Genome-Wide Identification and Expression Analysis of C3H Zinc Finger Family in Potato (Solanum tuberosum L.)
Abstract
:1. Introduction
2. Results
2.1. Identification of C3H Gene Family in Potato
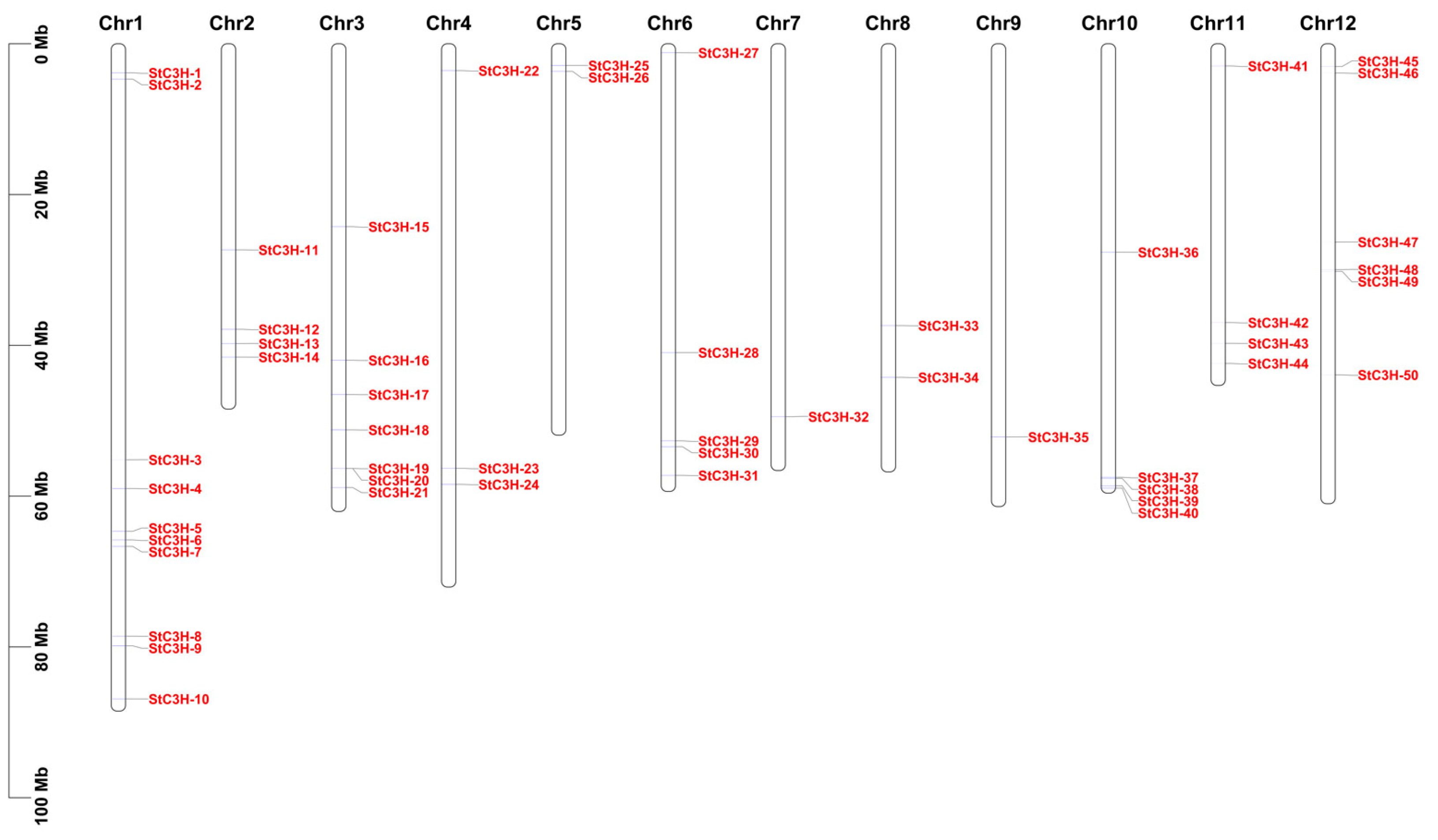
2.2. Location Analysis of C3H Gene Family Members on the Chromosome in Potato
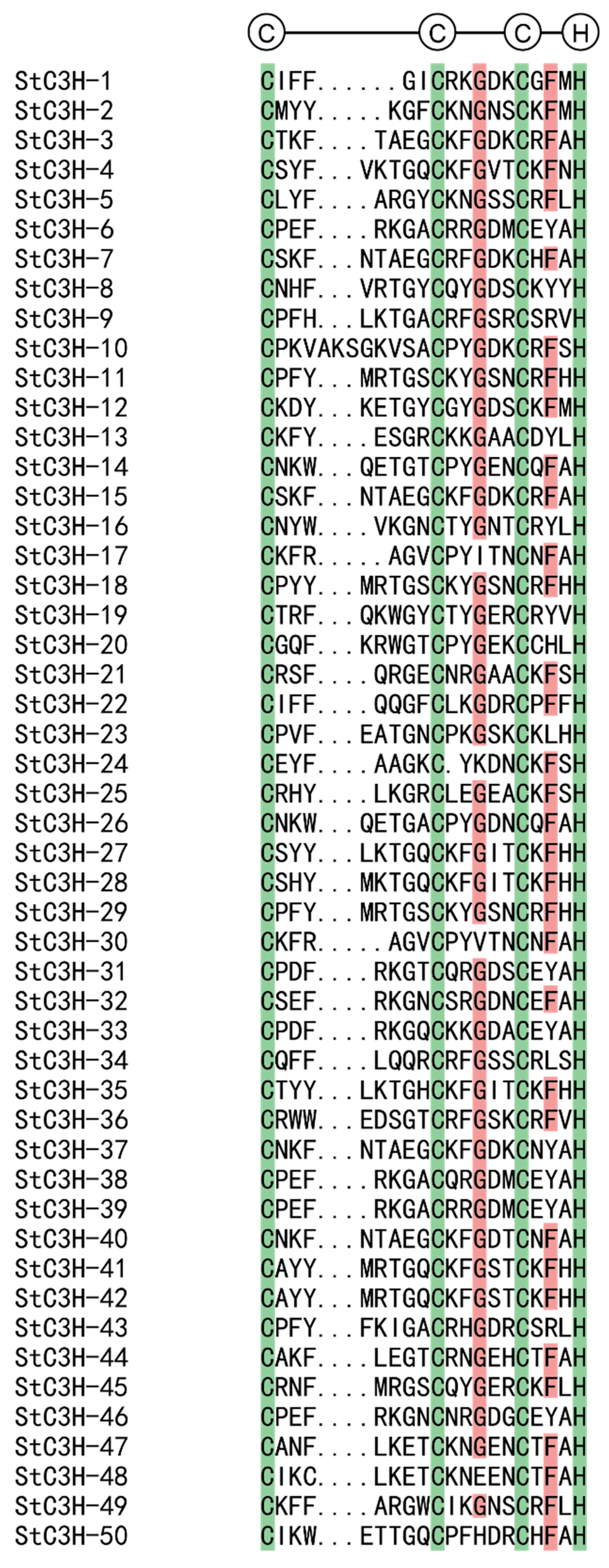
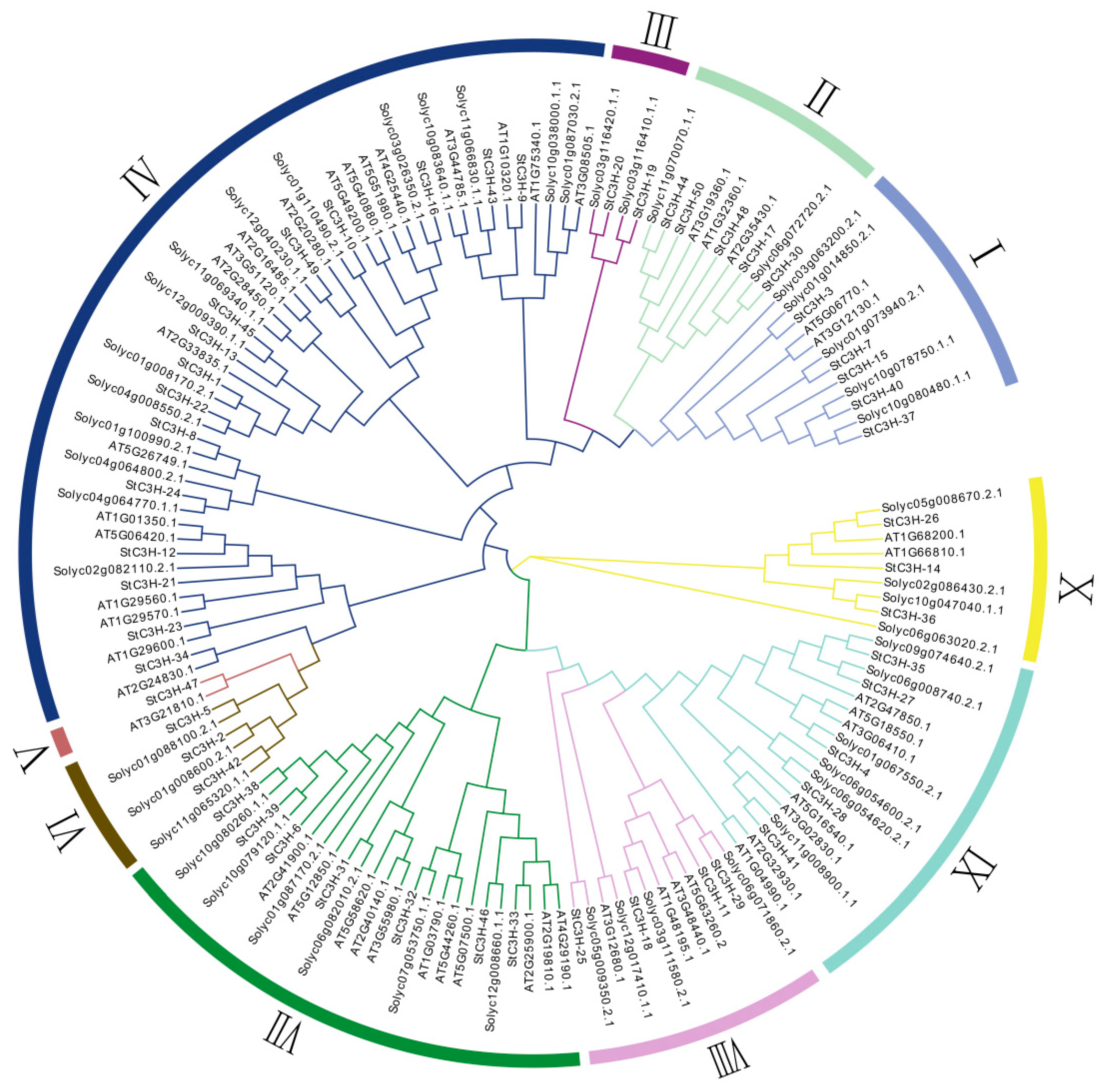
2.3. Multiple Sequence Alignment and Phylogenetic Analysis of C3H Gene Family in Potato
2.4. Analysis of Gene and Protein Structure and Function Prection of C3H Family in Potato
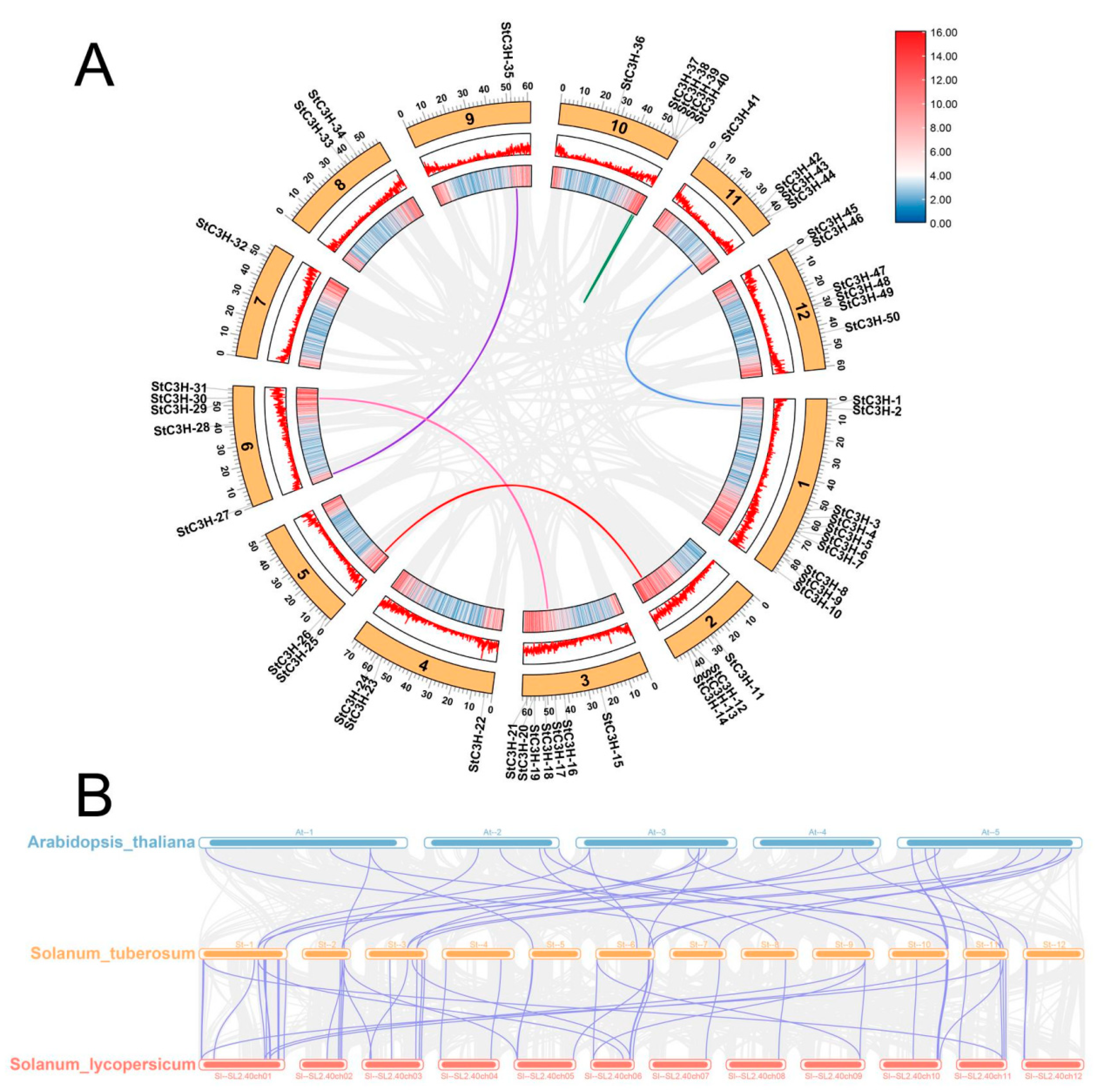
2.5. Collinearity Analysis of C3H Gene Family
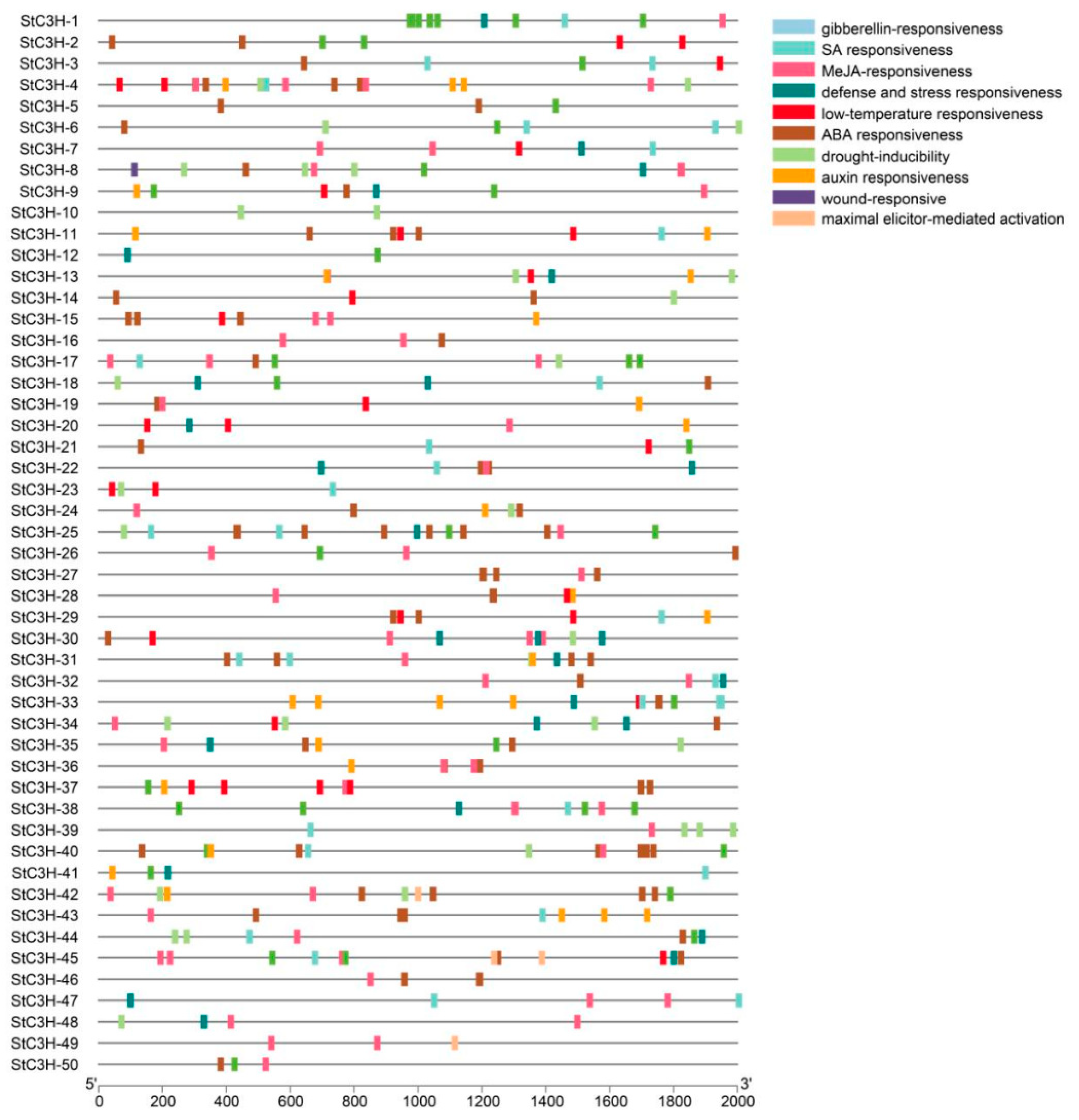
2.6. Cis-Acting Elements Analysis
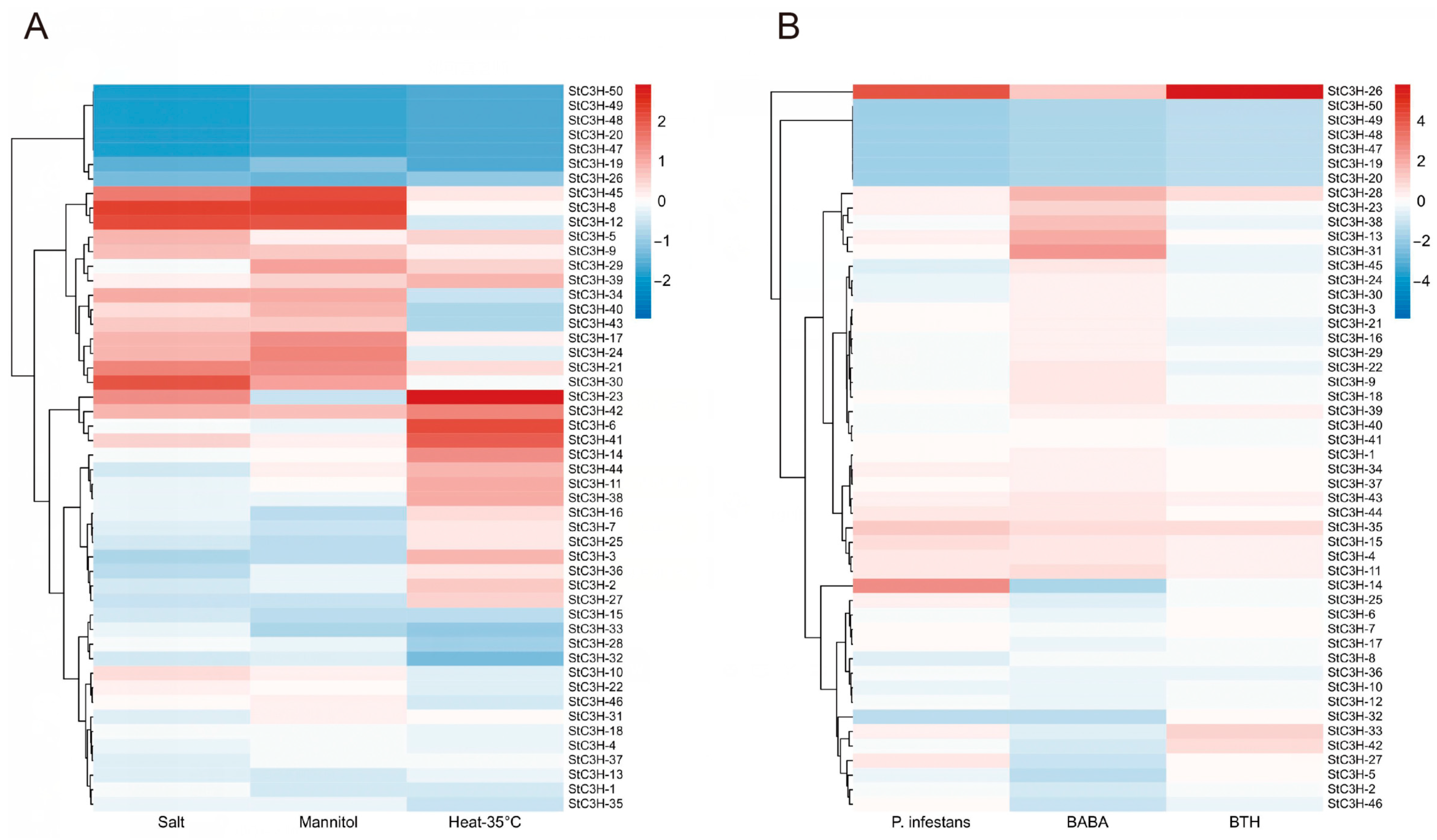
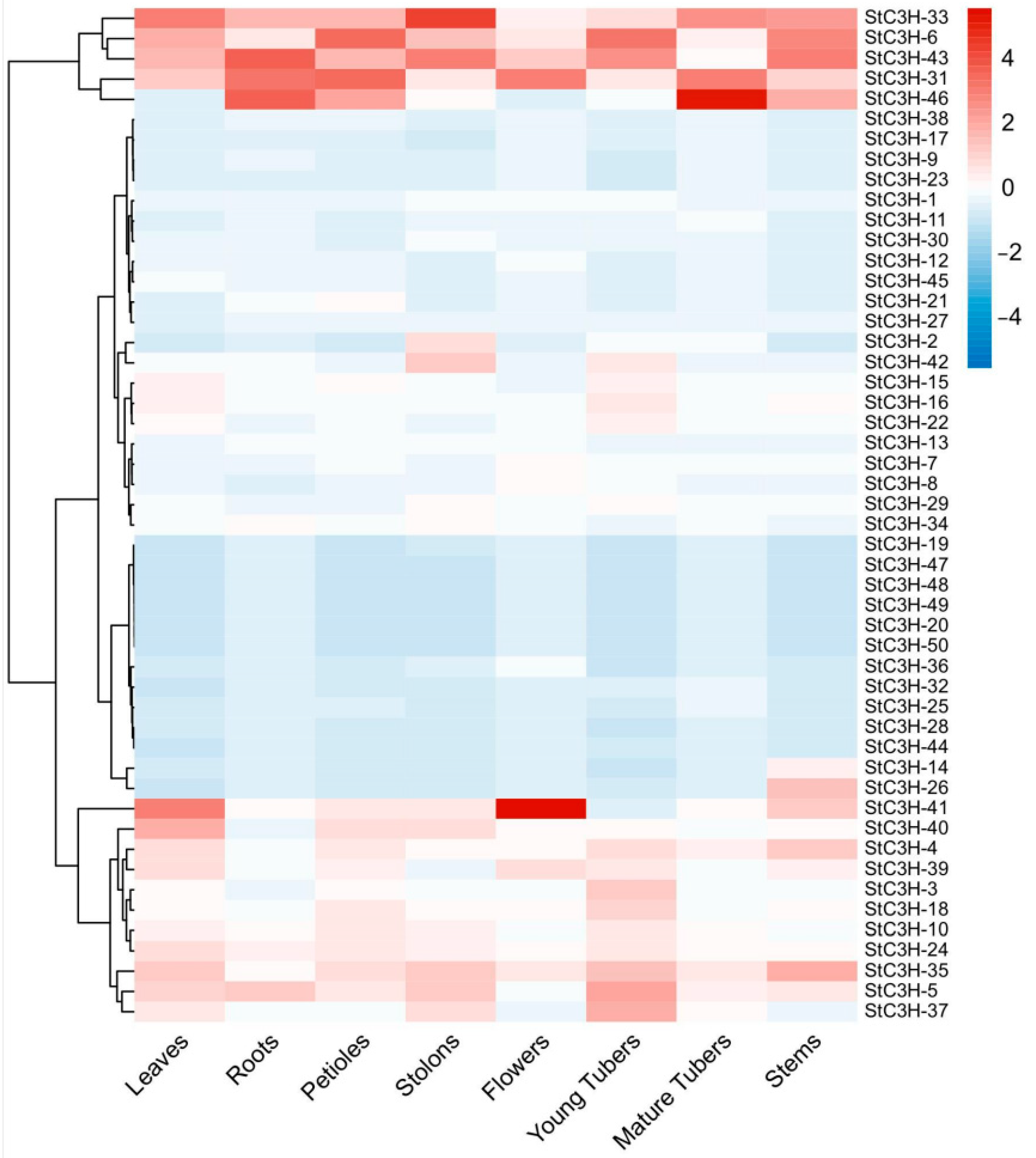
2.7. Expression Analysis of C3H Gene Family in Potato
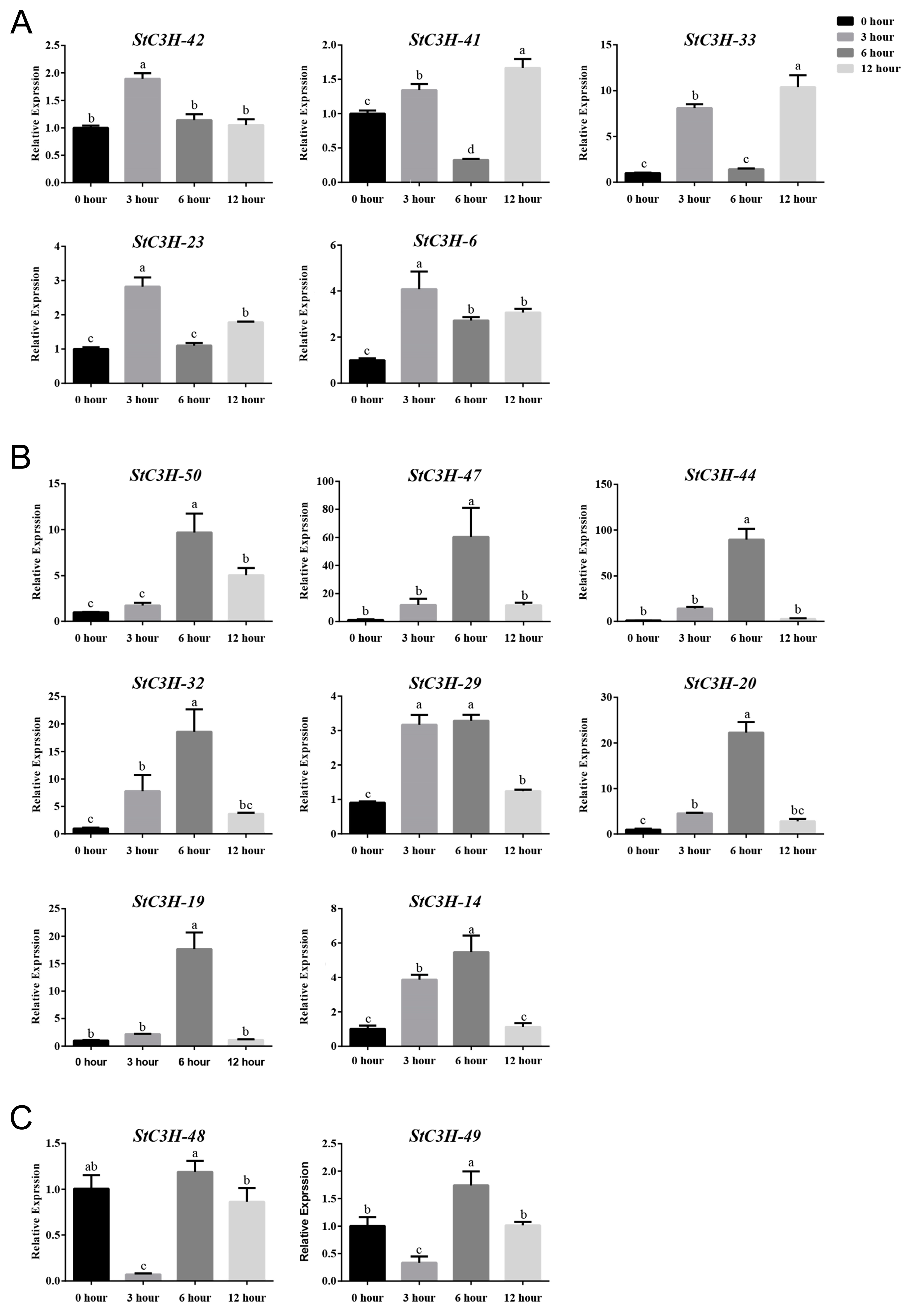
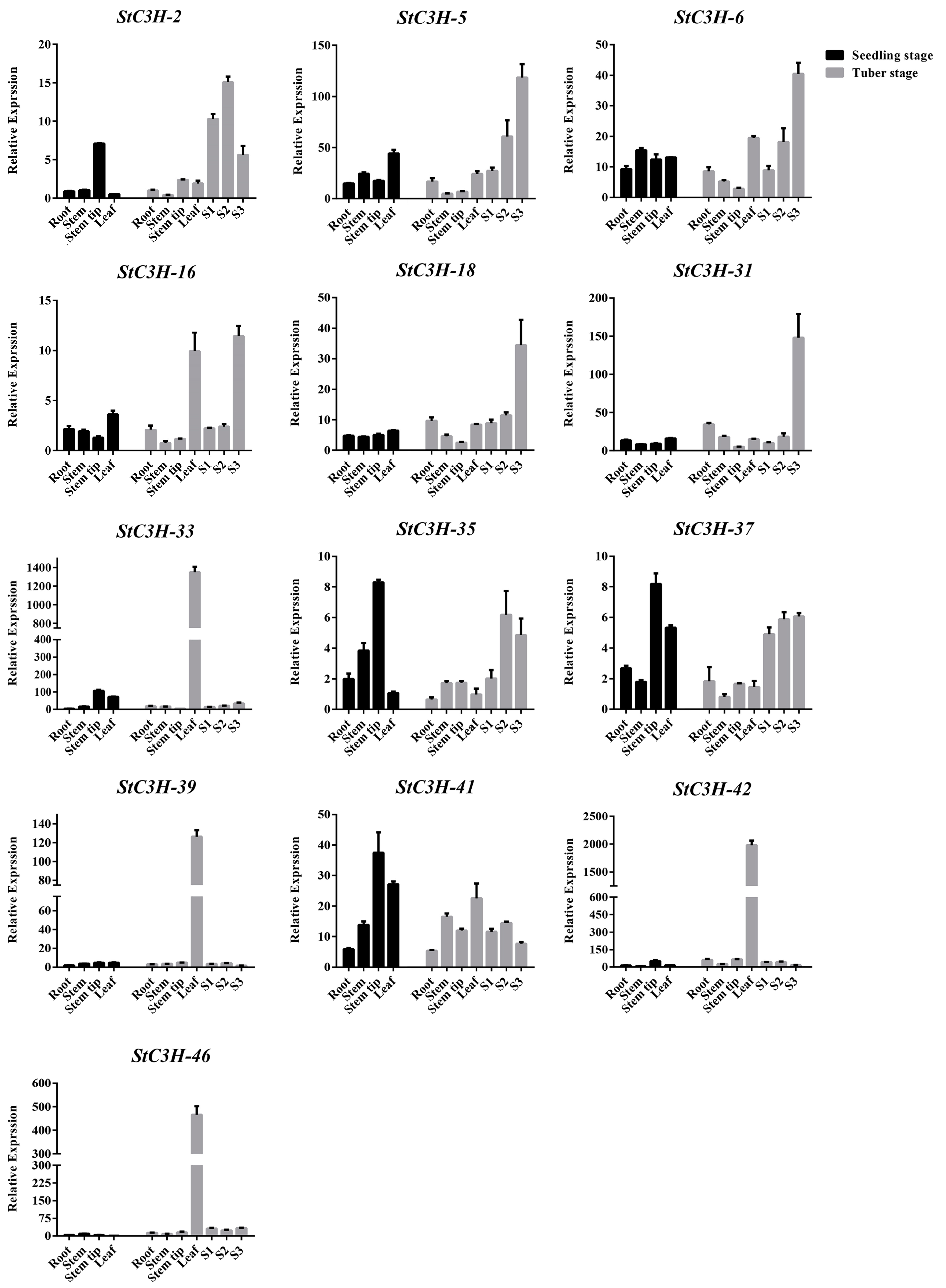
2.8. RT-qPCR Verification of Expression Level of StC3H Gene Family in Potato
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Identification of C3H Gene Family in Potato
4.3. Location Analysis of C3H Gene Family Members on the Chromosome in Potato
4.4. Multiple Sequence Alignment and Phylogenetic Analysis of C3H Gene Family in Potato
4.5. Analysis of Gene and Protein Structure and Function Prection of C3H Family in Potato
4.6. Collinearity Analysis of StC3H Genes in Potato
4.7. Cis-Acting Elements Analysis
4.8. Expression of StC3H Genes in Different Tissues and Its Response to Different Stress
