Insight into the Global Negative Regulation of Iron Scavenger 7-HT Biosynthesis by the SigW/RsiW System in Pseudomonas donghuensis HYS
Abstract
:1. Introduction
2. Results
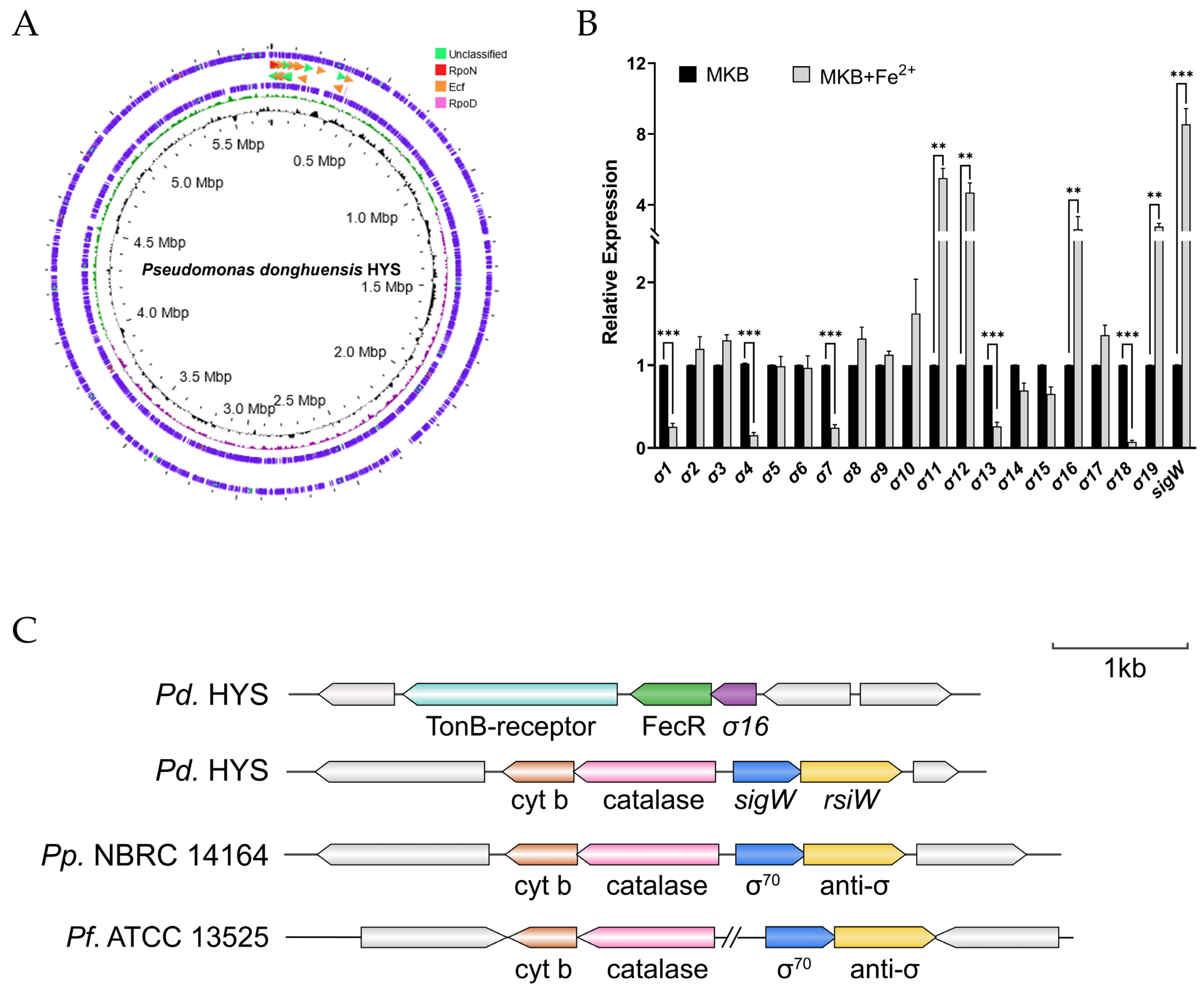
2.1. Response of ECF σ Factors to Extracellular Ferrous Iron in P. donghuensis HYS
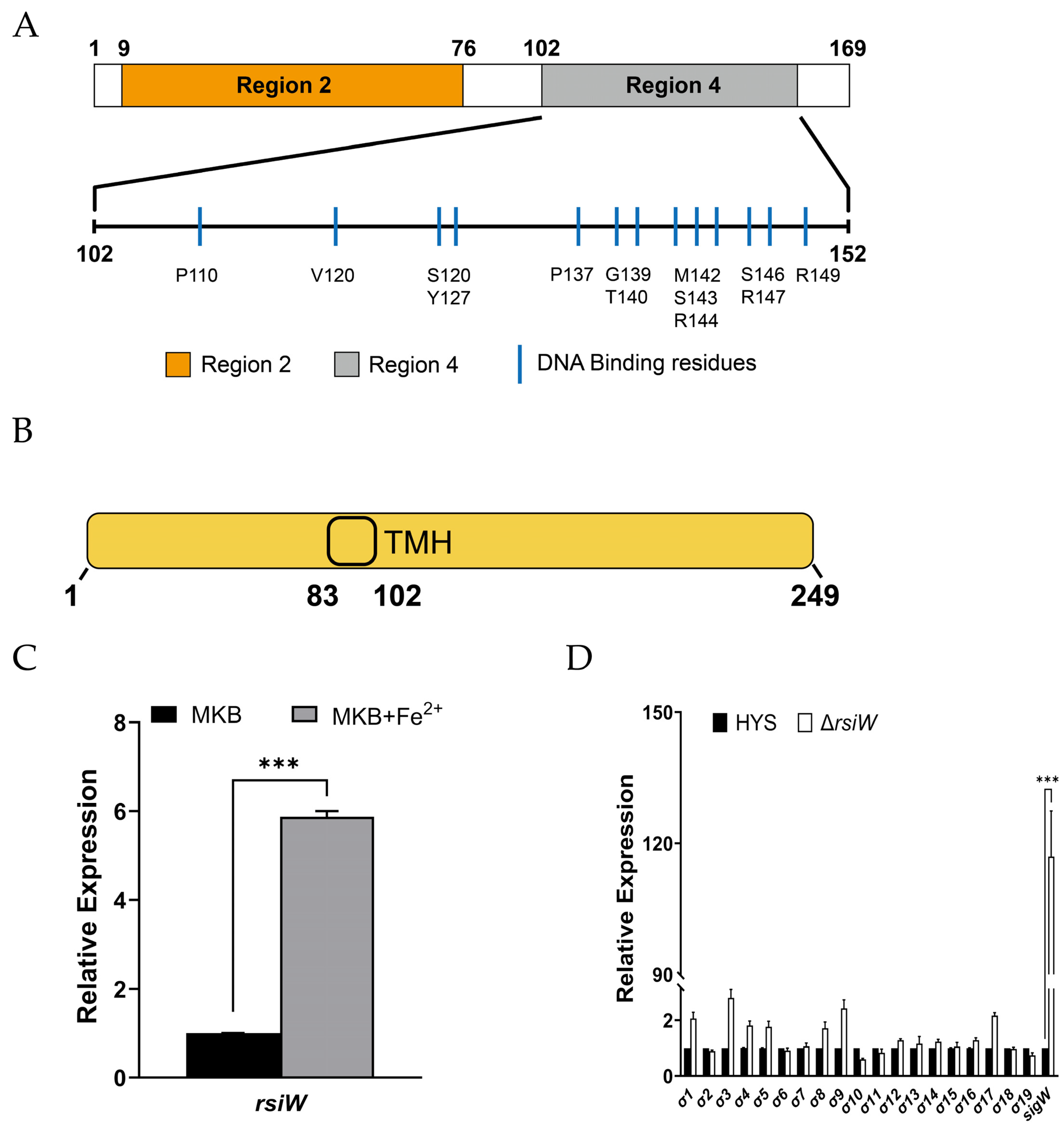
2.2. SigW Is Only Regulated by RsiW under Iron-Limiting Conditions in P. donghuensis HYS
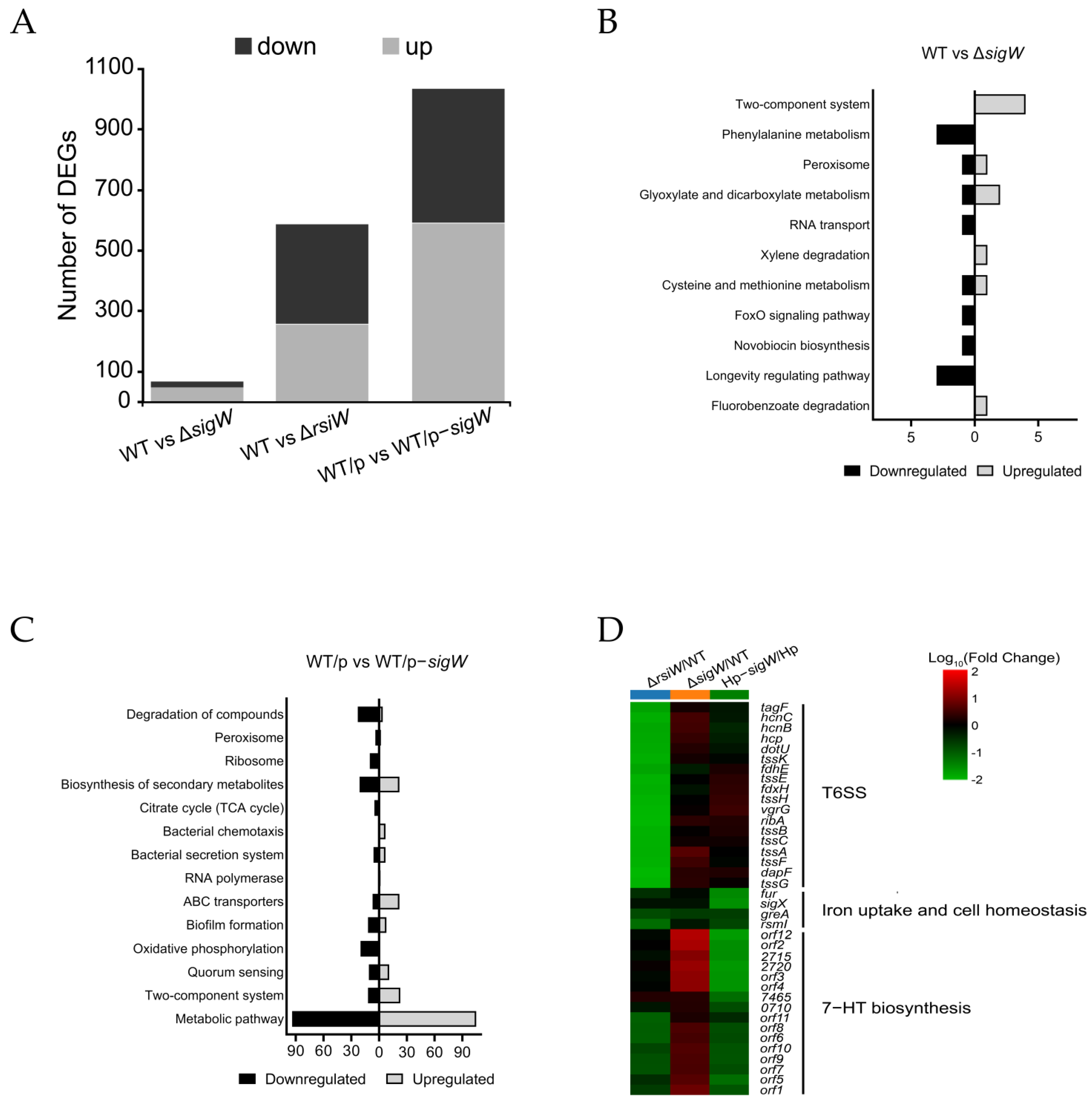
2.3. The SigW/RsiW System Is Involved in Iron Metabolism and 7-HT Biosynthesis
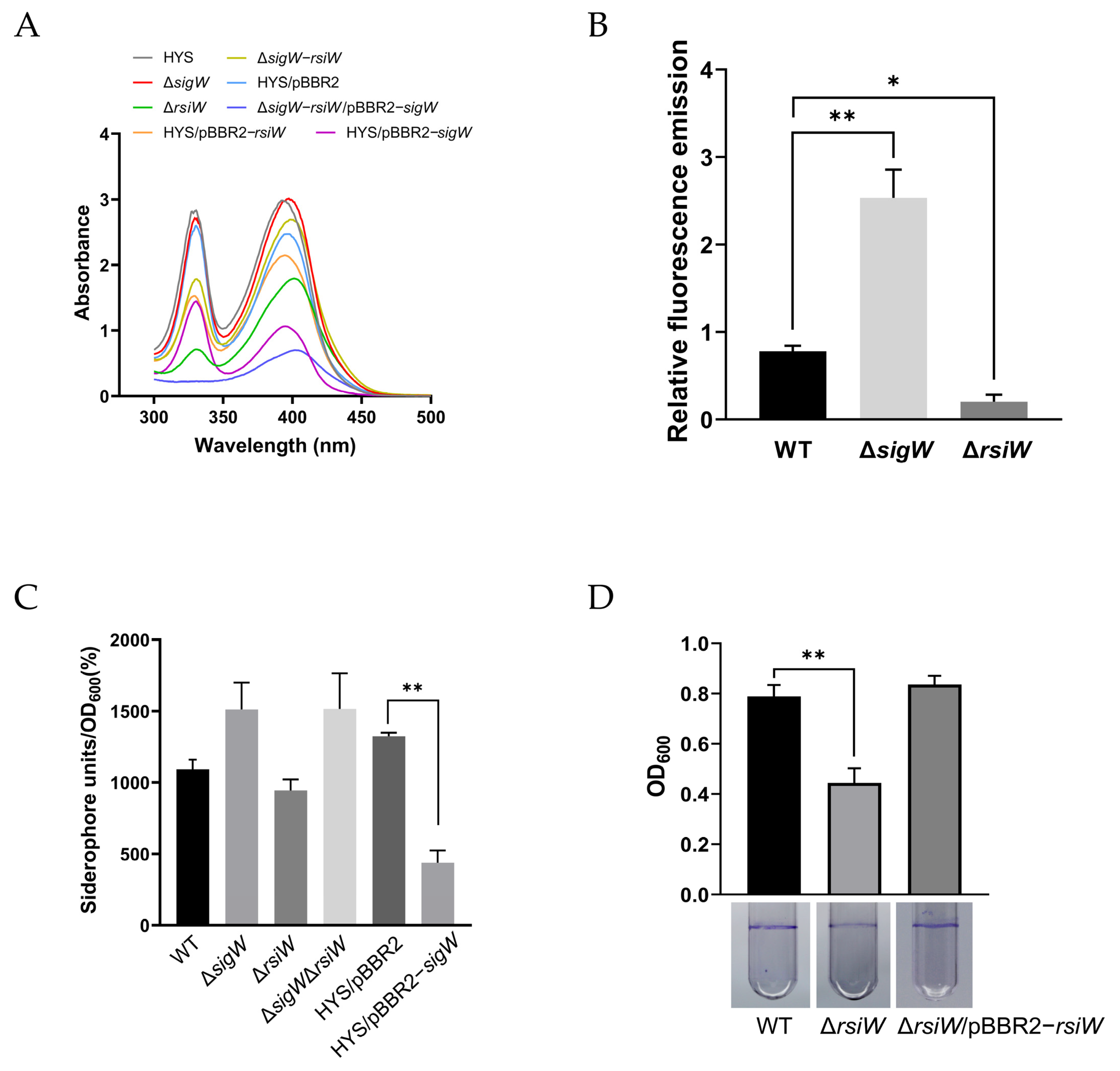
2.4. SigW Inhibits the Production of Two Siderophores: Pyoverdine and 7-HT
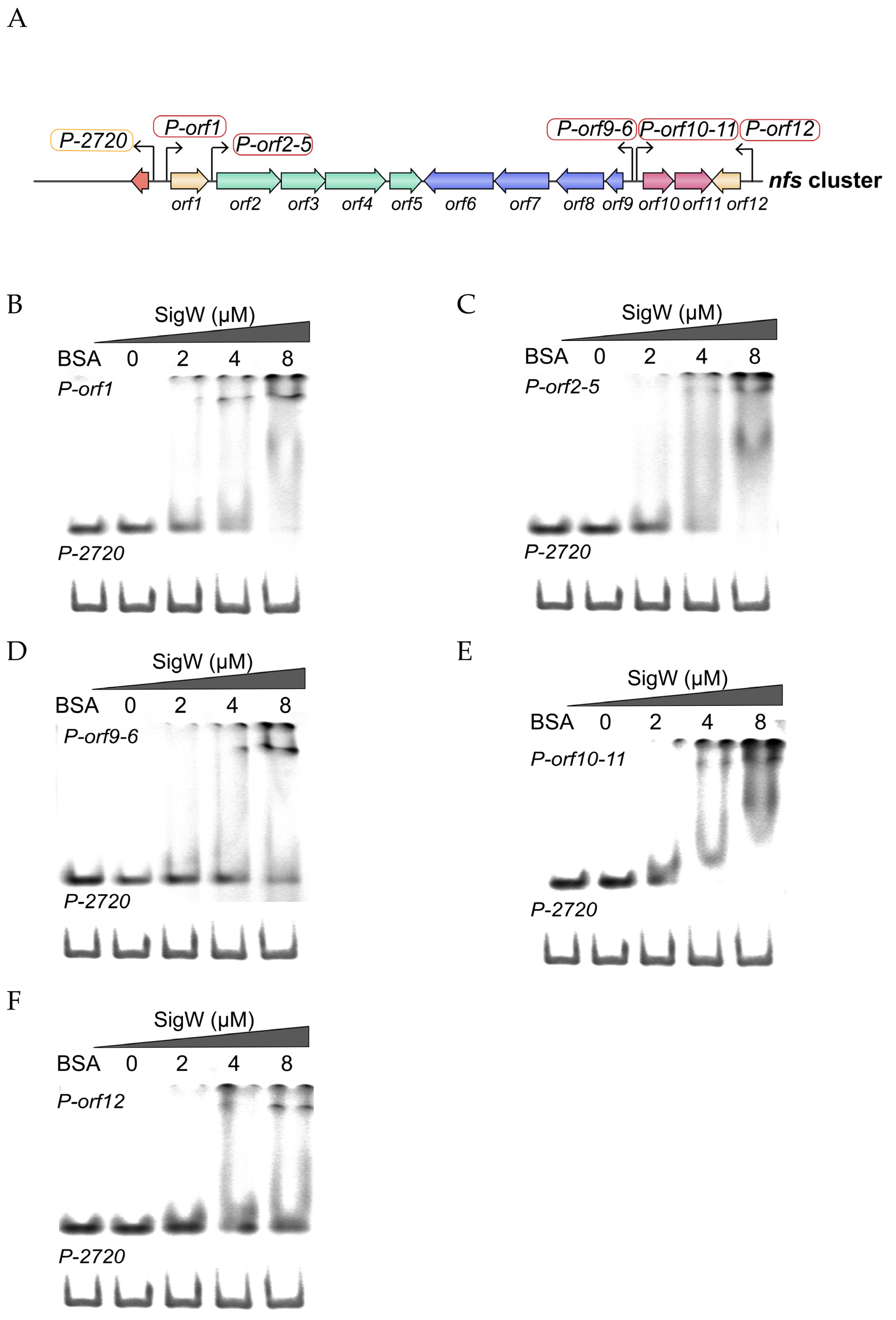
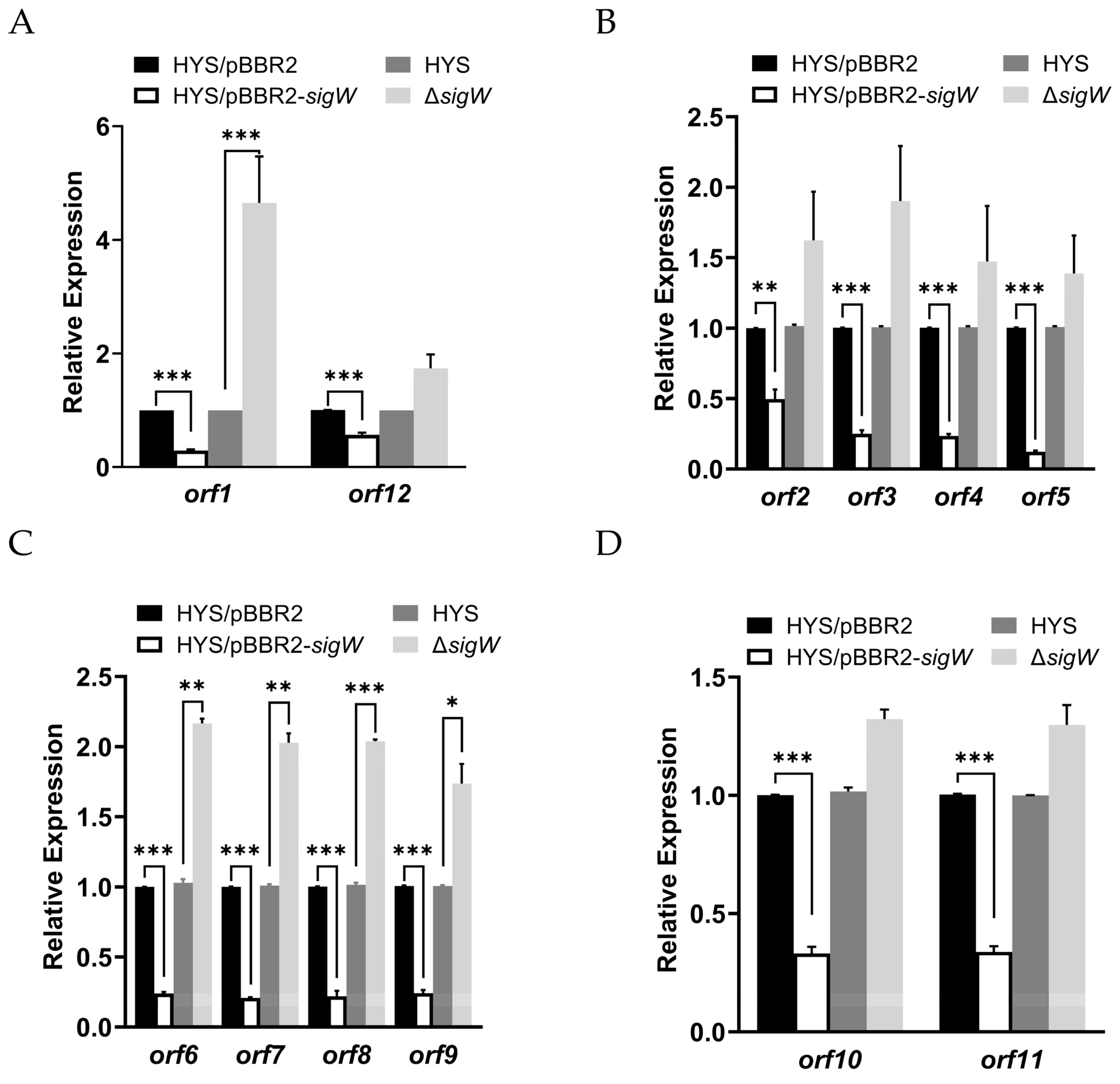
2.5. SigW Directly Regulates the Expression of the nfs Cluster
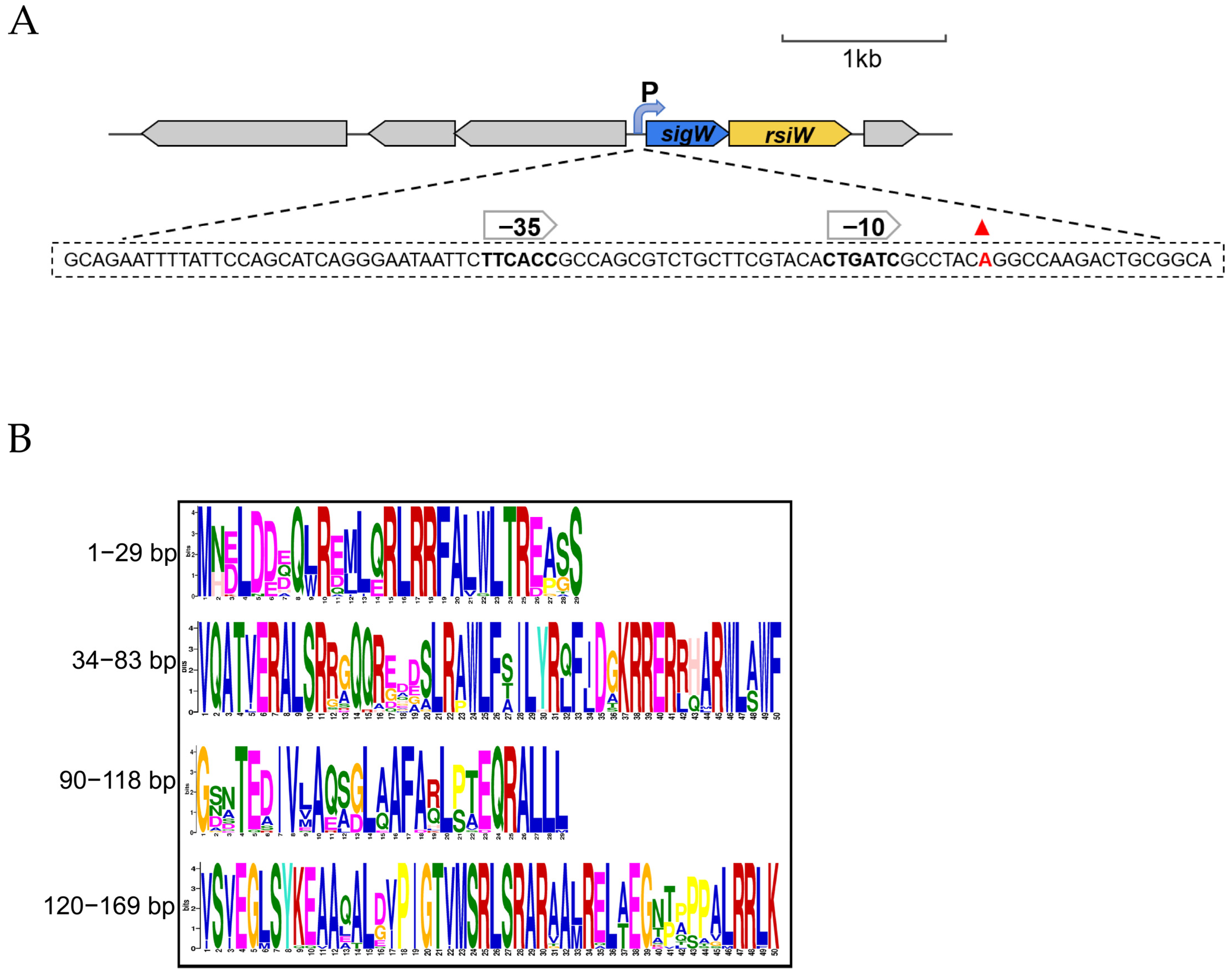
2.6. Transcription Start Sites for the SigW Operon
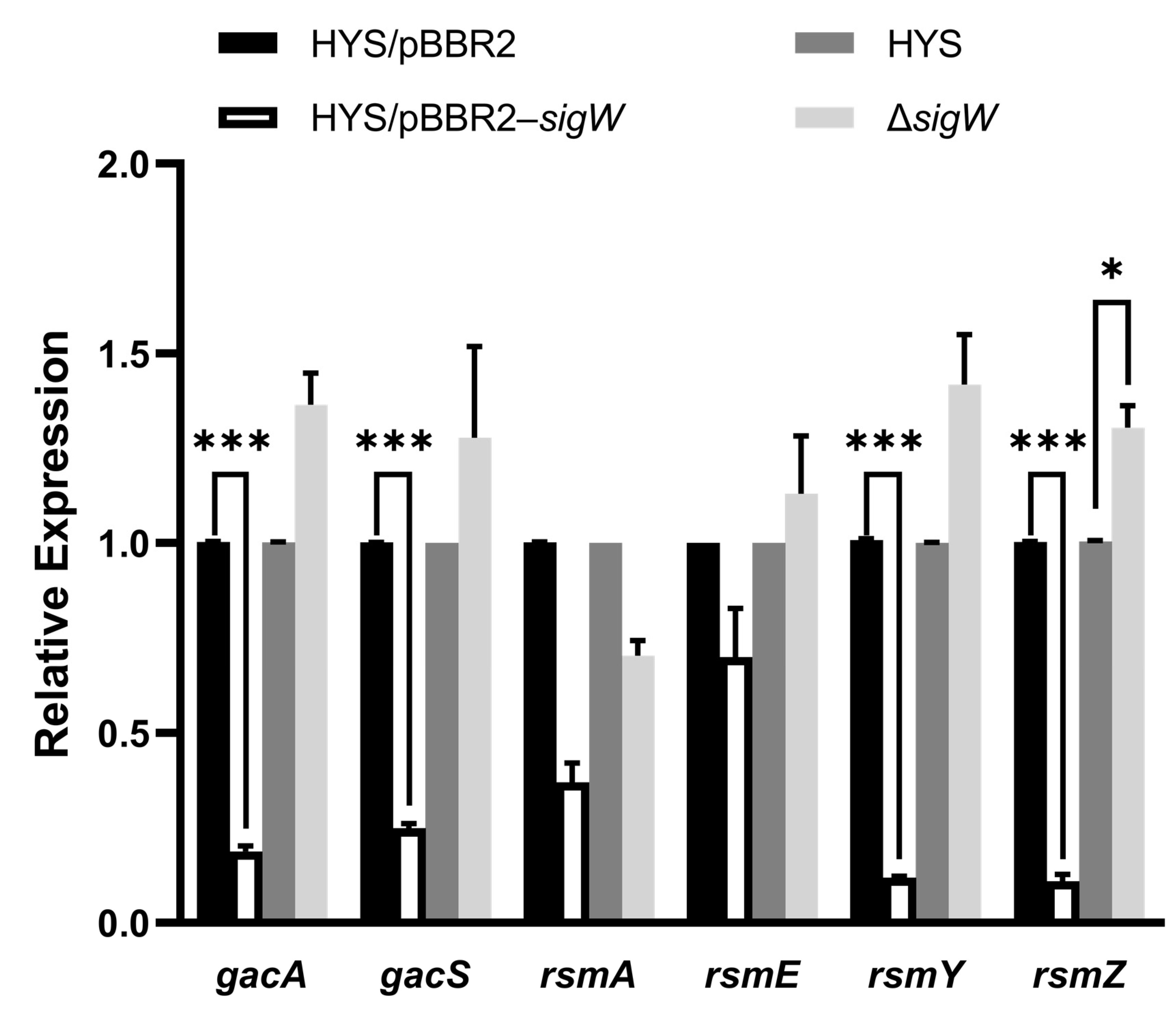
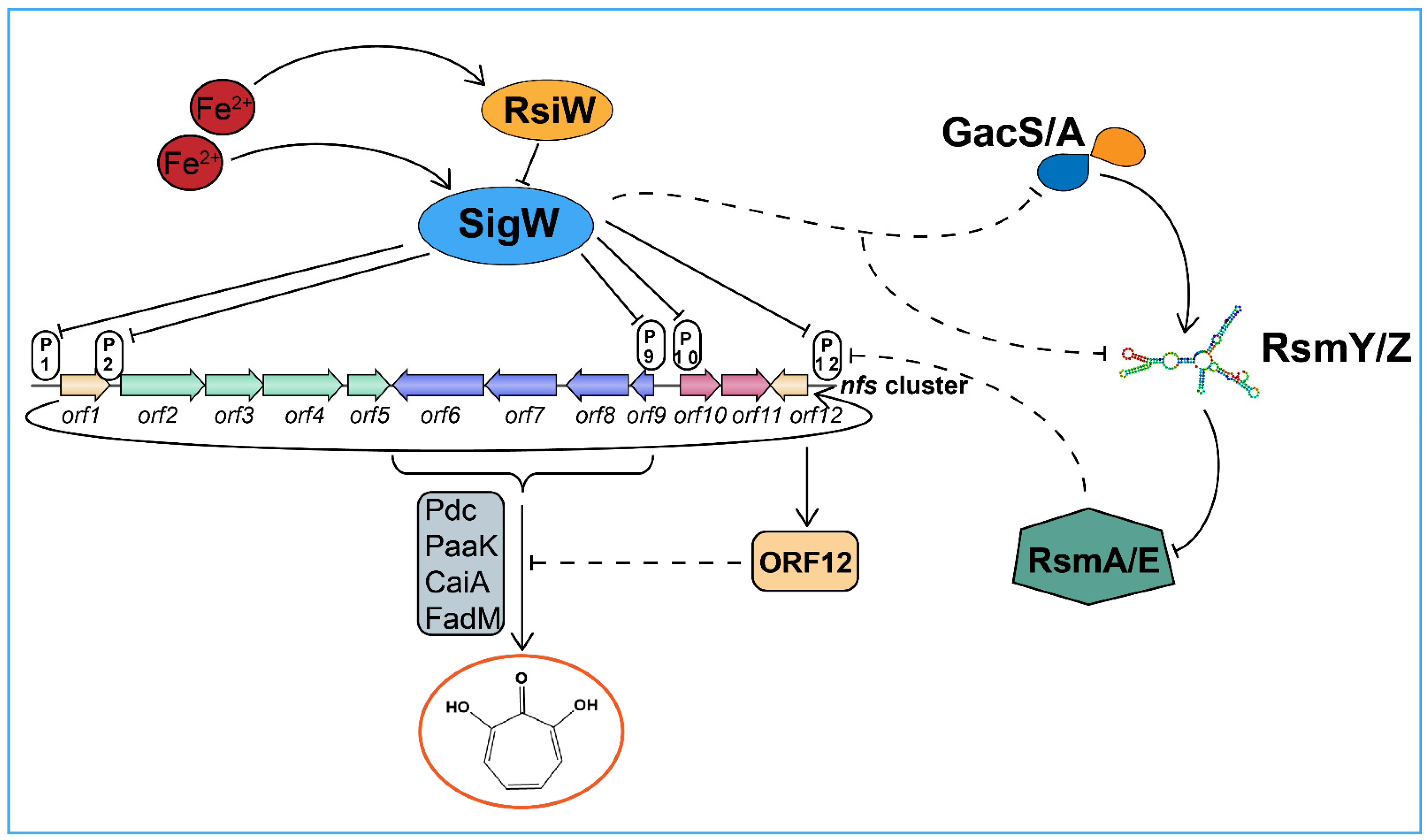
2.7. SigW Negatively Regulates the Gac/Rsm System to Inhibit 7-HT and Enhance Pyoverdine Production
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Culture Conditions
4.2. Construction of Deletion Mutants
4.3. Siderophore Determination Assays
4.4. Biofilm Formation Analysis
4.5. RNA Isolation and RT-PCR
4.6. Protein Expression and Purification
4.7. Promoter Activity Assay
4.8. Electrophoretic Mobility Shift Assay
4.9. 5′RACE PCR Analysis to Identify Transcription Start Sites
4.10. Bioinformatic Analysis
4.11. Accession Numbers
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Van Dingenen, J. Harder, better, faster, stronger: Iron strengthens pathogenic bacteria too! Plant Cell 2021, 33, 1853–1854. [Google Scholar] [CrossRef] [PubMed]
- Pang, Z.; Raudonis, R.; Glick, B.R.; Lin, T.J.; Cheng, Z. Antibiotic resistance in Pseudomonas aeruginosa: Mechanisms and alternative therapeutic strategies. Biotechnol. Adv. 2019, 37, 177–192. [Google Scholar] [CrossRef] [PubMed]
- Azam, M.W.; Khan, A.U. Updates on the pathogenicity status of Pseudomonas aeruginosa. Drug Discov. Today 2019, 24, 350–359. [Google Scholar] [CrossRef]
