Structural Insight into Polymerase Mechanism via a Chiral Center Generated with a Single Selenium Atom
Abstract
:1. Introduction
2. Results
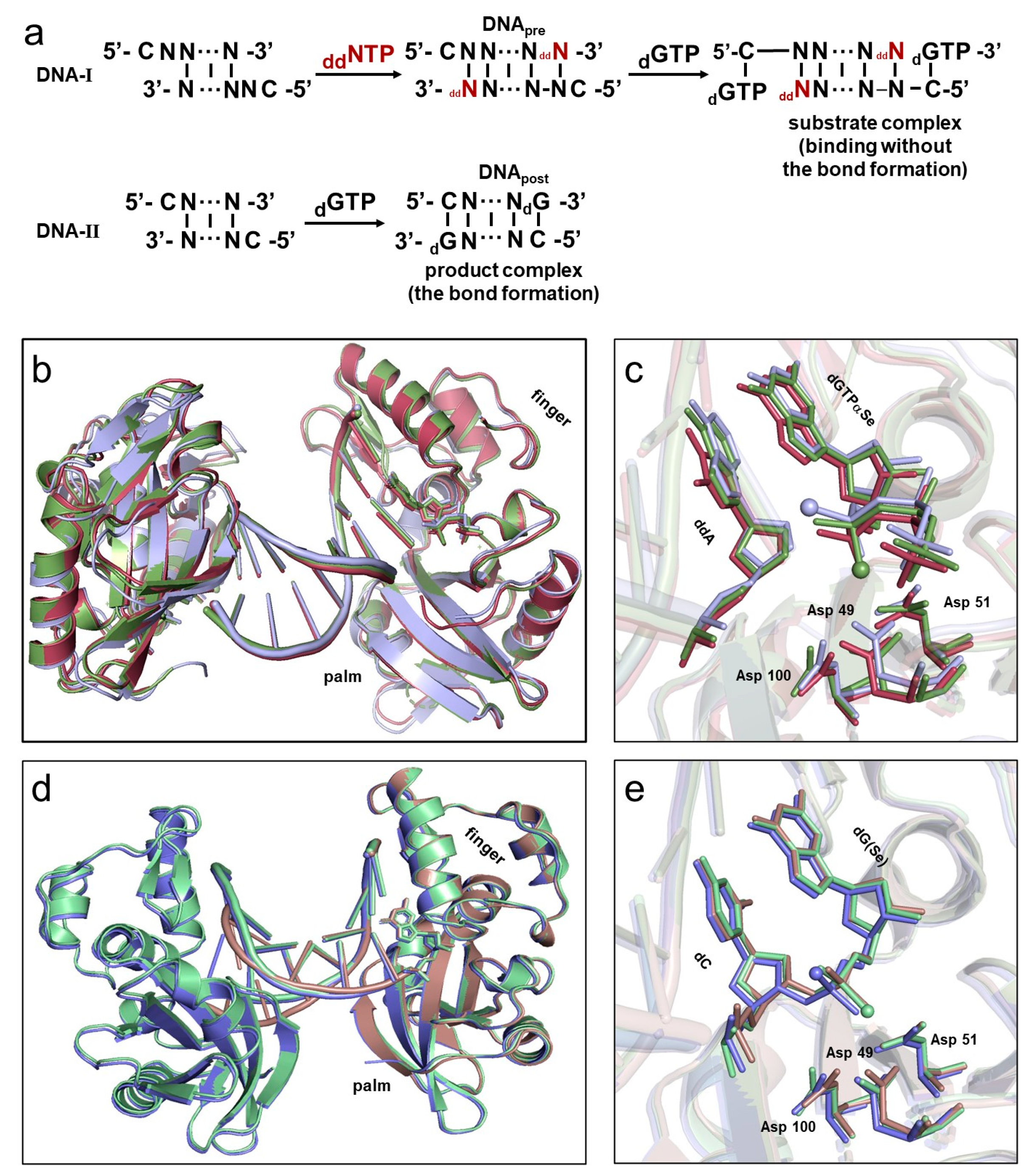
2.1. Overall Structures of the Complexes
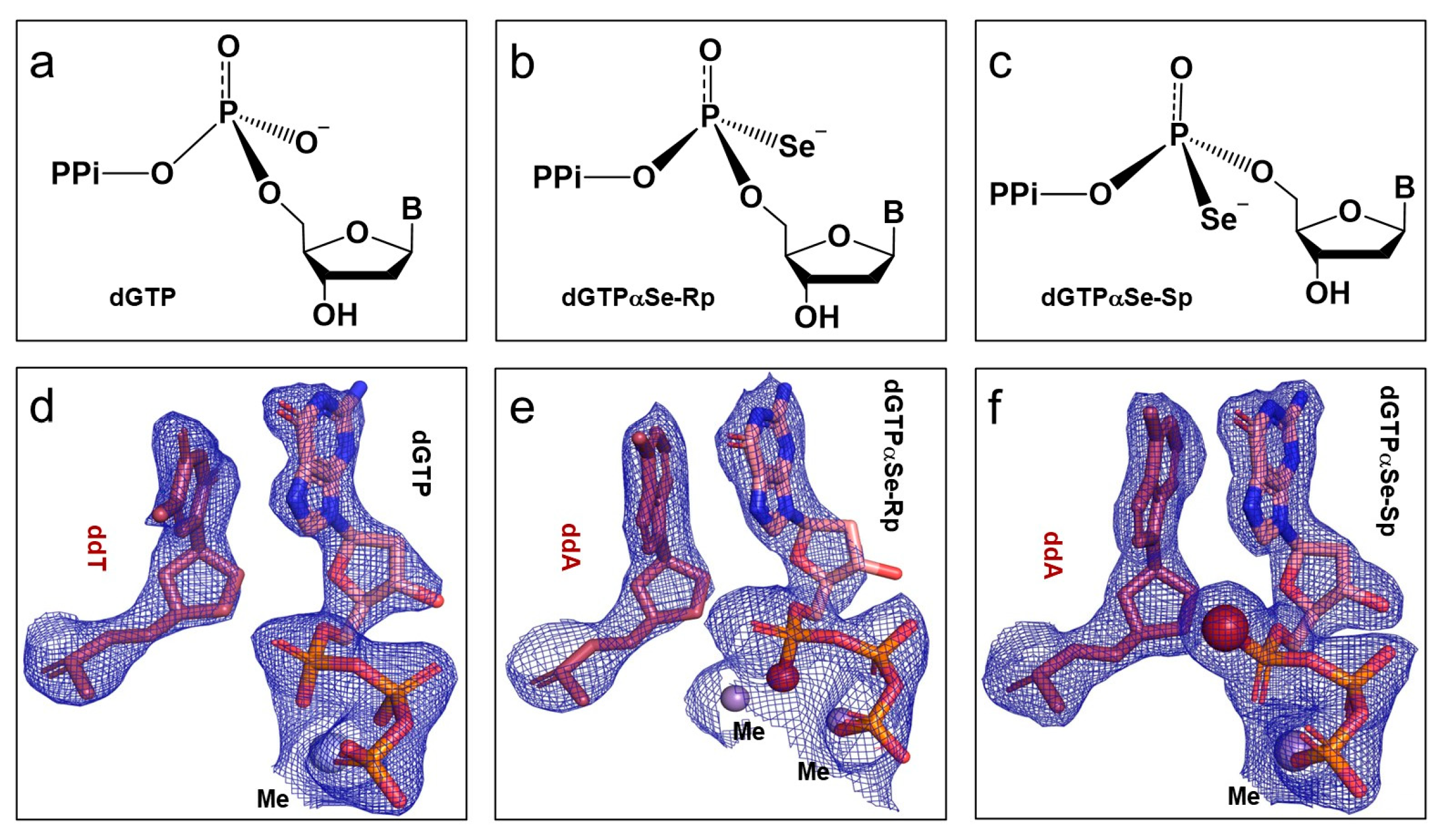
2.2. The Absolute Configurations of the Two dGTPαSe Diastereomers
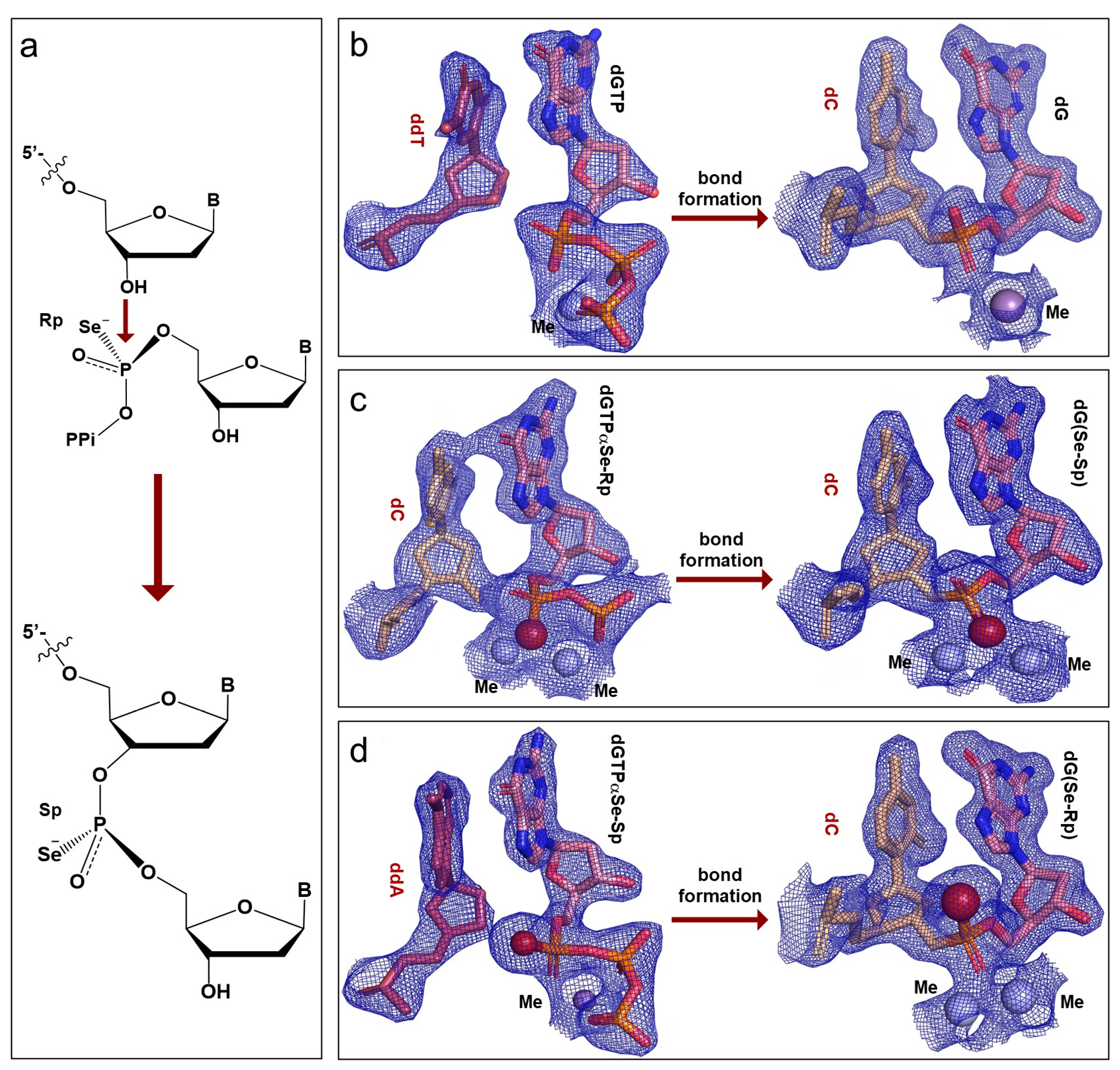
2.3. Configuration Inversion at the α-Phosphorus Center in the Polymerization Reaction
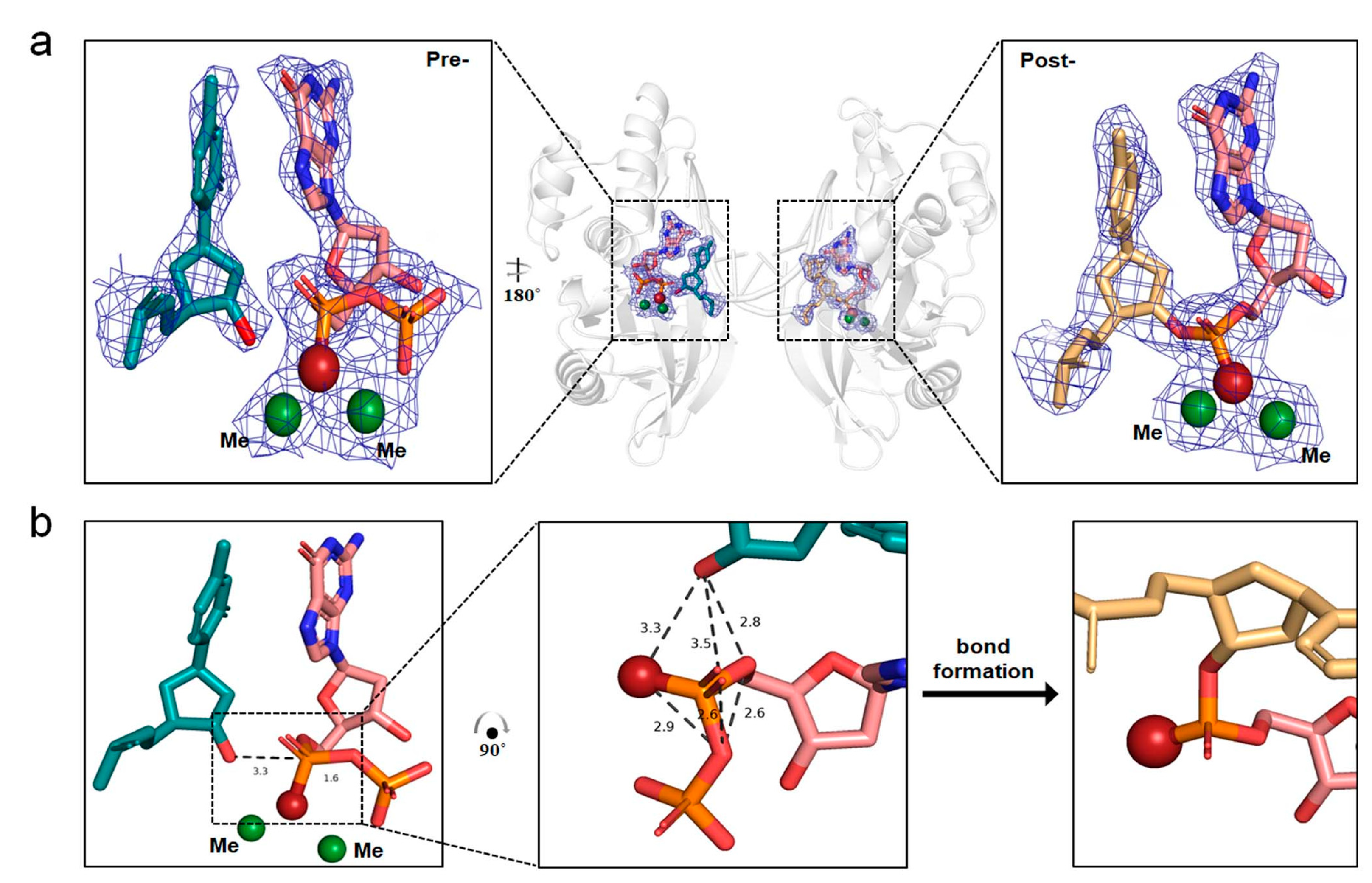
2.4. The In-Line SN2 Attacking Observed in One Single Crystal with dGTPαSe-Rp
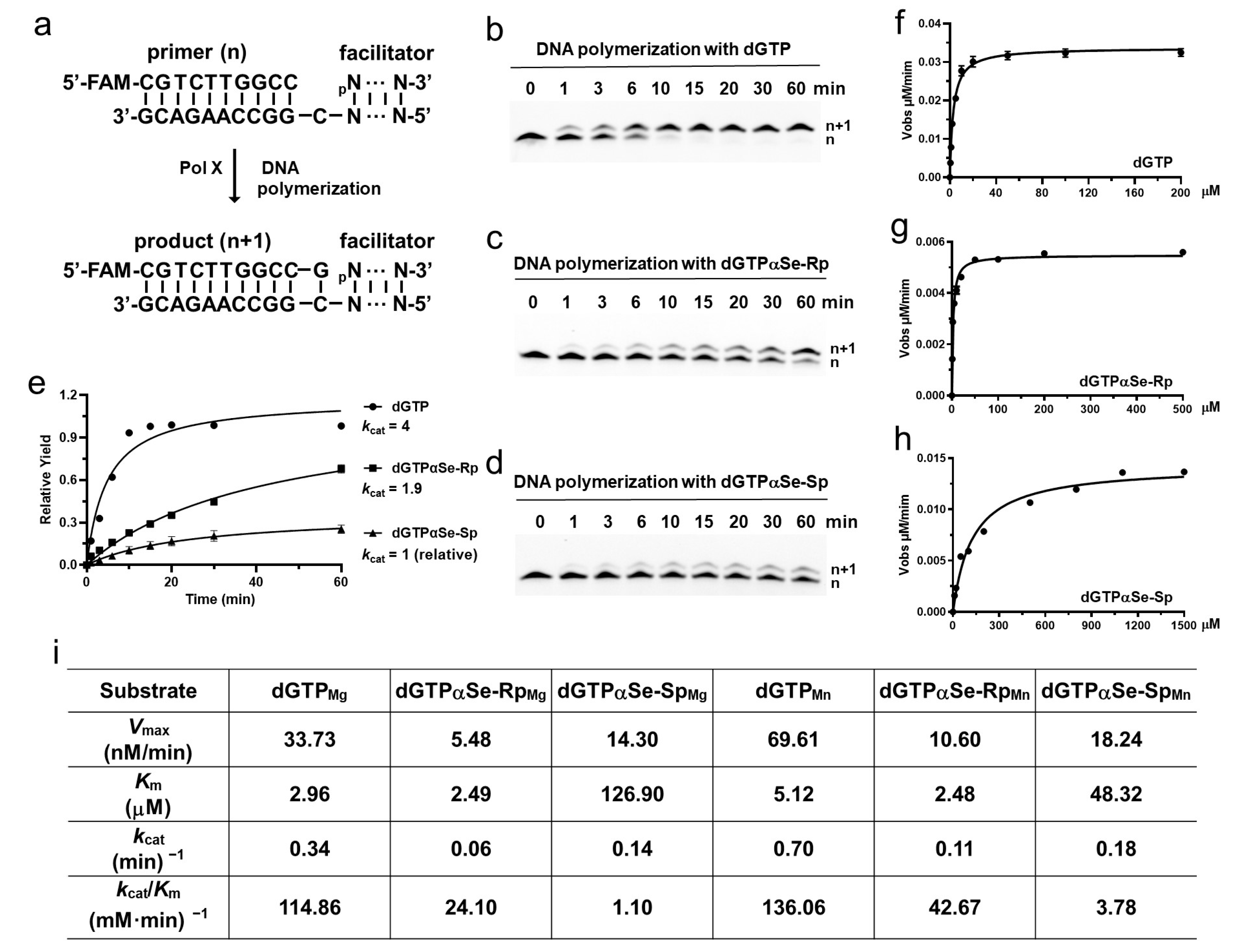
2.5. DNA Polymerase Recognizing dNTPαSe as the Substrates in Thermodynamic and Kinetic Studies
3. Discussion
4. Materials and Methods
4.1. Protein Expression and Purification
4.2. Crystallization and X-ray-Diffraction Data Collection
4.3. Structure Determination and Refinement
4.4. dNTP Binding Assay In Vitro
4.5. The Apparent and Standard Kinetic Study of Polymerase
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Guo, M.Z.; Marrone, K.A.; Spira, A.; Scott, S.C. Poly(ADP-Ribose) Polymerase Inhibition in Small Cell Lung Cancer: A Review. Cancer J. 2021, 27, 476–481. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.-J.; Yang, W.; Tsai, M.-D. How DNA polymerases catalyse replication and repair with contrasting fidelity. Nat. Rev. Chem. 2017, 1, 0068. [Google Scholar] [CrossRef]
- Bessman, M.J.; Kornberg, A.; Lehman, I.R.; Simms, E.S. Enzymic synthesis of deoxyribonucleic acid. Biochim. Biophys. Acta 1956, 21, 197–198. [Google Scholar] [CrossRef] [PubMed]
- Lehman, I.R.; Bessman, M.J.; Simms, E.S.; Kornberg, A. Enzymatic synthesis of deoxyribonucleic acid. I. Preparation of substrates and partial purification of an enzyme from Escherichia coli. J. Biol. Chem. 1958, 233, 163–170. [Google Scholar] [CrossRef]
