KCTD5 Forms Hetero-Oligomeric Complexes with Various Members of the KCTD Protein Family
Abstract
:1. Introduction
2. Results
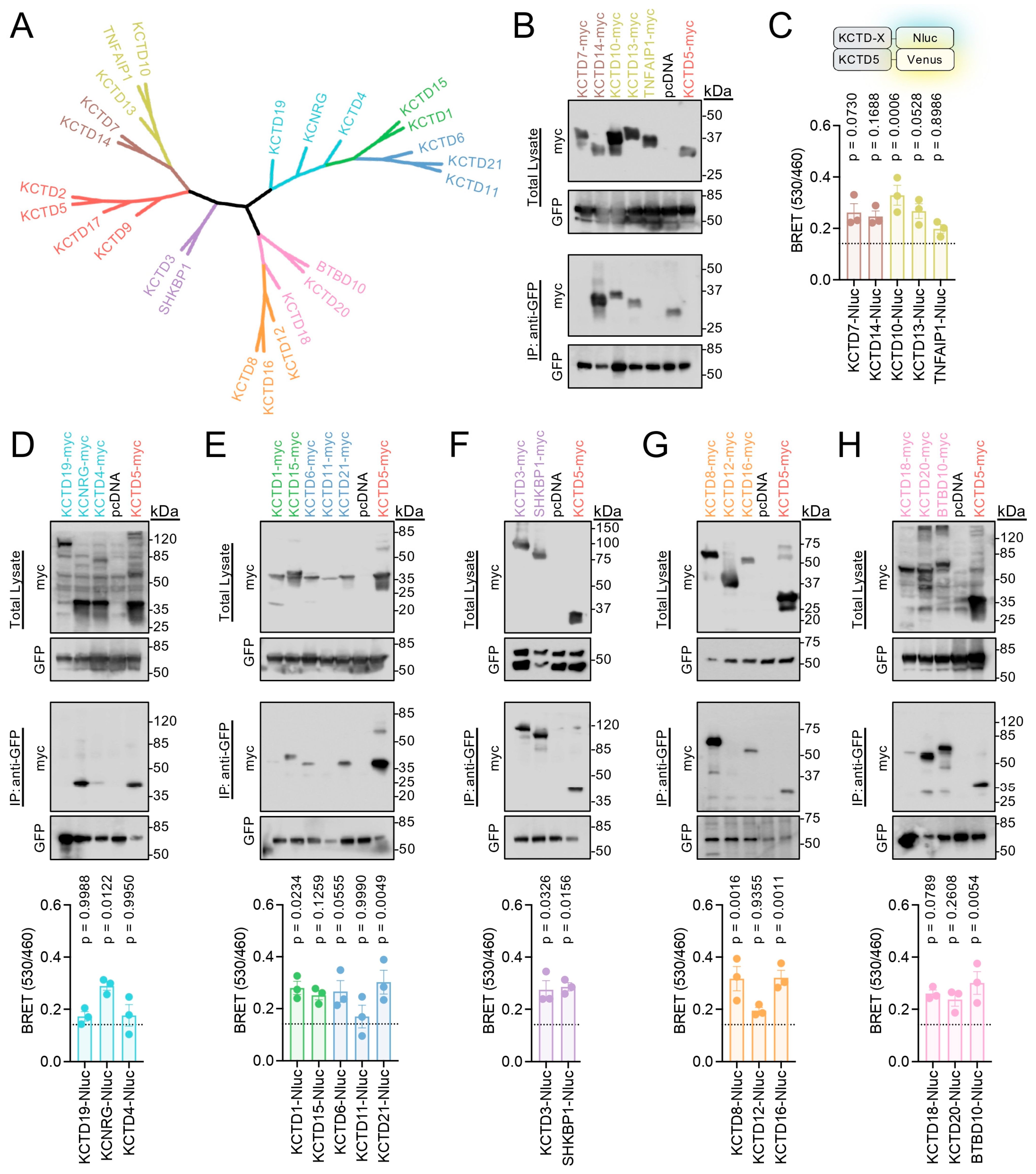
2.1. Assaying KCTD5’s Interaction with KCTD Proteins
2.2. Profiling KCTD5’s Interaction with Each KCTD Family Member
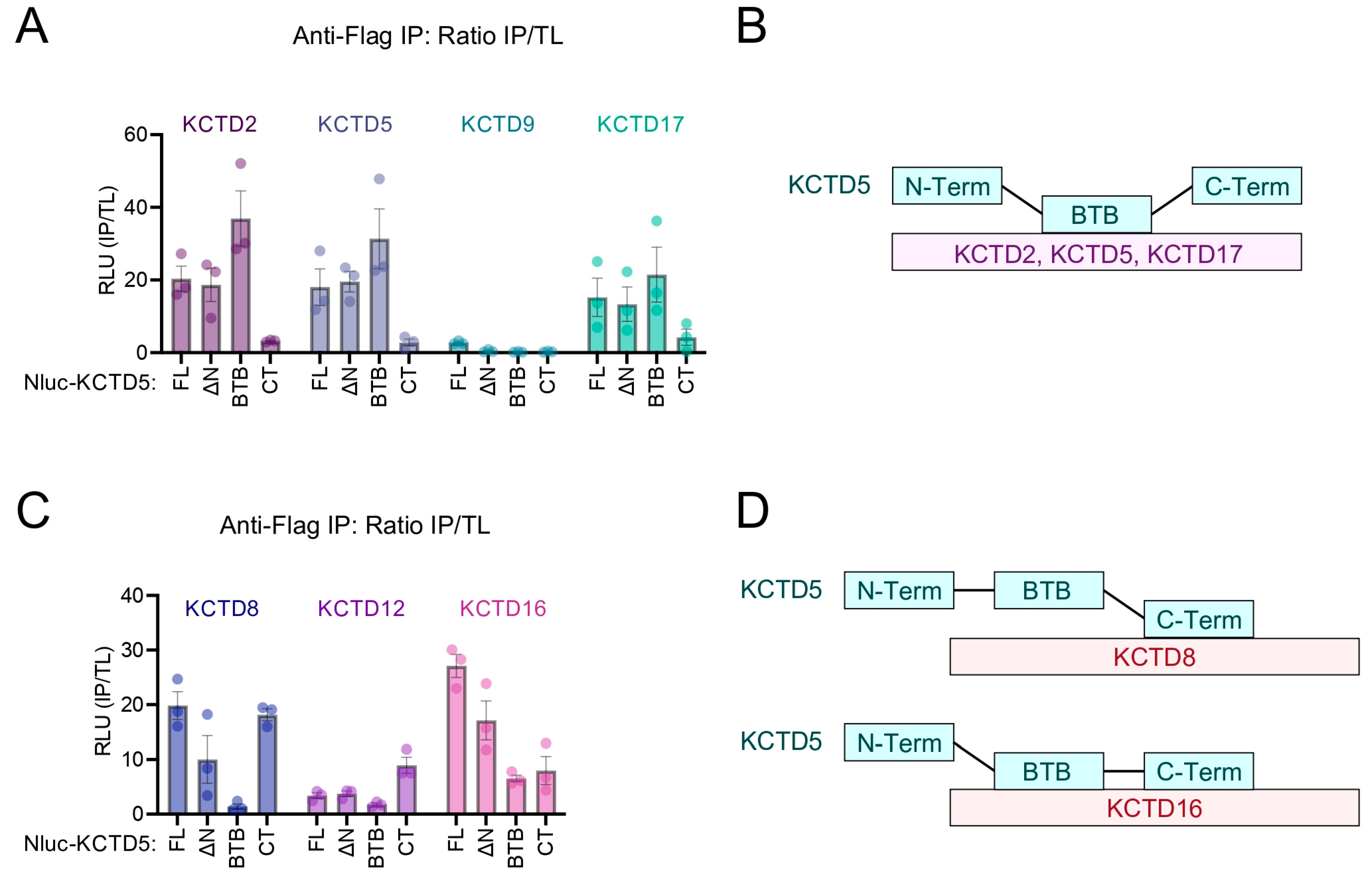
2.3. Determination of Which Region on KCTD5 Facilitates Interaction with Other KCTD Family Proteins
2.4. KCTD5 BTB and CT Domains Differentially Contribute to Interaction with KCTD Protein Family Members
3. Discussion
4. Materials and Methods
4.1. Bioinformatics
4.2. Cell Culture
4.3. Plasmids and Molecular Cloning
4.4. Co-Immunoprecipitation (Co-IP)
4.5. Western Blot
4.6. BRET Assay
4.7. IP-Luminescence Assay
4.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Liu, Z.; Xiang, Y.; Sun, G. The KCTD family of proteins: Structure, function, disease relevance. Cell Biosci. 2013, 3, 45. [Google Scholar] [CrossRef] [PubMed]
- Teng, X.; Aouacheria, A.; Lionnard, L.; Metz, K.A.; Soane, L.; Kamiya, A.; Hardwick, J.M. KCTD: A new gene family involved in neurodevelopmental and neuropsychiatric disorders. CNS Neurosci. Ther. 2019, 25, 887–902. [Google Scholar] [CrossRef] [PubMed]
- Angrisani, A.; Di Fiore, A.; De Smaele, E.; Moretti, M. The emerging role of the KCTD proteins in cancer. Cell Commun. Signal. 2021, 19, 56. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Abreu, N.; Levitz, J.; Kruse, A.C. Structural basis for KCTD-mediated rapid desensitization of GABA(B) signalling. Nature 2019, 567, 127–131. [Google Scholar] [CrossRef] [PubMed]
- Spiombi, E.; Angrisani, A.; Fonte, S.; De Feudis, G.; Fabretti, F.; Cucchi, D.; Izzo, M.; Infante, P.; Miele, E.; Po, A.; et al. KCTD15 inhibits the Hedgehog pathway in Medulloblastoma cells by increasing protein levels of the oncosuppressor KCASH2. Oncogenesis 2019, 8, 64. [Google Scholar] [CrossRef] [PubMed]
- Sloan, D.C.; Cryan, C.E.; Muntean, B.S. Multiple potassium channel tetramerization domain (KCTD) family members interact with Gbetagamma, with effects on cAMP signaling. J. Biol. Chem. 2023, 299, 102924. [Google Scholar] [CrossRef] [PubMed]
- Muntean, B.S.; Marwari, S.; Li, X.; Sloan, D.C.; Young, B.D.; Wohlschlegel, J.A.; Martemyanov, K.A. Members of the KCTD family are major regulators of cAMP signaling. Proc. Natl. Acad. Sci. USA 2022, 119, e2119237119. [Google Scholar] [CrossRef] [PubMed]