- Kang, D.; Kirienko, N.V. Interdependence between iron acquisition and biofilm formation in Pseudomonas aeruginosa. J. Microbiol. 2018, 56, 449–457. [Google Scholar] [CrossRef]
- Leinweber, A.; Weigert, M.; Kummerli, R. The bacterium Pseudomonas aeruginosa senses and gradually responds to interspecific competition for iron. Evolution 2018, 72, 1515–1528. [Google Scholar] [CrossRef]
- Bashir, A.; Tian, T.; Yu, X.; Meng, C.; Ali, M.; Li, L. Pyoverdine-Mediated Killing of Caenorhabditis elegans by Pseudomonas syringae MB03 and the Role of Iron in Its Pathogenicity. Int. J. Mol. Sci. 2020, 21, 2198–2214. [Google Scholar] [CrossRef] [Green Version]
- Soares, E.V. Perspective on the biotechnological production of bacterial siderophores and their use. Appl. Microbiol. Biotechnol. 2022, 106, 3985–4004. [Google Scholar] [CrossRef]
- Kramer, J.; Ozkaya, O.; Kummerli, R. Bacterial siderophores in community and host interactions. Nat. Rev. Microbiol. 2020, 18, 152–163. [Google Scholar] [CrossRef]
- De Serrano, L.O.; Camper, A.K.; Richards, A.M. An overview of siderophores for iron acquisition in microorganisms living in the extreme. Biometals 2016, 29, 551–571. [Google Scholar] [CrossRef] [Green Version]
- Cornelis, P. Iron uptake and metabolism in Pseudomonads. Appl. Microbiol. Biotechnol. 2010, 86, 1637–1645. [Google Scholar] [CrossRef]
- Taguchi, F.; Suzuki, T.; Inagaki, Y.; Toyoda, K.; Shiraishi, T.; Ichinose, Y. The siderophore pyoverdine of Pseudomonas syringae pv. tabaci 6605 is an intrinsic virulence factor in host tobacco infection. J. Bacteriol. 2010, 192, 117–126. [Google Scholar] [CrossRef] [Green Version]
- Sass, G.; Nazik, H.; Penner, J.; Shah, H.; Ansari, S.R.; Clemons, K.V.; Groleau, M.C.; Dietl, A.M.; Visca, P.; Haas, H.; et al. Studies of Pseudomonas aeruginosa Mutants Indicate Pyoverdine as the Central Factor in Inhibition of Aspergillus fumigatus Biofilm. J. Bacteriol. 2018, 200, e00345-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hazotte, A.; Péron, O.; Gaudin, P.; Abdelouas, A.; Lebeau, T. Effect of Pseudomonas fluorescens and pyoverdine on the phytoextraction of cesium by red clover in soil pots and hydroponics. Environ. Sci. Pollut. Res. Int. 2018, 25, 20680–20690. [Google Scholar] [CrossRef]
