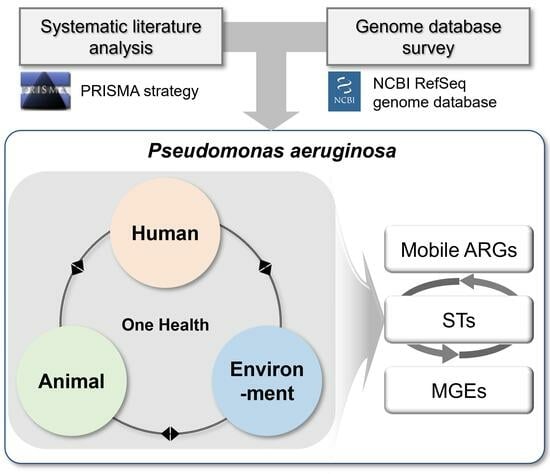
Prioritization of Critical Factors for Surveillance of the Dissemination of Antibiotic Resistance in Pseudomonas aeruginosa: A Systematic Review
Abstract
:1. Introduction
2. Methods
2.1. Systematic Literature Analysis
2.2. Genome Database Survey
3. Results
3.1. Systematic Literature Analysis
3.1.1. MGEs Linked to ARGs in P. aeruginosa
3.1.2. Major STs Involved in ARG Dissemination via MGEs
3.1.3. Mobile ARGs
3.2. Genome Database Survey
3.2.1. Intrinsic and Acquired ARGs in P. aeruginosa
3.2.2. Acquired ARGs in One Health Sectors
3.2.3. ARG Repertoires and the Correlation with One-Health Sector, ST, and MGEs
3.2.4. Mobile ARGs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Moradali, M.F.; Ghods, S.; Rehm, B.H. Pseudomonas aeruginosa lifestyle: A paradigm for adaptation, survival, and persistence. Front. Cell. Infect. Microbiol. 2017, 7, 39. [Google Scholar] [CrossRef]
- Gellatly, S.L.; Hancock, R.E. Pseudomonas aeruginosa: New insights into pathogenesis and host defenses. Pathog. Dis. 2013, 67, 159–173. [Google Scholar] [CrossRef]
- Qin, S.; Xiao, W.; Zhou, C.; Pu, Q.; Deng, X.; Lan, L.; Liang, H.; Song, X.; Wu, M. Pseudomonas aeruginosa: Pathogenesis, virulence factors, antibiotic resistance, interaction with host, technology advances and emerging therapeutics. Signal Transduct. Target. Ther. 2022, 7, 199. [Google Scholar] [CrossRef] [PubMed]
- Cramer, N.; Klockgether, J.; Tümmler, B. Microevolution of Pseudomonas aeruginosa in the airways of people with cystic fibrosis. Curr. Opin. Immunol. 2023, 83, 102328. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, D.; Kollef, M. The epidemiology and pathogenesis and treatment of Pseudomonas aeruginosa infections: An update. Drugs 2021, 81, 2117–2131. [Google Scholar] [CrossRef] [PubMed]
- Oliver, A.; Mulet, X.; López-Causapé, C.; Juan, C. The increasing threat of Pseudomonas aeruginosa high-risk clones. Drug Resist. Updat. 2015, 21–22, 41–59. [Google Scholar] [CrossRef]
- Waters, C.M.; Goldberg, J.B. Pseudomonas aeruginosa in cystic fibrosis: A chronic cheater. Proc. Natl. Acad. Sci. USA 2019, 116, 6525–6527. [Google Scholar] [CrossRef] [PubMed]
- Glen, K.A.; Lamont, I.L. β-Lactam resistance in Pseudomonas aeruginosa: Current status, future prospects. Pathogens 2021, 10, 1638. [Google Scholar] [CrossRef] [PubMed]
- Mesaros, N.; Nordmann, P.; Plésiat, P.; Roussel-Delvallez, M.; Van Eldere, J.; Glupczynski, Y.; Van Laethem, Y.; Jacobs, F.; Lebecque, P.; Malfroot, A.; et al. Pseudomonas aeruginosa: Resistance and therapeutic options at the turn of the new millennium. Clin. Microbiol. Infect. 2007, 13, 560–578. [Google Scholar] [CrossRef]
- Oliver, A.; Cantón, R.; Campo, P.; Baquero, F.; Blázquez, J. High frequency of hypermutable Pseudomonas aeruginosa in cystic fibrosis lung infection. Science 2000, 288, 1251–1254. [Google Scholar] [CrossRef]
- Lister, P.D.; Wolter, D.J.; Hanson, N.D. Antibacterial-resistant Pseudomonas aeruginosa: Clinical impact and complex regulation of chromosomally encoded resistance mechanisms. Clin. Microbiol. Rev. 2009, 22, 582–610. [Google Scholar] [CrossRef] [PubMed]
- Boucher, H.W.; Talbot, G.H.; Bradley, J.S.; Edwards, J.E.; Gilbert, D.; Rice, L.B.; Scheld, M.; Spellberg, B.; Bartlett, J. Bad bugs, no drugs: No ESKAPE! an update from the Infectious Diseases Society of America. Clin. Infect. Dis. 2009, 48, 1–12. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, D.M.P.; Forde, B.M.; Kidd, T.J.; Harris, P.N.A.; Schembri, M.A.; Beatson, S.A.; Paterson, D.L.; Walker, M.J. Antimicrobial resistance in ESKAPE pathogens. Clin. Microbiol. Rev. 2020, 33, e00181-19. [Google Scholar] [CrossRef]
- Tacconelli, E.; Carrara, E.; Savoldi, A.; Harbarth, S.; Mendelson, M.; Monnet, D.L.; Pulcini, C.; Kahlmeter, G.; Kluytmans, J.; Carmeli, Y.; et al. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 2018, 18, 318–327. [Google Scholar] [CrossRef]
- Pang, Z.; Raudonis, R.; Glick, B.R.; Lin, T.J.; Cheng, Z. Antibiotic resistance in Pseudomonas aeruginosa: Mechanisms and alternative therapeutic strategies. Biotechnol. Adv. 2019, 37, 177–192. [Google Scholar] [CrossRef] [PubMed]
- Torrens, G.; Hernández, S.B.; Ayala, J.A.; Moya, B.; Juan, C.; Cava, F.; Oliver, A. Regulation of AmpC-driven β-lactam resistance in Pseudomonas aeruginosa: Different pathways, different signaling. mSystems 2019, 4, e00524-19. [Google Scholar] [CrossRef] [PubMed]
- Morita, Y.; Tomida, J.; Kawamura, Y. MexXY multidrug efflux system of Pseudomonas aeruginosa. Front. Microbiol. 2012, 3, 408. [Google Scholar] [CrossRef]
- Kung, V.L.; Ozer, E.A.; Hauser, A.R. The accessory genome of Pseudomonas aeruginosa. Microbiol. Mol. Biol. Rev. 2010, 74, 621–641. [Google Scholar] [CrossRef]
- López-Causapé, C.; Cabot, G.; Del Barrio-Tofiño, E.; Oliver, A. The versatile mutational resistome of Pseudomonas aeruginosa. Front. Microbiol. 2018, 9, 685. [Google Scholar] [CrossRef]
- de Sousa, T.; Hébraud, M.; Dapkevicius, M.; Maltez, L.; Pereira, J.E.; Capita, R.; Alonso-Calleja, C.; Igrejas, G.; Poeta, P. Genomic and metabolic characteristics of the pathogenicity in Pseudomonas aeruginosa. Int. J. Mol. Sci. 2021, 22, 12892. [Google Scholar] [CrossRef]
- Klockgether, J.; Cramer, N.; Wiehlmann, L.; Davenport, C.F.; Tümmler, B. Pseudomonas aeruginosa genomic structure and diversity. Front. Microbiol. 2011, 2, 150. [Google Scholar] [CrossRef] [PubMed]
- Mathee, K.; Narasimhan, G.; Valdes, C.; Qiu, X.; Matewish, J.M.; Koehrsen, M.; Rokas, A.; Yandava, C.N.; Engels, R.; Zeng, E.; et al. Dynamics of Pseudomonas aeruginosa genome evolution. Proc. Natl. Acad. Sci. USA 2008, 105, 3100–3105. [Google Scholar] [CrossRef] [PubMed]
- Del Barrio-Tofiño, E.; López-Causapé, C.; Oliver, A. Pseudomonas aeruginosa epidemic high-risk clones and their association with horizontally-acquired β-lactamases: 2020 update. Int. J. Antimicrob. Agents 2020, 56, 106196. [Google Scholar] [CrossRef]
- Hu, Y.; Peng, W.; Wu, Y.; Li, H.; Wang, Q.; Yi, H.; Zhang, R.; Shao, B.; Zhu, K. A potential high-risk clone of Pseudomonas aeruginosa ST463. Front. Microbiol. 2021, 12, 670202. [Google Scholar] [CrossRef] [PubMed]
- Cholley, P.; Ka, R.; Guyeux, C.; Thouverez, M.; Guessennd, N.; Ghebremedhin, B.; Frank, T.; Bertrand, X.; Hocquet, D. Population structure of clinical Pseudomonas aeruginosa from West and Central African countries. PLoS ONE 2014, 9, e107008. [Google Scholar] [CrossRef]
- Laborda, P.; Sanz-García, F.; Hernando-Amado, S.; Martínez, J.L. Pseudomonas aeruginosa: An antibiotic resilient pathogen with environmental origin. Curr. Opin. Microbiol. 2021, 64, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.W.; Cha, C.J. Antibiotic resistome from the One-Health perspective: Understanding and controlling antimicrobial resistance transmission. Exp. Mol. Med. 2021, 53, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Page, M.J.; Moher, D.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. PRISMA 2020 explanation and elaboration: Updated guidance and exemplars for reporting systematic reviews. BMJ 2021, 372, n160. [Google Scholar] [CrossRef]
- Jeon, J.H.; Jang, K.M.; Lee, J.H.; Kang, L.W.; Lee, S.H. Transmission of antibiotic resistance genes through mobile genetic elements in Acinetobacter baumannii and gene-transfer prevention. Sci. Total Environ. 2023, 857, 159497. [Google Scholar] [CrossRef]
