Structure and Dynamics of Compact Dinucleosomes: Analysis by Electron Microscopy and spFRET
Abstract
1. Introduction
2. Results
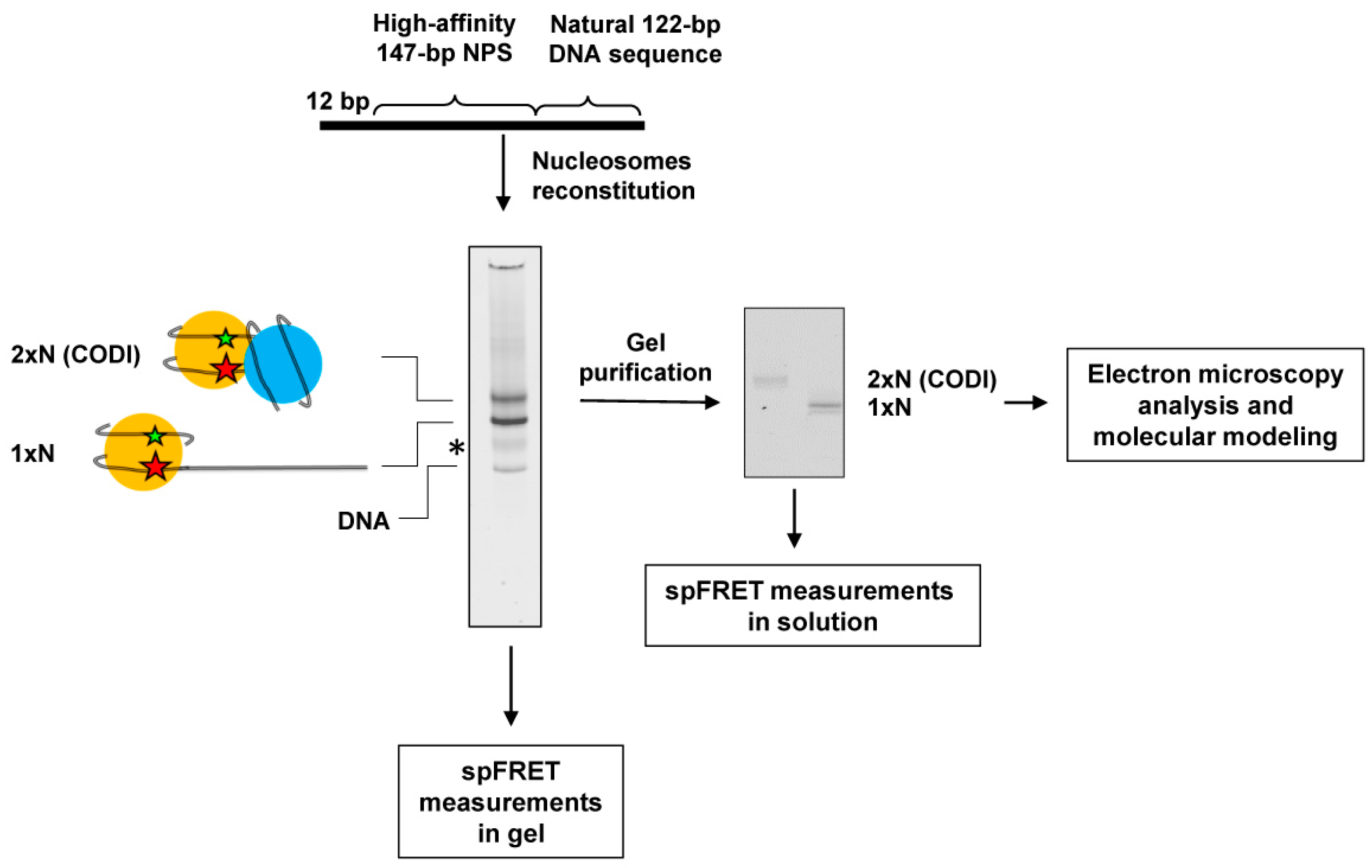
2.1. The Primary Experimental Approach
2.2. Nucleosomal DNA Is Partially Uncoiled in Dinucleosomes
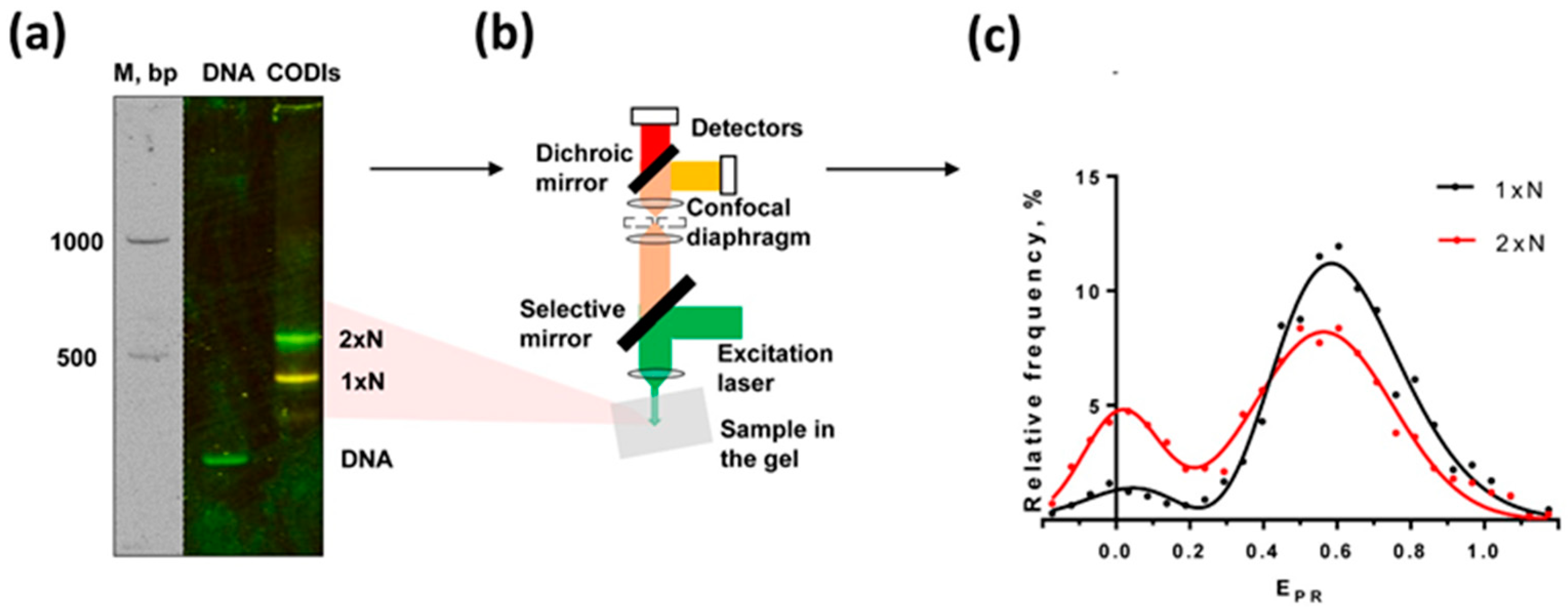
2.3. Multiple Conformations of CODIs Are Present within a Single Band on a Native Gel: spFRET in-Gel (spFRETG) Approach
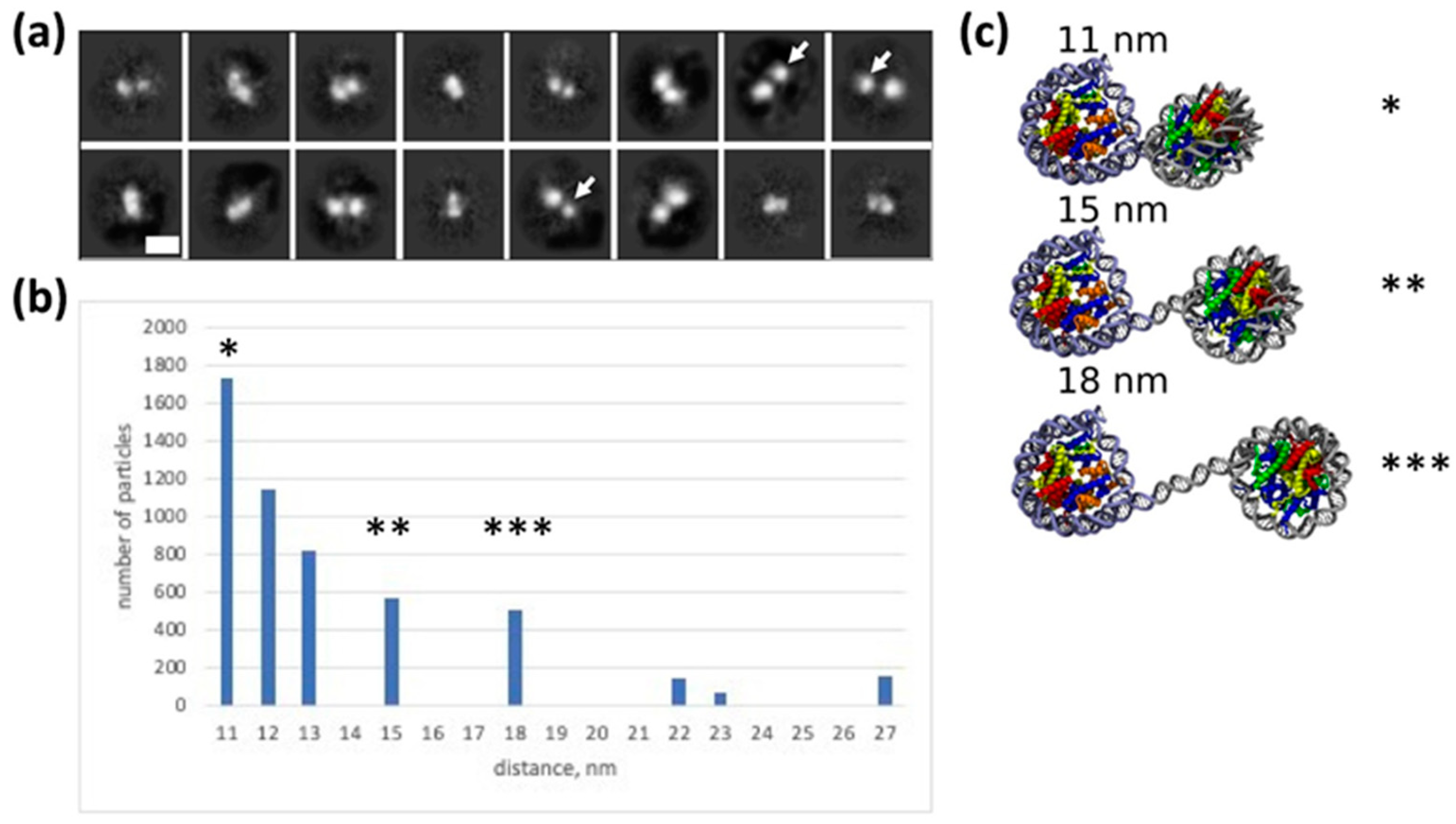
2.4. Heterogeneity in the Structure of CODIs: Analysis by Electron Microscopy
2.5. Modeling Structure and Dynamics of the Dinucleosome
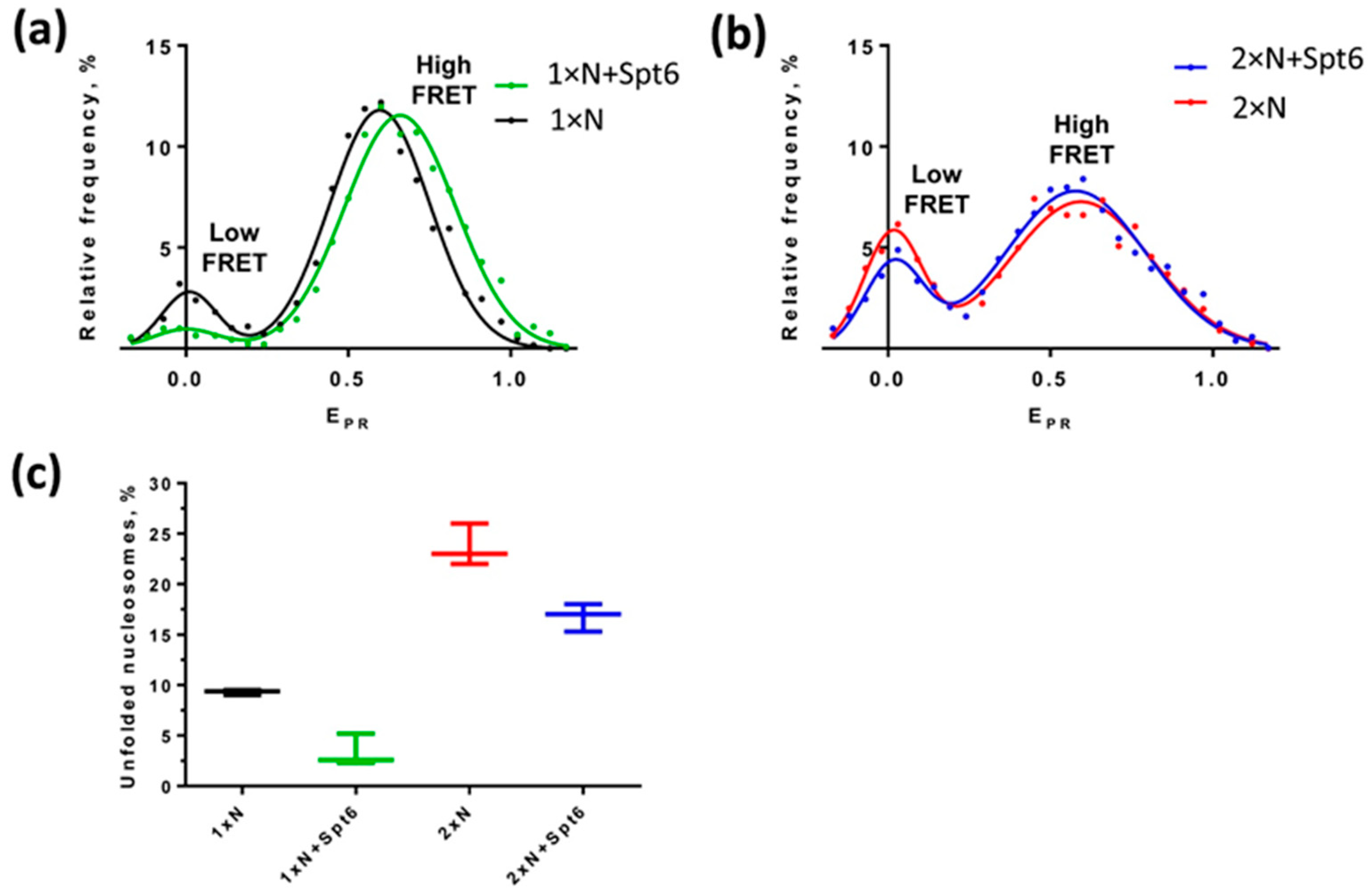
2.6. DNA Uncoiling Is Differently Affected in Mono- and Dinucleosomes by Histone Chaperone Spt6
3. Discussion
4. Materials and Methods
4.1. Proteins and Nucleosomes
4.2. spFRET Measurements in Solution
4.3. FRET and spFRET Measurements in Gel
4.4. Electron Microscopy
4.5. Incubation of Nucleosomes in the Presence of Histone Chaperones
4.6. Modeling Dinucleosome Structures
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bruno, M.; Flaus, A.; Stockdale, C.; Rencurel, C.; Ferreira, H.; Owen-Hughes, T. Histone H2A/H2B Dimer Exchange by ATP-Dependent Chromatin Remodeling Activities. Mol. Cell 2003, 12, 1599–1606. [Google Scholar] [CrossRef] [PubMed]
- Bouazoune, K.; Miranda, T.B.; Jones, P.A.; Kingston, R.E. Analysis of Individual Remodeled Nucleosomes Reveals Decreased Histone-DNA Contacts Created by HSWI/SNF. Nucleic Acids Res. 2009, 37, 5279–5294. [Google Scholar] [CrossRef] [PubMed]
- Clapier, C.R.; Cairns, B.R. The Biology of Chromatin Remodeling Complexes. Annu. Rev. Biochem. 2009, 78, 273–304. [Google Scholar] [CrossRef] [PubMed]