- Ringel, M.T.; Brüser, T. The biosynthesis of pyoverdines. Microb. Cell 2018, 5, 424–437. [Google Scholar] [CrossRef]
- Frangipani, E.; Visaggio, D.; Heeb, S.; Kaever, V.; Cámara, M.; Visca, P.; Imperi, F. The Gac/Rsm and cyclic-di-GMP signalling networks coordinately regulate iron uptake in Pseudomonas aeruginosa. Environ. Microbiol. 2014, 16, 676–688. [Google Scholar] [CrossRef]
- Hassan, K.A.; Johnson, A.; Shaffer, B.T.; Ren, Q.; Kidarsa, T.A.; Elbourne, L.D.; Hartney, S.; Duboy, R.; Goebel, N.C.; Zabriskie, T.M.; et al. Inactivation of the GacA response regulator in Pseudomonas fluorescens Pf-5 has far-reaching transcriptomic consequences. Environ. Microbiol. 2010, 12, 899–915. [Google Scholar] [CrossRef]
- Kong, H.S.; Roberts, D.P.; Patterson, C.D.; Kuehne, S.A.; Heeb, S.; Lakshman, D.K.; Lydon, J. Effect of overexpressing rsmA from Pseudomonas aeruginosa on virulence of select phytotoxin-producing strains of P. syringae. Phytopathology 2012, 102, 575–587. [Google Scholar] [CrossRef]
- Ferreiro, M.D.; Gallegos, M.T. Distinctive features of the Gac-Rsm pathway in plant-associated Pseudomonas. Environ. Microbiol. 2021, 23, 5670–5689. [Google Scholar] [CrossRef]
- Yu, X.; Chen, M.; Jiang, Z.; Hu, Y.; Xie, Z. The two-component regulators GacS and GacA positively regulate a nonfluorescent siderophore through the Gac/Rsm signaling cascade in high-siderophore-yielding Pseudomonas sp. strain HYS. J. Bacteriol. 2014, 196, 3259–3270. [Google Scholar] [CrossRef] [Green Version]
- Tan, S.Y.; Liu, Y.; Chua, S.L.; Vejborg, R.M.; Jakobsen, T.H.; Chew, S.C.; Li, Y.; Nielsen, T.E.; Tolker-Nielsen, T.; Yang, L.; et al. Comparative systems biology analysis to study the mode of action of the isothiocyanate compound Iberin on Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2014, 58, 6648–6659. [Google Scholar] [CrossRef]