- Jeukens, J.; Freschi, L.; Kukavica-Ibrulj, I.; Emond-Rheault, J.G.; Tucker, N.P.; Levesque, R.C. Genomics of antibiotic-resistance prediction in Pseudomonas aeruginosa. Ann. N. Y. Acad. Sci. 2019, 1435, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Freschi, L.; Vincent, A.T.; Jeukens, J.; Emond-Rheault, J.G.; Kukavica-Ibrulj, I.; Dupont, M.J.; Charette, S.J.; Boyle, B.; Levesque, R.C. The Pseudomonas aeruginosa pan-genome provides new insights on its population structure, horizontal gene transfer, and pathogenicity. Genome Biol. Evol. 2019, 11, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Alcock, B.P.; Huynh, W.; Chalil, R.; Smith, K.W.; Raphenya, A.R.; Wlodarski, M.A.; Edalatmand, A.; Petkau, A.; Syed, S.A.; Tsang, K.K.; et al. CARD 2023: Expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids. Res. 2023, 51, D690–D699. [Google Scholar] [CrossRef] [PubMed]
- Jolley, K.A.; Bray, J.E.; Maiden, M.C.J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 3, 124. [Google Scholar] [CrossRef]
- Gillings, M.; Boucher, Y.; Labbate, M.; Holmes, A.; Krishnan, S.; Holley, M.; Stokes, H.W. The evolution of class 1 integrons and the rise of antibiotic resistance. J. Bacteriol. 2008, 190, 5095–5100. [Google Scholar] [CrossRef]
- Bardou, P.; Mariette, J.; Escudié, F.; Djemiel, C.; Klopp, C. jvenn: An interactive Venn diagram viewer. BMC Bioinform. 2014, 15, 293. [Google Scholar] [CrossRef]
- Li, H.; Luo, Y.F.; Williams, B.J.; Blackwell, T.S.; Xie, C.M. Structure and function of OprD protein in Pseudomonas aeruginosa: From antibiotic resistance to novel therapies. Int. J. Med. Microbiol. 2012, 302, 63–68. [Google Scholar] [CrossRef]
- Wolter, D.J.; Hanson, N.D.; Lister, P.D. Insertional inactivation of oprD in clinical isolates of Pseudomonas aeruginosa leading to carbapenem resistance. FEMS Microbiol. Lett. 2004, 236, 137–143. [Google Scholar] [CrossRef]
- Martínez, J.L.; Coque, T.M.; Baquero, F. What is a resistance gene? Ranking risk in resistomes. Nat. Rev. Microbiol. 2015, 13, 116–123. [Google Scholar] [CrossRef]
- Zhang, A.N.; Gaston, J.M.; Dai, C.L.; Zhao, S.; Poyet, M.; Groussin, M.; Yin, X.; Li, L.G.; van Loosdrecht, M.C.M.; Topp, E.; et al. An omics-based framework for assessing the health risk of antimicrobial resistance genes. Nat. Commun. 2021, 12, 4765. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Kim, D.W.; Cha, C.J. Overview of bioinformatic methods for analysis of antibiotic resistome from genome and metagenome data. J. Microbiol. 2021, 59, 270–280. [Google Scholar] [CrossRef]
- Naas, T.; Oueslati, S.; Bonnin, R.A.; Dabos, M.L.; Zavala, A.; Dortet, L.; Retailleau, P.; Iorga, B.I. Beta-lactamase database (BLDB)—Structure and function. J. Enzyme Inhib. Med. Chem. 2017, 32, 917–919. [Google Scholar] [CrossRef]
- Snesrud, E.; Maybank, R.; Kwak, Y.I.; Jones, A.R.; Hinkle, M.K.; McGann, P. Chromosomally encoded mcr-5 in colistin-nonsusceptible Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2018, 62, e00679-18. [Google Scholar] [CrossRef]
- Colque, C.A.; Albarracín Orio, A.G.; Feliziani, S.; Marvig, R.L.; Tobares, A.R.; Johansen, H.K.; Molin, S.; Smania, A.M. Hypermutator Pseudomonas aeruginosa exploits multiple genetic pathways to develop multidrug resistance during long-term infections in the airways of cystic fibrosis patients. Antimicrob. Agents Chemother. 2020, 64, e02142-19. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Zhou, J.; Zhang, T.; Li, C.; Chen, J.; Fan, J.; Qu, L.; Su, X. Comparative genomics of the sequential Pseudomonas aeruginosa isolates obtained from the continuous imipenem stress evolution. Appl. Microbiol. Biotechnol. 2020, 104, 10655–10667. [Google Scholar] [CrossRef] [PubMed]
- Schick, A.; Shewaramani, S.; Kassen, R. Genomics of diversification of Pseudomonas aeruginosa in cystic fibrosis lung-like conditions. Genome Biol. Evol. 2022, 14, evac074. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Martínez, J.; Rocha-Gracia, R.D.C.; Bello-López, E.; Cevallos, M.A.; Castañeda-Lucio, M.; Sáenz, Y.; Jiménez-Flores, G.; Cortés-Cortés, G.; López-García, A.; Lozano-Zarain, P. Comparative genomics of Pseudomonas aeruginosa strains isolated from different ecological niches. Antibiotics 2023, 12, 866. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Xie, L.; Wang, C.; Zhou, Q.; Jelsbak, L. Comparative whole-genome analysis of China and global epidemic Pseudomonas aeruginosa high-risk clones. J. Glob. Antimicrob. Resist. 2023, 35, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Pelegrin, A.C.; Palmieri, M.; Mirande, C.; Oliver, A.; Moons, P.; Goossens, H.; van Belkum, A. Pseudomonas aeruginosa: A clinical and genomics update. FEMS Microbiol. Rev. 2021, 45, fuab026. [Google Scholar] [CrossRef]
- Langendonk, R.F.; Neill, D.R.; Fothergill, J.L. The building blocks of antimicrobial resistance in Pseudomonas aeruginosa: Implications for current resistance-breaking therapies. Front. Cell. Infect. Microbiol. 2021, 11, 665759. [Google Scholar] [CrossRef]
- Galetti, R.; Andrade, L.N.; Varani, A.M.; Darini, A.L.C. SPM-1-producing Pseudomonas aeruginosa ST277 carries a chromosomal pack of acquired resistance genes: An example of high-risk clone associated with ‘intrinsic resistome’. J. Glob. Antimicrob. Resist. 2019, 16, 183–186. [Google Scholar] [CrossRef]
- Maravić, A.; Šamanić, I.; Šprung, M.; Fredotović, Ž.; Ilić, N.; Dragičević, J.; Puizina, J. Broad-spectrum resistance of Pseudomonas aeruginosa from shellfish: Infrequent acquisition of novel resistance mechanisms. Environ. Monit. Assess. 2018, 190, 81. [Google Scholar] [CrossRef] [PubMed]
- Jeong, J.H.; Shin, K.S.; Lee, J.W.; Park, E.J.; Son, S.Y. Analysis of a novel class 1 integron containing metallo-β-lactamase gene VIM-2 in Pseudomonas aeruginosa. J. Microbiol. 2009, 47, 753–759. [Google Scholar] [CrossRef] [PubMed]
- Feng, W.; Lv, J.; Wang, H.; Yao, P.; Xiong, L.; Xia, P.; Yuan, Q.; Sun, F. The first report of the blaIMP-10 gene and complete sequence of the IMP-10-encoding plasmid p12NE515 from Pseudomonas aeruginosa in China. Acta Trop. 2022, 228, 106326. [Google Scholar] [CrossRef] [PubMed]
- Rosales-Reyes, R.; Esposito, F.; Fontana, H.; Lezana-Fernández, J.L.; Lascurain, R.; De la Cruz, M.A.; Fuga, B.; Lincopan, N.; Santos-Preciado, J.I. Emergence of GES-19-producing Pseudomonas aeruginosa exoU+ belonging to the global high-risk clone ST235 in cystic fibrosis infection. Diagn. Microbiol. Infect. Dis. 2021, 101, 115454. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, X.; Cai, H.; Zhu, Y.; Ji, J.; Qu, T.; Tu, Y.; Zhou, H.; Yu, Y. Overexpression of blaGES-1 due to a strong promoter in the class 1 integron contributes to decreased ceftazidime-avibactam susceptibility in carbapenem-resistant Pseudomonas aeruginosa ST235. Drug Resist. Updat. 2023, 69, 100973. [Google Scholar] [CrossRef]
- Kiddee, A.; Henghiranyawong, K.; Yimsabai, J.; Tiloklurs, M.; Niumsup, P.R. Nosocomial spread of class 1 integron-carrying extensively drug-resistant Pseudomonas aeruginosa isolates in a Thai hospital. Int. J. Antimicrob. Agents 2013, 42, 301–306. [Google Scholar] [CrossRef] [PubMed]
- Jaillard, M.; van Belkum, A.; Cady, K.C.; Creely, D.; Shortridge, D.; Blanc, B.; Barbu, E.M.; Dunne, W.M., Jr.; Zambardi, G.; Enright, M.; et al. Correlation between phenotypic antibiotic susceptibility and the resistome in Pseudomonas aeruginosa. Int. J. Antimicrob. Agents 2017, 50, 210–218. [Google Scholar] [CrossRef]
- Xu, T.; Wang, J.; Ying, J.; Zhu, T.; Liu, Y.; Xu, L.; Li, P.; Li, P.; Ying, J.; Li, K.; et al. Characterisation of a class 1 integron associated with the formation of quadruple blaGES-5 cassettes from an IncP-1β group plasmid in Pseudomonas aeruginosa. Int. J. Antimicrob. Agents 2018, 52, 485–491. [Google Scholar] [CrossRef]
- Fonseca É, L.; Morgado, S.M.; Caldart, R.V.; Freitas, F.; Vicente, A.C.P. Emergence of a VIM-2-producing extensively drug-resistant (XDR) Pseudomonas aeruginosa ST309 in South America: A comparative genomic analysis. Int. J. Antimicrob. Agents 2022, 59, 106507. [Google Scholar] [CrossRef]