- Brockmann, M.; Blomen, V.A.; Nieuwenhuis, J.; Stickel, E.; Raaben, M.; Bleijerveld, O.B.; Altelaar, A.F.M.; Jae, L.T.; Brummelkamp, T.R. Genetic wiring maps of single-cell protein states reveal an off-switch for GPCR signalling. Nature 2017, 546, 307–311. [Google Scholar] [CrossRef]
- Skoblov, M.; Marakhonov, A.; Marakasova, E.; Guskova, A.; Chandhoke, V.; Birerdinc, A.; Baranova, A. Protein partners of KCTD proteins provide insights about their functional roles in cell differentiation and vertebrate development. Bioessays 2013, 35, 586–596. [Google Scholar] [CrossRef]
- Dementieva, I.S.; Tereshko, V.; McCrossan, Z.A.; Solomaha, E.; Araki, D.; Xu, C.; Grigorieff, N.; Goldstein, S.A. Pentameric assembly of potassium channel tetramerization domain-containing protein 5. J. Mol. Biol. 2009, 387, 175–191. [Google Scholar] [CrossRef]
- Esposito, L.; Balasco, N.; Vitagliano, L. Alphafold Predictions Provide Insights into the Structural Features of the Functional Oligomers of All Members of the KCTD Family. Int. J. Mol. Sci. 2022, 23, 13346. [Google Scholar] [CrossRef]
- Smaldone, G.; Pirone, L.; Pedone, E.; Marlovits, T.; Vitagliano, L.; Ciccarelli, L. The BTB domains of the potassium channel tetramerization domain proteins prevalently assume pentameric states. FEBS Lett. 2016, 590, 1663–1671. [Google Scholar] [CrossRef] [PubMed]
- Fritzius, T.; Stawarski, M.; Isogai, S.; Bettler, B. Structural Basis of GABA(B) Receptor Regulation and Signaling. Curr. Top. Behav. Neurosci. 2022, 52, 19–37. [Google Scholar] [CrossRef] [PubMed]
- Fritzius, T.; Turecek, R.; Seddik, R.; Kobayashi, H.; Tiao, J.; Rem, P.D.; Metz, M.; Kralikova, M.; Bouvier, M.; Gassmann, M.; et al. KCTD Hetero-oligomers Confer Unique Kinetic Properties on Hippocampal GABAB Receptor-Induced K+ Currents. J. Neurosci. 2017, 37, 1162–1175. [Google Scholar] [CrossRef] [PubMed]
- Bayon, Y.; Trinidad, A.G.; de la Puerta, M.L.; Del Carmen Rodriguez, M.; Bogetz, J.; Rojas, A.; De Pereda, J.M.; Rahmouni, S.; Williams, S.; Matsuzawa, S.; et al. KCTD5, a putative substrate adaptor for cullin3 ubiquitin ligases. FEBS J. 2008, 275, 3900–3910. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Kellner, D.A.; Hatch, H.A.M.; Yumita, T.; Sanchez, S.; Machold, R.P.; Frank, C.A.; Stavropoulos, N. Conserved properties of Drosophila Insomniac link sleep regulation and synaptic function. PLoS Genet. 2017, 13, e1006815. [Google Scholar] [CrossRef]
