Reference Genes Screening and Gene Expression Patterns Analysis Involved in Gelsenicine Biosynthesis under Different Hormone Treatments in Gelsemium elegans
Abstract
:1. Introduction
2. Results
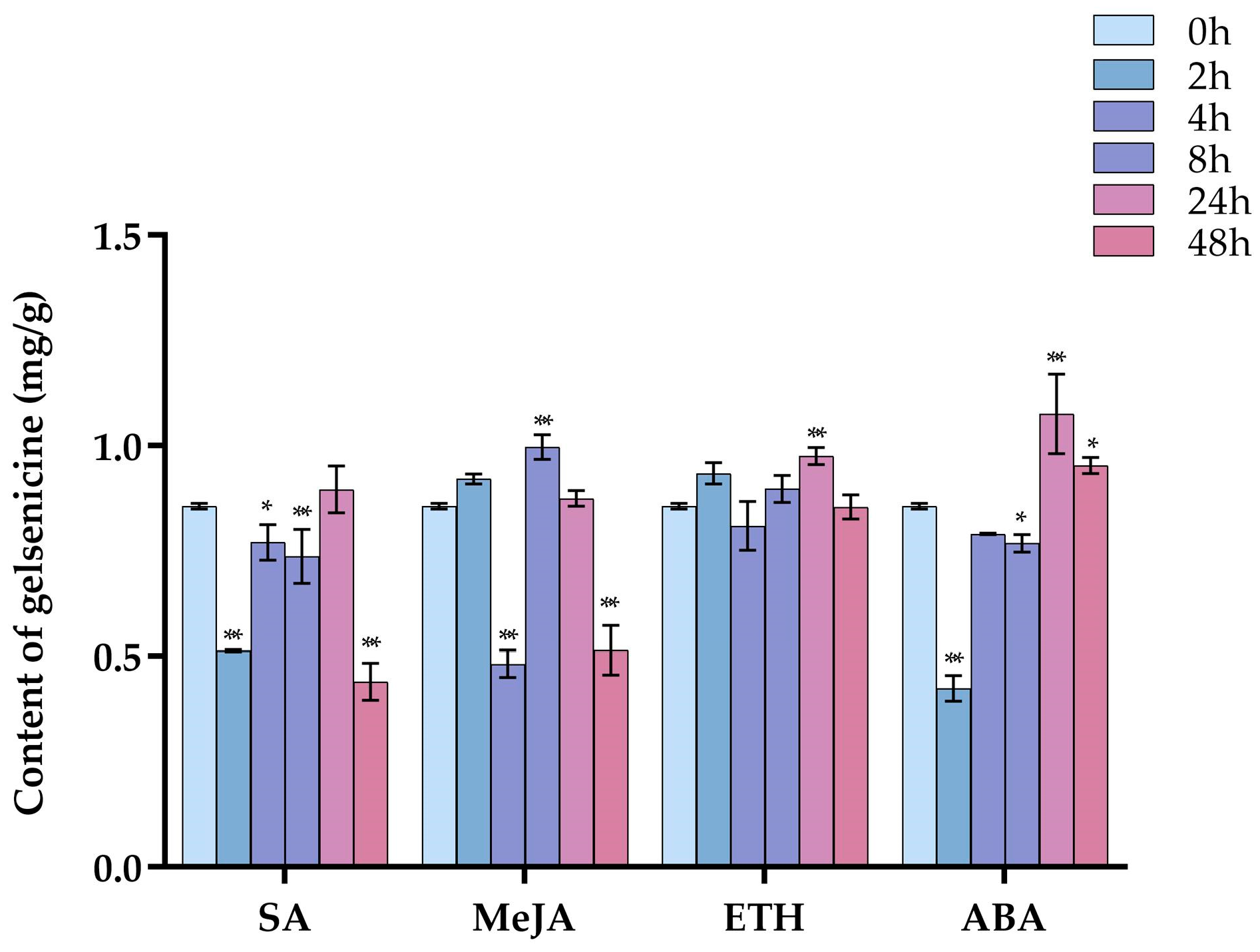
2.1. The Content of Gelsenicine under Four Hormone Treatments
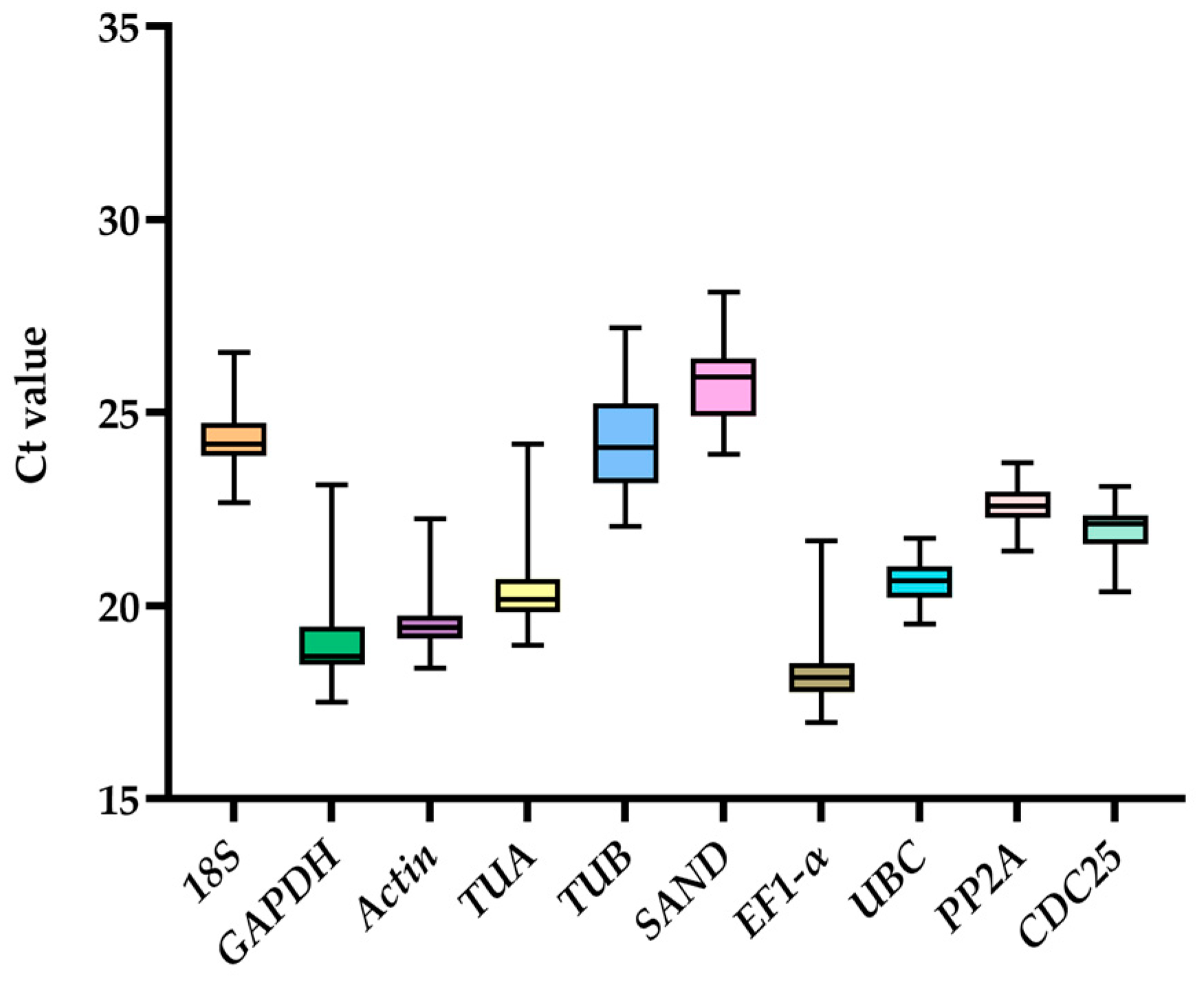
2.2. Amplification Specificity, Efficiency, and Expression Profile Analysis of Reference Genes in G. elegans
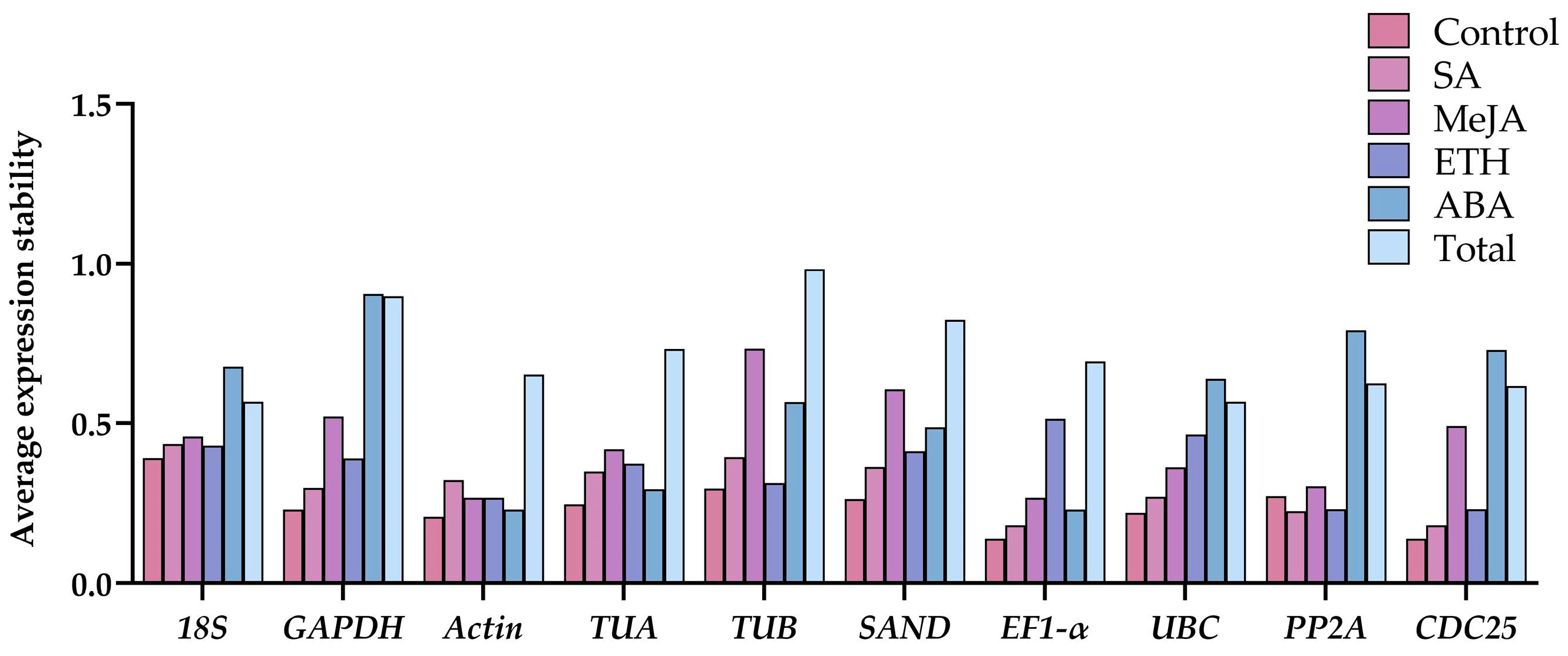
2.3. Stability Analysis of Candidate Reference Genes
2.3.1. GeNorm Analysis of Reference Genes in G. elegans
2.3.2. NormFinder Analysis of Reference Genes in G. elegans
2.3.3. BestKeeper Analysis of Reference Genes in G. elegans
2.3.4. ΔCT Method of Reference Genes in G. elegans
2.3.5. RefFinder Analysis of Reference Genes in G. elegans
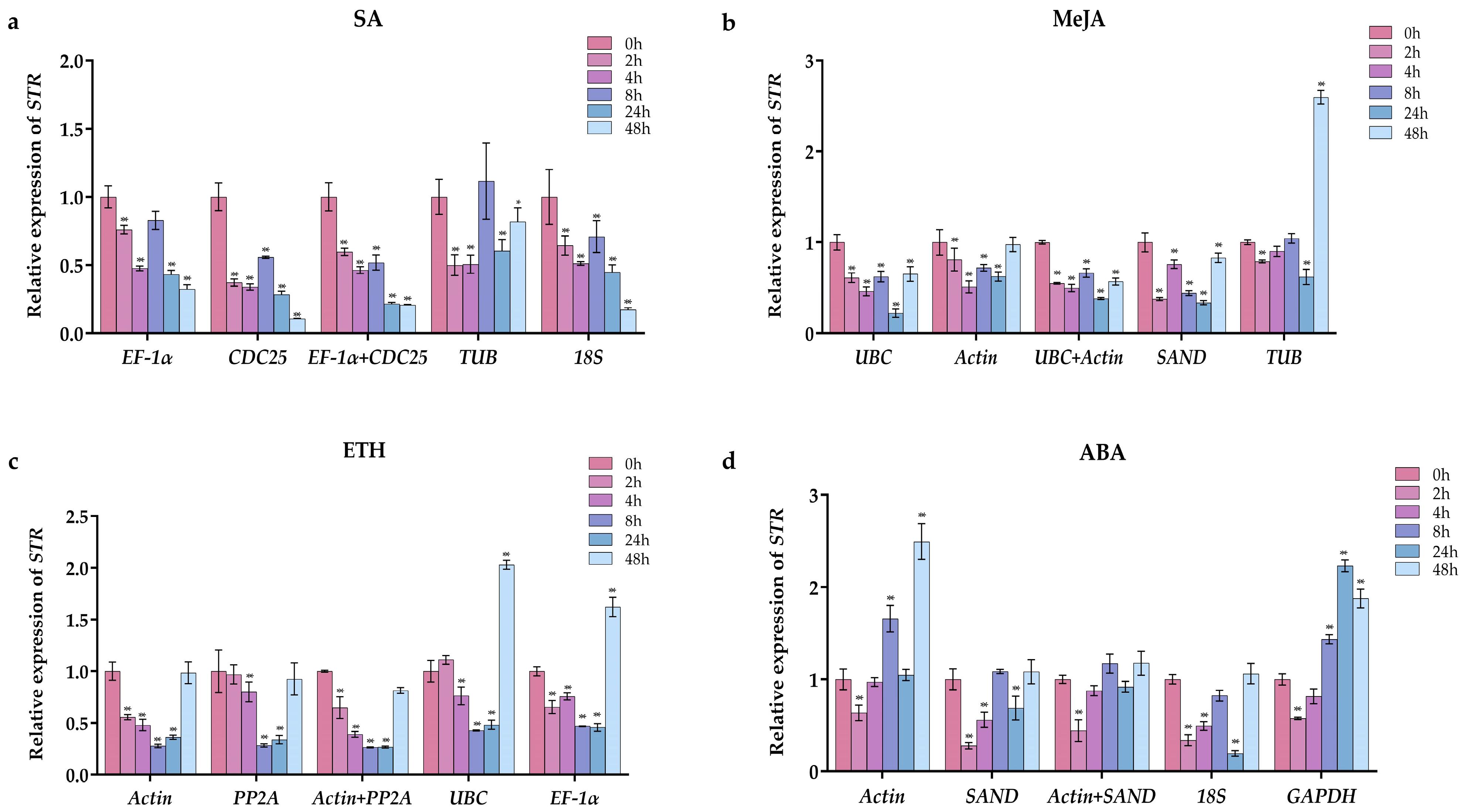
2.4. Validation of the Stability of Reference Genes
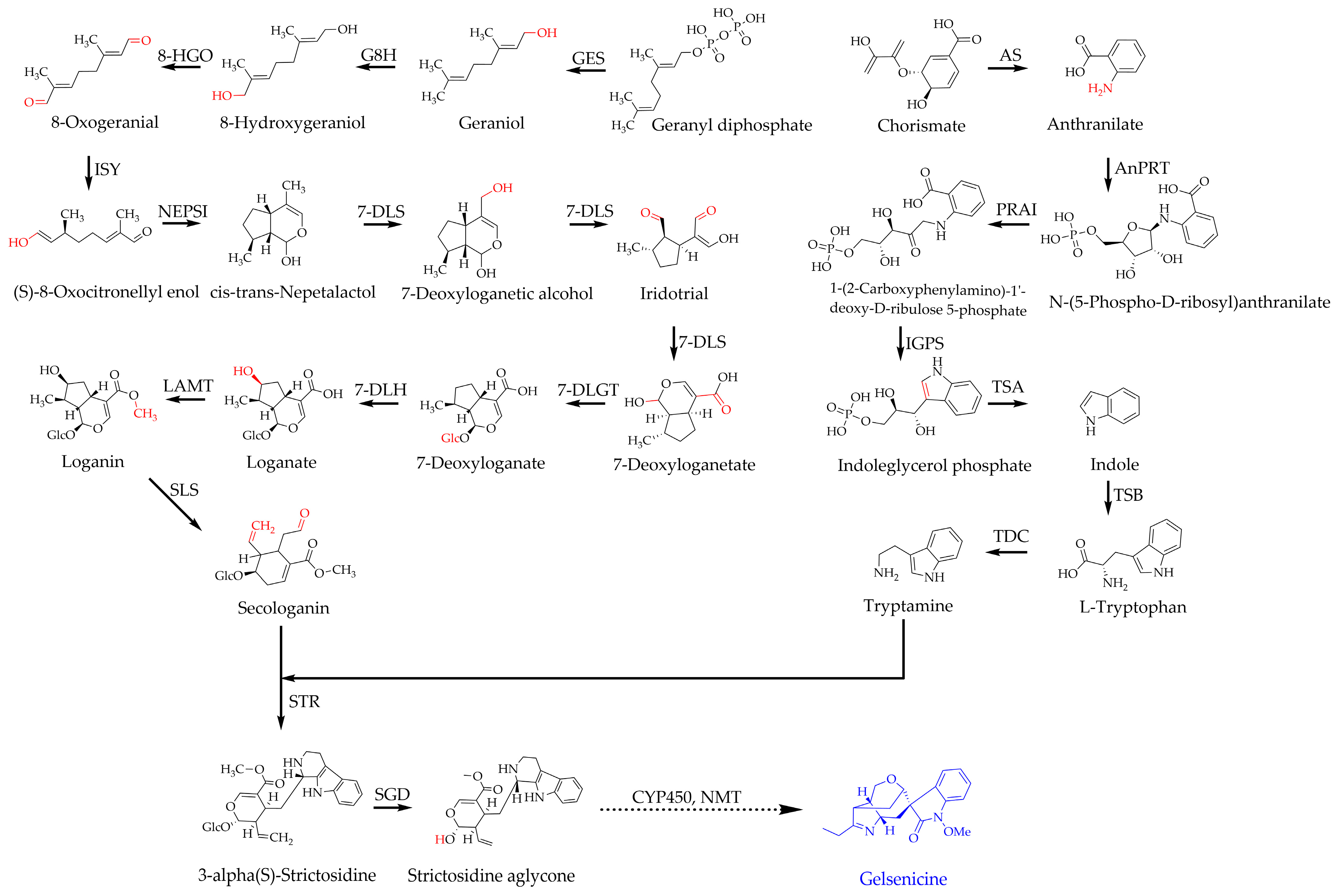
2.5. Expression Patterns of Pathway Genes Involved in the Biosynthesis of Gelsenicine
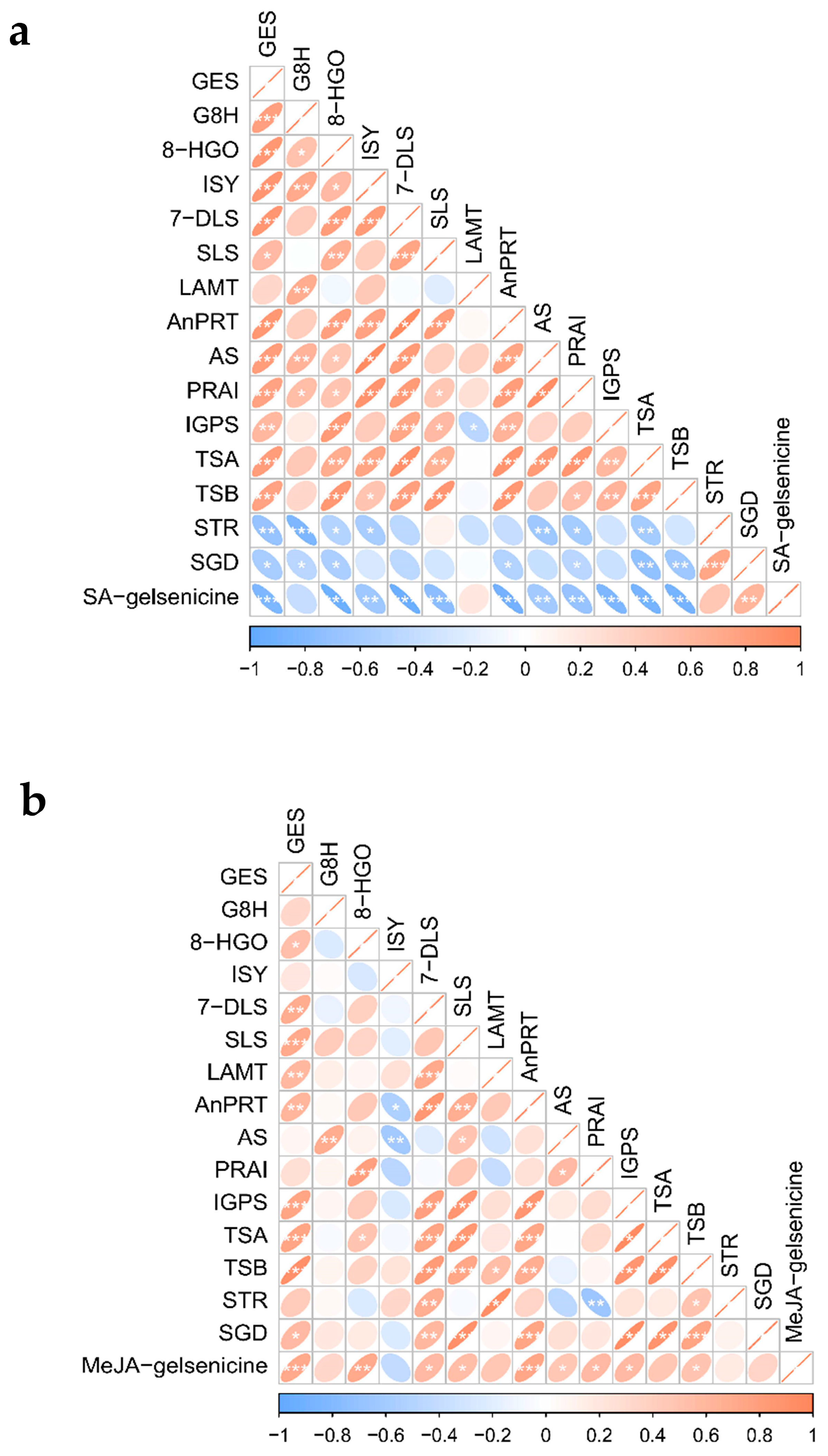
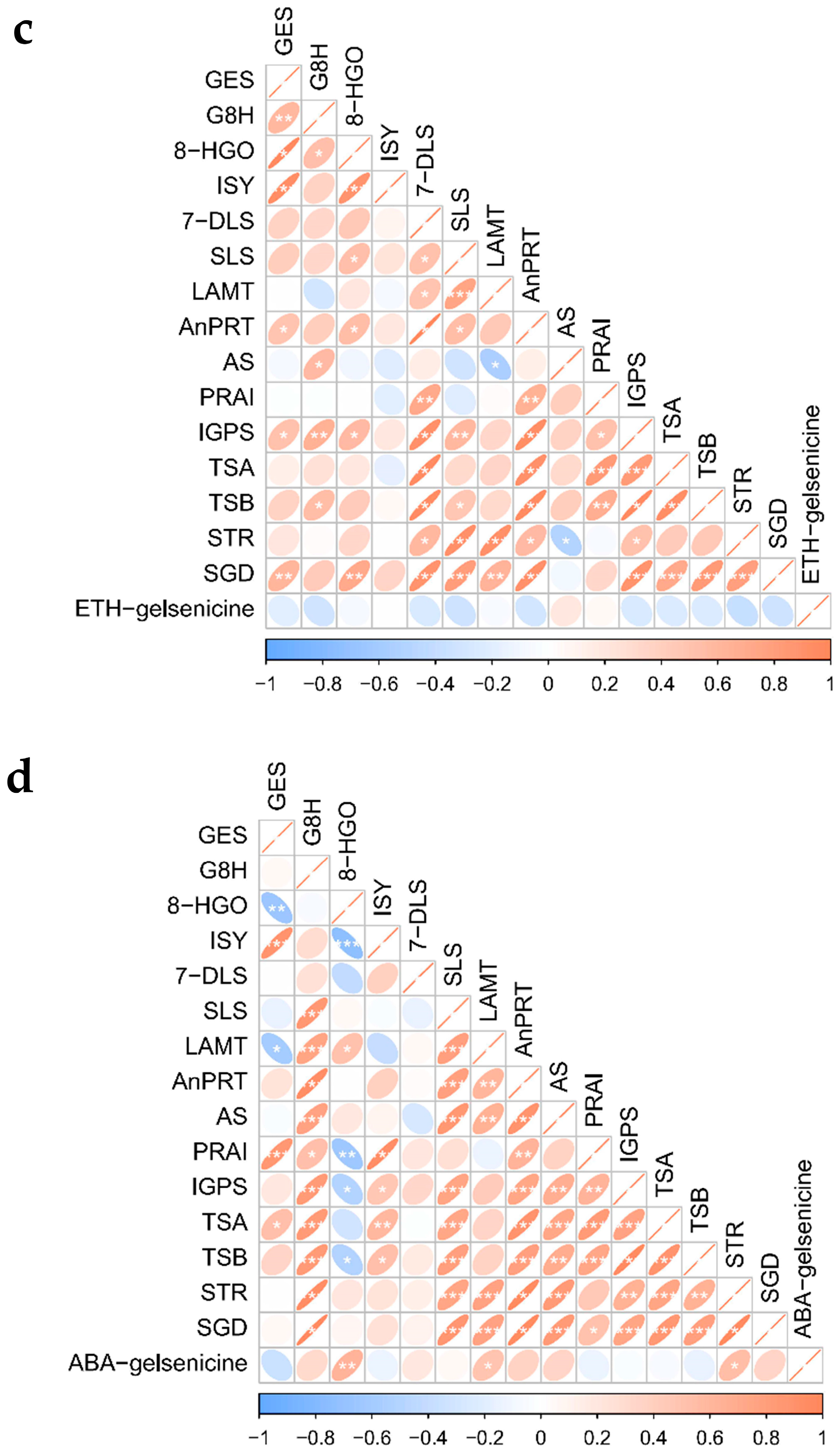
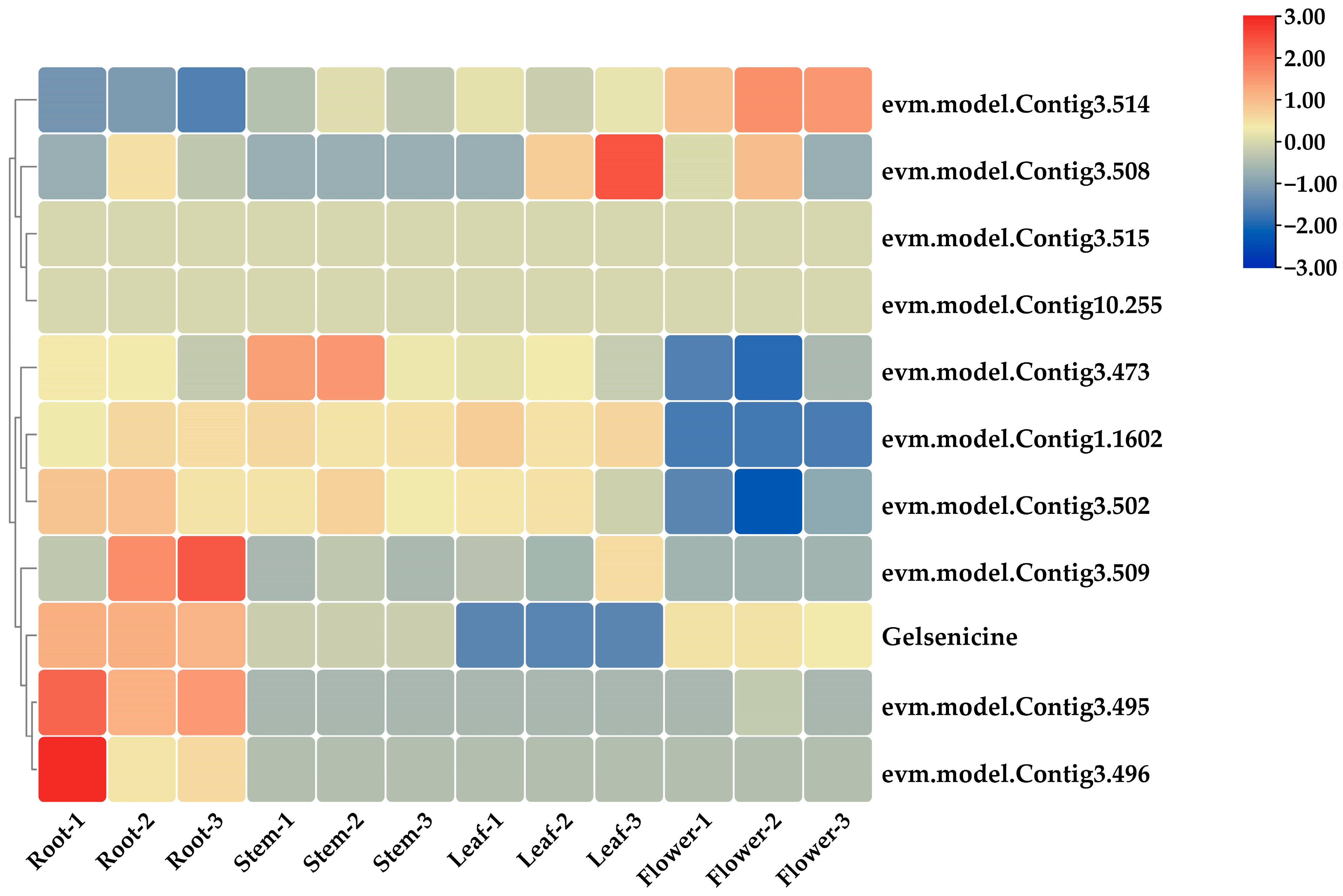
2.5.1. Co-Expression Screening for MIA Pathway Genes
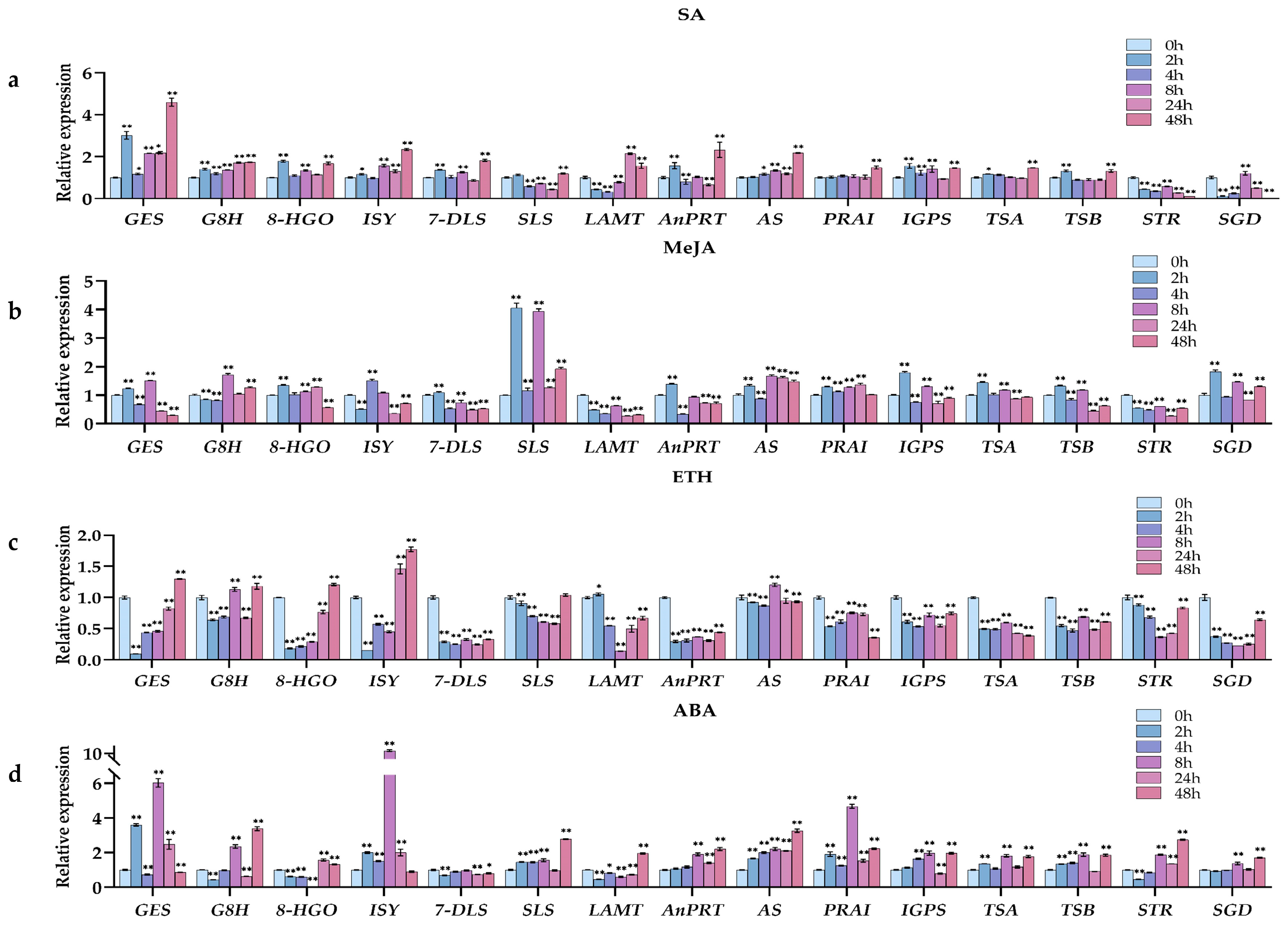
2.5.2. Expression Pattern Analysis of Gelsenicine-Related Genes under SA Treatments
2.5.3. Expression Pattern Analysis of Gelsenicine-Related Genes under MeJA Treatments
2.5.4. Expression Pattern Analysis of Gelsenicine-Related Genes under ETH Treatments
2.5.5. Expression Pattern Analysis of Gelsenicine-Related Genes under ABA Treatments
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Determination of the Content of Gelsenicine by HPLC
4.3. RNA Extraction and cDNA Synthesis
4.4. Candidate Gene Selection and Primer Design
4.5. RT-qPCR
4.6. Evaluation of Reference Genes
4.7. Validation of Reference Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhao, X.; Geng, Y.; Hu, T.; Zhao, Y.; Yang, S.; Hao, D. Evaluation of Optimal Reference Genes for qRT-PCR Analysis in Hyphantria cunea (Drury). Insects 2022, 13, 97. [Google Scholar] [CrossRef] [PubMed]
- Freitas, B.L.; Leach, L.; Chaturvedi, V.; Chaturvedi, S. Reverse Transcription-Quantitative Real-Time PCR (RT-qPCR) Assay for the Rapid Enumeration of Live Candida auris Cells from the Health Care Environment. J. Clin. Microbiol. 2022, 60, e0077921. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Yang, J.; Wang, X.; Xiong, Y.; Xiong, Y.; Dong, Z.; Lei, X.; Yan, L.; Ma, X. Selection and Validation of Reference Genes for qRT-PCR Gene Expression Analysis in Kengyilia melanthera. Genes 2022, 13, 1445. [Google Scholar] [CrossRef]
- Tang, J.; Liang, G.; Dong, S.; Shan, S.; Zhao, M.; Guo, X. Selection and Validation of Reference Genes for Quantitative Real-Time PCR Normalization in Athetis dissimilis (Lepidoptera: Noctuidae) Under Different Conditions. Front. Physiol. 2022, 13, 842195. [Google Scholar] [CrossRef]
- Deguchi, M.; Potlakayala, S.; Spuhler, Z.; George, H.; Sheri, V.; Agili, R.; Patel, A.; Rudrabhatla, S. Selection and validation of reference genes for normalization of qRT-PCR data to study the cannabinoid pathway genes in industrial hemp. PLoS ONE 2021, 16, e0260660. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.D.; Wang, B.; Li, Y.P.; Zeng, M.J.; Liu, J.T.; Ye, X.R.; Zhu, H.S.; Wen, Q.F. Reference gene selection for qRT-PCR analyses of luffa (Luffa cylindrica) plants under abiotic stress conditions. Sci. Rep. 2021, 11, 3161. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhu, L.; Xue, J.; Yang, J.; Hu, H.; Cui, J.; Xu, J. Selection and Verification of Appropriate Reference Genes for Expression Normalization in Cryptomeria fortunei under Abiotic Stress and Hormone Treatments. Genes 2021, 12, 791. [Google Scholar] [CrossRef]