4.9. Total RNA Extraction and RT-qPCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Droge-Laser, W.; Snoek, B.L.; Snel, B.; Weiste, C. The Arabidopsis bZIP transcription factor family—An update. Curr. Opin. Plant Biol. 2018, 45, 36–49. [Google Scholar] [CrossRef] [PubMed]
- Perez-Rodriguez, P.; Riano-Pachon, D.M.; Correa, L.G.G.; Rensing, S.A.; Kersten, B.; Mueller-Roeber, B. PInTFDB: Updated content and new features of the plant transcription factor database. Nucleic Acids Res. 2010, 38, D822–D827. [Google Scholar] [CrossRef] [PubMed]
- Netti, F.; Malgieri, G.; Esposito, S.; Palmieri, M.; Baglivo, I.; Isernia, C.; Omichinski, J.G.; Pedone, P.V.; Lartillot, N.; Fattorusso, R. An Experimentally Tested Scenario for the Structural Evolution of Eukaryotic Cys2His2 Zinc Fingers from Eubacterial Ros Homologs. Mol. Biol. Evol. 2013, 30, 1504–1513. [Google Scholar] [CrossRef]
- Hall, T.M.T. Multiple modes of RNA recognition by zinc finger proteins. Curr. Opin. Struct. Biol. 2005, 15, 367–373. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.S.; Zhou, X. Bioinformatic Analyses of CCCH-Zinc Finger Family in Zebrafish (Danio rerio). Pak. J. Zool. 2014, 46, 989–995. [Google Scholar]
- Liu, S.; Khan, M.R.G.; Li, Y.; Zhang, J.; Hu, C. Comprehensive analysis of CCCH-type zinc finger gene family in citrus (Clementine mandarin) by genome-wide characterization. Mol. Genet. Genom. 2014, 289, 855–872. [Google Scholar] [CrossRef]
