Circulating miRNAs as Noninvasive Biomarkers for PDAC Diagnosis and Prognosis in Mexico
Abstract
1. Introduction
2. Results
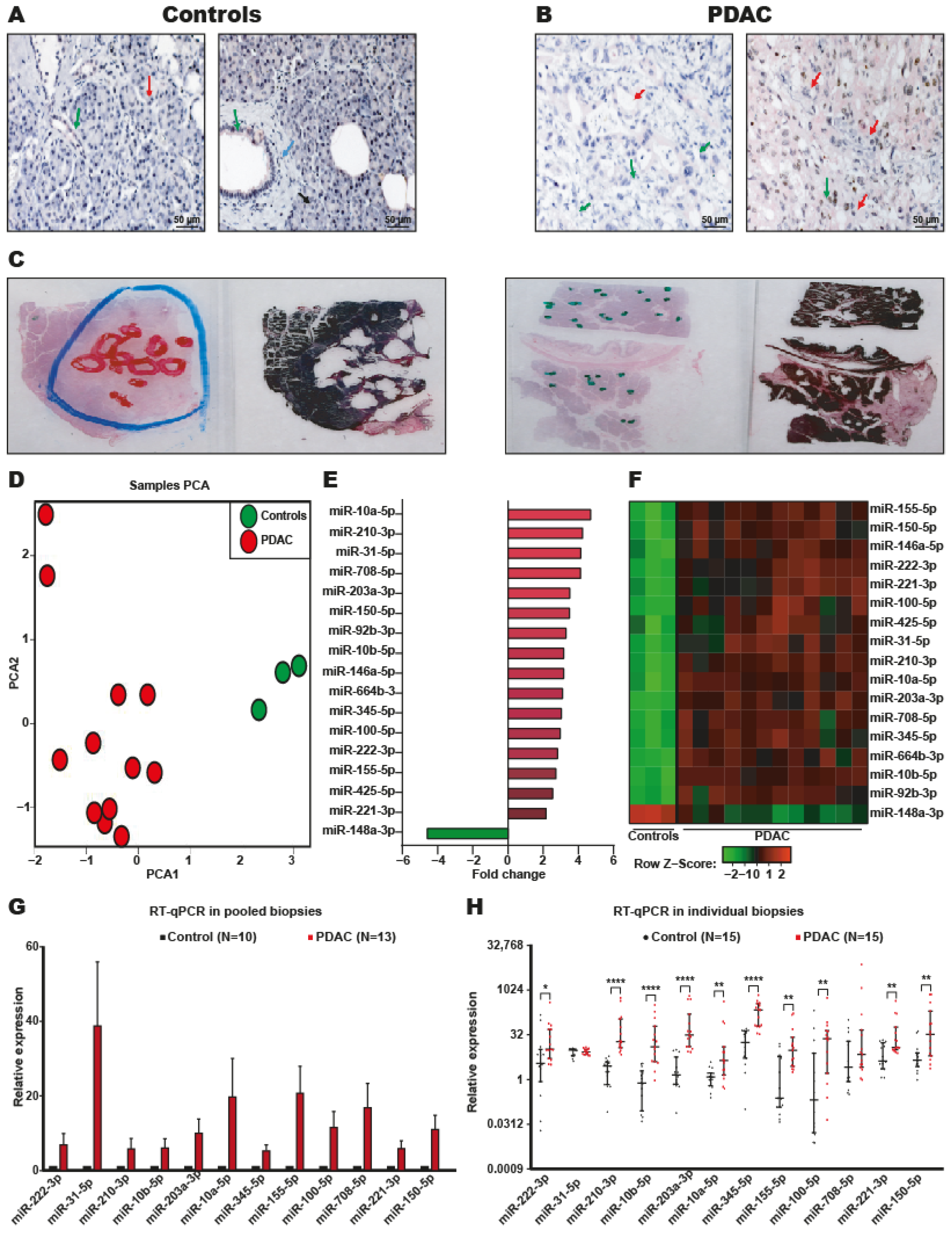
2.1. Identification of DEmiRNAs in PDAC
2.2. Systematic Validation of DEmiRNAs in PDAC
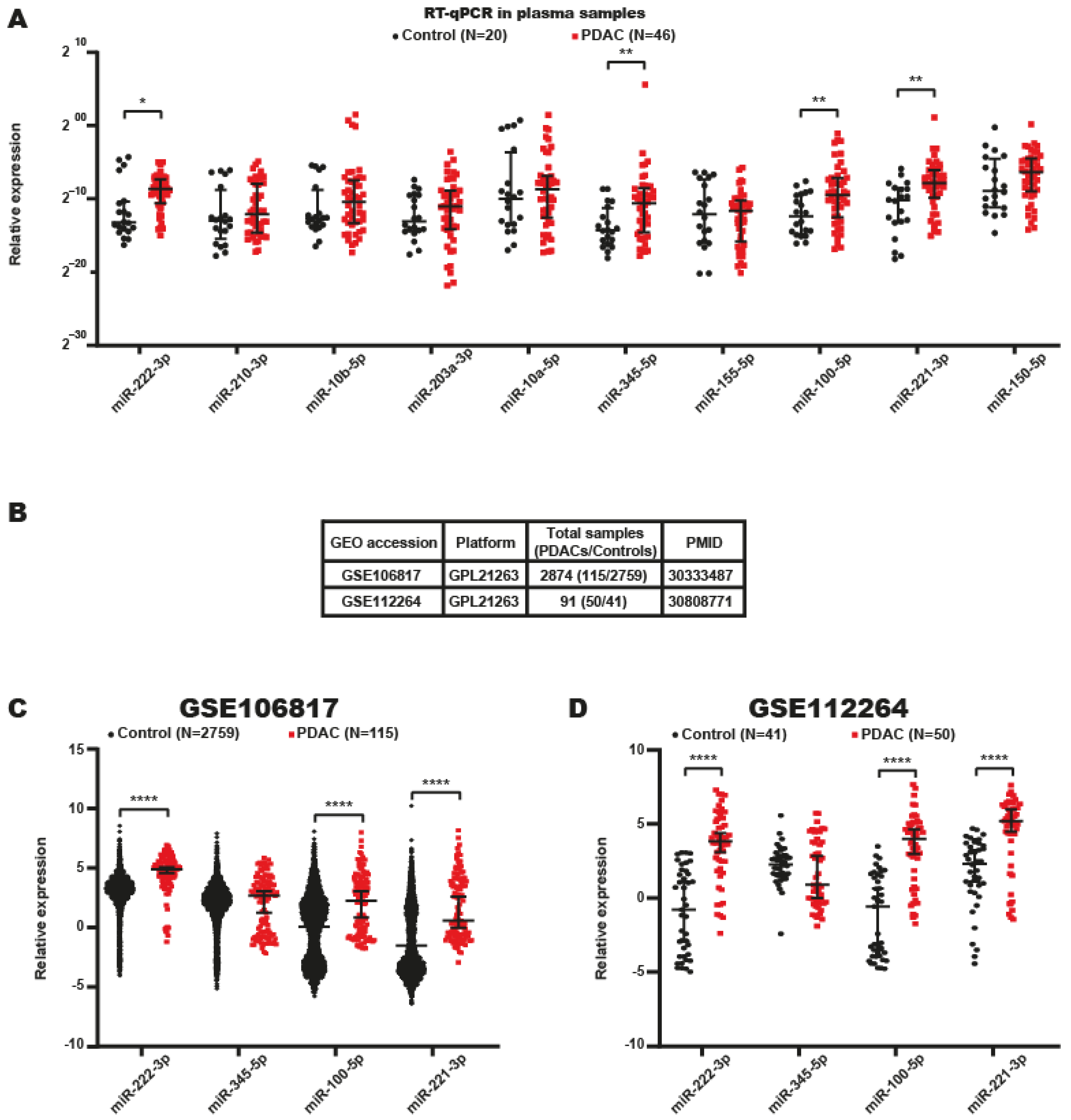
2.3. Identification of miRNA Signatures in the Plasma of Patients with PDAC
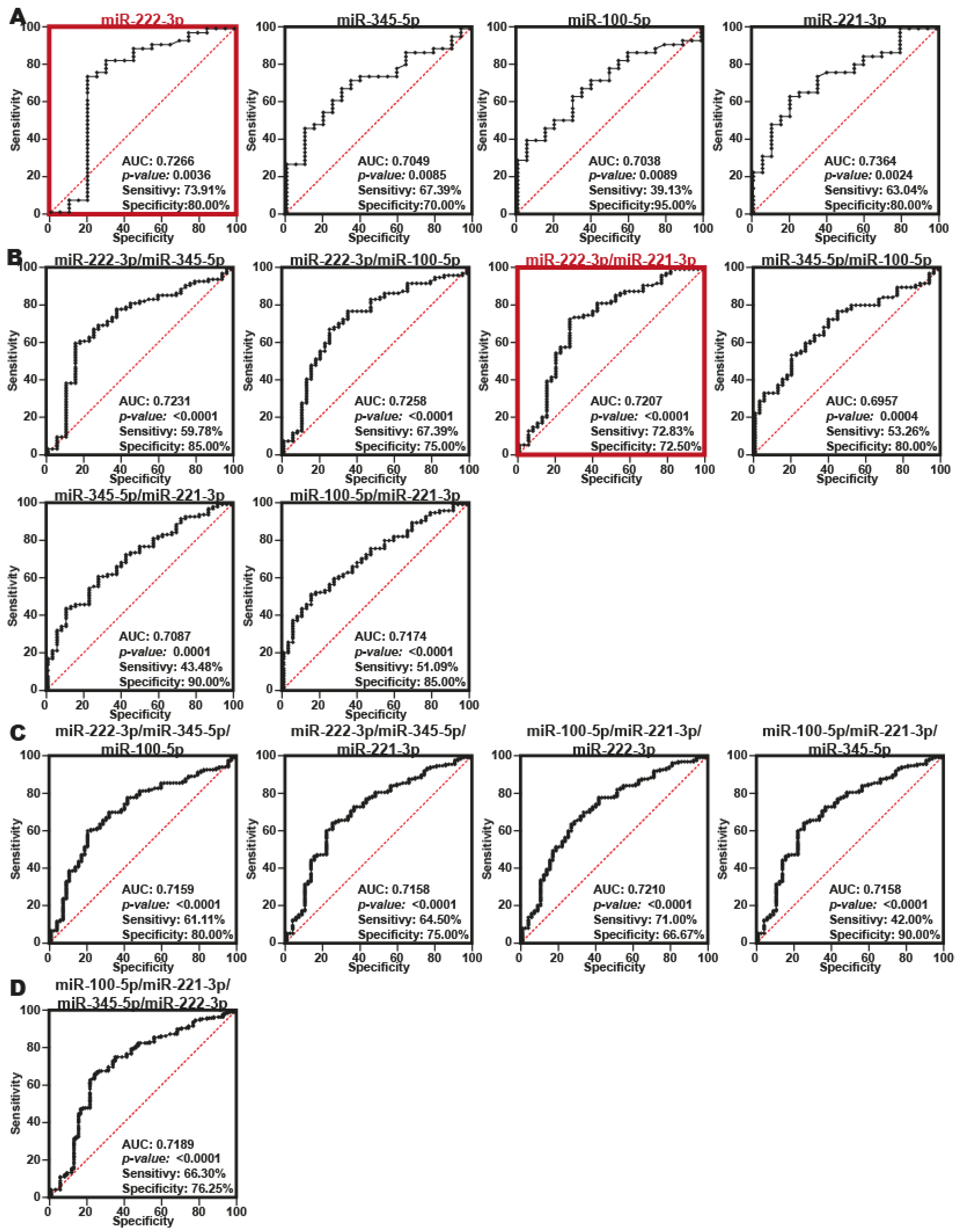
2.4. Serum miRNA Signatures with Potential Value in the Diagnosis of PDAC
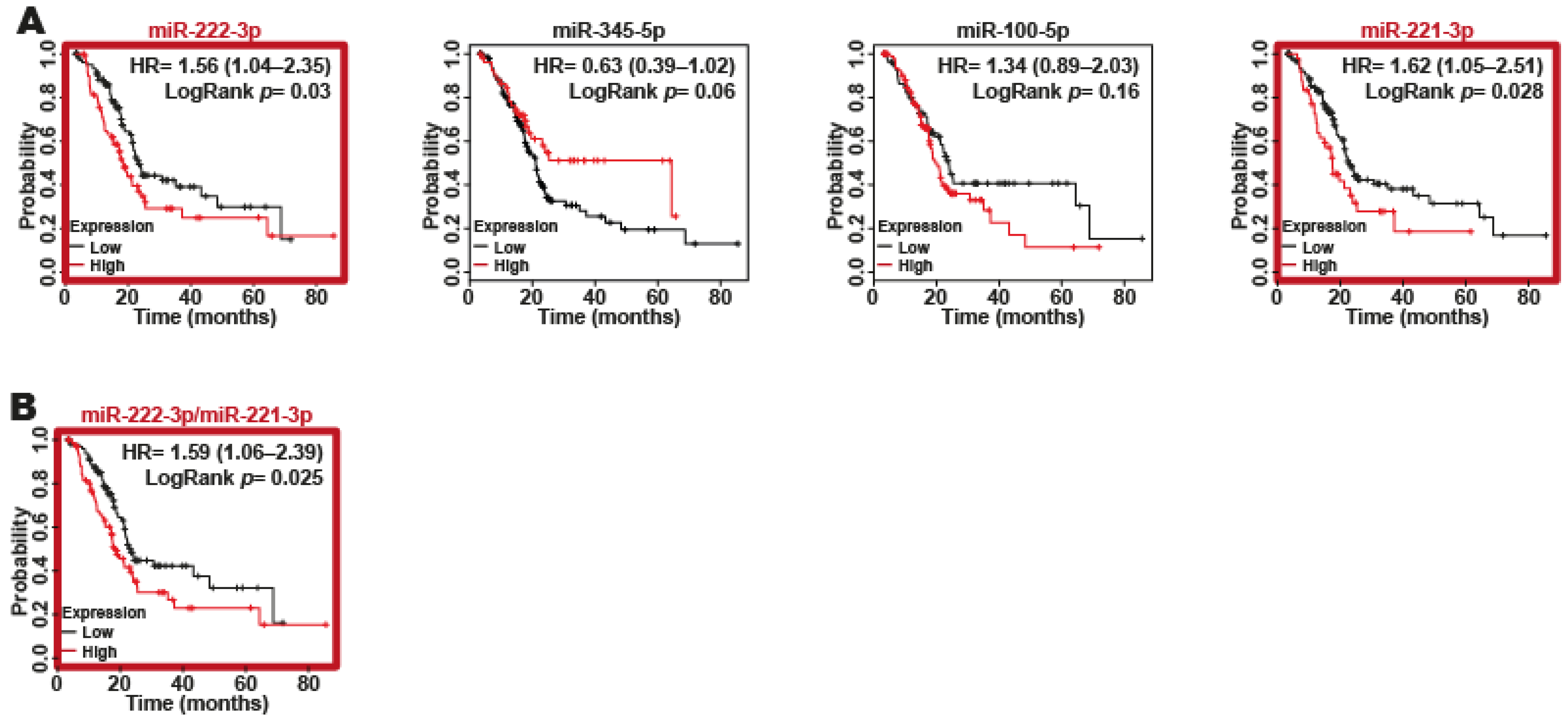
2.5. miR-221-3p and miR-222-3p as Biomarkers of Poor Survival in Patients with PDAC
2.6. Target Gene Prediction and Functional Analysis of Four DEmiRNAs in PDAC
2.7. Putative Target Genes of Four DEmiRNAs in PDAC Tumors Represent Prognostic Factors in Patients with PDAC
3. Discussion
4. Materials and Methods
4.1. The Tissue and Plasma Samples
4.2. Ethics Approval and Consent to Participate
4.3. Tissue Macrodissection and Total RNA Extraction
4.4. Plasma miRNA Isolation and cDNA Synthesis
4.5. miRNA Microarray Assays
4.6. miRNA Differential Expression Analysis
4.7. miRNA Expression by RT-qPCR
4.8. Bioinformatics Analysis of miRNAs and Gene Expression
4.9. miRNA Target Gene Identification
4.10. ROC Curve Analysis
4.11. ROC Curve Combinations
4.12. Survival Analysis
4.13. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E. Jemal AJCacjfc. Cancer Stat. 2022, 72, 7–33. [Google Scholar]
- Rahib, L.; Wehner, M.R.; Matrisian, L.M.; Nead, K.T. Estimated Projection of US Cancer Incidence and Death to 2040. JAMA Netw. Open 2021, 4, e214708. [Google Scholar] [CrossRef] [PubMed]
- Chiu, J.W.; Wong, H.; Leung, R.; Pang, R.; Cheung, T.T.; Fan, S.T.; Poon, R.; Yau, T. Advanced pancreatic cancer: Flourishing novel approaches in the era of biological therapy. Oncologist 2014, 19, 937–950. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rhim, A.D.; Mirek, E.T.; Aiello, N.M.; Maitra, A.; Bailey, J.M.; McAllister, F.; Reichert, M.; Beatty, G.L.; Rustgi, A.K.; Vonderheide, R.H.; et al. EMT and dissemination precede pancreatic tumor formation. Cell 2012, 148, 349–361. [Google Scholar] [CrossRef] [PubMed]
- Ryan, D.P.; Hong, T.S.; Bardeesy, N. Pancreatic adenocarcinoma. N. Engl. J. Med. 2014, 371, 1039–1049. [Google Scholar] [CrossRef]
- O’Neill, R.S.; Stoita, A. Biomarkers in the diagnosis of pancreatic cancer: Are we closer to finding the golden ticket? World J. Gastroenterol. 2021, 27, 4045–4087. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef]
- Smolarz, B.; Durczyński, A.; Romanowicz, H.; Szyłło, K.; Hogendorf, P. miRNAs in Cancer (Review of Literature). Int. J. Mol. Sci. 2022, 23, 2805. [Google Scholar] [CrossRef]
