Metabolomic Analysis of Trehalose Alleviating Oxidative Stress in Myoblasts
Abstract
:1. Introduction
2. Results
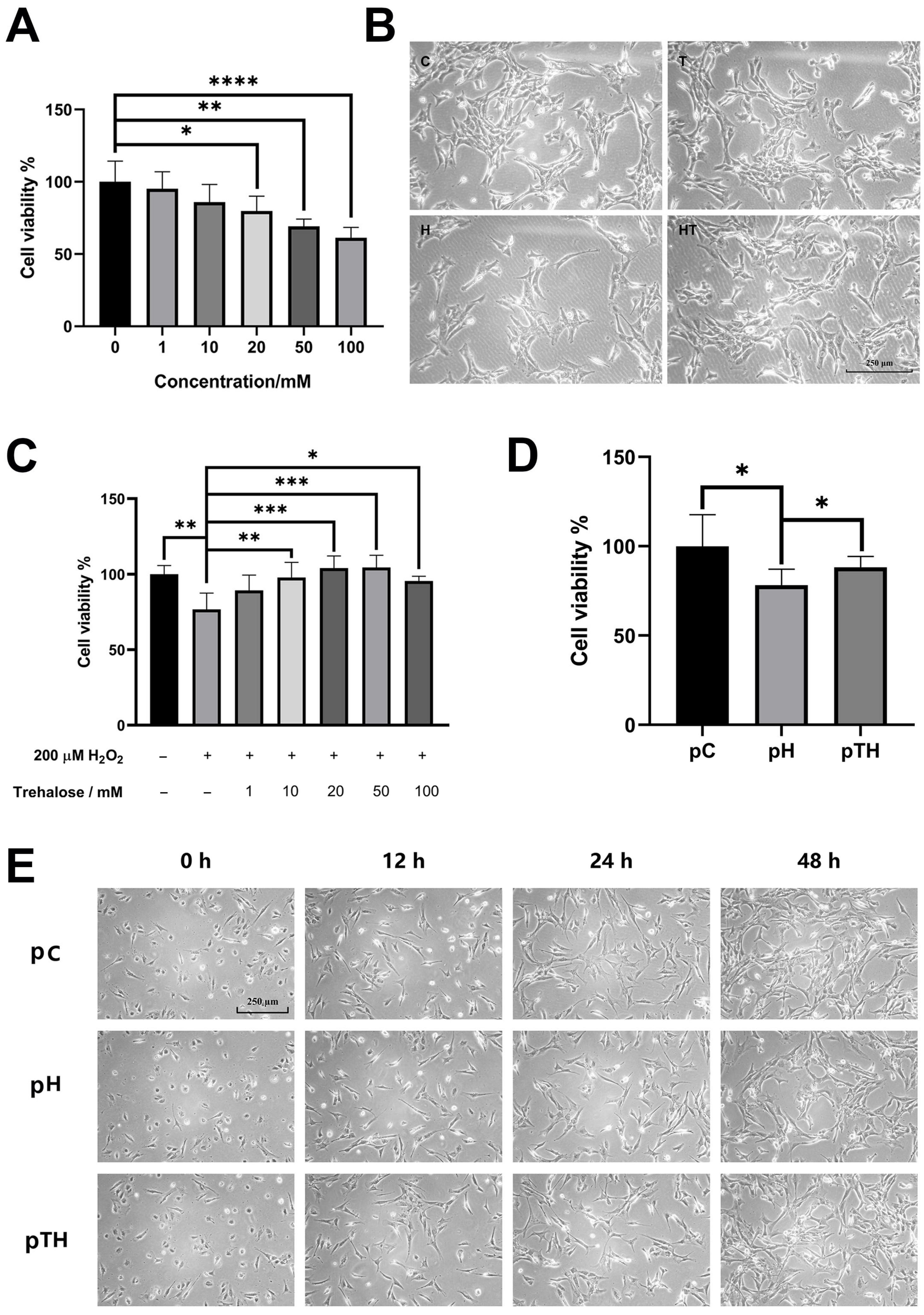
2.1. Trehalose Attenuated H2O2-Induced Decrease in Myoblast Viability
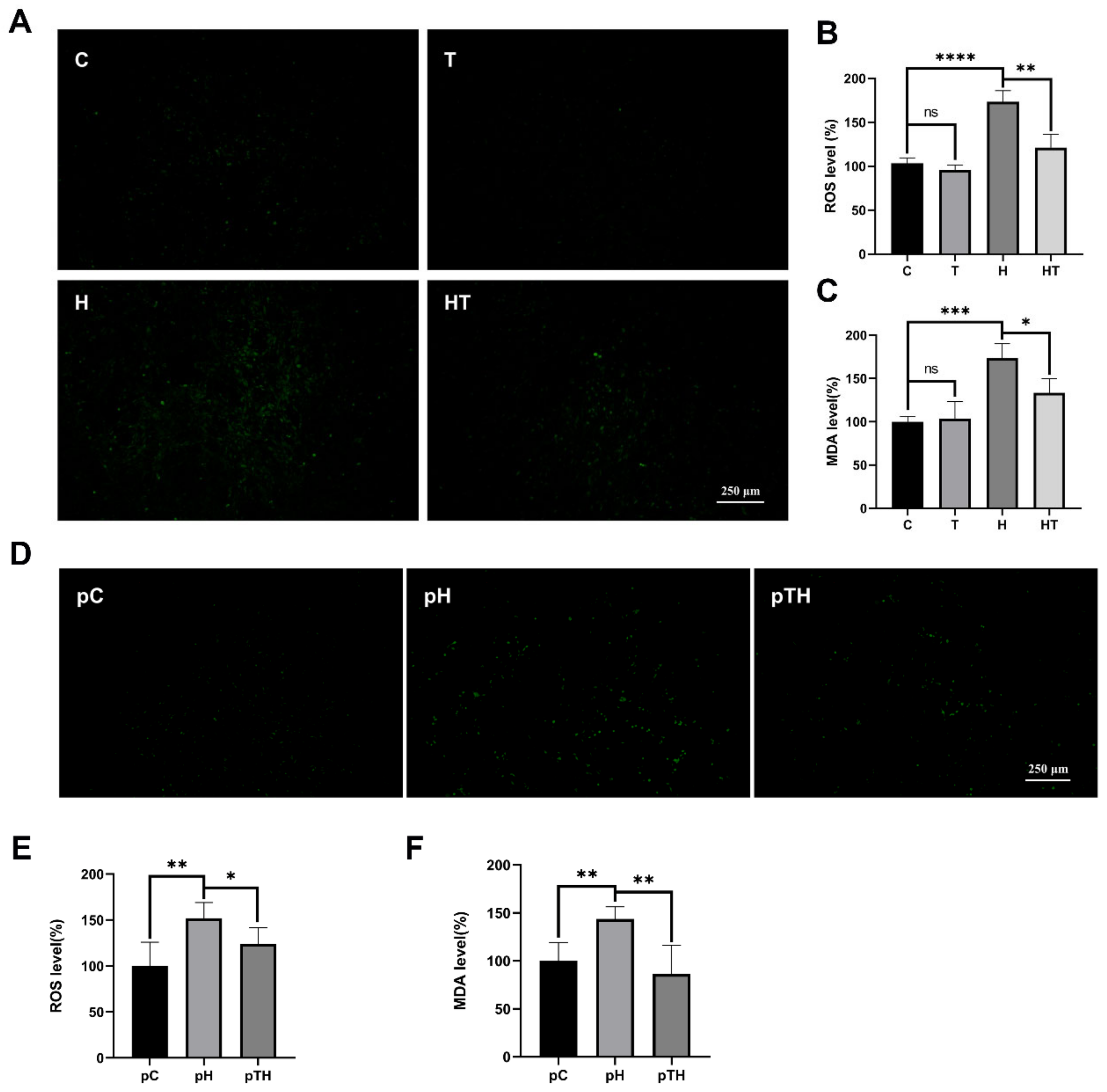
2.2. Trehalose Facilitated ROS Scavenging in Myoblasts
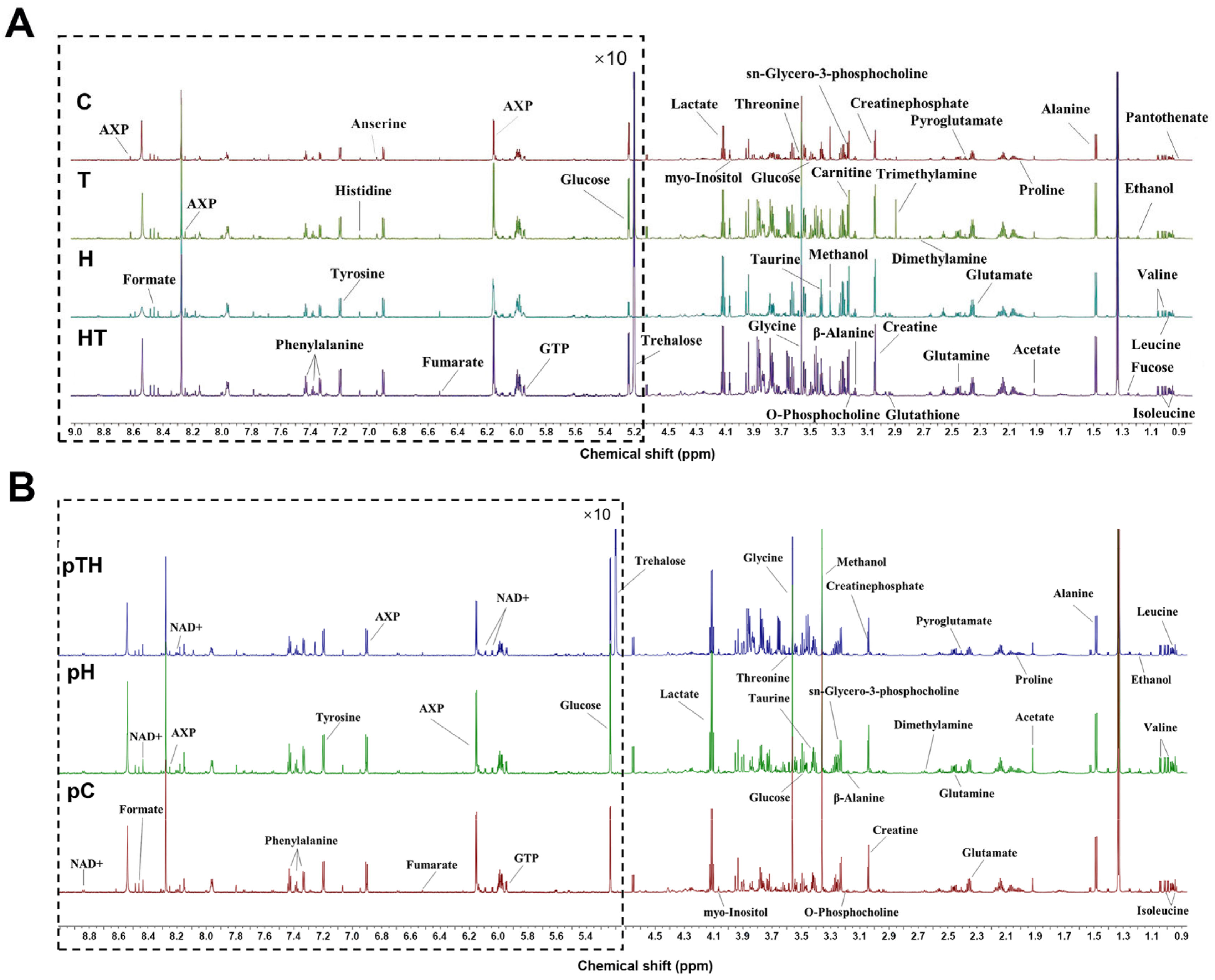
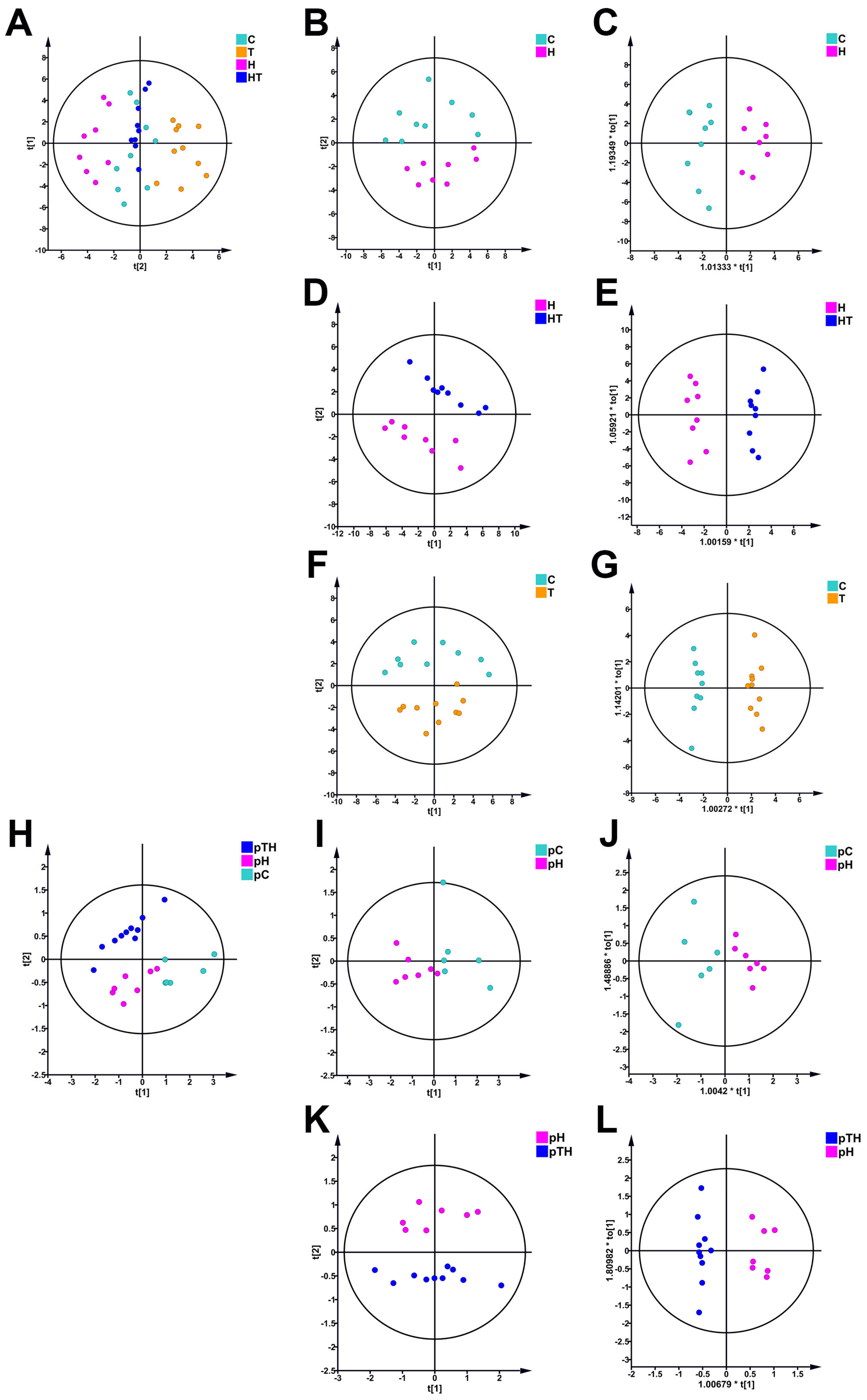
2.3. Trehalose Altered the Metabolic Profile of Myoblasts
2.4. Identification of Significant Metabolites in Myoblasts
2.5. Trehalose Changed Metabolites Levels in Myoblasts
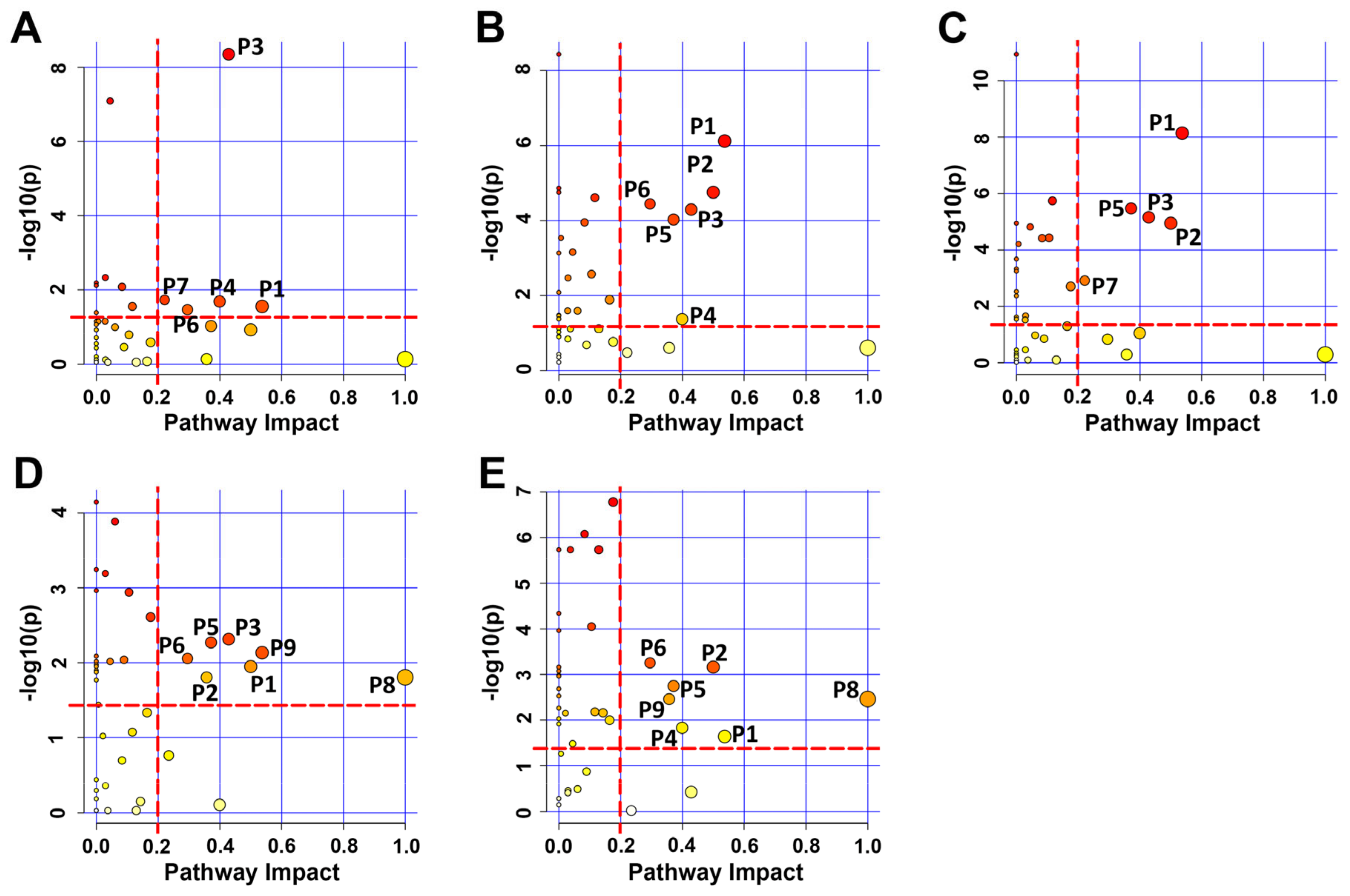
2.6. Trehalose Altered Metabolic Pathways in Myoblasts
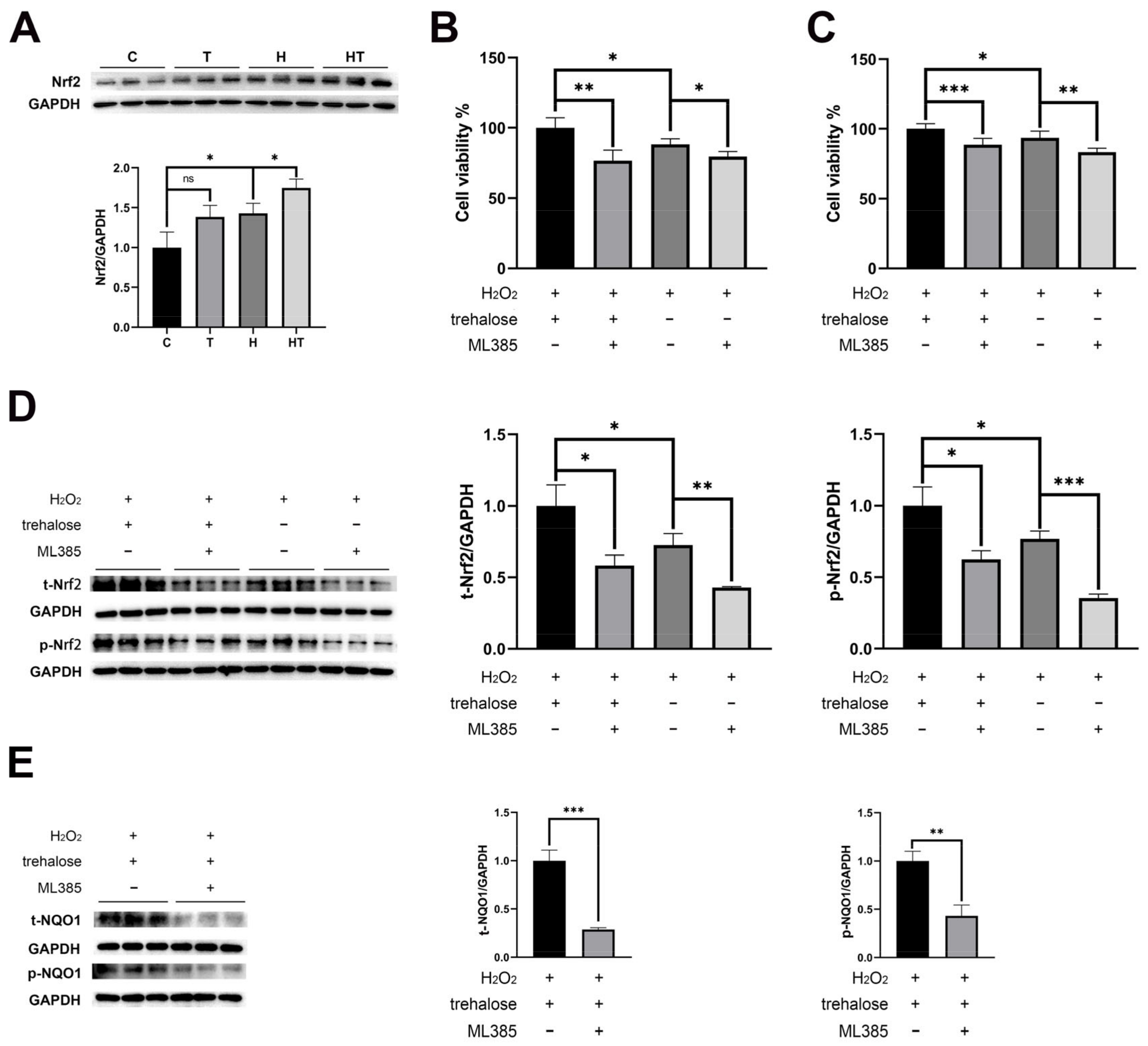
2.7. Trehalose Activated the Nrf2-NQO1 Axis and Promoted the Nuclear Translocation of Nrf2
3. Discussion
3.1. Trehalose Regulates Oxidant Metabolism and Facilitates ROS Scavenging
3.2. Trehalose Regulates Amino Acid Metabolism and the TCA Cycle
3.3. Trehalose Increases the Nuclear Localization of Nrf2 by Increasing the Levels of P62 and Nrf2
4. Materials and Methods
4.1. Reagents
4.2. Cell Culture
4.3. Cell Viability Measurements
4.4. Intracellular ROS Measurements
4.5. Lipid Peroxidation Assessed Using the MDA Assay
4.6. Assay of Cellular T-AOC
4.7. Extraction of Cellular Metabolites
4.8. Acquisition of NMR Spectra
4.9. NMR Data Preprocessing and Resonance Assignments
4.10. Multivariate Analysis and Univariate Analysis of NMR Data
4.11. Metabolic Pathway Analysis
4.12. Inhibition of Nrf2 Activity
4.13. Extraction of Nuclear and Cytoplasmic Proteins
4.14. Measurements of Protein Expressions
4.15. General Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Storz, G.; Imlay, J.A. Oxidative stress. Curr. Opin. Microbiol. 1999, 2, 188–194. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.P. Redefining Oxidative Stress. Antioxid. Redox Signal. 2006, 8, 1865–1879. [Google Scholar] [CrossRef] [PubMed]
- Helmut, S.; Carsten, B.; Jones, D.P. Oxidative Stress. In Annual Review of Biochemistry; Kornberg, R.D., Ed.; Annual Reviews: San Mateo, CA, USA, 2017; pp. 715–748. [Google Scholar]
- Sies, H.; Jones, D.P. Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat. Rev. Mol. Cell Biol. 2020, 21, 363–383. [Google Scholar] [CrossRef] [PubMed]
- Parascandolo, A.; Laukkanen, M.O. Carcinogenesis and Reactive Oxygen Species Signaling: Interaction of the NADPH Oxidase NOX1–5 and Superoxide Dismutase 1–3 Signal Transduction Pathways. Antioxid. Redox Signal. 2019, 30, 443–486. [Google Scholar] [CrossRef] [PubMed]
- Clanton, T.L.; Zuo, L.; Klawitter, P. Oxidants and Skeletal Muscle Function: Physiologic and Pathophysiologic Implications. Proc. Soc. Exp. Boil. Med. 1999, 222, 253–262. [Google Scholar] [CrossRef] [PubMed]
- Daenen, K.; Andries, A.; Mekahli, D.; Van Schepdael, A.; Jouret, F.; Bammens, B. Oxidative stress in chronic kidney disease. Pediatr. Nephrol. 2018, 34, 975–991. [Google Scholar] [CrossRef]