- Wei, X.; Huang, X.; Tang, L.; Wu, D.; Xu, Y. Global control of GacA in secondary metabolism, primary metabolism, secretion systems, and motility in the rhizobacterium Pseudomonas aeruginosa M18. J. Bacteriol. 2013, 195, 3387–3400. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gui, Z.; You, J.; Xie, G.; Qin, Y.; Wu, T.; Xie, Z. Pseudomonas donghuensis HYS 7-hydroxytropolone contributes to pathogenicity toward Caenorhabditis elegans and is influenced by pantothenic acid. Biochem. Biophys. Res. Commun. 2020, 533, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Muzio, F.M.; Agaras, B.C.; Masi, M.; Tuzi, A.; Evidente, A.; Valverde, C. 7-hydroxytropolone is the main metabolite responsible for the fungal antagonism of Pseudomonas donghuensis strain SVBP6. Environ. Microbiol. 2020, 22, 2550–2563. [Google Scholar] [CrossRef]
- Jiang, Z.; Chen, M.; Yu, X.; Xie, Z. 7-Hydroxytropolone produced and utilized as an iron-scavenger by Pseudomonas donghuensis. Biometals 2016, 29, 817–826. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Wang, P.; Xie, X. A Complex Mechanism Involving LysR and TetR/AcrR That Regulates Iron Scavenger Biosynthesis in Pseudomonas donghuensis HYS. J. Bacteriol. 2018, 200, e00087-18. [Google Scholar] [CrossRef] [Green Version]
- Thieffry, H.S.D.; Huerta, A.M.; Collado-Vides, J. Prediction of transcriptional regulatory sites in the complete genome sequence of Escherichia coli K-12. Bioinformatics 1998, 14, 391–400. [Google Scholar] [CrossRef] [Green Version]
- Raivio, T.L.; Silhavy, T.J. Periplasmic stress and ECF sigma factors. Annu. Rev. Microbiol. 2001, 55, 591–624. [Google Scholar] [CrossRef]
- Mourino, S.; Wilks, A. Extracellular haem utilization by the opportunistic pathogen Pseudomonas aeruginosa and its role in virulence and pathogenesis. Adv. Microb. Physiol. 2021, 79, 89–132. [Google Scholar] [CrossRef]
- Cornelis, P.; Tahrioui, A.; Lesouhaitier, O.; Bouffartigues, E.; Feuilloley, M.; Baysse, C.; Chevalier, S. High affinity iron uptake by pyoverdine in Pseudomonas aeruginosa involves multiple regulators besides Fur, PvdS, and FpvI. Biometals 2022, 22, 369–376. [Google Scholar] [CrossRef]
- Peng, J.; Chen, G.; Xu, X.; Wang, T.; Liang, H. Iron facilitates the RetS-Gac-Rsm cascade to inversely regulate protease IV (piv) expression via the sigma factor PvdS in Pseudomonas aeruginosa. Environ. Microbiol. 2020, 22, 5402–5413. [Google Scholar] [CrossRef]