- Ollis, D.L.; Brick, P.; Hamlin, R.; Xuong, N.G.; Steitz, T.A. Structure of large fragment of Escherichia coli DNA polymerase I complexed with dTMP. Nature 1985, 313, 762–766. [Google Scholar] [CrossRef]
- Ling, H.; Boudsocq, F.; Woodgate, R.; Yang, W. Crystal structure of a Y-family DNA polymerase in action: A mechanism for error-prone and lesion-bypass replication. Cell 2001, 107, 91–102. [Google Scholar] [CrossRef]
- Chen, Y.; Zhang, J.; Liu, H.; Gao, Y.; Li, X.; Zheng, L.; Cui, R.; Yao, Q.; Rong, L.; Li, J.; et al. Unique 5′-P recognition and basis for dG:dGTP misincorporation of ASFV DNA polymerase X. PLoS Biol. 2017, 15, e1002599. [Google Scholar] [CrossRef]
- Yang, W.; Gao, Y. Translesion and Repair DNA Polymerases: Diverse Structure and Mechanism. Annu. Rev. Biochem. 2018, 87, 239–261. [Google Scholar] [CrossRef]
- Garcia-Diaz, M.; Bebenek, K.; Krahn, J.M.; Kunkel, T.A.; Pedersen, L.C. A closed conformation for the Pol lambda catalytic cycle. Nat. Struct. Mol. Biol. 2005, 12, 97–98. [Google Scholar] [CrossRef]
- Garcia-Diaz, M.; Bebenek, K. Multiple functions of DNA polymerases. CRC Crit. Rev. Plant Sci. 2007, 26, 105–122. [Google Scholar] [CrossRef]
- Pelletier, H.; Sawaya, M.R.; Kumar, A.; Wilson, S.H.; Kraut, J. Structures of ternary complexes of rat DNA polymerase beta, a DNA template-primer, and ddCTP. Science 1994, 264, 1891–1903. [Google Scholar] [CrossRef] [PubMed]
- Doublié, S.; Sawaya, M.R.; Ellenberger, T. An open and closed case for all polymerases. Structure 1999, 7, R31–R35. [Google Scholar] [CrossRef] [PubMed]
- Batra, V.K.; Beard, W.A.; Shock, D.D.; Krahn, J.M.; Pedersen, L.C.; Wilson, S.H. Magnesium-induced assembly of a complete DNA polymerase catalytic complex. Structure 2006, 14, 757–766. [Google Scholar] [CrossRef] [PubMed]
