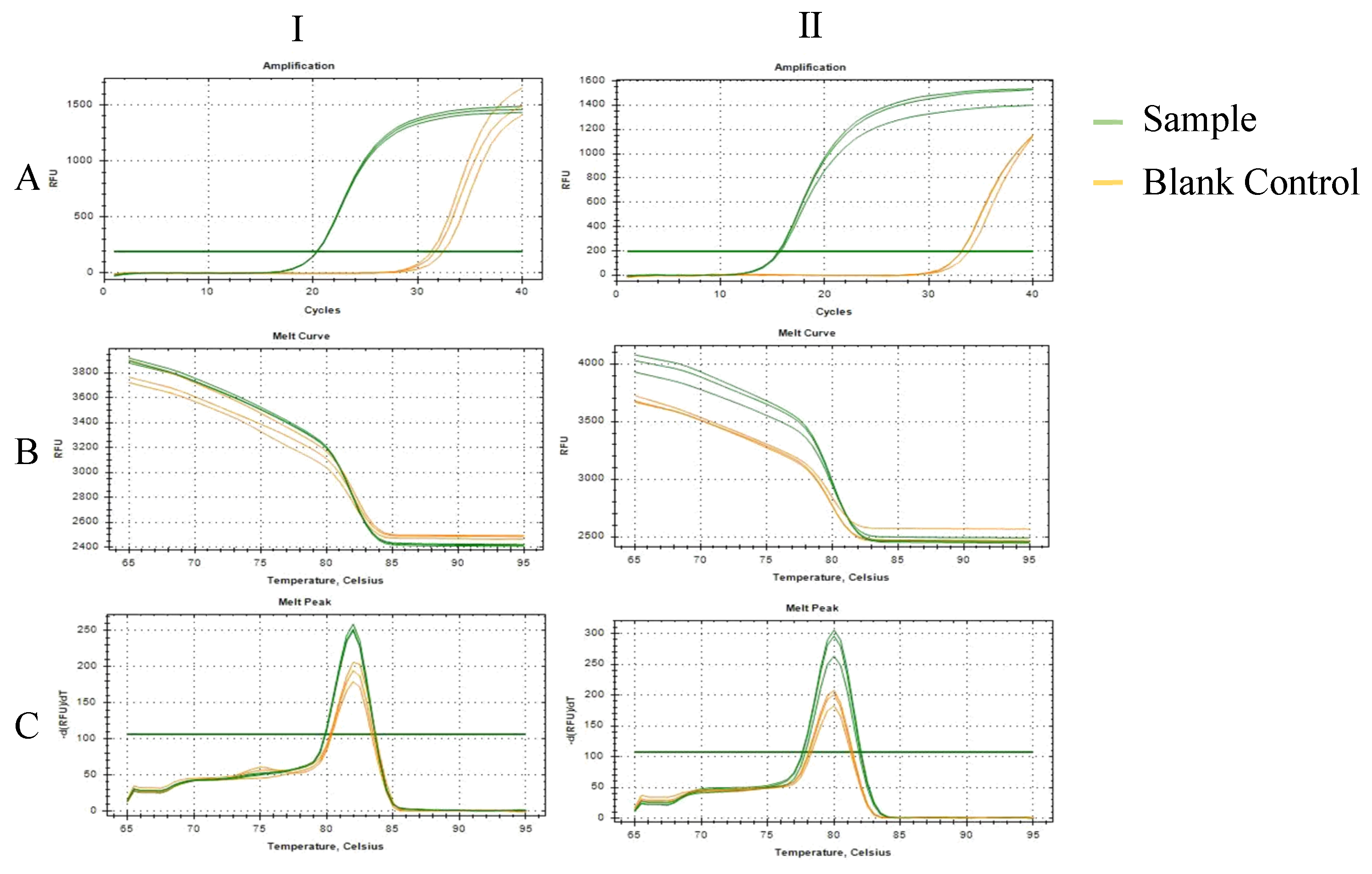
2.1. Results of Primer Screening for PCR and RT-qPCR Amplification
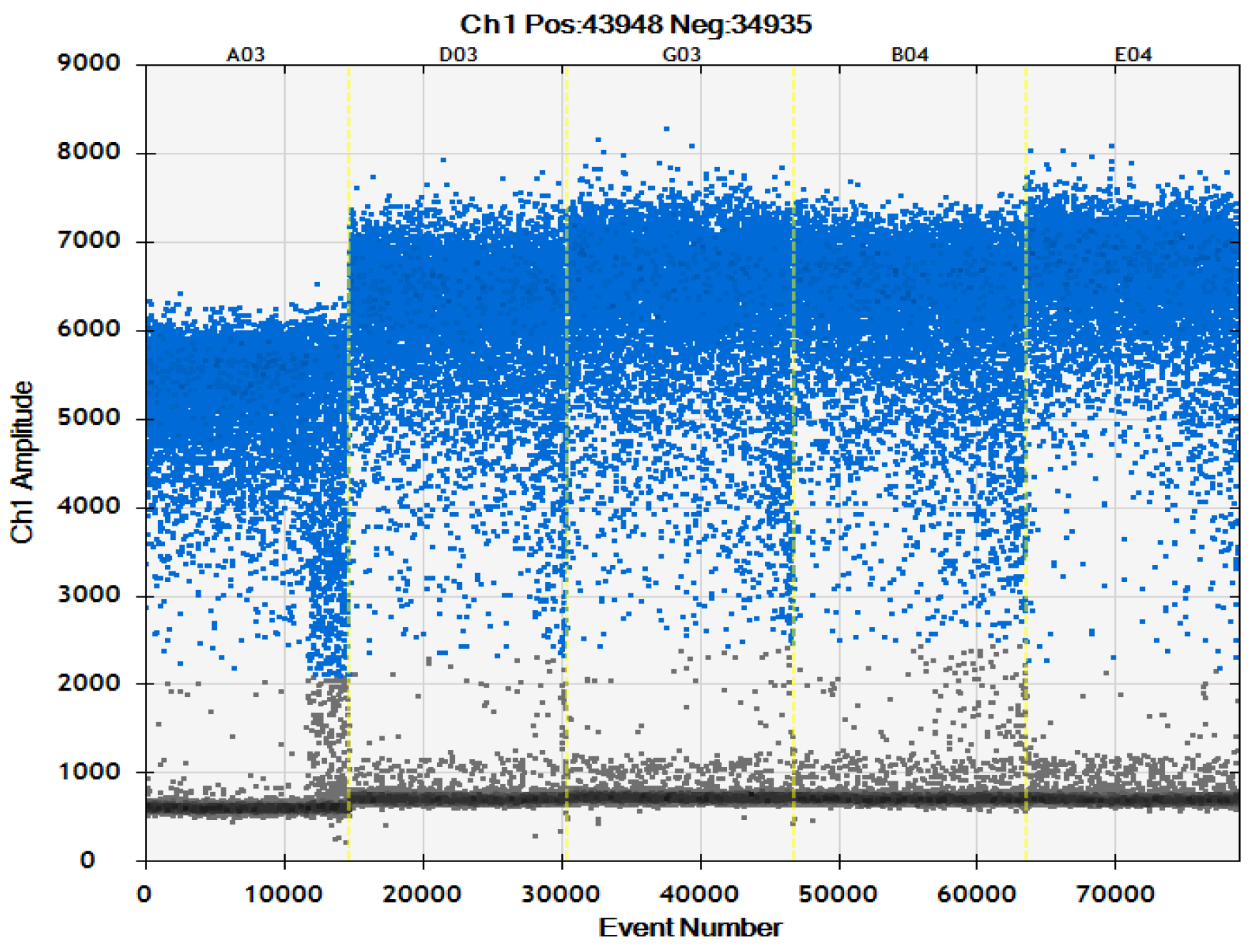
2.2. Optimization of ddPCR System
2.2.1. Optimization of Primer Concentration
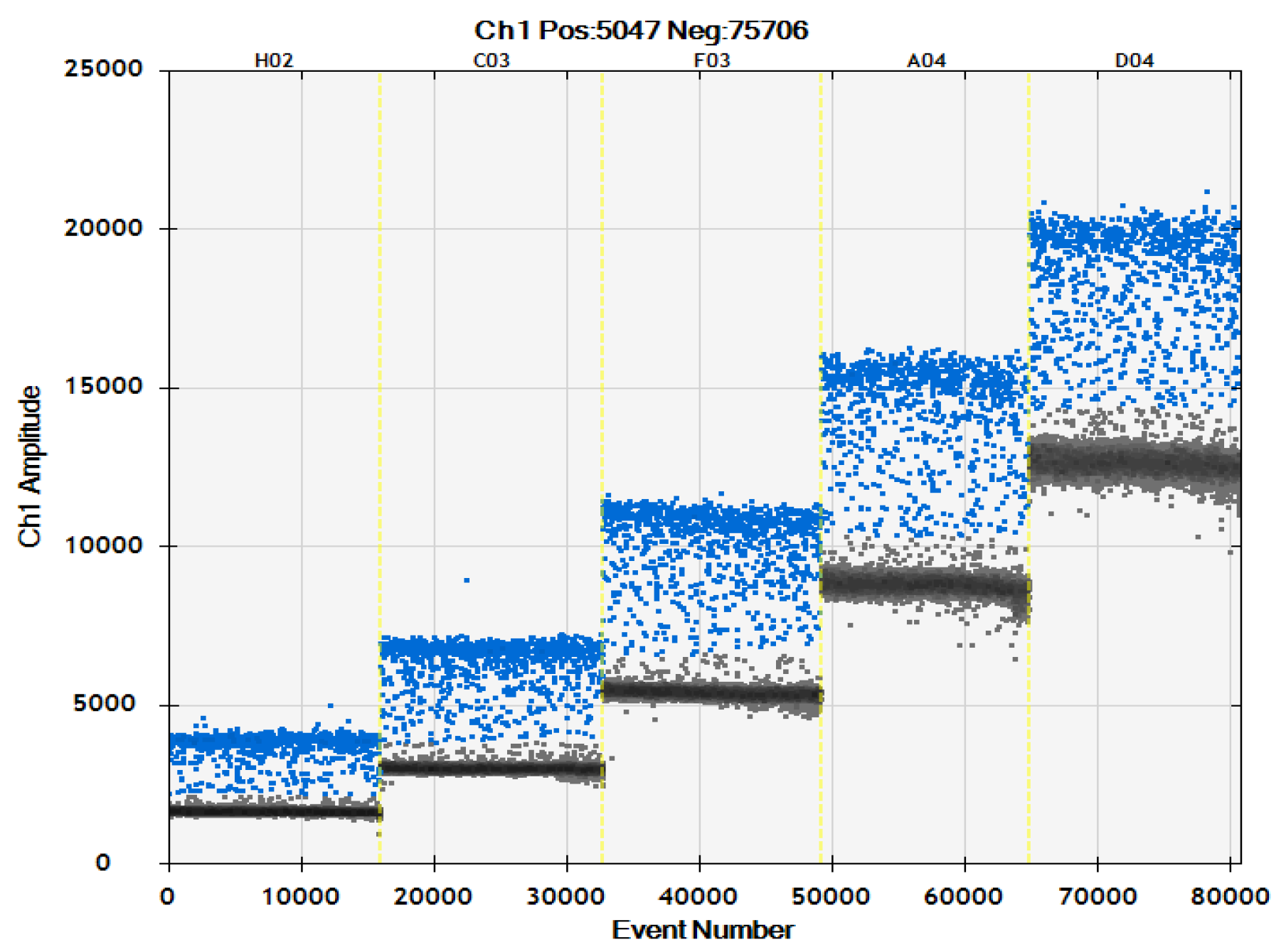
2.2.2. Optimization of Probe Concentration
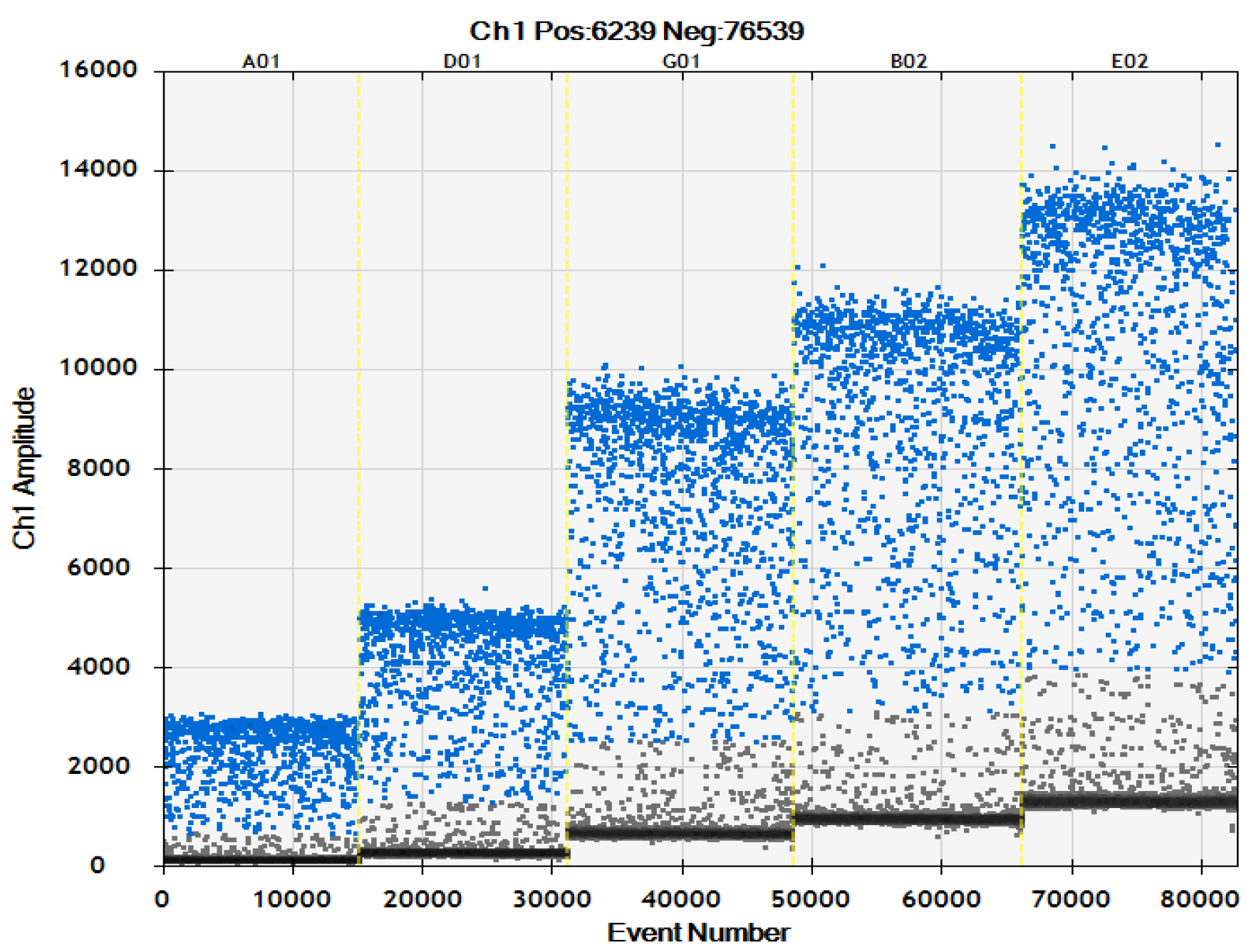
2.3. ddPCR Sensitivity Test
2.4. ddPCR Specificity Test
2.5. ddPCR Repeatability Test
3. Discussion
4. Materials and Methods
4.1. Collection and Culture of Insects
4.2. Specific Experiment
4.2.1. Principal Reagent
4.2.2. Main Instruments and Materials
4.3. Genome Extraction of Symbiotic Fungi
4.4. Design and Screening of Primer and Probe
4.4.1. PCR Amplification and Sequencing Alignment
4.4.2. SYBR Green Dye Real-Time Quantitative PCR
4.4.3. Optimization of Annealing Temperature Using Taq Man Probe Real-Time Quantitative PCR
4.5. Plasmid Sample Preparation
4.6. ddPCR System Optimization
4.7. Sensitivity Test
4.8. Specific Test
4.9. Data Statistics and Analysis
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Frago, E.; Dicke, M.; Godfray, H.C.J. Insect symbionts as hidden players in insect–plant interactions. Trends Ecol. Evol. 2012, 27, 705–711. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.P.; Chu, D.; Liu, B.M.; Xie, W.; Wang, S.L.; Wu, Q.J.; Xu, B.Y.; Zhang, Y.J. Relative Amount of Symbionts in Insect Hosts Changes with Host-Plant Adaptation and Insecticide Resistance. Environ. Entomol. 2013, 42, 74–78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vásquez, A.; Forsgren, E.; Fries, I.; Paxton, R.J.; Flaberg, E.; Szekely, L.; Olofsson, T.C. Symbionts as major modulators of insect health: Lactic acid bacteria and honeybees. PLoS ONE 2017, 7, e33188. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xuan, Z.; Xing, W.; Zikun, G.; Xueying, L.; Ping, W.; Xiangqun, Y.; Yiping, L. Antibiotic Treatment Reduced the Gut Microbiota Diversity, Prolonged the Larval Development Period and Lessened Adult Fecundity of Grapholita molesta (Lepidoptera: Tortricidae). Insects 2022, 13, 838. [Google Scholar]
- Lü, D.; Dong, Y.; Yan, Z.; Liu, X.; Zhang, Y.; Yang, D.; He, K.; Wang, Z.; Wang, P.; Yuan, X.; et al. Dynamics of gut microflora across the life cycle of Spodoptera frugiperda and its effects on the feeding and growth of larvae. Pest Manag. Sci. 2022, 79, 173–182. [Google Scholar] [CrossRef]