- Annese, T.; Tamma, R.; De Giorgis, M.; Ribatti, D. microRNAs Biogenesis, Functions and Role in Tumor Angiogenesis. Front. Oncol. 2020, 10, 581007. [Google Scholar] [CrossRef]
- Galvão-Lima, L.J.; Morais, A.H.F.; Valentim, R.A.M.; Barreto, E.J.S.S. miRNAs as biomarkers for early cancer detection and their application in the development of new diagnostic tools. BioMed. Eng. OnLine 2021, 20, 21. [Google Scholar] [CrossRef]
- Cronin, M.; Pho, M.; Dutta, D.; Stephans, J.C.; Shak, S.; Kiefer, M.C.; Esteban, J.M.; Baker, J.B. Measurement of gene expression in archival paraffin-embedded tissues: Development and performance of a 92-gene reverse transcriptase-polymerase chain reaction assay. Am. J. Pathol. 2004, 164, 35–42. [Google Scholar] [CrossRef]
- Manterola, L.; Guruceaga, E.; Gallego Perez-Larraya, J.; Gonzalez-Huarriz, M.; Jauregui, P.; Tejada, S.; Diez-Valle, R.; Segura, V.; Sampron, N.; Barrena, C.; et al. A small noncoding RNA signature found in exosomes of GBM patient serum as a diagnostic tool. Neuro-Oncology 2014, 16, 520–527. [Google Scholar] [CrossRef]
- Chen, K.; Wang, Q.; Kornmann, M.; Tian, X.; Yang, Y. The Role of Exosomes in Pancreatic Cancer From Bench to Clinical Application: An Updated Review. Front. Oncol. 2021, 11, 644358. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.; Deng, C.X. Current Progresses of Exosomes as Cancer Diagnostic and Prognostic Biomarkers. Int. J. Biol. Sci. 2019, 15, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Chhatriya, B.; Mukherjee, M.; Ray, S.; Sarkar, P.; Chatterjee, S.; Nath, D.; Das, K.; Goswami, S. Comparison of tumour and serum specific microRNA changes dissecting their role in pancreatic ductal adenocarcinoma: A meta-analysis. BMC Cancer 2019, 19, 1175. [Google Scholar] [CrossRef] [PubMed]
- Dittmar, R.L.; Liu, S.; Tai, M.C.; Rajapakshe, K.; Huang, Y.; Longton, G.; DeCapite, C.; Hurd, M.W.; Paris, P.L.; Kirkwood, K.S.; et al. Plasma miRNA Biomarkers in Limited Volume Samples for Detection of Early-stage Pancreatic Cancer. Cancer Prev. Res. 2021, 14, 729–740. [Google Scholar] [CrossRef]
- Gablo, N.; Trachtova, K.; Prochazka, V.; Hlavsa, J.; Grolich, T.; Kiss, I.; Srovnal, J.; Rehulkova, A.; Lovecek, M.; Skalicky, P.; et al. Identification and Validation of Circulating Micrornas as Prognostic Biomarkers in Pancreatic Ductal Adenocarcinoma Patients Undergoing Surgical Resection. J. Clin. Med. 2020, 9, 2440. [Google Scholar] [CrossRef]
- Gao, Z.; Jiang, W.; Zhang, S.; Li, P. The State of the Art on Blood MicroRNAs in Pancreatic Ductal Adenocarcinoma. Anal. Cell. Pathol. 2019, 2019, 9419072. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Cao, Z.; Liu, W.; You, L.; Zhou, L.; Wang, C.; Lou, W.; Sun, B.; Miao, Y.; Liu, X.; et al. Plasma miRNAs Effectively Distinguish Patients With Pancreatic Cancer From Controls: A Multicenter Study. Ann. Surg. 2016, 263, 1173–1179. [Google Scholar] [CrossRef]