- Mueller-Planitz, F.; Klinker, H.; Becker, P.B. Nucleosome Sliding Mechanisms: New Twists in a Looped History. Nat. Struct. Mol. Biol. 2013, 20, 1026–1032. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, S.K.; Hailu, S.G.; Olufemi, L.; Brahma, S.; Kundu, S.; Hota, S.K.; Persinger, J.; Bartholomew, B. Dinucleosome Specificity and Allosteric Switch of the ISW1a ATP-Dependent Chromatin Remodeler in Transcription Regulation. Nat. Commun. 2020, 11, 5913. [Google Scholar] [CrossRef]
- Gkikopoulos, T.; Schofield, P.; Singh, V.; Pinskaya, M.; Mellor, J.; Smolle, M.; Workman, J.L.; Barton, G.J.; Owen-Hughes, T. A Role for Snf2-Related Nucleosome-Spacing Enzymes in Genome-Wide Nucleosome Organization. Science 2011, 333, 1758–1760. [Google Scholar] [CrossRef]
- Ocampo, J.; Chereji, R.V.; Eriksson, P.R.; Clark, D.J. The ISW1 and CHD1 ATP-Dependent Chromatin Remodelers Compete to Set Nucleosome Spacing in Vivo. Nucleic Acids Res. 2016, 44, 4625–4635. [Google Scholar] [CrossRef]
- Ocampo, J.; Chereji, R.V.; Eriksson, P.R.; Clark, D.J. Contrasting Roles of the RSC and ISW1/CHD1 Chromatin Remodelers in RNA Polymerase II Elongation and Termination. Genome Res. 2019, 29, 407–417. [Google Scholar] [CrossRef]
- Ulyanova, N.P.; Schnitzler, G.R. Human SWI/SNF Generates Abundant, Structurally Altered Dinucleosomes on Polynucleosomal Templates. Mol. Cell. Biol. 2005, 25, 11156–11170. [Google Scholar] [CrossRef]
- Engeholm, M.; de Jager, M.; Flaus, A.; Brenk, R.; van Noort, J.; Owen-Hughes, T. Nucleosomes Can Invade DNA Territories Occupied by Their Neighbors. Nat. Struct. Mol. Biol. 2009, 16, 151–158. [Google Scholar] [CrossRef]