- Di Meo, S.; Napolitano, G.; Venditti, P. Mediators of Physical Activity Protection against ROS-Linked Skeletal Muscle Damage. Int. J. Mol. Sci. 2019, 20, 3024. [Google Scholar] [CrossRef] [PubMed]
- El Assar, M.; Angulo, J.; Rodríguez-Mañas, L. Frailty as a phenotypic manifestation of underlying oxidative stress. Free. Radic. Biol. Med. 2020, 149, 72–77. [Google Scholar] [CrossRef]
- Lian, D.; Chen, M.-M.; Wu, H.; Deng, S.; Hu, X. The Role of Oxidative Stress in Skeletal Muscle Myogenesis and Muscle Disease. Antioxidants 2022, 11, 755. [Google Scholar] [CrossRef]
- Yamamoto, M.; Kensler, T.W.; Motohashi, H. The KEAP1-NRF2 System: A Thiol-Based Sensor-Effector Apparatus for Maintaining Redox Homeostasis. Physiol. Rev. 2018, 98, 1169–1203. [Google Scholar] [CrossRef]
- Suzuki, T.; Muramatsu, A.; Saito, R.; Iso, T.; Shibata, T.; Kuwata, K.; Kawaguchi, S.-I.; Iwawaki, T.; Adachi, S.; Suda, H.; et al. Molecular Mechanism of Cellular Oxidative Stress Sensing by Keap1. Cell Rep. 2019, 28, 746–758. [Google Scholar] [CrossRef]
- Yu, C.; Xiao, J.-H. The Keap1-Nrf2 System: A Mediator between Oxidative Stress and Aging. Oxidative Med. Cell. Longev. 2021, 2021, 6635460. [Google Scholar] [CrossRef] [PubMed]
- Itoh, K.; Wakabayashi, N.; Katoh, Y.; Ishii, T.; Igarashi, K.; Engel, J.D.; Yamamoto, M. Keap1 represses nuclear activation of antioxidant responsive elements by Nrf2 through binding to the amino-terminal Neh2 domain. Genes Dev. 1999, 13, 76–86. [Google Scholar] [CrossRef]
- Magesh, S.; Chen, Y.; Hu, L. Small molecule modulators of Keap1-Nrf2-ARE pathway as potential preventive and therapeutic agents. Med. Res. Rev. 2012, 32, 687–726. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Feng, C.; Jiang, H. Novel target for treating Alzheimer’s Diseases: Crosstalk between the Nrf2 pathway and autophagy. Ageing Res. Rev. 2021, 65, 101207–101218. [Google Scholar] [CrossRef]
- Kalyanaraman, B. Teaching the basics of redox biology to medical and graduate students: Oxidants, antioxidants and disease mechanisms. Redox Biol. 2013, 1, 244–257. [Google Scholar] [CrossRef] [PubMed]
- Elbein, A.D.; Pan, Y.T.; Pastuszak, I.; Carroll, D. New insights on trehalose: A multifunctional molecule. Glycobiology 2003, 13, 17R–27R. [Google Scholar] [CrossRef]
- Richards, A.; Krakowka, S.; Dexter, L.; Schmid, H.; Wolterbeek, A.; Waalkens-Berendsen, D.; Shigoyuki, A.; Kurimoto, M. Trehalose: A review of properties, history of use and human tolerance, and results of multiple safety studies. Food Chem. Toxicol. 2002, 40, 871–898. [Google Scholar] [CrossRef]
- Khalifeh, M.; Barreto, G.E.; Sahebkar, A. Trehalose as a promising therapeutic candidate for the treatment of Parkinson’s disease. Br. Pharmacol. Soc. 2019, 176, 1173–1189. [Google Scholar] [CrossRef]