- Edgar, R.; Xu, X.; Shirley, M.; Konings, A.; Martin, L.; Ackerley, D. Interactions between an anti-sigma protein and two sigma factors that regulate the pyoverdine signaling pathway in Pseudomonas aeruginosa. BMC Microbiol. 2014, 14, 287. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H.; Hu, S.; Yan, X.; Yang, Y.; Liu, W.; Bu, Z.; Li, G.; Cai, W. An Extracytoplasmic Function Sigma/Anti-Sigma Factor System Regulates Hypochlorous Acid Resistance and Impacts Expression of the Type IV Secretion System in Brucella melitensis. J. Bacteriol. 2021, 203, e00127-21. [Google Scholar] [CrossRef] [PubMed]
- Greenwell, R.; Nam, T.W.; Donohue, T.J. Features of Rhodobacter sphaeroides ChrR required for stimuli to promote the dissociation of sigma(E)/ChrR complexes. J. Mol. Biol. 2011, 407, 477–491. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Yang, Z.; Swingle, B.; Kvitko, B.H. AlgU, a Conserved Sigma Factor Regulating Abiotic Stress Tolerance and Promoting Virulence in Pseudomonas syringae. Mol. Plant Microbe Interact. 2021, 34, 326–336. [Google Scholar] [CrossRef]
- Schofield, M.C.; Rodriguez, D.Q.; Kidman, A.A.; Cassin, E.K.; Michaels, L.A.; Campbell, E.A.; Jorth, P.A.; Tseng, B.S. The anti-sigma factor MucA is required for viability in Pseudomonas aeruginosa. Mol. Microbiol. 2021, 116, 550–563. [Google Scholar] [CrossRef]
- Potvin, E.; Sanschagrin, F.; Levesque, R.C. Sigma factors in Pseudomonas aeruginosa. FEMS Microbiol. Rev. 2008, 32, 38–55. [Google Scholar] [CrossRef] [Green Version]
- Martínez-Bueno, M.A.; Tobes, R.; Rey, M.; Ramos, J.L. Detection of multiple extracytoplasmic function (ECF) sigma factors in the genome of Pseudomonas putida KT2440 and their counterparts in Pseudomonas aeruginosa PA01. Environ. Microbiol. 2002, 4, 842–855. [Google Scholar] [CrossRef]
- Butcher, B.G.; Bao, Z.; Wilson, J.; Stodghill, P.; Swingle, B.; Filiatrault, M.; Schneider, D.; Cartinhour, S. The ECF sigma factor, PSPTO_1043, in Pseudomonas syringae pv. tomato DC3000 is induced by oxidative stress and regulates genes involved in oxidative stress response. PLoS ONE 2017, 12, e0180340–e0180360. [Google Scholar] [CrossRef]