- Wang, Q.; Guo, C.; Yang, S.; Zhong, Q.; Tian, J. Screening and Verification of Reference Genes for Analysis of Gene Expression in Garlic (Allium sativum L.) under Cold and Drought Stress. Plants 2023, 12, 763. [Google Scholar] [CrossRef]
- Adeola, F. Normalization of Gene Expression by Quantitative RT-PCR in Human Cell Line: Comparison of 12 Endogenous Reference Genes. Ethiop. J. Health Sci. 2018, 28, 741–748. [Google Scholar] [CrossRef]
- Sundaram, V.K.; Sampathkumar, N.K.; Massaad, C.; Grenier, J. Optimal use of statistical methods to validate reference gene stability in longitudinal studies. PLoS ONE 2019, 14, e0219440. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pair-wise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Wang, M.; Gan, Y.; Gong, H.; Li, J.; Zheng, X.; Liu, X.; Zhao, S.; Luo, J.; Wu, H. Identification of suitable reference genes for quantitative reverse transcription PCR in Luffa (Luffa cylindrica). Physiol. Mol. Biol. Plants 2022, 28, 737–747. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.; Wang, J.; Zhang, B. RefFinder: A web-based tool for comprehensively analyzing and identifying reference genes. Funct. Integr. Genom. 2023, 23, 125. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.; Xiao, P.; Chen, D.; Xu, L.; Zhang, B. miRDeepFinder: A miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol. Biol. 2012, 80, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, research0034.1. [Google Scholar] [CrossRef]
- Hu, W.; DiGennaro, P.M. Identification of Suitable Meloidogyne spp. Housekeeping Genes. J. Nematol. 2019, 51, e2019–e2055. [Google Scholar] [CrossRef]
- Wang, J.J.; Han, S.; Yin, W.; Xia, X.; Liu, C. Comparison of Reliable Reference Genes Following Different Hormone Treatments by Various Algorithms for qRT-PCR Analysis of Metasequoia. Int. J. Mol. Sci. 2018, 20, 34. [Google Scholar] [CrossRef]
- Ma, R.; Xu, S.; Zhao, Y.; Xia, B.; Wang, R. Selection and Validation of Appropriate Reference Genes for Quantitative Real-Time PCR Analysis of Gene Expression in Lycoris aurea. Front. Plant Sci. 2016, 7, 536. [Google Scholar] [CrossRef]
- Silver, N.; Best, S.; Jiang, J.; Thein, S.L. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol. Biol. 2006, 7, 33. [Google Scholar] [CrossRef]
- Yin, H.; Yin, D.; Zhang, M.; Gao, Z.; Tuluhong, M.; Li, X.; Li, J.; Li, B.; Cui, G. Validation of Appropriate Reference Genes for qRT-PCR Normalization in Oat (Avena sativa L.) under UV-B and High-Light Stresses. Int. J. Mol. Sci. 2022, 23, 11187. [Google Scholar] [CrossRef]
- Que, W.; Chen, M.; Yang, L.; Zhang, B.; Zhao, Z.; Liu, M.; Cheng, Y.; Qiu, H. A network pharmacology-based investigation on the bioactive ingredients and molecular mechanisms of Gelsemium elegans Benth against colorectal cancer. BMC Complement. Med. Ther. 2021, 21, 99. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.; Qiu, H.; Cheng, Y.; Liu, M.; Chen, M.; Que, Y.; Que, W. Gelsemium elegans Benth: Chemical Components, Pharmacological Effects, and Toxicity Mechanisms. Molecules 2021, 26, 7145. [Google Scholar] [CrossRef]