- Chen, J.; Su, Z.; Liu, Y.; Wang, S.; Dai, X.; Li, Y.; Peng, S.; Shao, Q.; Zhang, H.; Wen, P.; et al. Identification and characterization of class 1 integrons among Pseudomonas aeruginosa isolates from patients in Zhenjiang, China. Int. J. Infect. Dis. 2009, 13, 717–721. [Google Scholar] [CrossRef]
- Rajabal, V.; Taner, F.; Sanlidag, T.; Suer, K.; Guler, E.; Sayan, M.; Petrovski, S. Genetic characterisation of antibiotic resistance transposons Tn6608 and Tn6609 isolated from clinical Pseudomonas strains in Cyprus. J. Glob. Antimicrob. Resist. 2021, 26, 330–334. [Google Scholar] [CrossRef]
- Vásquez-Ponce, F.; Higuera-Llantén, S.; Parás-Silva, J.; Gamboa-Acuña, N.; Cortés, J.; Opazo-Capurro, A.; Ugalde, J.A.; Alcalde-Rico, M.; Olivares-Pacheco, J. Genetic characterization of clinically relevant class 1 integrons carried by multidrug resistant bacteria (MDRB) isolated from the gut microbiota of highly antibiotic treated Salmo salar. J. Glob. Antimicrob. Resist. 2022, 29, 55–62. [Google Scholar] [CrossRef]
- Bean, D.C.; Wareham, D.W. Draft genome sequence of a multidrug-resistant Pseudomonas aeruginosa producing blaSIM metallo-β-lactamase: London, UK. J. Glob. Antimicrob. Resist. 2022, 29, 222–224. [Google Scholar] [CrossRef] [PubMed]
- Mihara, T.; Kimura, T.; Momiyama, K.; Kainuma, A.; Akiyama, K.; Ohara, J.; Inoue, K.; Kinoshita, M.; Shimizu, M.; Moriyama, K.; et al. Secondary in-hospital epidemiological investigation after an outbreak of Pseudomonas aeruginosa ST357. J. Infect. Chemother. 2020, 26, 257–265. [Google Scholar] [CrossRef] [PubMed]
- Nemec, A.; Krizova, L.; Maixnerova, M.; Musilek, M. Multidrug-resistant epidemic clones among bloodstream isolates of Pseudomonas aeruginosa in the Czech Republic. Res. Microbiol. 2010, 161, 234–242. [Google Scholar] [CrossRef] [PubMed]
- Martinez, E.; Pérez, J.E.; Buelvas, F.; Tovar, C.; Vanegas, N.; Stokes, H.W. Establishment and multi drug resistance evolution of ST235 Pseudomonas aeruginosa strains in the intensive care unit of a Colombian hospital. Res. Microbiol. 2014, 165, 852–856. [Google Scholar] [CrossRef]
- Rubin, J.; Walker, R.D.; Blickenstaff, K.; Bodeis-Jones, S.; Zhao, S. Antimicrobial resistance and genetic characterization of fluoroquinolone resistance of Pseudomonas aeruginosa isolated from canine infections. Vet. Microbiol. 2008, 131, 164–172. [Google Scholar] [CrossRef]
- Cazares, A.; Moore, M.P.; Hall, J.P.J.; Wright, L.L.; Grimes, M.; Emond-Rhéault, J.G.; Pongchaikul, P.; Santanirand, P.; Levesque, R.C.; Fothergill, J.L.; et al. A megaplasmid family driving dissemination of multidrug resistance in Pseudomonas. Nat. Commun. 2020, 11, 1370. [Google Scholar] [CrossRef] [PubMed]
- Molina-Mora, J.A.; Campos-Sánchez, R.; Rodríguez, C.; Shi, L.; García, F. High quality 3C de novo assembly and annotation of a multidrug resistant ST-111 Pseudomonas aeruginosa genome: Benchmark of hybrid and non-hybrid assemblers. Sci. Rep. 2020, 10, 1392. [Google Scholar] [CrossRef]
- Mobaraki, S.; Aghazadeh, M.; Soroush Barhaghi, M.H.; Yousef Memar, M.; Goli, H.R.; Gholizadeh, P.; Samadi Kafil, H. Prevalence of integrons 1, 2, 3 associated with antibiotic resistance in Pseudomonas aeruginosa isolates from Northwest of Iran. Biomedicine 2018, 8, 2. [Google Scholar] [CrossRef] [PubMed]
- Chairat, S.; Ben Yahia, H.; Rojo-Bezares, B.; Sáenz, Y.; Torres, C.; Ben Slama, K. High prevalence of imipenem-resistant and metallo-β-lactamase-producing Pseudomonas aeruginosa in the Burns Hospital in Tunisia: Detection of a novel class 1 integron. J. Chemother. 2019, 31, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Phoon, H.Y.P.; Hussin, H.; Hussain, B.M.; Thong, K.L. Molecular Characterization of extended-spectrum β-lactamase- and carbapenemase-producing Pseudomonas aeruginosa strains from a Malaysian tertiary hospital. Microb. Drug Resist. 2018, 24, 1108–1116. [Google Scholar] [CrossRef]
- Hong, J.S.; Choi, N.; Kim, S.J.; Choi, K.H.; Roh, K.H.; Lee, S. Molecular characteristics of GES-Type carbapenemase-producing Pseudomonas aeruginosa clinical isolates from long-term care facilities and general hospitals in South Korea. Microb. Drug Resist. 2020, 26, 605–610. [Google Scholar] [CrossRef]
- Liu, M.; Ma, J.; Jia, W.; Li, W. Antimicrobial resistance and molecular characterization of gene cassettes from Class 1 Integrons in Pseudomonas aeruginosa strains. Microb. Drug Resist. 2020, 26, 670–676. [Google Scholar] [CrossRef]
- Pincus, N.B.; Bachta, K.E.R.; Ozer, E.A.; Allen, J.P.; Pura, O.N.; Qi, C.; Rhodes, N.J.; Marty, F.M.; Pandit, A.; Mekalanos, J.J.; et al. Long-term persistence of an extensively drug-resistant subclade of globally distributed Pseudomonas aeruginosa clonal complex 446 in an academic medical center. Clin. Infect. Dis. 2020, 71, 1524–1531. [Google Scholar] [CrossRef] [PubMed]
- Maniati, M.; Ikonomidis, A.; Mantzana, P.; Daponte, A.; Maniatis, A.N.; Pournaras, S. A highly carbapenem-resistant Pseudomonas aeruginosa isolate with a novel blaVIM-4/blaP1b integron overexpresses two efflux pumps and lacks OprD. J. Antimicrob. Chemother. 2007, 60, 132–135. [Google Scholar] [CrossRef]
- Botelho, J.; Grosso, F.; Quinteira, S.; Mabrouk, A.; Peixe, L. The complete nucleotide sequence of an IncP-2 megaplasmid unveils a mosaic architecture comprising a putative novel blaVIM-2-harbouring transposon in Pseudomonas aeruginosa. J. Antimicrob. Chemother. 2017, 72, 2225–2229. [Google Scholar] [CrossRef] [PubMed]
- Zhan, Z.; Hu, L.; Jiang, X.; Zeng, L.; Feng, J.; Wu, W.; Chen, W.; Yang, H.; Yang, W.; Gao, B.; et al. Plasmid and chromosomal integration of four novel blaIMP-carrying transposons from Pseudomonas aeruginosa, Klebsiella pneumoniae and an Enterobacter sp. J. Antimicrob. Chemother. 2018, 73, 3005–3015. [Google Scholar] [CrossRef]
- Jahan, M.I.; Rahaman, M.M.; Hossain, M.A.; Sultana, M. Occurrence of intI1-associated VIM-5 carbapenemase and co-existence of all four classes of β-lactamase in carbapenem-resistant clinical Pseudomonas aeruginosa DMC-27b. J. Antimicrob. Chemother. 2020, 75, 86–91. [Google Scholar] [CrossRef]
- Roy Chowdhury, P.; Scott, M.; Worden, P.; Huntington, P.; Hudson, B.; Karagiannis, T.; Charles, I.G.; Djordjevic, S.P. Genomic islands 1 and 2 play key roles in the evolution of extensively drug-resistant ST235 isolates of Pseudomonas aeruginosa. Open. Biol. 2016, 6, 150175. [Google Scholar] [CrossRef] [PubMed]
- Khosravi, Y.; Tay, S.T.; Vadivelu, J. Analysis of integrons and associated gene cassettes of metallo-β-lactamase-positive Pseudomonas aeruginosa in Malaysia. J. Med. Microbiol. 2011, 60, 988–994. [Google Scholar] [CrossRef]
- Fusté, E.; López-Jiménez, L.; Segura, C.; Gainza, E.; Vinuesa, T.; Viñas, M. Carbapenem-resistance mechanisms of multidrug-resistant Pseudomonas aeruginosa. J. Med. Microbiol. 2013, 62, 1317–1325. [Google Scholar] [CrossRef] [PubMed]
- Lin, D.; Foley, S.L.; Qi, Y.; Han, J.; Ji, C.; Li, R.; Wu, C.; Shen, J.; Wang, Y. Characterization of antimicrobial resistance of Pseudomonas aeruginosa isolated from canine infections. J. Appl. Microbiol. 2012, 113, 16–23. [Google Scholar] [CrossRef]
- Arais, L.R.; Barbosa, A.V.; Carvalho, C.A.; Cerqueira, A.M. Antimicrobial resistance, integron carriage, and gyrA and gyrB mutations in Pseudomonas aeruginosa isolated from dogs with otitis externa and pyoderma in Brazil. Vet. Dermatol. 2016, 27, 113-7e31. [Google Scholar] [CrossRef] [PubMed]
- Hyun, J.E.; Chung, T.H.; Hwang, C.Y. Identification of VIM-2 metallo-β-lactamase-producing Pseudomonas aeruginosa isolated from dogs with pyoderma and otitis in Korea. Vet. Dermatol. 2018, 29, 186-e168. [Google Scholar] [CrossRef] [PubMed]