- Yu, W.; Zha, W.; Peng, H.; Wang, Q.; Zhang, S.; Ren, J. Trehalose Protects against Insulin Resistance-Induced Tissue Injury and Excessive Autophagy in Skeletal Muscles and Kidney. Curr. Pharm. Des. 2019, 25, 2077–2085. [Google Scholar] [CrossRef]
- Ichihara, H.; Kuwabara, K.; Matsumoto, Y. Trehalose liposomes induce apoptosis of breast tumor cells in vitro and in vivo. Biochem. Biophys. Res. Commun. 2020, 532, 505–512. [Google Scholar] [CrossRef] [PubMed]
- Chaitanya, N.S.; Devi, A.; Sahu, S.; Alugoju, P. Molecular mechanisms of action of Trehalose in cancer: A comprehensive review. Life Sci. 2021, 269, 118968. [Google Scholar] [CrossRef]
- Gao, Z.; Wang, H.; Zhang, B.; Wu, X.; Zhang, Y.; Ge, P.; Chi, G.; Liang, J. Trehalose inhibits H2O2-induced autophagic death in dopaminergic SH-SY5Y cells via mitigation of ROS-dependent endoplasmic reticulum stress and AMPK activation. Int. J. Med. Sci. 2018, 15, 1014–1024. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-F.; Kuo, Y.-T.; Chen, T.-Y.; Chien, C.-T. Quercetin-Rich Guava (Psidium guajava) Juice in Combination with Trehalose Reduces Autophagy, Apoptosis and Pyroptosis Formation in the Kidney and Pancreas of Type II Diabetic Rats. Molecules 2016, 21, 334. [Google Scholar] [CrossRef] [PubMed]
- Aoki, N.; Furukawa, S.; Sato, K.; Kurokawa, Y.; Kanda, S.; Takahashi, Y.; Mitsuzumi, H.; Itabashi, H. Supplementation of the diet of dairy cows with trehalose results in milk with low lipid peroxide and high antioxidant content. J. Dairy Sci. 2010, 93, 4189–4195. [Google Scholar] [CrossRef] [PubMed]
- Rani, V.; Deep, G.; Singh, R.K.; Palle, K.; Yadav, U.C.S. Oxidative stress and metabolic disorders: Pathogenesis and therapeutic strategies. Life Sci. 2016, 148, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Alonso, A.; Marsal, S.; Juliã, A. Analytical Methods in Untargeted Metabolomics: State of the Art in 2015. Front. Bioeng. Biotechnol. 2015, 3, 23. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.H.; Ivanisevic, J.; Siuzdak, G. Metabolomics: Beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 2016, 17, 451–459. [Google Scholar] [CrossRef]
- Emwas, A.-H.; Roy, R.; McKay, R.T.; Tenori, L.; Saccenti, E.; Gowda, G.A.N.; Raftery, D.; Alahmari, F.; Jaremko, L.; Jaremko, M.; et al. NMR Spectroscopy for Metabolomics Research. Metabolites 2019, 9, 123. [Google Scholar] [CrossRef]
- Sies, H. Hydrogen peroxide as a central redox signaling molecule in physiological oxidative stress: Oxidative eustress. Redox Biol. 2017, 11, 613–619. [Google Scholar] [CrossRef]
- Mao, C.-Y.; Lu, H.-B.; Kong, N.; Li, J.-Y.; Liu, M.; Yang, C.-Y.; Yang, P. Levocarnitine protects H9c2 rat cardiomyocytes from H2O2-induced mitochondrial dysfunction and apoptosis. Int. J. Med. Sci. 2014, 11, 1107–1115. [Google Scholar] [CrossRef] [PubMed]