- Mönttinen, H.A.; Ravantti, J.J.; Stuart, D.I.; Poranen, M.M. Automated structural comparisons clarify the phylogeny of the right-hand-shaped polymerases. Mol. Biol. Evol. 2014, 31, 2741–2752. [Google Scholar] [CrossRef]
- Yin, Y.W.; Steitz, T.A. The structural mechanism of translocation and helicase activity in T7 RNA polymerase. Cell 2004, 116, 393–404. [Google Scholar] [CrossRef]
- Westover, K.D.; Bushnell, D.A.; Kornberg, R.D. Structural basis of transcription: Nucleotide selection by rotation in the RNA polymerase II active center. Cell 2004, 119, 481–489. [Google Scholar] [CrossRef]
- Steitz, T.A. DNA polymerases: Structural diversity and common mechanisms. J. Biol. Chem. 1999, 274, 17395–17398. [Google Scholar] [CrossRef]
- Kuznetsova, A.A.; Fedorova, O.S.; Kuznetsov, N.A. Structural and Molecular Kinetic Features of Activities of DNA Polymerases. Int. J. Mol. Sci. 2022, 23, 6373. [Google Scholar] [CrossRef]
- Gao, Y.; Yang, W. Capture of a third Mg2+ is essential for catalyzing DNA synthesis. Science 2016, 352, 1334–1337. [Google Scholar] [CrossRef]
- Freudenthal, B.D.; Beard, W.A.; Perera, L.; Shock, D.D.; Kim, T.; Schlick, T.; Wilson, S.H. Uncovering the polymerase-induced cytotoxicity of an oxidized nucleotide. Nature 2015, 517, 635–639. [Google Scholar] [CrossRef]
- Nakamura, T.; Zhao, Y.; Yamagata, Y.; Hua, Y.J.; Yang, W. Watching DNA polymerase η make a phosphodiester bond. Nature 2012, 487, 196–201. [Google Scholar] [CrossRef] [PubMed]
- Steitz, T.A. DNA-Dependent and Rna-Dependent DNA-Polymerases. Curr. Opin. Struct. Biol. 1993, 3, 31–38. [Google Scholar] [CrossRef]
- Jamsen, J.A.; Beard, W.A.; Pedersen, L.C.; Shock, D.D.; Moon, A.F.; Krahn, J.M.; Bebenek, K.; Kunkel, T.A.; Wilson, S.H. Time-lapse crystallography snapshots of a double-strand break repair polymerase in action. Nat. Commun. 2017, 8, 253. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Weng, P.J.; Gao, Y. A new paradigm of DNA synthesis: Three-metal-ion catalysis. Cell Biosci. 