- Li, N.P.; Liu, J.S.; Liu, J.W.; Tian, H.Y.; Zhou, H.L.; Zheng, Y.R.; Huang, X.J.; Cao, J.Q.; Ye, W.C.; Wang, L. Monoterpenoid indole alkaloids from the fruits of Gelsemium elegans and their anti-inflammatory activities. Bioorganic Chem. 2021, 107, 104624. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Xu, H.L.; Liang, J.W.; Ding, Y.Y.; Meng, F.H. An Integrated Network, RNA Sequencing, and Experiment Pharmacology Approach Reveals the Active Component, Potential Target, and Mechanism of Gelsemium elegans in the Treatment of Colorectal Cancer. Front. Oncol. 2020, 10, 616628. [Google Scholar] [CrossRef] [PubMed]
- Xiong, B.; Zhong, Z.; Chen, C.; Huang, H.; Lin, J.; Xu, Y.; Yang, J.; Yu, C. The anxiolytic effect of koumine on a predatory sound stress-induced anxiety model and its associated molecular mechanisms. Phytomedicine 2022, 103, 154225. [Google Scholar] [CrossRef]
- Ye, Q.; Feng, Y.; Wang, Z.; Zhou, A.; Xie, S.; Zhang, Y.; Xiang, Q.; Song, E.; Zou, J. Effects of dietary Gelsemium elegans alkaloids on growth performance, immune responses and disease resistance of Megalobrama amblycephala. Fish. Shellfish. Immunol. 2019, 91, 29–39. [Google Scholar] [CrossRef]
- Xiong, B.; You, W.; Luo, Y.; Jin, G.; Wu, M.; Xu, Y.; Yang, J.; Huang, H.; Yu, C. Investigation of the Possible Allostery of Koumine Extracted From Gelsemium elegans Benth. And Analgesic Mechanism Associated With Neurosteroids. Front. Pharmacol. 2021, 12, 739618. [Google Scholar] [CrossRef]
- Wang, L.; Chen, S.; Gao, X.; Liang, X.; Lv, W.; Zhang, D.; Jin, X. Recent progress in chemistry and bioactivity of monoterpenoid indole alkaloids from the genus gelsemium: A comprehensive review. J. Enzym. Inhib. Med. Chem. 2023, 38, 2155639. [Google Scholar] [CrossRef]
- Li, Y.; Hu, P.; Zhang, Z.; Yuan, Z.; Yang, K.; Sun, Z. Protective autophagy alleviates neurotoxin-gelsenicine induced apoptosis through PERK signaling pathway in Neuro-2a cells. Toxicology 2022, 474, 153210. [Google Scholar] [CrossRef]
- Patra, B.; Pattanaik, S.; Schluttenhofer, C.; Yuan, L. A network of jasmonate-responsive bHLH factors modulate monoterpenoid indole alkaloid biosynthesis in Catharanthus roseus. New Phytol. 2018, 217, 1566–1581. [Google Scholar] [CrossRef]
- Zhang, J.; Hansen, L.G.; Gudich, O.; Viehrig, K.; Lassen, L.M.M.; Schrubbers, L.; Adhikari, K.B.; Rubaszka, P.; Carrasquer-Alvarez, E.; Chen, L.; et al. A microbial supply chain for production of the anti-cancer drug vinblastine. Nature 2022, 609, 341–347. [Google Scholar] [CrossRef]
- Liu, Y.; Patra, B.; Singh, S.K.; Paul, P.; Zhou, Y.; Li, Y.; Wang, Y.; Pattanaik, S.; Yuan, L. Terpenoid indole alkaloid biosynthesis in Catharanthus roseus: Effects and prospects of environmental factors in metabolic engineering. Biotechnol. Lett. 2021, 43, 2085–2103. [Google Scholar] [CrossRef] [PubMed]
- Shahsavarani, M.; Utomo, J.C.; Kumar, R.; Paz-Galeano, M.; Garza-Garcia, J.J.O.; Mai, Z.; Ro, D.K.; Qu, Y. Improved protein glycosylation enabled heterologous biosynthesis of monoterpenoid indole alkaloids and their unnatural derivatives in yeast. Metab. Eng. Commun. 2023, 16, e00215. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, D.; Lateef, A.; Natarajan, P.; Palanisamy, S. De novo transcriptome analysis of Justicia adhatoda reveals candidate genes involved in major biosynthetic pathway. Mol. Biol. Rep. 2022, 49, 10307–10314. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Zhang, D.; Yang, B.; Su, N.; Cui, J. Salicylic Acid Regulates Indole-3-Carbinol Biosynthesis Under Blue Light in Broccoli Sprouts (Brassica oleracea L.). Front. Plant Sci. 2022, 13, 848454. [Google Scholar] [CrossRef]