- Tarpey, M.M.; Fridovich, I. Methods of Detection of Vascular Reactive Species. Circ. Res. 2001, 89, 224–236. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zhong, Z.; Xu, Z.; Chen, L.; Wang, Y. 2′,7′-Dichlorodihydrofluorescein as a fluorescent probe for reactive oxygen species measurement: Forty years of application and controversy. Free Radic. Res. 2010, 44, 587–604. [Google Scholar] [CrossRef] [PubMed]
- Proctor, D.N.; O’brien, P.C.; Atkinson, E.J.; Nair, K.S.; Clark, R.V.; Walker, A.C.; O’Connor-Semmes, R.L.; Leonard, M.S.; Miller, R.R.; Stimpson, S.A.; et al. Comparison of techniques to estimate total body skeletal muscle mass in people of different age groups. Am. J. Physiol. Metab. 1999, 277, E489–E495. [Google Scholar] [CrossRef] [PubMed]
- Dumont, N.A.; Bentzinger, C.F.; Sincennes, M.C.; Rudnicki, M.A. Satellite Cells and Skeletal Muscle Regeneration. Compr. Physiol. 2015, 5, 1027–1059. [Google Scholar] [PubMed]
- Kjøbsted, R.; Hingst, J.R.; Fentz, J.; Foretz, M.; Sanz, M.N.; Pehmøller, C.; Shum, M.; Marette, A.; Mounier, R.; Treebak, J.T.; et al. AMPK in skeletal muscle function and metabolism. Fed. Am. Soc. Exp. Biol. 2018, 32, 1741–1777. [Google Scholar] [CrossRef] [PubMed]
- Ohtake, S.; Wang, Y.J. Trehalose: Current use and future applications. J. Pharm. Sci. 2011, 100, 2020–2053. [Google Scholar] [CrossRef]
- Benaroudj, N.; Lee, D.H.; Goldberg, A.L. Trehalose accumulation during cellular stress protects cells and cellular proteins from damage by oxygen radicals. J. Biol. Chem. 2001, 276, 24261–24267. [Google Scholar] [CrossRef]
- Luo, Y.; Li, W.-M.; Wang, W. Trehalose: Protector of antioxidant enzymes or reactive oxygen species scavenger under heat stress? Environ. Exp. Bot. 2008, 63, 378–384. [Google Scholar] [CrossRef]
- Honma, Y.; Sato-Morita, M.; Katsuki, Y.; Mihara, H.; Baba, R.; Hino, K.; Kawashima, A.; Ariyasu, T.; Harada, M. Trehalose alleviates oxidative stress-mediated liver injury and Mallory-Denk body formation via activating autophagy in mice. Med. Mol. Morphol. 2020, 54, 41–51. [Google Scholar] [CrossRef]
- Bastin, A.R.; Nazari-Robati, M.; Sadeghi, H.; Doustimotlagh, A.H.; Sadeghi, A. Trehalose and N-Acetyl Cysteine Alleviate Inflammatory Cytokine Production and Oxidative Stress in LPS-Stimulated Human Peripheral Blood Mononuclear Cells. Immunol. Investig. 2021, 51, 963–979. [Google Scholar] [CrossRef] [PubMed]
- Qu, K.-C.; Wang, Z.-Y.; Tang, K.-K.; Zhu, Y.-S.; Fan, R.-F. Trehalose suppresses cadmium-activated Nrf2 signaling pathway to protect against spleen injury. Ecotoxicol. Environ. Saf. 2019, 181, 224–230. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Yu, M.; Tian, Z.; Cui, Y.; Deng, D.; Rong, T.; Liu, Z.; Song, M.; Li, Z.; Ma, X.; et al. Exogenous Glutathione Protects IPEC-J2 Cells against Oxidative Stress through a Mitochondrial Mechanism. Molecules 2022, 27, 2416. [Google Scholar] [CrossRef] [PubMed]