- Berg, J.M.; Shi, Y. The galvanization of biology: A growing appreciation for the roles of zinc. Science 1996, 271, 1081–1085. [Google Scholar] [CrossRef]
- Rameneni, J.J.; Dhandapani, V.; Paul, P.; Devaraj, S.P.; Choi, S.R.; Yi, S.Y.; Kim, M.-S.; Hong, S.; Oh, S.H.; Oh, M.-H.; et al. Comprehensive analysis of CCCH zinc-finger-type transcription factors in the Brassica rapa genome. Hortic. Environ. Biotechnol. 2018, 59, 729–747. [Google Scholar] [CrossRef]
- Pi, B.; Pan, J.; Xiao, M.; Hu, X.; Zhang, L.; Chen, M.; Liu, B.; Ruan, Y.; Huang, Y. Systematic analysis of CCCH zinc finger family in Brassica napus showed that BnRR-TZF s are involved in stress resistance. BMC Plant Biol. 2021, 21, 555. [Google Scholar] [CrossRef]
- Seok, H.-Y.; Nguyen, L.V.; Park, H.-Y.; Tarte, V.N.; Ha, J.; Lee, S.-Y.; Moon, Y.-H. Arabidopsis non-TZF gene AtC3H17 functions as a positive regulator in salt stress response. Biochem. Biophys. Res. Commun. 2018, 498, 954–959. [Google Scholar] [CrossRef]
- Wang, D.; Yao, S.; Agassin, R.H.; Zhang, M.; Lou, X.; Huang, Z.; Zhang, J.; Ji, K. Transcriptome-Wide Identification of CCCH-Type Zinc Finger Proteins Family in Pinus massoniana and RR-TZF Proteins in Stress Response. Genes 2022, 13, 1639. [Google Scholar] [CrossRef] [PubMed]
- Bogamuwa, S.P.; Jang, J.C. Tandem CCCH zinc finger proteins in plant growth, development and stress response. Plant Cell Physiol. 2014, 55, 1367–1375. [Google Scholar] [CrossRef] [PubMed]
- Han, G.; Qiao, Z.; Li, Y.; Wang, C.; Wang, B. The Roles of CCCH Zinc-Finger Proteins in Plant Abiotic Stress Tolerance. Int. J. Mol. Sci. 2021, 22, 8327. [Google Scholar] [CrossRef]