- Akhgari, A.; Laakso, I.; Maaheimo, H.; Choi, Y.H.; Seppanen-Laakso, T.; Oksman-Caldentey, K.M.; Rischer, H. Methyljasmonate Elicitation Increases Terpenoid Indole Alkaloid Accumulation in Rhazya stricta Hairy Root Cultures. Plants 2019, 8, 534. [Google Scholar] [CrossRef]
- Markowski, M.; Alsoufi, A.S.M.; Szakiel, A.; Dlugosz, M. Effect of Ethylene and Abscisic Acid on Steroid and Triterpenoid Synthesis in Calendula officinalis Hairy Roots and Saponin Release to the Culture Medium. Plants 2022, 11, 303. [Google Scholar] [CrossRef]
- Ma, Y.N.; Xu, D.B.; Yan, X.; Wu, Z.K.; Kayani, S.I.; Shen, Q.; Fu, X.Q.; Xie, L.H.; Hao, X.L.; Hassani, D.; et al. Jasmonate- and abscisic acid-activated AaGSW1-AaTCP15/AaORA transcriptional cascade promotes artemisinin biosynthesis in Artemisia annua. Plant Biotechnol. J. 2021, 19, 1412–1428. [Google Scholar] [CrossRef]
- Bahieldin, A.; Atef, A.; Edris, S.; Gadalla, N.O.; Al-Matary, M.; Al-Kordy, M.A.; Ramadan, A.M.; Bafeel, S.; Alharbi, M.G.; Al-Quwaie, D.A.H.; et al. Stepwise response of MeJA-induced genes and pathways in leaves of C. roseus. Comptes Rendus Biol. 2018, 341, 411–420. [Google Scholar] [CrossRef]
- Jiao, C.; Wei, M.; Fan, H.; Song, C.; Wang, Z.; Cai, Y.; Jin, Q. Transcriptomic analysis of genes related to alkaloid biosynthesis and the regulation mechanism under precursor and methyl jasmonate treatment in Dendrobium officinale. Front. Plant Sci. 2022, 13, 941231. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, Y.; Bai, H.; Han, Y.; Yu, F. An ABCG-Type Transporter Facilitates ABA Influx and Regulates Camptothecin Biosynthesis in Camptotheca acuminata. Int. J. Mol. Sci. 2022, 23, 16120. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Tang, Q.; Cheng, P.; Zhu, M.; Zhang, H.; Liu, J.; Zuo, M.; Huang, C.; Wu, C.; Sun, Z.; et al. Whole-genome sequencing and analysis of the Chinese herbal plant Gelsemium elegans. Acta Pharm. Sin. B 2020, 10, 374–382. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, D.; Yang, L.; Zeng, X.; Zhang, Q.; Liu, G.; Pan, Y. Cuproptosis related genes associated with Jab1 shapes tumor microenvironment and pharmacological profile in nasopharyngeal carcinoma. Front. Immunol. 2022, 13, 989286. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, L.F.; Piovezani, A.R.; Ivanov, D.A.; Yoshida, L.; Segal Floh, E.I.; Kato, M.J. Selection and validation of reference genes for measuring gene expression in Piper species at different life stages using RT-qPCR analysis. Plant Physiol. Biochem. 2022, 171, 201–212. [Google Scholar] [CrossRef]
- Udvardi, M.K.; Czechowski, T.; Scheible, W.R. Eleven golden rules of quantitative RT-PCR. Plant Cell 2008, 20, 1736–1737. [Google Scholar] [CrossRef]
- Ferreira, M.J.; Silva, J.; Pinto, S.C.; Coimbra, S. I Choose You: Selecting Accurate Reference Genes for qPCR Expression Analysis in Reproductive Tissues in Arabidopsis thaliana. Biomolecules 2023, 13, 463. [Google Scholar] [CrossRef]
- Zhu, X.; Wang, B.; Wang, X.; Wei, X. Screening of stable internal reference gene of Quinoa under hormone treatment and abiotic stress. Physiol. Mol. Biol. Plants 2021, 27, 2459–2470. [Google Scholar] [CrossRef]