- Dechassa, M.L.; Sabri, A.; Pondugula, S.; Kassabov, S.R.; Chatterjee, N.; Kladde, M.P.; Bartholomew, B. SWI/SNF Has Intrinsic Nucleosome Disassembly Activity That Is Dependent on Adjacent Nucleosomes. Mol. Cell 2010, 38, 590–602. [Google Scholar] [CrossRef] [PubMed]
- Clapier, C.R.; Kasten, M.M.; Parnell, T.J.; Viswanathan, R.; Szerlong, H.; Sirinakis, G.; Zhang, Y.; Cairns, B.R. Regulation of DNA Translocation Efficiency within the Chromatin Remodeler RSC/Sth1 Potentiates Nucleosome Sliding and Ejection. Mol. Cell 2016, 62, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Kato, D.; Osakabe, A.; Arimura, Y.; Mizukami, Y.; Horikoshi, N.; Saikusa, K.; Akashi, S.; Nishimura, Y.; Park, S.-Y.; Nogami, J.; et al. Crystal Structure of the Overlapping Dinucleosome Composed of Hexasome and Octasome. Science 2017, 356, 205–208. [Google Scholar] [CrossRef] [PubMed]
- Lowary, P.T.; Widom, J. New DNA Sequence Rules for High Affinity Binding to Histone Octamer and Sequence-Directed Nucleosome Positioning. J. Mol. Biol. 1998, 276, 19–42. [Google Scholar] [CrossRef]
- Gaykalova, D.A.; Kulaeva, O.I.; Volokh, O.; Shaytan, A.K.; Hsieh, F.-K.; Kirpichnikov, M.P.; Sokolova, O.S.; Studitsky, V.M. Structural Analysis of Nucleosomal Barrier to Transcription. Proc. Natl. Acad. Sci. USA 2015, 112, E5787–E5795. [Google Scholar] [CrossRef]
- Kulaeva, O.I.; Gaykalova, D.A.; Pestov, N.A.; Golovastov, V.V.; Vassylyev, D.G.; Artsimovitch, I.; Studitsky, V.M. Mechanism of Chromatin Remodeling and Recovery during Passage of RNA Polymerase II. Nat. Struct. Mol. Biol. 2009, 16, 1272–1278. [Google Scholar] [CrossRef] [PubMed]