- Young, B.D.; Sha, J.; Vashisht, A.A.; Wohlschlegel, J.A. Human Multisubunit E3 Ubiquitin Ligase Required for Heterotrimeric G-Protein beta-Subunit Ubiquitination and Downstream Signaling. J. Proteome Res. 2021, 20, 4318–4330. [Google Scholar] [CrossRef]
- Shi, Y.X.; Yan, J.H.; Liu, W.; Deng, J. Identifies KCTD5 as a novel cancer biomarker associated with programmed cell death and chemotherapy drug sensitivity. BMC Cancer 2023, 23, 408. [Google Scholar] [CrossRef]
- Shi, Y.X.; Zhang, W.D.; Dai, P.H.; Deng, J.; Tan, L.H. Comprehensive analysis of KCTD family genes associated with hypoxic microenvironment and immune infiltration in lung adenocarcinoma. Sci. Rep. 2022, 12, 9938. [Google Scholar] [CrossRef]
- Stogios, P.J.; Downs, G.S.; Jauhal, J.J.; Nandra, S.K.; Prive, G.G. Sequence and structural analysis of BTB domain proteins. Genome Biol. 2005, 6, R82. [Google Scholar] [CrossRef]
- Bardwell, V.J.; Treisman, R. The POZ domain: A conserved protein-protein interaction motif. Genes Dev. 1994, 8, 1664–1677. [Google Scholar] [CrossRef] [PubMed]
- Zollman, S.; Godt, D.; Prive, G.G.; Couderc, J.L.; Laski, F.A. The BTB domain, found primarily in zinc finger proteins, defines an evolutionarily conserved family that includes several developmentally regulated genes in Drosophila. Proc. Natl. Acad. Sci. USA 1994, 91, 10717–10721. [Google Scholar] [CrossRef] [PubMed]
- Bonchuk, A.; Balagurov, K.; Georgiev, P. BTB domains: A structural view of evolution, multimerization, and protein-protein interactions. Bioessays 2023, 45, e2200179. [Google Scholar] [CrossRef] [PubMed]
- Pinkas, D.M.; Sanvitale, C.E.; Bufton, J.C.; Sorrell, F.J.; Solcan, N.; Chalk, R.; Doutch, J.; Bullock, A.N. Structural complexity in the KCTD family of Cullin3-dependent E3 ubiquitin ligases. Biochem. J. 2017, 474, 3747–3761. [Google Scholar] [CrossRef] [PubMed]
- Sereikaite, V.; Fritzius, T.; Kasaragod, V.B.; Bader, N.; Maric, H.M.; Schindelin, H.; Bettler, B.; Stromgaard, K. Targeting the gamma-Aminobutyric Acid Type B (GABA(B)) Receptor Complex: Development of Inhibitors Targeting the K(+) Channel Tetramerization Domain (KCTD) Containing Proteins/GABA(B) Receptor Protein-Protein Interaction. J. Med. Chem. 2019, 62, 8819–8830. [Google Scholar] [CrossRef] [PubMed]