- Li, M.Y.; Song, X.; Wang, F.; Xiong, A.S. Suitable Reference Genes for Accurate Gene Expression Analysis in Parsley (Petroselinum crispum) for Abiotic Stresses and Hormone Stimuli. Front. Plant Sci. 2016, 7, 1481. [Google Scholar] [CrossRef]
- Feng, K.; Liu, J.X.; Xing, G.M.; Sun, S.; Li, S.; Duan, A.Q.; Wang, F.; Li, M.Y.; Xu, Z.S.; Xiong, A.S. Selection of appropriate reference genes for RT-qPCR analysis under abiotic stress and hormone treatment in celery. PeerJ 2019, 7, e7925. [Google Scholar] [CrossRef]
- Zhang, K.; Fan, W.; Chen, D.; Jiang, L.; Li, Y.; Yao, Z.; Yang, Y.; Qiu, D. Selection and validation of reference genes for quantitative gene expression normalization in Taxus spp. Sci. Rep. 2020, 10, 22205. [Google Scholar] [CrossRef]
- Li, L.; Li, N.; Fang, H.; Qi, X.; Zhou, Y. Selection and Validation of Reference Genes for Normalisation of Gene Expression in Glehnia littoralis. Sci. Rep. 2020, 10, 7374. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Zhou, Y.J.; Hillwig, M.L.; Shen, Y.; Yang, L.; Wang, Y.; Zhang, X.; Liu, W.; Peters, R.J.; Chen, X.; et al. CYP76AH1 catalyzes turnover of miltiradiene in tanshinones biosynthesis and enables heterologous production of ferruginol in yeasts. Proc. Natl. Acad. Sci. USA 2013, 110, 12108–12113. [Google Scholar] [CrossRef] [PubMed]
- Tang, Q.; Ma, X.; Mo, C.; Wilson, I.W.; Song, C.; Zhao, H.; Yang, Y.; Fu, W.; Qiu, D. An efficient approach to finding Siraitia grosvenorii triterpene biosynthetic genes by RNA-seq and digital gene expression analysis. BMC Genom. 2011, 12, 343. [Google Scholar] [CrossRef] [PubMed]
- Dai, L.; Liu, C.; Zhu, Y.; Zhang, J.; Men, Y.; Zeng, Y.; Sun, Y. Functional Characterization of Cucurbitadienol Synthase and Triterpene Glycosyltransferase Involved in Biosynthesis of Mogrosides from Siraitia grosvenorii. Plant Cell Physiol. 2015, 56, 1172–1182. [Google Scholar] [CrossRef]
- Zhang, J.; Dai, L.; Yang, J.; Liu, C.; Men, Y.; Zeng, Y.; Cai, Y.; Zhu, Y.; Sun, Y. Oxidation of Cucurbitadienol Catalyzed by CYP87D18 in the Biosynthesis of Mogrosides from Siraitia grosvenorii. Plant Cell Physiol. 2016, 57, 1000–1007. [Google Scholar] [CrossRef]
- Soltani, N.; Firouzabadi, F.N.; Shafeinia, A.; Shirali, M.; Sadr, A.S. De Novo transcriptome assembly and differential expression analysis of catharanthus roseus in response to salicylic acid. Sci. Rep. 2022, 12, 17803. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Yang, J.; Han, F.; Yang, L.; Wang, J.; Jin, F.; Luo, A.; Zhao, F. Identification of Reference Genes for RT-qPCR Analysis in Gleditsia microphylla under Abiotic Stress and Hormone Treatment. Genes 2022, 13, 1227. [Google Scholar] [CrossRef]
- Hellemans, J.; Mortier, G.; De Paepe, A.; Speleman, F.; Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 2007, 8, R19. [Google Scholar] [CrossRef]
- Yi, S.; Lu, H.; Tian, C.; Xu, T.; Song, C.; Wang, W.; Wei, P.; Gu, F.; Liu, D.; Cai, Y.; et al. Selection of Suitable Reference Genes for Gene Expression Normalization Studies in Dendrobium huoshanense. Genes 2022, 13, 1486. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, C.; Zhang, J.; Chen, F.; Gong, Y.; Li, Y.; Su, Y.; Wei, Y.; Zhao, Y. Selection of the Reference Gene for Expression Normalization in Papaver somniferum L. under Abiotic Stress and Hormone Treatment. Genes 2020, 11, 124. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Qi, X.; Yan, H.; Huang, L.; Nie, G.; Zhang, X. Reference Gene Selection for Quantitative Real-Time Reverse-Transcriptase PCR in Annual Ryegrass (Lolium multiflorum) Subjected to Various Abiotic Stresses. Molecules 2018, 23, 172. [Google Scholar] [CrossRef] [PubMed]