- Jeon, S.-H.; Lee, M.-Y.; Rahman, M.; Kim, S.-J.; Kim, G.-B.; Park, S.-Y.; Hong, C.-U.; Kim, S.-Z.; Kim, J.-S.; Kang, H.-S. The antioxidant, taurine reduced lipopolysaccharide (LPS)-induced generation of ROS, and activation of MAPKs and Bax in cultured pneumocytes. Pulm. Pharmacol. Ther. 2009, 22, 562–566. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Dai, S.; Tao, J.; He, Z.; Liu, Q.; Zhao, J.; Deng, Y.; Kang, J.; Zhang, X.; Yang, S.; et al. Taurine suppresses ROS-dependent autophagy via activating Akt/mTOR signaling pathway in calcium oxalate crystals-induced renal tubular epithelial cell injury. Aging 2021, 13, 3161. [Google Scholar] [CrossRef] [PubMed]
- Liddell, J.R.; Zwingmann, C.; Schmidt, M.M.; Thiessen, A.; Leibfritz, D.; Robinson, S.R.; Dringen, R. Sustained hydrogen peroxide stress decreases lactate production by cultured astrocytes. J. Neurosci. Res. 2009, 87, 2696–2708. [Google Scholar] [CrossRef] [PubMed]
- Storz, P.; Toker, A. NF-kappaB signaling-an alternate pathway for oxidative stress responses. Cell Cycle 2003, 2, 9–10. [Google Scholar] [CrossRef] [PubMed]
- McCubrey, J.A.; LaHair, M.M.; Franklin, R.A.; Choi, S.-I.; Lee, J.S.; Lee, S.; Cho, B.-Y.; Choi, S.-H.; Han, X.; Sim, W.-S.; et al. Reactive Oxygen Species-Induced Activation of the MAP Kinase Signaling Pathways. Antioxid. Redox Signal. 2006, 8, 1775–1789. [Google Scholar] [CrossRef]
- Matsumaru, D.; Motohashi, H. The KEAP1-NRF2 System in Healthy Aging and Longevity. Antioxidants 2021, 10, 1929. [Google Scholar] [CrossRef]
- Kobayashi, M.; Li, L.; Iwamoto, N.; Nakajima-Takagi, Y.; Kaneko, H.; Nakayama, Y.; Eguchi, M.; Wada, Y.; Kumagai, Y.; Yamamoto, M. The antioxidant defense system Keap1-Nrf2 comprises a multiple sensing mechanism for responding to a wide range of chemical compounds. Mol. Cell. Biol. 2009, 29, 493–502. [Google Scholar] [CrossRef]
- Gong, F.; Ge, T.; Liu, J.; Xiao, J.; Wu, X.; Wang, H.; Zhu, Y.; Xia, D.; Hu, B. Trehalose inhibits ferroptosis via NRF2/HO-1 pathway and promotes functional recovery in mice with spinal cord injury. Aging 2022, 14, 3216–3232. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Zhao, Q.; Xiao, Y.; Liu, X.; Li, Y.; Zhang, J.; Pan, J.; Zhang, Z. Trehalose targets Nrf2 signal to alleviate d-galactose induced aging and improve behavioral ability. Biochem. Biophys. Res. Commun. 2020, 521, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, M.; Yasukawa, H.; Arikawa, T.; Deguchi, Y.; Mizushima, N.; Sakurai, M.; Onishi, S.; Tagawa, R.; Sudo, Y.; Okita, N.; et al. Trehalose induces SQSTM1/p62 expression and enhances lysosomal activity and antioxidative capacity in adipocytes. FEBS Open Bio. 2021, 11, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Mizunoe, Y.; Kobayashi, M.; Sudo, Y.; Watanabe, S.; Yasukawa, H.; Natori, D.; Hoshino, A.; Negishi, A.; Okita, N.; Komatsu, M.; et al. Trehalose protects against oxidative stress by regulating the Keap1-Nrf2 and autophagy pathways. Redox Biol. 2017, 15, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.-J.; Yoon, Y.-S.; Lee, S.-J. Mechanism of neuroprotection by trehalose: Controversy surrounding autophagy induction. Cell Death Dis. 2018, 9, 712–723. [Google Scholar] [CrossRef] [PubMed]