- Strigini, M.; Leulier, F. The role of the microbial environment in Drosophila post-embryonic development. Dev. Comp. Immunol. 2016, 64, 39–52. [Google Scholar] [CrossRef]
- Li, Y.; Su, W.; Zhu, X.; Hui, X.; Fan, X.; Yao, H.; Chang, X.; Liu, W. Isolation of Acetobacter orientalis and their promotion of the growth and development of Drosophila melanogaster. Acta Microbiol. Sin. 2017, 57, 1536–1545. [Google Scholar]
- Michael, B.-Y.; Yael, A.; Edouard, J.; Boaz, Y. Give us the tools and we will do the job: Symbiotic bacteria affect olive fly fitness in a diet-dependent fashion. Proc. Biol. Sci. 2010, 277, 1545–1552. [Google Scholar]
- Qiwen, Z.; Pumo, C.; Bo, W.; Xuxiang, L.; Jia, L.; Ruohan, H.; Hehe, Z.; Chuandong, Y.; Xuesen, S.; Qinge, J.; et al. Manipulation of Gut Symbionts for Improving the Sterile Insect Technique: Quality Parameters of Bactrocera dorsalis (Diptera: Tephritidae) Genetic Sexing Strain Males After Feeding on Bacteria-Enriched Diets. J. Econ. Entomol. 2021, 114, 560–570. [Google Scholar]
- Akami, M.; Ren, X.-M.; Qi, X.; Mansour, A.; Gao, B.; Cao, S.; Niu, C.-Y. Symbiotic bacteria motivate the foraging decision and promote fecundity and survival of Bactrocera dorsalis (Diptera: Tephritidae). BMC Microbiol. 2019, 19, 229. [Google Scholar] [CrossRef] [Green Version]
- Bilodeau, E.; Guay, J.-F.; Turgeon, J.; Cloutier, C. Survival to parasitoids in an insect hosting defensive symbionts: A multivariate approach to polymorphic traits affecting host use by its natural enemy. PLoS ONE 2018, 8, e60708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, S.; Dos-Santos, A.L.A.; Huang, W.; Liu, K.C.; Oshaghi, M.A.; Wei, G.; Agre, P.; Jacobs-Lorena, M. Driving mosquito refractoriness to Plasmodium falciparum with engineered symbiotic bacteria. Science 2017, 357, 1399–1402. [Google Scholar] [CrossRef] [PubMed]
- Xue, J.; Zhang, X.-Q.; Xu, H.-J.; Fan, H.-W.; Huang, H.-J.; Ma, X.-F.; Wang, C.-Y.; Chen, J.-G.; Cheng, J.-A.; Zhang, C.-X. Molecular characterization of the flightin gene in the wing-dimorphic planthopper, Nilaparvata lugens, and its evolution in Pancrustacea. Insect Biochem. Mol. Biol. 2013, 43, 433–443. [Google Scholar] [CrossRef] [PubMed]
- Hongoh, Y.; Sasaki, T.; Ishikawa, H. Cloning, sequence analysis and expression in Escherichia coli of the gene encoding a uricase from the yeast-like symbiont of the brown planthopper, Nilaparvata lugens. Insect Biochem. Mol. Biol. 2000, 30, 173–182. [Google Scholar] [CrossRef]