- Kitao, T.; Miyoshi-Akiyama, T.; Kirikae, T. AAC(6′)-Iaf, a novel aminoglycoside 6′-N-acetyltransferase from multidrug-resistant Pseudomonas aeruginosa clinical isolates. Antimicrob. Agents Chemother. 2009, 53, 2327–2334. [Google Scholar] [CrossRef] [PubMed]
- Perez, F.; Hujer, A.M.; Marshall, S.H.; Ray, A.J.; Rather, P.N.; Suwantarat, N.; Dumford, D., 3rd; O’Shea, P.; Domitrovic, T.N.; Salata, R.A.; et al. Extensively drug-resistant Pseudomonas aeruginosa isolates containing blaVIM-2 and elements of Salmonella genomic island 2: A new genetic resistance determinant in Northeast Ohio. Antimicrob. Agents Chemother. 2014, 58, 5929–5935. [Google Scholar] [CrossRef]
- Pournaras, S.; Tsakris, A.; Maniati, M.; Tzouvelekis, L.S.; Maniatis, A.N. Novel variant (blaVIM-4) of the metallo-β-lactamase gene blaVIM-1 in a clinical strain of Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2002, 46, 4026–4028. [Google Scholar] [CrossRef] [PubMed]
- Prakki, S.R.S.; Hon, P.Y.; Lim, Z.Q.; Thevasagayam, N.M.; Loy, S.Q.D.; De, P.P.; Marimuthu, K.; Vasoo, S.; Ng, O.T. Dissemination of Pseudomonas aeruginosa blaNDM-1-positive ST308 clone in Singapore. Microbiol. Spectr. 2023, 11, e0403322. [Google Scholar] [CrossRef]
- Liakopoulos, A.; Mavroidi, A.; Katsifas, E.A.; Theodosiou, A.; Karagouni, A.D.; Miriagou, V.; Petinaki, E. Carbapenemase-producing Pseudomonas aeruginosa from central Greece: Molecular epidemiology and genetic analysis of class I integrons. BMC Infect. Dis. 2013, 13, 505. [Google Scholar] [CrossRef]
- Mano, Y.; Saga, T.; Ishii, Y.; Yoshizumi, A.; Bonomo, R.A.; Yamaguchi, K.; Tateda, K. Molecular analysis of the integrons of metallo-β-lactamase-producing Pseudomonas aeruginosa isolates collected by nationwide surveillance programs across Japan. BMC Microbiol. 2015, 15, 41. [Google Scholar] [CrossRef] [PubMed]
- Botelho, J.; Roberts, A.P.; León-Sampedro, R.; Grosso, F.; Peixe, L. Carbapenemases on the move: It’s good to be on ICEs. Mob. DNA 2018, 9, 37. [Google Scholar] [CrossRef]
- Böhm, M.E.; Razavi, M.; Marathe, N.P.; Flach, C.F.; Larsson, D.G.J. Discovery of a novel integron-borne aminoglycoside resistance gene present in clinical pathogens by screening environmental bacterial communities. Microbiome 2020, 8, 41. [Google Scholar] [CrossRef] [PubMed]
- Hocquet, D.; Llanes, C.; Thouverez, M.; Kulasekara, H.D.; Bertrand, X.; Plésiat, P.; Mazel, D.; Miller, S.I. Evidence for induction of integron-based antibiotic resistance by the SOS response in a clinical setting. PLoS Pathog. 2012, 8, e1002778. [Google Scholar] [CrossRef] [PubMed]
- López-García, A.; Rocha-Gracia, R.D.C.; Bello-López, E.; Juárez-Zelocualtecalt, C.; Sáenz, Y.; Castañeda-Lucio, M.; López-Pliego, L.; González-Vázquez, M.C.; Torres, C.; Ayala-Nuñez, T.; et al. Characterization of antimicrobial resistance mechanisms in carbapenem-resistant Pseudomonas aeruginosa carrying IMP variants recovered from a Mexican Hospital. Infect. Drug Resist. 2018, 11, 1523–1536. [Google Scholar] [CrossRef]
- Ahmadian, L.; Haghshenas, M.R.; Mirzaei, B.; Norouzi Bazgir, Z.; Goli, H.R. Distribution and molecular characterization of resistance gene cassettes containing class 1 integrons in multi-drug resistant (MDR) clinical isolates of Pseudomonas aeruginosa. Infect. Drug Resist. 2020, 13, 2773–2781. [Google Scholar] [CrossRef] [PubMed]
- Di Pilato, V.; Antonelli, A.; Giani, T.; Henrici De Angelis, L.; Rossolini, G.M.; Pollini, S. Identification of a novel plasmid lineage associated with the dissemination of metallo-β-lactamase genes among Pseudomonads. Front. Microbiol. 2019, 10, 1504. [Google Scholar] [CrossRef] [PubMed]
- Doumith, M.; Alhassinah, S.; Alswaji, A.; Alzayer, M.; Alrashidi, E.; Okdah, L.; Aljohani, S.; Balkhy, H.H.; Alghoribi, M.F. Genomic characterization of carbapenem-non-susceptible Pseudomonas aeruginosa clinical isolates from Saudi Arabia revealed a global dissemination of GES-5-producing ST235 and VIM-2-producing ST233 sub-lineages. Front. Microbiol. 2021, 12, 765113. [Google Scholar] [CrossRef]
- Lin, H.; Feng, C.; Zhu, T.; Li, A.; Liu, S.; Zhang, L.; Li, Q.; Zhang, X.; Lin, L.; Lu, J.; et al. Molecular mechanism of the β-lactamase mediated β-lactam antibiotic resistance of Pseudomonas aeruginosa isolated from a Chinese teaching hospital. Front. Microbiol. 2022, 13, 855961. [Google Scholar] [CrossRef]
- Gómez-Martínez, J.; Rocha-Gracia, R.D.C.; Bello-López, E.; Cevallos, M.A.; Castañeda-Lucio, M.; López-García, A.; Sáenz, Y.; Jiménez-Flores, G.; Cortés-Cortés, G.; Lozano-Zarain, P. A Plasmid carrying blaIMP-56 in Pseudomonas aeruginosa belonging to a novel resistance plasmid family. Microorganisms 2022, 10, 1863. [Google Scholar] [CrossRef]
- Colinon, C.; Jocktane, D.; Brothier, E.; Rossolini, G.M.; Cournoyer, B.; Nazaret, S. Genetic analyses of Pseudomonas aeruginosa isolated from healthy captive snakes: Evidence of high inter- and intrasite dissemination and occurrence of antibiotic resistance genes. Environ. Microbiol. 2010, 12, 716–729. [Google Scholar] [CrossRef] [PubMed]
- Peter, S.; Bosio, M.; Gross, C.; Bezdan, D.; Gutierrez, J.; Oberhettinger, P.; Liese, J.; Vogel, W.; Dörfel, D.; Berger, L.; et al. Tracking of antibiotic resistance transfer and rapid plasmid evolution in a hospital setting by nanopore sequencing. mSphere 2020, 5, e00525-20. [Google Scholar] [CrossRef]
- Amoureux, L.; Riedweg, K.; Chapuis, A.; Bador, J.; Siebor, E.; Péchinot, A.; Chrétien, M.L.; de Curraize, C.; Neuwirth, C. Nosocomial infections with IMP-19-producing Pseudomonas aeruginosa linked to contaminated sinks, France. Emerg. Infect. Dis. 2017, 23, 304–307. [Google Scholar] [CrossRef]
- Libisch, B.; Balogh, B.; Füzi, M. Identification of two multidrug-resistant Pseudomonas aeruginosa clonal lineages with a countrywide distribution in Hungary. Curr. Microbiol. 2009, 58, 111–116. [Google Scholar] [CrossRef]
- Sánchez-Martinez, G.; Garza-Ramos, U.J.; Reyna-Flores, F.L.; Gaytán-Martínez, J.; Lorenzo-Bautista, I.G.; Silva-Sanchez, J. In169, a new class 1 integron that encoded blaIMP-18 in a multidrug-resistant Pseudomonas aeruginosa isolate from Mexico. Arch. Med. Res. 2010, 41, 235–239. [Google Scholar] [CrossRef]
- Meradji, S.; Barguigua, A.; Bentakouk, M.C.; Nayme, K.; Zerouali, K.; Mazouz, D.; Chettibi, H.; Timinouni, M. Epidemiology and virulence of VIM-4 metallo-β-lactamase-producing Pseudomonas aeruginosa isolated from burn patients in eastern Algeria. Burns 2016, 42, 906–918. [Google Scholar] [CrossRef]
- Mózes, J.; Szűcs, I.; Molnár, D.; Jakab, P.; Fatemeh, E.; Szilasi, M.; Majoros, L.; Orosi, P.; Kardos, G. A potential role of aminoglycoside resistance in endemic occurrence of Pseudomonas aeruginosa strains in lower airways of mechanically ventilated patients. Diagn. Microbiol. Infect. Dis. 2014, 78, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Balero de Paula, S.; Cayô, R.; Streling, A.P.; Silva Nodari, C.; Pereira Matos, A.; Eches Perugini, M.R.; Gales, A.C.; Carrara-Marroni, F.E.; Yamada-Ogatta, S.F. Detection of blaVIM-7 in an extensively drug-resistant Pseudomonas aeruginosa isolate belonging to ST1284 in Brazil. Diagn. Microbiol. Infect. Dis. 2017, 89, 80–82. [Google Scholar] [CrossRef]
- Lee, M.F.; Peng, C.F.; Hsu, H.J.; Chen, Y.H. Molecular characterisation of the metallo-β-lactamase genes in imipenem-resistant Gram-negative bacteria from a university hospital in southern Taiwan. Int. J. Antimicrob. Agents 2008, 32, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Martínez, L.; López-Jiménez, L.; Fusté, E.; Vinuesa, T.; Martínez, J.P.; Viñas, M. Class 1 integrons in environmental and clinical isolates of Pseudomonas aeruginosa. Int. J. Antimicrob. Agents 2011, 38, 398–402. [Google Scholar] [CrossRef]