- Visca, P.; Leoni, L.; Wilson, M.J.; Lamont, I.L. Iron transport and regulation cell signalling and genomics lessons from Escherichia coli and Pseudomonas. Mol. Microbiol. 2002, 45, 1177–1190. [Google Scholar] [CrossRef]
- Otero-Asman, J.R.; Wettstadt, S.; Bernal, P.; Llamas, M.A. Diversity of extracytoplasmic function sigma (σECF) factor-dependent signaling in Pseudomonas. Mol. Microbiol. 2019, 112, 356–373. [Google Scholar] [CrossRef]
- Krzyzanowska, D.M.; Ossowicki, A.; Rajewska, M.; Maciag, T.; Jablonska, M.; Obuchowski, M.; Heeb, S.; Jafra, S. When Genome-Based Approach Meets the “Old but Good”: Revealing Genes Involved in the Antibacterial Activity of Pseudomonas sp. P482 against Soft Rot Pathogens. Front. Microbiol. 2016, 7, 782–800. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Agaras, B.C.; Iriarte, A.; Valverde, C.F. Genomic insights into the broad antifungal activity, plant-probiotic properties, and their regulation, in Pseudomonas donghuensis strain SVBP6. PLoS ONE 2018, 13, e0194088. [Google Scholar] [CrossRef] [Green Version]
- Xiao, Y.; Wang, P.; Zhu, X.; Xie, Z. Pseudomonas donghuensis HYS gtrA/B/II Gene Cluster Contributes to Its Pathogenicity toward Caenorhabditis elegans. Int. J. Mol. Sci. 2021, 22, 10741–10759. [Google Scholar] [CrossRef]
- Höfte, M.; Buysens, S.; Koedam, N.; Cornelis, P. Zinc affects siderophore-mediated high affinity iron uptake systems in the rhizosphere Pseudomonas aeruginosa 7NSK2. Biometals 1993, 6, 85–91. [Google Scholar] [CrossRef]
- Miyamoto, K.; Kawano, H.; Okai, N.; Hiromoto, T.; Miyano, N.; Tomoo, K.; Tsuchiya, T.; Komano, J.; Tanabe, T.; Funahashi, T.; et al. Iron-Utilization System in Vibrio vulnificus M2799. Mar. Drugs 2021, 19, 710–721. [Google Scholar] [CrossRef] [PubMed]
- Matuszewska, M.; Maciag, T.; Rajewska, M.; Wierzbicka, A.; Jafra, S. The carbon source-dependent pattern of antimicrobial activity and gene expression in Pseudomonas donghuensis P482. Sci. Rep. 2021, 11, 10994–11011. [Google Scholar] [CrossRef]