- Seong, S.Y.; Shim, J.S.; Bang, S.W.; Kim, J.-K. Overexpression of OsC3H10, a CCCH-zinc finger, improves drought tolerance in rice by regulating stress-related genes. Plants 2020, 9, 1298. [Google Scholar] [CrossRef] [PubMed]
- Jamil, M.; Iqbal, W.; Bangash, A.; Rehman, S.U.; Imran, Q.M.; Rha, E.S. Constitutive expression of OSC3H33, OSC3H50 and OSC3H37 genes in rice under salt stress. Pak. J. Bot. 2010, 42, 4003–4009. [Google Scholar]
- Seok, H.-Y.; Woo, D.-H.; Park, H.-Y.; Lee, S.-Y.; Tran, H.T.; Lee, E.-H.; Vu Nguyen, L.; Moon, Y.-H.J.P.; Physiology, C. AtC3H17, a non-tandem CCCH zinc finger protein, functions as a nuclear transcriptional activator and has pleiotropic effects on vegetative development, flowering and seed development in Arabidopsis. Plant Cell Physiol. 2016, 57, 603–615. [Google Scholar] [CrossRef]
- Lin, P.C.; Pomeranz, M.C.; Jikumaru, Y.; Kang, S.G.; Hah, C.; Fujioka, S.; Kamiya, Y.; Jang, J.C. The Arabidopsis tandem zinc finger protein AtTZF1 affects ABA-and GA-mediated growth, stress and gene expression responses. Plant J. 2011, 65, 253–268. [Google Scholar] [CrossRef]
- Seok, H.-Y.; Bae, H.; Kim, T.; Mehdi, S.M.M.; Nguyen, L.V.; Lee, S.-Y.; Moon, Y.-H. Non-TZF protein AtC3H59/ZFWD3 is involved in seed germination, seedling development, and seed development, interacting with PPPDE family protein Desi1 in Arabidopsis. Int. J. Mol. Sci. 2021, 22, 4738. [Google Scholar] [CrossRef]