- Bocharova, Y.; Savinova, T.; Lazareva, A.; Polikarpova, S.; Gordinskaya, N.; Mayanskiy, N.; Chebotar, I. Genotypes, carbapenemase carriage, integron diversity and oprD alterations among carbapenem-resistant Pseudomonas aeruginosa from Russia. Int. J. Antimicrob. Agents 2020, 55, 105899. [Google Scholar] [CrossRef]
- Rojo-Bezares, B.; Estepa, V.; Cebollada, R.; de Toro, M.; Somalo, S.; Seral, C.; Castillo, F.J.; Torres, C.; Sáenz, Y. Carbapenem-resistant Pseudomonas aeruginosa strains from a Spanish hospital: Characterization of metallo-β-lactamases, porin OprD and integrons. Int. J. Med. Microbiol. 2014, 304, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Kostyanev, T.; Nguyen, M.N.; Markovska, R.; Stankova, P.; Xavier, B.B.; Lammens, C.; Marteva-Proevska, Y.; Velinov, T.; Cantón, R.; Goossens, H.; et al. Emergence of ST654 Pseudomonas aeruginosa co-harbouring blaNDM-1 and blaGES-5 in novel class I integron In1884 from Bulgaria. J. Glob. Antimicrob. Resist 2020, 22, 672–673. [Google Scholar] [CrossRef] [PubMed]
- Murugan, N.; Malathi, J.; Umashankar, V.; Madhavan, H.N. Unraveling genomic and phenotypic nature of multidrug-resistant (MDR) Pseudomonas aeruginosa VRFPA04 isolated from keratitis patient. Microbiol. Res. 2016, 193, 140–149. [Google Scholar] [CrossRef]
- Caetano, T.; Ferreira, S.; Mondego, A.P.; Correia, A.; Mendo, S. In99, an In100-related integron, its occurrence and prevalence in clinical Pseudomonas aeruginosa strains from a central region of Portugal. Epidemiol. Infect. 2007, 135, 502–504. [Google Scholar] [CrossRef]
- Girlich, D.; Naas, T.; Leelaporn, A.; Poirel, L.; Fennewald, M.; Nordmann, P. Nosocomial spread of the integron-located veb-1-like cassette encoding an extended-pectrum β-lactamase in Pseudomonas aeruginosa in Thailand. Clin. Infect. Dis. 2002, 34, 603–611. [Google Scholar] [CrossRef] [PubMed]
- Zaidi, F.Z.; Dali-Yahia, R.; Zenati, K.; Yazi, L.; Lounes, M.; Aberkane, S.; Jean Pierre, H.; Barraud, O.; Godreuil, S.; Touati, A. Characterization of VIM-4 producing clinical Pseudomonas aeruginosa isolates from western Algeria: Sequence type and class 1 integron description. Microb. Drug Resist. 2020, 26, 1437–1441. [Google Scholar] [CrossRef]
- San Millan, A.; Toll-Riera, M.; Escudero, J.A.; Cantón, R.; Coque, T.M.; MacLean, R.C. Sequencing of plasmids pAMBL1 and pAMBL2 from Pseudomonas aeruginosa reveals a blaVIM-1 amplification causing high-level carbapenem resistance. J. Antimicrob. Chemother. 2015, 70, 3000–3003. [Google Scholar] [CrossRef]
- Rojo-Bezares, B.; Cavalié, L.; Dubois, D.; Oswald, E.; Torres, C.; Sáenz, Y. Characterization of carbapenem resistance mechanisms and integrons in Pseudomonas aeruginosa strains from blood samples in a French hospital. J. Med. Microbiol. 2016, 65, 311–319. [Google Scholar] [CrossRef]
- Toval, F.; Guzmán-Marte, A.; Madriz, V.; Somogyi, T.; Rodríguez, C.; García, F. Predominance of carbapenem-resistant Pseudomonas aeruginosa isolates carrying blaIMP and blaVIM metallo-β-lactamases in a major hospital in Costa Rica. J. Med. Microbiol. 2015, 64, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Llanes, C.; Neuwirth, C.; El Garch, F.; Hocquet, D.; Plésiat, P. Genetic analysis of a multiresistant strain of Pseudomonas aeruginosa producing PER-1 β-lactamase. Clin. Microbiol. Infect. 2006, 12, 270–278. [Google Scholar] [CrossRef] [PubMed]
- Edalucci, E.; Spinelli, R.; Dolzani, L.; Riccio, M.L.; Dubois, V.; Tonin, E.A.; Rossolini, G.M.; Lagatolla, C. Acquisition of different carbapenem resistance mechanisms by an epidemic clonal lineage of Pseudomonas aeruginosa. Clin. Microbiol. Infect. 2008, 14, 88–90. [Google Scholar] [CrossRef] [PubMed]
- Quinones-Falconi, F.; Galicia-Velasco, M.; Marchiaro, P.; Mussi, M.A.; Ballerini, V.; Vila, A.J.; Viale, A.M.; Bermejo-Morales, K.; Limansky, A.S. Emergence of Pseudomonas aeruginosa strains producing metallo-β-lactamases of the IMP-15 and VIM-2 types in Mexico. Clin. Microbiol. Infect. 2010, 16, 126–131. [Google Scholar] [CrossRef]
- Henrichfreise, B.; Wiegand, I.; Pfister, W.; Wiedemann, B. Resistance mechanisms of multiresistant Pseudomonas aeruginosa strains from Germany and correlation with hypermutation. Antimicrob. Agents Chemother. 2007, 51, 4062–4070. [Google Scholar] [CrossRef] [PubMed]
- Janice, J.; Agyepong, N.; Owusu-Ofori, A.; Govinden, U.; Essack, S.Y.; Samuelsen, Ø.; Sundsfjord, A.; Pedersen, T. Carbapenem resistance determinants acquired through novel chromosomal integrations in extensively drug-resistant Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2021, 65, e0028921. [Google Scholar] [CrossRef]
- Xiong, J.; Alexander, D.C.; Ma, J.H.; Déraspe, M.; Low, D.E.; Jamieson, F.B.; Roy, P.H. Complete sequence of pOZ176, a 500-kilobase IncP-2 plasmid encoding IMP-9-mediated carbapenem resistance, from outbreak isolate Pseudomonas aeruginosa 96. Antimicrob. Agents Chemother. 2013, 57, 3775–3782. [Google Scholar] [CrossRef]
- El Garch, F.; Bogaerts, P.; Bebrone, C.; Galleni, M.; Glupczynski, Y. OXA-198, an acquired carbapenem-hydrolyzing class D β-lactamase from Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2011, 55, 4828–4833. [Google Scholar] [CrossRef]
- Gutiérrez, O.; Juan, C.; Cercenado, E.; Navarro, F.; Bouza, E.; Coll, P.; Pérez, J.L.; Oliver, A. Molecular epidemiology and mechanisms of carbapenem resistance in Pseudomonas aeruginosa isolates from Spanish hospitals. Antimicrob. Agents Chemother. 2007, 51, 4329–4335. [Google Scholar] [CrossRef] [PubMed]
- Castanheira, M.; Bell, J.M.; Turnidge, J.D.; Mathai, D.; Jones, R.N. Carbapenem resistance among Pseudomonas aeruginosa strains from India: Evidence for nationwide endemicity of multiple metallo-β-lactamase clones (VIM-2, -5, -6, and -11 and the newly characterized VIM-18). Antimicrob. Agents Chemother. 2009, 53, 1225–1227. [Google Scholar] [CrossRef]
- Tada, T.; Miyoshi-Akiyama, T.; Shimada, K.; Shimojima, M.; Kirikae, T. Novel 6′-N-aminoglycoside acetyltransferase AAC(6′)-Iaj from a clinical isolate of Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2013, 57, 96–100. [Google Scholar] [CrossRef] [PubMed]
- Maurya, A.P.; Talukdar, A.D.; Chanda, D.D.; Chakravarty, A.; Bhattacharjee, A. Integron-borne transmission of VEB-1 extended-spectrum β-lactamase in Pseudomonas aeruginosa in a tertiary care hospital in India. Antimicrob. Agents Chemother. 2014, 58, 6966–6969. [Google Scholar] [CrossRef]
- Bonnin, R.A.; Bogaerts, P.; Girlich, D.; Huang, T.D.; Dortet, L.; Glupczynski, Y.; Naas, T. Molecular Characterization of OXA-198 Carbapenemase-producing Pseudomonas aeruginosa clinical isolates. Antimicrob. Agents Chemother. 2018, 62, e02496-17. [Google Scholar] [CrossRef]
- Martínez, T.; Vazquez, G.J.; Aquino, E.E.; Goering, R.V.; Robledo, I.E. Two novel class I integron arrays containing IMP-18 metallo-β-lactamase gene in Pseudomonas aeruginosa clinical isolates from Puerto Rico. Antimicrob. Agents Chemother. 2012, 56, 2119–2121. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Lambert, T.; Türkoglü, S.; Ronco, E.; Gaillard, J.; Nordmann, P. Characterization of Class 1 integrons from Pseudomonas aeruginosa that contain the blaVIM-2 carbapenem-hydrolyzing β-lactamase gene and of two novel aminoglycoside resistance gene cassettes. Antimicrob. Agents Chemother. 2001, 45, 546–552. [Google Scholar] [CrossRef]
- Mendes, R.E.; Toleman, M.A.; Ribeiro, J.; Sader, H.S.; Jones, R.N.; Walsh, T.R. Integron carrying a novel metallo-β-lactamase gene, blaIMP-16, and a fused form of aminoglycoside-resistant gene aac(6′)-30/aac(6′)-Ib′: Report from the SENTRY antimicrobial surveillance program. Antimicrob. Agents Chemother. 2004, 48, 4693–4702. [Google Scholar] [CrossRef] [PubMed]