2016, 6, 51. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Tsai, M.D. DNA polymerase beta: Pre-steady-state kinetic analyses of dATP alpha S stereoselectivity and alteration of the stereoselectivity by various metal ions and by site-directed mutagenesis. Biochemistry 2001, 40, 9014–9022. [Google Scholar] [CrossRef] [PubMed]
- Di Pasquale, F.; Fischer, D.; Grohmann, D.; Restle, T.; Geyer, A.; Marx, A. Opposed steric constraints in human DNA polymerase beta and E. coli DNA polymerase I. J. Am. Chem. Soc. 2008, 130, 10748–10757. [Google Scholar] [CrossRef]
- Obeid, S.; Busskamp, H.; Welte, W.; Diederichs, K.; Marx, A. Snapshot of a DNA polymerase while incorporating two consecutive C5-modified nucleotides. J. Am. Chem. Soc. 2013, 135, 15667–15669. [Google Scholar] [CrossRef]
- Raper, A.T.; Reed, A.J.; Suo, Z. Kinetic Mechanism of DNA Polymerases: Contributions of Conformational Dynamics and a Third Divalent Metal Ion. Chem. Rev. 2018, 118, 6000–6025. [Google Scholar] [CrossRef]
- Fiala, K.A.; Abdel-Gawad, W.; Suo, Z. Pre-steady-state kinetic studies of the fidelity and mechanism of polymerization catalyzed by truncated human DNA polymerase lambda. Biochemistry 2004, 43, 6751–6762. [Google Scholar] [CrossRef]
- Wang, J.; Konigsberg, W.H. Two-Metal-Ion Catalysis: Inhibition of DNA Polymerase Activity by a Third Divalent Metal Ion. Front. Mol. Biosci. 2022, 9, 824794. [Google Scholar] [CrossRef]
- Wang, J.; Smithline, Z.B. Crystallographic evidence for two-metal-ion catalysis in human pol eta. Protein Sci. 2019, 28, 439–447. [Google Scholar] [CrossRef]
- Maga, G.; Hubscher, U. Repair and translesion DNA polymerases as anticancer drug targets. Anticancer Agents Med. Chem. 2008, 8, 431–447. [Google Scholar] [CrossRef]