- Zuo, H.; Glaaser, I.; Zhao, Y.; Kurinov, I.; Mosyak, L.; Wang, H.; Liu, J.; Park, J.; Frangaj, A.; Sturchler, E.; et al. Structural basis for auxiliary subunit KCTD16 regulation of the GABA(B) receptor. Proc. Natl. Acad. Sci. USA 2019, 116, 8370–8379. [Google Scholar] [CrossRef] [PubMed]
- Cathomas, F.; Sigrist, H.; Schmid, L.; Seifritz, E.; Gassmann, M.; Bettler, B.; Pryce, C.R. Behavioural endophenotypes in mice lacking the auxiliary GABA(B) receptor subunit KCTD16. Behav. Brain Res. 2017, 317, 393–400. [Google Scholar] [CrossRef] [PubMed]
- Schwenk, J.; Metz, M.; Zolles, G.; Turecek, R.; Fritzius, T.; Bildl, W.; Tarusawa, E.; Kulik, A.; Unger, A.; Ivankova, K.; et al. Native GABA(B) receptors are heteromultimers with a family of auxiliary subunits. Nature 2010, 465, 231–235. [Google Scholar] [CrossRef]
- Turecek, R.; Schwenk, J.; Fritzius, T.; Ivankova, K.; Zolles, G.; Adelfinger, L.; Jacquier, V.; Besseyrias, V.; Gassmann, M.; Schulte, U.; et al. Auxiliary GABAB receptor subunits uncouple G protein betagamma subunits from effector channels to induce desensitization. Neuron 2014, 82, 1032–1044. [Google Scholar] [CrossRef]
- Kasahara, K.; Kawakami, Y.; Kiyono, T.; Yonemura, S.; Kawamura, Y.; Era, S.; Matsuzaki, F.; Goshima, N.; Inagaki, M. Ubiquitin-proteasome system controls ciliogenesis at the initial step of axoneme extension. Nat. Commun. 2014, 5, 5081. [Google Scholar] [CrossRef]
- Esposito, L.; Balasco, N.; Smaldone, G.; Berisio, R.; Ruggiero, A.; Vitagliano, L. AlphaFold-Predicted Structures of KCTD Proteins Unravel Previously Undetected Relationships among the Members of the Family. Biomolecules 2021, 11, 1862. [Google Scholar] [CrossRef] [PubMed]
- Balasco, N.; Pirone, L.; Smaldone, G.; Di Gaetano, S.; Esposito, L.; Pedone, E.M.; Vitagliano, L. Molecular recognition of Cullin3 by KCTDs: Insights from experimental and computational investigations. Biochim. Biophys. Acta 2014, 1844, 1289–1298. [Google Scholar] [CrossRef] [PubMed]
- Correale, S.; Pirone, L.; Di Marcotullio, L.; De Smaele, E.; Greco, A.; Mazza, D.; Moretti, M.; Alterio, V.; Vitagliano, L.; Di Gaetano, S.; et al. Molecular organization of the cullin E3 ligase adaptor KCTD11. Biochimie 2011, 93, 715–724. [Google Scholar] [CrossRef] [PubMed]