- Zhou, X.; Lu, Z.; Wang, T.; Huang, Z.; Zhu, W.; Miao, Y. Plasma miRNAs in diagnosis and prognosis of pancreatic cancer: A miRNA expression analysis. Gene 2018, 673, 181–193. [Google Scholar] [CrossRef] [PubMed]
- Wnuk, J.; Strzelczyk, J.K.; Gisterek, I. Clinical Value of Circulating miRNA in Diagnosis, Prognosis, Screening and Monitoring Therapy of Pancreatic Ductal Adenocarcinoma-A Review of the Literature. Int. J. Mol. Sci. 2023, 24, 5113. [Google Scholar] [CrossRef] [PubMed]
- Kabiraj, L.; Kundu, A. Potential role of microRNAs in pancreatic cancer manifestation: A review. J. Egypt. Natl. Cancer Inst. 2022, 34, 26. [Google Scholar] [CrossRef] [PubMed]
- Yokoi, A.; Matsuzaki, J.; Yamamoto, Y.; Yoneoka, Y.; Takahashi, K.; Shimizu, H.; Uehara, T.; Ishikawa, M.; Ikeda, S.I.; Sonoda, T.; et al. Integrated extracellular microRNA profiling for ovarian cancer screening. Nat. Commun. 2018, 9, 4319. [Google Scholar] [CrossRef] [PubMed]
- Urabe, F.; Matsuzaki, J.; Yamamoto, Y.; Kimura, T.; Hara, T.; Ichikawa, M.; Takizawa, S.; Aoki, Y.; Niida, S.; Sakamoto, H.; et al. Large-scale Circulating microRNA Profiling for the Liquid Biopsy of Prostate Cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2019, 25, 3016–3025. [Google Scholar] [CrossRef]
- Mandrekar, J.N. Receiver operating characteristic curve in diagnostic test assessment. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2010, 5, 1315–1316. [Google Scholar] [CrossRef]
- Yang, S.; He, P.; Wang, J.; Schetter, A.; Tang, W.; Funamizu, N.; Yanaga, K.; Uwagawa, T.; Satoskar, A.R.; Gaedcke, J.; et al. A Novel MIF Signaling Pathway Drives the Malignant Character of Pancreatic Cancer by Targeting NR3C2. Cancer Res. 2016, 76, 3838–3850. [Google Scholar] [CrossRef]
- Zhang, G.; He, P.; Tan, H.; Budhu, A.; Gaedcke, J.; Ghadimi, B.M.; Ried, T.; Yfantis, H.G.; Lee, D.H.; Maitra, A.; et al. Integration of metabolomics and transcriptomics revealed a fatty acid network exerting growth inhibitory effects in human pancreatic cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2013, 19, 4983–4993. [Google Scholar] [CrossRef]
- Carlsen, A.L.; Joergensen, M.T.; Knudsen, S.; de Muckadell, O.B.; Heegaard, N.H. Cell-free plasma microRNA in pancreatic ductal adenocarcinoma and disease controls. Pancreas 2013, 42, 1107–1113. [Google Scholar] [CrossRef]
- Alemar, B.; Gregorio, C.; Ashton-Prolla, P. miRNAs As Diagnostic and Prognostic Biomarkers in Pancreatic Ductal Adenocarcinoma and Its Precursor Lesions: A Review. Biomark. Insights 2015, 10, 113–124. [Google Scholar] [CrossRef]