| Gene | Gene Description | Primer Sequence | Slope/k | E/% | R2 | Accession Number |
|---|---|---|---|---|---|---|
| 18S | 18S Ribosomal RNA | F: GATGGAGTCCCGAAGTTGC R: TCCAGATCGCATGGCATAG | −3.37 | 97.9 | 0.994 | OR413515 |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | F: AAGGGTGGTGCCAAGAAGG R: CAGTGGGAACACGGAAAGC | −3.11 | 109.8 | 0.998 | OR413520 |
| Actin | Actin | F: GTTGCCCAGAAGTCCTATT R: TTCCTGTGGACGATTGATG | −3.12 | 109.3 | 0.997 | OR413517 |
| TUA | α-Tubulin | F: ATGAAGTTAGAACAGGGACA R: CAAGCAGGGAGTGAGTAGA | −3.15 | 107.6 | 0.992 | OR413519 |
| TUB | β-Tubulin | F: TGTCCGTAAAGAAGCCGAGAA R: CAGGGAAACGAAGGCAACA | −3.26 | 102.8 | 0.996 | OR413518 |
| SAND | SAND family protein | F: CATCCGACCCACCTACCGT R: ACTCTGCCAACTCCGCTCC | −3.13 | 108.5 | 0.996 | OR413523 |
| EF1-α | Elongation factor 1α | F: AAGCCACTCCGTCTCCCACT R: TCGGCAAACTTGACAGCAATA | −3.1 | 110.1 | 0.991 | OR413516 |
| UBC | Ubiquitin C | F: CAAAGGTGGTGAGGAGGAT R: ACAGAGCAGCGACTGAATG | −3.33 | 99.6 | 0.994 | OR413521 |
| PP2A | Protein phosphatases 2A | F: TGATTACCTGCCTCTGAC R: TGTGGAACCTCCTGTATG | −3.19 | 105.9 | 0.999 | OR413514 |
| CDC25 | Cell division cyclin 25 homolog C | F: CAGGGATGACGAAAGGAGT R: CGCAATGGAAAACAAGAGT | −3.11 | 109.8 | 0.999 | OR413522 |
| Group | Rank | GeNorm | NormFinder | BestKeeper | ΔCT | RefFinder | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Gene | Stab. | Gene | Stab. | Gene | CV ± SD | Gene | Stab. | Gene | Stab. | ||
| Control | 1 | CDC25 | 0.139 | TUA | 0.016 | Actin | 0.41 ± 0.08 | TUA | 0.30 | TUA | 2.55 |
| 2 | EF1-α | 0.139 | GAPDH | 0.126 | UBC | 0.42 ± 0.09 | GAPDH | 0.32 | GAPDH | 2.78 | |
| 3 | Actin | 0.207 | SAND | 0.147 | GAPDH | 0.73 ± 0.13 | UBC | 0.32 | Actin | 2.94 | |
| 4 | UBC | 0.220 | UBC | 0.176 | CDC25 | 0.68 ± 0.15 | SAND | 0.33 | UBC | 3.13 | |
| 5 | GAPDH | 0.230 | Actin | 0.232 | PP2A | 0.69 ± 0.16 | Actin | 0.35 | CDC25 | 3.72 | |
| 6 | TUA | 0.246 | PP2A | 0.263 | EF1A | 0.90 ± 0.16 | CDC25 | 0.37 | EF1-α | 4.56 | |
| 7 | SAND | 0.263 | TUB | 0.266 | TUA | 0.92 ± 0.19 | PP2A | 0.38 | SAND | 5.09 | |
| 8 | PP2A | 0.272 | CDC25 | 0.291 | SAND | 0.80 ± 0.20 | EF1-α | 0.38 | PP2A | 6.40 | |
| 9 | TUB | 0.296 | EF1-α | 0.293 | TUB | 1.42 ± 0.33 | TUB | 0.40 | TUB | 8.45 | |
| 10 | 18S | 0.391 | 18S | 0.749 | 18S | 2.82 ± 0.70 | 18S | 0.77 | 18S | 10.00 | |
| SA | 1 | CDC25 | 0.181 | EF1-α | 0.164 | GAPDH | 0.90 ± 0.17 | EF1-α | 0.35 | EF1-α | 1.32 |
| 2 | EF1-α | 0.181 | GAPDH | 0.191 | CDC25 | 0.77 ± 0.17 | GAPDH | 0.37 | CDC25 | 2.06 | |
| 3 | PP2A | 0.225 | CDC25 | 0.236 | EF1-α | 1.04 ± 0.19 | CDC25 | 0.38 | GAPDH | 2.11 | |
| 4 | UBC | 0.270 | UBC | 0.254 | Actin | 1.20 ± 0.24 | UBC | 0.40 | PP2A | 4.40 | |
| 5 | GAPDH | 0.298 | PP2A | 0.286 | PP2A | 1.16 ± 0.26 | PP2A | 0.42 | UBC | 4.43 | |
| 6 | Actin | 0.322 | Actin | 0.294 | UBC | 1.65 ± 0.34 | Actin | 0.42 | Actin | 5.42 | |
| 7 | TUA | 0.349 | TUA | 0.326 | TUA | 1.85 ± 0.38 | TUA | 0.44 | TUA | 7.00 | |
| 8 | SAND | 0.363 | SAND | 0.338 | SAND | 1.56 ± 0.40 | SAND | 0.45 | SAND | 8.00 | |
| 9 | TUB | 0.394 | TUB | 0.411 | TUB | 1.72 ± 0.41 | TUB | 0.51 | TUB | 9.00 | |
| 10 | 18S | 0.435 | 18S | 0.524 | 18S | 1.93 ± 0.49 | 18S | 0.60 | 18S | 10.00 | |
| MeJA | 1 | Actin | 0.267 | UBC | 0.174 | UBC | 1.08 ± 0.22 | UBC | 0.56 | UBC | 1.41 |