- Quinteira, S.; Sousa, J.C.; Peixe, L. Characterization of In100, a new integron carrying a metallo-β-lactamase and a carbenicillinase, from Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2005, 49, 451–453. [Google Scholar] [CrossRef]
- Sekiguchi, J.; Asagi, T.; Miyoshi-Akiyama, T.; Fujino, T.; Kobayashi, I.; Morita, K.; Kikuchi, Y.; Kuratsuji, T.; Kirikae, T. Multidrug-resistant Pseudomonas aeruginosa strain that caused an outbreak in a neurosurgery ward and its aac(6′)-Iae gene cassette encoding a novel aminoglycoside acetyltransferase. Antimicrob. Agents Chemother. 2005, 49, 3734–3742. [Google Scholar] [CrossRef] [PubMed]
- Villegas, M.V.; Lolans, K.; del Rosario Olivera, M.; Suarez, C.J.; Correa, A.; Queenan, A.M.; Quinn, J.P. First detection of metallo-β-lactamase VIM-2 in Pseudomonas aeruginosa isolates from Colombia. Antimicrob. Agents Chemother. 2006, 50, 226–229. [Google Scholar] [CrossRef] [PubMed]
- Naas, T.; Aubert, D.; Lambert, T.; Nordmann, P. Complex genetic structures with repeated elements, a sul-type class 1 integron, and the blaVEB extended-spectrum β-lactamase gene. Antimicrob. Agents Chemother. 2006, 50, 1745–1752. [Google Scholar] [CrossRef]
- Pagani, L.; Colinon, C.; Migliavacca, R.; Labonia, M.; Docquier, J.D.; Nucleo, E.; Spalla, M.; Li Bergoli, M.; Rossolini, G.M. Nosocomial outbreak caused by multidrug-resistant Pseudomonas aeruginosa producing IMP-13 metallo-β-lactamase. J. Clin. Microbiol. 2005, 43, 3824–3828. [Google Scholar] [CrossRef]
- Rao, J.; Adenikinju, A.; Kerkering, T.M.; Garner, D.C.; Jensen, R.V. Complete genome sequence of Pseudomonas aeruginosa CMC-097, isolated from a ventilator-associated pneumonia patient, containing a novel carbapenem resistance class 1 integron. Microbiol. Resour. Announc. 2021, 10, e0077421. [Google Scholar] [CrossRef] [PubMed]
- Krasauskas, R.; Labeikytė, D.; Markuckas, A.; Povilonis, J.; Armalytė, J.; Plančiūnienė, R.; Kavaliauskas, P.; Sužiedėlienė, E. Purification and characterization of a new β-lactamase OXA-205 from Pseudomonas aeruginosa. Ann. Clin. Microbiol. Antimicrob. 2015, 14, 52. [Google Scholar] [CrossRef]
- Paul, D.; Dhar, D.; Maurya, A.P.; Mishra, S.; Sharma, G.D.; Chakravarty, A.; Bhattacharjee, A. Occurrence of co-existing blaVIM-2 and blaNDM-1 in clinical isolates of Pseudomonas aeruginosa from India. Ann. Clin. Microbiol. Antimicrob. 2016, 15, 31. [Google Scholar] [CrossRef] [PubMed]
- Meradji, S.; Barguigua, A.; Zerouali, K.; Mazouz, D.; Chettibi, H.; Elmdaghri, N.; Timinouni, M. Epidemiology of carbapenem non-susceptible Pseudomonas aeruginosa isolates in Eastern Algeria. Antimicrob. Resist. Infect. Control 2015, 4, 27. [Google Scholar] [CrossRef]
- Fang, Y.; Baloch, Z.; Zhang, W.; Hu, Y.; Zheng, R.; Song, Y.; Tai, W.; Xia, X. Emergence of carbapenem-resistant ST244, ST292, and ST2446 Pseudomonas aeruginosa clones in burn patients in Yunnan province. Infect. Drug Resist. 2022, 15, 1103–1114. [Google Scholar] [CrossRef]
- van der Zee, A.; Kraak, W.B.; Burggraaf, A.; Goessens, W.H.F.; Pirovano, W.; Ossewaarde, J.M.; Tommassen, J. Spread of carbapenem resistance by transposition and conjugation among Pseudomonas aeruginosa. Front. Microbiol. 2018, 9, 2057. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.H.; Kwon, G.C.; Kim, S.; Koo, S.H. Distribution of Pseudomonas-derived cephalosporinase and metallo-β-lactamases in carbapenem-resistant Pseudomonas aeruginosa isolates from Korea. J. Microbiol. Biotechnol. 2015, 25, 1154–1162. [Google Scholar] [CrossRef]
- Mohanam, L.; Menon, T. Coexistence of metallo-β-lactamase-encoding genes in Pseudomonas aeruginosa. Indian J. Med. Res. 2017, 146, S46–S52. [Google Scholar]
- Elena, A.; Quinteros, M.; Di Conza, J.; Gutkind, G.; Cejas, D.; Radice, M.A. Full characterization of an IncR plasmid harboring qnrS1 recovered from a VIM-11-producing Pseudomonas aeruginosa. Rev. Argent. Microbiol. 2020, 52, 298–304. [Google Scholar] [CrossRef]
- Cejas, D.; Elena, A.; González-Espinosa, F.E.; Pallecchi, L.; Vay, C.; Rossolini, G.M.; Gutkind, G.; Di Pilato, V.; Radice, M. Characterisation of blaKPC-2-harbouring plasmids recovered from Pseudomonas aeruginosa ST654 and ST235 high-risk clones. J. Glob. Antimicrob. Resist. 2022, 29, 310–312. [Google Scholar] [CrossRef]
- Zhang, B.; Xu, X.; Song, X.; Wen, Y.; Zhu, Z.; Lv, J.; Xie, X.; Chen, L.; Tang, Y.W.; Du, H. Emerging and re-emerging KPC-producing hypervirulent Pseudomonas aeruginosa ST697 and ST463 between 2010 and 2021. Emerg. Microbes Infect. 2022, 11, 2735–2745. [Google Scholar] [CrossRef]
- Wang, L.J.; Chen, E.Z.; Yang, L.; Feng, D.H.; Xu, Z.; Chen, D.Q. Emergence of clinical Pseudomonas aeruginosa isolate Guangzhou-PaeC79 carrying crpP, blaGES-5, and blaKPC-2 in Guangzhou of China. Microb. Drug Resist. 2021, 27, 965–970. [Google Scholar] [CrossRef]
- Chanawong, A.; M’Zali, F.H.; Heritage, J.; Lulitanond, A.; Hawkey, P.M. SHV-12, SHV-5, SHV-2a and VEB-1 extended-spectrum β-lactamases in Gram-negative bacteria isolated in a university hospital in Thailand. J. Antimicrob. Chemother. 2001, 48, 839–852. [Google Scholar] [CrossRef]
- Li, Z.; Cai, Z.; Cai, Z.; Zhang, Y.; Fu, T.; Jin, Y.; Cheng, Z.; Jin, S.; Wu, W.; Yang, L.; et al. Molecular genetic analysis of an XDR Pseudomonas aeruginosa ST664 clone carrying multiple conjugal plasmids. J. Antimicrob. Chemother. 2020, 75, 1443–1452. [Google Scholar] [CrossRef] [PubMed]
- Naas, T.; Bonnin, R.A.; Cuzon, G.; Villegas, M.V.; Nordmann, P. Complete sequence of two KPC-harbouring plasmids from Pseudomonas aeruginosa. J. Antimicrob. Chemother. 2013, 68, 1757–1762. [Google Scholar] [CrossRef]
- Hagemann, J.B.; Pfennigwerth, N.; Gatermann, S.G.; von Baum, H.; Essig, A. KPC-2 carbapenemase-producing Pseudomonas aeruginosa reaching Germany. J. Antimicrob. Chemother. 2018, 73, 1812–1814. [Google Scholar] [CrossRef] [PubMed]
- Olalekan, A.; Bader, B.K.; Iwalokun, B.; Wolf, S.; Lalremruata, A.; Dike, A.; Mannie-Udoh, M.; Lo Presti, L.; Liese, J.; Guther, J.; et al. High incidence of carbapenemase-producing Pseudomonas aeruginosa clinical isolates from Lagos, Nigeria. JAC Antimicrob. Resist. 2023, 5, dlad038. [Google Scholar] [CrossRef] [PubMed]
- Rajaee Behbahani, M.; Keshavarzi, A.; Pirbonyeh, N.; Javanmardi, F.; Khoob, F.; Emami, A. Plasmid-related β-lactamase genes in Pseudomonas aeruginosa isolates: A molecular study in burn patients. J. Med. Microbiol. 2019, 68, 1740–1746. [Google Scholar] [PubMed]
- Pincus, N.B.; Rosas-Lemus, M.; Gatesy, S.W.M.; Bertucci, H.K.; Brunzelle, J.S.; Minasov, G.; Shuvalova, L.A.; Lebrun-Corbin, M.; Satchell, K.J.F.; Ozer, E.A.; et al. Functional and structural characterization of OXA-935, a novel OXA-10-family β-lactamase from Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2022, 66, e0098522. [Google Scholar] [CrossRef]
- Schlüter, A.; Szczepanowski, R.; Kurz, N.; Schneiker, S.; Krahn, I.; Pühler, A. Erythromycin resistance-conferring plasmid pRSB105, isolated from a sewage treatment plant, harbors a new macrolide resistance determinant, an integron-containing Tn402-like element, and a large region of unknown function. Appl. Environ. Microbiol. 2007, 73, 1952–1960. [Google Scholar] [CrossRef] [PubMed]
- Dong, N.; Liu, C.; Hu, Y.; Lu, J.; Zeng, Y.; Chen, G.; Chen, S.; Zhang, R. Emergence of an extensive drug resistant Pseudomonas aeruginosa strain of chicken origin carrying blaIMP-45, tet(X6), and tmexCD3-toprJ3 on an IncpRBL16 Plasmid. Microbiol. Spectr. 2022, 10, e0228322. [Google Scholar] [CrossRef] [PubMed]