- Valieva, M.E.; Armeev, G.A.; Kudryashova, K.S.; Gerasimova, N.S.; Shaytan, A.K.; Kulaeva, O.I.; McCullough, L.L.; Formosa, T.; Georgiev, P.G.; Kirpichnikov, M.P.; et al. Large-Scale ATP-Independent Nucleosome Unfolding by a Histone Chaperone. Nat. Struct. Mol. Biol. 2016, 23, 1111–1116. [Google Scholar] [CrossRef] [PubMed]
- Valieva, M.E.; Gerasimova, N.S.; Kudryashova, K.S.; Kozlova, A.L.; Kirpichnikov, M.P.; Hu, Q.; Botuyan, M.V.; Mer, G.; Feofanov, A.V.; Studitsky, V.M. Stabilization of Nucleosomes by Histone Tails and by FACT Revealed by SpFRET Microscopy. Cancers 2017, 9, 3. [Google Scholar] [CrossRef] [PubMed]
- Sultanov, D.C.; Gerasimova, N.S.; Kudryashova, K.S.; Maluchenko, N.V.; Kotova, E.Y.; Langelier, M.-F.; Pascal, J.M.; Kirpichnikov, M.P.; Feofanov, A.V.; Studitsky, V.M. Unfolding of Core Nucleosomes by PARP-1 Revealed by SpFRET Microscopy. AIMS Genet. 2017, 4, 21–31. [Google Scholar] [CrossRef]
- Kudryashova, K.S.; Chertkov, O.V.; Nikitin, D.V.; Pestov, N.A.; Kulaeva, O.I.; Efremenko, A.V.; Solonin, A.S.; Kir-pichnikov, M.P.; Studitsky, V.M.; Feofanov, A.V. Preparation of Mononucleosomal Templates for Analysis of Transcription with RNA Polymerase Using SpFRET. Methods Mol. Biol. 2015, 1288, 395–412. [Google Scholar] [CrossRef]
- Vasudevan, D.; Chua, E.Y.D.; Davey, C.A. Crystal Structures of Nucleosome Core Particles Containing the “601” Strong Positioning Sequence. J. Mol. Biol. 2010, 403, 1–10. [Google Scholar] [CrossRef]
- Sivkina, A.L.; Karlova, M.G.; Valieva, M.E.; McCullough, L.L.; Formosa, T.; Shaytan, A.K.; Feofanov, A.V.; Kirpichnikov, M.P.; Sokolova, O.S.; Studitsky, V.M. Electron Microscopy Analysis of ATP-Independent Nucleosome Unfolding by FACT. Commun. Biol. 2022, 5, 2. [Google Scholar] [CrossRef]