- Cheng, D.J.; Hou, R.F. Histological observations on transovarial transmission of a yeast-like symbiote in Nilaparvata lugens Stål (Homoptera, Delphacidae). Tissue Cell 2001, 33, 273–279. [Google Scholar] [CrossRef]
- Dong, S.; Pang, K.; Bai, X.; Yu, X.; Hao, P. Identification of two species of yeast-like symbiotes in the brown planthopper, Nilaparvata lugens. Curr. Microbiol. 2011, 62, 1133–1138. [Google Scholar] [CrossRef]
- Xue, J.; Zhou, X.; Zhang, C.-X.; Yu, L.-L.; Fan, H.-W.; Wang, Z.; Xu, H.-J.; Xi, Y.; Zhu, Z.-R.; Zhou, W.-W.; et al. Genomes of the rice pest brown planthopper and its endosymbionts reveal complex complementary contributions for host adaptation. Genome Biol. 2014, 15, 521. [Google Scholar] [CrossRef] [Green Version]
- Sasaki, T.; Kawamura, M.; Ishikawa, H. Nitrogen recycling in the brown planthopper, Nilaparvata lugens: Involvement of yeast-like endosymbionts in uric acid metabolism. J. Insect Physiol. 1996, 42, 125–129. [Google Scholar] [CrossRef]
- Koyama, K. Nutritional Physiology of the Brown Rice Planthopper, Nilaparvata lugens STAL (Hemiptera: Delphacidae): III. Essential Vitamins for Nymphal Development. Appl. Entomol. Zool. 1986, 21, 252–257. [Google Scholar] [CrossRef] [Green Version]
- Hiroaki, N.; Kojiro, W.; Tetsuo, S. Sterols in Laodelphax striatellus with special reference to the intracellular yeastlike symbiotes as a sterol source. J. Insect Physiol. 1979, 25, 443–447. [Google Scholar]
- Xu, H.; Zhen, X.; Tong, Z.; Lu, Z.; Chen, J.; Yu, X.; Tao, L. Effects of insecticides on the symbiotes in brown planthopper. Acta Agric. Zhejiangensis 2000, 12, 126–128. [Google Scholar]
- Liu, M.; Hong, G.; Li, H.; Bing, X.; Chen, Y.; Jing, X.; Gershenzon, J.; Lou, Y.; Baldwin, I.T.; Li, R. Sakuranetin protects rice from brown planthopper attack by depleting its beneficial endosymbionts. Proc. Natl. Acad. Sci. USA 2023, 120, e2305007120. [Google Scholar] [CrossRef] [PubMed]
- Nasu, S. Studies on some leafhoppers and planthoppers which transmit virus disease of rice plant in Japan. Bull. Kyushu Agric. Exp. Sta. 1963, 8, 1533349. [Google Scholar]
- Chen, F.; Zhang, J.; Chen, J. Change in size and abundance of the yeast-like endosymbiote during the interaction between brown planthopper, Nilaparvata lugens (Stål) and resistant-variety rice. Acta Agric. Zhejiangensis 2006, 18, 294–298. [Google Scholar]