- Ge, H.; Qiao, J.; Zheng, J.; Xu, H.; Liu, R.; Zhao, J.; Chen, R.; Li, C.; Guo, X.; Zheng, B. Emergence and clonal dissemination of KPC-3-producing Pseudomonas aeruginosa in China with an IncP-2 megaplasmid. Ann. Clin. Microbiol. Antimicrob. 2023, 22, 31. [Google Scholar] [CrossRef] [PubMed]
- Bi, D.; Xie, Y.; Tai, C.; Jiang, X.; Zhang, J.; Harrison, E.M.; Jia, S.; Deng, Z.; Rajakumar, K.; Ou, H.Y. A site-specific integrative plasmid found in Pseudomonas aeruginosa clinical isolate HS87 along with a plasmid carrying an aminoglycoside-resistant gene. PLoS One 2016, 11, e0148367. [Google Scholar] [CrossRef] [PubMed]
- Subedi, D.; Kohli, G.S.; Vijay, A.K.; Willcox, M.; Rice, S.A. Accessory genome of the multi-drug resistant ocular isolate of Pseudomonas aeruginosa PA34. PLoS One 2019, 14, e0215038. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Xu, T.; Ying, J.; Zhou, W.; Chen, Q.; Qian, C.; Zhu, X.; Shen, K.; Li, P.; Li, K.; et al. PAU-1, a novel plasmid-encoded Ambler class A β-Lactamase identified in a clinical Pseudomonas aeruginosa isolate. Infect. Drug Resist. 2019, 12, 3827–3834. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Zhu, Y.; Hu, D.; Li, Y.; Leptihn, S.; Loh, B.; Hua, X.; Yu, Y. Co-harboring of novel blaKPC-2 plasmid and integrative and conjugative element carrying Tn6203 in multidrug-resistant Pseudomonas aeruginosa. Front. Microbiol. 2021, 12, 674974. [Google Scholar] [CrossRef]
- Patil, S.; Chen, X.; Dong, S.; Mai, H.; Lopes, B.S.; Liu, S.; Wen, F. Resistance genomics and molecular epidemiology of high-risk clones of ESBL-producing Pseudomonas aeruginosa in young children. Front. Cell. Infect. Microbiol. 2023, 13, 1168096. [Google Scholar] [CrossRef]
- Woegerbauer, M.; Kuffner, M.; Domingues, S.; Nielsen, K.M. Involvement of aph(3′)-IIa in the formation of mosaic aminoglycoside resistance genes in natural environments. Front. Microbiol. 2015, 6, 442. [Google Scholar] [CrossRef]
- Sun, F.; Zhou, D.; Wang, Q.; Feng, J.; Feng, W.; Luo, W.; Zhang, D.; Liu, Y.; Qiu, X.; Yin, Z.; et al. The first report of detecting the blaSIM-2 gene and determining the complete sequence of the SIM-encoding plasmid. Clin. Microbiol. Infect. 2016, 22, 347–351. [Google Scholar] [CrossRef]
- Yang, Q.; Li, Y.; Fang, L.; Lei, T.; Cai, H.; Hua, X.; Zheng, M.; Yu, Y. A novel KPC-113 variant conferring carbapenem and ceftazidime-avibactam resistance in a multidrug-resistant Pseudomonas aeruginosa isolate. Clin. Microbiol. Infect. 2023, 29, 387. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira Santos, I.C.; Albano, R.M.; Asensi, M.D.; D’Alincourt Carvalho-Assef, A.P. Draft genome sequence of KPC-2-producing Pseudomonas aeruginosa recovered from a bloodstream infection sample in Brazil. J. Glob. Antimicrob. Resist. 2018, 15, 99–100. [Google Scholar] [CrossRef]
- Tartari, D.C.; Zamparette, C.P.; Martini, G.; Christakis, S.; Costa, L.H.; Silveira, A.C.O.; Sincero, T.C.M. Genomic analysis of an extensively drug-resistant Pseudomonas aeruginosa ST312 harbouring IncU plasmid-mediated blaKPC-2 isolated from ascitic fluid. J. Glob. Antimicrob. Resist. 2021, 25, 151–153. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Tang, M.; Xu, Y.; Xu, M.; Qian, C.; Zheng, X.; Zhou, T.; Wu, Q. Characteristics of rare ST463 carbapenem-resistant Pseudomonas aeruginosa clinical isolates from blood. J. Glob. Antimicrob. Resist. 2023, 32, 122–130. [Google Scholar] [CrossRef] [PubMed]
- Silveira, M.C.; Albano, R.M.; Rocha-de-Souza, C.M.; Leão, R.S.; Marques, E.A.; Picão, R.C.; Kraychete, G.B.; de Oliveira Santos, I.C.; Oliveira, T.; Tavares-Teixeira, C.B.; et al. Description of a novel IncP plasmid harboring blaKPC-2 recovered from a SPM-1-producing Pseudomonas aeruginosa from ST277. Infect. Genet. Evol. 2022, 102, 105302. [Google Scholar] [CrossRef]
- Feng, W.; Zhou, D.; Wang, Q.; Luo, W.; Zhang, D.; Sun, Q.; Tong, Y.; Chen, W.; Sun, F.; Xia, P. Dissemination of IMP-4-encoding pIMP-HZ1-related plasmids among Klebsiella pneumoniae and Pseudomonas aeruginosa in a Chinese teaching hospital. Sci. Rep. 2016, 6, 33419. [Google Scholar] [CrossRef]
- Botelho, J.; Grosso, F.; Quinteira, S.; Brilhante, M.; Ramos, H.; Peixe, L. Two decades of blaVIM-2-producing Pseudomonas aeruginosa dissemination: An interplay between mobile genetic elements and successful clones. J. Antimicrob. Chemother. 2018, 73, 873–882. [Google Scholar] [CrossRef]
- Poirel, L.; Weldhagen, G.F.; Naas, T.; De Champs, C.; Dove, M.G.; Nordmann, P. GES-2, a class A β-lactamase from Pseudomonas aeruginosa with increased hydrolysis of imipenem. Antimicrob. Agents Chemother. 2001, 45, 2598–2603. [Google Scholar] [CrossRef]
- Hu, Y.; Liu, C.; Wang, Q.; Zeng, Y.; Sun, Q.; Shu, L.; Lu, J.; Cai, J.; Wang, S.; Zhang, R.; et al. Emergence and expansion of a carbapenem-resistant Pseudomonas aeruginosa clone are associated with plasmid-borne blaKPC-2 and virulence-related genes. mSystems 2021, 6, e00154-21. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Cai, H.; Li, Y.; Wang, N.; Zhang, P.; Hua, X.; Yu, Y.; Sun, R. Plasmid-borne AFM alleles in Pseudomonas aeruginosa clinical isolates from China. Microbiol. Spectr. 2022, 10, e0203522. [Google Scholar] [CrossRef] [PubMed]
- García, P.; Brito, B.; Alcalde-Rico, M.; Munita, J.M.; Martínez, J.R.W.; Olivares-Pacheco, J.; Quiroz, V.; Wozniak, A. Acquisition of resistance to ceftazidime-avibactam during infection treatment in Pseudomonas aeruginosa through D179Y mutation in one of two blaKPC-2 gene copies without losing carbapenem resistance. Front. Cell. Infect. Microbiol. 2022, 12, 981792. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Zhou, D.; Xiong, W.; Feng, J.; Luo, W.; Luo, G.; Wang, H.; Sun, F.; Zhou, X. The IncP-6 plasmid p10265-KPC from Pseudomonas aeruginosa carries a novel ΔISEc33-associated blaKPC-2 gene cluster. Front. Microbiol. 2016, 7, 310. [Google Scholar] [CrossRef] [PubMed]
- Galetti, R.; Andrade, L.N.; Varani, A.M.; Darini, A.L.C. A phage-like plasmid carrying blaKPC-2 Gene in carbapenem-resistant Pseudomonas aeruginosa. Front. Microbiol. 2019, 10, 572. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Guan, H.; Sha, D.; Cao, W.; Song, X.; Che, J.; Kan, B.; Li, J. Characterization of blaKPC-2-carrying plasmid pR31-KPC from a Pseudomonas aeruginosa strain isolated in China. Antibiotics 2021, 10, 1234. [Google Scholar] [CrossRef]
- Khajuria, A.; Praharaj, A.K.; Kumar, M.; Grover, N. Emergence of NDM-1 in the clinical isolates of Pseudomonas aeruginosa in India. J. Clin. Diagn. Res. 2013, 7, 1328–1331. [Google Scholar] [CrossRef]
- Shahid, M.; Malik, A. Resistance due to aminoglycoside modifying enzymes in Pseudomonas aeruginosa isolates from burns patients. Indian J. Med. Res. 2005, 122, 324–329. [Google Scholar] [PubMed]
- Khan, M.; Willcox, M.D.P.; Rice, S.A.; Sharma, S.; Stapleton, F. Development of antibiotic resistance in the ocular Pseudomonas aeruginosa clone ST308 over twenty years. Exp. Eye Res. 2021, 205, 108504. [Google Scholar] [CrossRef] [PubMed]
- Fortunato, G.; Vaz-Moreira, I.; Gajic, I.; Manaia, C.M. Insight into phylogenomic bias of blaVIM-2 or blaNDM-1 dissemination amongst carbapenem-resistant Pseudomonas aeruginosa. Int. J. Antimicrob. Agents 2023, 61, 106788. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; Summers, S.; Rice, S.A.; Stapleton, F.; Willcox, M.D.P.; Subedi, D. Acquired fluoroquinolone resistance genes in corneal isolates of Pseudomonas aeruginosa. Infect. Genet. Evol. 2020, 85, 104574. [Google Scholar] [CrossRef]