- Chertkov, O.V.; Valieva, M.E.; Malyuchenko, N.V.; Feofanov, A.V. Analysis of Nucleosome Structure in Polyacrylamide Gel by the Förster Resonance Energy Transfer Method. Mosc. Univ. Biol. Sci. Bull. 2017, 72, 196–200. [Google Scholar] [CrossRef]
- Bilokapic, S.; Strauss, M.; Halic, M. Structural Rearrangements of the Histone Octamer Translocate DNA. Nat. Commun. 2018, 9, 1330. [Google Scholar] [CrossRef] [PubMed]
- McCullough, L.; Connell, Z.; Petersen, C.; Formosa, T. The Abundant Histone Chaperones Spt6 and FACT Collaborate to Assemble, Inspect, and Maintain Chromatin Structure in Saccharomyces cerevisiae. Genetics 2015, 201, 1031–1045. [Google Scholar] [CrossRef] [PubMed]
- Jeronimo, C.; Robert, F. Histone Chaperones FACT and Spt6 Prevent Histone Variants from Turning into Histone Deviants. BioEssays 2016, 38, 420–426. [Google Scholar] [CrossRef]
- Duina, A.A. Histone Chaperones Spt6 and FACT: Similarities and Differences in Modes of Action at Transcribed Genes. Genet. Res. Int. 2011, 2011, 625210. [Google Scholar] [CrossRef]
- Close, D.; Johnson, S.J.; Sdano, M.A.; McDonald, S.M.; Robinson, H.; Formosa, T.; Hill, C.P. Crystal Structures of the S. Cerevisiae Spt6 Core and C-Terminal Tandem SH2 Domain. J. Mol. Biol. 2011, 408, 697–713. [Google Scholar] [CrossRef]
- Shaytan, A.K.; Armeev, G.A.; Goncearenco, A.; Zhurkin, V.B.; Landsman, D.; Panchenko, A.R. Coupling between Histone Conformations and DNA Geometry in Nucleosomes on a Microsecond Timescale: Atomistic Insights into Nucleosome Functions. J. Mol. Biol. 2016, 428, 221–237. [Google Scholar] [CrossRef]