- Shah, N.; Mohammad, A.S.; Saralkar, P.; Sprowls, S.A.; Vickers, S.D.; John, D.; Tallman, R.M.; Lucke-Wold, B.P.; Jarrell, K.E.; Pinti, M.; et al. Investigational chemotherapy and novel pharmacokinetic mechanisms for the treatment of breast cancer brain metastases. Pharmacol. Res. 2018, 132, 47–68. [Google Scholar] [CrossRef]
- Jiang, Y.; Yin, W.; Xu, H.E. RNA-dependent RNA polymerase: Structure, mechanism, and drug discovery for COVID-19. Biochem. Biophys. Res. Commun. 2021, 538, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Miura, S.; Izuta, S. DNA polymerases as targets of anticancer nucleosides. Curr. Drug Targets 2004, 5, 191–195. [Google Scholar] [CrossRef] [PubMed]
- Jordheim, L.P.; Durantel, D.; Zoulim, F.; Dumontet, C. Advances in the development of nucleoside and nucleotide analogues for cancer and viral diseases. Nat. Rev. Drug Discov. 2013, 12, 447–464. [Google Scholar] [CrossRef] [PubMed]
- Abdur, R.; Gerlits, O.O.; Gan, J.; Jiang, J.; Salon, J.; Kovalevsky, A.Y.; Chumanevich, A.A.; Weber, I.T.; Huang, Z. Novel complex MAD phasing and RNase H structural insights using selenium oligonucleotides. Acta Crystallogr. D Biol. Crystallogr. 2014, 70, 354–361. [Google Scholar] [CrossRef] [PubMed]
- Sheng, J.; Gan, J.; Soares, A.S.; Salon, J.; Huang, Z. Structural insights of non-canonical U*U pair and Hoogsteen interaction probed with Se atom. Nucleic Acids Res. 2013, 41, 10476–10487. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Sheng, J.; Hassan, A.E.; Jiang, S.; Gan, J.; Huang, Z. Novel RNA base pair with higher specificity using single selenium atom. Nucleic Acids Res. 2012, 40, 5171–5179. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Sheng, J.; Huang, Z. Nucleic acid X-ray crystallography via direct selenium derivatization. Chem. Soc. Rev. 2011, 40, 4591–4602. [Google Scholar] [CrossRef]
- Showalter, A.K.; Byeon, I.J.; Su, M.I.; Tsai, M.D. Solution structure of a viral DNA polymerase X and evidence for a mutagenic function. Nat. Struct. Biol. 2001, 8, 942–946. [Google Scholar] [CrossRef] [PubMed]