- Hawa, Z.; Haque, I.; Ghosh, A.; Banerjee, S.; Harris, L.; Banerjee, S.K. The miRacle in Pancreatic Cancer by miRNAs: Tiny Angels or Devils in Disease Progression. Int. J. Mol. Sci. 2016, 17, 809. [Google Scholar] [CrossRef]
- Wang, D.; Sang, Y.; Sun, T.; Kong, P.; Zhang, L.; Dai, Y.; Cao, Y.; Tao, Z.; Liu, W. Emerging roles and mechanisms of microRNA-222-3p in human cancer (Review). Int. J. Oncol. 2021, 58, 1–12. [Google Scholar] [CrossRef]
- Wang, J.; Chen, J.; Chang, P.; LeBlanc, A.; Li, D.; Abbruzzesse, J.L.; Frazier, M.L.; Killary, A.M.; Sen, S. MicroRNAs in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer Prev. Res. 2009, 2, 807–813. [Google Scholar] [CrossRef]
- Wei, L.; Yao, K.; Gan, S.; Suo, Z. Clinical utilization of serum- or plasma-based miRNAs as early detection biomarkers for pancreatic cancer: A meta-analysis up to now. Medicine 2018, 97, e12132. [Google Scholar] [CrossRef]
- Liu, J.; Gao, J.; Du, Y.; Li, Z.; Ren, Y.; Gu, J.; Wang, X.; Gong, Y.; Wang, W.; Kong, X. Combination of plasma microRNAs with serum CA19-9 for early detection of pancreatic cancer. Int. J. Cancer 2012, 131, 683–691. [Google Scholar] [CrossRef]
- Yan, Q.; Hu, D.; Li, M.; Chen, Y.; Wu, X.; Ye, Q.; Wang, Z.; He, L.; Zhu, J. The Serum MicroRNA Signatures for Pancreatic Cancer Detection and Operability Evaluation. Front. Bioeng. Biotechnol. 2020, 8, 379. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Ouyang, Y.; Che, J.; Li, X.; Zhao, Y.; Yang, K.; Zhao, X.; Chen, Y.; Fan, C.; Yuan, W. Potential Value of miR-221/222 as Diagnostic, Prognostic, and Therapeutic Biomarkers for Diseases. Front. Immunol. 2017, 8, 56. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Xu, J.W.; Wang, L.; Liu, H.; Yan, Y.; Hu, S.Y. MicroRNA-221-3p is up-regulated and serves as a potential biomarker in pancreatic cancer. Artif. Cells Nanomed. Biotechnol. 2018, 46, 482–487. [Google Scholar] [CrossRef]
- Cote, G.A.; Gore, A.J.; McElyea, S.D.; Heathers, L.E.; Xu, H.; Sherman, S.; Korc, M. A pilot study to develop a diagnostic test for pancreatic ductal adenocarcinoma based on differential expression of select miRNA in plasma and bile. Am. J. Gastroenterol. 2014, 109, 1942–1952. [Google Scholar] [CrossRef]
- Zhang, J.; Zhao, C.Y.; Zhang, S.H.; Yu, D.H.; Chen, Y.; Liu, Q.H.; Shi, M.; Ni, C.R.; Zhu, M.H. Upregulation of miR-194 contributes to tumor growth and progression in pancreatic ductal adenocarcinoma. Oncol. Rep. 2014, 31, 1157–1164. [Google Scholar] [CrossRef] [PubMed]
- Sever, R.; Brugge, J.S. Signal transduction in cancer. Cold Spring Harb. Perspect. Med. 2015, 5, a006098. [Google Scholar] [CrossRef] [PubMed]
- Faubert, B.; Solmonson, A.; DeBerardinis, R.J. Metabolic reprogramming and cancer progression. Science 2020, 368, eaaw5473. [Google Scholar] [CrossRef]