- Khodabandeh, F.; Fatemi, H.; Mohammad-Rafiee, F. Insight into the Unwrapping of the Dinucleosome. Soft Matter 2020, 16, 4806–4813. [Google Scholar] [CrossRef]
- Ulyanova, N.P.; Schnitzler, G.R. Inverted Factor Access and Slow Reversion Characterize SWI/SNF-Altered Nucleosome Dimers. J. Biol. Chem. 2007, 282, 1018–1028. [Google Scholar] [CrossRef]
- Krajewski, W.A.; Vassiliev, O.L. The Saccharomyces cerevisiae Swi/Snf Complex Can Catalyze Formation of Dimeric Nucleosome Structures in Vitro. Biochemistry 2010, 49, 6531–6540. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Fletcher, A.G.L.; Cheung, V.; Winston, F.; Stargell, L.A. Spn1 Regulates the Recruitment of Spt6 and the Swi/Snf Complex during Transcriptional Activation by RNA Polymerase II. Mol. Cell. Biol. 2008, 28, 1393–1403. [Google Scholar] [CrossRef] [PubMed]
- Erkina, T.Y.; Erkine, A. ASF1 and the SWI/SNF Complex Interact Functionally during Nucleosome Displacement, While FACT Is Required for Nucleosome Reassembly at Yeast Heat Shock Gene Promoters during Sustained Stress. Cell Stress Chaperones 2015, 20, 355–369. [Google Scholar] [CrossRef] [PubMed]
- Martin, B.J.E.; Chruscicki, A.T.; Howe, L.J. Transcription Promotes the Interaction of the FAcilitates Chromatin Transactions (FACT) Complex with Nucleosomes in Saccharomyces cerevisiae. Genetics 2018, 210, 869–881. [Google Scholar] [CrossRef]
- Jeronimo, C.; Angel, A.; Nguyen, V.Q.; Kim, J.M.; Poitras, C.; Lambert, E.; Collin, P.; Mellor, J.; Wu, C.; Robert, F. FACT Is Recruited to the +1 Nucleosome of Transcribed Genes and Spreads in a Chd1-Dependent Manner. Mol. Cell 2021, 81, 3542–3559.e11. [Google Scholar] [CrossRef]