- Beckonert, O.; Keun, H.C.; Ebbels, T.M.D.; Bundy, J.; Holmes, E.; Lindon, J.C.; Nicholson, J.K. Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat. Protoc. 2007, 2, 2692–2703. [Google Scholar] [CrossRef] [PubMed]
- Bhinderwala, F.; Roth, H.E.; Noel, H.; Feng, D.; Powers, R. Chemical shift variations in common metabolites. J. Magn. Reson. 2022, 345, 107335–107344. [Google Scholar] [CrossRef] [PubMed]
- Ito, K.; Tsutsumi, Y.; Date, Y.; Kikuchi, J. Fragment Assembly Approach Based on Graph/Network Theory with Quantum Chemistry Verifications for Assigning Multidimensional NMR Signals in Metabolite Mixtures. ACS Chem. Biol. 2016, 11, 1030–1038. [Google Scholar] [CrossRef]
- Ran, H.; Liu, H.; Wu, P. Echinatin mitigates H2O2-induced oxidative damage and apoptosis in lens epithelial cells via the Nrf2/HO-1 pathway. Adv. Clin. Exp. Med. 2021, 30, 1195–1203. [Google Scholar] [CrossRef]
- Dong, W.; Liu, G.; Zhang, K.; Tan, Y.; Zou, H.; Yuan, Y.; Gu, J.; Song, R.; Zhu, J.; Liu, Z. Cadmium exposure induces rat proximal tubular cells injury via p62-dependent Nrf2 nucleus translocation mediated activation of AMPK/AKT/mTOR pathway. Ecotoxicol. Environ. Saf. 2021, 214, 112058. [Google Scholar] [CrossRef]
- Huang, X.; Liu, Y.; Shen, H.; Fu, T.; Guo, Y.; Qiu, S. Chlorogenic acid attenuates inflammation in LPS-induced Human gingival fibroblasts via CysLT1R/Nrf2/NLRP3 signaling. Int. Immunopharmacol. 2022, 107, 108706. [Google Scholar] [CrossRef]


| Metabolite | Mean ± SD | t-Test | |||||
|---|---|---|---|---|---|---|---|
| C | T | H | HT | H vs. C | T vs. C | HT vs. H | |
| Pantothenate | 0.038 ± 0.004 | 0.045 ± 0.005 | 0.039 ± 0.006 | 0.042 ± 0.003 | ns | ** | ns |
| Leucine | 0.487 ± 0.023 | 0.496 ± 0.023 | 0.488 ± 0.021 | 0.488 ± 0.018 | ns | ns | ns |
| Isoleucine | 0.244 ± 0.016 | 0.245 ± 0.010 | 0.247 ± 0.014 | 0.248 ± 0.012 | ns | ns | ns |
| Valine | 0.249 ± 0.017 | 0.245 ± 0.011 | 0.252 ± 0.012 | 0.249 ± 0.012 | ns | ns | ns |
| Fucose | 0.199 ± 0.017 | 0.198 ± 0.015 | 0.213 ± 0.015 | 0.221 ± 0.011 | ns | ns | ns |
| Alanine | 1.411 ± 0.069 | 1.654 ± 0.050 | 1.330 ± 0.046 | 1.563 ± 0.060 | * | **** | **** |
| Acetate | 0.213 ± 0.068 | 0.237 ± 0.032 | 0.196 ± 0.033 | 0.210 ± 0.038 | ns | ns | ns |
| Proline | 0.109 ± 0.004 | 0.102 ± 0.005 | 0.109 ± 0.005 | 0.104 ± 0.005 | ns | ** | ns |
| Glutamate | 0.384 ± 0.024 | 0.414 ± 0.019 | 0.373 ± 0.017 | 0.376 ± 0.022 | ns | ** | ns |
| Pyroglutamate | 0.116 ± 0.009 | 0.077 ± 0.005 | 0.116 ± 0.008 | 0.080 ± 0.004 | ns | **** | **** |