- Chen, Y.; Sun, A.; Wang, M.; Zhu, Z.; Ouwerkerk, P.B. Functions of the CCCH type zinc finger protein OsGZF1 in regulation of the seed storage protein GluB-1 from rice. Plant Mol. Biol. 2014, 84, 621–634. [Google Scholar] [CrossRef]
- Guo, Y.H.; Yu, Y.P.; Wang, D.; Wu, C.A.; Yang, G.D.; Huang, J.G.; Zheng, C.C. GhZFP1, a novel CCCH-type zinc finger protein from cotton, enhances salt stress tolerance and fungal disease resistance in transgenic tobacco by interacting with GZIRD21A and GZIPR5. New Phytol. 2009, 183, 62–75. [Google Scholar] [CrossRef]
- Peng, X.; Zhao, Y.; Cao, J.; Zhang, W.; Jiang, H.; Li, X.; Ma, Q.; Zhu, S.; Cheng, B. CCCH-type zinc finger family in maize: Genome-wide identification, classification and expression profiling under abscisic acid and drought treatments. PLoS ONE 2012, 7, e40120. [Google Scholar] [CrossRef] [PubMed]
- Tully, J.P.; Hill, A.E.; Ahmed, H.M.; Whitley, R.; Skjellum, A.; Mukhtar, M.S. Expression-based network biology identifies immune-related functional modules involved in plant defense. BMC Genom. 2014, 15, 421. [Google Scholar] [CrossRef]
- Eriksson, D.; Carlson-Nilsson, U.; Ortíz, R.; Andreasson, E. Overview and breeding strategies of table potato production in Sweden and the Fennoscandian region. Potato Res. 2016, 59, 279–294. [Google Scholar] [CrossRef]
- Wikipedia Contributors. 2008: International Year of the Potato. Wikipedia, The Free Encyclopedia. Available online: https://en.wikipedia.org/w/index.php?title=International_Year_of_the_Potato&oldid=1073524679 (accessed on 3 August 2023).