- Kudryashova, K.S.; Nikitin, D.V.; Chertkov, O.V.; Gerasimova, N.S.; Valieva, M.E.; Studitsky, V.M.; Feofanov, A.V. Development of fluorescently labeled mononucleosomes to study transcription mechanisms by method of microscopy of single complexes. Mosc. Univ. Biol. Sci. Bull. 2015, 70, 189–193. [Google Scholar] [CrossRef]
- Gaykalova, D.A.; Kulaeva, O.I.; Bondarenko, V.A.; Studitsky, V.M. Preparation and Analysis of Uniquely Positioned Mononucleosomes. Methods Mol. Biol. Clifton NJ 2009, 523, 109–123. [Google Scholar] [CrossRef]
- Ruone, S.; Rhoades, A.R.; Formosa, T. Multiple Nhp6 Molecules Are Required to Recruit Spt16-Pob3 to Form YFACT Complexes and to Reorganize Nucleosomes. J. Biol. Chem. 2003, 278, 45288–45295. [Google Scholar] [CrossRef]
- Paull, T.T.; Johnson, R.C. DNA Looping by Saccharomyces cerevisiae High Mobility Group Proteins NHP6A/B. Consequences for Nucleoprotein Complex Assembly and Chromatin Condensation. J. Biol. Chem. 1995, 270, 8744–8754. [Google Scholar] [CrossRef]
- Biswas, D.; Yu, Y.; Prall, M.; Formosa, T.; Stillman, D.J. The Yeast FACT Complex Has a Role in Transcriptional Initiation. Mol. Cell. Biol. 2005, 25, 5812–5822. [Google Scholar] [CrossRef] [PubMed]
- Wittmeyer, J.; Joss, L.; Formosa, T. Spt16 and Pob3 of Saccharomyces cerevisiae Form an Essential, Abundant Heterodimer That Is Nuclear, Chromatin-Associated, and Copurifies with DNA Polymerase Alpha. Biochemistry 1999, 38, 8961–8971. [Google Scholar] [CrossRef] [PubMed]
- Jensen, G. Cryo-EM Part B: 3-D Reconstruction; Academic Press: Cambridge, MA, USA, 2010; ISBN 978-0-12-384992-2. [Google Scholar]