| 2 | EF1-α | 0.267 | TUA | 0.267 | GAPDH | 1.61 ± 0.30 | TUA | 0.61 | Actin | 2.63 | |
| 3 | PP2A | 0.303 | GAPDH | 0.292 | TUA | 1.58 ± 0.32 | Actin | 0.61 | TUA | 2.78 | |
| 4 | UBC | 0.362 | Actin | 0.360 | Actin | 1.73 ± 0.33 | GAPDH | 0.64 | EF1-α | 3.50 | |
| 5 | TUA | 0.419 | 18S | 0.418 | EF1-α | 2.06 ± 0.37 | EF1-α | 0.64 | GAPDH | 3.72 | |
| 6 | 18S | 0.458 | EF1-α | 0.420 | PP2A | 1.65 ± 0.37 | PP2A | 0.66 | PP2A | 5.24 | |
| 7 | CDC25 | 0.491 | PP2A | 0.459 | 18S | 1.55 ± 0.37 | 18S | 0.68 | 18S | 6.19 | |
| 8 | GAPDH | 0.521 | CDC25 | 0.657 | CDC25 | 2.37 ± 0.51 | CDC25 | 0.79 | CDC25 | 7.74 | |
| 9 | SAND | 0.606 | SAND | 0.738 | SAND | 2.08 ± 0.55 | SAND | 0.90 | SAND | 9.00 | |
| 10 | TUB | 0.733 | TUB | 1.175 | TUB | 3.54 ± 0.90 | TUB | 1.24 | TUB | 10.00 | |
| ETH | 1 | CDC25 | 0.231 | Actin | 0.183 | Actin | 0.68 ± 0.13 | Actin | 0.42 | Actin | 1.32 |
| 2 | PP2A | 0.231 | PP2A | 0.208 | PP2A | 1.01 ± 0.23 | PP2A | 0.43 | PP2A | 1.68 | |
| 3 | Actin | 0.267 | CDC25 | 0.269 | CDC25 | 1.19 ± 0.26 | CDC25 | 0.45 | CDC25 | 2.28 | |
| 4 | TUB | 0.313 | TUB | 0.322 | 18S | 1.34 ± 0.33 | TUB | 0.48 | TUB | 4.43 | |
| 5 | TUA | 0.373 | TUA | 0.330 | GAPDH | 1.80 ± 0.33 | TUA | 0.49 | TUA | 5.62 | |
| 6 | GAPDH | 0.390 | GAPDH | 0.340 | TUB | 1.52 ± 0.35 | GAPDH | 0.49 | GAPDH | 5.73 | |
| 7 | SAND | 0.412 | SAND | 0.347 | SAND | 1.46 ± 0.36 | SAND | 0.51 | 18S | 6.73 | |
| 8 | 18S | 0.430 | 18S | 0.408 | TUA | 1.89 ± 0.38 | 18S | 0.54 | SAND | 7.00 | |
| 9 | UBC | 0.465 | UBC | 0.521 | EF1-α | 2.13 ± 0.39 | UBC | 0.62 | UBC | 9.24 | |
| 10 | EF1-α | 0.514 | EF1-α | 0.633 | UBC | 2.77 ± 0.56 | EF1-α | 0.71 | EF1-α | 9.74 | |
| ABA | 1 | Actin | 0.230 | SAND | 0.313 | PP2A | 1.72 ± 0.40 | Actin | 0.73 | Actin | 2.14 |
| 2 | EF1-α | 0.230 | TUB | 0.337 | CDC25 | 1.87 ± 0.42 | SAND | 0.76 | SAND | 2.51 | |
| 3 | TUA | 0.294 | Actin | 0.353 | UBC | 2.49 ± 0.52 | TUB | 0.77 | TUB | 3.31 | |
| 4 | SAND | 0.487 | UBC | 0.480 | TUB | 2.47 ± 0.61 | UBC | 0.79 | EF1-α | 3.76 | |
| 5 | TUB | 0.566 | EF1-α | 0.540 | SAND | 2.84 ± 0.75 | EF1-α | 0.81 | UBC | 4.12 | |
| 6 | UBC | 0.639 | 18S | 0.631 | 18S | 3.34 ± 0.81 | 18S | 0.88 | PP2A | 5.20 | |
| 7 | 18S | 0.677 | TUA | 0.701 | Actin | 4.50 ± 0.92 | TUA | 0.91 | CDC25 | 5.66 | |
| 8 | CDC25 | 0.729 | CDC25 | 0.764 | EF1-α | 5.49 ± 1.06 | CDC25 | 0.96 | TUA | 6.03 | |
| 9 | PP2A | 0.790 | PP2A | 0.891 | TUA | 5.64 ± 1.22 | PP2A | 1.07 | 18S | 6.24 | |
| 10 | GAPDH | 0.904 | GAPDH | 1.251 | GAPDH | 6.03 ± 1.26 | GAPDH | 1.36 | GAPDH | 10.00 | |
| Total | 1 | 18S | 0.567 | Actin | 0.420 | PP2A | 1.77 ± 0.40 | Actin | 0.81 | UBC | 1.68 |
| 2 | UBC | 0.567 | UBC | 0.453 | UBC | 2.16 ± 0.45 | UBC | 0.85 | Actin | 2.24 | |
| 3 | CDC25 | 0.616 | TUA | 0.554 | CDC25 | 2.07 ± 0.45 | TUA | 0.90 | 18S | 2.83 | |
| 4 | PP2A | 0.624 | 18S | 0.601 | 18S | 2.02 ± 0.49 | 18S | 0.91 | PP2A | 3.16 | |
| 5 | Actin | 0.652 | PP2A | 0.606 | Actin | 2.93 ± 0.57 | PP2A | 0.91 | TUA | 4.58 | |
| 6 | EF1-α | 0.693 | EF1-α | 0.612 | EF1-α | 3.70 ± 0.68 | EF1-α | 0.91 | CDC25 | 4.58 | |
| 7 | TUA | 0.732 | CDC25 | 0.715 | TUA | 3.82 ± 0.78 | CDC25 | 0.96 | EF1-α | 6.00 | |
| 8 | SAND | 0.823 | SAND | 0.792 | SAND | 3.43 ± 0.88 | SAND | 1.06 | SAND | 8.00 | |
| 9 | GAPDH | 0.897 | GAPDH | 0.989 | TUB | 4.33 ± 1.05 | GAPDH | 1.19 | GAPDH | 9.24 | |
| 10 | TUB | 0.982 | TUB | 1.167 | GAPDH | 5.46 ± 1.05 | TUB | 1.32 | TUB | 9.74 | |