| Glutamine | 0.189 ± 0.014 | 0.221 ± 0.015 | 0.168 ± 0.014 | 0.206 ± 0.008 | ** | **** | **** |
| Dimethylamine | 0.022 ± 0.005 | 0.029 ± 0.022 | 0.009 ± 0.005 | 0.004 ± 0.000 | **** | ns | ns |
| Aspartate | 0.041 ± 0.003 | 0.043 ± 0.004 | 0.032 ± 0.007 | 0.029 ± 0.003 | ** | ns | ns |
| Trimethylamine | 0.170 ± 0.060 | 0.306 ± 0.337 | 0.045 ± 0.053 | 0.014 ± 0.004 | **** | ns | |
| Glutathione | 0.110 ± 0.014 | 0.093 ± 0.010 | 0.125 ± 0.012 | 0.113 ± 0.009 | * | ** | * |
| Lysine | 0.106 ± 0.009 | 0.115 ± 0.008 | 0.113 ± 0.012 | 0.123 ± 0.009 | ns | * | ns |
| Creatine | 1.568 ± 0.139 | 1.621 ± 0.140 | 1.547 ± 0.143 | 1.551 ± 0.073 | ns | ns | ns |
| Phosphocreatine | 1.216 ± 0.173 | 1.133 ± 0.176 | 1.107 ± 0.150 | 0.992 ± 0.099 | ns | ns | ns |
| Beta-Alanine | 0.205 ± 0.024 | 0.198 ± 0.017 | 0.200 ± 0.016 | 0.172 ± 0.012 | ns | ns | ** |
| Choline | 0.083 ± 0.022 | 0.077 ± 0.013 | 0.152 ± 0.042 | 0.076 ± 0.009 | ** | ns | **** |
| PC | 1.976 ± 0.139 | 1.812 ± 0.105 | 1.950 ± 0.189 | 1.642 ± 0.149 | ns | ** | ** |
| GPC | 1.484 ± 0.169 | 1.281 ± 0.066 | 1.268 ± 0.189 | 1.132 ± 0.059 | * | ** | ns |
| Taurine | 0.438 ± 0.023 | 0.398 ± 0.017 | 0.481 ± 0.013 | 0.433 ± 0.024 | **** | *** | **** |
| Glucose | 0.395 ± 0.057 | 0.365 ± 0.052 | 0.276 ± 0.085 | 0.318 ± 0.056 | ** | ns | ns |
| Glycine | 2.996 ± 0.165 | 2.896 ± 0.117 | 2.909 ± 0.215 | 2.746 ± 0.117 | ns | ns | ns |
| Threonine | 0.183 ± 0.023 | 0.196 ± 0.016 | 0.174 ± 0.004 | 0.186 ± 0.004 | ns | ns | **** |
| Myo-inositol | 1.343 ± 0.140 | 1.341 ± 0.078 | 1.287 ± 0.092 | 1.305 ± 0.075 | ns | ns | ns |
| Lactate | 1.447 ± 0.102 | 1.429 ± 0.071 | 1.595 ± 0.093 | 1.602 ± 0.072 | ** | ns | ns |
| GTP | 0.103 ± 0.008 | 0.099 ± 0.007 | 0.097 ± 0.006 | 0.091 ± 0.006 | ns | ns | ns |
| Fumarate | 0.008 ± 0.002 | 0.010 ± 0.001 | 0.008 ± 0.002 | 0.009 ± 0.001 | ns | * | * |
| Histidine | 0.023 ± 0.002 | 0.022 ± 0.002 | 0.025 ± 0.002 | 0.024 ± 0.001 | ns | ns | ns |
| Tyrosine | 0.169 ± 0.010 | 0.162 ± 0.009 | 0.165 ± 0.006 | 0.162 ± 0.005 | ns | ns | ns |
| Phenylalanine | 0.088 ± 0.007 | 0.087 ± 0.004 | 0.086 ± 0.005 | 0.088 ± 0.004 | ns | ns | ns |
| AXP | 0.456 ± 0.031 | 0.437 ± 0.019 | 0.417 ± 0.028 | 0.400 ± 0.028 | * | ns | ns |
| Formate | 0.021 ± 0.003 | 0.024 ± 0.005 | 0.018 ± 0.001 | 0.020 ± 0.002 | * | ns | * |
| Metabolite | Mean ± SD | t-Test | |||
|---|---|---|---|---|---|
| pC | pH | pTH | pH vs. pC | pTH vs. pH | |
| Leucine | 0.338 ± 0.008 | 0.343 ± 0.011 | 0.316 ± 0.011 | ns | **** |
| Isoleucine | 0.326 ± 0.009 | 0.336 ± 0.011 | 0.310 ± 0.011 | ns | **** |
| Valine | 0.325 ± 0.007 | 0.337 ± 0.011 | 0.310 ± 0.009 | * | **** |
| Alanine | 3.369 ± 0.142 | 2.929 ± 0.160 | 2.905 ± 0.231 | **** | ns |
| Acetate | 0.397 ± 0.027 | 0.492 ± 0.032 | 0.456 ± 0.030 | **** | * |
| Proline | 0.181 ± 0.017 | 0.165 ± 0.012 | 0.157 ± 0.012 | ns | ns |
| Glutamate | 2.126 ± 0.094 | 1.819 ± 0.116 | 1.625 ± 0.103 | **** | ** |