- Noda, H.; Nakashima, N.; Koizumi, M. Phylogenetic position of yeast-like symbiotes of rice planthoppers based on partial 18S rDNA Sequences. Insect Biochem. Mol. Biol. 1995, 25, 639–646. [Google Scholar] [CrossRef] [PubMed]
- Pang, K.; Dong, S.Z.; Hou, Y.; Bian, Y.L.; Yang, K.; Yu, X.P. Cultivation, identification and quantification of one species of yeast-like symbiotes, Candida, in the rice brown planthopper, Nilaparvata lugens. Insect Sci. 2012, 19, 477–484. [Google Scholar] [CrossRef]
- Zhang, J.-F.; Chen, J.-M.; Chen, F.-J.; Zheng, X.-S.; Chen, L.-Z.; Yu, X.-P. The Isolation of Yeast-Like-Symbiots in the Brown Planthopper and the Sequences Analysis of Its 26S rDNA. Sci. Agric. Sin. 2009, 42, 2211–2216. [Google Scholar]
- Wang, T.Z. Gut Microbial Diversity and Difference Analysis in Nilaparvata lugens from Different Developement Stages and Virulence Populations. Master’s Thesis, China Jiliang University, Hangzhou, China, 2019. [Google Scholar]
- Hindson, C.M.; Chevillet, J.R.; Briggs, H.A.; Gallichotte, E.N.; Ruf, I.K.; Hindson, B.J.; Vessella, R.L.; Tewari, M. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat. Methods 2013, 10, 1003–1005. [Google Scholar] [CrossRef]
- Whale, A.S.; Huggett, J.F.; Cowen, S.; Speirs, V.; Shaw, J.; Ellison, S.; Foy, C.A.; Scott, D.J. Comparison of microfluidic digital PCR and conventional quantitative PCR for measuring copy number variation. Nucleic Acids Res. 2012, 40, e82. [Google Scholar] [CrossRef]
- Eischeid, A.C. Optimised multiplex droplet digital PCR is more precise, but not more sensitive, than real-time PCR for the detection of allergenic peanut. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2022, 39, 1797–1805. [Google Scholar] [CrossRef]
- Zhang, M.; Xiao, J.; Lin, X.; Yin, W.; Liang, M.; Zhou, L.; Sun, X. Simultaneous Quantification Method of DNA Copy Number for Three Food-borne Pathogens by Multiplex Droplet Digital PCR. J. Agric. Biotechnol. 2022, 30, 606–618. [Google Scholar]
- Pai, T.; Shetty, O.; Joshi, P.; Gurav, M.; Dhanavade, D.; Desai, S. Validation and Comprehensive Analysis of T790M mutation in Liquid Biopsy by Droplet Digital PCR. Lab. Investig. 2022, 102 (Suppl. S1), 1334–1335. [Google Scholar]
- Konstantinos, M.; Maria, P.K.; Maria, R.; Aris, I.; Kleita, M.; Chris, B.; Thomas, V.L.; Anastasia, T.; John, V. Multiple TaqMan qPCR and droplet digital PCR (ddPCR) diagnostics for pesticide resistance monitoring and management, in the major agricultural pest Tetranychus urticae. Pest Manag. Sci. 2021, 78, 263–273. [Google Scholar]