- Ruedas-López, A.; Alonso-García, I.; Lasarte-Monterrubio, C.; Guijarro-Sánchez, P.; Gato, E.; Vázquez-Ucha, J.C.; Vallejo, J.A.; Fraile-Ribot, P.A.; Fernández-Pérez, B.; Velasco, D.; et al. Selection of ampC β-Lactamase variants and metallo-β-lactamases leading to ceftolozane/tazobactam and ceftazidime/avibactam resistance during treatment of MDR/XDR Pseudomonas aeruginosa infections. Antimicrob. Agents Chemother. 2022, 66, e0206721. [Google Scholar] [CrossRef] [PubMed]
- Talat, A.; Blake, K.S.; Dantas, G.; Khan, A.U. Metagenomic insight into microbiome and antibiotic resistance genes of high clinical concern in urban and rural hospital wastewater of Northern India origin: A major reservoir of antimicrobial resistance. Microbiol. Spectr. 2023, 11, e0410222. [Google Scholar] [CrossRef]
- Yu, T.; Yang, H.; Li, J.; Chen, F.; Hu, L.; Jing, Y.; Luo, X.; Yin, Z.; Zou, M.; Zhou, D. Novel chromosome-borne accessory genetic elements carrying multiple antibiotic resistance genes in Pseudomonas aeruginosa. Front. Cell Infect. Microbiol. 2021, 11, 638087. [Google Scholar] [CrossRef] [PubMed]
- Papa-Ezdra, R.; Cordeiro, N.F.; Outeda, M.; Garcia-Fulgueiras, V.; Araújo, L.; Seija, V.; Ayala, J.A.; Bado, I.; Vignoli, R. Novel resistance regions carrying TnaphA6, blaVIM-2, and blaPER-1, embedded in an ISPa40-derived transposon from two multi-resistant Pseudomonas aeruginosa clinical isolates. Antibiotics 2023, 12, 304. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Yang, H.; Yin, Z.; Jing, Y.; Zhao, Y.; Fu, H.; Du, H.; Zhou, D. Diversification and prevalence of the quinolone resistance crpP genes and the crpP-carrying Tn6786-related integrative and conjugative elements in Pseudomonas aeruginosa. Virulence 2021, 12, 2162–2170. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, L.; Li, D.; Wang, C.; Guo, Q.; Wang, M. Characterization of the novel plasmid-encoded MBL gene blaAFM-1, integrated into a blaIMP-45-bearing transposon Tn6485e in a carbapenem-resistant Pseudomonas aeruginosa clinical isolate. J. Antimicrob. Chemother. 2021, 77, 83–88. [Google Scholar] [CrossRef]
- Bour, M.; Fournier, D.; Jové, T.; Pouzol, A.; Miltgen, G.; Janvier, F.; Jeannot, K.; Plésiat, P. Acquisition of class C β-lactamase PAC-1 by ST664 strains of Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2019, 63, e01375-19. [Google Scholar] [CrossRef]
- Sun, Y.; Han, R.; Ding, L.; Yang, Y.; Guo, Y.; Wu, S.; Hu, F.; Yin, D. First report of blaOXA-677 with enhanced meropenem-hydrolyzing ability in Pseudomonas aeruginosa in China. Infect. Drug Resist. 2021, 14, 5725–5733. [Google Scholar] [CrossRef]
- Wołkowicz, T.; Patzer, J.A.; Kamińska, W.; Gierczyński, R.; Dzierżanowska, D. Distribution of carbapenem resistance mechanisms in Pseudomonas aeruginosa isolates among hospitalised children in Poland: Characterisation of two novel insertion sequences disrupting the oprD gene. J. Glob. Antimicrob. Resist. 2016, 7, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Subedi, D.; Vijay, A.K.; Kohli, G.S.; Rice, S.A.; Willcox, M. Comparative genomics of clinical strains of Pseudomonas aeruginosa strains isolated from different geographic sites. Sci. Rep. 2018, 8, 15668. [Google Scholar] [CrossRef]
- Vincent, A.T.; Freschi, L.; Jeukens, J.; Kukavica-Ibrulj, I.; Emond-Rheault, J.G.; Leduc, A.; Boyle, B.; Jean-Pierre, F.; Groleau, M.C.; Déziel, E.; et al. Genomic characterisation of environmental Pseudomonas aeruginosa isolated from dental unit waterlines revealed the insertion sequence ISPa11 as a chaotropic element. FEMS Microbiol. Ecol. 2017, 93, fix106. [Google Scholar]
- Sentausa, E.; Basso, P.; Berry, A.; Adrait, A.; Bellement, G.; Couté, Y.; Lory, S.; Elsen, S.; Attrée, I. Insertion sequences drive the emergence of a highly adapted human pathogen. Microb. Genom. 2020, 6, mgen000265. [Google Scholar] [CrossRef] [PubMed]
- Fowler, R.C.; Hanson, N.D. Emergence of carbapenem resistance due to the novel insertion sequence ISPa8 in Pseudomonas aeruginosa. PLoS One 2014, 9, e91299. [Google Scholar] [CrossRef] [PubMed]
- Diene, S.M.; L’Homme, T.; Bellulo, S.; Stremler, N.; Dubus, J.C.; Mely, L.; Leroy, S.; Degand, N.; Rolain, J.M. ISPa46, a novel insertion sequence in the oprD porin gene of an imipenem-resistant Pseudomonas aeruginosa isolate from a cystic fibrosis patient in Marseille, France. Int. J. Antimicrob. Agents 2013, 42, 268–271. [Google Scholar] [CrossRef]
- Sun, Q.; Ba, Z.; Wu, G.; Wang, W.; Lin, S.; Yang, H. Insertion sequence ISRP10 inactivation of the oprD gene in imipenem-resistant Pseudomonas aeruginosa clinical isolates. Int. J. Antimicrob. Agents 2016, 47, 375–379. [Google Scholar] [CrossRef]
- Bocharova, Y.; Savinova, T.; Shagin, D.A.; Shelenkov, A.A.; Mayanskiy, N.A.; Chebotar, I.V. Inactivation of the oprD porin gene by a novel insertion sequence ISPa195 associated with large deletion in a carbapenem-resistant Pseudomonas aeruginosa clinical isolate. J. Glob. Antimicrob. Resist. 2019, 17, 309–311. [Google Scholar] [CrossRef]
- Al-Bayssari, C.; Valentini, C.; Gomez, C.; Reynaud-Gaubert, M.; Rolain, J.M. First detection of insertion sequence element ISPa1328 in the oprD porin gene of an imipenem-resistant Pseudomonas aeruginosa isolate from an idiopathic pulmonary fibrosis patient in Marseille, France. New Microbes New Infect. 2015, 7, 26–27. [Google Scholar] [CrossRef] [PubMed]
- Shariati, A.; Azimi, T.; Ardebili, A.; Chirani, A.S.; Bahramian, A.; Pormohammad, A.; Sadredinamin, M.; Erfanimanesh, S.; Bostanghadiri, N.; Shams, S.; et al. Insertional inactivation of oprD in carbapenem-resistant Pseudomonas aeruginosa strains isolated from burn patients in Tehran, Iran. New Microbes New Infect. 2018, 21, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Boutoille, D.; Corvec, S.; Caroff, N.; Giraudeau, C.; Espaze, E.; Caillon, J.; Plésiat, P.; Reynaud, A. Detection of an IS21 insertion sequence in the mexR gene of Pseudomonas aeruginosa increasing β-lactam resistance. FEMS Microbiol. Lett. 2004, 230, 143–146. [Google Scholar] [CrossRef] [PubMed]
- Evans, J.C.; Segal, H. A novel insertion sequence, ISPA26, in oprD of Pseudomonas aeruginosa is associated with carbapenem resistance. Antimicrob. Agents Chemother. 2007, 51, 3776–3777. [Google Scholar] [CrossRef]
- Ruiz-Martínez, L.; López-Jiménez, L.; d’Ostuni, V.; Fusté, E.; Vinuesa, T.; Viñas, M. A mechanism of carbapenem resistance due to a new insertion element (ISPa133) in Pseudomonas aeruginosa. Int. Microbiol. 2011, 14, 51–58. [Google Scholar] [PubMed]
- Yin, S.; Chen, P.; You, B.; Zhang, Y.; Jiang, B.; Huang, G.; Yang, Z.; Chen, Y.; Chen, J.; Yuan, Z.; et al. Molecular typing and carbapenem resistance mechanisms of Pseudomonas aeruginosa isolated from a chinese burn center from 2011 to 2016. Front. Microbiol. 2018, 9, 1135. [Google Scholar] [CrossRef] [PubMed]







Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, J.H.; Kim, N.-H.; Jang, K.-M.; Jin, H.; Shin, K.; Jeong, B.C.; Kim, D.-W.; Lee, S.H. Prioritization of Critical Factors for Surveillance of the Dissemination of Antibiotic Resistance in Pseudomonas aeruginosa: A Systematic Review. Int. J. Mol. Sci. 2023, 24, 15209. https://doi.org/10.3390/ijms242015209
Lee JH, Kim N-H, Jang K-M, Jin H, Shin K, Jeong BC, Kim D-W, Lee SH. Prioritization of Critical Factors for Surveillance of the Dissemination of Antibiotic Resistance in Pseudomonas aeruginosa: A Systematic Review. International Journal of Molecular Sciences. 2023; 24(20):15209. https://doi.org/10.3390/ijms242015209
Chicago/Turabian StyleLee, Jung Hun, Nam-Hoon Kim, Kyung-Min Jang, Hyeonku Jin, Kyoungmin Shin, Byeong Chul Jeong, Dae-Wi Kim, and Sang Hee Lee. 2023. "Prioritization of Critical Factors for Surveillance of the Dissemination of Antibiotic Resistance in Pseudomonas aeruginosa: A Systematic Review" International Journal of Molecular Sciences 24, no. 20: 15209. https://doi.org/10.3390/ijms242015209