- Phan, L.M.; Yeung, S.C.; Lee, M.H. Cancer metabolic reprogramming: Importance, main features, and potentials for precise targeted anti-cancer therapies. Cancer Biol. Med. 2014, 11, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Perera, R.M.; Bardeesy, N. Pancreatic Cancer Metabolism: Breaking It Down to Build It Back Up. Cancer Discov. 2015, 5, 1247–1261. [Google Scholar] [CrossRef] [PubMed]
- Qin, C.; Yang, G.; Yang, J.; Ren, B.; Wang, H.; Chen, G.; Zhao, F.; You, L.; Wang, W.; Zhao, Y. Metabolism of pancreatic cancer: Paving the way to better anticancer strategies. Mol. Cancer 2020, 19, 50. [Google Scholar] [CrossRef] [PubMed]
- Cairns, R.A.; Harris, I.S.; Mak, T.W. Regulation of cancer cell metabolism. Nat. Rev. Cancer 2011, 11, 85–95. [Google Scholar] [CrossRef]
- DeBerardinis, R.J.; Chandel, N.S. Fundamentals of cancer metabolism. Sci. Adv. 2016, 2, e1600200. [Google Scholar] [CrossRef]
- Gonzalez, A.; Hall, M.N.; Lin, S.C.; Hardie, D.G. AMPK and TOR: The Yin and Yang of Cellular Nutrient Sensing and Growth Control. Cell Metab. 2020, 31, 472–492. [Google Scholar] [CrossRef]
- Yuan, H.X.; Xiong, Y.; Guan, K.L. Nutrient sensing, metabolism, and cell growth control. Mol. Cell 2013, 49, 379–387. [Google Scholar] [CrossRef]
- Kang, J.M.; Park, S.; Kim, S.J.; Kim, H.; Lee, B.; Kim, J.; Park, J.; Kim, S.T.; Yang, H.K.; Kim, W.H.; et al. KIAA1324 Suppresses Gastric Cancer Progression by Inhibiting the Oncoprotein GRP78. Cancer Res. 2015, 75, 3087–3097. [Google Scholar] [CrossRef]
- Cho, J.M.; Moon, K.T.; Lee, H.J.; Shin, S.C.; Choi, J.D.; Kang, J.Y.; Yoo, T.K. Nucleobindin 2 expression is an independent prognostic factor for bladder cancer. Medicine 2020, 99, e19597. [Google Scholar] [CrossRef]
- Manoochehri, M.; Wu, Y.; Giese, N.A.; Strobel, O.; Kutschmann, S.; Haller, F.; Hoheisel, J.D.; Moskalev, E.A.; Hackert, T.; Bauer, A.S. SST gene hypermethylation acts as a pan-cancer marker for pancreatic ductal adenocarcinoma and multiple other tumors: Toward its use for blood-based diagnosis. Mol. Oncol. 2020, 14, 1252–1267. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Choi, Y.; Namkung, J.; Yi, S.G.; Kim, H.; Yu, J.; Kim, Y.; Kwon, M.S.; Kwon, W.; Oh, D.Y.; et al. Diagnostic performance enhancement of pancreatic cancer using proteomic multimarker panel. Oncotarget 2017, 8, 93117–93130. [Google Scholar] [CrossRef] [PubMed]
- Murray, M.J.; Bell, E.; Raby, K.L.; Rijlaarsdam, M.A.; Gillis, A.J.; Looijenga, L.H.; Brown, H.; Destenaves, B.; Nicholson, J.C.; Coleman, N. A pipeline to quantify serum and cerebrospinal fluid microRNAs for diagnosis and detection of relapse in paediatric malignant germ-cell tumours. Br. J. Cancer 2016, 114, 151–162. [Google Scholar] [CrossRef]