- Wan, P.-J.; Yang, L.; Wang, W.-X.; Fan, J.-M.; Fu, Q.; Li, G.-Q. Constructing the major biosynthesis pathways for amino acids in the brown planthopper, Nilaparvata lugens Stål (Hemiptera: Delphacidae), based on the transcriptome data. Insect Mol. Biol. 2014, 23, 152–164. [Google Scholar] [CrossRef]
- Fan, H.-W.; Noda, H.; Xie, H.-Q.; Suetsugu, Y.; Zhu, Q.-H.; Zhang, C.-X. Genomic Analysis of an Ascomycete Fungus from the Rice Planthopper Reveals How It Adapts to an Endosymbiotic Lifestyle. Genome Biol. Evol. 2015, 7, 2623–2634. [Google Scholar] [CrossRef]
- Chen, Y.H.; Bernal, C.C.; Tan, J.; Horgan, F.G.; Fitzgerald, M.A. Planthopper “adaptation” to resistant rice varieties: Changes in amino acid composition over time. J. Insect Physiol. 2011, 57, 1375–1384. [Google Scholar] [CrossRef]
- Wan, P.-J.; Yang, L.; Yuan, S.-Y.; Tang, Y.-H.; Fu, Q.; Li, G.-Q. RNA interference-aided knockdown of a putative saccharopine dehydrogenase leads to abnormal ecdysis in the brown planthopper, Nilaparvata lugens (Stål) (Hemiptera: Delphacidae). Bull. Entomol. Res. 2015, 105, 390–398. [Google Scholar] [CrossRef]
- Wang, X.L. Study on the Relationship between the Release og Yeast-like Symbionts and Vitellogenin in the Brown Planthopper, Nilaparvata lugens(Stål). Bachelor’s Thesis, China Jiliang University, Hangzhou, China, 2019. [Google Scholar]
- Fan, H.W. The Genomic Analysis Reveals the Symbiotic Relationship between the Brown Planthopper and Ite Endosymbionts. Ph.D. Thesis, ZheJiang University, Hangzhou, China, 2015. [Google Scholar]
- Zhang, J.; He, Y.; Chen, J. Effects of nystatin on the feeding behavior, nutrition and yeast-like symbiont number of the brown planthopper, Nilaparvata lugens (Stål). J. Plant Prot. 2015, 42, 264–270. [Google Scholar]
- Chen, F.J.; Zhang, J.F.; Xia, Z.E.; Lu, Z.X.; Yu, X.P. Morphological observation on the Yeast-like endosymbiotes in brown planthopper, Nilaparvata lugens. Zool. Syst. 2006, 31, 55–62. [Google Scholar]
- Xiong, Z.Z.; Shi, J.T.; Song, Y.; Shentu, X.P.; Yu, X.P. The number changes of endosymbionts in the fat body and gut of the brown planthopper, Nilaparvata Lugens (Stål) at different developmental stages. J. China Univ. Metrol. 2022, 33, 100–105. [Google Scholar]
- Ion, G.-A.; Nejc, R.; Tanja, D.; Maja, R. Droplet digital PCR for absolute quantification of pathogens. Methods Mol. Biol. 2015, 1302, 331–347. [Google Scholar]