- Xin, H.; Takahata, S.; Blanksma, M.; McCullough, L.; Stillman, D.J.; Formosa, T. YFACT Induces Global Accessibility of Nucleosomal DNA without H2A-H2B Displacement. Mol. Cell 2009, 35, 365–376. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stefanova, M.E.; Volokh, O.I.; Chertkov, O.V.; Armeev, G.A.; Shaytan, A.K.; Feofanov, A.V.; Kirpichnikov, M.P.; Sokolova, O.S.; Studitsky, V.M. Structure and Dynamics of Compact Dinucleosomes: Analysis by Electron Microscopy and spFRET. Int. J. Mol. Sci. 2023, 24, 12127. https://doi.org/10.3390/ijms241512127
Stefanova ME, Volokh OI, Chertkov OV, Armeev GA, Shaytan AK, Feofanov AV, Kirpichnikov MP, Sokolova OS, Studitsky VM. Structure and Dynamics of Compact Dinucleosomes: Analysis by Electron Microscopy and spFRET. International Journal of Molecular Sciences. 2023; 24(15):12127. https://doi.org/10.3390/ijms241512127
Chicago/Turabian StyleStefanova, Maria E., Olesya I. Volokh, Oleg V. Chertkov, Grigory A. Armeev, Alexey K. Shaytan, Alexey V. Feofanov, Mikhail P. Kirpichnikov, Olga S. Sokolova, and Vasily M. Studitsky. 2023. "Structure and Dynamics of Compact Dinucleosomes: Analysis by Electron Microscopy and spFRET" International Journal of Molecular Sciences 24, no. 15: 12127. https://doi.org/10.3390/ijms241512127
APA StyleStefanova, M. E., Volokh, O. I., Chertkov, O. V., Armeev, G. A., Shaytan, A. K., Feofanov, A. V., Kirpichnikov, M. P., Sokolova, O. S., & Studitsky, V. M. (2023). Structure and Dynamics of Compact Dinucleosomes: Analysis by Electron Microscopy and spFRET. International Journal of Molecular Sciences, 24(15), 12127. https://doi.org/10.3390/ijms241512127