- Oliveros, M.; Yanez, R.J.; Salas, M.L.; Salas, J.; Vinuela, E.; Blanco, L. Characterization of an African swine fever virus 20-kDa DNA polymerase involved in DNA repair. J. Biol. Chem. 1997, 272, 30899–30910. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Xie, M.; Wu, W.; Chen, Z. Structures and Functional Diversities of ASFV Proteins. Viruses 2021, 13, 2124. [Google Scholar] [CrossRef]
- Garcia-Escudero, R.; Garcia-Diaz, M.; Salas, M.L.; Blanco, L.; Salas, J. DNA polymerase X of African swine fever virus: Insertion fidelity on gapped DNA substrates and AP lyase activity support a role in base excision repair of viral DNA. J. Mol. Biol. 2003, 326, 1403–1412. [Google Scholar] [CrossRef]
- Hu, B.; Wang, Y.; Sun, S.; Yan, W.; Zhang, C.; Luo, D.; Deng, H.; Hu, L.R.; Huang, Z. Synthesis of Selenium-Triphosphates (dNTPalphaSe) for More Specific DNA Polymerization. Angew. Chem. Int. Ed. Engl. 2019, 58, 7835–7839. [Google Scholar] [CrossRef]
- Brody, R.S.; Frey, P.A. Unambiguous determination of the stereochemistry of nucleotidyl transfer catalyzed by DNA polymerase I from Escherichia coli. Biochemistry 1981, 20, 1245–1252. [Google Scholar] [CrossRef] [PubMed]
- Mizrahi, V.; Henrie, R.N.; Marlier, J.F.; Johnson, K.A.; Benkovic, S.J. Rate-limiting steps in the DNA polymerase I reaction pathway. Biochemistry 1985, 24, 4010–4018. [Google Scholar] [CrossRef] [PubMed]
- Dahlberg, M.E.; Benkovic, S.J. Kinetic mechanism of DNA polymerase I (Klenow fragment): Identification of a second conformational change and evaluation of the internal equilibrium constant. Biochemistry 1991, 30, 4835–4843. [Google Scholar] [CrossRef]
- Washington, M.T.; Prakash, L.; Prakash, S. Yeast DNA polymerase eta utilizes an induced-fit mechanism of nucleotide incorporation. Cell 2001, 107, 917–927. [Google Scholar] [CrossRef]