- Katarina, C.; Dejan, Š.; Tanja, D.; Jana, Ž.; Kristina, G. Critical points of DNA quantification by real-time PCR—Effects of DNA extraction method and sample matrix on quantification of genetically modified organisms. BMC Biotechnol. 2006, 6, 37. [Google Scholar]
- Yang, R.; Paparini, A.; Monis, P.; Ryan, U. Comparison of next-generation droplet digital PCR (ddPCR) with quantitative PCR (qPCR) for enumeration of Cryptosporidium oocysts in faecal samples. Int. J. Parasitol. 2014, 44, 1105–1113. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Engel, P.; Martinson, V.G.; Moran, N.A. Functional diversity within the simple gut microbiota of the honey bee. Proc. Natl. Acad. Sci. USA 2012, 109, 11002–11007. [Google Scholar] [CrossRef] [PubMed]
- Schillewaert, S.; Parmentier, T.; Vantaux, A.; den Ende, W.V.; Vorburger, C.; Wenseleers, T. The influence of facultative endosymbionts on honeydew carbohydrate and amino acid composition of the black bean aphid Aphis fabae. Physiol. Entomol. 2017, 42, 125–133. [Google Scholar] [CrossRef] [Green Version]
- Hadapad, A.B.; Prabhakar, C.S.; Chandekar, S.C.; Tripathi, J.; Hire, R.S. Diversity of bacterial communities in the midgut of Bactrocera cucurbitae (Diptera: Tephritidae) populations and their potential use as attractants. Pest Manag. Sci. 2016, 72, 1222–1230. [Google Scholar] [CrossRef]
- Pagadala, D.K.J.; Arthikirubha, A.; Vivek, K. Commensal Bacteria Aid Mate-selection in the Fruit Fly, Bactrocera dorsalis. Microb. Ecol. 2016, 72, 725–729. [Google Scholar]
- Ruxin, D.; Lei, M.; Ningxin, W. Research Progresses on Insecticide Resistance Mediated by Symbiotic Bacteria. Biotechnol. Bluuetin 2019, 35, 29–34. [Google Scholar]
- Wang, Z.-Y.; Wang, W.-F.; Lu, Y.-J. Symbiotic microbiota and insecticide resistance in insects. Chin. J. Appl. Entomol. 2021, 58, 265–276. [Google Scholar]
- Bahar, H.; Wist, T.J.; Bekkaoui, D.R.; Hegedus, D.D.; Olivier, C.Y. Aster leafhopper survival and reproduction, and Aster yellows transmission under static and fluctuating temperatures, using ddPCR for phytoplasma quantification. Sci. Rep. 2018, 8, 227. [Google Scholar] [CrossRef] [Green Version]
- Morella, N.M.; Zhang, X.; Koskella, B. Tomato Seed-Associated Bacteria Confer Protection of Seedlings Against Foliar Disease Caused by Pseudomonas syringae. Phytobiomes J. 2019, 3, 177–190. [Google Scholar] [CrossRef] [Green Version]