- Cirera, S.; Busk, P.K. Quantification of miRNAs by a simple and specific qPCR method. Methods Mol. Biol. 2014, 1182, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Felix, T.F.; Lopez Lapa, R.M.; de Carvalho, M.; Bertoni, N.; Tokar, T.; Oliveira, R.A.; MA, M.R.; Hasimoto, C.N.; Oliveira, W.K.; Pelafsky, L.; et al. MicroRNA modulated networks of adaptive and innate immune response in pancreatic ductal adenocarcinoma. PLoS ONE 2019, 14, e0217421. [Google Scholar] [CrossRef] [PubMed]
- Irizarry, R.A.; Hobbs, B.; Collin, F.; Beazer-Barclay, Y.D.; Antonellis, K.J.; Scherf, U.; Speed, T.P. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 2003, 4, 249–264. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Busk, P.K. A tool for design of primers for microRNA-specific quantitative RT-qPCR. BMC Bioinform. 2014, 15, 29. [Google Scholar] [CrossRef]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for functional genomics data sets--update. Nucleic Acids Res. 2013, 41, D991–D995. [Google Scholar] [CrossRef]
- Tang, Z.; Li, C.; Kang, B.; Gao, G.; Li, C.; Zhang, Z. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017, 45, W98–W102. [Google Scholar] [CrossRef]
- Colwill, K.; Renewable Protein Binder Working Group; Graslund, S. A roadmap to generate renewable protein binders to the human proteome. Nat. Meth. 2011, 8, 551–558. [Google Scholar] [CrossRef] [PubMed]
- Kehl, T.; Kern, F.; Backes, C.; Fehlmann, T.; Stockel, D.; Meese, E.; Lenhof, H.P.; Keller, A. miRPathDB 2.0: A novel release of the miRNA Pathway Dictionary Database. Nucleic Acids Res. 2020, 48, D142–D147. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Bailey, A.; Kuleshov, M.V.; Clarke, D.J.B.; Evangelista, J.E.; Jenkins, S.L.; Lachmann, A.; Wojciechowicz, M.L.; Kropiwnicki, E.; Jagodnik, K.M.; et al. Gene Set Knowledge Discovery with Enrichr. Curr. Protoc. 2021, 1, e90. [Google Scholar] [CrossRef] [PubMed]
- Loraine, A.E.; Blakley, I.C.; Jagadeesan, S.; Harper, J.; Miller, G.; Firon, N. Analysis and visualization of RNA-Seq expression data using RStudio, Bioconductor, and Integrated Genome Browser. Methods Mol. Biol. 2015, 1284, 481–501. [Google Scholar] [CrossRef] [PubMed]
- Youden, W.J. Index for rating diagnostic tests. Cancer 1950, 3, 32–35. [Google Scholar] [CrossRef]


| Type of Sample | Characteristic | Controls n = 48 | Patients n = 86 |
|---|---|---|---|
| Age | |||
| Tissue (n = 68) | Median (range) | 50 (26–78) | 63 (40–86) |
| Plasma (n = 66) | Median (range) | 46 (27–62) | 63 (41–83) |
| Gender | |||
| Tissue (n= 68) | Male | 17 | 17 |
| Female | 11 | 23 | |
| Plasma (n = 66) | Male | 9 | 21 |
| Female | 11 | 25 | |
| Alcohol | |||
| Tissue (n = 68) | Yes | 0 | 0 |
| No | 0 | 0 | |