| Pyroglutamate | 0.218 ± 0.027 | 0.221 ± 0.013 | 0.210 ± 0.009 | ns | ns |
| Glutamine | 1.395 ± 0.101 | 1.352 ± 0.079 | 1.418 ± 0.086 | ns | ns |
| Dimethylamine | 0.015 ± 0.003 | 0.014 ± 0.003 | 0.011 ± 0.003 | ns | ns |
| Aspartate | 0.021 ± 0.008 | 0.021 ± 0.008 | 0.020 ± 0.007 | ns | ns |
| Glutathione | 0.266 ± 0.042 | 0.185 ± 0.044 | 0.169 ± 0.047 | ** | ns |
| lysine | 0.143 ± 0.021 | 0.170 ± 0.014 | 0.164 ± 0.012 | * | ns |
| Creatine | 1.275 ± 0.139 | 1.027 ± 0.084 | 1.093 ± 0.086 | ** | ns |
| Phosphocreatine | 0.806 ± 0.050 | 0.779 ± 0.069 | 0.855 ± 0.095 | ns | ns |
| Beta-Alanine | 0.030 ± 0.003 | 0.027 ± 0.003 | 0.031 ± 0.003 | ns | * |
| Choline | 0.047 ± 0.010 | 0.044 ± 0.008 | 0.052 ± 0.008 | ns | ns |
| PC | 1.302 ± 0.174 | 1.072 ± 0.094 | 1.271 ± 0.101 | * | *** |
| GPC | 1.023 ± 0.150 | 0.918 ± 0.087 | 1.061 ± 0.106 | ns | ** |
| Taurine | 0.563 ± 0.045 | 0.489 ± 0.029 | 0.481 ± 0.035 | ** | ns |
| Glucose | 0.484 ± 0.122 | 0.607 ± 0.077 | 0.611 ± 0.092 | ns | ns |
| Glycine | 3.758 ± 0.439 | 3.215 ± 0.254 | 2.951 ± 0.247 | * | * |
| Threonine | 0.209 ± 0.007 | 0.190 ± 0.011 | 0.197 ± 0.010 | ** | ns |
| Myo-inositol | 0.409 ± 0.058 | 0.384 ± 0.038 | 0.482 ± 0.034 | ns | **** |
| Lactate | 5.156 ± 0.214 | 5.786 ± 0.164 | 5.812 ± 0.220 | **** | ns |
| GTP | 0.103 ± 0.010 | 0.085 ± 0.011 | 0.079 ± 0.010 | ** | ns |
| AXP | 0.584 ± 0.056 | 0.469 ± 0.055 | 0.429 ± 0.047 | ** | ns |
| Fumarate | 0.005 ± 0.001 | 0.004 ± 0.001 | 0.004 ± 0.001 | ns | ns |
| Tyrosine | 0.268 ± 0.005 | 0.270 ± 0.005 | 0.250 ± 0.004 | ns | **** |
| Phenylalanine | 0.206 ± 0.004 | 0.209 ± 0.007 | 0.217 ± 0.008 | ns | * |
| Tryptophan | 0.017 ± 0.001 | 0.017 ± 0.001 | 0.015 ± 0.001 | ns | ** |
| NAD+ | 0.035 ± 0.005 | 0.029 ± 0.004 | 0.030 ± 0.003 | * | ns |
| Formate | 0.011 ± 0.003 | 0.009 ± 0.001 | 0.010 ± 0.001 | ns | ns |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, S.; Qiu, X.; Zhang, Y.; Huang, C.; Lin, D. Metabolomic Analysis of Trehalose Alleviating Oxidative Stress in Myoblasts. Int. J. Mol. Sci. 2023, 24, 13346. https://doi.org/10.3390/ijms241713346
Zhang S, Qiu X, Zhang Y, Huang C, Lin D. Metabolomic Analysis of Trehalose Alleviating Oxidative Stress in Myoblasts. International Journal of Molecular Sciences. 2023; 24(17):13346. https://doi.org/10.3390/ijms241713346
Chicago/Turabian StyleZhang, Shuya, Xu Qiu, Yue Zhang, Caihua Huang, and Donghai Lin. 2023. "Metabolomic Analysis of Trehalose Alleviating Oxidative Stress in Myoblasts" International Journal of Molecular Sciences 24, no. 17: 13346. https://doi.org/10.3390/ijms241713346
APA StyleZhang, S., Qiu, X., Zhang, Y., Huang, C., & Lin, D. (2023). Metabolomic Analysis of Trehalose Alleviating Oxidative Stress in Myoblasts. International Journal of Molecular Sciences, 24(17), 13346. https://doi.org/10.3390/ijms241713346