- Xie, S.; Yu, H.; Wang, Q.; Cheng, Y.; Ding, T. Two rapid and sensitive methods based on TaqMan qPCR and droplet digital PCR assay for quantitative detection of Bacillus subtilis in rhizosphere. J. Appl. Microbiol. 2020, 128, 518–527. [Google Scholar] [CrossRef] [PubMed]
- Zhukova, M.; Sapountzis, P.; Schiøtt, M.; Boomsma, J.J. Diversity and Transmission of Gut Bacteria in Atta and Acromyrmex Leaf-Cutting Ants during Development. Front. Microbiol. 2017, 8, 1942. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Richardson, K.M.; Griffin, P.C.; Lee, S.F.; Ross, P.A.; Endersby-Harshman, N.M.; Schiffer, M.; Hoffmann, A.A. A Wolbachia infection from Drosophila that causes cytoplasmic incompatibility despite low prevalence and densities in males. Heredity 2019, 122, 428–440. [Google Scholar] [CrossRef] [Green Version]
- Hickin, M.L.; Kakumanu, M.L.; Schal, C. Effects of Wolbachia elimination and B-vitamin supplementation on bed bug development and reproduction. Sci. Rep. 2022, 12, 10270. [Google Scholar] [CrossRef]

| Symbionts | Primers | Sequences (5′ → 3′) | Amplification Product Size (bp) |
|---|---|---|---|
| Ascomycetes symbionts | AsF | GTCGTAGTCTTAACCATAA | 145 |
| AsR | CTTCCGTCAATTTCTTTAAG | ||
| AsP | TCAGCCTTGCGACCATA | ||
| Pichia guilliermondii | PgF | CCTCTCAATGTATTAGGTTTA | 101 |
| PgR | TGAGGTCAAACTTGTTTG | ||
| PgP | CCAACAATACCAGAAATATCCCGCC |
| Symbionts | Primer Concentration (nM) | |||||
|---|---|---|---|---|---|---|
| 600 | 700 | 800 | 900 | 1000 | ||
| Ascomycetes symbionts | Copy number copies/μL | 903.00 | 962.00 | 965.00 | 963.00 | 965.00 |
| Pichia guilliermondii | Copy number copies/μL | 73.00 | 80.00 | 79.60 | 134.00 | 76.00 |
| Symbionts | Probe Concentration (nM) | |||||
|---|---|---|---|---|---|---|
| 62.5 | 125 | 250 | 375 | 500 | ||
| Ascomycetes symbionts | Copy number copies/μL | 81.10 | 89.00 | 100.00 | 92.20 | 97.00 |
| Pichia guilliermondii | Copy number copies/μL | 79.20 | 78.90 | 74.90 | 68.80 | 77.50 |
| Dilution Ratio | ddPCR/(copies·μL−1) | ||
|---|---|---|---|
| Result 1 | Result 2 | Result 3 | |
| 10−5 | 401.00 | 468.00 | 630.00 |
| 10−6 | 42.90 | 46.40 | 54.00 |
| 10−7 | 9.30 | 10.30 | 13.60 |
| 10−8 | 1.80 | 1.70 | 1.50 |
| 10−9 | 1.30 | 1.40 | No call |
| Dilution Ratio | ddPCR/(copies·μL−1) | ||
|---|---|---|---|
| Result 1 | Result 2 | Result 3 | |
| 10−5 | 475.00 | 464.00 | 520.00 |