| No data | 28 | 40 | |
| Plasma (n = 66) | Yes | 0 | 9 |
| No | 0 | 31 | |
| No data | 20 | 6 | |
| Smoking | |||
| Tissue (n = 68) | Yes | 0 | 0 |
| No | 0 | 0 | |
| No data | 28 | 40 | |
| Plasma (n = 66) | Yes | 0 | 16 |
| No | 0 | 20 | |
| No data | 20 | 10 | |
| Ethnicity | |||
| Tissue (n = 68) | Mexican-Mestizo | 28 | 40 |
| Plasma (n = 66) | Mexican-Mestizo | 20 | 46 |
| Tumor size | |||
| Tissue (n = 68) | ≤4 cm | - | 9 |
| >4 cm | - | 18 | |
| No data | - | 13 | |
| Plasma (n = 66) | ≤4 cm | - | 1 |
| >4 cm | - | 0 | |
| No data | - | 45 | |
| TNM | |||
| Tissue (n = 68) | I | - | 0 |
| II | - | 18 | |
| III | - | 8 | |
| IV | - | 11 | |
| No data | - | 3 | |
| Plasma (n = 66) | I | - | 1 |
| II | - | 5 | |
| III | - | 18 | |
| IV | - | 20 | |
| No data | - | 2 | |
| Differentiation grade | |||
| Tissue (n = 68) | G1 | - | 2 |
| G2 | - | 30 | |
| G3 | - | 7 | |
| No data | - | 1 | |
| Plasma (n = 66) | G1 | - | 0 |
| G2 | - | 0 | |
| G3 | - | 0 | |
| No data | - | 46 | |
| Survival (Tissue/Plasma) | |||
| Tissue (n = 68) | Short: ≤14 months | - | 15 |
| Long: >14 months | - | 7 | |
| No data | - | 18 | |
| Plasma (n = 66) | Short: ≤14 months | - | 0 |
| Long: >14 months | - | 0 | |
| No data | - | 46 |
| Transcript_ID | logFC | AveExpr | t | p-Value | FDR |
|---|---|---|---|---|---|
| Overexpressed miRNAs | |||||
| miR-222-3p | 2.8844 | 10.8590 | 10.7525 | 9.96 × 10−9 | 6.41 × 10−7 |
| miR-31-5p | 4.1967 | 9.7829 | 10.7844 | 9.56 × 10−9 | 6.53 × 10−7 |
| miR-210-3p | 4.3097 | 8.0389 | 10.3553 | 1.69 × 10−8 | 7.75 × 10−7 |
| miR-10b-5p | 3.2291 | 6.3356 | 10.5711 | 1.27 × 10−8 | 9.64 × 10−7 |
| miR-203a | 3.5790 | 6.0323 | 9.9259 | 3.06 × 10−8 | 1.30 × 10−6 |
| miR-10a-5p | 4.4917 | 8.5372 | 9.5532 | 5.19 × 10−8 | 2.38 × 10−6 |
| miR-345-5p | 3.0996 | 5.9820 | 8.6866 | 1.88 × 10−7 | 2.88 × 10−6 |
| miR-155-5p | 2.7886 | 9.5953 | 8.4357 | 2.78 × 10−7 | 3.96 × 10−6 |
| miR-100-5p | 3.0294 | 10.4069 | 8.2603 | 3.66 × 10−7 | 4.38 × 10−6 |
| miR-708-5p | 4.1888 | 6.7173 | 8.0084 | 5.49 × 10−7 | 6.25 × 10−6 |
| miR-221-3p | 2.2323 | 10.7475 | 7.6056 | 1.06 × 10−6 | 1.10 × 10−5 |
| miR-146a-5p * | 3.2269 | 8.8496 | 7.5014 | 1.27 × 10−6 | 1.26 × 10−5 |
| miR-150-5p | 3.5555 | 8.7285 | 7.4770 | 1.32 × 10−6 | 1.27 × 10−5 |
| miR-664b-3p | 3.1629 | 5.9450 | 8.1304 | 4.51 × 10−7 | 1.35 × 10−5 |
| miR-92b-3p | 3.3605 | 5.8241 | 7.3678 | 1.59 × 10−6 | 1.45 × 10−5 |
| miR-425-5p | 2.6038 | 8.3962 | 6.4191 | 8.51 × 10−6 | 7.36 × 10−5 |
| Downregulated miRNA | |||||
| miR-148a-3p | −4.5795 | 7.7280 | −9.7445 | 3.95 × 10−8 | 9.37 × 10−7 |
| miRNA | Forward | Reverse |
|---|---|---|
| miR-222-3p | GCAGAGCTACATCTGGCT | CCAGTTTTTTTTTTTTTTTACCCAGT |
| miR-31-5p | GCGCAGCTGTGCGTGTGACA | GTCCAGTTTTTTTTTTTTTTTAGCTATG |