- Lal, M.K.; Tiwari, R.K.; Kumar, A.; Dey, A.; Kumar, R.; Kumar, D.; Jaiswal, A.; Changan, S.S.; Raigond, P.; Dutt, S. Mechanistic concept of physiological, biochemical, and molecular responses of the potato crop to heat and drought stress. Plants 2022, 11, 2857. [Google Scholar] [CrossRef] [PubMed]
- Begum, S.; Jing, S.; Yu, L.; Sun, X.; Wang, E.; Abu Kawochar, M.; Qin, J.; Liu, J.; Song, B. Modulation of JA signalling reveals the influence of StJAZ1-like on tuber initiation and tuber bulking in potato. Plant J. 2022, 109, 952–964. [Google Scholar] [CrossRef]
- He, H.; He, L.-F. Advances in Potato Breeding for Abiotic Stress Tolerance. In Smart Plant Breeding for Vegetable Crops in Post-Genomics Era; Springer: Berlin/Heidelberg, Germany, 2023; pp. 383–407. [Google Scholar]
- Dahal, K.; Li, X.-Q.; Tai, H.; Creelman, A.; Bizimungu, B. Improving potato stress tolerance and tuber yield under a climate change scenario–a current overview. Front. Plant Sci. 2019, 10, 563. [Google Scholar] [CrossRef]
- Zhang, Q.; Zhang, J.; Wei, F.; Fu, X.; Wei, H.; Lu, J.; Ma, L.; Wang, H. The CCCH-Type Zinc-Finger Protein GhC3H20 Enhances Salt Stress Tolerance in Arabidopsis thaliana and Cotton through ABA Signal Transduction Pathway. Int. J. Mol. Sci. 2023, 24, 5057. [Google Scholar] [CrossRef]
- Xie, Z.; Lin, W.; Yu, G.; Cheng, Q.; Xu, B.; Huang, B. Improved cold tolerance in switchgrass by a novel CCCH-type zinc finger transcription factor gene, PvC3H72, associated with ICE1-CBF-COR regulon and ABA-responsive genes. Biotechnol. Biofuels 2019, 12, 224. [Google Scholar] [CrossRef]
- Xu, L.; Liu, T.; Xiong, X.; Liu, W.; Yu, Y.; Cao, J. Overexpression of Two CCCH-type Zinc-Finger Protein Genes Leads to Pollen Abortion in Brassica campestris ssp. chinensis. Genes 2020, 11, 1287. [Google Scholar] [CrossRef]
- Xu, L.; Xiong, X.; Liu, W.; Liu, T.; Yu, Y.; Cao, J. BcMF30a and BcMF30c, Two Novel Non-Tandem CCCH Zinc-Finger Proteins, Function in Pollen Development and Pollen Germination in Brassica campestris ssp. chinensis. Int. J. Mol. Sci. 2020, 21, 6428. [Google Scholar] [CrossRef]
- Matsushita, K.; Takeuchi, O.; Standley, D.M.; Kumagai, Y.; Kawagoe, T.; Miyake, T.; Satoh, T.; Kato, H.; Tsujimura, T.; Nakamura, H. Zc3h12a is an RNase essential for controlling immune responses by regulating mRNA decay. Nature 2009, 458, 1185–1190. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-J.; Jung, H.J.; Kang, H.; Kim, S.Y. Arabidopsis zinc finger proteins AtC3H49/AtTZF3 and AtC3H20/AtTZF2 are involved in ABA and JA responses. Plant Cell Physiol. 2012, 53, 673–686. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Jia, J.; Yan, X.; Shi, H.; Han, Y. Arabidopsis KHZ1 and KHZ2, two novel non-tandem CCCH zinc-finger and K-homolog domain proteins, have redundant roles in the regulation of flowering and senescence. Plant Mol. Biol. 2017, 95, 549–565. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wei, X.; Jiao, G.; Chen, W.; Wu, Y.; Sheng, Z.; Hu, S.; Xie, L.; Wang, J.; Tang, S.J. GBSS-BINDING PROTEIN, encoding a CBM48 domain-containing protein, affects rice quality and yield. J. Integr. Plant Biol. 2020, 62, 948–966. [Google Scholar] [CrossRef] [PubMed]
- Holub, E.B. The arms race is ancient history in Arabidopsis, the wildflower. Nat. Rev. Genet. 2001, 2, 516–527. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Liang, J.; Song, W.; Tromp, G.; Kolattukudy, P.E.; Fu, M. Genome-wide survey and expression profiling of CCCH-zinc finger family reveals a functional module in macrophage activation. PLoS ONE 2008, 3, e2880. [Google Scholar] [CrossRef]
- Flagel, L.E.; Wendel, J.F. Gene duplication and evolutionary novelty in plants. New Phytol. 2009, 183, 557–564. [Google Scholar] [CrossRef]
- Obiero, C.O.; Milroy, S.P.; Bell, R.W. Importance of whole plant dry matter dynamics for potato (Solanum tuberosum L.) tuber yield response to an episode of high temperature. Environ. Exp. Bot. 2019, 162, 560–571. [Google Scholar] [CrossRef]
- Ahmad, M.; Yan, X.; Li, J.; Yang, Q.; Jamil, W.; Teng, Y.; Bai, S. Genome wide identification and predicted functional analyses of NAC transcription factors in Asian pears. BMC Plant Biol. 2018, 18, 214. [Google Scholar] [CrossRef] [PubMed]
- Riechmann, J.L.; Heard, J.; Martin, G.; Reuber, L.; Jiang, C.-Z.; Keddie, J.; Adam, L.; Pineda, O.; Ratcliffe, O.; Samaha, R. Arabidopsis transcription factors: Genome-wide comparative analysis among eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar] [CrossRef] [PubMed]
- Wray, G.A.; Hahn, M.W.; Abouheif, E.; Balhoff, J.P.; Pizer, M.; Rockman, M.V.; Romano, L.A. The evolution of transcriptional regulation in eukaryotes. Mol. Biol. Evol. 2003, 20, 1377–1419. [Google Scholar] [CrossRef] [PubMed]
- Ai, Q.; Pan, W.; Zeng, Y.; Li, Y.; Cui, L. CCCH Zinc finger genes in Barley: Genome-wide identification, evolution, expression and haplotype analysis. BMC Plant Biol. 2022, 22, 117. [Google Scholar] [CrossRef]
- Tang, W.; Hao, Y.; Ma, X.; Shi, Y.; Dang, Y.; Dong, Z.; Zhao, Y.; Zhao, T.; Zhu, S.; Zhang, Z.; et al. Genome-wide analysis and identification of stress-responsive genes of the CCCH zinc finger family in Capsicum annuum L. Front. Plant Sci. 2023, 14, 1189038. [Google Scholar] [CrossRef]
- Liu, C.; Xu, X.; Kan, J.; Cheng, Z.M.; Chang, Y.; Lin, J.; Li, H. Genome-wide analysis of the C3H zinc finger family reveals its functions in salt stress responses of Pyrus betulaefolia. PeerJ 2020, 8, e9328. [Google Scholar] [CrossRef]