| 10−6 | 40.90 | 84.00 | 85.00 |
| 10−7 | 7.70 | 9.10 | 9.70 |
| 10−8 | 4.60 | 5.90 | 6.90 |
| 10−9 | 1.70 | No call | 1.20 |
| Sample | Ascomycetes Symbionts | Arsenophonus Symbionts | Acinetobacter soli | Pichia guilliermondii | ddH2O |
|---|---|---|---|---|---|
| Copy number copies/μL | 20.50 | 2.60 | 1.70 | 3.60 | 0.80 |
| 22.10 | 1.40 | 1.40 | 5.00 | 0.65 | |
| 23.10 | 1.80 | 1.80 | 5.30 | No call |
| Sample | Ascomycetes Symbionts | Arsenophonus Symbionts | Acinetobacter soli | Pichia guilliermondii | ddH2O |
|---|---|---|---|---|---|
| Copy number copies/μL | 2.20 | 2.40 | 0.66 | 64.20 | 0.56 |
| 1.50 | No call | 0.90 | 63.00 | 0.80 | |
| 2.00 | 1.70 | 1.30 | 71.00 | 0.90 |
| Symbionts | N | Average Value (copies/μL) | Standard Deviation (copies/μL) | Relative Standard Deviation (%) | Results of One-Way ANOVA |
|---|---|---|---|---|---|
| Ascomycetes symbionts | 1 | 84.00 | 2.65 | 3.15 | F = 1.915 p = 0.227 |
| 2 | 80.00 | 2.00 | 2.50 | ||
| 3 | 76.33 | 2.52 | 3.30 | ||
| Pichia guilliermondii | 1 | 35.67 | 1.53 | 4.28 | F = 1.126 p = 0.385 |
| 2 | 35.50 | 0.53 | 1.49 | ||
| 3 | 36.62 | 0.54 | 1.47 |
| Symbiotic Bacteria | Primer | Sequence (5′-3′) | Length of Amplified Product (bp) |
|---|---|---|---|
| Arsenophonus symbionts | ArF | GGGAATATTGCACAATGG | 125 |
| ArR | CGTCAATTGCTAAGGTTA | ||
| ArP | AACCTTAACACCTTCCTCACGACT | ||
| Acinetobacter soli | AcF | GCCAATTAAGTCAAATGTG | 102 |
| AcR | GCTACACCTGGAATTCTA | ||
| AcP | CCACACTCTAGCCAACCAGTATCG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Lai, C.; Shentu, X.; Hao, P.; Pang, K.; Yu, X. Establishment of a Rapid Detection Method for Yeast-like Symbionts in Brown Planthopper Based on Droplet Digital PCR Technology. Int. J. Mol. Sci. 2023, 24, 11071. https://doi.org/10.3390/ijms241311071
Zhang J, Lai C, Shentu X, Hao P, Pang K, Yu X. Establishment of a Rapid Detection Method for Yeast-like Symbionts in Brown Planthopper Based on Droplet Digital PCR Technology. International Journal of Molecular Sciences. 2023; 24(13):11071. https://doi.org/10.3390/ijms241311071
Chicago/Turabian StyleZhang, Jun, Chengling Lai, Xuping Shentu, Peiying Hao, Kun Pang, and Xiaoping Yu. 2023. "Establishment of a Rapid Detection Method for Yeast-like Symbionts in Brown Planthopper Based on Droplet Digital PCR Technology" International Journal of Molecular Sciences 24, no. 13: 11071. https://doi.org/10.3390/ijms241311071