- Abril, A.G.; Rama, J.L.R.; Sanchez-Perez, A.; Villa, T.G. Prokaryotic sigma factors and their transcriptional counterparts in Archaea and Eukarya. Appl. Microbiol. Biotechnol. 2020, 104, 4289–4302. [Google Scholar] [CrossRef]
- Passmore, I.J.; Dow, J.M.; Coll, F.; Cuccui, J.; Palmer, T.; Wren, B.W. Ferric Citrate Regulator FecR Is Translocated across the Bacterial Inner Membrane via a Unique Twin-Arginine Transport-Dependent Mechanism. J. Bacteriol. 2020, 202, e00541-19. [Google Scholar] [CrossRef] [Green Version]
- Bastiaansen, K.C.; Civantos, C.; Bitter, W.; Llamas, M.A. New Insights into the Regulation of Cell-Surface Signaling Activity Acquired from a Mutagenesis Screen of the Pseudomonas putida IutY Sigma/Anti-Sigma Factor. Front. Microbiol. 2017, 8, 747–762. [Google Scholar] [CrossRef] [Green Version]
- Quesada, J.M.; Otero-Asman, J.R.; Bastiaansen, K.C.; Civantos, C.; Llamas, M.A. The Activity of the Pseudomonas aeruginosa Virulence Regulator σ(VreI) Is Modulated by the Anti-σ Factor VreR and the Transcription Factor PhoB. Front. Microbiol. 2016, 7, 1159–1175. [Google Scholar] [CrossRef]
- Hughes, K.T.; Mathee, K. The anti-sigma factors. Annu. Rev. Microbiol. 1998, 52, 231–286. [Google Scholar] [CrossRef]
- Chevalier, S.; Bouffartigues, E.; Bazire, A.; Tahrioui, A.; Duchesne, R.; Tortuel, D.; Maillot, O.; Clamens, T.; Orange, N.; Feuilloley, M.G.J.; et al. Extracytoplasmic function sigma factors in Pseudomonas aeruginosa. Biochim. Biophys. Acta Gene Regul. Mech. 2019, 1862, 706–721. [Google Scholar] [CrossRef]
- Tettmann, B.; Dotsch, A.; Armant, O.; Fjell, C.D.; Overhage, J. Knockout of extracytoplasmic function sigma factor ECF-10 affects stress resistance and biofilm formation in Pseudomonas putida KT2440. Appl. Environ. Microbiol. 2014, 80, 4911–4919. [Google Scholar] [CrossRef] [Green Version]
- Sun, C.; Guo, Y.; Tang, K.; Wen, Z.; Li, B.; Zeng, Z.; Wang, X. MqsR/MqsA Toxin/Antitoxin System Regulates Persistence and Biofilm Formation in Pseudomonas putida KT2440. Front. Microbiol. 2017, 8, 840–856. [Google Scholar] [CrossRef] [Green Version]