| miR-210-3p | GCGCAGCTGTGCGTGTGACA | GTTTTTTTTTTTTTTTCAGCCGCT |
| miR-10b-5p | CAGTACCCTGTAGAACCGA | GGTCCAGTTTTTTTTTTTTTTTCAC |
| miR-203a | CAGGTGAAATGTTTAGGACCA | GGTCCAGTTTTTTTTTTTTTTTCTAGT |
| miR-10a-5p | GCAGTACCCTGTAGATCCGA | GGTCCAGTTTTTTTTTTTTTTTCAC |
| miR-345-5p | GGCTGACTCCTAGTCCAG | GGTCCAGTTTTTTTTTTTTTTTGAG |
| miR-155-5p | CGCAGTTAATGCTAATCGTGATAG | GGTCCAGTTTTTTTTTTTTTTTAACC |
| miR-100-5p | CAGAACCCGTAGATCCGA | GTCCAGTTTTTTTTTTTTTTTACAAG |
| miR-708-5p | CAGAAGGAGCTTACAATCTAGC | GTCCAGTTTTTTTTTTTTTTTCCCA |
| miR-221-3p | GCAGAGCTACATTGTCTGCT | CAGTTTTTTTTTTTTTTTGAAACCCA |
| miR-150-5p | AGTCTCCCAACCCTTGTACCA | GGTCCAGTTTTTTTTTTTTTTTCACT |
| RT-primer | CAGGTCCAGTTTTTTTTTTTTTTTVN | |
| RNU6 | CTCGCTTCGGCAGCACATATACT | ACGCTTCACGAATTTGCGTGTC |
| miR-39-3p | GTCACCGGGTGTAAATCAG | GGTCCAGTTTTTTTTTTTTTTTTTCAAG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Álvarez-Hilario, L.G.; Salmerón-Bárcenas, E.G.; Ávila-López, P.A.; Hernández-Montes, G.; Aréchaga-Ocampo, E.; Herrera-Goepfert, R.; Albores-Saavedra, J.; Manzano-Robleda, M.d.C.; Saldívar-Cerón, H.I.; Martínez-Frías, S.P.; et al. Circulating miRNAs as Noninvasive Biomarkers for PDAC Diagnosis and Prognosis in Mexico. Int. J. Mol. Sci. 2023, 24, 15193. https://doi.org/10.3390/ijms242015193
Álvarez-Hilario LG, Salmerón-Bárcenas EG, Ávila-López PA, Hernández-Montes G, Aréchaga-Ocampo E, Herrera-Goepfert R, Albores-Saavedra J, Manzano-Robleda MdC, Saldívar-Cerón HI, Martínez-Frías SP, et al. Circulating miRNAs as Noninvasive Biomarkers for PDAC Diagnosis and Prognosis in Mexico. International Journal of Molecular Sciences. 2023; 24(20):15193. https://doi.org/10.3390/ijms242015193
Chicago/Turabian StyleÁlvarez-Hilario, Lissuly Guadalupe, Eric Genaro Salmerón-Bárcenas, Pedro Antonio Ávila-López, Georgina Hernández-Montes, Elena Aréchaga-Ocampo, Roberto Herrera-Goepfert, Jorge Albores-Saavedra, María del Carmen Manzano-Robleda, Héctor Iván Saldívar-Cerón, Sandra Paola Martínez-Frías, and et al. 2023. "Circulating miRNAs as Noninvasive Biomarkers for PDAC Diagnosis and Prognosis in Mexico" International Journal of Molecular Sciences 24, no. 20: 15193. https://doi.org/10.3390/ijms242015193
APA StyleÁlvarez-Hilario, L. G., Salmerón-Bárcenas, E. G., Ávila-López, P. A., Hernández-Montes, G., Aréchaga-Ocampo, E., Herrera-Goepfert, R., Albores-Saavedra, J., Manzano-Robleda, M. d. C., Saldívar-Cerón, H. I., Martínez-Frías, S. P., Thompson-Bonilla, M. D. R., Vargas, M., & Hernández-Rivas, R. (2023). Circulating miRNAs as Noninvasive Biomarkers for PDAC Diagnosis and Prognosis in Mexico. International Journal of Molecular Sciences, 24(20), 15193. https://doi.org/10.3390/ijms242015193