- Su, L.-Y.; Xiao, X.-C.; Jiang, M.-Q.; Huang, S.-Q.; Xue, X.-D.; Xue, L.; Lai, Z.-X.; Lin, Y.-L. Genome-wide analysis of the CCCH zinc finger family in longan: Characteristic identification and expression profiles in Dimocarpus longan Lour. J. Integr. Agric. 2022, 21, 113–130. [Google Scholar] [CrossRef]
- Wang, D.; Guo, Y.; Wu, C.; Yang, G.; Li, Y.; Zheng, C. Genome-wide analysis of CCCH zinc finger family in Arabidopsis and rice. BMC Genom. 2008, 9, 44. [Google Scholar] [CrossRef]
- Chai, G.; Hu, R.; Zhang, D.; Qi, G.; Zuo, R.; Cao, Y.; Chen, P.; Kong, Y.; Zhou, G. Comprehensive analysis of CCCH zinc finger family in poplar (Populus trichocarpa). BMC Genom. 2012, 13, 253. [Google Scholar] [CrossRef]
- Zhang, C.; Zhang, H.; Zhao, Y.; Jiang, H.; Zhu, S.; Cheng, B.; Xiang, Y. Genome-wide analysis of the CCCH zinc finger gene family in Medicago truncatula. Plant Cell Rep. 2013, 32, 1543–1555. [Google Scholar] [CrossRef]
- Deng, J.; Chen, Y.; Chen, J.; Wang, Y.; Ou, X.; Sun, L.; Zhang, T.; Jian, W. Genome-wide identification and expression analysis of the tomato C3H gene family. Acta Plant Physiol. 2023, 59, 619–631. [Google Scholar] [CrossRef]
- Li, C.-H.; Fang, Q.-X.; Zhang, W.-J.; Li, Y.-H.; Zhang, J.-Z.; Chen, S.; Yin, Z.-G.; Li, W.-J.; Liu, W.-D.; Yi, Z.; et al. Genome-wide identification of the CCCH gene family in rose (Rosa chinensis Jacq.) reveals its potential functions. Biotechnol. Biotechnol. Equip. 2021, 35, 517–526. [Google Scholar] [CrossRef]
- Pi, B.; He, X.; Ruan, Y.; Jang, J.-C.; Huang, Y. Genome-wide analysis and stress-responsive expression of CCCH zinc finger family genes in Brassica rapa. BMC Plant Biol. 2018, 18, 373. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Zuo, J. The CCCH zinc finger family of soybean (Glycine max L.): Genome-wide identification, expression, domestication, GWAS and haplotype analysis. BMC Genom. 2021, 22, 511. [Google Scholar] [CrossRef] [PubMed]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef]
- Ouna, B.A.; Stewart, M.; Helbig, C.; Clayton, C. The Trypanosoma brucei CCCH zinc finger proteins ZC3H12 and ZC3H13. Mol. Biochem. Parasitol. 2012, 183, 184–188. [Google Scholar] [CrossRef]
- Zhang, D.; Xu, Z.; Cao, S.; Chen, K.; Li, S.; Liu, X.; Gao, C.; Zhang, B.; Zhou, Y. An uncanonical CCCH-tandem zinc-finger protein represses secondary wall synthesis and controls mechanical strength in rice. Mol. Plant 2018, 11, 163–174. [Google Scholar] [CrossRef]
- Pomeranz, M.C.; Hah, C.; Lin, P.-C.; Kang, S.G.; Finer, J.J.; Blackshear, P.J.; Jang, J.-C. The Arabidopsis tandem zinc finger protein AtTZF1 traffics between the nucleus and cytoplasmic foci and binds both DNA and RNA. Plant Physiol. 2010, 152, 151–165. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, L.; Xing, X.; Sun, L.; Pan, J.; Kong, X.; Zhang, M.; Li, D. ZmLEA3, a Multifunctional Group 3 LEA Protein from Maize (Zea mays L.), is Involved in Biotic and Abiotic Stresses. Plant Cell Physiol. 2013, 54, 944–959. [Google Scholar] [CrossRef]
- Park, J.S.; Park, S.J.; Kwon, S.Y.; Shin, A.Y.; Moon, K.B.; Park, J.M.; Cho, H.S.; Park, S.U.; Jeon, J.H.; Kim, H.S.; et al. Temporally distinct regulatory pathways coordinate thermo-responsive storage organ formation in potato. Cell Rep. 2022, 38, 110579. [Google Scholar] [CrossRef]
- Xu, W.; Jian, S.; Li, J.; Wang, Y.; Zhang, M.; Xia, K.J. Genomic Identification of CCCH-Type Zinc Finger Protein Genes Reveals the Role of HuTZF3 in Tolerance of Heat and Salt Stress of Pitaya (Hylocereus polyrhizus). Int. J. Mol. Sci. 2023, 24, 6359. [Google Scholar] [CrossRef]
- Navarro, C.; Abelenda, J.A.; Cruz-Oró, E.; Cuéllar, C.A.; Tamaki, S.; Silva, J.; Shimamoto, K.; Prat, S. Control of flowering and storage organ formation in potato by FLOWERING LOCUS T. Nature 2011, 478, 119–122. [Google Scholar] [CrossRef]
- Wang, M.; Zhang, H.; Dai, S.; Feng, S.; Gong, S.; Wang, J.; Zhou, A. AaZFP3, a Novel CCCH-Type Zinc Finger Protein from Adonis amurensis, Promotes Early Flowering in Arabidopsis by Regulating the Expression of Flowering-Related Genes. Int. J. Mol. Sci. 2022, 23, 8166. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Xiao, S.; Sui, S.; Huang, R.; Wang, X.; Wu, H.; Liu, X. A tandem CCCH type zinc finger protein gene CpC3H3 from Chimonanthus praecox promotes flowering and enhances drought tolerance in Arabidopsis. BMC Plant Biol. 2022, 22, 506. [Google Scholar] [CrossRef] [PubMed]
- Chao, Y.; Zhang, T.; Yang, Q.; Kang, J.; Sun, Y.; Gruber, M.Y.; Qin, Z. Expression of the alfalfa CCCH-type zinc finger protein gene MsZFN delays flowering time in transgenic Arabidopsis thaliana. Plant Sci. 2014, 215, 92–99. [Google Scholar] [CrossRef] [PubMed]
- Jang, J.-C. Arginine-rich motif-tandem CCCH zinc finger proteins in plant stress responses and post-transcriptional regulation of gene expression. Plant Sci. 2016, 252, 118–124. [Google Scholar] [CrossRef] [PubMed]
- Bogamuwa, S.; Jang, J.C. The A rabidopsis tandem CCCH zinc finger proteins AtTZF4, 5 and 6 are involved in light-, abscisic acid-and gibberellic acid-mediated regulation of seed germination. Plant Cell Environ. 2013, 36, 1507–1519. [Google Scholar] [CrossRef]