- Ji, A.X.; Chu, A.; Nielsen, T.K.; Benlekbir, S.; Rubinstein, J.L.; Prive, G.G. Structural Insights into KCTD Protein Assembly and Cullin3 Recognition. J. Mol. Biol. 2016, 428, 92–107. [Google Scholar] [CrossRef] [PubMed]
- Smaldone, G.; Pirone, L.; Balasco, N.; Di Gaetano, S.; Pedone, E.M.; Vitagliano, L. Cullin 3 Recognition Is Not a Universal Property among KCTD Proteins. PLoS ONE 2015, 10, e0126808. [Google Scholar] [CrossRef] [PubMed]
- Balasco, N.; Smaldone, G.; Vitagliano, L. The Structural Versatility of the BTB Domains of KCTD Proteins and Their Recognition of the GABA(B) Receptor. Biomolecules 2019, 9, 323. [Google Scholar] [CrossRef]
- Muntean, B.S.; Martemyanov, K.A. Association with the Plasma Membrane Is Sufficient for Potentiating Catalytic Activity of Regulators of G Protein Signaling (RGS) Proteins of the R7 Subfamily. J. Biol. Chem. 2016, 291, 7195–7204. [Google Scholar] [CrossRef] [PubMed]
- Coutant, E.P.; Gagnot, G.; Hervin, V.; Baatallah, R.; Goyard, S.; Jacob, Y.; Rose, T.; Janin, Y.L. Bioluminescence Profiling of NanoKAZ/NanoLuc Luciferase Using a Chemical Library of Coelenterazine Analogues. Chemistry 2020, 26, 948–958. [Google Scholar] [CrossRef]
- Coutant, E.P.; Goyard, S.; Hervin, V.; Gagnot, G.; Baatallah, R.; Jacob, Y.; Rose, T.; Janin, Y.L. Gram-scale synthesis of luciferins derived from coelenterazine and original insights into their bioluminescence properties. Org. Biomol. Chem. 2019, 17, 3709–3713. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liao, Y.; Sloan, D.C.; Widjaja, J.H.; Muntean, B.S. KCTD5 Forms Hetero-Oligomeric Complexes with Various Members of the KCTD Protein Family. Int. J. Mol. Sci. 2023, 24, 14317. https://doi.org/10.3390/ijms241814317
Liao Y, Sloan DC, Widjaja JH, Muntean BS. KCTD5 Forms Hetero-Oligomeric Complexes with Various Members of the KCTD Protein Family. International Journal of Molecular Sciences. 2023; 24(18):14317. https://doi.org/10.3390/ijms241814317
Chicago/Turabian StyleLiao, Yini, Douglas C. Sloan, Josephine H. Widjaja, and Brian S. Muntean. 2023. "KCTD5 Forms Hetero-Oligomeric Complexes with Various Members of the KCTD Protein Family" International Journal of Molecular Sciences 24, no. 18: 14317. https://doi.org/10.3390/ijms241814317