- Shaw, B.R.; Dobrikov, M.; Wang, X.; Wan, J.; He, K.; Lin, J.L.; Li, P.; Rait, V.; Sergueeva, Z.A.; Sergueev, D. Reading, writing, and modulating genetic information with boranophosphate mimics of nucleotides, DNA, and RNA. Ann. N. Y. Acad. Sci. 2003, 1002, 12–29. [Google Scholar] [CrossRef]
- Wilds, C.J.; Pattanayek, R.; Pan, C.; Wawrzak, Z.; Egli, M. Selenium-assisted nucleic acid crystallography: Use of phosphoroselenoates for MAD phasing of a DNA structure. J. Am. Chem. Soc. 2002, 124, 14910–14916. [Google Scholar] [CrossRef]
- Nakamura, T.; Zhao, Y.; Yamagata, Y.; Hua, Y.J.; Yang, W. Mechanism of the nucleotidyl-transfer reaction in DNA polymerase revealed by time-resolved protein crystallography. Biophysics Nagoya-shi 2013, 9, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Steitz, T.A.; Steitz, J.A. A general two-metal-ion mechanism for catalytic RNA. Proc. Natl. Acad. Sci. USA 1993, 90, 6498–6502. [Google Scholar] [CrossRef] [PubMed]
- Eckstein, F.; Armstrong, V.; Sternbach, H. Stereochemistry of polymerization by DNA-dependent RNA-polymerase from Escherichia coli: An investigation with a diastereomeric ATP-analogue. Proc. Natl. Acad. Sci. USA 1976, 73, 2987–2990. [Google Scholar] [CrossRef] [PubMed]
- Carrasco, N.; Huang, Z. Enzymatic synthesis of phosphoroselenoate DNA using thymidine 5′-(alpha-P-seleno)triphosphate and DNA polymerase for X-ray crystallography via MAD. J. Am. Chem. Soc. 2004, 126, 448–449. [Google Scholar] [CrossRef]
- Cowan, J.A. Metal Activation of Enzymes in Nucleic Acid Biochemistry. Chem. Rev. 1998, 98, 1067–1088. [Google Scholar] [CrossRef]
- Jamsen, J.A.; Sassa, A.; Shock, D.D.; Beard, W.A.; Wilson, S.H. Watching a double strand break repair polymerase insert a pro-mutagenic oxidized nucleotide. Nat. Commun. 2021, 12, 2059. [Google Scholar] [CrossRef]