- Jing, S.; Sun, X.; Yu, L.; Wang, E.; Cheng, Z.; Liu, H.; Jiang, P.; Qin, J.; Begum, S.; Song, B. Transcription factor StABI5-like 1 binding to the FLOWERING LOCUS T homologs promotes early maturity in potato. Plant Physiol. 2022, 189, 1677–1693. [Google Scholar] [CrossRef]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.E.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server; Springer: Berlin/Heidelberg, Germany, 2005. [Google Scholar]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chen, Q.; Liu, B. iDRBP_MMC: Identifying DNA-Binding Proteins and RNA-Binding Proteins Based on Multi-Label Learning Model and Motif-Based Convolutional Neural Network. J. Mol. Biol. 2020, 432, 5860–5875. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]


| Gene Name | Gene ID | Protein | Subcellular Location | |||
|---|---|---|---|---|---|---|
| Number of Amino Acids (aa) | Molecular Weight (Da) | Theoretical pI | Grand Average of Hydropathicity | |||
| StC3H-1 | PGSC0003DMG400016400 | 707 | 79,152.04 | 5.80 | −1.060 | Nucleus |
| StC3H-2 | PGSC0003DMG400010493 | 650 | 71,922.33 | 6.00 | −0.570 | Nucleus |
| StC3H-3 | PGSC0003DMG400024054 | 223 | 23,792.81 | 9.01 | −0.480 | Nucleus |
| StC3H-4 | PGSC0003DMG400022651 | 492 | 51,506.80 | 8.41 | −0.390 | Nucleus |
| StC3H-5 | PGSC0003DMG400006828 | 739 | 80,448.74 | 5.93 | −0.503 | Nucleus |
| StC3H-6 | PGSC0003DMG400006803 | 730 | 79,277.70 | 6.02 | −0.468 | Nucleus |
| StC3H-7 | PGSC0003DMG400006330 | 289 | 30,656.69 | 9.33 | −0.438 | Nucleus |
| StC3H-8 | PGSC0003DMG400002748 | 129 | 14,562.29 | 8.61 | −0.888 | Cytoplasm |
| StC3H-9 | PGSC0003DMG400020705 | 761 | 89,133.84 | 6.62 | −1.436 | Peroxisome |
| StC3H-10 | PGSC0003DMG400001658 | 702 | 77,141.25 | 6.67 | −0.512 | Nucleus |
| StC3H-11 | PGSC0003DMG400010434 | 477 | 53,542.39 | 5.78 | −0.882 | Nucleus |
| StC3H-12 | PGSC0003DMG400013655 | 338 | 38,113.47 | 7.12 | −1.036 | Nucleus |
| StC3H-13 | PGSC0003DMG400003622 | 858 | 91,533.33 | 5.48 | −0.722 | Nucleus |
| StC3H-14 | PGSC0003DMG400030591 | 341 | 38,323.55 | 6.02 | −0.748 | Nucleus |
| StC3H-15 | PGSC0003DMG400009821 | 284 | 30,014.31 | 9.57 | −0.381 | Cytoplasm |
| StC3H-16 | PGSC0003DMG400014305 | 430 | 47,162.96 | 7.88 | −0.374 | Chloroplast |
| StC3H-17 | PGSC0003DMG400026471 | 349 | 38,383.75 | 6.19 | −0.690 | Nucleus |
| StC3H-18 | PGSC0003DMG400015168 | 518 | 58,326.50 | 5.14 | −1.168 | Nucleus |
| StC3H-19 | PGSC0003DMG400000604 | 224 | 24,991.78 | 8.80 | −0.724 | Nucleus |
| StC3H-20 | PGSC0003DMG400038576 | 212 | 24,095.85 | 6.14 | −0.979 | Nucleus |
| StC3H-21 | PGSC0003DMG400005636 | 352 | 41,316.02 | 8.51 | −1.540 | Nucleus |
| StC3H-22 | PGSC0003DMG400031374 | 708 | 79,259.80 | 5.93 | −1.084 | Nucleus |
| StC3H-23 | PGSC0003DMG402025318 | 159 | 17,809.09 | 6.43 | −0.834 | Chloroplast |
| StC3H-24 | PGSC0003DMG400024801 | 858 | 96,138.64 | 9.10 | −1.114 | Nucleus |
| StC3H-25 | PGSC0003DMG400014565 | 832 | 91,063.68 | 6.40 | −0.707 | Nucleus |
| StC3H-26 | PGSC0003DMG400030567 | 338 | 37,972.62 | 7.72 | −0.684 | Nucleus |
| StC3H-27 | PGSC0003DMG400005333 | 397 | 43,110.49 | 8.51 | −0.455 | Nucleus |
| StC3H-28 | PGSC0003DMG400030758 | 396 | 42,133.40 | 6.10 | −0.380 | Nucleus |
| StC3H-29 | PGSC0003DMG402027067 | 603 | 67,135.00 | 5.15 | −0.862 | Nucleus |
| StC3H-30 | PGSC0003DMG400026937 | 357 | 39,088.35 | 6.79 | −0.743 | Nucleus |
| StC3H-31 | PGSC0003DMG400030361 | 668 | 73,112.19 | 5.97 | −0.434 | Nucleus |
| StC3H-32 | PGSC0003DMG400032829 | 427 | 47,977.30 | 7.59 | −0.725 | Nucleus |
| StC3H-33 | PGSC0003DMG400025459 | 350 | 39,389.36 | 6.46 | −0.579 | Nucleus |
| StC3H-34 | PGSC0003DMG400014784 | 495 | 55,654.30 | 5.53 | −0.779 | Nucleus |
| StC3H-35 | PGSC0003DMG400011383 | 435 | 46,976.53 | 8.81 | −0.315 | Nucleus |
| StC3H-36 | PGSC0003DMG400024171 | 336 | 36,065.91 | 7.59 | −0.534 | Nucleus |
| StC3H-37 | PGSC0003DMG400023710 | 294 | 30,815.82 | 9.30 | −0.499 | Cytoplasm |
| StC3H-38 | PGSC0003DMG400023719 | 697 | 76,264.47 | 6.83 | −0.478 | Nucleus |
| StC3H-39 | PGSC0003DMG400008150 | 701 | 76,411.82 | 6.61 | −0.444 | Nucleus |
| StC3H-40 | PGSC0003DMG400011414 | 301 | 31,381.47 | 9.35 | −0.503 | Chloroplast |
| StC3H-41 | PGSC0003DMG400016236 | 433 | 47,084.25 | 8.85 | −0.463 | Nucleus |
| StC3H-42 | PGSC0003DMG400029993 | 701 | 76,760.82 | 5.82 | −0.489 | Nucleus |
| StC3H-43 | PGSC0003DMG400026524 | 331 | 38,988.95 | 9.62 | −1.358 | Nucleus |
| StC3H-44 | PGSC0003DMG400025515 | 335 | 38,362.12 | 6.76 | −0.885 | Nucleus |
| StC3H-45 | PGSC0003DMG400002893 | 425 | 45,207.41 | 8.58 | −0.687 | Nucleus |
| StC3H-46 | PGSC0003DMG400000350 | 372 | 42,312.18 | 5.84 | −0.815 | Nucleus |
| StC3H-47 | PGSC0003DMG400046840 | 215 | 24,845.06 | 6.50 | −0.899 | Cytoplasm |
| StC3H-48 | PGSC0003DMG400005394 | 385 | 43,943.14 | 6.08 | −0.497 | Cytoskeleton |
| StC3H-49 | PGSC0003DMG400045292 | 414 | 47,260.84 | 6.08 | −0.710 | Nucleus |
| StC3H-50 | PGSC0003DMG400043565 | 281 | 32,097.45 | 6.56 | −0.702 | Cytoskeleton |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Deng, Z.; Yang, Z.; Liu, X.; Dai, X.; Zhang, J.; Deng, K. Genome-Wide Identification and Expression Analysis of C3H Zinc Finger Family in Potato (Solanum tuberosum L.). Int. J. Mol. Sci. 2023, 24, 12888. https://doi.org/10.3390/ijms241612888
Deng Z, Yang Z, Liu X, Dai X, Zhang J, Deng K. Genome-Wide Identification and Expression Analysis of C3H Zinc Finger Family in Potato (Solanum tuberosum L.). International Journal of Molecular Sciences. 2023; 24(16):12888. https://doi.org/10.3390/ijms241612888
Chicago/Turabian StyleDeng, Zeyi, Zhijiang Yang, Xinyan Liu, Xiumei Dai, Jiankui Zhang, and Kexuan Deng. 2023. "Genome-Wide Identification and Expression Analysis of C3H Zinc Finger Family in Potato (Solanum tuberosum L.)" International Journal of Molecular Sciences 24, no. 16: 12888. https://doi.org/10.3390/ijms241612888