- Hoang, T.T.; Karkhoff-Schweizer, R.R.; Kutchma, A.J.; Schweizer, H.P. A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: Application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene 1998, 212, 77–86. [Google Scholar] [CrossRef]
- Mulcahy, H.; Charron-Mazenod, L.; Lewenza, S. Extracellular DNA chelates cations and induces antibiotic resistance in Pseudomonas aeruginosa biofilms. PLoS Pathog. 2008, 4, e1000213–e1000225. [Google Scholar] [CrossRef] [Green Version]
- Kong, W.; Zhao, J.; Kang, H.; Zhu, M.; Zhou, T.; Deng, X.; Liang, H. ChIP-seq reveals the global regulator AlgR mediating cyclic di-GMP synthesis in Pseudomonas aeruginosa. Nucleic Acids Res. 2015, 43, 8268–8282. [Google Scholar] [CrossRef] [Green Version]
- Kurien, B.T.; Scofield, R.H. Western blotting. Methods 2006, 38, 283–293. [Google Scholar] [CrossRef]
- Griffith, K.L.; Wolf, R.E., Jr. Measuring β-galactosidase activity in bacteria: Cell growth, permeabilization, and enzyme assays in 96-well arrays. Biochem. Biophys. Res. Commun. 2002, 290, 397–402. [Google Scholar] [CrossRef] [Green Version]
- Ortet, P.; De Luca, G.; Whitworth, D.E.; Barakat, M. P2TF: A comprehensive resource for analysis of prokaryotic transcription factors. BMC Genom. 2012, 13, 628–636. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Teng, S.; Wu, T.; Gao, D.; Wu, S.; Xiao, Y.; Long, Y.; Xie, Z. Insight into the Global Negative Regulation of Iron Scavenger 7-HT Biosynthesis by the SigW/RsiW System in Pseudomonas donghuensis HYS. Int. J. Mol. Sci. 2023, 24, 1184. https://doi.org/10.3390/ijms24021184
Teng S, Wu T, Gao D, Wu S, Xiao Y, Long Y, Xie Z. Insight into the Global Negative Regulation of Iron Scavenger 7-HT Biosynthesis by the SigW/RsiW System in Pseudomonas donghuensis HYS. International Journal of Molecular Sciences. 2023; 24(2):1184. https://doi.org/10.3390/ijms24021184
Chicago/Turabian StyleTeng, Shiyu, Tingting Wu, Donghao Gao, Siyi Wu, Yaqian Xiao, Yan Long, and Zhixiong Xie. 2023. "Insight into the Global Negative Regulation of Iron Scavenger 7-HT Biosynthesis by the SigW/RsiW System in Pseudomonas donghuensis HYS" International Journal of Molecular Sciences 24, no. 2: 1184. https://doi.org/10.3390/ijms24021184