| Experimental Treatments | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Control | SA | MeJA | ETH | ABA | Total | ||||||
| Most | Least | Most | Least | Most | Least | Most | Least | Most | Least | Most | Least |
| TUA GAPDH | 18S | EF1-α CDC25 | 18S | UBC Actin | TUB | Actin PP2A | EF1-α | Actin SAND | GAPDH | UBC Actin 18S | TUB |
| Gene | Gene Description | Primer Sequence | Slope/k | E/% | R2 | Accession Number |
|---|---|---|---|---|---|---|
| GES | Geraniol synthase | F: GGCTGCGTTTCAGGTTGCT R: CTTTAGGTGGGCTTGGGTG | −3.20 | 105.2 | 0.983 | OR413524 |
| G8H | Geraniol 8-hydroxylase | F: GTTTGGCGGAACAGACACC R: CGCTGAAATCCCACTTGCT | −3.28 | 102.0 | 0.995 | OR413525 |
| 8-HGO | 8-hydroxygeraniol dehydrogenase | F: CTGTCTTCCCGCTGCTTGC R: CGTTCCATTGCCGTGTTGA | −3.24 | 103.5 | 0.993 | OR413526 |
| ISY | (S)-8-oxocitronellyl enol synthase | F: CCGACCTGCTCTGGTTTTC R: AGGCTCACCTGTTCTTTGC | −3.01 | 114.8 | 0.992 | OR413527 |
| 7-DLS | 7-deoxyloganetic acid synthase | F: TGGCTGAGGTGTTGTTTG R: TGAATACCAGGCGAGTTT | −3.24 | 103.3 | 0.995 | OR413528 |
| LAMT | Loganate methyltransferase | F: CTGCTCCACAGGTCCCAATA R: GTGCCATCAACCCTCCGT | −3.43 | 95.7 | 0.996 | OR413529 |
| SLS | Secologanin synthase | F: TAGGCTGCTATTTGGGGATT R: ATGAGCACGGCAGGTTTT | −3.38 | 97.7 | 0.995 | OR413530 |
| AS | Anthranilate synthase | F: GGCGAATCCCGTTGTTGT R: TTGAGGCGTTCCAGGTCC | −3.36 | 98.5 | 0.996 | OR413531 |
| AnPRT | Anthranilate phosphoribosyltransferase | F: ACGGCAATCCTCCTTCCAA R: TTCGCCTGAGCATCCAACA | −3.21 | 105.0 | 0.999 | OR413532 |
| PRAI | Phosphoribosylanthranilate isomerase | F: GGGGCTTGGCTATTCTTGT R: GTTTTCCTCAGAGCAGCGT | −3.39 | 97.3 | 0.993 | OR413533 |
| IGPS | Indole-3-glycerol phosphate synthase | F: TGGTCCCTTTGAGTTTCGG R: AGGCAACCCAGTTCGTGAG | −3.21 | 105.0 | 0.998 | OR413534 |
| TSA | Tryptophan synthase alpha chain | F: CAAGCGTGGTGTTGAAAAG R: GCTCTGGGAGTTGTGGGTG | −3.25 | 103.2 | 0.998 | OR413535 |
| TSB | Tryptophan synthase beta chain | F: CAGTTCATTCTGGGACCGC R: TTCCAATGCCTGCTTCCTT | −3.21 | 104.8 | 0.998 | OR413536 |
| STR | Strictosidine synthase | F: AAGGAAGAGGGCGTGGAA R: GCAACAGGCAATGCAGAA | −3.33 | 99.8 | 0.987 | OR413537 |
| SGD | Strictosidine-β-D- glucosidase | F: TATGGTTATGCGTCGGGTGT R: AAGGCTCTGTGCCAGGGTT | −3.39 | 97.1 | 0.996 | OR413538 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Mu, D.; Wang, L.; Wang, X.; Wilson, I.W.; Chen, W.; Wang, J.; Liu, Z.; Qiu, D.; Tang, Q. Reference Genes Screening and Gene Expression Patterns Analysis Involved in Gelsenicine Biosynthesis under Different Hormone Treatments in Gelsemium elegans. Int. J. Mol. Sci. 2023, 24, 15973. https://doi.org/10.3390/ijms242115973
Zhang Y, Mu D, Wang L, Wang X, Wilson IW, Chen W, Wang J, Liu Z, Qiu D, Tang Q. Reference Genes Screening and Gene Expression Patterns Analysis Involved in Gelsenicine Biosynthesis under Different Hormone Treatments in Gelsemium elegans. International Journal of Molecular Sciences. 2023; 24(21):15973. https://doi.org/10.3390/ijms242115973
Chicago/Turabian StyleZhang, Yao, Detian Mu, Liya Wang, Xujun Wang, Iain W. Wilson, Wenqiang Chen, Jinghan Wang, Zhaoying Liu, Deyou Qiu, and Qi Tang. 2023. "Reference Genes Screening and Gene Expression Patterns Analysis Involved in Gelsenicine Biosynthesis under Different Hormone Treatments in Gelsemium elegans" International Journal of Molecular Sciences 24, no. 21: 15973. https://doi.org/10.3390/ijms242115973