- Saha, P.; Mandal, T.; Talukdar, A.D.; Kumar, D.; Kumar, S.; Tripathi, P.P.; Wang, Q.E.; Srivastava, A.K. DNA polymerase eta: A potential pharmacological target for cancer therapy. J. Cell. Physiol. 2021, 236, 4106–4120. [Google Scholar] [CrossRef]
- Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 125–132. [Google Scholar] [CrossRef]
- Emsley, P.; Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 2004, 60, 2126–2132. [Google Scholar] [CrossRef]
- Adams, P.D.; Afonine, P.V.; Bunkoczi, G.; Chen, V.B.; Davis, I.W.; Echols, N.; Headd, J.J.; Hung, L.W.; Kapral, G.J.; Grosse-Kunstleve, R.W.; et al. PHENIX: A comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 213–221. [Google Scholar] [CrossRef] [PubMed]
- Murshudov, G.N.; Skubak, P.; Lebedev, A.A.; Pannu, N.S.; Steiner, R.A.; Nicholls, R.A.; Winn, M.D.; Long, F.; Vagin, A.A. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 2011, 67, 355–367. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qin, T.; Hu, B.; Zhao, Q.; Wang, Y.; Wang, S.; Luo, D.; Lyu, J.; Chen, Y.; Gan, J.; Huang, Z. Structural Insight into Polymerase Mechanism via a Chiral Center Generated with a Single Selenium Atom. Int. J. Mol. Sci. 2023, 24, 15758. https://doi.org/10.3390/ijms242115758
Qin T, Hu B, Zhao Q, Wang Y, Wang S, Luo D, Lyu J, Chen Y, Gan J, Huang Z. Structural Insight into Polymerase Mechanism via a Chiral Center Generated with a Single Selenium Atom. International Journal of Molecular Sciences. 2023; 24(21):15758. https://doi.org/10.3390/ijms242115758
Chicago/Turabian StyleQin, Tong, Bei Hu, Qianwei Zhao, Yali Wang, Shaoxin Wang, Danyan Luo, Jiazhen Lyu, Yiqing Chen, Jianhua Gan, and Zhen Huang. 2023. "Structural Insight into Polymerase Mechanism via a Chiral Center Generated with a Single Selenium Atom" International Journal of Molecular Sciences 24, no. 21: 15758. https://doi.org/10.3390/ijms242115758




